Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
The accuracy of carbon-13 NMR magnetic-shielding tensors calculated using periodic DFT: a case study on the distinction of crystalline serine phases
Kathrin
Rübartsch
a,
Leeroy
Hendrickx
 ab,
Ettore
Bartalucci
ab,
Ettore
Bartalucci
 ab,
Maksim
Plekhanov
c,
Mirijam
Zobel
ab,
Maksim
Plekhanov
c,
Mirijam
Zobel
 c,
Carsten
Bolm
c,
Carsten
Bolm
 d,
Maria
Fyta
d,
Maria
Fyta
 *ef and
Thomas
Wiegand
*ef and
Thomas
Wiegand
 *ab
*ab
aInstitute of Technical and Macromolecular Chemistry, RWTH Aachen University, Worringerweg 2, 52074 Aachen, Germany
bMax Planck Institute for Chemical Energy Conversion, Stiftstr. 34-36, 45470 Mülheim/Ruhr, Germany. E-mail: thomas.wiegand@cec.mpg.de
cInstitute of Crystallography, RWTH Aachen University, Jägerstraße 17-19, 52066 Aachen, Germany
dInstitute of Organic Chemistry, RWTH Aachen University, Landoltweg 1, 52074 Aachen, Germany
eComputational Biotechnology, RWTH Aachen University, Worringerweg 3, 52074 Aachen, Germany. E-mail: mfyta@biotec.rwth-aachen.de
fCenter for Computational Life Sciences (CCLS), RWTH Aachen University, Aachen, Germany
First published on 17th February 2026
Abstract
Carbon-13 nuclear magnetic resonance (NMR) spectroscopic fingerprints enable the unambiguous identification of a chemical compound. Ab initio calculations of NMR magnetic-shielding tensors are crucial for facilitating spectral resonance assignment, but require an accuracy on the order of one ppm or even less for carbon-13 for distinguishing structurally rather similar compounds, such as structural isomers and polymorphs. Using quantum-mechanical calculations within the density functional theory (DFT) approach and solid-state NMR spectroscopy under magic-angle spinning (MAS) conditions, we herein explore whether the accuracy of current DFT levels is sufficient to distinguish between three distinct solid phases of the amino acid serine using computer simulations. In that vein, enantiopure L-serine, L-serine monohydrate, and racemic DL-serine have been studied. Using solid-state calculations employing periodic boundary conditions, we computed NMR observables, such as 13C isotropic magnetic-shielding values and 13C magnetic-shielding anisotropy parameters of these phases. Different levels of DFT theory utilizing distinct exchange correlation functionals were tested in optimizing the hydrogen-atom positions prior to NMR observable calculations, which significantly improved the agreement with the experimental values. The comparison of the measured and the calculated observables confirmed the distinction between the serine phases not only experimentally but also by DFT calculations. We also compare the DFT-calculated NMR observables with predictions from a recently proposed machine learning (ML) approach. Our analysis clearly revealed that a rather low-cost exchange correlation functional in periodic solid-state DFT calculations reaches the desired accuracy of 13C magnetic-shielding tensor calculations, provided that highly accurate initial structures are applied. These results underline that complementary experimental and computational studies can provide key insights into molecular systems and the interactions therein, enabling the distinction of structurally similar compounds (often polymorphs), which is of particular importance in pharmaceutics.
Introduction
The unambiguous distinction of polymorphs (or sometimes even pseudo-polymorphs, where solvent molecules are embedded in the crystal structure) is of paramount importance, for instance, in the field of pharmaceutics.1,2 NMR crystallography3–11 enables the determination of the structures of such polymorphic species. Solid-state NMR spectroscopy under magic-angle spinning (MAS) conditions plays an important role in the NMR crystallography toolbox and generally serves as a valuable tool for spectroscopically distinguishing polymorphic species, as illustrated for various examples.12–22 Typically, the isotropic magnetic-shielding value (or in the case of spectra that have been referenced to a standard compound, the chemical shift value), which is defined as the average value of the trace of the 3 × 3 magnetic-shielding tensor, is sufficient to allow the spectroscopic distinction of those, although the magnetic-shielding anisotropy provides additional information about the electronic structure of the nucleus-of-interest. The latter can, for instance, be reported in the Herzfeld–Berger convention,23 in which the span (Ω) reports the strength of the anisotropy (the widths of the static powder NMR lineshape) and the skew (κ) is a measure of the asymmetry of the tensor.The accuracy of ab initio calculations of NMR magnetic-shielding tensors is crucial for supporting spectral resonance assignment, particularly for distinguishing polymorphs, often with minor structural differences.13,24Ab initio calculations of NMR observables are typically performed in the DFT framework25 using either Gaussian- or Slater-type orbitals.26 In most molecular species, high-level gas phase calculations on monomers (or smaller molecular clusters) are often sufficient, since the NMR observables are mainly dominated by local properties. However, particularly in cases where intermolecular interactions are of importance, solid-state calculations using plane-wave basis sets need to be performed,27–30 which involves significantly higher computational costs, often restricting the DFT calculations to a lower level-of-theory and limiting the accuracy of the results.31 In such methods, the gauge-including projector-augmented wave (GIPAW) approach is used.32–35 Alternatively, quantum mechanics/molecular mechanics calculations have also been reported to compute solid-state NMR observables for molecular solids.31 In the past few years, machine learning models have also entered the field, enabling a rather efficient prediction of magnetic-shielding parameters.36–38
Some of us have recently studied three distinct phases of the amino acid serine in the context of mechanochemical molecular-recognition processes.39 The enantiopure phase of serine (L- and D-serine, respectively, both giving identical 13C solid-state NMR spectra40) shows a different 13C spectroscopic fingerprint than the racemic phase (DL-serine) with average 13C chemical-shift differences on the order of 2 ppm.39 The third serine phase we have observed in the course of our studies was the monohydrate L-serine phase (L-serine·H2O),41 which has been subjected to detailed solid-state NMR investigations before.42 The chemical-shift values (1H and 13C) of several amino acids43 have been calculated using GIPAW DFT with the PBE functional in previous work.44,45 These values were further corrected using the isolated-molecule correction scheme, in which the differences in isotropic magnetic-shielding values at a PBE and PBE0 level-of-theory for a single molecule are calculated.38,46 A mean absolute error of around 1.5 ppm was determined for 13C isotropic magnetic-shielding values.43 Recently, a very detailed study employed the experimentally determined 13C isotropic chemical-shielding values of 20 solid amino acids (the L-enantiomers have been studied) for benchmarking periodic GIPAW calculations with GGA, meta-GGA, and hybrid DFT functionals.20 Interestingly, best agreement between the experimental and calculated values has been achieved for the GGA level-of-theory (PBE functional).20
We herein study whether the current accuracy of plane-wave DFT calculations is sufficient to unambiguously distinguish the three serine phases mentioned above using 13C magnetic-shielding tensor calculations. Based on initial benchmarks on the main DFT parameters and detailed test calculations for the different structures and exchange correlation functionals, we propose a strategy for periodic DFT calculations of interacting molecular systems. We do not intend to benchmark, as recently described in an excellent article,20 but aim at illustrating, for our specific example, the potential of plane-wave periodic DFT calculations in calculating 13C magnetic-shielding tensors, enabling the distinction of polymorphs. Our studies particularly highlight the importance of an accurate starting structure for such calculations (as often provided by single-crystal X-ray or neutron diffraction),47 which indeed allows the distinction between the separate serine phases, thus reproducing experimentally observed trends.
