 Open Access Article
Open Access ArticleNavigating in the chemical space of peptides: computational strategies and molecular features to unveil their functional and drug-like properties
Ewerton
Cristhian Lima de Oliveira
 a,
Juliana
Auzier
a,
Juliana
Auzier
 b,
Gabriel
Pereira Coelho
b,
Gabriel
Pereira Coelho
 c,
Lidiane Diniz
do Nascimento
c,
Lidiane Diniz
do Nascimento
 c,
Anderson Henrique
Lima e Lima
c,
Anderson Henrique
Lima e Lima
 d,
Caio Marcos Flexa
Rodrigues
d,
Caio Marcos Flexa
Rodrigues
 c,
Anton
De Spiegeleer
c,
Anton
De Spiegeleer
 fg,
Evelien
Wynendaele
fg,
Evelien
Wynendaele
 eg,
Claudomiro
Sales
eg,
Claudomiro
Sales
 b,
Bart
De Spiegeleer
b,
Bart
De Spiegeleer
 *eg and
Kauê
Santana
*eg and
Kauê
Santana
 *c
*c
aInstituto Tecnológico Vale, 66055-090 Belém, Pará, Brasil
bLaboratório de Inteligência Computacional e Pesquisa Operacional, Instituto de Tecnologia, Universidade Federal do Pará, Campos Belém, 66075-110 Belém, Pará, Brasil
cLaboratório de Simulação Computacional, Instituto de Biodiversidade, Universidade Federal do Oeste do Pará, Campus Santarém, Santarém, Pará, 68.040-070, Brasil. E-mail: kaue.costa@ufopa.edu.br
dLaboratório de Planejamento e Desenvolvimento de Fármacos, Instituto de Ciências Exatas e Naturais, Universidade Federal do Pará, Belém, Pará 66075-110, Brasil
eDrug Quality and Registration (DruQuaR) group, Faculty of Pharmaceutical Sciences, Ghent University, Ottergemsesteenweg 460, B-9000 Ghent, Belgium. E-mail: Bart.DeSpiegeleer@ugent.be
fDepartment of Geriatrics, Faculty of Medicine and Health Sciences, Ghent University Hospital, Ghent, Belgium
gTranslational Research in Immunosenescence, Gerontology and Geriatrics (TRIGG) group, Ghent University Hospital, Ghent, Belgium
First published on 6th April 2026
Abstract
Peptides, short chains of amino acids linked by peptide bonds, typically ranging from 2 to 50 residues, are fundamental to diverse biological processes and represent a valuable source for the development of novel bioactive compounds. In this work, we provide a comprehensive and conceptual overview of approaches to exploring the peptide chemical space. We emphasize intrinsic challenges in their chemical space investigation, particularly the complex interplay among peptide conformation, bioactivity, and bioavailability, as well as the role of sequence- and structure-derived molecular features in elucidating structure–activity relationships. Furthermore, we examine computational strategies, such as dimensionality reduction techniques, machine learning models, and similarity-based complex networks for classifying and characterizing this chemical space. Finally, we underscore the importance of interdisciplinary frameworks in advancing peptide research, highlighting how integrative approaches can uncover intersections of bioactivity across different peptide classes and leverage alternative chemical spaces to optimize and characterize peptide structures.
Professor Kauê Santana da Costa is an Adjunct Professor at the Federal University of Western Pará (UFOPA) (H-index: 16, ORCID: https://orcid.org/0000-0002-2735-8016, Web of Science: https://www.webofscience.com/wos/author/record/1851974), where he coordinates the Laboratory of Computational Simulation & Scientific Education and leads the Interdisciplinary Group for the Application and Development of Biomolecular Technologies. His research intersects theoretical and computational chemistry, bioinformatics, and artificial intelligence, with a particular focus on self-assembling, quorum-sensing activity, and membrane-penetrating peptides for nanocarrier and biotechnological applications. Over the last years, Professor Costa has become a reference in the development of machine-learning models for peptide bioactivity, especially cell-penetrating and blood–brain–barrier-penetrating peptides. He is a co-author of the Scientific Reports article “Predicting cell-penetrating peptides using machine learning algorithms and navigating in their chemical space” (2021), which introduced supervised models to classify CPPs and explore their chemical space. Building on this line, he contributed to the comprehensive review “Biological Membrane-Penetrating Peptides: Computational Prediction and Applications” in Frontiers in Cellular and Infection Microbiology (2022), and to “BrainPepPass: A Framework Based on Supervised Dimensionality Reduction for Predicting Blood–Brain Barrier-Penetrating Peptides” in Journal of Chemical Information and Modeling (2023), which couples supervised dimensionality reduction with peptide classification. Most recently, he co-authored the deep-learning study “Investigating molecular descriptors in cell-penetrating peptides prediction with deep learning: Employing N, O, and hydrophobicity according to the Eisenberg scale” in PLOS One (2024), refining descriptor selection for peptide prediction, and the ACS Omega article “Beyond Molecular Weight: Peptide Characteristics Influencing the Sensitivity of Retention to Changes in Organic Solvent in Reversed-Phase Chromatography” (2025), which deepens the understanding of peptide physicochemical behavior. He also coordinates the PepSpace, an international collaborative project that aims to map and organize the chemical space of peptides. The web server is under development, integrating supervised and unsupervised learning to analyze the chemical space and bioactivity of quorum-sensing, cell-penetrating, and blood–blood–brain–barrier-penetrating peptides, consolidating his role at the interface of peptide science and machine learning. |
Introducing the main concepts of chemical spaces and their applications to peptide science
The term ‘chemical space’ is used in two complementary ways. In a broad conceptual sense, it refers to the theoretical universe of chemically feasible molecules. In practical chemoinformatics, however, chemical space is operationally defined for a given dataset by representing molecules as vectors in a multidimensional descriptor or fingerprint space, where inter-compound distances reflect similarity relationships within the chosen representation.1,2 The choice of representation, i.e., how the peptides are encoded into a form that computers can handle, influences which analyses are efficiently possible.3 These molecular descriptors or fingerprints encode structural and physicochemical properties, enabling visualization (often via 2D/3D projections) and analysis of structure–property/activity relationships relevant to bioactivity, encompassing pharmacodynamics and pharmacokinetics.4 In this work, we use ‘chemical space’ primarily in this operational, dataset-driven sense, as it enables visualization and analysis of structure–property/activity relationships for peptide sets.Determining the chemical space of molecules aims to classify compounds, identify potential bioactive molecules, design and improve lead candidates, and understand their molecular properties.5,6 Independent of the method, mapping the chemical space can significantly enhance the efficiency of novel discoveries across various areas of chemistry, including chemical synthesis,7–9 quantum chemistry,10 materials science,11–13 and drug discovery.5,14,15
The chemical space of compounds can be explored by analyzing libraries of organic molecules using two-dimensional (2D) or three-dimensional (3D) visual representations of multidimensional descriptor spaces plotted in Cartesian coordinates, which often require dimensionality-reduction or clustering techniques.16,17
Fig. 1 illustrates the application of a computational workflow to map the chemical space of peptides. Chemical space mapping integrates heterogeneous peptide inputs—either sequence-based representations (primary structure) or 3D structural models (tertiary structure)—to derive informative molecular descriptors. The visual exploration of peptide chemical space can be performed using computational tools that efficiently provide a visual correlation between molecules exhibiting similar chemical and/or functional properties.18–20 The most straightforward approach to accomplish this task involves the development of pipelines that integrate: the calculation of molecular descriptors using chemical packages, such as RDKit,21 iFeature,22,23 and Mordred;24 the application of feature correlations tests; and finally clustering methods, such as k-means and density-based spatial clustering of applications with noise (DBSCAN); as well as dimensionality reduction techniques including t-distributed stochastic neighbor embedding (t-SNE)25 and uniform manifold approximation and projection (UMAP);26 and finally the graphical representations for the 2D and 3D visualization of the numerical representations generated from these projections.18,20
Several open-source computational tools have been developed to facilitate the exploration of molecular chemical space based on molecular descriptor calculations. ChemPlot is a Python-based tool designed for both static and interactive visualization of chemical space in molecular datasets, encompassing dimensionality reduction techniques and similarity computations.27 TMAP is a tool developed for Python and employed to visualize high-dimensional chemical space by organizing molecules into a minimum spanning tree structure through molecular fingerprint comparisons.28 Similarly, KNIME, an open-source software developed for data science and visual analytics, in which computational workflows are structured as flowchart-based pipelines, has dedicated extensions to support chemical space visualization and other cheminformatics applications.29
Fig. 2 illustrates an example of how the chemical space of blood–brain barrier penetrating peptides (B3PPs) and quorum-sensing peptides (QSPs) can be represented in a 2D chart with the results of the dimensionality reduction provided by PCA and the clustering of the peptides predicted by the k-means algorithm.
In contrast to coordinate-based representations, chemical space networks (CSNs) have been introduced for chemical space analyses, allowing the exploration of molecular properties without reducing dimensionality.30–33 Various similarity-based complex networks, including half-space proximal networks (HSPNs), metadata networks, and CSNs, have been utilized to study the bioactivity of compounds and their associated chemical space. Fig. 3 represents an overview of the applications of these methods in peptide science.
