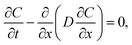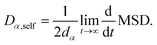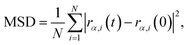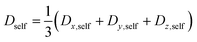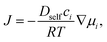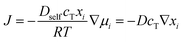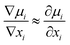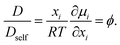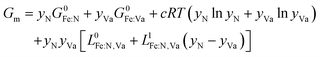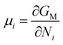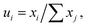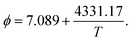Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Molecular dynamics simulation of nitrogen diffusion in iron and iron nitrides using ab initio data trained machine learning potentials
Peijie
Feng
 a,
Aditya Dilip
Lele
a,
Aditya Dilip
Lele
 bc,
Minhyeok
Lee
bc,
Minhyeok
Lee
 a,
Yiguang
Ju
a,
Yiguang
Ju
 cd and
Yuji
Suzuki
cd and
Yuji
Suzuki
 *a
*a
aDepartment of Mechanical Engineering, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-8656, Japan. E-mail: ysuzuki@mesl.t.u-tokyo.ac.jp
bDepartment of Mechanical Engineering, Rowan University, Glassboro, New Jersey 08028, USA
cMechanical & Aerospace Engineering, Princeton University, Princeton, New Jersey 08544, USA
dInstitute of Fluid Science, Tohoku University, Sendai, Miyagi 980-8576, Japan
First published on 23rd March 2026
Abstract
Ammonia is receiving increasing attention as a hydrogen energy carrier and a green fuel for achieving carbon neutrality. Combustors utilizing ammonia as a fuel are subject to “unwanted” surface nitriding, which hardens and embrittles the metal walls of combustors. Predicting the nitridation rate and nitride layer thicknesses with Fick's diffusion law requires temperature- and concentration-dependent nitrogen atoms’ diffusion coefficients in the relevant iron and iron nitrides. Experimentally determining the diffusion coefficients is challenging due to steep N-gradient, and concurrent phase transformations. In this study, we calculated the nitrogen atom diffusion coefficients using molecular dynamics (MD) driven by a machine learning interatomic potential (MLP). The MLP was trained on snapshots extracted from ab initio molecular dynamics (AIMD) trajectories covering different temperatures and nitrogen atom concentrations, and the contribution of each subset to the model's accuracy was quantified. The results show that the MLP reproduces independently the calculated diffusion energy barriers obtained using density functional theory (DFT) in α-, γ-iron and γ′-, ε-iron nitrides. For ε-iron nitride, the MLP-driven MD-computed self-diffusion coefficient was converted into chemical (Fickian) diffusion coefficients, enabling direct comparison with experiments. Our MLP-driven MD simulation delivers diffusion coefficients approaching ab initio accuracy, enabling precise concentration- and temperature-resolved modeling of nitrogen diffusion in iron and its nitrides. Arrhenius fitting to the 830–1500 K chemical diffusion coefficients reproduce the experimental activation energy and pre-exponential factor. Extrapolation to 823 K, the temperature at which most of the experiments are performed, yields a coefficient that falls within the experimental uncertainty. The present MLP–MD model will facilitate the mechanistic understanding of iron surface nitridation and quantitative predictions of “unwanted” nitriding of iron-based combustor wall materials induced by ammonia at high temperatures.
1 Introduction
To address the key challenges of carbon neutrality, it is imperative to explore alternative energy carriers that can significantly reduce global reliance on fossil fuels. Ammonia (NH3) is increasingly recognized both as a hydrogen carrier and as a direct fuel source due to its carbon-free nature.1–3 Ammonia can be readily liquefied at room temperature under 0.9 MPa and possesses an overall twofold volumetric H2 density than pressurized hydrogen at 70 MPa. These favorable properties facilitate both the transportation and storage of ammonia. Additionally, electrified “green” ammonia produced via renewable-powered water electrolysis4,5 and electrocatalysis6,7 could position ammonia as a zero-carbon alternative to emissions-intensive hydrocarbon fuels. Thus, to meet our increasing power generation demand with fulfillment of carbon neutrality, direct combustion of ammonia is becoming a promising solution including internal combustion engines,8,9 gas turbines, and industrial furnaces.10,11Combustors that are designed for burning hydrocarbon fuels at high temperatures have typically employed stainless steel. However, our recent study12 shows that austenite stainless steel used in such combustors suffers from cumulative nitriding, which progressively modifies the surface chemical composition, hardens and embrittles the steel, potentially resulting in mechanical fractures during long-time operation. Moreover, the “unwanted nitriding” results in surface modifications, accompanied by the enhancement of surface reactivity13,14 that promotes the decomposition of ammonia (2NH3 → 3H2 + N2). These findings indicate that processes involving ammonia at high temperatures exhibit significant surface reactivity. Consequently, employing ammonia in ferrous combustors with extended operational durations requires an understanding of these surface effects.
In fact, nitriding is a conventional surface modification treatment used to enhance the hardness and wear resistance of iron-based materials. There are several nitriding approaches, including gas, salt bath, fluidized-bed, and plasma nitriding.15 Among these, gas nitriding is the most commonly used, and is typically conducted at temperatures ranging 773–853 K (500–580 °C) in gaseous ammonia over a period of a few tens of hours.16,17 The gas-nitriding mechanism of ferrous materials involves the absorption of ammonia onto the iron-based material surface, decomposition to nitrogen and hydrogen atoms, and diffusion of a small portion of the absorbed atoms into the solid, whereas the majority of nitrogen and hydrogen atoms recombine and desorb back into the gas phase. Once the molar concentration of diffused atoms becomes high enough, the material properties are modified.
The diffusion process can be modeled through Fick's second law:
On the experimental side, direct measurement of the diffusion coefficient at a prescribed nitrogen concentration has significant challenges. The experimental determination of the diffusion coefficient typically involves either weighing the nitriding sample21 or tracking the compound layer growth22 over time. These methods yield an averaged chemical diffusion coefficient that does not account for variations in local nitrogen concentration. In ammonia-fueled combustion systems, such as gas turbine combustors and industrial furnaces, the wall materials are exposed to ammonia for extended periods under various operating temperatures. For example, experiments conducted with a micro gas-turbine that produces 50 kW of net electrical power fueled by pure ammonia recorded combustor-liner metal temperatures, ranging from approximately 500 to 1000 °C.23 Therefore, in order to obtain accurate predictions for nitrogen profiles and layer thicknesses in these applications, a concentration- and temperature-dependent diffusion coefficient must be obtained.
Ab initio molecular dynamics (AIMD) calculations have been employed to investigate diffusional characteristics of systems such as Li-ion conductivity in solid electrolytes,24 lattice-vacancy diffusions,25 and surface adsorbates’ diffusions and reactions.26 However, most of these applications focus on processes with approximately 0.2 eV activation barriers, which can be observed in AIMD simulations with a system size of hundreds of atoms on a timescale of hundreds of picoseconds. On the other hand, the experimental apparent activation energy of nitrogen diffusion in iron nitrides typically exceeds 105 J mol−1, corresponding to a minimum activation energy of 1.0 eV, leading to slower reaction kinetics. These rare events associated with higher activation energy are even harder to observe directly in AIMD due to the formidable computational cost to model the required longer simulation timescale. Although the constrained MD method27,28 and the biased potential method29,30 have been applied to accelerate the sampling of rare events, they require meticulous selection of collective variables, which depend significantly on the reactive system.
Recently, the development of machine learning potentials (MLP, or machine learning force fields) has provided an opportunity to predict the potential energy surface (PES) with accuracy nearly comparable to quantum mechanical methods and efficiency comparable to classical molecular dynamics (MD).31 MLP enables the simulation of atomic systems over significantly larger scales and longer time spans as compared to AIMD,32 which offers a possibility to investigate reactive processes with activation energies from approximately 0.5 eV33 to 1.0 eV34 while the associated uncertainties on the diffusion barriers remain modest (roughly 0.04 eV to 0.15 eV relative to ground-truth DFT values). Consequently, accuracy loss is minimized compared with classical force field methods, which can exhibit errors of up to 0.5 eV during the same process.34
However, training and validating a MLP is not trivial, especially for reactive systems that undergo chemical bond cleavage and formation, typically in the nitriding process. Proper sampling of configurations and chemical space is essential for building high-quality MLPs. In fact, advanced sampling methods have been shown to effectively explore the PES and enhance the predictability of MLPs for reactive and diffusional systems. Notable methods include umbrella sampling35 and metadynamics.36 But unlike gaseous reactions, diffusion problems in solid-state materials involve reactants and products that differ based on the local environments of the diffusing atoms, which is hard to formulate in reaction coordinates. For example, in the interstitial solution of nitrogen in ε iron nitride, every two iron atoms offer an octahedral site that can host one nitrogen atom,37 and the many possible arrangements of neighboring nitrogen atoms create a rich variety of local atomic environments. This diversity poses challenges for sampling local environments of the diffusing atoms during MLP training. Recently, another alternative, the active learning method, has drawn significant attention.35,38,39 Active learning strategically samples conformations associated with high model uncertainty, thereby requiring a minimal number of first-principles calculations to achieve a prescribed threshold of accuracy.