Experimental
All chemicals employed in this work were obtained from commercial sources (see Table 1). The materials (except for DL-serine used for the experiments at 16.4 T) were ball milled for 20 min at 25 Hz in a 10 mL stainless-steel jar with one 10 mm stainless steel milling ball. The ball mill used was a MM400 shaker ball mill (RETSCH, Haan, Germany). DL-Serine measured at a magnetic-field strength of 16.4 T was milled for 20 min at 25 Hz in a 10 mL stainless-steel multi-cavity jar with three 7 mm stainless-steel milling balls. The ball mill used was a mixer mill MM500 nano (RETSCH, Haan, Germany). The L-serine monohydrate samples were prepared by mixing 50 mg of L-serine with 15 µL of de-ionized water in a 2 mL snap cap vial followed by processing in the resonant acoustic mixer LabRAM I (Resodyn Acoustic Mixers, Butte, Montana, USA) for 20 min at 100 g.| CAS number | Purity | Manufacturer | |
|---|---|---|---|
| L-Serine | 56-45-1 | 99% | abcr GmbH (Karlsruhe, Germany) |
| DL-Serine | 302-84-1 | 99% | abcr GmbH (Karlsruhe, Germany) |
| Water | — | De-ionized | — |
Solid-state NMR experiments
Solid-state NMR experiments were performed in 3.2 mm zirconia rotors with Vespel caps (Bruker BioSpin) in a 3.2 mm standard Bruker triple-resonance HXY probe on a 500 MHz Bruker NMR-spectrometer (AVANCE III HD console) as well as in a 3.2 mm standard Bruker double-resonance HX probe on a 700 MHz Bruker NMR-spectrometer (AVANCE NEO console). For each slow-spinning 1H–13C CP-MAS spectrum recorded with an MAS frequency of 2.0 kHz, a lineshape simulation using the software ssNake (version 1.5)48 was performed to determine the chemical-shift anisotropy parameters. All details of the performed experiments are given in Tables S1 and S2, SI.X-ray crystallography
The powder X-ray diffraction (PXRD) patterns were collected on a D8 Advance diffractometer (Bruker, Karlsruhe, Germany) operating in Bragg–Brentano geometry, using a flat circular sample holder with a sample thickness of 0.5 mm and a diameter of 10 mm. Cu-tube and a Ni-filter (Cu Kα radiation λ = 1.5406 Å) at a tube voltage of 40 kV together with a LYNXEYE semiconductor strip detector were used. Measurements were carried out at room temperature over a 2θ range of 10–60° (corresponding to a momentum transfer Q range of 0.71–4.1 Å−1), with a step size of 0.0105° and a counting time of 3 s per step. Structure refinements were performed using GSAS-II software (version 5609).49Computational details
DFT calculations were performed as implemented in the Vienna ab initio Simulation Package (VASP) (version 6.4.3).50–53 For the geometry optimizations using the conjugate gradient algorithm, an energy cutoff of 1 × 10−6 eV, Gaussian smearing with a width of 0.01 eV, and a regular 3 × 3 × 3 k-point mesh for integration in the Brillouin zone were chosen as convergence criteria after careful benchmarks, as can be seen in Fig. S1–S3.54 The Kohn–Sham orbitals were expanded in a plane-wave basis set with a kinetic energy cutoff of 500 eV. Different approaches for the exchange–correlation effects were applied, including the GGA PBE, meta-GGA TPSS, or hybrid functionals HSE06 and B3LYP.44,45,55–57 The DFT-D3 dispersion correction with the zero-damping function was used.58 Magnetic-shielding tensors were computed using the GIPAW method and PBE-functional. The isotropic chemical-shift values δ reported using VASP for nucleus in the molecule-of-interest correspond to the negative of the calculated magnetic-shielding values σ (δcalc,VASPiso ≈ −σcalc,VASPiso + σcalc,VASPiso,ref, with σcalc,VASPiso,ref representing the calculated isotropic magnetic-shielding value of the considered nucleus in a reference molecule, which was not further considered in this work).32,44,45,59 The “G = 0 including” chemical-shift values that consideronly the valence contribution are reported, with G = 0 taking the induced magnetic-field caused by the macroscopic magnetic-susceptibility tensor into account.60 Standard plane augmented wave (PAW) pseudopotentials were used for all atomic species.61The crystal structures of serine considered in this work were taken from the Cambridge Structural Database (CSD), with access codes LSERIN18, LSERMH10, and DLSERN11.62,63Table 2 provides an overview of the corresponding CSD IDs and original publications, showing which structure was used to represent the crystalline phases L-serine, L-serine monohydrate, and DL-serine, respectively.