The similarity-based complex networks are graphical representations of the chemical space of peptides, where nodes represent the peptides and the edges between two nodes denote their pairwise similarity or dissimilarity relationships in the space.34 The distance between the compounds is often measured using similarity (or dissimilarity) distance metrics, such as Euclidean, Manhattan, Tanimoto, and Soergel coefficients. In these networks, the relevance of the elements is investigated using centrality measures (betweenness, closeness, and edge betweenness), as well as global network properties and their corresponding global measures, such as modularity, connectivity, density, and size.35,36 For example, the StarPep Toolbox is a platform to explore the chemical space of antimicrobial peptides (AMPs) through molecular network-based representations and similarity-search methods to support peptide drug repurposing, as well as the development and optimization of novel sequences.37 Recently, antiviral peptides (AVPs) were mapped into a chemical space using HSPNs and contextualized with metadata networks using the StarPep toolbox. The analyses revealed eight chemically distinct, biologically coherent AVP communities without fixed similarity thresholds and linking them to origins, functions, and viral targets through metadata networks. The authors performed a centrality-guided scaffold extraction, which revealed four non-redundant subsets suitable for modeling and multi-query searches. The mapping of motifs against non-AVP datasets indicated that motif burden correlates with higher predicted AVP probabilities, with peptides carrying four to five motifs achieving the highest scores across independent predictors, suggesting that the motif-driven design is an interesting strategy to expand AVP chemical space.33
Recently, similarity-based complex networks and machine-learning algorithms have been used to map the landscape of some classes of peptides.38–40 Half-space proximal networks, metadata networks, and chemical space networks are examples of computational methods that leverage graph theory to analyze and explore relationships among chemical entities based on a given molecular property.41,42 These methods aim to simplify and analyze the vast complexity of chemical data, associating this information with the desired properties. For example, Ayala-Ruano et al. (2022) used network analyses and similarity-guided screening to investigate the chemical space of antiparasitic peptides, emphasizing the challenge of discovering new therapeutic peptides from this vast chemical space. The authors combined HSPNs, CSNs, and metadata networks to identify central peptides and to perform multi-query similarity searches against the StarPepDB database. Although the model reported strong performance (Matthews Correlation Coefficient, MCC, values ranging from 0.834 to 0.965), challenges remained, especially regarding the high sequence diversity of peptides, the need for effective toxicity filtering, and the reliance on computational methods that may not fully capture the complexity of peptide interactions and biological functions.38 Similarly, a study conducted by Castillo-Mendieta et al. (2024) used chemical space complex networks to map the chemical space of hemolytic peptides and to enhance the design of safe peptide-based therapeutics. By analyzing a database of 2004 hemolytic peptides, the authors identified 12 consensus hemolytic motifs. They developed multi-query similarity searching models that outperformed the existing machine learning models in predicting hemolytic activity.39
In another study, Wang et al. (2024) improved a computational framework using a reinforcement learning (RL)-driven generative model integrated with graph attention mechanisms, which captured the connectivity structure between amino acid residues in peptides and used it to guide the search for optimal peptide sequences. The algorithm incorporates bioactivity and ADMET (absorption, distribution, metabolism, excretion, and toxicity) properties, ensuring that the generated peptides meet drug-like criteria.43
In contrast to the consensus chemical space, the concept of the chemical multiverse was introduced as a collection of multiple chemical spaces, each defined by different descriptors. This concept emphasizes that there is no single chemical space; rather, various representations of the same set of molecules can yield distinct chemical spaces.44 The chemical multiverse allows for a more comprehensive analysis of compound datasets using multiple descriptors, which can capture different aspects of molecular structures and properties. This approach contrasts with the idea of a consensus chemical space widely applied in medicinal chemistry, which attempts to combine various descriptors into a single representation, potentially losing valuable information in the process. The chemical multiverse does not rely on multiple descriptor combinations; instead, it consists of various alternative graphical representations of the chemical space, incorporating different molecular properties derived from the investigated compounds, such as molecular fingerprints, structure-based or sequence-based descriptors.44
The description of the chemical space of peptides has faced significant challenges, partly due to their chemical structure, vast physicochemical properties, and their intrinsic polymer-like features related to the amide backbone repetition, which tends to mask the prediction of their bioactive properties, hindering the analysis of their properties in the chemical space.45,46 In addition, it has been demonstrated that some unrelated classes of peptides show unexplained intersections between their bioactivities,47,48 thus evidencing that some previous distinct chemical spaces exhibit molecular similarities that must be better explored to find possible intersections in the intervals of the molecular descriptors applied to characterize them.
The chemical space usually contains some distinct clusters named ‘constellations’, which are populated by molecules with specific properties that can be identified using scaffold-based analysis due to the presence of a common structural core.49,50 The Murcko framework has been widely applied to investigate the structural core of drugs, revealing structural information and distinguishing molecules by their ring systems, linkers, and side chain atoms.51 However, Murcko frameworks can only represent molecules containing ring systems; therefore, acyclic (linear) peptides are usually omitted from these analyses.50 Moreover, peptides are characterized by their large molecular size and shape, as well as by the presence of polar groups, which usually put them beyond the conventional predictors of drug-likeness for molecules52,53 and impose more complexity in evaluating their conformational changes and pharmacophore properties.54
Scrambled peptides contain similar amino acid composition (AAC) and sequence length; however, they could acquire different conformations, which confer them different biological activities.55,56 The study of scrambled peptides has demonstrated some of these peculiarities of peptides compared to other small molecules.
Revealing the complexity behind the chemical space of peptides: investigating the interplay between conformation, bioactivity, and bioavailability
The chemical space of peptides encompasses a multidimensional subset of molecular properties linked to functional and drug-like characteristics,57 due to their unique physicochemical and structural properties that differentiate them from traditional small molecules used in the discovery and design of new drugs.15,58,59 These properties present a central challenge in characterizing a general chemical space of peptides, as they are crucial for predicting their biological, pharmacokinetic, and pharmacodynamic properties.60 This comprehension aids medicinal chemists in accurately classifying peptides and identifying suitable applications.Chemically modified residues that undergo post-translational modifications are likely to be extensive. The peptide chemical space thus encompasses a wide array of changes that can significantly alter the properties and functions of peptides.61 Understanding the chemical space of peptides also involves the comprehension of the three-dimensional conformations adopted by these molecules in the solvation medium, as their geometries are closely related to the mechanisms of action associated with membrane permeability and stereoselectivity against the molecular target.60,62,63 The conformation of peptides refers to the spatial arrangement of their atoms adopted due to the rotation around a single bond over time. It intrinsically depends on the peptide sequence and the external environment, and it is related to the formation of the secondary structure propensity. Peptides dynamically adopt a collection of conformations distributed across a free energy landscape, with their occurrence governed by Boltzmann-weighted probabilities.64
Some conformational adaptability in peptides can also modulate their bioactivity, because changes in the conformational ensemble alter the positions of pharmacophoric side chains and the population of binding-competent states, thereby affecting potency, selectivity, and the recognition to the target.65–67 This is consistent with observations that scrambled variants of peptides with similar composition may adopt distinct conformations and exhibit different biological activities.56 As the secondary structure is a determinant of peptide bioactivity, some strategies have been developed to impose constraints on their conformation to control their bioactivity.68,69 For example, some peptide design strategies for AMPs include the incorporation of restrictors, such as lactam and disulfide bridges, which act as conformational inducers (promoting β-like structures) and enhance resistance to protease degradation.70,71
Two classes of peptides that naturally cross biological barriers illustrate the relevance of conformational changes: the cell-penetrating peptides (CPPs) and blood–brain barrier-penetrating (B3PPs).72,73 Both classes exhibit a conformational characteristic that influences their ability to cross these barriers, which is called chameleonic properties. This conformational property refers to their ability to change conformation in response to environmental conditions, particularly to expose or hide polar groups when crossing biological membranes.74,75 Some well-reported examples of chameleonic properties of peptides include cyclosporin A75 and some of its derivatives76 (Fig. 4, panel A), as well as some cyclic peptides.75,77 This property has significant implications for their chemical space and the overall functionality of biomembrane-penetrating peptides. By altering their conformation, these peptides can effectively navigate through the hydrophobic core of cell membranes, improving their bioavailability.77
 | ||
| Fig. 4 Chameleonicity and backbone N-methylation as determinants of passive membrane permeability in macrocyclic peptides. Panel (A) Cyclosporin A, a chameleonic molecule, shown in representative open (greater polar surface exposed) and closed (polar surface partially buried) conformations, with electrostatic potential maps (scale in kBT/e) that illustrate environment-dependent exposure/burial of polar groups through conformational switching and intramolecular hydrogen bonding. Center: Conceptual schematic of passive diffusion through a lipid bilayer: lower-permeability variants tend to retain solvent-exposed hydrogen-bond donors and acceptors, whereas higher-permeability variants more effectively mask polarity. Panel (B) Parent scaffold cyclo[Leu1, D-Leu2, Leu3, Leu4, D-Pro5, Tyr6] (compound 1) and the corresponding trimethylated analogue (compound 3), bearing backbone N-methyl groups at D-Leu2, Leu3, and Tyr6 (Me)—a modification pattern associated with higher permeability, as reported by White et al. (2011).93 | ||
Some computational models may struggle to accurately represent the conformational variability of peptides, posing challenges for predicting their pharmacokinetic properties.58 For example, the calculation of topological polar surface area (tPSA) does not depend on the three-dimensional characteristics of the molecules, and it has been widely applied to correlate with the hydrogen bond pattern of molecules in the aqueous phase.78 This property has been associated with the prediction models of solubility and passive diffusion through cell membranes.79–82 Elevated tPSA values are associated with complexation with water molecules and increased molecular volume, which can hinder membrane permeability.83 Typically, the penetration of compounds across cell membranes is restricted when tPSA exceeds 140 Å2.84 However, higher values are generally acceptable for macrocyclic peptides (tPSA = 220 Å2) and peptides exhibiting chameleonic properties (tPSA = 280 Å2).81,85 The Molecular 3D PSA (MPSA) has emerged as a more accurate measure of compound solubility and membrane permeability than the tPSA, as it considers the three-dimensional conformation of the compound in a given environment.86 The tPSA, however, does not depend on the tridimensional structure and could reach satisfactory prediction, especially when associated with the molecular weight of compounds.87 The macrocycle peptide cyclosporine A is an example of a natural peptide that exhibits chameleonic activity. Cyclosporin A has a high tPSA value of 279 Å2 and an MPSA value equal to 105 Å2 with approximately 62% of its PSA concealed in nonpolar environments.87,88
The permeability of peptides into cell membranes could be significantly influenced by their secondary structure.89 Studies have demonstrated the impact of peptide conformation on arginine-mediated internalization in cell membranes.90 Similarly, other studies have shown that for CPPs, some helices stabilized by hydrocarbon cross-links can effectively enter the cells.91,92 The N-methylation of cyclic peptides has been widely reported as an interesting strategy to improve the permeability in cell membranes.79,93 White et al. (2011), for example, demonstrated that methylated analogues generated by on-resin N-methylation improved their membrane intake. The Figure X, panel (B) shows the regioselective backbone N-methylation of a cyclic hexapeptide scaffold cyclo[Leu1, D-Leu2, Leu3, Leu4, D-Pro5, Tyr6] (compound 1) and its trimethylated analogue (compound 3) generated by on-resin N-methylation. In compound 3, D-Leu2, Leu3, and Tyr6 are N-methylated (Me), a pattern associated with markedly improved passive permeability in the study by White et al. 2011.93
Currently, most in silico models—including cheminformatics filters and machine-learning approaches used in peptide science—assume that these molecules cross biological membranes primarily via passive diffusion, implying that membrane penetration occurs mainly through biophysical interactions between the peptide's structure and the membrane.58 Consequently, active transport pathways, like receptor-mediated transcytosis, active influx transport, and carrier-mediated transcytosis, are often overlooked in the design of these models, primarily due to the intricate binding processes of membrane proteins associated with the conformational accommodation related to receptor binding.58,94 Furthermore, some cheminformatics filters that use a set of molecular descriptors and their intervals to characterize drug-like molecules, such as the BOILED-Egg model and Lipinski's Rule of Five, often fail to accurately predict the permeability of peptides due to their distinct chemical space.82,95 These classical rules and empirical models were largely developed and calibrated using small molecules and therefore delineate a region of property space enriched for passive permeability and oral bioavailability.79,95 When applied to peptides, these filters often fail to accurately predict peptide permeability and overall drug-like behavior because peptides typically present higher molecular weight, larger polar surface area, multiple hydrogen-bond donors/acceptors, and higher conformational flexibility—features that shift them outside conventional small-molecule boundaries.83,95 Therefore, applying small-molecule drug-likeness rules can artificially truncate peptide chemical space and may lead to misleading conclusions to their bioactivity or bioavailability.
