Open Access Article
Open Access ArticleMolecular dynamics insights into the interactions of a potential neurotherapeutic peptide with model liposomes
Gulsah
Gul

Department of Chemical and Biological Engineering, Koç University, İstanbul, Turkey. E-mail: gugul@ku.edu.tr
First published on 12th March 2026
Abstract
Peptide molecules capable of disrupting toxic protein aggregates implicated in neurodegenerative diseases hold significant therapeutic potential; however, their clinical translation is constrained by rapid proteolysis and poor penetration into brain tissue. Lipid-based nanoparticles provide a promising delivery platform due to their ability to encapsulate diverse therapeutic cargos, reduce toxicity, and offer high biocompatibility. Here, therefore, we investigated the interactions of a cationic inhibitor peptide, KR, originally developed against Alzheimer's disease, with PC/PG lipid bilayers containing varying cholesterol concentrations using both atomistic and coarse-grained molecular dynamics (MD) simulations. Atomistic simulations revealed that the membrane response to KR is concentration-dependent: higher peptide loadings enhance lipid mobility and slightly increase the area per lipid, especially in cholesterol-free membranes, where deeper insertion facilitates local membrane loosening. KR peptides were preferentially associated with lipid headgroups through electrostatic and hydrogen-bond interactions, predominantly mediated by C-terminal Arg residues. Cholesterol reduced membrane permeability and, in coarse-grained simulations, strengthened both van der Waals and electrostatic interactions with PG lipids, resulting in peptides forming roughly three times more contacts with PG than with PC lipids. Across all systems, KR could not traverse the hydrophobic membrane core from bulk solution, yet peptides were efficiently encapsulated when partially embedded within the bilayer interior. Our study constitutes one of the first multiscale MD investigations of a potential neurotherapeutic peptide at the molecular level and provides mechanistic insights for designing liposomal nanocarriers for peptide delivery to the brain.
Introduction
Liposomes are vesicular structures composed of one or more concentric lipid bilayers, typically consisting of phospholipids and cholesterol, with sizes ranging from tens of nanometers to several micrometers.1 They were first discovered when the spontaneous self-assembly of lipids in aqueous media was observed by Alec Bangham and his colleagues.2 Since then, liposomes have been employed to deliver a wide variety of bioactive molecules, from small-molecule anticancer drugs to nucleic acids, proteins, and antibodies.3 Their growing popularity stems largely from their structural versatility: liposomes can encapsulate hydrophilic compounds within their aqueous core, incorporate hydrophobic drugs into their lipid bilayers, and accommodate amphiphilic molecules in the interfacial regions between the two.4 Furthermore, they are able to enhance the stability and biodistribution of therapeutic compounds, facilitate cellular uptake, minimize systemic toxicity and enable controlled release.3,5Despite these advantages, most liposomal drug-delivery efforts have centered on anticancer and antifungal drugs, whereas applications targeting neurological diseases remain relatively underexplored. Neurological disorders, however, pose a growing threat to public health, and the increased incidence of neurodegeneration observed in the post-COVID era highlights the urgent need for novel therapeutic strategies.6,7 In this context, peptide-based therapeutics have shown great potential, as they can disrupt pathogenic protein fibrils implicated in disorders such as Alzheimer's and Parkinson's disease.8–10 Yet, they face major pharmacokinetic hurdles, including rapid enzymatic breakdown and limited penetration across the blood–brain barrier (BBB). Given these challenges, liposomal encapsulation offers an effective strategy to enhance peptide stability, prolong the circulation time and improve brain delivery. For instance, Zheng et al. showed that H102, a β-sheet breaker peptide, can be delivered to the brain via intranasal administration using liposomes, resulting in improved memory in rats through reduced amyloid deposition.11 Likewise, Rezaei et al. combined experimental and modeling approaches to demonstrate that encapsulation of an antiangiogenic endostatin-derived peptide (ES-SS) into POPC liposomes improved both structural stability and cell viability compared to free peptide formulations.12
To rationally design such systems and optimize their performance, molecular-level insights into lipid organization and drug–membrane interactions are crucial. Molecular dynamics (MD) simulations have therefore become indispensable tools for exploring the structural and dynamical properties of lipid assemblies at atomic resolution.13 Previously, atomistic MD tools were utilized to investigate how antimicrobial peptides interact with model membranes,14–17 providing mechanistic information on lipid binding, peptide conformation, translocation, and membrane perturbation that are often inaccessible to experimental observation. However, these simulations are typically restricted to planar bilayers and become computationally prohibitive when extended to vesicular geometries, which require much larger system sizes and longer timescales to capture lipid rearrangement, curvature effects, and encapsulation events. Coarse-grained (CG) molecular dynamics methods overcome these limitations by reducing the level of atomic detail, thereby enabling the study of complex phenomena such as vesicle formation, fusion, and drug encapsulation over microsecond to millisecond timescales.18,19
Accordingly, prior work has demonstrated the capability of coarse-grained simulations to determine the physicochemical behavior of liposomal systems. Vesicle-like structures of lipid-based nanoparticles have been characterized to elucidate their drug delivery potential,20 while variations in temperature and solvent composition (ethanol concentration) have been shown to modulate liposome self-assembly and stability.21 Drug–membrane interactions have also been explored though only sporadically; for example, the distribution and orientation of hypericin molecules within lipid bilayers were resolved on the microsecond timescale, providing mechanistic insight into molecular alignment and partitioning behavior.22 Similarly, simulations of naproxen and palmatine HCl, representing poorly and highly water-soluble drugs, respectively, highlighted the critical influence of drug physicochemical properties on encapsulation efficiency and membrane stability.23 More recently, reduced-scale CG models have been developed to investigate marketed liposomal formulations, revealing structural details of both passive and active loading formulations.24 However, coarse-grained studies focusing on the encapsulation and membrane organization of neurotherapeutic peptides within liposomal carriers remain scarce.
So far, we extended the strategy of peptide delivery via liposomes by examining the interactions of β-sheet breaker peptides, which exhibit inhibitory activity against neurological targets, with liposomal nanocarriers.25 Through molecular docking and dynamics analyses, we identified that the KR peptide, which is a cationic 12-amino acid sequence derived from amyloid-β(16–20),26 displays strong binding affinity and structural stability toward both amyloid-β and α-synuclein fibrillar assemblies. In the present study, we further examine the interaction and encapsulation behavior of KR peptides within liposomes, considering both bilayer and vesicular forms to determine their structural and dynamical characteristics in lipid environments. Our work sheds light on significant interactions governing peptide–lipid association and reveals the effect of the lipid composition, peptide concentration and initial configuration on the encapsulation process across multiple scales under physiologically relevant conditions.
Computational methodology
Atomistic MD simulations
| Composition | DMPC | DMPG | CHOL | W | NA | CL | KR |
|---|---|---|---|---|---|---|---|
| DMPC/DMPG | 486 | 26 | — | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 720 720 |
104 | 78 | — |
| DMPC/DMPG/CHOL15 | 410 | 26 | 76 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 720 720 |
104 | 78 | — |
| DMPC/DMPG/CHOL30 | 334 | 26 | 152 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 720 720 |
105 | 79 | — |
| DMPC/DMPG/KR | 486 | 26 | — | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 720 720 |
104 | 84 | 1 |
| DMPC/DMPG/CHOL15/KR | 410 | 26 | 76 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 720 720 |
104 | 84 | 1 |
| DMPC/DMPG/CHOL30/KR | 334 | 26 | 152 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 720 720 |
105 | 85 | 1 |
| DMPC/DMPG/5KR | 486 | 26 | — | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 696 696 |
104 | 108 | 5 |
| DMPC/DMPG/CHOL15/5KR | 410 | 26 | 76 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 696 696 |
104 | 108 | 5 |
| DMPC/DMPG/CHOL30/5KR | 334 | 26 | 152 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 696 696 |
105 | 109 | 5 |
For peptide incorporation, KR peptide conformations were obtained from cluster analysis of our previous peptide–protein complex simulations.25 Either one or five peptide molecules were placed randomly in the bulk water region of the equilibrated membrane systems using VMD,31 and simulations were carried out following the same protocol for bare membranes.