If not reported otherwise, all three structures were geometrically optimized by allowing only the hydrogen atoms to move freely. For the calculation of NMR parameters, two distinct approaches were employed: (i) direct computation of magnetic-shielding tensors using the database structure without any relaxation with DFT and (ii) prior optimization of hydrogen-atom positions using various levels of theory, that is, various exchange correlation (XC) DFT functionals, while keeping the unit-cell parameters fixed. Specifically, we performed various calculations of the three structures and the different XC functionals without or with structural optimization. We refer to these as ‘no opt.’ or ‘X’ (X = functional), respectively. For comparison with the DFT calculations, we applied the machine learning (ML) model ShiftML3 to compute magnetic shielding tensors,36 using either the structures as directly obtained from the CSD or geometries resulting from hydrogen-atom optimization at the B3LYP level-of-theory using VASP.
Results and discussion
Solid-state NMR characterization of three distinct serine phases
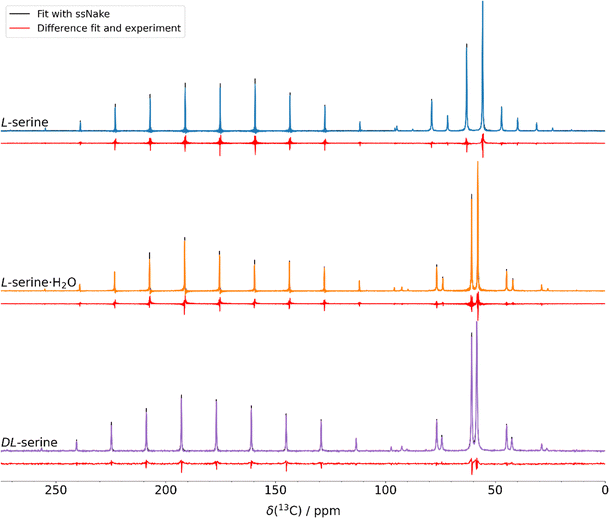
Upon studying mechanochemically induced molecular-recognition events, we came across the example of amino acids, for which ball milling or resonant acoustic mixing (RAM) of an equimolar ratio of the D- and L-amino acids provided the corresponding racemic phases (DL-amino acid), with the latter requiring liquid-assisted RAM conditions.39,41 This has been illustrated for the two examples of alanine and serine.39,41 Carbon-13 detected solid-state NMR spectroscopy has been used as the central analytical tool to distinguish such phases and to monitor the underlying solid-state molecular-recognition events.Fig. 1 shows the 1H–13C cross-polarization (CP) spectra we have reported previously,39,41 indeed showing that DL-serine and L-serine can be distinguished based on their characteristic spectral fingerprints. Chemical-shift differences of up to 3 ppm have been observed for the individual carbon atoms (for the isotropic 13C chemical-shift values, see Table 3). We also prepared the L-serine monohydrate sample (in the following denoted as L-serine·H2O) using a RAM device, whose Cα and Cβ chemical-shift values resemble those of DL-serine. The carboxylic acid group carbon atoms, though, allow an unambiguous distinction of the three phases. Herein, we also explore the accuracy of plane-wave periodic DFT calculations in distinguishing such phases in computer simulations, requiring accuracies in the calculated 13C isotropic chemical-shift values of around 1–2 ppm.
 | ||
| Fig. 1 1H–13C CP-MAS NMR spectra of DL-serine (purple, bottom), L-serine·H2O (orange, middle) and L-serine (blue, top) revealing distinct differences in the isotropic 13C chemical-shift values of the phases. All spectra have been recorded at a magnetic-field strength of 11.7 T and a MAS frequency of 17.0 kHz. L-Serine·H2O contains a small amount of L-serine. The spectra have already been published in our previous work, see ref. 39 and 41. The truncation visible in some of the spectra is caused by too short data acquisition times caused by probe limitations during high power 1H decoupling. In addition, representative structures of the different serine phases based on the single-crystal structures (CSD codes DLSERN11, LSERMH10 and LSERIN18 for DL-serine, L-serine·H2O and L-serine, respectively) are given. In the molecular stuctures hydrogen, carbon, oxygen, and nitrogen atoms are depicted in white, grey, red, and blue, respectively. | ||
| δ exp.iso (COO−)/ppm | δ exp.iso (Cα)/ppm | δ exp.iso (Cβ)/ppm | |
|---|---|---|---|
| L-Serine | 175.2 | 55.9 | 63.0 |
| L-Serine·H2O | 176.0 | 58.6 | 61.4 |
| DL-Serine | 177.1 | 58.7 | 61.0 |
A central goal of this work is to compare the anisotropy of the magnetic-shielding tensor between experiment and DFT calculations. The anisotropy reveals important information about the local chemical environment of the nucleus-of-interest,64,65 can be linked directly to electronic structures,66,67 even enabling the understanding of chemical reactivities (for a recent review article, see ref. 68), and can reveal insights into molecular dynamics.69 The magnetic-shielding interaction is described by a 3 × 3 tensor, which contains in the laboratory frame (LAB, in which the external magnetic field, B0, points along the z-axis) nine independent components (see eqn (1)). By using rotation matrices involving the Euler angles α, β, and γ, this matrix can be diagonalized in a molecule-fixed frame (the principal axis system, PAS) and three independent components remain:
 | (1) |
In solution, due to the fast Brownian motion, only the isotropic chemical-shift value (note that the chemical shift is obtained from the magnetic shielding by referencing to a standard compound) can be measured (which is 1/3 of the trace of the above reported tensor). In contrast to this, solid-state NMR allows the determination of all three principal-axis system components, for instance, by recording slow-spinning MAS spectra and simulating their lineshapes to envelopes of the static powder spectra70 or by recording multidimensional isotropic-anisotropic correlation experiments.71–74 In that vein, according to eqn (1), we determined the 13C chemical-shift anisotropy (CSA) by recording slow-spinning MAS experiments (employing an MAS frequency of 2.0 kHz) at two different magnetic-field strengths (for the spectra at 11.7 T, see Fig. 2, and the spectra at 16.4 T are provided in Fig. S4). Lineshape simulations allow the determination of the span (Ω) and skew (κ) parameters (using the Herzfeld–Berger convention23). The respective results are summarized in Table 4, reporting only the values for the carboxylic group carbon, which exhibits the highest CSA. Rather similar Ω-values of around 130 ppm have been observed for the three different serine phases, whereas the κ-values range between −0.04 and 0.22. The results for Cα and Cβ are displayed in Tables S3 and S4 of the SI, respectively.