Besides the challenges in sampling, most MLPs are developed to achieve an accuracy of less than tens of meV per atom in comparison with the ground-truth DFT energies on their test datasets, thereby serving as a sanity check of the predicted potential energy. However, low training and testing errors alone are not a reliable indicator of the applicability of the MLP, given the excellent interpolation achieved by ML algorithms against the training data. Even if some macroscopic properties, such as diffusion coefficient or conductivity, predicted by MLPs agree well with experimental observations, this consistency should not be interpreted as verification that these MLPs accurately represent the underlying atomic-level mechanisms. Especially for reactive potentials, without demonstration of the MLP's ability to reproduce the energy profile from the reactant to the product through the critical transition state, as validated against independent first-principles calculations, there are no guarantees that the MLP will reliably predict reactions not explicitly included in the training and testing data. Consequently, there is a possibility that MLPs may fortuitously approximate macroscopic measurements without faithfully capturing microscale phenomena.
Among all classical potentials, the only available Fe–N potential was developed using the modified embedded atom method (MEAM),40 which is not validated for diffusion of nitrogen in iron nitrides. On the other hand, MLPs have been used to investigate Fe–N systems, such as N2 dissociation on body-centered cubic (BCC) iron (111) facet,41 ammonia cracking on BCC iron (110) and (111) facets,42 and the diffusion of nitrogen atoms along the (100) direction into the BCC iron (110) facet.43 To the best of our knowledge, diffusion pathways and barriers for nitrogen in iron nitrides have not yet been studied using first-principles methods, nor has any MLP been demonstrated to model such phenomena. Thus, this study focuses on developing an MLP, independently validated for predicting diffusion barriers and energy profiles consistent with DFT calculations, with the goal of obtaining temperature- and concentration-dependent diffusion coefficients that are directly comparable with experimental observations.
This paper is arranged as follows: first, we describe the methods used to generate AIMD snapshots as training data using DFT, along with the methods used for training and validation of the MLP. Then, we outline the calculation procedure for determining diffusion coefficients from molecular dynamics simulations. In the results section, we first verify the accuracy and validity of the MLP to model nitrogen diffusion. Afterwards, we discuss the calculation of diffusion coefficients using this MLP and its comparison with existing experimental data. Finally, we conclude our work by outlining the key findings and observations from our analysis.
2 Methodology
The methodology in this study includes the generation of training data, training of MLPs, validation of MLPs, and calculation of diffusion coefficients. We first introduce the atomic models and describe the DFT calculation setups used to run AIMD and obtain the training data. The method for the independent validation of the diffusional energy profiles using a static DFT-based approach is also included. Then, we discuss the MLP training setups. Finally, we present the method for calculating the diffusion coefficient from MD simulations using MLP.2.1 MLP training data generation
In all the DFT calculations, the electronic self-consistent loop converges to a tolerance of 10−6 eV for energy. In DFT calculations for structure relaxations, the wavefunctions in a planewave basis set with a kinetic energy cutoff of 550 eV are set to minimize the effect of Pulay stress, and a Γ-centered k-point spacing of 0.25–0.3 Å−1 is adopted. In calculations with fixed volume supercells, the wavefunctions in a planewave basis set with a kinetic energy cutoff of 450 eV are set, and the Brillouin zone is sampled by a 1 × 1 × 1 Γ-centered k-point mesh to facilitate efficient AIMD calculations.
| ID | C(N) wt% | Atoms Fe![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N N |
Minimum cell size | AIMD@Temp [K] | AIMD with N add/removal | AIMD with Exp./Comp. | |
|---|---|---|---|---|---|---|---|
| 01 | Fe (mp-13) | 0.00% | 250![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 0 |
15.0 Å | 500, 1000, 1500 | +1/2 N | ±5% |
| 02 | Fe (mp-150) | 0.00% | 108![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 0 |
10.9 Å | 500, 1000, 1500 | +2/4 N | ±5% |
| 03 | Fe8N (mp-555) | 3.03% | 432![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 54 54 |
16.7 Å | 500, 1000, 1500 | ±2 N | ±5% |
| 04 | Fe4N (mp-535) | 5.88% | 256![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 64 64 |
14.8 Å | 500, 1000, 1500 | ±2 N | ±5% |
| 05 | Fe3N (mp-1804) | 7.69% | 384![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 128 128 |
15.9 Å | 500, 1000, 1500 | ±2 N | ±5% |
| 06 | Fe12N5 (mp-27908) | 9.43% | 288![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 120 120 |
12.7 Å | 500, 1000, 1500 | ±4 N | −5%/+3% |
| 07 | Fe2N (mp-248) | 11.11% | 162![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 81 81 |
12.7 Å | 500, 1000, 1500 | ±2 N | ±5% |
| 08 | Fe2N (mp-21476) | 11.11% | 144![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 72 72 |
12.7 Å | 500, 1000, 1500 | None | ±5% |
| 09 | FeN (mp-12120) | 20.00% | 64![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 64 64 |
10.0 Å | 500, 1000, 1500 | None | ±5% |
| 10 | FeN (mp-6988) | 20.00% | 256![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 256 256 |
16.8 Å | 500, 1000, 1500 | None | ±5% |
The conformations used to train the MLP in this work are those generated from AIMD runs using the following initial structures. Supercells for AIMD are constructed based on the relaxed conventional cells as detailed in Table 1, resulting in a total of 10 initial conformations. They are constructed to ensure a minimum cell periodicity is two times larger than the cutoff radius of MLP (in this work, rcutoff = 5.0 Å). Within the specified cutoff radius, each atom uniquely exhibits a distinct local atomic environment, ensuring that no two atoms share identical neighbors. Subsequently, these supercells constructed directly from conventional cells are subjected to a random nitrogen atom addition or removal in their octahedral site, as shown in Table 1 in column “AIMD with N add/removal”. For pure BCC iron (ID 01), either one or two nitrogen atoms are introduced into the octahedral sites formed by iron atoms, creating a supercell composed of 250 iron atoms plus either 1 or 2 nitrogen atoms. This is because the solubility of nitrogen in BCC iron can be as small as 0.1 wt%.49 Similar operations are performed for FCC iron (ID 02), but with two or four additions of nitrogen atoms. For iron nitrides (ID 03–08), two (or four for ID 06) nitrogen atoms are randomly subtracted from or added to the octahedral site, each of them generating a unique supercell. For iron nitrides (ID 08 and 09), no addition or removal of nitrogen atoms is performed, due to the extremely high nitrogen contents that are not expected to be achieved during exposure to ammonia at high temperatures. By slightly modifying the nitrogen concentration in each of these eight supercells, a degree of freedom on the nitrogen concentration can be explored by MLP in its local environment. The conformations are subsequently relaxed, in total adding 16 initial conformations for AIMD.
Since iron nitride is generated on the bulk iron, the lattice of iron nitride cannot be fully relaxed but is slightly compressed. Resembling the residual stress hardening effect, the high hardness of iron nitrides is attributed in part to the constrained or skewed cell, where the cell size is smaller than that of the relaxed ones. To address this, we introduce another degree of freedom in the cell volumes related to energies; an additional 20 conformations are constructed by scaling the lattice constant 5% smaller or 5% larger (3% for ID 06). The initial conformations undergo relaxation of their atomic positions with the cell size held constant, while the convergence tolerance for the atomic-force component is set at 0.02 eV Å−1.
2.2 DFT calculations for independent validation of MLPs
Despite achieving high training accuracy, MLPs are not guaranteed to have similar accuracies for data not explicitly included in the training. Thus, validation should include independent, application-relevant quantities.57 We use nitrogen diffusion barriers as an independent DFT benchmark: our training trajectories contain no nitrogen hopping, so these barriers are out-of-distribution for the MLP. Moreover, because diffusion coefficients depend exponentially on the migration barrier, capturing the nitrogen diffusion barriers is essential for predicting transport properties.To compute the diffusion barriers on the minimum energy path, climbing image nudged elastic band (CI-NEB) calculations are performed using the VTST tools58,59 in conjunction with VASP. The diffusion pathways of one additional nitrogen atom from an octahedral site to another adjacent one are computed in BCC iron (ID 01), FCC iron (ID 02), γ′ iron nitride (ID 04), and ε iron nitride (ID 05 and 06). The atomic models are built based on the initial supercells used to run AIMD at multiple temperatures, with the only difference being the addition of an interstitial nitrogen, considering the lattice symmetry. The cell size is kept constant, and the convergence tolerance for the atomic-force component is set at 0.02 eV Å−1 for all images. The spring constant between the images is set as 5.0 eV Å−2. It should be noted that the diffusion pathways calculated by DFT in this study are comprehensive but not exhaustive, due to the diversities associated with the local environment of the diffusing nitrogen mentioned in Section 1. Thus, the diffusional energy profiles obtained here serve as ground truths for validating MLPs and provide a quantitative assessment of some of the possible diffusion barriers in various iron nitrides. The CI-NEB calculations were likewise conducted with the present MLP and the MEAM40 in the large-scale atomic/molecular massively parallel simulator (LAMMPS)60 release 2Aug2023, using identical force-convergence tolerances and spring constants.