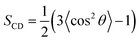 | (1) |
Peptide–lipid hydrogen bonds were analyzed using VMD with a 3.5 Å distance cutoff and a 35° angle criterion.
Coarse-grained MD simulations
For peptide encapsulation tests, KR peptides were modeled using Martini 3 force fields with neutral terminal groups.39 Two initial configurations were considered: (i) peptides randomly distributed in the bulk water phase outside the pre-formed vesicles and (ii) peptides positioned approximately half inside and half outside the vesicles. The peptide-to-lipid mass ratio was set to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40, consistent with experimental reports42,43 and atomistic 5KR systems, corresponding to a total of 35 molecules. Peptide insertion into the equilibrated liposomal systems was conducted according to the specified scenarios using the gmx insert-molecules module, and all system composition details are presented in Table 2.
40, consistent with experimental reports42,43 and atomistic 5KR systems, corresponding to a total of 35 molecules. Peptide insertion into the equilibrated liposomal systems was conducted according to the specified scenarios using the gmx insert-molecules module, and all system composition details are presented in Table 2.
| Composition | DLPC | DLPG | CHOL | W | NA | CL | KR |
|---|---|---|---|---|---|---|---|
| DLPC/DLPG | 3376 | 177 | — | 230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 236 236 |
2676 | 2499 | — |
| DLPC/DLPG/CHOL15 | 2859 | 178 | 535 | 230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 268 268 |
2677 | 2499 | — |
| DLPC/DLPG/CHOL30 | 2337 | 179 | 1078 | 230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 376 376 |
2678 | 2499 | — |
| DLPC/DLPG/KR | 3376 | 177 | — | 230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 446 446 |
2466 | 2499 | 35 |
| DLPC/DLPG/CHOL15/KR | 2859 | 178 | 535 | 230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 445 445 |
2467 | 2499 | 35 |
| DLPC/DLPG/CHOL30/KR | 2337 | 179 | 1078 | 230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 444 444 |
2468 | 2499 | 35 |
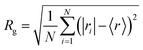 | (2) |
Results and discussion
Atomistic MD simulation results
Prior to this study, we compared the binding characteristics of β-sheet breaker peptides against those of amyloid-β and α-synuclein fibrils, two key targets in neurodegenerative diseases. Of 50 peptides examined, the cationic peptide KR exhibited the strongest binding affinity along with robust interactions with both targets.25 Since peptides are prone to easy degradation and poor permeability from the BBB, we proposed a strategy to deliver them via liposome nanocarriers. For this aim, in this study, we performed molecular dynamics simulations of KR peptides at two different concentrations across DMPC/DMPG liposomes at 0, 15 and 30% CHOL levels to understand peptide–lipid interactions, translocation behavior and encapsulation potential.The resulting membrane properties of both bare and peptide-containing systems are summarized in Table 3. The area-per-lipid and bilayer thickness values obtained for the DMPC/DMPG membranes agree well with previous computational and experimental reports on DMPC, which describe APL values of 0.60–0.61 nm2 and thicknesses of 3.60–3.61 nm under comparable thermal conditions.46–48 Lipid lateral diffusivities also follow literature trends which yield ∼9 and ∼11 µm2 s−1 for DMPC membranes at 30 °C and 35 °C, respectively.49,50
| Lipid composition | Area per lipid (nm2) | Bilayer thickness (nm) | Lipid lateral diffusion constant (µm2 s−1) |
|---|---|---|---|
| DMPC/DMPG | 0.607 ± 0.007 | 3.60 ± 0.03 | 10.5 ± 0.2 |
| DMPC/DMPG/CHOL15 | 0.507 ± 0.006 | 3.99 ± 0.03 | 4.9 ± 0.3 |
| DMPC/DMPG/CHOL30 | 0.429 ± 0.004 | 4.34 ± 0.02 | 2.4 ± 1.5 |
| DMPC/DMPG/KR | 0.607 ± 0.007 | 3.60 ± 0.04 | 10.3 ± 0.4 |
| DMPC/DMPG/CHOL15/KR | 0.511 ± 0.005 | 3.96 ± 0.03 | 7.8 ± 1.4 |
| DMPC/DMPG/CHOL30/KR | 0.429 ± 0.004 | 4.34 ± 0.03 | 1.6 ± 0.1 |
| DMPC/DMPG/5KR | 0.617 ± 0.007 | 3.56 ± 0.03 | 12.1 ± 0.9 |
| DMPC/DMPG/CHOL15/5KR | 0.515 ± 0.006 | 3.93 ± 0.03 | 5.0 ± 0.2 |
| DMPC/DMPG/CHOL30/5KR | 0.433 ± 0.004 | 4.32 ± 0.03 | 1.9 ± 0.2 |
In the fluid state, where temperatures exceed the gel–fluid transition, cholesterol enhances acyl chain ordering and bilayer rigidity.51 Accordingly, the addition of cholesterol in our systems reduced both the area per lipid and lipid mobility, while increasing the overall membrane thickness. The APL decreased to ∼0.51 and ∼0.43 nm2 at 15% and 30% CHOL, respectively, whereas the bilayer thickness increased to ∼4.0 and ∼4.3 nm. Lateral diffusion coefficients simultaneously dropped from 10.5 µm2 s−1 to 4.9 and ∼2.4 µm2 s−1, reflecting reduced permeability and restricted molecular motion due to increased lipid ordering. Similar concentration-dependent effects of cholesterol have been reported for DMPC membranes at 30 °C, with an APL value of ∼0.44 nm2 and a lateral diffusion constant of ∼2 µm2 s−1 at 30% cholesterol, consistent with our results.49,52 These findings indicate that the presence of 5% DMPG in mixtures produces similar results to DMPC-alone membranes.
Furthermore, as shown in Table 3, the presence of peptides did not significantly alter membrane properties at either concentration. However, in systems containing five KR molecules, a modest increase in APL was observed across all membrane models, with the effect being more pronounced in the cholesterol-free bilayer. In the DMPC/DMPG membrane, peptides penetrated more deeply into the bilayer, which, in turn, promoted higher lipid lateral diffusivity. A similar trend was observed in the DMPC/DMPG/CHOL15/KR system, where the peptide remained embedded at the membrane surface, leading to enhanced diffusivity. Taken together, these results suggest that when peptides insert along their full length rather than interacting only through their termini, lipid mobility is enhanced in parallel with the accompanying increase in APL.
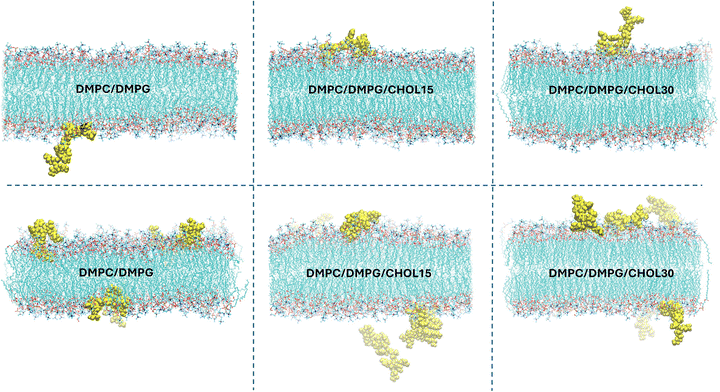
Lipid order parameters are another important indicator of the membrane structure, reflecting how regularly the lipid tails are packed. In this sense, deuterium order parameters for DMPC and DMPG lipids across different membrane models are shown in Fig. 2. As the cholesterol concentration increases, its condensing effect led to higher lipid ordering in both PC and PG components.53 In the presence of a single KR molecule, the order parameter profiles nearly overlapped with those of the peptide-free membranes. However, the addition of five peptides introduced a slight perturbation in acyl-chain ordering, with PG lipids exhibiting a more pronounced decrease. Nevertheless, peptides did not exert a major influence on the structural or dynamic properties of the membranes. Spatial distribution analysis (Fig. 3) further confirmed that peptides interact predominantly with the membrane surface rather than penetrating deeply into the bilayer. As a result, their presence did not substantially alter main characteristics such as the area per lipid, thickness, lateral diffusion, or order parameters, displaying that the membrane architecture remains intact.