| Ω exp. (COO−)/ppm | κ exp. (COO−) | |
|---|---|---|
| L-Serine | 130.8 [131.3] | −0.04 [−0.03] |
| L-Serine·H2O | 129.9 [n.d.] | 0.2 [n.d.] |
| DL-Serine | 135.8 [133.3] | 0.09 [0.09] |
We next attempted to establish a protocol for determining the agreement between the experimental data provided in Tables 3 and 4 and calculations of NMR magnetic-shielding tensors using DFT calculations under periodic-boundary conditions. As a matter of fact, finding a decent starting structure remains crucial for the calculations. We thus recorded the PXRD pattern of all three serine phases for which NMR spectra have been recorded. Crystal structures taken from the Cambridge Crystallographic database75 were used for comparing the experimental and back-predicted patterns (Fig. 3). For L-serine, the structure LSERIN1862 matches the diffraction pattern. For DL-serine, the best match has been observed for DLSERN11.63 For L-serine·H2O, a good agreement with the structure LSERMH10 has been found, taken into account its decomposition into L-serine over time.63 All samples exhibit strong texture effects, resulting in changes in relative peak intensities. We concluded that the identified crystal structures revealed good agreement with the experimental PXRD pattern and decided to use those geometries in the complementary DFT calculations. The hydrogen-atom positions, however, were optimized in most cases prior to the NMR parameter calculations, as will be outlined further below. Note that while the structure of L-serine has been determined by X-ray diffraction, the structures of DL-serine and L-serine·H2O were determined by neutron diffraction, already allowing us to assume more accurate hydrogen-atom positions (vide infra).
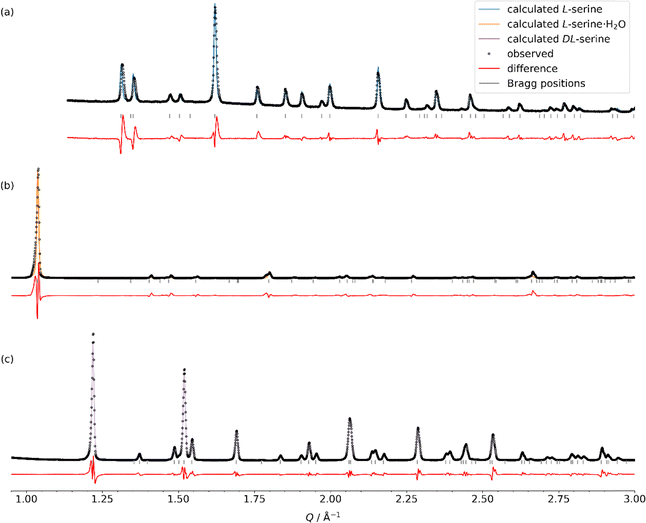 | ||
| Fig. 3 Comparison of experimental PXRD patterns of the three serine phases (a) L-serine, (b) L-serine·H2O and (c) DL-serine to the back-predicted ones using published single-crystal structures (see Table 2 for the corresponding CSD entries used). | ||
DFT-assisted calculation of solid-state NMR observables
After ensuring the accuracy of the database structures, we used these in periodic DFT calculations in order to extract the NMR observables. A comparison of the isotropic 13C chemical-shift values and the CSA parameters for the carboxylic acid group (COO−) is provided in Fig. 4 for the different DFT exchange correlation functionals used for optimizing the hydrogen-atom positions prior to the calculation of NMR observables. Specifically, this figure provides the comparison between experimental and calculated isotropic 13C chemical-shift values for the difference in the chemical-shift values between the carboxyl acid group carbon and Cα (Fig. 4a), between the carboxyl acid group carbon and Cβ (Fig. 4b) and between Cβ and Cα (Fig. 4c) for the three different serine phases introduced above. These differences serve as a measure for the deviation between the experiment and plane-wave DFT calculations using various level-of-theories for the optimization of the hydrogen-atom positions prior to the magnetic-shielding tensor calculations. The data reveal that based on the calculations without structural relaxation, each crystal structure exhibited an overall distinct deviation from the experimental chemical-shift differences of up to 4.7 ppm for L-serine (LSERIN18), with DL-serine (DLSERN11) demonstrating the best match for the three isotropic chemical-shift differences (0.2 ppm for COO−–Cα, −1.2 ppm for COO−–Cβ and 1.4 ppm for Cβ–Cα).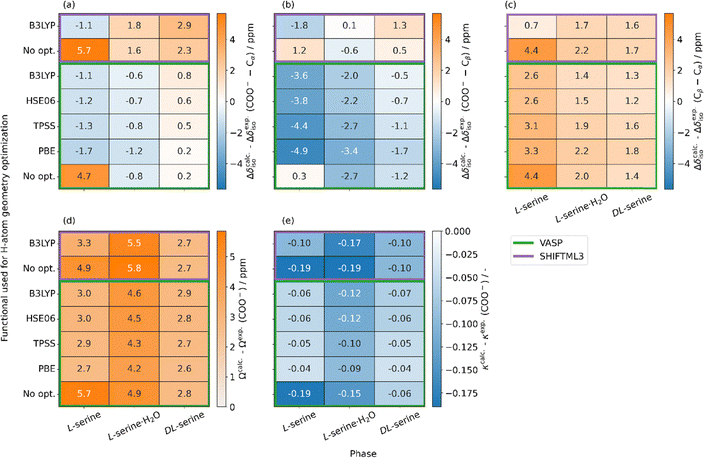 | ||
| Fig. 4 “Heat maps” comparing experimental and DFT-calculated isotropic 13C chemical-shift values. The differences between experimental and DFT-calculated values for the difference in the chemical-shift values between the carboxyl acid group carbon and Cα (a), between the carboxyl acid group carbon and Cβ (b) and between Cβ and Cα (c) for the three different serine phases are plotted in units of ppm. “Heat maps” of the CSA parameters Ω (d) and κ (e) only for COO− as a function of the DFT functional. In all cases (except no opt.) the hydrogen-atom positions were geometrically optimized in VASP before the calculation of the 13C magnetic-shielding tensors using different DFT functionals, as indicated in the figure. In VASP (highlighted by the green lines), calculations of 13C magnetic-shielding tensors have been performed in all cases with the PBE DFT functional. As comparison and separated by the purple lines, the results obtained by using the machine learning model ShiftML3 are shown.36 | ||
In general, the calculated chemical-shift difference between COO− and Cα was found to be in better agreement with the experimental data than the corresponding difference between COO− and Cβ and between Cβ and Cα. For calculations involving the relaxation of the hydrogen-atom positions only, the deviation between calculated and experimental chemical-shift differences varied depending on the level-of-theory used during the optimization. Except for DL-serine, the best agreement with experiment was achieved for hybrid functionals, such as B3LYP. The minimal deviations between computation and the experiment for the Cβ–Cα chemical-shift difference have been determined to 2.6 ppm for L-serine (B3LYP), 1.4 ppm for L-serine·H2O (B3LYP) and 1.2 ppm for DL-serine (HSE06). As noted above, smaller deviations ranging between 0.2 and −1.1 ppm for the minimal values are obtained for the COO−–Cα difference. Larger deviations involving Cβ chemical-shift values are attributed to higher uncertainties in their DFT-computed isotropic