Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A desolvation-based molecular crowding mechanism revealed through α,β,γ,δ-tetrakis(1-methylpyridinium-4-yl)porphyrin p-toluenesulfonate–Zn complexation in alcohols
Akihisa
Miyagawa
 *a,
Chisa
Ito
b,
Shigenori
Nagatomo
b and
Kiyoharu
Nakatani
b
*a,
Chisa
Ito
b,
Shigenori
Nagatomo
b and
Kiyoharu
Nakatani
b
aDepartment of Chemistry, Graduate School of Advanced Science and Engineering, Hiroshima University, 1-3-1 Kagamiyama, Higashi-Hiroshima, 739-8526, Japan. E-mail: miyagawaaki@hiroshima-u.ac.jp
bDepartment of Chemistry, Faculty of Pure and Applied Sciences, University of Tsukuba, Tsukuba, Ibaraki 305-8571, Japan
First published on 17th February 2026
Abstract
Molecular crowding can substantially alter chemical reactivity, yet its mechanistic influence in organic solvents remains largely unexplored. Here, we quantitatively and mechanistically extend this concept to non-aqueous media by evaluating the elementary-step kinetics of TMPyP–Zn2+ complexation in methanol (MeOH), ethanol (EtOH), and 1-propanol (PrOH). Rate constants for ion-pair formation, ion-pair dissociation, and metal insertion were extracted from time-resolved absorbance changes and analyzed as a function of polyethylene glycol concentration (CPEG). Our analysis reveals that across all solvents and concentrations, crowding enhances reaction rates primarily through osmotic-pressure-driven desolvation. Notably, ion-pair formation is influenced by both volume exclusion and osmotic pressure effects, for which the excluded-volume contribution becomes negligible (Γ ≈ 1). These findings demonstrate that molecular crowding can effectively accelerate reactions in organic media and provide a unified physico-chemical interpretation of crowding effects beyond aqueous systems, establishing solvent activity engineering as a generalizable framework for controlling reaction kinetics in non-aqueous environments.
Introduction
Understanding and controlling chemical reaction rates is a central objective in chemistry as reaction kinetics determine efficiency, selectivity, and overall feasibility across diverse applications.1,2 Molecular crowding refers to a solvent environment observed in living cells, and the unique reactivity under such conditions has attracted much attention.3–6 The densely packed environment of macromolecules, such as DNAs, RNAs, proteins, and carbohydrates, modified both the kinetics and thermodynamics of chemical reactions compared to those in diluted solutions. In cellular contexts, macromolecular crowding is commonly rationalized by excluded-volume effects—i.e., the reduction of accessible space because two molecules cannot occupy the same location—leading to changes in thermodynamic activity and thereby impacting equilibria and kinetics. At the same time, even widely used synthetic polymers are rarely completely inert; therefore, weak or specific interactions between the crowder and the solute can complicate interpretation unless hard-core and chemical contributions are disentangled experimentally. For example, studies have reported a 48-fold increase in the complexation constant of hydroxyquinol with Zn2+,7 a 4-fold enhancement of the rate constant in the Hantzsch pyridine reaction,8 and accelerated folding of G-quadruplex structures in aqueous media.9 These findings highlight the significant potential of molecular crowding environments to improve reaction efficiency. However, while the effects of molecular crowding on reaction behavior have been extensively investigated in aqueous solutions, little to no attention has been given to such phenomena in organic solvents even though many chemical reactions are conducted in organic media. Despite their central role in synthetic chemistry, organic solvents have remained essentially unexplored as media for molecular crowding. This gap arises because most crowding studies have been motivated by the desire to mimic intracellular environments, which are overwhelmingly aqueous.3–6 Furthermore, crowding in organic solvents is conceptually more complex: solvation energies are much larger, preferential interactions between polymeric crowders and solvents vary widely, and desolvation often dominates activation barriers. These factors have made it difficult to experimentally isolate crowding effects in organic media, leaving a major mechanistic gap in the field.From a theoretical standpoint, early descriptions of crowding originated from solution thermodynamics and depletion/excluded-volume concepts, and were later formalized for biomacromolecules by Minton and coworkers.10 Classic crowding theories often approximate crowders and solutes as hard particles, predicting that crowding effects are athermal, purely entropic, and generally stabilizing because hard-core repulsions reduce the available volume.11 However, accumulating experimental evidence has established that macromolecular crowding is frequently more than hard-core repulsions: crowding can even destabilize proteins and complexes, and crowding effects often involve nonzero enthalpic contributions. These observations require accounting for soft (chemical) interactions and preferential interactions among solute, crowder, and solvent, beyond the hard-sphere picture.12 Modern frameworks emphasize that crowding and cosolute effects can be interpreted through preferential interactions and solution thermodynamics, including approaches based on Kirkwood–Buff theory,13 where changes in hydration and cosolute distributions around solutes provide a quantitative route to connect cosolute addition with stability and association energetics.