Moreover, some peptide classes can partially overcome these restrictions through structural adaptations that are not captured by simple 2D descriptors used in the cheminformatic filters. For example, macrocyclic and “chameleonic” peptides can conceal polar surface area via intramolecular hydrogen bonding and environment-dependent conformational changes, thereby improving passive permeability despite high tPSA values. In addition, descriptors that incorporate 3D conformation may be more informative than purely 2D metrics for some peptide subclasses.62,74
Considering these limitations, efforts have been made to accurately represent their geometries by analyzing the rotamers and possible conformational changes of peptides to enhance the prediction of their molecular activities.96,97
An overview of molecular descriptors applied to peptides is demonstrated in Fig. 5.
 | ||
| Fig. 5 Classes of molecular descriptors commonly used to represent peptide sequences and structures and to analyze peptide structure–activity relationships. | ||
Structure-based features to describe the chemical space of peptides
Many chemical properties of peptides, particularly topological, stereochemical, electronic, aromatic, and physicochemical features, can only be calculated by examining their atomic structure. This is particularly true for synthetically modified peptides or those with post-translational modifications, where chemical changes to the amino acid side chains (such as the addition or removal of functional groups) create new molecular functions and properties.5,98 These descriptors can be calculated from the 2D or 3D representation of peptides in structural data files, such as SDF, CDX, and MDL Molfiles.Many descriptors widely used in peptide science and medicinal chemistry are computed from 2D connectivity or composition and therefore do not require a specific 3D conformer (e.g., MW, HBA/HBD counts, tPSA, atom counts, fragment-based log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, and 2D fingerprints). In contrast, geometry-, surface-, and shape-based properties are conformation-dependent and should be computed from an explicit 3D structure or a conformational ensemble (e.g., SASA/3D PSA such as MPSA, WHIM, GETAWAY, and 3D-MoRSE descriptors). The 2D representation is often sufficient to calculate most descriptors since these descriptors capture features like atomic constitution, the presence of specific chemical groups, physicochemical properties, and molecular topology. The selection of the most appropriate molecular descriptors usually depends on the type of peptide under investigation and its associated biological activity.58
P, and 2D fingerprints). In contrast, geometry-, surface-, and shape-based properties are conformation-dependent and should be computed from an explicit 3D structure or a conformational ensemble (e.g., SASA/3D PSA such as MPSA, WHIM, GETAWAY, and 3D-MoRSE descriptors). The 2D representation is often sufficient to calculate most descriptors since these descriptors capture features like atomic constitution, the presence of specific chemical groups, physicochemical properties, and molecular topology. The selection of the most appropriate molecular descriptors usually depends on the type of peptide under investigation and its associated biological activity.58
Descriptors used in large peptide libraries are broadly borrowed from medicinal chemistry, originally designed for small molecules, where they proved useful in predicting drug-likeness and bioavailability. Key examples include tPSA, MW, HBA, and HBD, number of aromatic rings (NAR), the fraction of sp3-hybridized carbon atoms (Fsp3), lipophilicity calculated by the logarithm of 1-octanol/water partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), number of chiral centers (NCC), and the number of rotatable bonds (NRB).79,84,99,100 Additional descriptors capture structural features, such as secondary structure composition, ionization state, topology, shape, and hydrophilicity.
P), number of chiral centers (NCC), and the number of rotatable bonds (NRB).79,84,99,100 Additional descriptors capture structural features, such as secondary structure composition, ionization state, topology, shape, and hydrophilicity.
The importance of these descriptors lies in their connection to bioavailability, i.e., how efficiently a compound can dissolve in aqueous environments and cross biological membranes. Two widely used descriptors are log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and tPSA, which reflect lipophilicity and polar surface area, respectively.82,83 Similarly, intrinsic solubility can be explored through lipophilicity (e.g., log
P and tPSA, which reflect lipophilicity and polar surface area, respectively.82,83 Similarly, intrinsic solubility can be explored through lipophilicity (e.g., log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and the logarithm of the distribution partition coefficient (log
P and the logarithm of the distribution partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D) at pH 7.4), structural constitution (e.g., NAR), and molecular flexibility (e.g., Fsp3, NCC, and NRB).53,79,81,82,99,101
D) at pH 7.4), structural constitution (e.g., NAR), and molecular flexibility (e.g., Fsp3, NCC, and NRB).53,79,81,82,99,101
Molecular descriptors associated with the ionization state are informative of aqueous solubility (hydrophilicity) and include the isoelectric point (pI) and the logarithmic value of the acid dissociation constant (pKa). Molecular hydrophobicity (lipophilicity) is usually quantified by the calculations of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values, and alternative computational methods, such as X
P values, and alternative computational methods, such as X![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P,102C
P,102C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, and A
P, and A![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P103 calculate some of its derivative values. The A
P103 calculate some of its derivative values. The A![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P has demonstrated superior predictive accuracy for peptides compared with other calculated log
P has demonstrated superior predictive accuracy for peptides compared with other calculated log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values.104
P values.104
Certain molecular descriptors that capture the shape and topology of peptides can be linked to their stereoselectivity towards molecular receptors. These descriptors reflect the spatial rearrangement and connectivity of the molecules. Examples include Kier's Kappa indices,105 Balaban indices,106 Burden eigenvalues,107 and Randić shape indices.108 Other descriptors focus on molecular complexity. For instance, Basak's Indices provide a numerical representation of structural features such as connectivity, branching, and overall topology.109
Among these shape- and topology-oriented metrics, the Kappa descriptors stand out as topological indices derived from the hydrogen-suppressed molecular graph. They quantify molecular shape and the intricacy of branching by considering paths of different lengths through the structure, comparing the observed branching to an idealized linear or maximally branched reference. Because they are sensitive to the global shape rather than the size of the molecule, Kappa indices offer a refined picture of molecular flexibility and compactness. In QSAR applications, they are particularly useful for highlighting steric and topological features that influence biological activity and receptor interaction.110
Another important topological descriptor known for its low degeneracy and strong correlation with physicochemical properties is the Balaban index. It is calculated from the distance matrix of the molecular graph, integrating information from distance sums and the number of edges. By encoding details related to cyclicity, branching, and structural compactness while remaining largely independent of molecular size, the Balaban index stands out at distinguishing structural isomers, which is especially valuable in studies involving complex ring systems and detailed structure–property relationships.111
To study the geometry of compounds, researchers often rely on 3D molecular descriptors, which require the explicit 3D conformation of a molecule to capture the spatial arrangement of its atoms. These descriptors are widely used in structure–activity relationship (SAR) analyses. Notable examples include WHIM (Weighted Holistic Invariant Molecular descriptors), GETAWAY (GEometry, Topology, and WAter Accessibility),112 and 3D-MoRSE (Molecular Representation of Structures based on Electronic diffraction).113
The 3D-MoRSE encodes fundamental 3D atomic coordinates of the molecular structure using a fixed-size vector, drawing on concepts akin to electron diffraction. The 3D-MoRSE descriptors differ from purely topological indices because they incorporate three-dimensional structure along with electronic information. They are generated from simplified theoretical scattering curves—conceptually similar to electron diffraction—using atomic coordinates and weighted properties such as mass or partial charges. The resulting values represent the distribution of electron density across a range of scattering angles. In doing so, the 3D-MoRSE descriptors capture steric effects, electrostatic interactions, and other conformation-dependent features that cannot be inferred from 2D graph topology alone.113
The WHIM (Weighted Holistic Invariant Molecular descriptors)114 is a set of 3D molecular descriptors that capture the spatial arrangement of atoms in a molecule.114 The WHIM descriptors91 are also based on 3D atomic coordinates but summarize them through a weighted principal component analysis. It's used a covariance matrix built from atomic positions, with weights that may reflect atomic mass, polarizability, electronegativity, or other physicochemical attributes. Because they derive from molecular invariants, WHIM descriptors remain consistent under translation and rotation of the molecule, providing a global statistical representation of size, shape, symmetry, and atom distribution relative to three orthogonal axes.115
The GETAWAY (GEometry, Topology, and Atom-Weights AssemblY) is a 3D-molecular geometry provided by the molecular influence matrix and atom-relatedness by a molecular topology using different atomic weighting.112 The GETAWAY descriptors complement these approaches by integrating geometric features with molecular topology through the influence matrix and various atom-relatedness measures. Together, this family of 3D descriptors enables characterization of steric, electronic, and conformational aspects that strongly influence molecular recognition processes.112
Measures of molecular complexity lack a universal concept, but they have been frequently associated with synthetic accessibility and, in the context of drug design, with the specificity to a molecular target.116 Currently, different descriptors capture various aspects of molecular complexity, and their complementary use may provide an overview of this molecular property, such as topological and physicochemical descriptors (e.g.: NCC and Fsp3)99,101 and some substructure-based descriptors (number of rings, unsaturations, and heteroatoms). For peptides, the difficulty of synthesis has been associated with long amino acid chains and functional groups associated with their side chains.117 A non-exhaustive list of the most applied structure-based descriptors to investigate peptides is presented in Table 1.