2.3 Training machine learning potentials
We employ DeePMD-kit61–63 version v3.0.0b0 to construct the MLP. DeePMD-kit exploits the idea of Behler–Parrinello-type MLP,64 where the energy of each atom within a structure is predicted by accounting only for the neighbors within a prescribed cutoff radius by a neural network. These individual energy predictions are then summed to determine the total energy of the structure.We chose the descriptor of deep potential smooth edition (DeepPot-SE) and cutoff radius rcutoff = 5.0 Å, with atom type embedding disabled. The descriptor embedding network is composed of three layers with sizes 30, 60, and 120 neurons, respectively, and the axis neuron count is set at 16. The multi-layer perceptron used for fitting the local energy consists of three layers, each containing 220 neurons. The hyperbolic tangent (tan![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) h) activation function was applied for each hidden layer, and the ResNet architecture without timestep is adopted. The loss function is built with perfectors on energy and force starting from 0.02 and 1000, respectively, and all decay to 1 at the end of the training. The learning rate starts at 0.001 and decays exponentially to 3.5 × 10−8. The model is trained over a total of 106 batches, with a batch size of 1 due to the large supercell, using the Adam optimizer.65 Further increasing the training steps does not yield obvious improvement in the validation error. For more detailed information on the deep potential-smooth edition framework, readers are encouraged to refer to the documentation provided with the DeePMD-kit.66
h) activation function was applied for each hidden layer, and the ResNet architecture without timestep is adopted. The loss function is built with perfectors on energy and force starting from 0.02 and 1000, respectively, and all decay to 1 at the end of the training. The learning rate starts at 0.001 and decays exponentially to 3.5 × 10−8. The model is trained over a total of 106 batches, with a batch size of 1 due to the large supercell, using the Adam optimizer.65 Further increasing the training steps does not yield obvious improvement in the validation error. For more detailed information on the deep potential-smooth edition framework, readers are encouraged to refer to the documentation provided with the DeePMD-kit.66
2.4 Determine the diffusion coefficient
The atomic model used for running MD to calculate the diffusion coefficient is constructed from the supercell detailed in Section 2.2. The supercell of Fe12N5, with four nitrogen atoms added, is selected to represent ε-iron nitride with a nitrogen content of 9.72 wt%, a compound frequently observed in gas nitriding experiments.67 This configuration also corresponds to the nitrogen concentration at the surface in our previous flame nitriding experiments.12 We constructed a larger supercell by scaling the original supercell dimensions by factors of 3, 3, and 4 along the a, b, and c lattice vectors’ direction, respectively. This results in a supercell measuring 56.1 Å × 56.1 Å × 50.8 Å and consisting of 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 832 atoms in total, including 10
832 atoms in total, including 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 368 iron and 4464 nitrogen atoms. MD simulations were performed at temperatures higher than the classical gas nitriding temperature, ranging from 1000 to 1500 K, with an increment of 100 K, to obtain an efficient estimation through ensemble averaging.
368 iron and 4464 nitrogen atoms. MD simulations were performed at temperatures higher than the classical gas nitriding temperature, ranging from 1000 to 1500 K, with an increment of 100 K, to obtain an efficient estimation through ensemble averaging.
MD simulations of the nitrogen diffusion in the ε iron nitride were performed using LAMMPS60 (2Aug2023). The DeePMD pair style was used to drive MD with MLP, which was developed by the DeePMD-kit. A binning algorithm was used to create the neighbor list, and a 1.0 Å skin distance was incorporated. The equations of motion were integrated using a 1.0 fs timestep, chosen to facilitate a fair comparison across different temperatures while maintaining the stability of the simulation. Fe and N atom masses were set to 55.845 and 14.0068 µ, respectively.
The diffusion coefficients can be computed through two main types of MD simulations. First, the equilibrium MD, which employs the Stokes–Einstein–Sutherland equation68,69 or the Green–Kubo relations,70,71 operates without external driving forces and provides an estimation of the self-diffusion coefficient. Second, the non-equilibrium MD, utilizing the color-diffusion method72 determines the same self-diffusion coefficient by imposing an external body force on the solute to mimic the chemical-potential gradient. Some non-equilibrium MD73,74 directly imposed a concentration gradient, therefore yielding the chemical (Fickian) diffusion coefficient rather than the self-diffusion coefficient. Since all non-equilibrium MD simulations require an external driving force, they can facilitate the observation of mass transportation of slow species, particularly at low temperatures. However, capturing anisotropic diffusion in solid solutes requires directional computations and separate atomic models depending on the crystal symmetry.75 We therefore adopted the most straightforward approach – equilibrium MD using Stokes–Einstein–Sutherland equation68,69 –, since the diffusion coefficients of interest occur at high temperatures, where they can be effectively observed through equilibrium MD simulations.
We conducted the simulation using an NPT ensemble initially for 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 fs at the target temperature, allowing the supercell's volume to stabilize. Following the NPT simulation, an NVT ensemble was employed to perform simulated annealing to randomize the supercell further. This involves elevating the system temperature from the target temperature to 1500 K over 20
000 fs at the target temperature, allowing the supercell's volume to stabilize. Following the NPT simulation, an NVT ensemble was employed to perform simulated annealing to randomize the supercell further. This involves elevating the system temperature from the target temperature to 1500 K over 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 fs, maintaining it at 1500 K for another 20
000 fs, maintaining it at 1500 K for another 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 fs, and finally cooling it to the target temperature over 20
000 fs, and finally cooling it to the target temperature over 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 fs. The temperature was ramped linearly during the heating and cooling process using the Nosé–Hoover76,77 thermostat, and this method was consistently employed to control the temperature in all MD simulations presented in this work. Following the simulated annealing, equilibrium molecular dynamics simulations using the NVT ensemble were maintained to further run the system at the target temperature. Simulations above 1000 K were performed for over 1.25 ns, those at 1000 K for over 3 ns, and those at 830 K for 6 ns. Simulation data were archived every 100 fs for subsequent analysis.
000 fs. The temperature was ramped linearly during the heating and cooling process using the Nosé–Hoover76,77 thermostat, and this method was consistently employed to control the temperature in all MD simulations presented in this work. Following the simulated annealing, equilibrium molecular dynamics simulations using the NVT ensemble were maintained to further run the system at the target temperature. Simulations above 1000 K were performed for over 1.25 ns, those at 1000 K for over 3 ns, and those at 830 K for 6 ns. Simulation data were archived every 100 fs for subsequent analysis.
The Stokes–Einstein–Sutherland equation68,69 is used to calculate the self-diffusion coefficient from MD trajectories as:
The mean squared displacement (MSD) is defined by
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) self is the self-diffusion coefficient, t is the time, dα is the dimensionality (equals to 3 in this study for three-dimensional diffusion), N is the total number of atoms, r is the position, and α is the x, y or z Cartesian coordinates of the atomic center of mass. We discard a conservative burn-in window before MSD fitting, which is no more than half of the total simulation time for each run, detailed in the SI (Table S4). In practice, a weak dependence of the selection of r0 for the temporal slope of the MSD is observed. Therefore, instead of performing a single long MD simulation, four separate MD simulations are performed at each target temperature, each initialized with a different random distribution of velocities. This approach is used to adequately sample the phase space and estimate the ensemble average of the diffusion coefficient obtained.
self is the self-diffusion coefficient, t is the time, dα is the dimensionality (equals to 3 in this study for three-dimensional diffusion), N is the total number of atoms, r is the position, and α is the x, y or z Cartesian coordinates of the atomic center of mass. We discard a conservative burn-in window before MSD fitting, which is no more than half of the total simulation time for each run, detailed in the SI (Table S4). In practice, a weak dependence of the selection of r0 for the temporal slope of the MSD is observed. Therefore, instead of performing a single long MD simulation, four separate MD simulations are performed at each target temperature, each initialized with a different random distribution of velocities. This approach is used to adequately sample the phase space and estimate the ensemble average of the diffusion coefficient obtained.
Since the self-diffusion coefficient is a tensor, the diagonal components are averaged using the following equations for comparison with polycrystalline iron-based materials that do not exhibit orientation preference.
where Dself represent the self-diffusion coefficient that can be obtained through the present MD simulation by calculating MSD. μi is the chemical potential of the diffusing component i, J is the flux of particles. R is the gas constant, T is the temperature, and ci is the concentration of component (i) at which the self-diffusion occurs.
The diffusion coefficient is defined under a concentration gradient,
| J = −D∇ci, |
| ∇ci = cT∇xi |
Again, at a small gradient of other factors (temperature or pressure) influencing μi,
Thus, the relation between Dself and D is given by the thermodynamic factor ϕ as77
We adopted Frisk's two-sublattice model78 to formulate the chemical potential μi of ε-iron nitride, in the expression Fea(N,Va)c, where a and c are parameters to describe the component with the following conventions. In the first sublattice, iron atoms form an HCP structure with octahedral sites. In the second sublattice, octahedral sites accommodate either nitrogen (N) or vacancy (Va). For ε-iron nitride, a = 1 and c = 0.5, indicating that, at the maximum nitrogen concentration, each iron atom accommodates 0.5 nitrogen atoms, corresponding to Fe2N (Table 1, ID 07). Because this phase is above the convex hull, it will spontaneously transfer into another phase of Fe2N (Table 1, ID 08),79 known as ζ-iron nitride.