In addition to these thermodynamic effects, molecular crowding is also known to alter transport properties such as viscosity, viscoelasticity, and translational–rotational coupling, particularly for anisotropic crowders.14 Recent theoretical and simulation studies have demonstrated non-Gaussian and transiently anomalous diffusion in crowded systems, reflecting complex viscoelastic responses rather than simple hydrodynamic slowing.15 However, the extent to which such transport effects influence reaction-limited chemical kinetics remains largely unexplored, especially in organic solvents.
Primarily, three effects have been proposed to explain molecular crowding: the volume exclusion effect, the osmotic pressure effect, and structural changes.10,16–21 Minton et al. introduced the principle of volume exclusion based on a hard particle model, wherein the restricted volume available to reactants due to crowding agents increases their effective activity.10,22 Additionally, when molecular crowding agents are present at high concentrations, they elevate the osmotic pressure, leading to a decrease in solvent activity. Sugimoto et al. demonstrated that the reactivity of biomolecules, such as DNA, can be interpreted in terms of osmotic pressure effects, which are closely related to solvation and desolvation phenomena.21,23 At high concentrations, crowding agents can also induce conformational changes of macromolecules, including the active sites of proteins, thereby altering chemical reactivity. However, such structural changes are difficult to quantify using standard thermodynamic models. Therefore, simpler systems—such as metal complexation reactions that proceed without significant conformational changes—serve as useful models for studying molecular crowding effects, allowing the contributions of volume exclusion and osmotic pressure to be examined in isolation.7,8,24–27 Metal–ligand complexation reactions offer an ideal platform for dissecting crowding effects because they proceed without conformational changes and involve well-defined solvation/desolvation steps. In particular, the TMPyP–Zn system provides a rare opportunity to kinetically separate ion-pair formation, ion-pair dissociation, and metal insertion, allowing each elementary step to be independently evaluated under crowding.
In our previous study, we showed that PEG200-induced crowding enhances the overall complexation constant of 8-hydroxyquinoline with metal ions (Zn2+, Co2+, and Cd2+) in methanol.28 Although this result demonstrated that molecular crowding can significantly shift equilibrium in organic media, the analysis was limited to thermodynamic parameters. As a consequence, the kinetic origins of crowding—how individual elementary steps respond to crowding, whether volume exclusion or desolvation dominates, and how different solvents contribute—remained completely unresolved. Moreover, despite this initial insight, a comprehensive understanding of molecular crowding in organic solvents is still lacking. In particular, the influence of solvent identity, the reaction type, and the physicochemical nature of crowders has not been systematically examined. These knowledge gaps highlight that molecular crowding may provide a general strategy for tuning reaction kinetics in non-aqueous media, but the fundamental mechanisms governing such effects are far from established. This leads to several key unresolved questions:
(i) How does the nature of the solvent influence crowding-induced acceleration?
(ii) Can elementary reaction steps be individually modulated?
(iii) Is desolvation the dominant driving force in organic solvents?
Addressing these questions requires systems in which multi-step kinetics can be resolved—a challenge that has prevented the establishment of a unified framework for crowding in organic environments.
In this work, we provide the first quantitative and mechanistic demonstration of molecular crowding in organic solvents. By deconvoluting the TMPyP–Zn complexation into three elementary steps (ion-pair formation, ion-pair dissociation, and metal insertion), we show that crowding-driven desolvation—not volume exclusion—is the dominant contributor to rate enhancement in methanol (MeOH), ethanol (EtOH), and 1-propanol (PrOH). Importantly, this conclusion is based on step-resolved kinetic analysis demonstrating that for specific elementary steps the excluded-volume contribution vanishes (Γ = 1) while the rate enhancement persists, thereby directly excluding volume exclusion as the sole origin of the observed acceleration. This establishes “solvent activity engineering” as a new principle for controlling reaction kinetics in non-aqueous media. We evaluated the effects of molecular crowding on the complexation rate constants of a porphyrin derivative (TMPyP) with Zn2+ in MeOH, EtOH, and PrOH. PEG200 was employed as the molecular crowding agent. The complexation rate constants were determined from absorbance changes in each solvent at various concentrations of PEG200 (CPEG). These rate constants were then analyzed in the context of volume exclusion and osmotic pressure effects, allowing a quantitative separation of excluded-volume and desolvation contributions to each elementary step.