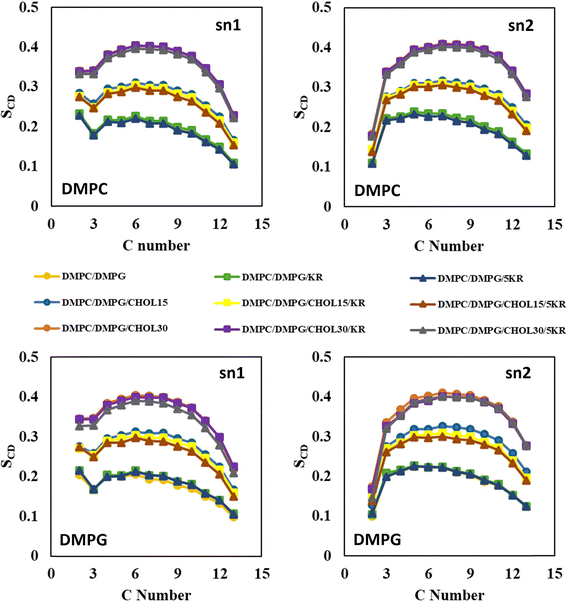 | ||
| Fig. 2 Lipid deuterium order parameters (SCD) of DMPC and DMPG in DMPC/DMPG membranes with varying cholesterol levels, in the absence and presence of KR peptides. | ||
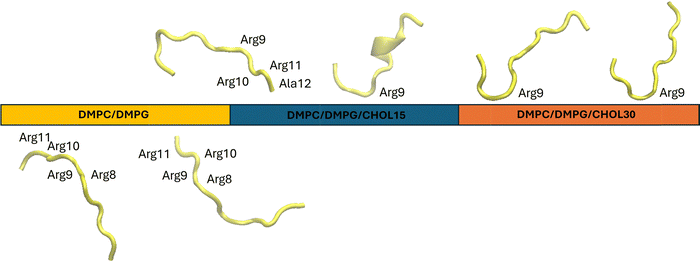
A detailed examination of peptide conformational behavior at the membrane surface revealed that peptides predominantly adopt a random coil conformation and interact with the lipid headgroups from their Arg-rich C-terminus (Fig. 4). In particular, they had a tendency to bend around Arg11 or Arg9 residues during surface adsorption. In systems containing five peptides, the peptides generally interacted more with the membrane rather than forming clusters among themselves. However, even though membrane binding is generally favored, hydrophobic interactions occasionally triggered peptide–peptide association. For example, in the CHOL15 system, a dimeric peptide cluster was observed, stabilized by reciprocal interactions between the Val4 and Phe5 residues of the two peptides (Fig. S1(a)).
In this dimeric state, both peptides continued to interact with the membrane through their C-terminal regions (Fig. S2), while residues 3–5 remained engaged in peptide–peptide contacts, indicating that dimer formation does not substantially interfere with membrane surface association. Consistently, dimerization is accompanied by an increase in short-ranged non-bonded peptide–peptide interactions, primarily driven by Lennard-Jones (van der Waals) contributions, as illustrated in Fig. S3. As no large peptide aggregates are observed within the simulated timescales and peptide concentrations, no significant disruption of the membrane structure or dynamics is detected under the conditions studied.
Peptide insertion was greater in DMPC/DMPG membranes, with the central region, including Phe5 and Phe6, appearing to play a key role in facilitating deeper penetration into the bilayer (cf. Fig. S1(b)). To further support this observation, we quantified the average distance of the peptides from the bilayer midplane and obtained values of 2.3 ± 0.2 nm, 2.5 ± 0.3 nm, and 3.0 ± 0.2 nm for membranes with single KR containing 0%, 15% and 30% CHOL, respectively. All independent trials, whether with a single peptide or five peptides, yielded the same trend (cf. Fig. S4 and S5), proving the diminished membrane permeability with increasing CHOL content. In line with this observation, the average total non-bonded interaction energies between the peptides and the membranes become disfavored in the presence of CHOL (Fig. S3), supporting the reduced membrane affinity observed in cholesterol-containing bilayers.
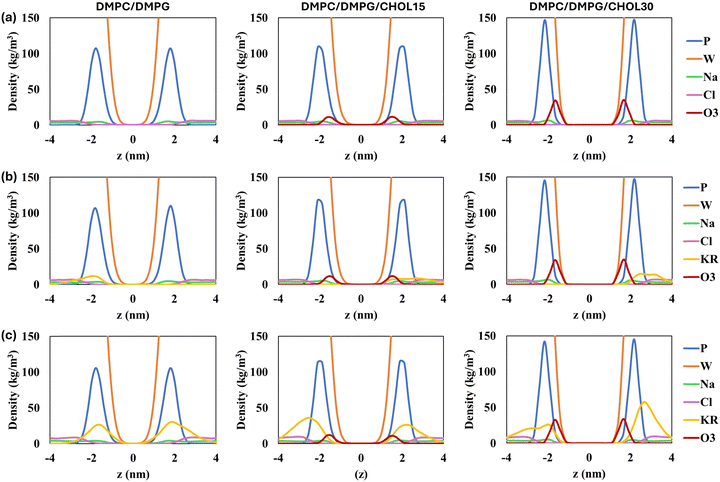
These findings are also supported by the density distributions of different molecular groups relative to the membrane center (z = 0), shown in Fig. 5. The peptide density profiles align with the insertion depths reported above: in CHOL-free membranes, peptide density peaks are located approximately ±2 nm from the bilayer center, while in CHOL-containing systems, these peaks broaden toward the bulk solvent. As the CHOL concentration increases, its peak height increases as expected and the phosphorus headgroup peaks become sharper due to enhanced lipid ordering. Furthermore, the water density exhibits a steeper decay toward the bilayer midplane, consistent with reduced water permeability.
Ion density profiles (Fig. S6) indicate that Na+ ions redistribute depending on the peptide position. For example, in membrane models containing a single KR peptide, Na+ density decreases in the leaflet where the peptide resides and increases in the opposite leaflet, reflecting repulsive interactions with the cationic peptide and the reduced accessible membrane surface area for Na+ in the peptide-occupied leaflet. In contrast, Cl− densities remain relatively unchanged. Because Na+ has a much smaller ionic radius than Cl−, its charge is concentrated over a significantly smaller surface area, resulting in a higher charge density. This higher charge density strengthens its electrostatic interactions with negatively charged membrane headgroups, explaining why Na+ responds more strongly than Cl− in our simulations.
In CHOL-containing membrane systems with five KR peptides on the other hand (Fig. S6(c)), Cl− density increases on the peptide-facing side of the membrane due to attractive interactions between the anions and the cationic peptides. The reduced degree of peptide insertion into the bilayer in these systems leaves more of the peptide exposed to the solvent, providing an additional opportunity for Cl− ions to interact with them. The relatively symmetric ion profiles observed in DMPC–DMPG membranes with five KR peptides arise from the deeper insertion of the peptides into the bilayer.
The preferential interactions between cationic peptides and anionic lipid membranes are mainly driven by electrostatic interactions and hydrogen bonding.14 Although DMPC is zwitterionic, and DMPG is negatively charged, both lipids carry negatively charged groups in their headgroups, which facilitate interactions with the cationic residues of the peptides. However, in DMPC, the presence of both positive and negative charges partially balances these interactions, reducing the overall affinity. Additionally, the smaller headgroup volume and larger lipid area of DMPG make it energetically more favorable for peptides to associate with DMPG, as this allows the displacement of more water molecules from the lipid headgroups.54 Therefore, both enthalpic contributions through peptide–lipid interactions such as electrostatics and hydrogen bonding and entropic gain through water expulsion from structured hydration layers around lipid headgroups direct peptides more towards DMPG than DMPC.