For one molar formula unit of Fea(N,Va)c, the Gibbs energy is modeled as
From the definition of chemical potential,
For one molar formula unit of Fea(N,Va)c, NN can be written as yN/c. Considering yVa is also related to yN, we can derive the chemical potential of nitrogen μN
In the interstitial diffusion problems, since the interstitial component has little contribution to the total volume,22,80 the concentration variable (ui) is used instead of the molar fraction in the thermodynamics factor:
where ui is the concentration variable and xj is the component forming the first sublattice with significant contribution to the total volume. For one molar formula unit of Fea(N,Va)c, the concentration variable of nitrogen is
Note that yN + yVa = 1, we derived the thermodynamics factor ϕ as
 | (1) |
3 Result and discussion: machine learning potentials
3.1 Training the machine learning potentials
We used the snapshots of AIMD described in Section 2.1 to train the MLPs. Given the small timesteps used in the AIMD simulations, the training configurations over consecutive timesteps are geometrically and energetically similar. Hence, to make sure that the training data contains a wide range of configurations, we employed subsampling of the AIMD data. For each continuous AIMD run, 20% of the snapshots were randomly sampled, ensuring that the training data conformations are not geometrically and energetically similar. Of these sampled conformations, 80% were used for training and 20% used for validation. Given the additional CI-NEB calculations and subsequent diffusion analysis, we did not utilize a test dataset. We observed that, in some of the AIMD simulations conducted at 1500 K, less than 10% of the SCF calculations did not converge. For each of these steps, both the subsequent step and the preceding step were discarded before subsampling to minimize error propagation. This non-convergence would not affect the present MLP training, as the AIMD simulations themselves are not directly used for any statistical estimations.The conditions employed to run AIMD are detailed in Table 1. After subsampling, there are mainly 5 groups of datasets: (1) 500 K, (2) 1000 K, (3) 1500 K, (4) addition or removal of nitrogen atoms (shortened as add/removal N), and (5) expansion or compression of the cell size (shortened as Exp./Comp.). A detailed training data description is provided in SI (Tables S1–S3). To systematically assess the training accuracy and the validity of the MLPs, we trained four different MLPs with the same hyperparameters and initialization described in Section 2.3. For each MLP, MLP[500 + 1000 + 1500 K] refers to the MLP trained with the data generated under 500, 1000, and 1500 K, while MLP[all] simply refers to the MLP trained with all groups of data.
3.2 Validation of the machine learning potentials
Validation of the MLPs have been carried out primarily through mean absolute errors (MAEs) on the training and validation sets. Typical MAEs for well-converged MLPs exhibit ∼0.005 eV for energies and ∼0.05 eV Å−1 for forces.82 However, these values depend on dataset size and diversity, so they reveal only interpolation skill. To validate transferability to diffusion problems at finite temperature, besides the MAEs, we also tested the predicted lattice constants, energy-volume relations, and diffusion barriers using the present MLP, and compared them with those obtained through DFT. The barrier conformations are absent from the training data and therefore form the strictest benchmark. | ||
| Fig. 1 (a) Energy and (b) force mean absolute error (MAE) on 4 groups of validation data, tested on 3 MLP models. | ||
Comparing the MLP[500 + 1000 + 1500 K] with MLP[500 + 1000 K], the inclusion of high temperature data of 1500 K proves to be overall favourable. As shown from the test dataset of 1500 K, incorporating high temperature data facilitates the prediction of system energies and forces at high temperature. In addition, it also enhances the prediction accuracy of system energy for different nitrogen concentrations not included in the training of both MLP[500 + 1000 K] and MLP[500 + 1000 + 1500 K]; the inclusion of high-temperature data improves the accuracy for the test dataset involving the addition and removal of nitrogen atoms (add/remove N). Incorporating training data for the addition and removal of nitrogen atoms (add/remove N) further improves the prediction error of system energy from around 4 to 2 meV atom−1. On the other hand, the accuracy in the atomic force predictions remains unchanged.
By comparing the MLPs tested using the volume expansion/compression (Vol-Exp/Comp) dataset, it is found that the high temperature data at prescribed volume does not effectively improve the accuracy of cell volume-to-energy relations. The tested MAE of energy is kept as high as over 40 meV atom−1. Although using the high-temperature data improves the MAE of force, the poor performance of the prediction on energy remains noteworthy, thus necessitating the explicit inclusion of DFT data for different cell volumes.
In terms of the overall performance, the MLP[all] model reached MAE of less than 5 meV atom−1 and the root mean square error (RMSE) of approximately 8 meV atom−1 (not shown here). We observed that the largest atomic force is up to 13 eV Å−1 in the training dataset. The tested MAE on atomic force is less than 0.15 eV Å−1 as shown in Fig. 1b, and the RMSE of the atomic force is also less than 0.15 eV Å−1 (not shown here). The MAE is slightly larger than that of the previous MLP study83 targeting reactive process in the gas phase (MAE ∼ 0.12 eV Å−1), and the RMSE is slightly smaller than the results obtained for a reactive process in Fe–C alloys84 (RMSE ∼ 0.17 eV Å−1). Validation of the MLP against the AIMD reference data is provided in the SI. Energy and force parity plots are shown in Fig. S3 and S4, respectively. Quantitative test errors are summarized in Tables S5–S8 (force MAE/RMSE in Tables S5, S6 and energy MAE/RMSE in Tables S7 and S8).
 | ||
| Fig. 2 Compressed and expanded cell lattice constants to energy relation in (a) γ-Fe; FCC iron, (b) γ′-Fe4N iron nitride and (c) ε-Fe12N5 iron nitride. | ||
However, as also shown in Fig. 2, adding a small dataset accounting for the cell volume to energy relationship (Vol-Exp./Comp.) significantly enhances the ability of the MLP to predict a concave-upward relationship. The smooth concave-upwards profile also prevents the system from becoming trapped in the incorrectly predicted local minima (e.g., MLP[500 + 1000 + 1500 K, add/remove N] in Fig. 2(a), a/aeq = 0.925).
| Crystal system | DFT | MLP[all] | MEAM | ||
|---|---|---|---|---|---|
| 1 | Fe | Cubic | a = b = c = 2.799 | a = b = c = 2.834 | a = b = c = 2.864 |
| (mp12) | |||||
| 2 | Fe | Cubic | a = b = c = 3.419 | a = b = c = 3.432 | a = b = c = 3.619 |
| (mp-150) | |||||
| 3 | Fe8N | Tetragonal | a = b = 5.612 | a = b = 5.588 | a = b = 5.652 |
| (mp-555) | c = 6.185 | c = 6.123 | c = 6.246 | ||
| 4 | Fe4N | Cubic | a = b = c = 3.755 | a = b = c = 3.760 | a = b = c = 3.792 |
| (mp-535) | |||||
| 5 | Fe3N | Hexagonal | a = b = 4.618 | a = b = 4.606 | a = b = 4.635 |
| (mp-1804) | c = 4.247 | c = 4.279 | c = 4.289 | ||
| 6 | Fe12N5 | Trigonal | a = b = 9.353 | a = b = 9.321 | a = b = 9.497 |
| (mp-27908) | c = 4.230 | c = 4.269 | c = 4.342 | ||
| 7 | Fe2N (mp-248) | Trigonal | a = 4.271 | a = 4.304 | a = 4.350 |
| b = 4.716 | b = 4.732 | b = 4.794 | |||
| c = 5.430 | c = 5.421 | c = 5.494 | |||
| 8 | Fe2N | Orthorhombic | a = b = 4.722 | a = b = 4.726 | a = b = 4.800 |
| (mp-21476) | c = 4.245 | c = 4.277 | c = 4.323 | ||
| 9 | FeN | Cubic | a = b = c = 4.211 | a = b = c = 4.200 | a = b = c = 4.254 |
| (mp-6988) | |||||
| 10 | FeN | Hexagonal | a = b = 2.711 | a = b = 2.730 | a = b = 2.738 |
| (mp-12120) | c = 4.934 | c = 4.962 | c = 4.947 |
Nitrogen diffusion was calculated in a 5 × 5 × 5 supercell constructed from the α-Fe (ID 01, BCC) conventional cell. As shown in Fig. 3a, we simulated the diffusion of a single nitrogen atom within the supercell from one octahedral site to another adjacent site, representing diffusion in a diluted solution. MLP[500 + 1000 K] trained only with low-temperature data roughly duplicates the energy profile, but there is a discrepancy at the transition state. The discrepancy is reduced when the training data are added. Although the prediction of MLP agrees well with the DFT calculation in a relatively low energy range (0 eV to 0.4 eV), MLP[all] still underestimates the diffusion barrier by approximately 0.05 eV at the transition state. The MEAM potential gives a sharper profile than DFT, but predicts the diffusion barriers nearly identical to those from DFT in this case.
As shown in Fig. 4, nitrogen diffusion was calculated in a 4 × 4 × 4 γ-Fe (ID 02, FCC) supercell. Because the path is symmetric,52 only the segment from the octahedral site (reaction coordinate equals 0) to the tetrahedral site (reaction coordinate equals to 1) was evaluated illustratively shown in Fig. 4a. The MLP[500 + 1000 K], which is trained on the low-temperature data, significantly underestimates the barrier as shown in Fig. 4b. By adding the 1500 K, add/remove-N, and Vol-Exp/Comp dataset, the energy profile progressively approaches the DFT value in the low energy range (<1.5 eV) but still underestimated the diffusion barrier by 0.24 eV. The MLP[all] model correctly captures the tetrahedral local minimum. On the other hand, MEAM fails to capture the local minimum, underestimates energies near the octahedral site, and overestimates the barrier by 0.26 eV.