| Structure-based descriptors | Peptide information |
|---|---|
Notes: CC: number of chiral carbons; Fsp3: fraction of sp3-hybridized carbon atoms;101 GETAWAY: GEometry, topology, and atom-weights AssemblY;112 HBA: hydrogen bond acceptors, HBD: hydrogen bond donors; log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P: 1-octanol/water partition coefficient; 3D-MoRSE: 3D molecular representations of structure based on electron diffraction; MPSA: molecular 3D polar surface area; MW: molecular weight; MSA: molecular surface area; NCC: number of chiral centers; NPA: number of primary amino groups (–NH2); NHA: number of heavy atoms, NAR: number of aromatic rings, NG: number of guanidine groups; NNCAA: number of negatively charged amino acid groups; NRB: number of rotatable bonds; pKa: logarithm of the acid dissociation constant, pI: isoelectric point; tPSA: topological polar surface area;78 PSA: polar surface area; VdW: van der Waals volume; SASA: solvent accessible surface area; XLOGP3: log P: 1-octanol/water partition coefficient; 3D-MoRSE: 3D molecular representations of structure based on electron diffraction; MPSA: molecular 3D polar surface area; MW: molecular weight; MSA: molecular surface area; NCC: number of chiral centers; NPA: number of primary amino groups (–NH2); NHA: number of heavy atoms, NAR: number of aromatic rings, NG: number of guanidine groups; NNCAA: number of negatively charged amino acid groups; NRB: number of rotatable bonds; pKa: logarithm of the acid dissociation constant, pI: isoelectric point; tPSA: topological polar surface area;78 PSA: polar surface area; VdW: van der Waals volume; SASA: solvent accessible surface area; XLOGP3: log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P estimated from the atom/fragment contribution values;120 WHIM: weighted holistic invariant molecular descriptors.114 P estimated from the atom/fragment contribution values;120 WHIM: weighted holistic invariant molecular descriptors.114 |
|
| Bounds count, heavy atoms count, atoms type count (e.g., N-, O-, C-, S), NRB. | Atomic constitution |
| NPA, NG, NNCAA | Presence of molecular groups |
| MW, NHA, VdW, MSA | Molecular size |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, log P, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D (pH 7.4), X D (pH 7.4), X![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P3, A P3, A![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, C P, C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, log P, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kow Kow |
Lipophilicity/hydrophobicity |
| Kappa indices (Kappa1, Kappa2, and Kappa3), Burden eigenvalues, Basak's Indices, Balaban index, Weiner index, Randić indices, WHIM indices114 | Molecular shape and topology |
| Fsp3, NRB | Molecular flexibility |
| tPSA, MPSA, PSA | Polar surface (polarity) |
| WHIM indices,114 3D-MoRSE,113 GETAWAY112 | Molecular geometry |
| NCC,99 Fsp3,99,101 CC,118 | Molecular complexity |
HBA, HBD, tPSA, net charge, pKa, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, X P, X![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, A P, A![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, C P, C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, log P, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kow, SASA Kow, SASA |
Hydrophilicity (aqueous solubility) |
| pKa, pI | Ionization state |
| Number of α-helices, number of β-sheets, number of coils | Secondary structure |
| Eisenberg scale119 | Hydrophobic moment |
Sequence-based features to describe the chemical space of peptides
While all peptide properties ultimately originate from the primary structure (amino acid composition and sequence order) and the chemical modifications, only a subset of properties can be directly computed from the sequence.121–123 Accordingly, sequence-based descriptors are associated with information calculated from the primary structure of an oligopeptide sequence, assessing the amino acid composition, sequence motifs, amino acid arrangements, and their physicochemical patterns, including the presence of hydrophilic and hydrophobic regions, and are especially valuable when tertiary structure information is unavailable.58 Some encoders, scoring, and substitution matrices have been applied to categorize peptides into distinct classes, providing a statistical description of their sequences, and filling a critical gap when tertiary structure information is absent.58,124,125 Most of these sequence-based descriptors were developed to classify and analyze protein sequences, but have also been applied for peptides.15,126,127 These descriptors could be calculated from the string representation of amino acids represented by the FASTA,128 PLN, BILN,129 and HELM130 file formats that contain the peptide's primary structure.131Peptides contain both hydrophilic and hydrophobic regions, often influenced by the relative abundance of specific amino acid residues, which in turn shape their molecular mechanisms of action. A notable example is the CPPs, which are typically enriched in lysine and arginine residues. This composition accounts for their cationic or amphipathic nature at physiological pH, as well as their water solubility and cell membrane permeability.132,133 Studies have demonstrated that incorporating arginine into cyclic peptides and protein surfaces enhances cell penetration.134,135 Adapting the amino acid structure of a peptide might influence their biological activity and drug-like properties.5 Cyclization and N-methylation are examples of chemical modifications that shift the peptides' positions within the chemical space, enhancing their potential bioavailability and biological activity.5 For example, a study revealed significant overlaps between the chemical space of synthetic linear and cyclic pentapeptides containing N-methylation and some FDA-approved peptide drugs. Some studies have shown that machine-learning algorithms applied to predict some classes of peptides that employ an optimized integration of sequence- and structure-based descriptors in the feature composition achieve greater accuracy than those relying solely on sequence- or structure-based descriptors.15,136–138
The sequence-based properties have also been used to design novel domains and modulate switchable properties of peptides. Self-assembling peptides (SAPs) are short polypeptide chains that, in an aqueous solution, can spontaneously organize themselves into complex, well-ordered, and stable nano- and meso-structures through the formation of non-covalent interactions,139,140 thus forming versatile building blocks which have been extensively studied to create stimuli-sensitive supramolecular systems.141–143 The amino acid sequence composition and the orientation of the amino acids of SAPs could play a critical role in driving the self-assembling properties. For example, in the SAP sequence, aromatic amino acids, such as phenylalanine, tyrosine, and tryptophan, contribute mostly to aggregation through π-stacking as the main driving force for self-assembly.144 On the other hand, the presence of histidine, serine, and threonine amino acids has highly polarizable side chains, and thus, these peptide structures could promote aggregation through hydrogen bonding formation. Some SAP structures are characterized by their amphiphilicity, meaning their sequences contain hydrophilic and hydrophobic domains that facilitate self-assembly in aqueous solutions, forming non-covalent interactions between amino acid residues.145
Peptide molecules that self-assemble into peptide nanofibers are primarily amphiphilic molecules. These consist of hydrophilic heads containing active peptide segments, hydrophobic tails with alkyl chains, and several amino acids between these two regions, creating enough space to prevent spontaneous aggregation after introducing negative charges.146,147 Amphipathic peptides are more likely to self-assemble into amyloid-like β-sheet fibrils when their primary sequence shows a pattern of alternating hydrophobic and hydrophilic amino acids. These fibrils form a bilayer structure comprising two β-sheets that align to conceal the hydrophobic side chains within the bilayer's interior. In contrast, the hydrophilic side chains remain exposed on the surface of the bilayer.148,149 Recently, a study demonstrated that the SAP sequence significantly influences structural sensitivity to supramolecular polymerization pathways, affecting the resulting polymers' structural and functional properties.150 Yuan et al. (2022) demonstrated that the order of amino acids in the sequences AAEE and AEAE (A and E represent alanine and glutamic acid, respectively) impacts the driving forces involved in peptide polymerization, which directly correlates with mechanical properties and bioactivity.150
Some computational models have been developed using a combination of sequence-based and structure-based descriptors to predict the bioactivity of peptides. These models have improved performance compared to algorithms relying solely on one class of molecular descriptors.15,94,124,136,137 For example, Rajput et al. (2015) analyzed the QSPs according to their amino acid composition, residue position, physicochemical properties, and sequence motifs and identified that some aromatic residues, such as tryptophan, tyrosine, and phenylalanine play an important role in their characterization, as well as positional preferences of residues, such as serine at the N-terminal end and phenylalanine at the C-terminal end so that these sequence-based properties could be used for their identification.124 Physicochemical properties, such as aromaticity, molecular weight, and secondary structure, contribute to QSP identification.124 Recent approaches utilize propensity score representation learning to extract and combine the propensities of amino acids and dipeptides.151
Sequence-based descriptors provide a quantitative framework for analyzing peptides, capturing features such as amino acid composition, positional distribution, and sequence arrangement patterns. These descriptors enable the application of statistical and machine learning approaches to uncover structure–activity/property relationships in peptide research.127,152–154 In this context, we focus on molecular descriptors and scoring matrices that support large-scale, data-driven analyses. Several Python libraries, widely used for big data applications, offer built-in tools for calculating such descriptors, such as BioPython,155 RDkit,21 Mordred,24 and PyBioMed.156 Additionally, there are tools focused on peptide analysis, such as PepFun,157 iFeature,23 iFeatureOmega,22 Peptide.py (https://pypi.org/project/peptides/),158 and PepFuNN.159
Some sequence-based descriptors evaluate the amino acid or k-mers constitution, providing information about their relative abundance or scarcity of amino acids, such as AAC,160 dipeptide composition (DPC),161 tripeptide composition (TPC),152 and terminus composition (TC).162k-mers are substrings created by moving a window of length k along the sequence at a set interval. In addition to the constitution, they reflect the overall frequency of these amino acids.
Other sequence-based encoders offer information about group- and gap-based amino acid rearrangements. The group-based amino acid descriptors aim to mitigate the high-dimensional data derived from the existence of 20 amino acids, so this class of encoders groups or reduces the amino acid compositions to investigate the peptide sequences. High-dimensional data can lead to overfitting, compromising the prediction accuracy of the models when the number of features exceeds the number of independent samples.163 Thus, these descriptors extract characteristics that better reflect the relationships of groups of amino acid residues in the sequence. The group-based amino acid composition descriptors, for example, include the grouped tripeptide composition (GTPC),164 grouped dipeptide composition (GDPC),164 pseudo k-tuple reduced amino acid composition (PseKRAAC),165 and the grouped amino acid composition (GAAC).23 In contrast, the gap-based amino acid descriptors create bi-mers from peptide sequences using various gap sizes, and subsequently analyze the distribution of the resulting gap-based bi-mers. These sequence-based descriptors include composition of k-spaced amino acid pairs (CKSAAP),166 and adaptive skip dipeptide composition (ASDC).167,168
In addition to these descriptors, some libraries extract from the sequence the AAindex, a curated, literature-derived database that compiles numerical indices describing physicochemical properties of amino acids. In its core component (AAindex1), each “amino acid index” represents a single property as a set of 20 numerical values, one per standard amino acid, enabling sequences to be converted into quantitative property profiles.169
Several substitution and scoring matrices have also been developed to represent the variability, the physicochemical properties, and substitution patterns of polypeptide sequences, including position-specific scoring matrix (PSSM), residue pairwise energy content matrix (RECM),170 Z-scale,171 and BLOcks Substitution Matrix (BLOSUM).172 The BLOSUM and PAM matrices, for example, are derived from oligopeptide sequence alignments, and both are commonly used as encoders to characterize peptide sequences based on their evolutionary substitution profiles (Table 3),173 showing variations depending on the identities of the pre-computed datasets. For example, BLOSUM matrices come in different versions, such as BLOSUM50, BLOSUM62, and BLOSUM80, created using the observed frequencies of amino acids in peptide sequences. The 62% identity threshold (BLOSUM62) is widely used for peptide and protein sequence characterization.174,175 In contrast, the position-specific scoring matrix (PSSM), residue pairwise energy content matrix (RECM), and z-scale171 are classified as scoring matrices applied for amino acid sequences (see Table 3). The z-scale, for example, is an amino acid descriptor set used to numerically represent the physicochemical, hydrophobic, and polar properties of amino acids in protein or peptide sequences. This matrix is derived from a PCA of various amino acid and physicochemical properties, reducing them into a few orthogonal components.171
While the content of the secondary structure is dependent on the conformation of the peptide, and is more accurately calculated using information derived from the three-dimensional structure, several sequence-based prediction methods have been developed that demonstrate promising results for predicting secondary features and classifying oligopeptide sequences.176–178 For example, Zhang et al. (2011) developed a transition probability matrix to represent secondary structures,178 and Dai et al. (2013) introduced a statistical position-based feature of secondary structural elements to predict the structural classes of oligopeptide sequences.177 The secondary structure elements content (SSEC), for instance, is a molecular descriptor calculated from the primary structure predicted by the PSIPRED V4.0 and provides the content of three types of secondary structure elements.23
The correlation encoder quantifies the relationship between amino acids by calculating correlation coefficients that reflect differences in the molecular descriptors that reveal information about hydrophobicity, hydrophilicity, mass, shape, topology, constitution, etc. These descriptors reveal how specific properties of amino acids are interrelated to the sequence. Moran,179 Normalized Moreau-Broto, and Geary179 are autocorrelation descriptors that uses eight amino acid indexes by default for peptide sequences, according to the following: the DAYM780201 represents the the residue substitution profile, the CHOC760101 represents the residue accessible surface area in tripeptide, the CIDH920105 represents the normalized average hydrophobicity scales, the BHAR880101 represents the average flexibility indices, CHAM820101 represents the polarizability parameter, CHAM820102, represents the free energy in water, the BIGC670101 represents the volume of the residue, and the CHAM810101 the steric parameter.23,180
Binary encoders are descriptors that transform amino acid sequences into statistical vectors, with each amino acid encoded as a 20-dimensional binary vector consisting of 0 s and 1 s. The binary representation is subdivided into 3, 5, 6, and 20 bits, and they represent some groups of amino acids of the sequence depending on their physicochemical properties.181 For example, the binary 6-bit uses a six-element amino-acid groups {e1, e2, e3, e4, e5, e6} to encode the oligopeptide sequence, where e1 ∈ {H, R, K}, e2 ∈ {D, E, N}, e3 = C, e4 ∈ {S, T, P, A, G}, e5 ∈ {M, I, L, V}, e6 ∈ {F, Y, W}. These groups capture conservative substitutions that can occur over evolutionary time. They function as equivalence classes grouping amino acids by similarity, and their definitions are based on PAM-based relationships. Then, each group is represented by a 6-dimensional binary vector, e.g., e1 is encoded by (100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000), e2 is encoded by (010
000), e2 is encoded by (010![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000), and so on.22 In the sparse encoding approach, each peptide sequence is mapped to a fixed-length vector of 100 positions, corresponding to the maximum sequence length stored in the database. A reference list containing the 20 standard amino acids plus one additional symbol for gaps or empty positions is used. Each amino acid is converted into a one-hot vector of length 21, where a single element indicating its position in the list is set to “1”, and the remaining elements are set to “0”. Consequently, every position in the 100-length sequence corresponds to a 21-dimensional vector. This representation ensures that each amino acid is uniquely identified by its position within the encoding space.181
000), and so on.22 In the sparse encoding approach, each peptide sequence is mapped to a fixed-length vector of 100 positions, corresponding to the maximum sequence length stored in the database. A reference list containing the 20 standard amino acids plus one additional symbol for gaps or empty positions is used. Each amino acid is converted into a one-hot vector of length 21, where a single element indicating its position in the list is set to “1”, and the remaining elements are set to “0”. Consequently, every position in the 100-length sequence corresponds to a 21-dimensional vector. This representation ensures that each amino acid is uniquely identified by its position within the encoding space.181
A list of molecular descriptors derived from the sequence is described in Table 2. A list of applied scoring and substitution matrices is described in Table 3. A list of autocorrelation descriptors associated with the amino acid indices is presented in Table 4.