To further understand these enthalpic contributions, we analyzed hydrogen bonding patterns in our simulations, which reveal how peptide–lipid interactions stabilize the association with DMPG. Peptide residues involved in hydrogen bonds with lipids are summarized in Table 4, alongside the lipid atoms participating in these interactions. It is observed that more hydrogen bonds were formed in cholesterol-free systems, consistent with the greater accessibility and flexibility of lipid headgroups in the absence of cholesterol. Hydrogen bonds mainly occur between Arg side chains of peptides (–NH1 and –NH2) and the phosphate oxygens (O13 and O14) of DMPC and DMPG. Notably, the more C-terminal Arg residues (Arg9, Arg10, and Arg11) exhibited higher occupancies than Arg8. Although Arg residues dominate hydrogen bonding, Lys, Leu and Ala also contributed to peptide–membrane association though less prominently. This is attributed to the ability of Arg guanidinium groups to form multiple hydrogen bonds simultaneously, enhancing interaction stability.25,55,56
| Lipid composition | Amino acids participating in hydrogen bonding | Lipid atoms participating in hydrogen bonding | Total number of hydrogen bondsa |
|---|---|---|---|
| a The total hydrogen bonds obtained from the two independent trials are listed in the table as comma-separated values. | |||
| DMPC/DMPG/KR | Arg10, Arg9, Arg11 | O14, O11, O13 | 12, 15 |
| DMPC/DMPG/CHOL15/KR | Arg11, Arg8 | O14, O13 | 9, 13 |
| DMPC/DMPG/CHOL30/KR | Arg9, Arg11, Arg8, Arg10 | O14, O13, O11, O12 | 14, 14 |
| DMPC/DMPG/5KR | Arg10, Arg9, Arg11, Arg8, Ala7, Leu3, Lys2 | O14, O11, O13, O32 | 56, 58 |
| DMPC/DMPG/CHOL15/5KR | Arg9, Arg10, Arg8, Arg11 | O14, O13, O22, O11 | 41, 43 |
| DMPC/DMPG/CHOL30/5KR | Arg11, Ala12, Arg9, Arg10, Arg8, Ala7 | O13, O14, O11, O32, O22 | 30, 51 |
Coarse-grained MD simulation results
Coarse-grained MD simulations enable the investigation of realistic vesicular membrane systems which can mimic liposomal drug encapsulation, allowing access to larger spatial scales, membrane curvature, and longer timescales than atomistic simulations alone. To extend the mechanistic insights obtained from atomistic membrane systems, coarse-grained MD simulations were therefore carried out on DLPC/DLPG vesicular membranes at the same cholesterol concentrations, using two different initial peptide configurations. In the first set of simulations, all peptides were initially placed in the bulk solvent to monitor their translocation toward the lipid bilayer and to assess the energetic feasibility of membrane permeation. As shown in Fig. 6, none of the peptides penetrated the membrane core throughout the simulations; instead, they remained embedded in the headgroup region of the outer leaflet. In contrast, when peptide candidates were initially positioned partially inside and partially outside the vesicles, they became attached in the headgroups from both the inner and outer leaflets (Fig. 7). However, no transfer occurred from the membrane core to the extracellular region, nor from the extracellular region into the core.It should be noted that the vesicles simulated in this study have an estimated diameter of ∼20 nm, which falls within the commonly used scale for CG simulations21 but is smaller than that of typical therapeutic liposomes (generally 50–200 nm).57 Due to the computational limitation on modeling larger vesicles, the relatively small vesicle size results in a high membrane curvature which may influence peptide binding and insertion as well. Nonetheless, the agreement between atomistic and coarse-grained results supports the generality of the observed peptide–membrane interaction trends within the accessible simulation timescales.
Fig. 6 and 7 show that peptides progressively interact with the lipid headgroups and adsorb onto the membrane surface throughout the simulation period. Across all models, the vesicles exhibit only minor local shape deformations, and no permanent disruption of the bilayer structure is observed in the presence of peptides. The extent of adsorption depends on cholesterol levels, with up to 30% cholesterol promoting peptide binding. In both initial configurations, more peptides were observed to remain in the aqueous phase in DLPC/DLPG liposomes compared to cholesterol-containing systems. In CHOL-containing vesicles, by 1 µs, nearly all peptides were incorporated into the bilayer via either leaflet. Over the next 1 µs, the vast majority of peptides maintained interaction with the liposomes, although one or two peptides were occasionally found in the aqueous phase. These comparative interaction patterns are illustrated in the magnified final-frame snapshots shown in Fig. 8.
Time-dependent association behavior further reveals differences between the two initial configurations. As shown in Fig. 7, peptides initially placed within the membrane core began to disperse within the first 100 ns, moving toward the inner leaflet headgroups. Rather than forming aggregates, they tend to become separated over 1 µs simulation. Similar to the externally placed configuration, peptide penetration across the membranes remains limited due to surface interactions; however, the initial positioning allows peptides to orient toward the membrane surface more rapidly. In other words, peptides initially located entirely outside the membrane adsorb more slowly, whereas those partially embedded display faster and stronger surface interactions.
The radial distribution functions of the phosphate headgroups relative to the liposome center of mass (see Fig. 9) revealed two distinct peaks at approximately 8 nm (r1) and 11 nm (r2), corresponding to the inner and outer PO4 headgroup regions, respectively. The higher intensity of the first peak indicates a greater number of PO4 groups, and hence more PC lipids, present in the inner leaflet, as expected from the vesicle structure. The distance between these peaks was similar in all systems, giving a bilayer thickness between ∼3.1 and 3.2 nm. At first glance, this contradicts the bilayer thickening effect typically observed with increasing cholesterol content; however, the discrepancy stems from the unequal total lipid numbers obtained during vesicle construction. By keeping the vesicle radius constant across all systems, rather than maintaining the same lipid count, the calculated bilayer thicknesses converge to similar values. As a result, the intrinsic thickening effect of cholesterol is masked by the geometric constraint imposed during vesicle generation.
The RDF peaks of peptides corroborate the positional trends shown in Fig. 6 and 7. Upon addition of cholesterol, the peaks shift slightly inward, reflecting the increased compactness of the bilayer and causing peptides to position closer to the membrane surface. Fig. 9(a) demonstrates that when the peptides are initially placed in the bulk water phase, they do not traverse the membrane but instead remain localized near the outer PO4 headgroups. In contrast, Fig. 9(b) illustrates that peptides can populate both the inner and outer PO4 regions when they begin in a half-inserted configuration. Additionally, they approach distances below 8 nm in the DLPC/DLPG and DLPC/DLPG/CHOL15 systems, whereas in DLPC/DLPG/CHOL30, they show no tendency to penetrate toward the membrane core (also see Fig. S7). Although the intensity of the radial distribution profiles increases with cholesterol content, the overall bilayer structure remains preserved.
The thickness of pure DLPC bilayers at 30 °C has been reported to be 3.14 nm46 and 3.26 nm,58 with corresponding APL values of 0.632 nm2 and 0.608 nm2, respectively. For DLPG bilayers, Pan et al. reported a thickness of 2.91 nm and an APL of 0.656 nm2.59 When compared with these reference values at 30 °C, our APL and thickness values in the DLPC/DLPG bilayer obtained at 37 °C (0.673 nm2 and 3.20 nm, respectively) are consistent with the expected temperature dependence of lipid bilayers: a slight increase in APL accompanied by a slight decrease in bilayer thickness at higher temperatures.59 Upon addition of CHOL, a decrease in APL values was observed (see Fig. 10(a)), consistent with the well-known condensation effect that enhances lipid packing and promotes bilayer compactness. However, compared to atomistic DMPC/DMPG bilayers, the magnitude of the APL reduction appears less pronounced in the present DLPC/DLPG system. This behavior can be primarily attributed to the shorter acyl chain length of DLPC (C12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0), which results in reduced hydrophobic matching with the rigid sterol ring of cholesterol and limits its packing efficiency.60 In addition, the higher intrinsic fluidity of DLPC bilayers may further attenuate the condensation effect of cholesterol.61 Thus, the APL declined from 0.67 nm2 in the CHOL-free bilayer to 0.65 nm2 and 0.63 nm2 at 15% and 30% CHOL, respectively. Furthermore, peptide-induced effects also modulated membrane packing: when all peptides interacted with the membrane solely from the outer leaflet (“KR-out”), the APL values were higher than that in the partially inserted (“KR-half in”) configuration, indicating that superficial binding perturbs lipid packing more strongly than partial insertion.