Nitrogen diffusion in γ′-Fe4N (ID 04) iron nitride was calculated with a 4 × 4 × 4 supercell as shown in Fig. 5. In comparison with the γ-Fe, the metal lattice remains FCC, but the lattice nitrogen deflects the path of diffusion away from the tetrahedral site. Prediction of MLP[500 + 1000 K] matches DFT below 0.75 eV but overpredicts the barrier. Similarly, adding more training data yields a barrier essentially identical to DFT. MEAM significantly overestimates the energy profile and gives a barrier 0.5 eV higher than DFT. None of the potentials reproduces the energy plateau observed near the transition state in DFT.
Fig. 6 and 7 show the predictions of N diffusion in ε-Fe12N5 (2 × 2 × 3 supercell) and ε-Fe3N (3 × 3 × 3 supercell). The octahedral start and end sites of diffusion are shown in Fig. 6a and 7a. Every nearest-neighbor hop of an added nitrogen atom was calculated, within an HCP layer (Fig. 6b, d, and 7c) and across layers (Fig. 6c and 7b). The MLP[500 + 1000 K] closely reproduces the DFT barriers. Adding more training data yields minor improvements for in-layer hops but slightly degrades the accuracy of cross-layer predictions. MLP[all] still overestimates the barrier by 0.15 eV as shown in Fig. 6c and 0.23 eV in Fig. 7b. Again, the MEAM potential significantly overestimates both the energy difference between different octahedral sites and the activation barriers of all calculated pathways in the ε-iron nitride.
In summary, MLPs reproduce nitrogen diffusion barriers in iron and iron nitrides well, even though no diffusional conformations were included in training. The validated diffusion barriers exhibit a maximum error of 0.24 eV (in γ-Fe of 2.32 eV barrier) for the best performing MLP, whereas the MEAM potential exhibits a maximum error up to 2.74 eV (in ε-Fe12N5 of 2.75 eV barrier). Errors in the predicted barrier are therefore larger than the MAEs as described in Section 3.2.1, but remain moderate. A model trained on the low-temperature data (500 K and 1000 K) already closely follows DFT in most systems. Adding 1500 K, add/remove N, and Vol-Exp./Comp. consistently provides improvement. Energy wells are predicted accurately, but some transition states are still under- or overestimated. It is probably because those geometries are absent in the training. This gap could potentially be improved by a more advanced sampling method, such as meta-dynamics. Importantly, the CI-NEB optimizations were performed independently with DFT, MLP, and MEAM, all initialized with identical images and subjected to the same convergence criteria. The converged MLP[all] pathways closely reproduced the DFT minimum energy paths, capturing not only the barrier heights but also the overall shapes of the energy profile along the reaction coordinates. This shows that the learned PES provides sufficiently accurate forces to drive the optimizer to the same saddle points and local minima rather than merely matching single-point energies. This geometry-level agreement, achieved without any diffusional conformations in the training set, constitutes a stringent validation of MLP transferability. Such equivalent pathway comparisons—identical starting images, optimizer, and tolerances across methods—are rarely reported in the MLP literature, yet are crucial for validating true transferability beyond single-point energies. In contrast, MEAM systematically converged to markedly overestimated barriers.
The possible reasons of MEAM's barrier discrepancy is discussed below. Because the Fe–N MEAM potential40 was fitted mainly to equilibrium properties (heats of formation and lattice constants of iron nitrides), it is well constrained near stable minima but not necessarily in the migration saddle-point (bottleneck) region that governs diffusion barriers. Moreover, finite-temperature diffusivities can be influenced by anharmonic vibrational contributions to both the migration free energy and prefactor, which are not explicitly constrained by such equilibrium fitting. This is consistent with recent work highlighting the importance of higher-order anharmonicity in high-temperature lattice dynamics and transport.86
3.3 Thermodynamic factor for diffusion coefficient
To enable comparison between the self-diffusion coefficient obtained from the MLP driven MD and the chemical (Fickian) diffusion coefficient measured experimentally, we calculated the thermodynamic factor for a typical iron nitride system, as described in Section 2.4.1. Since the chemical potential is an extensive quantity, the calculation is performed based on the unscaled supercell rather than the one build for the MD simulations in Section 2.4.As introduced in Section 2.4.1, for ε-iron nitrides, a = 1 and c = 0.5, thus b = a/c = 2. The system includes additional 4 nitrogen atoms per supercell, which is constructed from a 2 × 2 × 3 arrangement of the conventional cell. The resulting supercell consists of 288 iron and 124 nitrogen atoms, therefore yN = 124/144. The concentration variable of nitrogen is then uN = cyN/a = 124/288. With the thermodynamics data for ε iron nitride from H. Du,81
L0Fe:N,Va = 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 012 − 19.9853·T 012 − 19.9853·T |
| L1Fe:N,Va = −9446.0 + 9.3471·T |
Plugging the L1Fe:N,Va, L0Fe:N,Va, c, b and uN into eqn (1), we derived the thermodynamic factor for ε-iron nitride containing 9.72 wt% nitrogen as
Table 3 presents the estimated thermodynamic factor for the ε-iron nitride described above at 1500 K, 1000 K, and 830 K. The thermodynamic factor is greater than unity, indicating strong solute–solute (N–N) repulsions when the nitrogen weight percent is as high as 9.72 wt%.
| Temperature | 1500 K | 1000 K | 830 K |
| Thermodynamic factor (dimensionless) | 9.98 | 11.42 | 12.31 |
3.4 Prediction of diffusion coefficient on ε-iron nitride
Self-diffusion coefficients were ensemble-averaged over four independent MD runs using MLP[all] at each temperature, as described in Section 2.4.1. The Pearson correlation coefficients obtained from linear regression of MSD versus time at each temperature were greater than 0.999 for temperatures above 1300 K, exceeded 0.997 at 1200 K and 1100 K, and remained higher than 0.99 at 1000 K, demonstrating good linearity. However, the correlation coefficients estimated at 830 K were 0.787, 0.796, 0.802, and 0.340 for the four independent runs, respectively, indicating relatively poor statistical sampling even with a 6 ns simulation. Their standard deviations were propagated to log(D), and converted to 95% confidence intervals with Student's statistics87 (ν = 3). Multiplying the averaged self-diffusion coefficients by the thermodynamic factors, obtained in Section 2.4.2 and III.C, yielded the chemical diffusion coefficients plotted in Fig. 8. The computed MSD, along with the self-diffusion coefficients and the thermodynamic factors at each temperature, are available in the SI (Table S4 and Fig. S1). As introduced in Section 2.4.1, MSD was computed using a single time origin for each trajectory and averaged over four independent runs. A sensitivity test using multiple time origins (up to 1000 origins at 1000 K) produced smoother MSD curves but did not significantly reduce the uncertainty of the MSD slope (and diffusivity) when temporal correlation was accounted for via block bootstrap; the dominant uncertainty arose from variability across independent runs. | ||
| Fig. 8 Calculated chemical diffusion coefficient of ε-iron nitride; corresponding nitrogen weight percent 9.72 wt%. In comparison with experimental works (refers to the main text). | ||
We observed that the uncertainty is minimized at 1300 K, with larger values at both higher and lower temperatures. Above 1300 K, the uncertainty increases because the PES was trained only up to 1500 K. Below 1300 K, the uncertainty increases because the phase space is sampled less comprehensively. The limited number of diffusion events at these temperatures introduces statistical errors, particularly in the diffusion properties derived from averages over sampled configurations. A detailed discussion about the error at low temperatures is provided in the SI. It is noted that we did not observe the finite size effect (shown in Fig. S2), which has been reported in the MLP-driven MD-based diffusion coefficient computation of liquid.88 The diffusion coefficient obtained from a system with 3906 atoms was consistent with that of 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 888 atoms from 1300 to 1500 K, despite greater fluctuations in the MSD.
888 atoms from 1300 to 1500 K, despite greater fluctuations in the MSD.
The calculated chemical (Fickian) diffusion coefficient of the ε-iron nitride as a function of temperature is shown by the Arrhenius plot in Fig. 8. The experimental data in previous studies89–91 are also shown for comparison. The temperature range in previous experiments is represented by solid lines, while dashed lines correspond to the extrapolated data. Extrapolating results from low to high temperatures should have higher uncertainty, as the solute's mobility is limited at low temperatures.
We performed weighted linear regression, using the inverse of the variance of each point as the weighting factor. The fitted Arrhenius expression is D = A![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) exp(−Ea/RT), where D is the chemical (Fickian) diffusion coefficient, A is the pre-exponential factor, Ea is the activation energy, R is the gas constant, and T is the absolute temperature. The fitted results are shown in Table 4. The activation energy obtained in this work is consistent with the experimental result obtained by Lakhtin and Kogan.90 The pre-exponential factor is slightly larger than that reported by Lakhtin and Kogan.90
exp(−Ea/RT), where D is the chemical (Fickian) diffusion coefficient, A is the pre-exponential factor, Ea is the activation energy, R is the gas constant, and T is the absolute temperature. The fitted results are shown in Table 4. The activation energy obtained in this work is consistent with the experimental result obtained by Lakhtin and Kogan.90 The pre-exponential factor is slightly larger than that reported by Lakhtin and Kogan.90
Using the Arrhenius expression, the diffusion coefficient at 823 K, where most gas nitriding experiments are conducted, is computed as shown in Table 5. The prediction in this work is in reasonable agreement with the experimental data.