Experimental
PEG200, MeOH, EtOH, PrOH, and zinc nitrate, were purchased from Kanto Chemical Co. Inc. (Tokyo, Japan). α,β,γ,δ-Tetrakis(1-methylpyridinium-4-yl)porphyrin p-toluenesulfonate (TMPyP) was purchased as a porphyrin derivative from Tokyo Chemical Industry Co., Ltd (Tokyo, Japan). All chemicals were used as received without further purification.The molar absorption coefficient of TMPyP–Zn complex (ε) was determined in MeOH, EtOH, and PrOH solutions containing PEG200 in order to evaluate the complex concentration. Mixed solutions containing Zn2+ (0.6 mM) and various concentrations of TMPyP (1.48, 2.96, 4.44, and 5.92 µM for MeOH and EtOH, and 293, 587, 880, and 1170 nM for PrOH) were prepared and allowed to stand overnight to reach complexation equilibrium. The absorption spectra of the TMPyP–Zn complex were recorded using absorption spectrometer (V-730, JASCO). The absorption maxima of the complex in MeOH, EtOH, and PrOH were 438, 440, and 443 nm, respectively. CPEG used in the present study were as follows: 0, 10, 20, 30, 40, and 50 vol% for MeOH, 0, 2.5, 5.0, 7.5, 10, and 20 vol% for EtOH, and 0, 2.5, 5.0, 7.5, 10, and 15 vol% for PrOH. The ε values of the TMPyP–Zn2+ complex in the organic solvents at various CPEG values are summarized in Table S1.
The rate constant of the complexation of TMPyP with Zn2+ was evaluated based on the time-dependent absorbance change of the TMPyP–Zn2+ complex. A 3 µL aliquot of Zn2+ solution (1 mM) was added to 3 mL of TMPyP solution (5.92 µM), resulting in a final Zn2+ concentration of 1 µM. The absorbance change of the TMPyP–Zn2+ complex was monitored over time under continuous stirring at room temperature (25 °C).
Molecular orbital calculations for TMPyP, TMPyP–Zn ionic pair, and PEG200 were performed using Gaussian 16. The molecular structures were optimized using density functional theory (DFT) at the B3LYP/def2-SVP level including solvent effects (SMD, methanol) under loose convergence criteria. For Zn2+, a tetra-coordinated [Zn(MeOH)4]2+ solvate was used as the initial model. For TMPyP, four p-toluenesulfonate (TsO−) counterions were included to reproduce the realistic ionic environment. The polymerization degree of PEG200 was assumed to be 4. The full optimized structure − including four TsO− counterions and solvated methanol − and the simplified representation in which TsO− and methanol molecules were omitted for clarity are shown in Fig. S1.
Macromolecular crowding is known to increase the viscosity of the medium, which can alter diffusion coefficients and thereby influence reaction kinetics, particularly in diffusion-limited systems.14,15 All kinetic measurements were performed under continuous stirring, and preliminary tests confirmed that the observed kinetics were reaction-limited rather than diffusion-limited under the present conditions. Therefore, viscosity-related transport effects were not expected to dominate the measured rate constants.
Results and discussion
Complexation rate constants of TMPyP with Zn2+ in organic solvents at various CPEG
Fig. S2 shows the normalized absorption spectra of free TMPyP and the TMPyP–Zn complex in MeOH, EtOH, and PrOH. The absorption peaks at approximately 420 and 500–650 nm in the spectra of free TMPyP were attributed to the Soret and Q bands, respectively, corresponding to π–π* transitions.29 Upon complexation with Zn2+, both bands exhibited red shifts. The maximum absorption wavelengths of the Soret band for the TMPyP–Zn complex in MeOH, EtOH, and PrOH were 438, 440, and 443 nm, respectively. To minimize the influence of unreacted TMPyP, absorbance values at 450, 455, and 450 nm were used for the kinetic analysis in MeOH, EtOH, and PrOH, respectively.
Fig. 1 shows a representative time-dependent concentration profile of the TMPyP–Zn complex in MeOH. At CPEG = 0 vol%, the concentration of the TMPyP complex approached equilibrium at t = 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 s. As CPEG increased, the time required to reach the equilibrium decreased: t = 3000 s at CPEG = 10 vol%, t = 1500 s at CPEG = 20 vol%, t = 1000 s at CPEG = 30 vol%, t = 1000 s at CPEG = 40 vol%, and t = 500 s at CPEG = 50 vol%. These results indicate that the complexation rate of TMPyP with Zn2+ was accelerated by the addition of PEG200.