chemical-shift value, possibly as a consequence of intermolecular hydrogen-bond formation of the neighbored hydroxyl group. The overall better agreement for DL-serine and L-serine·H2O compared to L-serine might point to the more accurate refinement of hydrogen-atom positions via neutron diffraction, leading to structures of higher quality than in the case of L-serine. These results highlight the crucial role of hydrogen-atom positioning in accurately predicting the isotropic 13C chemical-shift differences, as already indicated in previous studies (for some examples, see ref. 28–30, 34, 35, and 76–78). For instance, 13C magnetic-shielding tensor GIPAW calculations for α-glycine using different crystal structures revealed the best agreement with experimental values in most cases by only geometrically optimizing the hydrogen atom positions, instead of an all-atom optimization.76 Whether a full geometry optimization even improves the agreement with experimental chemical-shift values further is under debate (as, for instance, such calculations would yield the geometry only at 0 K) and should be decided on a case-to-case basis as proposed in ref. 30. In the case of the serine phase studied herein, optimizing the hydrogen-atom positions seems to be sufficient,77 as full geometry optimization even leads to larger deviations from experimental values (vide infra).
We have also compared our experimental values with those obtained by the ML model ShiftML3, in which a ML model was trained on a database including periodic DFT-calculated magnetic-shielding tensors achieved at the PBE level-of-theory.36 Similar to the observations for the DFT calculations, the deviations to the experimental values decrease for the geometry-optimized structures (Fig. 4a, b and c), pointing again to the importance of an accurate structure. Interestingly, the DFT calculated and the ML predicted data in this Fig. 4 show a different relationship relative to the experimental data. Specifically, the DFT calculations match better the experimental measurements in the case of the COO−–Cα differences, while the ML model better captures the COO−–Cβ differences. Both methods reveal a similar accuracy for the Cβ–Cα differences. Taking the low computational cost of ShiftML3 into consideration, though, this algorithm already pinpoints the power of ML in predicting 13C magnetic-shielding tensors and it will probably even further improve with the increase of the accuracy in DFT-calculations of magnetic-shielding tensors (and thus the training data sets).
We next turn to the anisotropy of the 13C magnetic-shielding tensors of the three serine phases. As mentioned above, only the carboxylic acid carbon is analyzed as the span-value is largest in magnitude and thus determined with much higher precision experimentally. The comparison in Ω- (Fig. 4c) and κ-values (Fig. 4d) between calculation and experiment, again using different DFT-functionals for the geometry optimization of the hydrogen-atom positions, reveals in all cases an overestimation of the Ω-values with the largest deviations of 4.9 and 5.7 ppm for L-serine·H2O and L-serine without initial constrained geometry optimization. While the deviations in Ω-values between calculation and experiment reduce significantly for L-serine upon optimizing the hydrogen atom positions (minimal deviation of 2.7 ppm, PBE), the values for L-serine·H2O and DL-serine do not change significantly (minimal deviations of 4.2 and 2.6 ppm, respectively). A similar trend is observed for the κ-values with minimal deviations of −0.04 (PBE), −0.09 (PBE) and −0.04 (PBE) for L-serine, L-serine·H2O and DL-serine, respectively. Interestingly, the PBE GGA-functional revealed the best agreement between simulation and experiment in all cases. ShiftML3, as in the case of the DFT, overestimates the Ω-values, whereas for the κ-values in all cases, larger deviations are obtained by such ML-approaches as compared to the DFT calculations. Note that a geometry optimization allowing all atoms in the molecules to relax (PBE level-of-theory) provides a poorer agreement with nearly all of the respective experimental NMR observables in our cases instead of optimizing the hydrogen atoms only (see Fig. S5, S8).
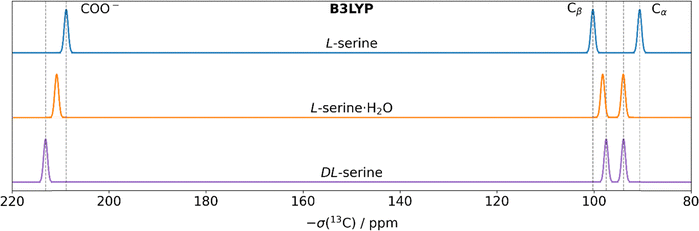
Fig. 5 shows schematically the 13C NMR spectra for all three serine phases using the DFT-calculated isotropic magnetic-shielding values obtained after optimizing the hydrogen atoms at a B3LYP level-of-theory (for the further DFT functionals employed herein, see Fig. S6). When compared with the experimental spectra in Fig. 1, the computed 13C chemical-shift values of the carboxylic acid group carbon atom reproduce the experimental order, with DL-serine having the highest chemical-shift value and L-serine the lowest. The same is observed for the Cβ–Cα chemical-shift difference, with L-serine showing the largest value experimentally and DL-serine the smallest. Even the small deviations observed experimentally between L-serine·H2O and DL-serine are qualitatively reproduced correctly. Although perfect reproduction of the experimental data is not possible (i.e. the differences in carboxyl group carbon chemical-shift values are overestimated; see Fig. 5), the three distinct serine phases can be qualitatively distinguished by means of DFT calculations. The same holds for the ShiftML3 predictions (Fig. S7). Note, though, that an optimization of the hydrogen atoms is required prior to the calculation of NMR observables, as discussed above (for the simulated spectra after full geometry optimization, see Fig. S8).