As part of estimating the error, this study is compared with prior research that calculated diffusion of nitrogen in BCC iron using the MEAM potential at 0.23–1.5 wt% nitrogen concentration.19 The standard deviation of the computed diffusion coefficient in that study was 7% of the mean value at 1173 K, while in this study, the standard deviation is 7.1% of the mean value at 1200 K. This suggested present study has a reasonable error range when estimating diffusion coefficient through ensemble averaging. In another previous research examining the diffusion of carbon in cementite (Fe3C), the MD-computed diffusion coefficients were approximately an order of magnitude smaller than the experimental value.92 By contrast, the present study shows smaller discrepancies that fall nearly within the experimental error range, which is attributed to the better overall accuracy of our MLP. Taken together, these results demonstrate that this work achieves a better computational accuracy and shows a good agreement with experimental data, which is often challenging to obtain in MD simulations due to the approximations and simplifications inherent in atomic models.
To the best of our knowledge, this is the first MLP trained and validated to capture nitrogen diffusion across diverse Fe–N phases (α-Fe, γ-Fe, γ′-Fe4N, ε-Fe3N/Fe12N5). Its fidelity was established through independent like-for-like CI-NEB pathway matching and energy-to-volume curves. Training drew on 8640 energy/forces-labeled snapshots (128–512 atoms each) spanning 500–1500 K, nitrogen atom addition/removal, and large volumetric strains. Given this scope, we defer broader parametric sweeps to future work. This MLP will serve as the foundation for mapping concentration/temperature-dependent nitrogen transport behavior in Fe–N systems.
4 Conclusions
We have developed a machine-learning interatomic potential (MLP) trained by the DFT data for molecular dynamics (MD) simulation of Fe–N systems and demonstrated the ability of the MLP–MD approach to predict the nitrogen atom diffusion rate in iron and iron nitrides. The results show that the machine-learning potential is accurate not only for predicting energy and forces against the training and testing data, but also for the diffusional profile related to the physical diffusion process of interest – an aspect often overlooked in previous studies. In addition, the MLP–MD model can well-reproduce the experimentally measured chemical diffusion coefficients of nitrogen in ε-iron nitride. Specifically, the following conclusions have been derived.– Incorporating problem-specific structural diversity into the training data is essential for constructing a robust machine-learning interatomic potential. Adding high-temperature configurations to the low-temperature baseline clearly reduces both energy and force prediction errors at high temperature and even improves extrapolation to unseen nitrogen contents. Introducing explicit N-addition/removal snapshots further halves the energy MAE from 4 to 2 meV atom−1, although the atomic force MAE remains unchanged. By contrast, high-temperature data alone fail to capture the cell-volume-energy relationship. Therefore, a dedicated volume-expansion/compression dataset is required to suppress these errors. The present composite dataset provides a good balance across thermodynamic, compositional, and volumetric perturbations, leading to overall MAE below 4.1 meV atom−1.
– The classical MEAM potential fails to predict the diffusional profile and significantly overestimates the barriers. The present MLP significantly outperforms the MEAM potential.
– MD simulations using the present MLP yield nitrogen atoms’ diffusivity in ε-iron that quantitatively agree with experimental data. Specifically, the Arrhenius analysis provides an activation energy of 1.57 × 105 J mol−1 and a pre-exponential factor of 1.96 × 10−4 s−1, both consistent with reported values. The predicted diffusion coefficient at 823 K is 2.17 × 10−14 m2 s−1, lying within the uncertainty interval of the experimental data, significantly outperforming earlier atomistic studies that underpredicted the diffusion coefficient by an order of magnitude.
Although the effects of phase morphology, grain boundaries, lattice dislocations and point defects in iron nitrides are not taken into account in the present study, the quantitative agreement of the present results with the experimental data indicates that the diffusion of nitrogen atoms’ in ε-iron nitrides should be primarily governed by the interstitial diffusional properties within the lattice over the existing experimental temperature range.
The present findings underscore the superior transferability and quantitative accuracy of the present MLP for nitrogen atoms’ diffusion in iron nitrides, paving the way for predicting concentration- and temperature-dependent diffusion coefficients in Fe–N systems based on ab initio data.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). The DFT dataset and trained machine-learning potential model (MLP[all]) are archived at Zenodo at https://doi.org/10.5281/zenodo.18220490. Supplementary information contains details of the AIMD training and validation datasets, additional MSD data and statistical analyses, and MLP validation through parity plots and tabulated energy and force errors. See DOI: https://doi.org/10.1039/d6cp00172f.Acknowledgements
This work was supported by JSPS KAKENHI Grant Numbers 21H04539 and NEDO Feasibility Study Program “Development of Carbon-neutral Industrial Furnace by Innovative Ammonia Combustion” funded by NEDO, Japan. PF is supported by JSPS KAKENHI Grant Numbers 24KJ0711. YJ and AL were supported by DOE BES research grant DE-SC0021135 and NSF ammonia nitridation grant and OIA-2428523. The computation in this work was performed using the facilities of the Supercomputer Center, the Institute for Solid State Physics, the University of Tokyo (ISSPkyodo-SC-2025-Ca-0007, 2025-Cb-0001).References
- B. Shiozawa, in CO2 Free Ammonia as an Energy Carrier, ed. K. Aika and H. Kobayashi, Springer Nature Singapore, Singapore, 2023, pp. 29–47 Search PubMed.
- M. B. Bertagni, R. H. Socolow, J. M. P. Martirez, E. A. Carter, C. Greig, Y. Ju, T. Lieuwen, M. E. Mueller, S. Sundaresan, R. Wang, M. A. Zondlo and A. Porporato, Minimizing the impacts of the ammonia economy on the nitrogen cycle and climate, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2311728120 CrossRef CAS PubMed.
- A. Valera-Medina, H. Xiao, M. Owen-Jones, W. I. F. David and P. J. Bowen, Ammonia for power, Prog. Energy Combust. Sci., 2018, 69, 63–102 CrossRef.
- H. Kobayashi, A. Hayakawa, K. D. K. A. Somarathne and E. C. Okafor, Science and technology of ammonia combustion, Proc. Combust. Inst., 2019, 37, 109–133 CrossRef CAS.
- A. Valera-Medina, F. Amer-Hatem, A. K. Azad, I. C. Dedoussi, M. De Joannon, R. X. Fernandes, P. Glarborg, H. Hashemi, X. He, S. Mashruk, J. McGowan, C. Mounaim-Rouselle, A. Ortiz-Prado, A. Ortiz-Valera, I. Rossetti, B. Shu, M. Yehia, H. Xiao and M. Costa, Review on Ammonia as a Potential Fuel: From Synthesis to Economics, Energy Fuels, 2021, 35, 6964–7029 CrossRef CAS.
- Y. Xu, N. Liu, Y. Lin, X. Mao, H. Zhong, Z. Chang, M. N. Shneider and Y. Ju, Enhancements of electric field and afterglow of non-equilibrium plasma by Pb(ZrxTi1−x)O3 ferroelectric electrode, Nat. Commun., 2024, 15, 3092 CrossRef CAS PubMed.
- Z. Zhang, C. Kondratowicz, J. Smith, P. Kucheryavy, J. Ouyang, Y. Xu, E. Desmet, S. Kurdziel, E. Tang, M. Adeleke, A. D. Lele, J. M. Martirez, M. Chi, Y. Ju and H. He, Plasma-Assisted Surface Nitridation of Proton Intercalatable WO3 for Efficient Electrocatalytic Ammonia Synthesis, ACS Energy Lett., 2025, 3349–3358 CrossRef PubMed.
- C. Lhuillier, P. Brequigny, F. Contino and C. Mounaïm-Rousselle, Experimental study on ammonia/hydrogen/air combustion in spark ignition engine conditions, Fuel, 2020, 269, 117448 CrossRef CAS.
- M.-C. Chiong, C. T. Chong, J.-H. Ng, S. Mashruk, W. W. F. Chong, N. A. Samiran, G. R. Mong and A. Valera-Medina, Advancements of combustion technologies in the ammonia-fuelled engines, Energy Convers. Manage., 2021, 244, 114460 CrossRef CAS.
- M. Tamura, T. Gotou, H. Ishii and D. Riechelmann, Experimental investigation of ammonia combustion in a bench scale 1.2 MW-thermal pulverised coal firing furnace, Appl. Energy, 2020, 277, 115580 CrossRef CAS.
- W. D. Monnery, K. A. Hawboldt, A. E. Pollock and W. Y. Svrcek, Ammonia Pyrolysis and Oxidation in the Claus Furnace, Ind. Eng. Chem. Res., 2001, 40, 144–151 CrossRef CAS.
- D. Wang, Y. Xing, M. Lee and Y. Suzuki, Effects of wall temperature and water vapor on the nitriding of stainless steel induced by ammonia flames, Proc. Combust. Inst., 2024, 40, 105562 CrossRef CAS.
- J. Tseng, D. Gu, C. Pistidda, C. Horstmann, M. Dornheim, J. Ternieden and C. Weidenthaler, Tracking the Active Catalyst for Iron-Based Ammonia Decomposition by In Situ Synchrotron Diffraction Studies, ChemCatChem, 2018, 10, 4465–4472 CrossRef CAS.