| Sequence-based descriptors | Peptide information |
|---|---|
| Notes.a GRAVY: grand average of hydropathy, corresponds to the value of the hydropathic index calculated by the Kyte–Doolittle method using the peptide sequence.b FLEX index: corresponds to the structural flexibility calculated from the peptide sequence according to the Vihinen et al., 1994. | |
| Amino acid composition (AAC)160 | Frequencies of the 20 types of native amino acids present over the peptide sequence |
| Pseudo-amino acid composition (PseAAC)182 | Frequencies of the discrete sequence correlation factors and the twenty components of the conventional amino acid composition |
| Amphiphilic pseudo-amino acid composition (APAAC) | Frequencies of the discrete sequence correlation factors related to the hydrophobicity and hydrophilicity |
| Dipeptide composition (DPC)161 | Frequencies of 400 types of dipeptides present over the sequence |
| Tripeptide composition (TPC)152 | Frequencies of 8000 types of tripeptides present over the sequence |
| Grouped amino acid composition (GAAC)23 | Frequencies of five groups of amino acids based on their physicochemical properties: negative charge (D, E), positive charge (H, R, K), aromatic group (F, Y, W), aliphatic group (A, G, I, L, M, V), and uncharged (C, N, P, Q, S, T). |
| Terminus composition (TC)162 | Frequencies of amino acids and dipeptides for 5, 10, and 15 residues present at the N- and C-terminus of the peptide sequence. |
| Composition of k-spaced amino acid pairs (CKSAAP)166 | Frequencies of 400 types of residue pairs separated by k other amino acids (k![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) = = ![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 2, 3) within a sequence or sequence fragment. 1, 2, 3) within a sequence or sequence fragment. |
| CTDT (composition/transition/distribution) | Distribution of amino acid composition patterns linked to specific chemical, physical, or structural properties within the peptide sequence. The composition (C) refers to the amino acid composition in sequence, the transition (T) corresponds to changes among three patterns: neutral, hydrophobic, and polar, and the distribution (D) refers to the pattern of distribution of these properties over the sequence. |
| Pseudo K-tuple reduced amino acids composition (PseKRAAC)165 | Frequencies of the 16 types of reduced K-tuple pseudo amino acids calculated from the sequence-order information for all dipeptides and the correlation between nth nearest residue. |
| Adaptive skip dipeptide composition (ASDC)167 | Frequencies of amino acid pairs separated by a variable (adaptive) number of intervening residues. |
| Quasi-sequence-order descriptors (QSOrder)183 | Frequencies of the amino acid sequence orders calculated using the sequence-order-coupling numbers that reflect the interactions between amino acids at various ranks of proximity. The coupling factor used to calculate these numbers is based on the physicochemical distance between amino acids, which considers properties like hydrophobicity, hydrophilicity, side-chain volume, and polarity. |
| Secondary structure elements content (SSEC) | Number of α-helices, β-sheets, and coils |
| Shannon information entropy | Scoring value that measures the degree of variability at a specific amino acid position in a multiple sequence alignment |
| AAindex169 | Compilation of literature-reported scales that quantify physicochemical tendencies of the standard amino acids. Each scale corresponds to one property and is encoded as a 20-element numeric vector, assigning a specific value to each amino acid |
| GRAVY index184,a | Hydropathic character |
| FLEX index185,b | Structural flexibility |
| Scoring and substitution matrices | Peptide information |
|---|---|
| Note.a Different identity thresholds can be applied in BLOSUM to characterize peptide sequences, with 62% typically used in most alignments. | |
| Position-specific scoring matrix (PSSM)186 | Scoring matrix containing the likelihood of each amino acid at a specific position in a peptide sequence. It is derived from multiple sequence alignments, aiding the identification of conserved regions. |
| Residue pairwise energy content matrix (RECM)170 | Scoring substitution 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) × × ![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 matrix containing residue pairwise energy for 20 standard amino acids derived from the primary structure of 674 proteins. 20 matrix containing residue pairwise energy for 20 standard amino acids derived from the primary structure of 674 proteins. |
| BLOcks Substitution Matrices (BLOSUM)172,a | Substitution 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) × × ![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 matrix based on observed substitutions in conserved blocks, with a threshold of identitya. 20 matrix based on observed substitutions in conserved blocks, with a threshold of identitya. |
| Grantham distance matrix187 | Scoring substitution 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) × × ![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 matrix that incorporate residue substitution frequencies that better correspond to the overall chemical differences including composition, polarity, and molecular volume. 20 matrix that incorporate residue substitution frequencies that better correspond to the overall chemical differences including composition, polarity, and molecular volume. |
| Point accepted mutation (PAM) matrices (also named Dayhoff matrices) | Substitution 20 × 20 matrix where each entry in the matrix represents the likelihood of one amino acid being replaced by another through accepted mutations over a specified evolutionary period |
| z-Scale171 | Scoring 87 × 26 matrix applied for amino acid sequences, where the 87 rows correspond to the different amino acids (including 20 standard amino acids plus many non-coded or unusual ones) and 26 columns correspond to different physicochemical descriptor scores. |
| Autocorrelation descriptors | Equations |
|---|---|
Notes.a I(d) is the Moran autocorrelation, d is the lag of the autocorrelation, nlag is the maximum value of the lag (default value: 30), Pi and Pi+d are the properties of the amino acids at positions i and i + d, respectively. ![[P with combining macron]](https://www.rsc.org/images/entities/i_char_0050_0304.gif) is the average of the considered property P over the entire sequence of length N.b C(d) is the Geary autocorrelation, d, P, Pi, and Pi+d, nlag, and N have the same definitions as defined for Moran.c ATS(d) is the normalized Moreau-Broto autocorrelation, AC(d) is the Moreau-Broto autocorrelation, d, P, Pi, and Pi+d, nlag, and N have the same definitions as defined for Moran. is the average of the considered property P over the entire sequence of length N.b C(d) is the Geary autocorrelation, d, P, Pi, and Pi+d, nlag, and N have the same definitions as defined for Moran.c ATS(d) is the normalized Moreau-Broto autocorrelation, AC(d) is the Moreau-Broto autocorrelation, d, P, Pi, and Pi+d, nlag, and N have the same definitions as defined for Moran. |
|
| Morana |
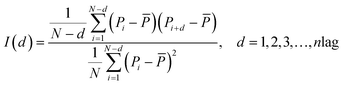
|
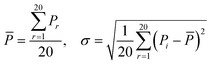
|
|
| Gearyb |
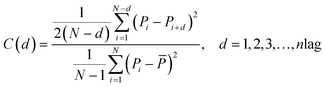
|
| Normalized Moreau-Broto autocorrelationc |
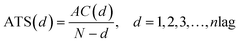
|
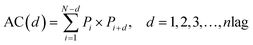
|
|
Molecular fingerprints for peptides
The selection of an appropriate molecular representation and the molecular properties most correlated with the investigated set of compounds plays a crucial role in analyzing structure–property relationships and exploring broader chemical space.188 Several computational strategies have emerged to capture peptides' chemical and functional space.6,15,189,190 These approaches have been applied to different bioactive peptide classes,189–191 peptides approved for human use,57 as well as molecules derived from peptides (peptide-like molecules, e.g., peptoids).46Molecular fingerprints provide a cost-efficient computational method for analyzing large compound libraries, due to their compact representation of complex molecular structures,6,192 which justifies their integration with computationally demanding virtual screening techniques.2,193 Molecular fingerprints serve as representations of a chemical structure that encode the presence or the absence of a particular molecular feature.193,194 These types of molecular representation are essential for analyzing large chemical libraries and comparing their structures using quantitative assessment of pairwise similarity.194 Currently, six main categories of molecular fingerprints are used to describe molecules: (1) descriptor-based, (2) substructure-based, (3) pharmacophore-based, (4) path-based (or hashed), (5) string-based, and (6) circular fingerprints.195
Descriptor-based fingerprints use molecular features derived from physicochemical properties, such as the van der Waals surface area (VSA) fingerprint. The substructure-based fingerprints are used to identify the presence of specific substructures, including functional groups and rings of certain sizes. This class includes the MACCS (Molecular ACCess System) key fingerprint.196 The pharmacophore fingerprints encode the pharmacophore groups present in molecules, and this class characterizes the interaction of the molecules with the protein environment. Belonging to this class is the MXFP, an atom-pair fingerprint that describes molecular shape and pharmacophores.192 The path-based (or hashed) fingerprints identify all types of subgraphs, including linear subgraphs representing the shortest paths between atom pairs and circular fingerprints that capture the neighborhoods of bonded atoms, hashing them inside a fixed-size vector. Atom-pairs are a subclass of path-based that describes a molecule by analyzing all possible triplets present in two atoms and the shortest path that connects them.197 These fingerprints include the E3FP,198 ECFP, and MAP4.6 The string-based fingerprints create molecular representations by analyzing the SMILES string of a compound rather than its graphical representation. Finally, the circular fingerprints decompose the analyzed compound into various fragments, similar to substructure-based fingerprints. However, instead of depending on predefined structural patterns, they dynamically generate these fragments from the molecular graph of each compound.