000 s. As CPEG increased, the time required to reach the equilibrium decreased: t = 3000 s at CPEG = 10 vol%, t = 1500 s at CPEG = 20 vol%, t = 1000 s at CPEG = 30 vol%, t = 1000 s at CPEG = 40 vol%, and t = 500 s at CPEG = 50 vol%. These results indicate that the complexation rate of TMPyP with Zn2+ was accelerated by the addition of PEG200.
Fig. S3 shows the time-dependent concentrations of the TMPyP–Zn2+ complex in EtOH and PrOH solutions. At CPEG = 0 vol%, the complexation was completed at t = 300 in EtOH and 600 s in PrOH, indicating that the reaction rate is influenced by the choice of solvent. In both EtOH and PrOH, the complexation of TMPyP was further accelerated at CPEG increased. These results obtained in MeOH, EtOH, and PrOH suggest that complexation reactions in organic solvents can be significantly accelerated by introducing a molecular crowding environment.
The complexation mechanism of TMPyP with Zn2+ has been previously reported by Toda et al.30 and is described by the following reaction scheme:
| H2P + Zn ⇄ H2P·Zn | (1) |
| H2P·Zn → 2H+ + ZnP | (2) |
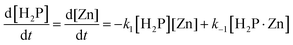 | (3) |
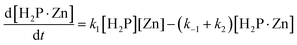 | (4) |
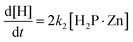 | (5) |
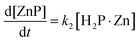 | (6) |
Eqn (3) and (4) allow us to evaluate the reaction behavior using k1, k−1, and k2 as fitting parameters, in which the Runge–Kutta method is employed in combination with the least-squares method. Fig. S4 shows the representative curve-fitted results of Zn complexation with TMPyP in MeOH at CPEG = 0 vol%. The experimental behavior was well explained based on eqn (3)–(6) using k1 = 4.72 × 10−3 M−1 s−1, k−1 = 0.456 s−1, and k2 = 1.0 × 10−4 s−1. Similar analyses were performed under other conditions, and the values of k1, k−1, and k2 were determined.
Table S2 summarizes the obtained k1, k−1, and k2 values for each solvent at various CPEG. Furthermore, Fig. 2 shows the relationship between CPEG and the log k values in MeOH, EtOH, and PrOH. The k1 values for MeOH and PrOH increased until CPEG = 20 and 7.5 vol%, respectively, and then decreased with CPEG. However, the k1 values at higher CPEG are larger than that for CPEG = 0 vol% for all solvents: k1 = (3.05 ± 1.44) × 10−3 M−1 s−1 at CPEG = 0 vol% and k1 = (4.06 ± 0.40) × 10−2 M−1 s−1 at CPEG = 50 vol% for MeOH, k1 = 0.204 ± 0.082 M−1 s−1 at CPEG = 0 vol% and k1 = 0.366 ± 0.113 M−1 s−1 at CPEG = 30 vol% for EtOH, and k1 = 0.116 ± 0.138 M−1 s−1 at CPEG = 0 vol% and k1 = 0.181 ± 0.020 M−1 s−1 at CPEG = 15 vol% for PrOH. In contrast, the k−1 and k2 values increased as CPEG increased. These behaviors will be explained later in terms of the molecular crowding effect.
The addition of PEG200 to the MeOH, EtOH, and PrOH may alter the solvent polarity, potentially affecting the observed k values. To evaluate the change in solvent polarity, we used Reichardt's dye, a well-known solvatochromic reagent.30,31 The Et value, calculated from the absorption maximum of the charge-transfer band in each solvent, was used to estimate the solvent polarity. Fig. S5 shows the absorption spectra of Reichardt's dye in MeOH, EtOH, PrOH, and PEG200. The maximum absorption wavelengths were observed at 515.5, 553.5, 568, and 550.5 nm for MeOH, EtOH, PrOH, and PEG200, respectively. The Et values were determined to be 55.5, 51.7, 50.3, and 51.9 kcal mol−1 for MeOH, EtOH, PrOH, and PEG200, respectively, based on Et = 2.859 × 10−3ν, where ν is the wavenumber of absorption maximum. The obtained Et values for MeOH, EtOH, and PrOH were consistent with previous reported reference values.32 Although the Et values of MeOH, EtOH, and PrOH were all close to that of PEG200, the deviation for MeOH was larger than those for EtOH and PrOH. This suggests that MeOH may be more sensitive to polarity changes induced by the addition of PEG200 compared to the other two solvents. Nevertheless, since the polarity parameters remain relatively similar overall, the increase in the k values shown in Fig. 2 is primarily attributed to the molecular crowding effect.