Computational time
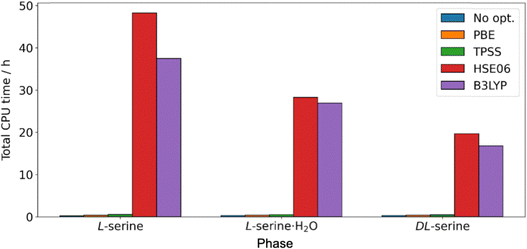
As a final remark on the DFT computations of NMR observables, we assess the computational demand of these, especially with respect to their accuracy as observed in Table 5 and Fig. 6. To this end, we compare in Fig. 6 the total computational times needed for performing the respective NMR calculations as well as geometry optimizations of hydrogen-atom positions in VASP. We focus only on the relaxation part, as the computation of the NMR observables was performed at the same level corresponding computational times on the order of 20 min (using 36 CPU cores). Intuitively, the most complex and non-local HSE06 reflected an almost 50 time increase in time than the case without structural optimization for the semi-local PBE functional. A comparison of these trends to the accuracy shown in Fig. 6 clearly reveals that the longer computational times for the structural optimization do not compensate in the accuracy of the NMR observables. Accordingly, the assessment of all these results points to the use of less complex and computationally less demanding exchange correlation functionals for DFT for the calculations of the NMR observables, provided the use of accurate and high-quality initial structures. For the calculation of magnetic-shielding tensors using ShiftML3, executing the Python script for the six magnetic-shielding tensors reported herein required a computational time of only 26.6 s (using 36 CPU cores). ShiftML3 was found to be approximately 271 times faster than calculating NMR observables with DFT. The very low computational cost thus makes ShiftML3 well suited for fast initial benchmarks, but still needs to occasionally be enhanced by additional insights from the DFT-calculated observables.Conclusions
Computer simulations within the DFT framework using periodic boundary conditions were compared against experimentally derived solid-state MAS NMR 13C magnetic-shielding tensors for three serine phases, namely, L- and DL-serine, as well as L-serine·H2O. Such phases can be distinguished experimentally based on 13C isotropic chemical-shift differences, which are, however, small, on the order of only 2–3 ppm. Our test study of this challenging system reveals the importance of optimizing the hydrogen-atom positions prior to the calculations of the NMR parameters, which is key particularly for crystal structures derived from X-ray diffraction, in which the hydrogen-atom positions remain less well defined. Using this protocol, the experimental trends in isotropic 13C magnetic-shielding values are reproduced correctly in the calculations, enabling an unambiguous distinction of the three phases. Similar conclusions hold for the anisotropy parameters, for which hydrogen-atom position optimization improves the agreement with experiment as well. Our work underlines that the use of a rather low-cost DFT exchange correlation functional for the constrained geometry optimizations already reveals decent agreement with experimental data, of similar magnitude to ShiftML3 predictions. For some observables, such as the COO−–Cα differences and the skew κ, DFT still shows better agreement with the experiment than the ML model, with a higher computational load. Indeed, the use of more complex semi-local or non-local exchange correlation functionals for structural optimization slightly enhances the accuracy, but at the same time significantly increases the computational cost. Most importantly, the insights obtained from this comparative study and the thereby proposed calculation protocol provide the means for efficiently complementing experimental efforts in fingerprinting structural polymorphs and emphasize the applicability of periodic plane-wave DFT simulations.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information is available. See DOI: https://doi.org/10.1039/d5cp04690d.The scripts and data used in this manuscript will be made available through a public Radar4Chem repository hosted at DOI: https://doi.org/10.22000/7022jhfh8mwq5tyk.
Acknowledgements
This research was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany's Excellence Strategy – Exzellenzcluster 2186 “The Fuel Science Center” (ID: 390919832). T. W. acknowledges further support from the DFG (project number 455240421 and Heisenberg fellowship, project number 455238107) and the Max Planck Society. K. R. is grateful to Sabrina Smid and Longlong Li for assisting with technical issues of the simulations and Dr Ridvan Ince for assisting with the ssNake analysis. The computing time provided in the NHR Center NHR4CES at RWTH Aachen University (project number p0024945) is greatly acknowledged. This study was also funded by the Federal Ministry of Education and Research and the state governments participating on the basis of the resolutions of the GWK for national high-performance computing at universities. Open Access funding provided by the Max Planck Society.References
- E. H. Lee, Asian J. Pharm. Sci., 2014, 9, 163–175 Search PubMed.
- Q. Shi, H. Chen, Y. Wang, J. Xu, Z. Liu and C. Zhang, Int. J. Pharm., 2022, 611, 121320 CrossRef CAS PubMed.
- P. Hodgkinson, Prog. Nucl. Magn. Reson. Spectrosc., 2020, 118–119, 10–53 Search PubMed.
- R. K. Harris, R. E. Wasylishen and M. J. Duer, NMR Crystallography, Wiley, 2012 Search PubMed.
- B. Elena and L. Emsley, J. Am. Chem. Soc., 2005, 127, 9140–9146 CrossRef CAS PubMed.
- C. J. Pickard, E. Salager, G. Pintacuda, B. Elena and L. Emsley, J. Am. Chem. Soc., 2007, 129, 8932–8933 CrossRef CAS PubMed.
- F. Taulelle, Solid State Sci., 2004, 6, 1053–1057 Search PubMed.
- J. Senker, L. Seyfarth and J. Voll, Solid State Sci., 2004, 6, 1039–1052 CrossRef CAS.