Currently, most virtual screening strategies or chemical space mapping of compounds applied in drug discovery use the MACCS key fingerprint, Morgan fingerprint – commonly referred to as the ECFP fingerprint,199 and MinHashed fingerprint MHFP6.200 Nevertheless, these molecular fingerprints usually struggle to accurately capture the overall characteristics of molecules, including their size and shape. Additionally, they are inadequate at recognizing structural variations that could be significant in larger molecules, such as distinguishing between linkers of varying lengths, identifying scrambled peptide sequences with the same amino acid composition and sequence length, or differentiating between regioisomers.52
A pharmacophore-based fingerprint derived from the 2D structure of peptides, termed 2DP, was developed to encode the molecular shape and pharmacophore properties of peptides. This fingerprint represents the peptides’ topology as a graph where nodes correspond to α-carbon atoms and edges represent bonds between them. This fingerprint captures key molecular features, including the number of hydrophobic, positively charged, negatively charged, and total non-hydrogen atoms in each residue. Distances between atom pairs are calculated along the shortest path in the peptide's topology, and Gaussian functions centered on these distances are used to generate a 136-dimensional chemical space. This fingerprinting method enables the exploration of peptides with unknown or flexible 3D structures, making it particularly suited for studying unconventional topologies like bicyclic peptides.201
It has been demonstrated that some 2D fingerprints can effectively distinguish between peptide-like molecules with varying degrees of biological activity. Eckert and Bajorath (2007) found that Molprint2D performed best in recovering active molecules with strong peptide character. However, the property descriptor-based fingerprint excelled in identifying compounds with lower peptide character, indicating its utility in transitioning from peptide-like compounds to non-peptide alternatives.45 Capecchi et al. (2020) developed the MAP4 which represents the relationships between pairs of atoms in a molecule, considering their types and the topological distance. This fingerprint was designed to handle large and complex molecules, such as peptides, proteins, and peptide-like compounds, while maintaining computational efficiency.52 Recently, Capecchi and Reymond (2021) used a genetic algorithm with the molecular fingerprint MAP4 to represent the chemical space of peptides, organizing them by sequence and size. The chemical space represents 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 531 peptides from eleven open-access peptide and peptide-containing databases, and the map obtained categorizes the peptides by activity type, indicating that the majority of the peptides in the investigated databases, comprising 17
531 peptides from eleven open-access peptide and peptide-containing databases, and the map obtained categorizes the peptides by activity type, indicating that the majority of the peptides in the investigated databases, comprising 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 260 sequences, or 43% of the total, are classified as antimicrobial and anticancer.6 The Reymond group also developed MAP4C, a chiral adaptation of the MAP4 fingerprint, to analyze the stereochemical properties of large molecules, such as peptides. This fingerprint generates MinHashes derived from character strings encoding the SMILES representations of all pairs of circular substructures with diameters of up to four bonds and the shortest topological distance between their central atoms. The MAP4C incorporates Cahn–Ingold–Prelog (CIP) annotations (R, S, r, or s) for chiral atoms at the center of circular substructures, uses a question mark for undefined stereocenters, and includes cis–trans information for double bonds when specified. In non-stereoselective virtual screening approaches, MAP4C performs slightly better than the achiral MAP4, ECFP, and AP fingerprints.202 To evaluate the chemical space of antimicrobial peptides (AMPs) and identify new candidates with therapeutic potential, Orsi et al. (2024)190 integrated cheminformatics, ligand-based virtual screening, and machine-learning techniques. Virtual peptide libraries, including bicyclic and dendritic structures, were constructed and analyzed using molecular fingerprints, such as MAP4 and its chiral variant, MAP4C. These fingerprints measure molecular similarities and facilitate the visualization of the chemical space through dimensionality reduction methods, including PCA. The ligand-based virtual screening was employed to prioritize AMP candidates based on their similarity or diversity to known bioactive molecules, significantly enhancing the efficiency and success rate compared to random selection. Furthermore, machine-learning models, such as support vector machines and recurrent neural networks, were trained on experimental AMP datasets to predict antimicrobial activity and toxicity, aiding in identifying promising peptides for experimental validation.190
260 sequences, or 43% of the total, are classified as antimicrobial and anticancer.6 The Reymond group also developed MAP4C, a chiral adaptation of the MAP4 fingerprint, to analyze the stereochemical properties of large molecules, such as peptides. This fingerprint generates MinHashes derived from character strings encoding the SMILES representations of all pairs of circular substructures with diameters of up to four bonds and the shortest topological distance between their central atoms. The MAP4C incorporates Cahn–Ingold–Prelog (CIP) annotations (R, S, r, or s) for chiral atoms at the center of circular substructures, uses a question mark for undefined stereocenters, and includes cis–trans information for double bonds when specified. In non-stereoselective virtual screening approaches, MAP4C performs slightly better than the achiral MAP4, ECFP, and AP fingerprints.202 To evaluate the chemical space of antimicrobial peptides (AMPs) and identify new candidates with therapeutic potential, Orsi et al. (2024)190 integrated cheminformatics, ligand-based virtual screening, and machine-learning techniques. Virtual peptide libraries, including bicyclic and dendritic structures, were constructed and analyzed using molecular fingerprints, such as MAP4 and its chiral variant, MAP4C. These fingerprints measure molecular similarities and facilitate the visualization of the chemical space through dimensionality reduction methods, including PCA. The ligand-based virtual screening was employed to prioritize AMP candidates based on their similarity or diversity to known bioactive molecules, significantly enhancing the efficiency and success rate compared to random selection. Furthermore, machine-learning models, such as support vector machines and recurrent neural networks, were trained on experimental AMP datasets to predict antimicrobial activity and toxicity, aiding in identifying promising peptides for experimental validation.190
Some molecular fingerprints applied for peptide analyses are described in Table 5. We focused on explaining the most commonly used fingerprints, which are applied for the analysis of peptides and are usually accessible on the most used C++, Java, and Python libraries, such as Scikit-fingerprints,203 RDKit,21 iFeatureOmega,22 and Open Babel.204
| Molecular fingerprint | Description | Category | Implemented libraries or websites |
|---|---|---|---|
| MinHashed atom-pair up to a diameter of four bonds fingerprint (MAP4)52 | The circular substructures (radii r = 1 and r = 2) around each atom in an atom pair are represented as two SMILES pairs linked by the topological distance between the central atoms. These atom-pair molecular shingles are hashed and then undergo MinHashing. | Circular and path-based fingerprint (subclass atom-pair) | scikit-fingerprints GitHub (https://github.com/reymond-group/map4) |
| Chiral MAP4 (MAP4C)6 | Chiral representation of the MAP4 fingerprint | String-based and path-based fingerprint | GitHub (https://github.com/reymond-group/mapchiral) |
| Molecular ACCess System (MACCS Key) | Consists of a fixed-length bit vector, typically 166 bits. Encodes molecules' predefined substructures or functional groups, such as rings, bonds, and specific atom types. | Substructure fingerprint | RDKit, OpeBabel, and scikit-fingerprints |
| Extended Connectivity Fingerprint 6/4 (ECFP6, ECFP4) | Encodes the environment of each atom circularly, capturing information about the atom and its neighboring atoms up to a specified radius. | Path-based fingerprint | RDKit, scikit-fingerprints, OpenBabel, and iFeatureOmega |
| Extended-connectivity count fingerprint (ECFC6) | A variant of ECFPs that not only indicates the presence of specific substructures but also counts the occurrences of each substructure within the molecule. The “6” refers to the maximum diameter considered during the fingerprint generation. | Path-based fingerprint | RDKit, scikit-fingerprints, and OpenBabel |
| DompeKeys205 | Set of substructure-based fingerprint descriptors designed to encode patterns of functional groups and chemical features within molecular structures. | Substructure fingerprint | Developer website (https://dompekeys.exscalate.eu), |
| MolPrint2D206 | Encodes molecular structures by representing the atom environment up to a specific distance. It generates exhaustive lists of substructures surrounding each atom, which are then indexed for similarity comparison. | Circular fingerprint | OpenBabel |
| Macromolecule eXtended FingerPrint (MXFP)192 | A 217-dimensional fuzzy fingerprint representing atom pairs from seven pharmacophore groups, which is ideal for comparing large molecules and facilitating scaffold hopping. | Pharmacophore fingerprint | Developer GitHub (https://github.com/markusorsi/mxfp_python) |
Embeddings for peptides
Embeddings are continuous numerical representations of discrete elements useful to compress conformation, shape, physicochemical features, and context-dependent exposure of polar and hydrophobic groups from amino-acid sequences or molecular graph-based representations related to peptides in a meaningful way for modern machine and deep learning algorithms.207–209 These embeddings can be derived from learned encoders, such as graph neural networks (GNNs), language models (LMs), and autoencoders (AEs), which extract informative features from raw sequence or structural data during training.210 Embeddings aim to produce representations of molecules in which similarity relationships and structural patterns become computationally exploitable.207,211 The embeddings support the training of machine-learning models, clustering methods, and the performance of similarity-driven analysis.131,212,213In peptide modeling, embeddings are generated through neural or deep-learning encoders trained on sequence and/or structural data. These encoders learn internal representations that capture regularities in biochemical composition, residue interactions, structural motifs, and context-dependent effects.207,209 The resulting latent space reflects patterns discovered from the data and organizes peptides according to shared properties and structural similarity.214,215 A well-trained embedding preserves chemically relevant information while structuring peptides in a way that facilitates similarity analysis, interpolation, clustering, screening, and predictive modeling.210,216 A schematic overview of an embedding-based workflow applied to peptide sciences is shown in Fig. 6.
The choice of representation of peptides is especially important because these molecules can be encoded at different hierarchical levels. A growing body of peptide cheminformatics literature emphasizes that amino acid-based representations often better reflect the functional building blocks of peptides than purely atom-level descriptions, since peptide activity is frequently driven by residue identity, order, and context.131 Accordingly, residue-level notations such as FASTA,128 PLN,217 HELM,130 and BILN129 are highly relevant for embedding workflows. Although FASTA provides a simple encoding for canonical peptide sequences, the PLN extends the representation to a broader range of modified peptides, and HELM offers a richer formalism for complex biomolecules, including cyclic, branched, and crosslinked peptides, while BILN improves the human readability of HELM-derived peptide descriptions.128,130
A good encoding scheme preserves the relevant chemical content while producing inputs that are compatible with modern deep-learning pipelines.218 For residue-based representations, the usual workflow begins by tokenizing the sequence into amino acids or modified monomers, followed by conversion into machine-readable vectors. In the simplest case, one-hot encoding represents each residue by a sparse binary vector, typically defined over the alphabet of canonical amino acids, although the vocabulary can be expanded to include non-canonical residues and common chemical modifications. More informative sequence encoders instead map each token to a dense learned vector, allowing the model to capture contextual dependencies, long-range interactions, and position-dependent effects across the peptide chain. In parallel, some peptide-focused pipelines use property-informed residue encodings, in which each amino acid is represented by physicochemical descriptors such as hydrophobicity, charge, steric parameters, or polarity-related indices, thereby injecting biochemical priors that can be especially helpful when data are limited or when interpretability is desired.