Molecular crowding effect on TMPyP complexation in organic solvents
As discussed above, the increase in the k values upon addition of PEG200 is primarily attributed to the molecular crowding effect. To further investigate this effect, we analyzed the influence of molecular crowding on the k values based on volume exclusion and osmotic pressure effects. Since TMPyP is a rigid molecule, it is reasonable to consider only these two factors in the present study.In the reaction scheme of eqn (1) and (2), the complexation rate constant is represented by k1, k−1, and k2. Under molecular crowding conditions, the molar free Gibbs energy change is shifted by −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) ln
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Γ. According to classical macromolecular crowding theory based on activity coefficients and transition-state arguments, the k value in the molecular crowding environment (kMC) is expressed as33
Γ. According to classical macromolecular crowding theory based on activity coefficients and transition-state arguments, the k value in the molecular crowding environment (kMC) is expressed as33
| kMC = Γkd | (7) |
Minton et al. proposed a theoretical estimation of the γ value in molecular crowding environments based on a hard particle model.10,22 This formulation is derived from scaled-particle theory applied to excluded-volume systems. This model suggests that the γ values of reactants, products, and crowding agents are determined by their respective molecular sizes. The activity coefficient of reactant i in molecular crowding is described by,
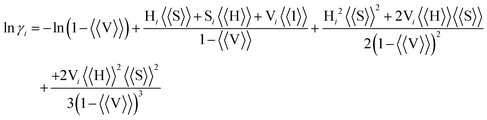 | (8) |
Fig. S6 shows the relationship between CPEG and ln γH2P or ln γH2P·Zn. The γ values for TMPyP and ion pair of TMPyP increased with increasing CPEG, indicating that the effective concentration of these molecules increases due to the volume exclusion by PEG200. At higher CPEG, the difference in the γ value becomes larger, indicating that the Γ1 becomes smaller as increasing CPEG. Therefore, as seen from eqn (7), the ion pair formation of TMPyP is suppressed by the volume exclusion effect.
The addition of a macromolecule such as PEG200 at high concentrations increases the osmotic pressure of the solvent, resulting in a decrease in solvent activity. This implies that reactions involving solvation and desolvation are influenced under such conditions.21,23 Consequently, the molar Gibbs free energy is altered by a term of −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) ln
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) γsol.34 This treatment follows the osmotic stress framework commonly applied to solvation thermodynamics. Incorporating the osmotic pressure effect into eqn (7) yields the following expression:
γsol.34 This treatment follows the osmotic stress framework commonly applied to solvation thermodynamics. Incorporating the osmotic pressure effect into eqn (7) yields the following expression:
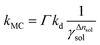 | (9) |
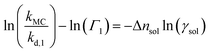 | (10) |
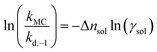 | (11) |
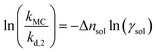 | (12) |
Eqn (10)–(12) are directly derived from eqn (9) and are used in this work to quantitatively extract Δnsol from experimental data. This equation indicates a linear relationship between the left hand side of the formula and ln(γsol), with a slope corresponding to Δnsol.
We attempted to quantify the extent of solvent activity in MeOH, EtOH, and PrOH by measuring the freezing point depression using differential scanning calorimetry. However, accurate determination proved challenging due to the intrinsically low freezing points and high volatilities of these solvents. Therefore, the γsol values were instead estimated using the van’t Hoff equation for osmotic pressure, expressed as35
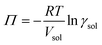 | (13) |
| γsol = exp(−CVsol) | (14) |
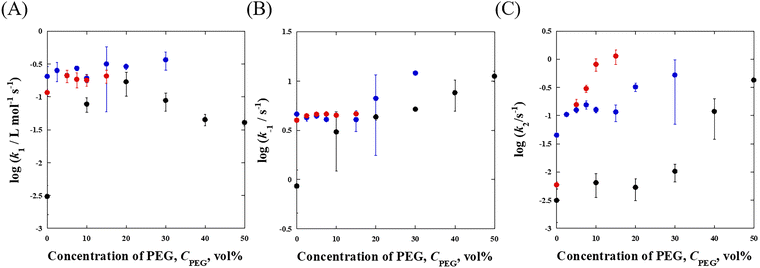
Fig. 3 shows the relationship between ln γsol and ln (kMC/kd) in MeOH, EtOH, and PrOH solutions. As ln γsol decreased, ln (kMC/kd) increased for all rate constants (k1, k−1, and k2) in all solvents. Especially, the k−1 and k2 are influenced only by the osmotic pressure based on eqn (11) and (12), indicating that the ion pair dissociation and TMPyP complexation are promoted by the osmotic pressure effect. Because the activity–coefficient ratio Γ reduces to unity for k−1 and k2, excluded-volume effects do not contribute to these steps. Therefore, the observed increase in k−1 and k2 cannot be explained by volume exclusion and must originate from osmotic-pressure-induced desolvation.