0), which results in reduced hydrophobic matching with the rigid sterol ring of cholesterol and limits its packing efficiency.60 In addition, the higher intrinsic fluidity of DLPC bilayers may further attenuate the condensation effect of cholesterol.61 Thus, the APL declined from 0.67 nm2 in the CHOL-free bilayer to 0.65 nm2 and 0.63 nm2 at 15% and 30% CHOL, respectively. Furthermore, peptide-induced effects also modulated membrane packing: when all peptides interacted with the membrane solely from the outer leaflet (“KR-out”), the APL values were higher than that in the partially inserted (“KR-half in”) configuration, indicating that superficial binding perturbs lipid packing more strongly than partial insertion.
Vesicle diameters were estimated as twice the radius of gyration (Rg) of the phosphate headgroups, as previously described.21 According to Rg values presented in Fig. 10(b), DLPC/DLPG liposomes exhibited diameters of around ∼20 nm, with a slight decrease upon CHOL addition, confirming the trend of bilayer thickness values obtained from RDF peaks. Notably, in cholesterol-containing vesicles, the Rg values for both peptide initial configurations were very close to those of the peptide-free controls, indicating that, at the concentrations studied, encapsulated peptides did not significantly alter the vesicle size.
As mentioned above, the diameter range studied here corresponds to the higher membrane curvature than that of typical experimental liposomes, which can induce asymmetry between the two leaflets.62 In particular, the inner leaflet becomes more tightly packed, while the outer leaflet becomes more loosely packed, as confirmed by the density profiles. This curvature-induced asymmetry may alter headgroup orientations and local charge distribution at the membrane interface.63 Nevertheless, the key peptide–lipid interaction behavior observed in the coarse-grained vesicle simulations (e.g., preferential binding to anionic lipids, dominant Arg–phosphate interactions, and no membrane crossing) was consistent with those obtained from atomistic bilayer simulations, indicating that the curvature differences do not fundamentally hinder cross-scale comparison within the scope of the present study.
To assess which lipid type the peptides preferentially interacted with, we analyzed the number of contacts formed with each lipid over time. After a rapid increase within the first 500 ns, the peptide–lipid contacts began to stabilize, reaching a plateau over the last ∼1 µs of the simulations, suggesting convergence (Fig. S8). When considering the scenario where peptides are initially outside the vesicles (Fig. 11(a)), interaction with DLPG accounted for ∼75% in the CHOL-free system, decreasing slightly to 66–67% in the CHOL-containing ones. Contacts with CHOL followed a similar trend in 15% and 30% concentration levels, reaching 12–13%. For partial peptide displacement shown in Fig. 11(b), in the DLPC/DLPG system, peptides initially had no close contact with DLPG, but within 100 ns, contacts with DLPC sharply decreased, while contacts with DLPG increased. By 2 µs, DLPG contacts (69%) were roughly twice those with DLPC (31%). Upon addition of CHOL, peptide–cholesterol contacts increased as expected, accompanied by a reduction in contacts with PC lipids. However, in both liposome models, contacts with PG lipids remained consistently high, remaining at or above 60%. These observations are supported by Fig. 12, where the radial distribution function of peptides relative to PO4 beads shows a pronounced peak at ∼0.5 nm for DLPG phosphate beads, indicating a higher binding affinity of the peptides for DLPG lipids. Such preference toward anionic lipids agrees with earlier findings for other cationic peptides, such as CM15, which similarly exhibited stronger binding to POPG-containing membranes than to zwitterionic PC bilayers due to strong electrostatic attraction.14
The time evolution of the non-bonded interaction energies over the final 1 µs of the simulations (Fig. S9) indicates that the interaction energies show equilibration with no systematic drift in this time window. Accordingly, average non-bonded interaction energies calculated during this period are presented in Fig. 13. Comparison with respect to the initial peptide placement reveals that partial insertion into the vesicle cores is energetically more favorable, as evidenced by the more negative interaction energies in all models. As cholesterol content increases, peptide–lipid interaction energies, like the number of peptide–lipid contacts, also increase. For example, the total non-bonded peptide–liposome energies in Fig. 13(b) are approximately −8500.2 kJ mol−1, −9900.3 kJ mol−1, and −10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 213.8 kJ mol−1 for 0, 15 and 30% CHOL levels, respectively.
213.8 kJ mol−1 for 0, 15 and 30% CHOL levels, respectively.
However, this trend contradicts with the atomistic simulations, where increasing CHOL content leads to a reduction in peptide–liposome interaction energies. This discrepancy may arise from several factors inherent to the modeling approaches and system specifications. In coarse-grained models such as MARTINI, electrostatic interactions can be amplified due to reduced dielectric screening and the localization of charges on coarse-grained beads, potentially leading to stronger peptide–lipid interactions, particularly with anionic lipids. Although peptide-to-lipid ratios are similar in both scales, CG simulations can still show stronger surface accumulation and collective binding effects due to larger system sizes and longer sampling times, which enhance the probability of peptide–lipid encounters. In addition, cholesterol may alter the membrane curvature and lipid packing more flexibly at the coarse-grained level, increasing the effective contact area between peptides and surrounding lipids. In contrast, atomistic simulations capture cholesterol-induced membrane ordering and stiffening more explicitly, which may restrict peptide insertion and reduce close peptide–lipid contacts, resulting in weaker non-bonded interaction energies.
As shown in Fig. 13, at 15% CHOL, both van der Waals and Coulomb interactions between peptides and DLPC reach their maximum, whereas at 30% CHOL, they decrease in parallel with the reduced DLPC concentration. This suggests that 15% CHOL enhances the interaction surface with DLPC, while 30% CHOL has the opposite effect. In contrast, peptide–DLPG interactions increase in both CHOL-containing systems despite DLPG representing only 5% of the lipids. Consistently, atomistic simulations reveal that peptide–PC interactions weaken with increasing CHOL content, whereas peptide–PG interactions strengthen (Fig. S3).
Although the total non-bonded interactions with DLPC are larger in magnitude, normalization per lipid (Fig. S10) reveals that van der Waals interactions are approximately threefold and Coulomb interactions about fivefold higher with DLPG across all models. As cholesterol lacks charged groups, it does not contribute to Coulomb interactions with peptides. van der Waals interactions with CHOL are also minimal, indicating that the enhancing effect of CHOL on peptide–lipid interactions is not driven by direct peptide–cholesterol contacts, but by the increased interaction surface with PC lipids. This is further supported by the peptide–peptide interactions: increasing CHOL content favors van der Waals interactions among peptides, whereas Coulomb interactions diminish, suggesting that electrostatic forces are primarily directed toward the membrane surface rather than between peptides. These results are consistent with our previous observations.
Lastly, to quantify lipid selectivity toward PG and to enable comparison with the CG results, we performed MM-PBSA binding free energy calculations64 for a single KR peptide interacting with pure DMPC and pure DMPG membranes at the atomistic level. Initially, the systems were analyzed analogously to a protein–ligand binding process,65 with the membrane acting as the receptor surface and the peptide as the ligand. However, this approach led to an artificial overestimation of electrostatic interactions (data not shown). Therefore, the peptide was subsequently treated as a membrane-associated protein, and periodic boundary conditions were applied to the membrane system using the geometric multigrid.
Based on these calculations, average binding free energies were estimated to be −140 kJ mol−1 and −392 kJ mol−1, for PC and PG membranes, respectively. While absolute MM-PBSA values should be interpreted with caution due to known methodological limitations, energy decomposition analysis (Table S1) reveals consistently stronger interactions with PG membranes, with approximately twofold higher van der Waals and ∼3.5-fold higher electrostatic contributions than those obtained for PC membranes. Thus, a deviation of around 30% from the relative trends of CG interaction energies was identified, highlighting qualitative agreement but quantitative differences between the two approaches.
Conclusions
In summary, we analyzed the interactions of KR peptides with PC/PG lipids at both atomistic and coarse-grained levels. Atomistic MD simulations were performed on DMPC/DMPG bilayers containing 0, 15, and 30% cholesterol at two peptide concentrations. The key findings from these simulations are summarized below:• While single-peptide addition did not significantly alter bilayer properties, insertion of five peptides increased the area per lipid and enhanced lipid lateral mobility along with a slight perturbation in the order parameter.