Atom-based molecular representation formed by strings of characters, such as tokenized SMILES, peptide/biopolymer notations like CHUCKLES (monomer-sequence SMILES translation), or robust grammars such as SELFIES, a typical workflow starts by converting the string into tokens and then mapping those tokens to numerical vectors suitable for neural networks.3 These tokens are processed by neural architectures that can capture contextual dependencies along the sequence, such as long-range residue interactions and position-dependent effects. Through training, the model internalizes statistical regularities of sequence composition and structural tendencies, yielding embeddings that reflect both local and global sequence organization (Fig. 7, panel A). Neural architectures specifically designed for graph processing operate directly on these topological structures, learning embeddings that encode local chemical environments as well as global connectivity patterns.219 These representations incorporate structural constraints, stereochemical relationships, and bond-level information, making them particularly suitable for computational tasks. In the one-hot encoding, each symbol in the amino acid alphabet is represented by a sparse binary vector with a single “1” at the index of that symbol and “0” elsewhere, and each peptide is represented by a vector of length N, where N is often the 20 canonical amino acids, but it can be expanded to include non-canonical residues and common chemical modifications. In parallel, peptide-focused pipelines often incorporate property-informed encodings, where each residue is mapped to a vector of physicochemical descriptors (e.g., hydrophobicity, charge-related indices, steric parameters, etc.). These encodings inject biochemical priors that can be helpful when data are limited or when interpretability of residue contributions is desired.220,221 For graph-based representations of peptides, commonly derived from structural formats such as SDF, MOL, MOL2, and CDX, the atoms are represented as nodes and the bonds as edges, the encoding typically requires building an adjacency matrix to capture atomic connectivity and a node feature matrix to describe atom-level attributes; in many cases, edge features are added as well to encode bond properties such as type, order, or aromaticity (Fig. 7, panel B).212
For interpretability and exploratory analysis, low-dimensional projection techniques such as PCA, UMAP, or t-SNE may be applied to the higher-dimensional embeddings (latent feature space). These methods are used solely for visualization purposes, enabling qualitative assessment of similarity relationships, neighborhood structures, and clustering tendencies.222,223 They do not define the embedding itself; rather, they provide a reduced-dimensional view of the latent space. The chemically meaningful representation remains in the higher-dimensional latent space produced by the trained neural encoder.222,223
Recent computational workflows increasingly rely on embeddings from learned representations rather than other classes of descriptors, because embeddings can encode sequence context in a way that better reflects both biological function and indirectly structural constraints.214,224 This shift was catalyzed by large protein language models (pLMs)225 such as ESM-1b207 and the Rostlab ProtTrans family (e.g., ProtBERT226 and ProtT5209), which transform a protein or peptide sequence into dense vectors that summarize informative patterns across the entire chain. In practice, these models are pretrained with self-supervised objectives, learning the likelihood of residues given their surrounding context and producing contextualized embeddings at the residue and/or sequence level.227 Because pLMs are trained on massive sequence collections such as UniRef50,228 the resulting embeddings can be reused as general-purpose features to be implemented in different computational tasks.229,230 Importantly, recent studies indicate that pLM-derived embeddings can also be effective for peptides, frequently matching or outperforming traditional representations based on composition and physicochemical descriptors in predictive modeling and similarity analyses.230,231 Recently, specific learned representations for peptides have been developed to predict peptide properties, including PeptideCLM229 and Multi-Peptide.232
The chemical spaces overlappings and molecular determinants of peptide bioactivities
Studies have revealed unexpected overlaps in the bioactivities of certain unrelated peptide classes,47,48,57 thus suggesting that previously distinct chemical spaces may share molecular similarities. QSPs, CPPs, and B3PPs are examples of such peptide classes.Quorum-sensing peptides are signaling molecules that enable communication within bacterial communities and coordinate their behavior based on population density. The QSPs regulate various physiological activities, including biofilm formation and virulence factor production. These peptides play a crucial role in this communication, often functioning as autoinducers that bind to specific receptors on neighboring cells, triggering a cascade of gene expression changes.47,124 Several studies have focused on analyzing their molecular properties to create prediction models of these peptides.124,151 However, these molecules have also been shown to have selective BBBP properties48 as well as to interact with mammalian cells, selectively promoting cancer metastasis,173,233 influencing immune,234 and muscle235 cells. According to Wynendaele et al. (2015), the chemical space of quorum-sensing peptides is divided into three main clusters, as indicated by analyses of principal components. The peptide size and compactness comprise the first cluster. The descriptors that illustrate these characteristics include the radial distribution function (RDF), Burden eigenvalues (BEH, BEL), Randic shape indices, autocorrelation descriptors (ATS, GATS), weighted holistic invariant molecular (WHIM) descriptors, Balaban index, and the lopping centric index. Furthermore, the chemical space is also influenced by lipophilicity and hydrophilicity, evaluated through log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values, tPSA, and the counts of HBD and HBA, along with connectivity indices that account for peptide cyclization and descriptors related to HOMA, AROM, and ARR aromaticity, which define the second principal component. The third principal component is characterized by S-evaluating descriptors representing thiol groups, thiolactones, or disulfides, while the fourth principal component emphasizes the presence and frequency of nitrogen bonds (N–N, N–O, and N–C). As a result, peptides high in cysteine and methionine cluster together, whereas those with basic amino acids and amides, such as asparagine form another cluster. Investigating the brain influx and efflux properties of three chemically diverse QSPs, Wynendaele et al. (2015) identified, according to clustering of the PCA results, three peptides named PhrCACET1, BIP-2, and PhrANTH2. These QSPs were investigated using a multiple-time regression technique in an in vivo mouse model (ICR-CD-1) to assess blood–brain transfer characteristics. The authors discovered that these peptides show blood–brain barrier (BBB) permeation, as well. The PhrCACET1 exhibited a notably high initial influx into the mouse brain (Kin = 20.87 µl g−1 min−1), whereas the brain penetrabilities of BIP-2 and PhrANTH2 were determined to be low (Kin = 2.68 µl g−1 min−1) and very low (Kin = 0.18 µl g−1 min−1), respectively.48 These findings directly implicate the chemical characterization of peptide space and demonstrate the existence of an intersection not been characterized previously.
P values, tPSA, and the counts of HBD and HBA, along with connectivity indices that account for peptide cyclization and descriptors related to HOMA, AROM, and ARR aromaticity, which define the second principal component. The third principal component is characterized by S-evaluating descriptors representing thiol groups, thiolactones, or disulfides, while the fourth principal component emphasizes the presence and frequency of nitrogen bonds (N–N, N–O, and N–C). As a result, peptides high in cysteine and methionine cluster together, whereas those with basic amino acids and amides, such as asparagine form another cluster. Investigating the brain influx and efflux properties of three chemically diverse QSPs, Wynendaele et al. (2015) identified, according to clustering of the PCA results, three peptides named PhrCACET1, BIP-2, and PhrANTH2. These QSPs were investigated using a multiple-time regression technique in an in vivo mouse model (ICR-CD-1) to assess blood–brain transfer characteristics. The authors discovered that these peptides show blood–brain barrier (BBB) permeation, as well. The PhrCACET1 exhibited a notably high initial influx into the mouse brain (Kin = 20.87 µl g−1 min−1), whereas the brain penetrabilities of BIP-2 and PhrANTH2 were determined to be low (Kin = 2.68 µl g−1 min−1) and very low (Kin = 0.18 µl g−1 min−1), respectively.48 These findings directly implicate the chemical characterization of peptide space and demonstrate the existence of an intersection not been characterized previously.
Similarly, the CPPs have been identified with blood–brain barrier permeation.47 For example, de Oliveira et al. (2021) identified that CPPs possess higher MW, tPSA, and NRB values compared to clinically approved peptides, suggesting that their mechanisms of membrane penetration may involve processes beyond passive diffusion, such as pore formation or endocytosis. Additionally, their findings emphasize the importance of molecular flexibility and specific structural features, such as hydrogen bond patterns and the presence of aromatic rings, in influencing the permeability of these peptides, which could be related to their stereoselectivity. Regarding the B3PPs, Cavaco et al. (2024) recently identified key molecular determinants for peptides effectively crossing the BBB: a slightly hydrophobic nature, with a mean hydrophobic residue content of approximately 35%; a small size, with an average molecular weight of 2046 g mol−1; few or no aromatic residues, indicated by an average molar absorptivity of 3790 M−1 cm−1 at 280 nm, which corresponds to 1–2 tyrosine or 0–1 tryptophan residues; and a slightly cationic charge, with an average net charge of +2. The study emphasizes that not all CPPs can function as B3PPs, as the overlap between these two families is minimal. Experimental validation demonstrated that four newly identified B3PPs exhibited high translocation abilities in vitro and greater brain accumulation in vivo than established B3PPs, highlighting the importance of specific physicochemical characteristics for effective brain targeting.236
Despite the cell membrane and BBB having highly diverse functional and chemical compositions, and various molecular mechanisms of permeation being described for molecules into these membranes, most predictive models typically attribute passive transport through the membrane as the most important mechanism.58 Furthermore, the biophysical interaction with these membranes has been explored as a key factor for permeation. Therefore, it is interesting to note that some molecular features usually applied to predict the CPPs have also been pointed out as relevant to predicting B3PPs. For example, Dichiara et al. (2019) established a set of chemical descriptors to facilitate the successful prediction of BBB permeation. They evaluated statistically 328 compounds, correlating their experimental in vivo log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BB values with various computed descriptors. They constructed contingency tables, calculated observed and expected distributions, and analyzed the relationships between descriptors and BBB permeation. The authors identified a significant influence of nine specific physicochemical properties on BBB permeation, including polar surface area, nitrogen and oxygen count, log
BB values with various computed descriptors. They constructed contingency tables, calculated observed and expected distributions, and analyzed the relationships between descriptors and BBB permeation. The authors identified a significant influence of nine specific physicochemical properties on BBB permeation, including polar surface area, nitrogen and oxygen count, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, nitrogen count, log
P, nitrogen count, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D, oxygen count, ionization state, hydrogen bond acceptors, and hydrogen bond donors.237
D, oxygen count, ionization state, hydrogen bond acceptors, and hydrogen bond donors.237
Despite both classes representing distinct biological activity, a previous study showed that chemically distinct CPPs named pVEC, SynB3, Tat 47–57, transportan 10 (TP10), and TP10-2 exhibit varying abilities to enter the BBB. Specifically, Tat 47–57, SynB3, and pVEC demonstrated significantly high rates of unidirectional influx, whereas the transportan variants displayed minimal to low brain penetration.47
Final considerations
Exploring chemical space and the concept of a chemical multiverse for peptides can provide a robust framework for understanding the diverse properties and biological activities of these biomolecules, thereby opening new avenues for the identification of novel chemical entities in screening strategies as well as for the design of new bioactive peptides.14,238The concept of chemical space gains particular significance when applied to peptides because their amino acid sequence intrinsically encodes and influences physicochemical and structural properties such as solubility, hydrophobicity, folding patterns, and three-dimensional conformation, which ultimately shape their biological activities. The unique characteristics of peptides, especially their conformational flexibility, further underscore the complexity of their chemical space, as these molecules can adopt distinct conformational states depending on the environment, which is often crucial for their biological function. This flexibility contributes to their ability to interact with diverse biological targets and, in some cases, to penetrate biological barriers. The choice of molecular representation strongly determines which aspects of peptide behavior become computationally accessible. Classical molecular descriptors and fingerprints remain essential for interpretable, scalable, and cost-effective analyses, particularly in virtual screening strategies. However, recent advances in machine learning have expanded this landscape by enabling embeddings derived from learned encoders, such as GNNs, LMs, and AEs, which can extract informative features directly from raw sequence or structural data. These learned representations organize peptides in latent spaces where similarity, clustering patterns, and predictive relationships become more readily exploitable, thus providing a powerful complement to conventional descriptor-based strategies.