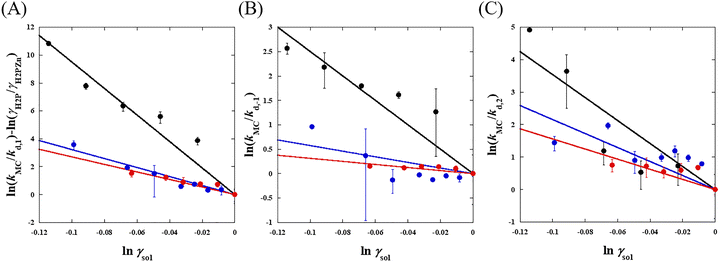 | ||
Fig. 3 Relationship between log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) γsol and log(kMC/kd) in MeOH (black), EtOH (blue), and PrOH (red) solutions. (A) ion pair formation, (B) ion pair dissociation, and (C) TMPyP complexation. Here, γsol represents the solvent activity coefficient estimated from osmotic pressure, and kMC/kd denotes the ratio of rate constants in crowded and dilute conditions. Solid lines correspond to linear least-squares fits based on eqn (10)–(12), from which the solvation number change (Δnsol) was determined. The linear dependence observed for all steps, particularly for k−1 and k2 where Γ = 1, demonstrates that the rate enhancement originates from osmotic-pressure-induced desolvation rather than excluded-volume effects. γsol and log(kMC/kd) in MeOH (black), EtOH (blue), and PrOH (red) solutions. (A) ion pair formation, (B) ion pair dissociation, and (C) TMPyP complexation. Here, γsol represents the solvent activity coefficient estimated from osmotic pressure, and kMC/kd denotes the ratio of rate constants in crowded and dilute conditions. Solid lines correspond to linear least-squares fits based on eqn (10)–(12), from which the solvation number change (Δnsol) was determined. The linear dependence observed for all steps, particularly for k−1 and k2 where Γ = 1, demonstrates that the rate enhancement originates from osmotic-pressure-induced desolvation rather than excluded-volume effects. | ||
The solid lines in Fig. 3 represent the regression line using the linear-squares method. We determined the Δnsol values from these slopes: Δnsol = 95.1 ± 6.1, 32.3 ± 1.9, and 27.0 ± 2.7 for k1 in MeOH, EtOH, and PrOH, respectively; Δnsol = 25.0 ± 2.4, 5.75 ± 0.48, and 3.14 ± 0.02 for k−1 in MeOH, EtOH, and PrOH, respectively; Δnsol = 35.4 ± 5.0, 21.6 ± 4.0, and 15.6 ± 0.4 for k2 in MeOH, EtOH, and PrOH, respectively. These Δnsol values are summarized in Table 1. All Δnsol values are positive, which indicates that the addition of PEG promotes desolvation in the reaction system. The Δnsol values for EtOH and PrOH were similar for all k, whereas MeOH consistently exhibited significantly larger Δnsol. This behavior can be rationalized in terms of solvent polarity. The empirical polarity parameter (Et) is markedly higher for MeOH (55.5 kcal mol−1), while EtOH and PrOH show comparable Et values (51.7 and 50.3 kcal mol−1). Therefore, the addition of PEG is expected to induce a larger change in the polarity of MeOH than in the other alcohols, thereby promoting more pronounced desolvation only in the MeOH system. Because the desolvation accompanying the forward reaction (k1 and k2) is substantially larger than that associated with the backward reaction (k−1), the overall reaction is promoted regardless of the type of organic solvent used. Therefore, we have demonstrated that molecular crowding in organic solvents promotes the rate of metal complexation reactions.
| Solvent | k 1 | k −1 | k 2 |
|---|---|---|---|
| Parentheses represent the fitting errors. | |||
| MeOH | 95.1 (6.1) | 25.0 (2.4) | 35.4 (5.0) |
| EtOH | 32.3 (1.9) | 5.75 (0.48) | 21.6 (4.0) |
| PrOH | 27.0 (2.7) | 3.14 (0.02) | 15.6 (0.4) |
In addition to thermodynamic contributions such as excluded-volume and solvent-activity effects, macromolecular crowding is also known to modify transport properties, including viscosity, viscoelasticity, and translational–rotational coupling, particularly in systems containing anisotropic crowders.14 Increased viscosity can alter diffusion coefficients and potentially influence reaction kinetics in diffusion-limited systems. Recent theoretical and simulation studies have further demonstrated that crowded environments may exhibit non-Gaussian and transiently anomalous diffusion due to complex viscoelastic responses rather than simple hydrodynamic slowing.