- P. Feng, M. Lee, D. Wang and Y. Suzuki, Ammonia thermal decomposition on quartz and stainless steel walls, Int. J. Hydrogen Energy, 2023, 48(75), 29209–29219 CrossRef CAS.
- J. L. Dossett and G. E. Totten, ASM handbook. 4A: Steel heat treating fundamentals and processes, ASM International, 2013 Search PubMed.
- Y.-L. Zhou, F. Xia, A.-J. Xie, H.-P. Peng, J.-H. Wang and Z.-W. Li, A Review—Effect of Accelerating Methods on Gas Nitriding: Accelerating Mechanism, Nitriding Behavior, and Techno-Economic Analysis, Coatings, 2023, 13, 1846 CrossRef CAS.
- N. Duong Nam, N. Anh Xuan, N. Van Bach, L. T. Nhung and L. T. Chieu, Control Gas Nitriding Process: A Review, J. Mech. Eng. Res. Dev, 2019, 42, 17–25 Search PubMed.
- M. A. J. Somers and E. J. Mittemeijer, Layer-growth kinetics on gaseous nitriding of pure iron: Evaluation of diffusion coefficients for nitrogen in iron nitrides, Metall. Mater. Trans. A, 1995, 26, 57–74 CrossRef.
- N. Razmara and R. Mohammadzadeh, Molecular dynamics study of nitrogen diffusion in nanocrystalline iron, J. Mol. Model., 2017, 23, 8 CrossRef PubMed.
- K. H. Jack, The iron–nitrogen system: the preparation and the crystal structures of nitrogen-austenite (γ) and nitrogen-martensite (α′), Proc. R. Soc. London, Ser. A, 1951, 208, 200–215 CAS.
- C. T. Cheung and G. Simkovich, Self-diffusivity of nitrogen in γ′-iron nitride (“Fe4N”), React. Solids, 1989, 7, 115–129 CrossRef CAS.
- H. Du and J. Ägren, Gaseous Nitriding Iron—Evaluation of Diffusion Data of N ın γ′ and ε Phases, Int. J. Mater. Res., 1995, 86, 522–529 CrossRef CAS.
- O. Kurata, N. Iki, T. Inoue, T. Matsunuma, T. Tsujimura, H. Furutani, M. Kawano, K. Arai, E. C. Okafor, A. Hayakawa and H. Kobayashi, Development of a wide range-operable, rich-lean low-NOx combustor for NH3 fuel gas-turbine power generation, Proc. Combust. Inst., 2019, 37, 4587–4595 CrossRef CAS.
- G. Ceder, S. P. Ong and Y. Wang, Predictive modeling and design rules for solid electrolytes, MRS Bull., 2018, 43, 746–751 CrossRef CAS.
- D. G. Sangiovanni, O. Hellman, B. Alling and I. A. Abrikosov, Efficient and accurate determination of lattice-vacancy diffusion coefficients via non equilibrium ab initio molecular dynamics, Phys. Rev. B, 2016, 93, 094305 CrossRef.
- L. Agosta, E. G. Brandt and A. P. Lyubartsev, Diffusion and reaction pathways of water near fully hydrated TiO2 surfaces from ab initio molecular dynamics, J. Chem. Phys., 2017, 147, 024704 CrossRef PubMed.
- M. Sprik and G. Ciccotti, Free energy from constrained molecular dynamics, J. Chem. Phys., 1998, 109, 7737–7744 CrossRef CAS.
- E. A. Carter, G. Ciccotti, J. T. Hynes and R. Kapral, Constrained reaction coordinate dynamics for the simulation of rare events, Chem. Phys. Lett., 1989, 156, 472–477 CrossRef CAS.
- A. Barducci, M. Bonomi and M. Parrinello, Metadynamics, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2011, 1, 826–843 CAS.
- K. M. Bal and E. C. Neyts, Merging Metadynamics into Hyperdynamics: Accelerated Molecular Simulations Reaching Time Scales from Microseconds to Seconds, J. Chem. Theory Comput., 2015, 11, 4545–4554 CrossRef CAS PubMed.
- Z. Shi, A. D. Lele, A. W. Jasper, S. J. Klippenstein and Y. Ju, Quasi-Classical Trajectory Calculation of Rate Constants Using an Ab Initio Trained Machine Learning Model (aML-MD) with Multifidelity Data, J. Phys. Chem. A, 2024, 128, 3449–3457 CrossRef CAS PubMed.
- L. Zhang, H. Wang, R. Car and W. E, Phase Diagram of a Deep Potential Water Model, Phys. Rev. Lett., 2021, 126, 236001 CrossRef CAS PubMed.
- T. Shi, W. Liu, C. Zhang, S. Lyu, Z. Sun, Q. Peng, Y. Li, F. Meng, C. Tang and C. Lu, An investigation of self-interstitial diffusion in α-zirconium by an on-the-fly machine learning force field, AIP Adv., 2024, 14, 055010 CrossRef CAS.
- Y. Wang, L. Zhang, B. Xu, X. Wang and H. Wang, A generalizable machine learning potential of Ag–Au nanoalloys and its application to surface reconstruction, segregation and diffusion, Modell. Simul. Mater. Sci. Eng., 2022, 30, 025003 CrossRef CAS.
- L. L. Schaaf, E. Fako, S. De, A. Schäfer and G. Csányi, Accurate energy barriers for catalytic reaction pathways: an automatic training protocol for machine learning force fields, npj Comput. Mater., 2023, 9, 180 CrossRef.
- D. Yoo, J. Jung, W. Jeong and S. Han, Metadynamics sampling in atomic environment space for collecting training data for machine learning potentials, npj Comput. Mater., 2021, 7, 131 CrossRef CAS.
- H. A. Wriedt, N. A. Gokcen and R. H. Nafziger, The Fe–N (Iron–Nitrogen) system, Bull. Alloy Phase Diagrams, 1987, 8, 355–377 CrossRef CAS.
- Y. Zhang, H. Wang, W. Chen, J. Zeng, L. Zhang, H. Wang and W. E, DP-GEN: A concurrent learning platform for the generation of reliable deep learning based potential energy models, Comput. Phys. Commun., 2020, 253, 107206 CrossRef CAS.
- D. Hedman, B. McLean, C. Bichara, S. Maruyama, J. A. Larsson and F. Ding, Dynamics of growing carbon nanotube interfaces probed by machine learning-enabled molecular simulations, Nat. Commun., 2024, 15, 4076 CrossRef CAS PubMed.
- B.-J. Lee, T.-H. Lee and S.-J. Kim, A modified embedded-atom method interatomic potential for the Fe–N system: A comparative study with the Fe–C system, Acta Mater., 2006, 54, 4597–4607 CrossRef CAS.
- L. Bonati, D. Polino, C. Pizzolitto, P. Biasi, R. Eckert, S. Reitmeier, R. Schlögl and M. Parrinello, The role of dynamics in heterogeneous catalysis: Surface diffusivity and N2 decomposition on Fe(111), Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2313023120 CrossRef CAS PubMed.
- S. Perego, L. Bonati, S. Tripathi and M. Parrinello, How Dynamics Changes Ammonia Cracking on Iron Surfaces, ACS Catal., 2024, 14, 14652–14664 CrossRef CAS.
- M. Purcel, S. Berendts, L. Bonati, S. Perego, A. Müller, M. Lerch, M. Parrinello and M. Muhler, Iron Nitride Formation and Decomposition during Ammonia Decomposition over a Wustite-Based Bulk Iron Catalyst, ACS Catal., 2024, 14, 13947–13957 CrossRef CAS.
- G. Kresse and J. Hafner, Ab initio molecular dynamics for liquid metals, Phys. Rev. B:Condens. Matter Mater. Phys., 1993, 47, 558–561 CrossRef CAS PubMed.
- G. Kresse and D. Joubert, From ultrasoft pseudopotentials to the projector augmented-wave method, Phys. Rev. B:Condens. Matter Mater. Phys., 1999, 59, 1758–1775 CrossRef CAS.
- G. Kresse and J. Furthmüller, Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set, Comput. Mater. Sci., 1996, 6, 15–50 CrossRef CAS.
- G. Kresse and J. Furthmüller, Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set, Phys. Rev. B:Condens. Matter Mater. Phys., 1996, 54, 11169–11186 CrossRef CAS PubMed.
- J. P. Perdew, K. Burke and M. Ernzerhof, Generalized Gradient Approximation Made Simple, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed.
- P. E. Blöchl, Projector augmented-wave method, Phys. Rev. B:Condens. Matter Mater. Phys., 1994, 50, 17953–17979 Search PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- ASM International Handbook Committee, ASM Handbook, Properties and Selection: Irons, Steels, and High-Performance Alloys, ASM International, Materials Park, OH, 10th edn, 1990, vol. 1 Search PubMed.
- D. E. Jiang and E. A. Carter, Carbon dissolution and diffusion in ferrite and austenite from first principles, Phys. Rev. B:Condens. Matter Mater. Phys., 2003, 67, 214103 CrossRef.
- K. Hirata, S. Iikubo, M. Koyama, K. Tsuzaki and H. Ohtani, First-Principles Study on Hydrogen Diffusivity in BCC, FCC, and HCP Iron, Metall. Mater. Trans. A, 2018, 49, 5015–5022 CrossRef CAS.