• The presence of cholesterol reduced membrane permeability and decreased the peptide insertion depth due to its condensing effect.
• KR peptides preferentially interacted with DMPG over DMPC, driven by stronger electrostatic attraction, favorable hydrogen bonding, and entropic gains from water displacement around PG headgroups.
• Peptide–lipid hydrogen bonding was dominated by Arg residues, particularly Arg9–Arg11, which formed stable interactions with phosphate oxygens; cholesterol reduced hydrogen-bond formation by limiting the number of accessible phosphate headgroups.
• Ion distributions also responded to peptide binding: Na+ was displaced from the peptide-facing leaflet and accumulated in the opposite leaflet, while Cl− increased near peptides in multi-peptide, cholesterol-containing systems due to electrostatic attraction.
To complement these atomistic insights and investigate longer timescale behaviors, coarse-grained MD simulations were conducted on DLPC/DLPG bilayers at the same cholesterol concentrations, with two different initial peptide placements. The main observations are as follows:
• When starting in the bulk solvent, peptides adsorbed onto the outer headgroup region and remained surface-bound. Partial initial insertion enabled encapsulation, allowing peptides to stably occupy both inner and outer leaflets, although no peptide fully crossed from one side to the other.
• Vesicles measured ∼20 nm in diameter, showing no significant expansion or contraction upon peptide encapsulation at the studied concentrations.
• Fully external peptides bound more slowly, whereas partially inserted peptides rapidly oriented toward headgroups and dispersed without forming aggregates, consistent with energetics reflecting partial insertion as more favorable with more negative total non-bonded energies.
• Interaction strength per lipid was dominated by DLPG, with Coulomb (∼5×) and van der Waals (∼3×) forces stronger than those with PC lipids, confirming the peptides’ preferential binding to anionic membranes.
Hence, our results demonstrate that coarse-grained MD simulations agree well with the atomistic findings and emphasize that proper encapsulation of KR peptides critically depends on both the lipid composition and initial peptide placement, highlighting the importance of the preparation strategy. Specifically, cationic peptides can be successfully encapsulated into anionic membranes; however, long-range electrostatic attractions with the membrane surface can have two key implications: (i) passive diffusion of peptides into pre-formed liposomes is unlikely due to adsorption and surface stacking and (ii) efficient drug release may be hindered by peptide trapping within the vesicle core, potentially slowing release. To address these challenges, active loading strategies, pH or ion gradients, could enhance encapsulation efficiency, while optimizing the proportion of anionic lipids or incorporating pH-responsive lipids may facilitate peptide delivery despite potential entrapment. These complex mechanisms warrant further investigation using blood–brain barrier models or experimental in vitro tests, guided by the mechanistic framework presented here for developing liposomal drug delivery systems targeting neurological diseases.
Author contributions
Gulsah Gul: conceptualization, data curation, formal analysis, funding acquisition, investigation, methodology, project administration, resources, supervision, validation, visualization, writing – original draft, and writing – review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Fig. S1. Representative conformations of aggregated peptides in DMPC/DMPG/CHOL15 membrane highlighting interacting residues (a) and peptide insertion in DMPC/DMPG membranes from two independent trials (b). Fig. S2. Peptide–lipid association for the peptide dimer in the DMPC/DMPG/CHOL15 membrane. C-Terminal Arg residues interact with the lipid headgroups, while Leu3, Val4 and Phe5 participate in the dimer interface. Fig. S3. Short-ranged non-bonded interaction energies between peptides (KR) and between peptides and lipid components (DMPC, DMPG, and CHOL) over time. Here, LJ stands for Lennard-Jones interactions, while Cou represents coulombic electrostatics. Fig. S4. Distribution of KR peptides across DMPC/DMPG membranes with varying cholesterol content, taken from the second independent trial. The upper panel shows systems containing a single KR peptide, while the lower panel shows systems with five KR peptides. Here, peptide molecules are represented by yellow van der Waals spheres. Fig. S5. Average center-of-mass (COM) distance between peptides and DMPC/DMPG membranes with varying cholesterol (CHOL) concentrations, calculated over the last 100 ns of the simulations. The left panel shows systems containing a single KR peptide, whereas the right panel shows systems containing five KR peptides. Fig. S6. Magnified density distributions of phosphorus (P), water (W), sodium (Na), chloride (Cl), peptide (KR), and cholesterol oxygen (O3) in DMPC/DMPG membranes with varying cholesterol levels, relative to the membrane center (z = 0). Profiles are shown for bare membranes (a), membranes containing a single KR peptide (b), and membranes containing five KR peptides (c). Fig. S7. Peptide distribution in the lower leaflet of DLPC/DLPG liposomes containing 0, 15, and 30% CHOL (left to right) at 2 µs. For clarity, only the inner-leaflet PO4 beads and peptides are shown, rendered as tan and yellow van der Waals spheres, respectively. Fig. S8. The time evolution of peptide–lipid contacts in DLPC/DLPG bilayers containing 0, 15, and 30% CHOL (left to right). The upper panel (a) corresponds to the external initial placement of peptides, whereas the lower panel (b) corresponds to their initial partial embedment. Fig. S9. Short-ranged non-bonded interaction energies between peptides (KR) and between peptides and lipid components (DMPC, DMPG, and CHOL) over time. The upper panel (a) corresponds to the external initial placement of peptides, whereas the lower panel (b) corresponds to their initial partial embedment. Fig. S10. Average van der Waals (left) and coulombic (right) interaction energies-per-lipid between peptides and each lipid type in DLPC/DLPG liposomes with varying CHOL levels. The upper panel (a) corresponds to the external initial placement of peptides, whereas the lower panel (b) corresponds to their initial partial embedment. Table S1. MM-PBSA binding free energies of the single KR peptide with pure DMPC and DMPG lipid membranes, including van der Waals (VDWAALS) and electrostatics (EEL) contributions. The Poisson–Boltzmann equation was solved using the geometric multigrid approach (solvopt = 2) with linear PB (ipb = 1) and periodic boundary conditions (bcopt = 10) to minimize unphysical edge effects. The parameter eneopt was set to 1 as the charge-view method (eneopt = 2) is not supported for protein-membrane systems; consequently, the EEL term includes both coulombic and reaction-field (EPB) energies. See DOI: https://doi.org/10.1039/d5cp04834f.Acknowledgements
The financial support was provided by the TÜBİTAK 2218 – National Postdoctoral Research Fellowship Program with project no 123C377. All-atom simulations reported in this paper were performed at TÜBİTAK ULAKBIM, High Performance and Grid Computing Center (TRUBA resources). Coarse-grained simulations were carried out using the MareNostrum 5 preexascale supercomputing system with GPU acceleration. The authors gratefully thank the Barcelona Supercomputing Center (BSC) and the Scientific and Technological Research Council of Turkey (TÜBİTAK) for providing access to these resources and supporting this research.References
- U. Bulbake, S. Doppalapudi, N. Kommineni and W. Khan, Liposomal formulations in clinical use: an updated review, Pharmaceutics, 2017, 9, 1–33, DOI:10.3390/pharmaceutics9020012.
- A. D. Bangham, M. M. Standish and J. C. Watkins, Diffusion of univalent ions across the lamellae of swollen phospholipids, J. Mol. Biol., 1965, 13, 238–252, DOI:10.1016/S0022-2836(65)80093-6.
- P. Liu, G. Chen and J. Zhang, A review of liposomes as a drug delivery system: current, Molecules, 2022, 27, 1372, DOI:10.3390/molecules27041372.
- M. Alavi, N. Karimi and M. Safaei, Application of various types of liposomes in drug delivery systems, Adv. Pharm. Bull., 2017, 7, 3–9, DOI:10.15171/apb.2017.002.
- H. Nsairat, D. Khater, U. Sayed, F. Odeh, A. Al and W. Alshaer, Liposomes: structure, composition, types, and clinical applications, Heliyon, 2022, 8, e09394, DOI:10.1016/j.heliyon.2022.e09394.