However, in the present system, all kinetic measurements were performed under continuous stirring, and the reaction was verified to proceed under reaction-limited rather than diffusion-limited conditions. Moreover, the linear dependence of ln(kMC/kd) on ln(γsol), together with the disappearance of the excluded-volume contribution (Γ = 1) for k−1 and k2 while rate enhancement persists, indicates that the dominant factor governing the observed acceleration is the change in activation free energy associated with desolvation. Therefore, although transport-related effects may become significant under highly crowded or diffusion-controlled conditions, they are unlikely to account for the systematic kinetic behavior observed in this study.
Conclusions
We have uncovered a unifying physical principle that governs molecular crowding in organic solvents: desolvation-driven acceleration. By resolving the TMPyP–Zn complexation into its three elementary steps, we provide a quantitative and step-resolved mechanistic demonstration that osmotic-pressure-induced desolvation—not volume exclusion—dominates the kinetics of ion pairing, its dissociation, and metal insertion across MeOH, EtOH, and PrOH. The disappearance of excluded-volume contributions (Γ = 1) in specific steps and the linear scaling between ln(kMC/kd) and ln(γsol) directly establish solvent activity as the key determinant of reaction rates under crowded conditions.The implications of this finding extend beyond the present system, as many synthetic transformations—including metal-mediated catalysis and ligand substitution—rely on desolvation as a fundamental activation process. Our results therefore introduce solvent activity engineering as a mechanistically grounded and potentially generalizable strategy for controlling reaction kinetics in non-aqueous media, thereby expanding the physico-chemical framework of molecular crowding beyond aqueous systems. The observed plateau in rate enhancement at high PEG concentrations highlights the role of crowder–solvent interactions and defines practical boundaries for crowding-based rate control. Further studies across diverse crowders, solvents, and reaction types will refine this framework and test its broader applicability.
Although the present study focuses on organic solvents, the underlying principle—that solvent activity governs desolvation barriers—may have broader implications beyond synthetic systems. In biological environments, macromolecular crowding and osmotic stress are known to influence protein association, ion pairing, metal insertion processes, and nucleic acid folding. These processes similarly involve changes in solvation and desolvation free energies. Therefore, our findings provide a conceptual bridge between crowding phenomena in synthetic organic media and those operating in cellular systems, highlighting solvent activity as a potentially universal regulator of reaction kinetics and reinforcing the broader physico-chemical significance of the present study.
Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting the findings of this study are available from the corresponding author upon reasonable request.The supplementary information includes the molar absorption coefficient of TMPyP, DFT-optimized molecular structures, absorption spectra of TMPyP and its complexes, time-dependent concentration profiles of the complexes, detailed kinetic analyses and numerical rate constants, solvent polarity changes evaluated using Reichardt's dye, and numerical γsol data supporting the discussion. See DOI: https://doi.org/10.1039/d5cp05054e.
Acknowledgements
This work was supported by a grand-in-aid from Toyota Physical and Chemical Research Institute (A. M.).References
- J. K. Norskov, T. Bligaard and J. Kleis, Science, 2009, 324, 1655–1656 CrossRef CAS PubMed.
- X. Sanchez-Vila, M. Dentz and L. D. Donado, Geophys. Res. Lett., 2007, 34, L10404 CrossRef.
- N. A. Chebotareva, B. I. Korganov and N. B. Livanova, BioChemistry, 2004, 69, 1522–1536 Search PubMed.
- J. A. Dix and A. S. Verkman, Annu. Rev. Biophys., 2008, 37, 247–263 CrossRef CAS PubMed.
- D. A. White, A. K. Buell, T. P. J. Knowles, M. E. Welland and C. M. Dobson, J. Am. Chem. Soc., 2010, 132, 5170–5175 CrossRef CAS PubMed.
- N. Ostrowska, M. Feig and J. Trylska, Front. Mol. Biosci., 2019, 6, 86 CrossRef CAS PubMed.