- P. M. J. Szell, S. O. Nilsson Lill, H. Blade, S. P. Brown and L. P. Hughes, Solid State Nucl. Magn. Reson., 2021, 116, 101761 CrossRef CAS PubMed.
- L. Emsley, Faraday Discuss., 2025, 255, 9–45 RSC.
- A. Hofstetter and L. Emsley, J. Am. Chem. Soc., 2017, 139, 2573–2576 CrossRef CAS PubMed.
- T. Wiegand, D. Ludeker, G. Brunklaus, K. Bussmann, G. Kehr, G. Erker and H. Eckert, Dalton Trans., 2014, 43, 12639–12647 Search PubMed.
- R. K. Harris, Analyst, 2006, 131, 351–373 Search PubMed.
- J. Thun, L. Seyfarth, J. Senker, R. E. Dinnebier and J. Breu, Angew. Chem., Int. Ed., 2007, 46, 6729–6731 Search PubMed.
- M. R. Chierotti and R. Gobetto, CrystEngComm, 2013, 15, 8599–8612 Search PubMed.
- A. Abraham, D. C. Apperley, T. Gelbrich, R. K. Harris and U. J. Griesser, Can. J. Chem., 2011, 89, 770–778 Search PubMed.
- H. Hamaed, J. M. Pawlowski, B. F. T. Cooper, R. Fu, S. H. Eichhorn and R. W. Schurko, J. Am. Chem. Soc., 2008, 130, 11056–11065 CrossRef CAS PubMed.
- S. A. Southern and D. L. Bryce, in Annual Reports on NMR Spectroscopy, ed. G. A. Webb, Academic Press, 2021, vol. 102, pp. 1–80 Search PubMed.
- R. Quiñones, R. J. Iuliucci, G. Behnke, R. Brown, D. Shoup, T. M. Riedel, C. Plavchak, B. E. Lininger and J. M. Spehar, J. Pharm. Biomed. Anal., 2018, 148, 163–169 CrossRef PubMed.
- E. Chaloupecká, V. Tyrpekl, K. Bártová, Y. Nishiyama and M. Dračínský, Solid State Nucl. Magn. Reson., 2024, 130, 101921 CrossRef PubMed.
- Z. Rehman, J. Lubay, W. T. Franks, A. P. Bartók, E. K. Corlett, B. Nguyen, G. Scrivens, B. M. Samas, H. Frericks-Schmidt and S. P. Brown, Faraday Discuss., 2025, 255, 222–243 RSC.
- Y. Dai, V. Terskikh, A. Brinmkmann and G. Wu, Cryst. Growth Des., 2020, 20, 7484–7491 CrossRef CAS.
- J. Herzfeld and A. E. Berger, J. Chem. Phys., 1980, 73, 6021–6030 CrossRef CAS.
- R. K. Harris, J. Pharm. Pharmacol., 2007, 59, 225–239 CrossRef CAS PubMed.
- P. E. Hansen, Molecules, 2024, 29, 336 CrossRef CAS PubMed.
- B. I. Dunlap and N. Rosch, in Adv. Quantum Chem., ed. P.-O. Löwdin, Academic Press, 1990, vol. 21, pp. 317–339 Search PubMed.
- M. Dračínský, in Modern NMR Crystallography: Concepts and Applications, ed. D. L. Bryce, Royal Society of Chemistry, 2025, vol. 36 Search PubMed.
- A. L. Reviglio, F. A. Martínez, M. D. A. Montero, Y. Garro-Linck, G. A. Aucar, N. R. Sperandeo and G. A. Monti, RSC Adv., 2021, 11, 7644–7652 RSC.
- E. K. Corlett, H. Blade, L. P. Hughes, P. J. Sidebottom, D. Walker, R. I. Walton and S. P. Brown, CrystEngComm, 2019, 21, 3502–3516 RSC.
- J. R. Yates, S. E. Dobbins, C. J. Pickard, F. Mauri, P. Y. Ghi and R. K. Harris, Phys. Chem. Chem. Phys., 2005, 7, 1402–1407 RSC.
- C. Poidevin, G. L. Stoychev, C. Riplinger and A. A. Auer, J. Chem. Theory Comput., 2022, 18, 2408–2417 Search PubMed.
- C. J. Pickard and F. Mauri, Phys. Rev. B:Condens. Matter Mater. Phys., 2001, 63, 245101 Search PubMed.
- C. Bonhomme, C. Gervais, F. Babonneau, C. Coelho, F. Pourpoint, T. Azaïs, S. E. Ashbrook, J. M. Griffin, J. R. Yates, F. Mauri and C. J. Pickard, Chem. Rev., 2012, 112, 5733–5779 CrossRef CAS PubMed.
- S. E. Ashbrook and D. McKay, Chem. Commun., 2016, 52, 7186–7204 Search PubMed.
- T. Charpentier, Solid State Nucl. Magn. Reson., 2011, 40, 1–20 Search PubMed.
- M. Kellner, J. B. Holmes, R. Rodriguez-Madrid, F. Viscosi, Y. Zhang, L. Emsley and M. Ceriotti, J. Phys. Chem. Lett., 2025, 16, 8714–8722 CrossRef CAS PubMed.
- F. M. Paruzzo, A. Hofstetter, F. Musil, S. De, M. Ceriotti and L. Emsley, Nat. Commun., 2018, 9, 4501 CrossRef PubMed.
- E. Chaloupecká, O. Socha and M. Dračínský, Solid State Nucl. Magn. Reson., 2025, 138, 102019 Search PubMed.
- C. Quaranta, I. d'Anciães Almeida Silva, S. Moos, E. Bartalucci, L. Hendrickx, B. M. D. Fahl, C. Pasqualini, F. Puccetti, M. Zobel, C. Bolm and T. Wiegand, Angew. Chem., Int. Ed., 2024, 63, e202410801 CAS.
- A.-A. Lafrance, M. Girard and D. L. Bryce, Solid State Nucl. Magn. Reson., 2024, 101925 CrossRef CAS PubMed.
- L. Hendrickx, C. Quaranta, E. Fuchs, M. Plekhanov, M. Zobel, C. Bolm and T. Wiegand, Molecules, 2025, 30, 3745 CrossRef CAS PubMed.