- E. Rudnicka-Drozak, P. Drozak, G. Mizerski, T. Zaborowski and B. Slusarska, Links between COVID-19 and Alzheimer's disease – What do we already know?, Int. J. Environ. Res. Public Health, 2023, 20, 2146, DOI:10.3390/ijerph20032146.
- D. Bonhenry, M. Charnley, J. Gonçalves, P. Hammarström, M. T. Heneka and R. Itzhaki, et al., SARS-CoV-2 infection as a cause of neurodegeneration, Lancet Neurol., 2024, 23, 562–563, DOI:10.1016/S1474-4422(24)00178-9.
- L. O. Tjernberg, J. Näslund, F. Lindqvist, J. Johansson, A. R. Karlström and J. Thyberg, et al., Arrest of β-amyloid fibril formation by a pentapeptide ligand, J. Biol. Chem., 1996, 271, 8545–8548, DOI:10.1074/jbc.271.15.8545.
- C. Soto, E. M. Sigurdsson, L. Morelli, R. A. Kumar, E. M. Castaño and B. Frangione, β-sheet breaker peptides inhibit fibrillogenesis in a rat brain model of amyloidosis: implications for Alzheimer's therapy, Nat. Med., 1998, 4, 822–826, DOI:10.1038/nm0798-822.
- O. M. A. El-Agnaf, K. E. Paleologou, B. Greer, A. M. Abogrein, J. E. King and S. A. Salem, et al., A strategy for designing inhibitors of α-synuclein aggregation and toxicity as a novel treatment for Parkinson's disease and related disorders, FASEB J., 2004, 18, 1315–1317, DOI:10.1096/fj.03-1346fje.
- X. Zheng, X. Shao, C. Zhang, Y. Tan, Q. Liu and X. Wan, et al., Intranasal H102 peptide-loaded liposomes for brain delivery to treat Alzheimer's disease, Pharm. Res., 2015, 32, 3837–3849, DOI:10.1007/s11095-015-1744-9.
- N. Rezaei, F. Mehrnejad, Z. Vaezi, M. Sedghi, S. M. Asghari and H. Naderi-Manesh, Encapsulation of an endostatin peptide in liposomes: stability, release, and cytotoxicity study, Colloids Surf., B, 2020, 185, 110552, DOI:10.1016/j.colsurfb.2019.110552.
- O. H. S. Ollila and G. Pabst, Atomistic resolution structure and dynamics of lipid bilayers in simulations and experiments, Biochim. Biophys. Acta, Biomembr., 2016, 1858, 2512–2528, DOI:10.1016/j.bbamem.2016.01.019.
- Y. Wang, D. E. Schlamadinger, J. E. Kim and J. A. McCammon, Comparative molecular dynamics simulations of the antimicrobial peptide CM15 in model lipid bilayers, Biochim. Biophys. Acta, Biomembr., 2012, 1818, 1402–1409, DOI:10.1016/j.bbamem.2012.02.017.
- M. Pachler, I. Kabelka, M. S. Appavou, K. Lohner, R. Vácha and G. Pabst, Magainin 2 and PGLa in bacterial membrane mimics I: peptide-peptide and lipid-peptide interactions, Biophys J., 2019, 117, 1858–1869, DOI:10.1016/j.bpj.2019.10.022.
- Z. Deng, X. Lu, C. Xu, B. Yuan and K. Yang, Lipid-specific interactions determine the organization and dynamics of membrane-active peptide melittin, Soft Matter, 2020, 16, 3498–3504, 10.1039/d0sm00046a.
- S. R. Bowers, D. K. Klimov and C. Lockhart, Mechanisms of binding of antimicrobial peptide PGLa to DMPC/DMPG membrane, J. Chem. Inf. Model., 2022, 62, 1525–1537, DOI:10.1021/acs.jcim.1c01518.
- H. I. Ingólfsson, C. A. Lopez, J. J. Uusitalo, D. H. de Jong, S. M. Gopal and X. Periole, et al., The power of coarse graining in biomolecular simulations, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2014, 4, 225–248, DOI:10.1002/wcms.1169.
- E. Khodadadi, E. Khodadadi, P. Chaturvedi and M. Moradi, Molecular dynamics simulations of liposomes: structure, dynamics, and applications, Membranes, 2025, 15, 1–37, DOI:10.3390/membranes15090259.
- M. Khalkhali, S. Mohammadinejad, F. Khoeini and K. Rostamizadeh, Vesicle-like structure of lipid-based nanoparticles as drug delivery system revealed by molecular dynamics simulations, Int. J. Pharm., 2019, 559, 173–181, DOI:10.1016/j.ijpharm.2019.01.036.
- T. Duran, A. P. Costa, J. Kneski, X. Xu, D. J. Burgess and H. Mohammadiarani, et al., Manufacturing process of liposomal formation: a coarse-grained molecular dynamics simulation, Int. J. Pharm., 2024, 659, 124288, DOI:10.1016/j.ijpharm.2024.124288.
- J. P. M. Jämbeck, E. S. E. Eriksson, A. Laaksonen, A. P. Lyubartsev and L. A. Eriksson, Molecular dynamics studies of liposomes as carriers for photosensitizing drugs: development, validation, and simulations with a coarse-grained model, J. Chem. Theory Comput., 2014, 10, 5–13, DOI:10.1021/ct400466m.
- R. Han, Z. Ye, Y. Zhang, Y. Cheng and Y. Zheng, Predicting liposome formulations by the integrated machine learning and molecular modeling, Asian J. Pharm. Sci., 2023, 18, 100811, DOI:10.1016/j.ajps.2023.100811.
- R. Wang, J. Zang, H. Zhong, Y. Zhang and D. Ouyang, Understanding the molecular insights of marketed liposomal drugs using molecular dynamics simulations of reduced scale coarse-grained models, npj Drug Discovery, 2025, 2, 11, DOI:10.1038/s44386-025-00014-5.
- G. Gul, Modeling β-sheet breaker peptides across multiple resolutions: from neurological targets to liposomal membranes, Nanoscale, 2026 10.1039/D5NR05060J.
- S. Jagota and J. Rajadas, The role of Pro, Gly, Lys, and Arg containing peptides on amyloid-beta aggregation, Int. J. Pept. Res. Ther., 2012, 18, 53–61, DOI:10.1007/s10989-011-9278-4.
- S. Jo, T. Kim, V. G. Iyer and W. Im, CHARMM-GUI: a web-based graphical user interface for CHARMM, J. Comput. Chem., 2008, 29, 1859–1865, DOI:10.1002/jcc.20945.
- J. Lee, X. Cheng, J. M. Swails, M. S. Yeom, P. K. Eastman and J. A. Lemkul, et al., CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field, J. Chem. Theory Comput., 2016, 12, 405–413, DOI:10.1021/acs.jctc.5b00935.
- J. B. Klauda, R. M. Venable, J. A. Freites, J. W. O’Connor, D. J. Tobias and C. Mondragon-Ramirez, et al., Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types, J. Phys. Chem. B, 2010, 114, 7830–7843, DOI:10.1021/jp101759q.
- J. Huang, S. Rauscher, G. Nawrocki, T. Ran, M. Feig and B. L. de Groot, et al., CHARMM36m: an improved force field for folded and intrinsically disordered proteins, Nat. Methods, 2017, 14, 71–73, DOI:10.1038/nmeth.4067.
- W. Humphrey, A. Dalke and K. Schulten, VMD: visual molecular dynamics, J. Mol. Graphics, 1996, 14, 33–38 CrossRef CAS PubMed.
- M. Bernetti and G. Bussi, Pressure control using stochastic cell rescaling, J. Chem. Phys., 2020, 153, 1–11, DOI:10.1063/5.0020514.
- G. Bussi, D. Donadio and M. Parrinello, Canonical sampling through velocity rescaling, J. Chem. Phys., 2007, 126, DOI:10.1063/1.2408420.
- T. Darden, D. York and L. Pedersen, Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems, J. Chem. Phys., 1993, 98, 10089–10092, DOI:10.1063/1.464397.
- B. Hess, H. Bekker, H. J. C. Berendsen and J. G. E. M. Fraaije, LINCS: a linear constraint solver for molecular simulations, J. Comput. Chem., 1997, 18, 1463–1472 CrossRef CAS.