- A. Jain, S. P. Ong, G. Hautier, W. Chen, W. D. Richards, S. Dacek, S. Cholia, D. Gunter, D. Skinner, G. Ceder and K. A. Persson, Commentary: The Materials Project: A materials genome approach to accelerating materials innovation, APL Mater., 2013, 1, 011002 CrossRef.
- H. A. Wriedt, N. A. Gokcen and R. H. Nafziger, The Fe–N (Iron–Nitrogen) system, Bull. Alloy Phase Diagrams, 1987, 8, 355–377 CrossRef CAS.
- T. Takahashi and W. A. Bassett, High-Pressure Polymorph of Iron, Science, 1964, 145, 483–486 CrossRef CAS PubMed.
- J. D. Morrow, J. L. A. Gardner and V. L. Deringer, How to validate machine-learned interatomic potentials, J. Chem. Phys., 2023, 158, 121501 Search PubMed.
- G. Henkelman and H. Jónsson, Improved tangent estimate in the nudged elastic band method for finding minimum energy paths and saddle points, J. Chem. Phys., 2000, 113, 9978–9985 Search PubMed.
- G. Henkelman, B. P. Uberuaga and H. Jónsson, A climbing image nudged elastic band method for finding saddle points and minimum energy paths, J. Chem. Phys., 2000, 113, 9901–9904 CrossRef CAS.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. In’T Veld, A. Kohlmeyer, S. G. Moore, T. D. Nguyen, R. Shan, M. J. Stevens, J. Tranchida, C. Trott and S. J. Plimpton, LAMMPS – a flexible simulation tool for particle-based materials modeling at the atomic, meso, and continuum scales, Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- J. Zeng, D. Zhang, D. Lu, P. Mo, Z. Li, Y. Chen, M. Rynik, L. Huang, Z. Li, S. Shi, Y. Wang, H. Ye, P. Tuo, J. Yang, Y. Ding, Y. Li, D. Tisi, Q. Zeng, H. Bao, Y. Xia, J. Huang, K. Muraoka, Y. Wang, J. Chang, F. Yuan, S. L. Bore, C. Cai, Y. Lin, B. Wang, J. Xu, J.-X. Zhu, C. Luo, Y. Zhang, R. E. A. Goodall, W. Liang, A. K. Singh, S. Yao, J. Zhang, R. Wentzcovitch, J. Han, J. Liu, W. Jia, D. M. York, W. E, R. Car, L. Zhang and H. Wang, DeePMD-kit v2: A software package for Deep Potential models, J. Chem. Phys., 2023, 159, 054801 Search PubMed.
- L. Zhang, J. Han, H. Wang, R. Car and W. E, Deep Potential Molecular Dynamics: A Scalable Model with the Accuracy of Quantum Mechanics, Phys. Rev. Lett., 2018, 120, 143001 CrossRef CAS PubMed.
- H. Wang, L. Zhang, J. Han and W. E, DeePMD-kit: A deep learning package for many-body potential energy representation and molecular dynamics, Comput. Phys. Commun., 2018, 228, 178–184 CrossRef CAS.
- J. Behler and M. Parrinello, Generalized Neural-Network Representation of High-Dimensional Potential-Energy Surfaces, Phys. Rev. Lett., 2007, 98, 146401 Search PubMed.
- D. P. Kingma and J. Ba, Adam: A Method for Stochastic Optimization, arXiv, 2017, preprint, arXiv:1412.6980, https://arxiv.org/abs/1412.6980 Search PubMed.
- DeePMD-kit's documentation—DeePMD-kit documentation, https://docs.deepmodeling.com/projects/deepmd/en/r2/index.html, (accessed December 11, 2024).
- C. Middendorf and W. Mader, Growth and microstructure of iron nitride layers and pore formation in ε-Fe3N, Int. J. Mater. Res., 2003, 94, 333–340 Search PubMed.
- W. Sutherland, LXXV. A dynamical theory of diffusion for non-electrolytes and the molecular mass of albumin, London, Edinburgh Dublin Philos. Mag. J. Sci., 1905, 9, 781–785 CrossRef CAS.
- A. Einstein, Über die von der molekularkinetischen Theorie der Wärme geforderte Bewegung von in ruhenden Flüssigkeiten suspendierten Teilchen, Ann. Phys., 1905, 322, 549–560 CrossRef.
- M. S. Green, Markoff Random Processes and the Statistical Mechanics of Time-Dependent Phenomena. II. Irreversible Processes in Fluids, J. Chem. Phys., 1954, 22, 398–413 CrossRef CAS.
- R. Kubo, Statistical-Mechanical Theory of Irreversible Processes. I. General Theory and Simple Applications to Magnetic and Conduction Problems, J. Phys. Soc. Jpn., 1957, 12, 570–586 CrossRef.
- D. J. Evans, W. G. Hoover, B. H. Failor, B. Moran and A. J. C. Ladd, Nonequilibrium molecular dynamics via Gauss's principle of least constraint, Phys. Rev. A:At., Mol., Opt. Phys., 1983, 28, 1016–1021 CrossRef CAS.
- G. S. Heffelfinger and D. M. Ford, Massively parallel dual control volume grand canonical molecular dynamics with LADERA I. Gradient driven diffusion in Lennard-Jones fluids, Mol. Phys., 1998, 94, 659–671 CrossRef CAS.
- J. M. D. MacElroy, Nonequilibrium molecular dynamics simulation of diffusion and flow in thin microporous membranes, J. Chem. Phys., 1994, 101, 5274–5280 CrossRef CAS.
- P. C. Aeberhard, S. R. Williams, D. J. Evans, K. Refson and W. I. F. David, Ab initio Nonequilibrium Molecular Dynamics in the Solid Superionic Conductor LiBH4, Phys. Rev. Lett., 2012, 108, 095901 CrossRef PubMed.
- W. G. Hoover, Canonical dynamics: Equilibrium phase-space distributions, Phys. Rev. A:At., Mol., Opt. Phys., 1985, 31, 1695–1697 CrossRef PubMed.
- S. Nosé, A unified formulation of the constant temperature molecular dynamics methods, J. Chem. Phys., 1984, 81, 511–519 CrossRef.
- K. Frisk, A thermodynamic evaluation of the Cr–N, Fe–N, Mo–N and Cr–Mo–N systems, CALPHAD:Comput. Coupling Phase Diagrams Thermochem., 1991, 15, 79–106 CrossRef CAS.
- K. H. Jack, The iron–nitrogen system: the crystal structures of ε-phase iron nitrides, Acta Crystallogr., 1952, 5, 404–411 CrossRef.
- J.-O. Andersson and J. Ågren, Models for numerical treatment of multicomponent diffusion in simple phases, J. Appl. Phys., 1992, 72, 1350–1355 CrossRef CAS.
- H. Du, A reevaluation of the Fe–N and Fe–C–N systems, J. Phase Equilib., 1993, 14, 682–693 CrossRef CAS.
- Y. Liu, X. He and Y. Mo, Discrepancies and error evaluation metrics for machine learning interatomic potentials, npj Comput. Mater., 2023, 9, 174 CrossRef.
- J. Zeng, L. Cao, M. Xu, T. Zhu and J. Z. H. Zhang, Complex reaction processes in combustion unraveled by neural network-based molecular dynamics simulation, Nat. Commun., 2020, 11, 5713 CrossRef CAS PubMed.
- X.-Y. Zhou, H.-H. Wu, J. Zhang, S. Ye, T. Lookman and X. Mao, Unveiling the mechanism of the carbon ordering and martensite tetragonality in Fe–C alloys studied via deep-potential molecular dynamics simulations, J. Mater. Sci. Technol., 2024, S1005030224010223 Search PubMed.
- K. Momma and F. Izumi, VESTA 3 for three-dimensional visualization of crystal, volumetric and morphology data, J. Appl. Crystallogr., 2011, 44, 1272–1276 CrossRef CAS.
- S. Malgope, M. K. Gupta, S. Bag, R. Mittal, S. Bhattacharya, A. Singh and S. L. Chaplot, Untangling high-temperature thermal expansion and lattice thermal conductivity behavior of vanadium using machine-learned molecular dynamics, Phys. Rev. B, 2024, 110, 054301 CrossRef CAS.
- Student, The Probable Error of a Mean, Biometrika, 1908, 6, 1 CrossRef.
- A. T. Celebi, S. H. Jamali, A. Bardow, T. J. H. Vlugt and O. A. Moultos, Finite-size effects of diffusion coefficients computed from molecular dynamics: a review of what we have learned so far, Mol. Simul., 2021, 47, 831–845 CrossRef CAS.
- S. R. Hosseini, F. Ashrafizadeh and A. Kermanpur, Calculation And Experimentation Of The Compound Layer Thickness In Gas And Plasma Nitriding Of Iron Search PubMed.
- Y. M. Lakhtin and Y. D. Kogan, Nitriding of steel, Met. Sci. Heat Treat., 1978, 20, 250–251 CrossRef.
- B. Prenosil, Kinetics of diffusion of nitrogen in epsilon- nitride, Kovove Mater., 1965, 3, 69–87 CAS.
- E. Levchenko, A. Evteev, I. Belova and G. Murch, Molecular dynamics simulation and theoretical analysis of carbon diffusion in cementite, Acta Mater., 2009, 57, 846–853 CrossRef CAS.
| This journal is © the Owner Societies 2026 |