- T. Kameda and H. Teramoto, Magn. Reson. Chem., 2006, 44, 318–324 CrossRef CAS PubMed.
- M. Dračínský, J. Vícha, K. Bártová and P. Hodgkinson, ChemPhysChem, 2020, 21, 2075–2083 CrossRef PubMed.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 Search PubMed.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1997, 78, 1396 Search PubMed.
- M. Dračínský, P. Unzueta and G. J. O. Beran, Phys. Chem. Chem. Phys., 2019, 21, 14992–15000 Search PubMed.
- S. A. Ramos, L. J. Mueller and G. J. O. Beran, Faraday Discuss., 2025, 255, 119–142 Search PubMed.
- S. G. J. van Meerten, W. M. J. Franssen and A. P. M. Kentgens, J. Magn. Reson., 2019, 301, 56–66 Search PubMed.
- B. H. Toby and R. B. Von Dreele, J. Appl. Crystallogr., 2013, 46, 544–549 Search PubMed.
- G. Kresse and J. Furthmüller, Phys. Rev. B:Condens. Matter Mater. Phys., 1996, 54, 11169–11186 Search PubMed.
- G. Kresse and J. Furthmüller, Comput. Mater. Sci., 1996, 6, 15–50 CrossRef CAS.
- G. Kresse and D. Joubert, Phys. Rev. B:Condens. Matter Mater. Phys., 1999, 59, 1758–1775 CrossRef CAS.
- G. Kresse and J. Hafner, Phys. Rev. B:Condens. Matter Mater. Phys., 1993, 47, 558–561 Search PubMed.
- W. S. Morgan, J. E. Christensen, P. K. Hamilton, J. J. Jorgensen, B. J. Campbell, G. L. W. Hart and R. W. Forcade, Comput. Mater. Sci., 2020, 173, 109340 Search PubMed.
- J. Tao, J. P. Perdew, V. N. Staroverov and G. E. Scuseria, Phys. Rev. Lett., 2003, 91, 146401 Search PubMed.
- A. V. Krukau, O. A. Vydrov, A. F. Izmaylov and G. E. Scuseria, J. Chem. Phys., 2006, 125, 224106 Search PubMed.
- P. J. Stephens, F. J. Devlin, C. F. Chabalowski and M. J. Frisch, J. Phys. Chem., 1994, 98, 11623–11627 Search PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 Search PubMed.
- J. R. Yates, C. J. Pickard and F. Mauri, Phys. Rev. B:Condens. Matter Mater. Phys., 2007, 76, 024401 CrossRef.
- G. A. de Wijs, R. Laskowski, P. Blaha, R. W. A. Havenith, G. Kresse and M. Marsman, J. Chem. Phys., 2017, 146, 064115 CrossRef CAS PubMed.
- P. E. Blöchl, Phys. Rev. B:Condens. Matter Mater. Phys., 1994, 50, 17953–17979 CrossRef PubMed.
- E. V. Boldyreva, E. N. Kolesnik, T. N. Drebushchak, H. Ahsbahs, J. A. Beukes and H.-P. Weber, Z. Kristallogr. – Cryst. Mater., 2005, 220, 58–65 Search PubMed.
- M. N. Frey, M. S. Lehmann, T. F. Koetzle and W. C. Hamilton, Acta Crystallogr., Sect. B:Struct. Sci., 1973, 29, 876–884 Search PubMed.
- M. Hong, J. Am. Chem. Soc., 2000, 122, 3762–3770 CrossRef CAS.
- Z. Wu, N. Tjandra and A. Bax, J. Am. Chem. Soc., 2001, 123, 3617–3618 Search PubMed.
- A. Venkatesh, D. Gioffrè, B. A. Atterberry, L. Rochlitz, S. L. Carnahan, Z. Wang, G. Menzildjian, A. Lesage, C. Copéret and A. J. Rossini, J. Am. Chem. Soc., 2022, 144, 13511–13525 Search PubMed.
- J. Koppe, A. V. Yakimov, D. Gioffre, M. E. Usteri, T. Vosegaard, G. Pintacuda, A. Lesage, A. J. Pell, S. Mitchell, J. Perez-Ramirez and C. Coperet, Nature, 2025, 642, 613–619 CrossRef CAS PubMed.
- Z. J. Berkson, W. Cao, D. Gioffrè, C. J. Kaul, L. Lätsch, Y. Kakiuchi, A. Yakimov and C. Copéret, JACS Au, 2025, 5, 2911–2931 CrossRef CAS PubMed.
- F. Blanc, J.-M. Basset, C. Copéret, A. Sinha, Z. J. Tonzetich, R. R. Schrock, X. Solans-Monfort, E. Clot, O. Eisenstein, A. Lesage and L. Emsley, J. Am. Chem. Soc., 2008, 130, 5886–5900 CrossRef CAS PubMed.
- M. M. Maricq and J. S. Waugh, J. Chem. Phys., 1979, 70, 3300–3316 CrossRef CAS.
- W. T. Dixon, J. Chem. Phys., 1982, 77, 1800–1809 CrossRef CAS.
- Z. Gan, J. Am. Chem. Soc., 1992, 114, 8307–8309 CrossRef CAS.
- O. N. Antzutkin, S. C. Shekar and M. H. Levitt, J. Magn. Reson., Ser. A, 1995, 115, 7–19 CrossRef CAS.
- S. L. Gann, J. H. Baltisberger and A. Pines, Chem. Phys. Lett., 1993, 210, 405–410 CrossRef CAS.
- F. Allen, Acta Crystallogr., Sect. B:Struct. Sci., 2002, 58, 380–388 CrossRef PubMed.
- Ł. Szeleszczuk, D. M. Pisklak and M. Zielińska-Pisklak, J. Comput. Chem., 2018, 39, 853–861 CrossRef PubMed.
- R. K. Harris, P. Hodgkinson, C. J. Pickard, J. R. Yates and V. Zorin, Magn. Reson. Chem., 2007, 45, S174–S186 Search PubMed.
- D. H. Brouwer, K. P. Langendoen and Q. Ferrant, Can. J. Chem., 2011, 89, 737–744 CrossRef.
| This journal is © the Owner Societies 2026 |