- M. Abraham, et al., GROMACS software (2024). GROMACS 2024.1 source code., 2024 DOI:10.5281/zenodo.10721181.
- S. Buchoux, FATSLiM: a fast and robust software to analyze MD simulations of membranes, Bioinformatics, 2017, 33, 133–134, DOI:10.1093/bioinformatics/btw563.
- G. Gul, R. Faller and N. Ileri-Ercan, Coarse-grained modeling of polystyrene-modified CNTs and their interactions with lipid bilayers, Biophys J., 2023, 122, 1–14, DOI:10.1016/j.bpj.2023.04.005.
- P. C. T. Souza, R. Alessandri, J. Barnoud, S. Thallmair, I. Faustino and F. Grünewald, et al., Martini 3: a general purpose force field for coarse-grained molecular dynamics, Nat. Methods, 2021, 18, 382–388, DOI:10.1038/s41592-021-01098-3.
- L. Borges-Araújo, A. C. Borges-Araújo, T. N. Ozturk, D. P. Ramirez-Echemendia, B. Fábián and T. S. Carpenter, et al., Martini 3 coarse-grained force field for cholesterol, J. Chem. Theory Comput., 2023, 19, 7387–7404, DOI:10.1021/acs.jctc.3c00547.
- Y. Qi, H. I. Ingólfsson, X. Cheng, J. Lee, S. J. Marrink and W. Im, CHARMM-GUI Martini maker for coarse-grained simulations with the Martini force field, J. Chem. Theory Comput., 2015, 11, 4486–4494, DOI:10.1021/acs.jctc.5b00513.
- G. Kogkos, F. Gkartziou, S. Mourtas, K. K. Barlos, P. Klepetsanis and K. Barlos, et al., Liposomal entrapment or chemical modification of Relaxin2 for prolongation of its stability and biological activity, Biomolecules, 2022, 12(10), 1362, DOI:10.3390/biom12101362.
- A. Vitali, P. Paolicelli, B. Bigi, J. Trilli, L. Di Muzio and V. C. Carriero, et al., Liposome encapsulation of the palmitoyl–KTTKS peptide: structural and functional characterization, Pharmaceutics, 2024, 16(2), 219, DOI:10.3390/pharmaceutics16020219.
- I. G. Tironi, R. Sperb, P. E. Smith and W. F. van Gunsteren, A generalized reaction field method for molecular dynamics simulations, J. Chem. Phys., 1995, 102, 5451–5459 CrossRef CAS.
- M. Abraham, et al., GROMACS software (2023). GROMACS 2023.3 source code, 2023 DOI:10.5281/zenodo.10017686.
- N. Kučerka, Y. Liu, N. Chu, H. I. Petrache, S. Tristram-Nagle and J. F. Nagle, Structure of fully hydrated fluid phase DMPC and DLPC lipid bilayers using X-ray scattering, Biophys. J., 2005, 88, 2626–2637, DOI:10.1529/biophysj.104.056606.
- W. C. Hung, M. T. Lee, F. Y. Chen and H. W. Huang, The condensing effect of cholesterol in lipid bilayers, Biophys J., 2007, 92, 3960–3967, DOI:10.1529/biophysj.106.099234.
- T. Broemstrup and N. Reuter, Molecular dynamics simulations of mixed acidic/zwitterionic phospholipid bilayers, Biophys J., 2010, 99, 825–833, DOI:10.1016/j.bpj.2010.04.064.
- G. Lindblom and G. Orädd, Lipid lateral diffusion and membrane heterogeneity, Biochim. Biophys. Acta, Biomembr., 2009, 1788, 234–244, DOI:10.1016/j.bbamem.2008.08.016.
- A. Filippov, G. Orädd and G. Lindblom, The effect of cholesterol on the lateral diffusion of phospholipids in oriented bilayers, Biophys J., 2003, 84, 3079–3086, DOI:10.1016/S0006-3495(03)70033-2.
- T. Róg, M. Pasenkiewicz-Gierula, I. Vattulainen and M. Karttunen, Ordering effects of cholesterol and its analogues, Biochim. Biophys. Acta, Biomembr., 2009, 1788, 97–121, DOI:10.1016/j.bbamem.2008.08.022.
- G. Orädd, G. Lindblom and P. W. Westerman, Lateral diffusion of cholesterol and dimyristoylphosphatidylcholine in a lipid bilayer measured by pulsed field gradient NMR spectroscopy, Biophys J., 2002, 83, 2702–2704, DOI:10.1016/S0006-3495(02)75279-X.
- G. Khelashvili, S. Mondal, O. S. Andersen and H. Weinstein, Cholesterol modulates the membrane effects and spatial organization of membrane-penetrating ligands for G-protein coupled receptors, J. Phys. Chem. B, 2010, 114, 12046–12057, DOI:10.1021/jp106373r.
- W. T. Heller, Molecular dynamics investigation of lipid-specific interactions with a fusion peptide, Biomolecules, 2024, 14, DOI:10.3390/biom14030285.
- N. Schmidt, A. Mishra, G. H. Lai and G. C. L. Wong, Arginine-rich cell-penetrating peptides, FEBS Lett., 2010, 584, 1806–1813, DOI:10.1016/j.febslet.2009.11.046.
- A. Mohan, S. R. C. K. Rajendran, J. Thibodeau, L. Bazinet and C. C. Udenigwe, Liposome encapsulation of anionic and cationic whey peptides: influence of peptide net charge on properties of the nanovesicles, LWT Food Sci. Technol., 2018, 87, 40–46, DOI:10.1016/j.lwt.2017.08.072.
- X. Guo, Y. Zhang, Q. Liu, M. Xu, J. Pang and B. Yang, et al., Progress on liposome delivery systems in the treatment of bladder cancer, RSC Adv., 2025, 15, 14315–14336, 10.1039/d5ra00746a.
- N. Kučerka, M. P. Nieh and J. Katsaras, Fluid phase lipid areas and bilayer thicknesses of commonly used phosphatidylcholines as a function of temperature, Biochim. Biophys. Acta, Biomembr., 2011, 1808, 2761–2771, DOI:10.1016/j.bbamem.2011.07.022.
- J. Pan, F. A. Heberle, S. Tristram-Nagle, M. Szymanski, M. Koepfinger and J. Katsaras, et al., Molecular structures of fluid phase phosphatidylglycerol bilayers as determined by small angle neutron and X-ray scattering, Biochim. Biophys. Acta, Biomembr., 2012, 1818, 2135–2148, DOI:10.1016/j.bbamem.2012.05.007.
- N. Kučerka, J. D. Perlmutter, J. Pan, S. Tristram-Nagle, J. Katsaras and J. N. Sachs, The effect of cholesterol on short- and long-chain monounsaturated lipid bilayers as determined by molecular dynamics simulations and X-ray scattering, Biophys J., 2008, 95, 2792–2805, DOI:10.1529/biophysj.107.122465.
- J. Pan, T. T. Mills, S. Tristram-Nagle and J. F. Nagle, Cholesterol perturbs lipid bilayers nonuniversally, Phys. Rev. Lett., 2008, 100, 198103, DOI:10.1103/PhysRevLett.100.198103.
- H. J. Risselada and S. J. Marrink, Curvature effects on lipid packing and dynamics in liposomes revealed by coarse grained molecular dynamics simulations, Phys. Chem. Chem. Phys., 2009, 11, 2056–2067, 10.1039/b818782g.
- P. Khunpetch, A. Majee and R. Podgornik, Curvature effects in charge-regulated lipid bilayers, Soft Matter, 2022, 18, 2597–2610, 10.1039/d1sm01665b.
- M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente and E. Moreno, Gmx_MMPBSA: a new tool to perform end-state free energy calculations with GROMACS, J. Chem. Theory Comput., 2021, 17, 6281–6291, DOI:10.1021/acs.jctc.1c00645.
- G. Gul, In silico screening of peptide inhibitors targeting α-synuclein for Parkinson's disease, J. Mol. Graphics Modell., 2025, 139, 109079, DOI:10.1016/j.jmgm.2025.109079.
| This journal is © the Owner Societies 2026 |