- A. Miyagawa, H. Komatsu, S. Nagatomo and K. Nakatani, J. Phys. Chem. B, 2021, 125, 9853–9859 Search PubMed.
- A. Miyagawa, Y. Ueda and K. Nakatani, Phys. Chem. Chem. Phys., 2024, 26, 5615–5620 RSC.
- D. Miyoshi, H. Karimata and N. Sugimoto, J. Am. Chem. Soc., 2006, 128, 7957–7963 CrossRef CAS PubMed.
- D. Hall and A. P. Minton, Biochim. Biophys. Acta, 2003, 1649, 127–139 CrossRef CAS PubMed.
- S. L. Speer, C. J. Stewart, L. Sapir, D. Harries and G. Pielak, Annu. Rev. Biophys., 2022, 51, 267–300 CrossRef CAS PubMed.
- I. M. Kuznetsova, K. K. Turoverov and V. N. Uversky, Int. J. Mol. Sci., 2014, 15, 23090–23140 CrossRef PubMed.
- M. B. Gee and P. E. Smith, J. Chem. Phys., 2009, 131, 165101 CrossRef PubMed.
- N. A. Bustos, C. M. Saad-Roy, A. G. Cherstvy and C. E. Wagner, Soft Matter, 2022, 18, 8572 RSC.
- K. Klett, A. G. Cherstvy, J. Shin, I. M. Sokolov and R. Metzler, Phys. Rev., 2021, 104, 064603 CAS.
- G. Rivas, F. Ferrone and J. Herzfeld, EMBO Rep., 2004, 5, 23–27 CrossRef CAS PubMed.
- D. Kilburn, J. H. Roh, L. Guo, R. M. Brinber and S. A. Woodson, J. Am. Chem. Soc., 2010, 132, 8690–8696 CrossRef CAS PubMed.
- J. A. Parsegian, R. P. Rand and D. C. Rau, Methods Enzymol., 1995, 259, 43–94 Search PubMed.
- F. Despa, D. P. Orgill and R. C. Lee, Ann. N. Y. Acad. Sci., 2005, 1066, 54–66 CrossRef CAS PubMed.
- D. Miyoshi and N. Sugimoto, Biochimie, 2008, 90, 1040–1051 CrossRef CAS PubMed.
- A. P. Minton, Mol. Cellular Biochem., 1983, 55, 119–140 CrossRef CAS PubMed.
- S. Nakano, D. Miyoshi and N. Sugimoto, Chem. Rev., 2014, 114, 2733–2758 CrossRef CAS PubMed.
- A. Miyagawa, H. Komatsu, S. Nagatomo and K. Nakatani, J. Mol. Liq., 2022, 360, 119526 CrossRef CAS.
- A. Miyagawa and K. Nakatani, Anal. Sci., 2022, 38, 1505–1512 CrossRef CAS PubMed.
- A. Miyagawa, H. Komatsu, S. Nagatomo and K. Nakatani, J. Mol. Liq., 2023, 372, 121181 CrossRef CAS.
- A. Miyagawa, H. Komatsu, S. Nagatomo and K. Nakatani, Bull. Chem. Soc. Jpn., 2024, 97, uoae064 CrossRef CAS.
- A. Miyagawa, Bull. Chem. Soc. Jpn., 2026, 99, uoaf116 CrossRef CAS.
- J. J. P. Stewart, J. Mol. Model., 2013, 19, 1–32 CrossRef CAS PubMed.
- L.-F. Wang, X.-W. Meng and F.-Q. Tang, J. Mol. Struct., 2010, 956, 26–32 CrossRef CAS.
- K. Toda, I. Sanemasa and T. Deguchi, Bunseki Kagaku, 1996, 45, 659–665 CrossRef CAS.
- A. Miyagawa, J. Eng, T. Okada, Y. Inoue, T. J. Penfold and G. Fukuhara, ACS Omega, 2020, 5, 897–903 CrossRef CAS PubMed.
- C. Reichardt, Chem. Rev., 1994, 94, 2319–2358 CrossRef CAS.
- S. B. Zimmerman and A. P. Minton, Annu. Rev. Biaphys. Biamal. Struct., 1993, 22, 27–65 CrossRef CAS PubMed.
- S. I. Sandler, Chemical, Biochemical, and Engineering Thermodynamics, 5th Edition, John Wiley & sons, Inc., Hoboken, New Jersey, 2017 Search PubMed.
- J. I. Carrero, ChemTexts, 2024, 10, 4 Search PubMed.
| This journal is © the Owner Societies 2026 |