In addition, peptide bioactivity should not be interpreted independently of conformation and context. Features such as chameleonicity, secondary-structure propensity, backbone flexibility, polarity masking, and membrane-interaction mechanisms reinforce the notion that peptide function emerges from a dynamic relationship between structure and environment. This complexity also helps explain why apparently unrelated peptide classes may partially overlap in chemical space and bioactivity, revealing intersections that are biologically meaningful and potentially useful for peptide discovery, repurposing, and design. Moreover, the overlap among distinct peptide classes with pleiotropic activities, such as QSPs, CPPs, and B3PPs, suggests shared regions of chemical space that warrant further investigation and may reveal new peptide functions as well as biotechnological and therapeutic opportunities.
Author contributions
ECLO, LDN, AHLL, CS, BDS, and KS conceptualized the study. ECLO, JA, GPC, LDN, AHLL, CMFR, ADS, EW, CS, BDS, and KS contributed to data curation, analysis, interpretation of the results, and scientific discussion. ECLO, JA, GPC, LDN, AHLL, and CMFR contributed to the visualization and preparation of figures, tables, and graphical materials. KS coordinated the study, supervised the project, wrote the manuscript, and critically reviewed the text. CMFR contributed to manuscript improvement, revision of visual materials, and scientific refinement of the work. ADS, EW, LDN, CS, and BDS contributed to the critical review of the manuscript and the intellectual improvement of the study. All authors reviewed, edited, and approved the final version of the manuscript.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this article.Data availability
No new primary data were generated or analysed in this study. All data discussed are available in the cited literature.Acknowledgements
The research was funded by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – CAPES (ROR ID: 00x0ma614). K. S. is grateful for the National Council for Scientific and Technological Development (CNPq, grant numbers: 408367/2024-5 and 442559/2025-9), a Brazilian funding agency, for the financial support of the study. The article processing charge for the publication of this research was funded by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – CAPES/Brazil (ROR ID: 00x0ma614). The authors have assigned the Creative Commons CC BY license to any accepted article version for open access. K. S. is grateful for the National Council for Scientific and Technological Development (CNPq, grant numbers: 408367/2024-5 and 442559/2025-9), a Brazilian funding agency, for the financial support of the study. The authors would like to thank the illustrator Miguel Silva for his countless revisions, which were essential to achieving the technical and scientific level of detail presented in the illustrative schemes.References
- J. L. Medina-Franco and F. I. Saldívar-González, Cheminformatics to Characterize Pharmacologically Active Natural Products, Biomolecules, 2020, 10(11), 1566, DOI:10.3390/biom10111566
.
- K. Santana, L. D. do Nascimento, A. Lima e Lima, V. Damasceno, C. Nahum, R. C. Braga and J. Lameira, Applications of Virtual Screening in Bioprospecting: Facts, Shifts, and Perspectives to Explore the Chemo-Structural Diversity of Natural Products, Front. Chem., 2021, 9, 662688, DOI:10.3389/fchem.2021.662688
.
- L. David, A. Thakkar, R. Mercado and O. Engkvist, Molecular Representations in AI-Driven Drug Discovery: A Review and Practical Guide, J. Cheminf., 2020, 12(1), 56, DOI:10.1186/s13321-020-00460-5
.
- J.-L. Reymond, The Chemical Space Project, Acc. Chem. Res., 2015, 48(3), 722–730, DOI:10.1021/ar500432k
.
- B. I. Díaz-Eufracio, O. Palomino-Hernández, R. A. Houghten and J. L. Medina-Franco, Exploring the Chemical Space of Peptides for Drug Discovery: A Focus on Linear and Cyclic Penta-Peptides, Mol. Diversity, 2018, 22(2), 259–267, DOI:10.1007/s11030-018-9812-9
.
- A. Capecchi and J.-L. L. Reymond, Peptides in Chemical Space, Med. Drug Discovery, 2021, 9, 100081, DOI:10.1016/j.medidd.2021.100081
.
- P. Schwaller, A. C. Vaucher, R. Laplaza, C. Bunne, A. Krause, C. Corminboeuf and T. Laino, Machine Intelligence for Chemical Reaction Space, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2022, 12(5), e1604, DOI:10.1002/wcms.1604
.
- M. Wang, S. Li, J. Wang, O. Zhang, H. Du, D. Jiang, Z. Wu, Y. Deng, Y. Kang, P. Pan, D. Li, X. Wang, X. Yao, T. Hou and C.-Y. Hsieh, ClickGen: Directed Exploration of Synthesizable Chemical Space via Modular Reactions and Reinforcement Learning, Nat. Commun., 2024, 15(1), 10127, DOI:10.1038/s41467-024-54456-y
.
- H. Kim, S. Ryu, N. Jung, J. Yang and C. Seok, CSearch: Chemical Space Search via Virtual Synthesis and Global Optimization, J. Cheminf., 2024, 16(1), 137, DOI:10.1186/s13321-024-00936-8
.
- K. Hansen, F. Biegler, R. Ramakrishnan, W. Pronobis, O. A. von Lilienfeld, K.-R. Müller and A. Tkatchenko, Machine Learning Predictions of Molecular Properties: Accurate Many-Body Potentials and Nonlocality in Chemical Space, J. Phys. Chem. Lett., 2015, 6(12), 2326–2331, DOI:10.1021/acs.jpclett.5b00831
.
- P. Gorai, P. Parilla, E. S. Toberer and V. Stevanović, Computational Exploration of the Binary A 1 B 1 Chemical Space for Thermoelectric Performance, Chem. Mater., 2015, 27(18), 6213–6221, DOI:10.1021/acs.chemmater.5b01179
.
- A. M. Mroz, V. Posligua, A. Tarzia, E. H. Wolpert and K. E. Jelfs, Into the Unknown: How Computation Can Help Explore Uncharted Material Space, J. Am. Chem. Soc., 2022, 144(41), 18730–18743, DOI:10.1021/jacs.2c06833
.
- A. Tudi, Z. Li, C. Xie, T. Baiheti, E. Tikhonov, F. Zhang, S. Pan and Z. Yang, Functional Modules Map of Unexplored Chemical Space: Guiding the Discovery of Giant Birefringent Materials, Adv. Funct. Mater., 2024, 34(51), 2409716, DOI:10.1002/adfm.202409716
.
- I. Di Bonaventura, S. Baeriswyl, A. Capecchi, B.-H. Gan, X. Jin, T. N. Siriwardena, R. He, T. Köhler, A. Pompilio, G. Di Bonaventura, C. van Delden, S. Javor and J.-L. Reymond, An Antimicrobial Bicyclic Peptide from Chemical Space against Multidrug Resistant Gram-Negative Bacteria, Chem. Commun., 2018, 54(40), 5130–5133, 10.1039/C8CC02412J
.
- E. C. L. de Oliveira, K. Santana, L. Josino, A. H. Lima e Lima and C. de Souza de Sales Júnior, Predicting Cell-Penetrating Peptides Using Machine Learning Algorithms and Navigating in Their Chemical Space, Sci. Rep., 2021, 11(1), 7628, DOI:10.1038/s41598-021-87134-w
.
- H. L. Barazorda-Ccahuana, K. E. Juárez-Mercado, J. L. Medina-Franco and M. A. Chavez-Fumagalli, Visualizing and Analyzing the Chemical Space of Natural Product Databases for Drug Discovery, J. Visualized Exp., 2024, 211, e66349, DOI:10.3791/66349
.
- F. I. Saldívar-González, E. Lenci, A. Trabocchi and J. L. Medina-Franco, Exploring the Chemical Space and the Bioactivity Profile of Lactams: A Chemoinformatic Study, RSC Adv., 2019, 9(46), 27105–27116, 10.1039/C9RA04841C
.
- E. López-López, J. P. Sánchez-Castañeda, M. S. Martinez-Cortés, C. de la Fuente-Nunez and J. L. Medina-Franco, Exploring and Expanding the Chemical Multiverse of Peptides, Chem. Sci., 2026, 17(3), 1461–1479, 10.1039/D5SC04465K
.
- V. Digiesi, V. de la Oliva Roque, M. Vallaro, G. Caron and G. Ermondi, Permeability Prediction in the Beyond-Rule-of 5 Chemical Space: Focus on Cyclic Hexapeptides, Eur. J. Pharm. Biopharm., 2021, 165, 259–270, DOI:10.1016/j.ejpb.2021.05.017
.
- J. M. Pelton, J. E. Hochuli, P. W. Sadecki, T. Katoh, H. Suga, L. M. Hicks, E. N. Muratov, A. Tropsha and A. A. Bowers, Cheminformatics-Guided Cell-Free Exploration of Peptide Natural Products, J. Am. Chem. Soc., 2024, 146(12), 8016–8030, DOI:10.1021/jacs.3c11306
.
- M. Lovrić, J. M. Molero and R. Kern, PySpark and RDKit: Moving towards Big Data in Cheminformatics, Mol. Inf., 2019, 38(6), 1800082, DOI:10.1002/minf.201800082
.
- Z. Chen, X. Liu, P. Zhao, C. Li, Y. Wang, F. Li, T. Akutsu, C. Bain, R. B. Gasser, J. Li, Z. Yang, X. Gao, L. Kurgan and J. Song, IFeatureOmega: An Integrative Platform for Engineering, Visualization and Analysis of Features from Molecular Sequences, Structural and Ligand Data Sets, Nucleic Acids Res., 2022, 50(W1), W434–W447, DOI:10.1093/nar/gkac351
.
- Z. Chen, P. Zhao, F. Li, A. Leier, T. T. Marquez-Lago, Y. Wang, G. I. Webb, A. I. Smith, R. J. Daly, K.-C. Chou and J. Song, IFeature: A Python Package and Web Server for Features Extraction and Selection from Protein and Peptide Sequences, Bioinformatics, 2018, 34(14), 2499–2502, DOI:10.1093/bioinformatics/bty140
.
- H. Moriwaki, Y.-S. S. Tian, N. Kawashita and T. Takagi, Mordred: A Molecular Descriptor Calculator, J. Cheminf., 2018, 10(1), 4, DOI:10.1186/s13321-018-0258-y
.
- L. Van Der Maaten and G. Hinton, Visualizing Data Using T-SNE, J. Mach. Learn. Res., 2008, 9, 2579–2625 Search PubMed
.
- L. McInnes, J. Healy, N. Saul and L. Großberger, UMAP: Uniform Manifold Approximation and Projection, J. Open Source Software, 2018, 3(29), 861, DOI:10.21105/joss.00861
.
- M. Cihan Sorkun, D. Mullaj, J. M. V. A. Koelman and S. Er, ChemPlot, a Python Library for Chemical Space Visualization**, Chem. – Methods, 2022, 2(7), e202200005, DOI:10.1002/cmtd.202200005
.
- D. Probst and J.-L. Reymond, Visualization of Very Large High-Dimensional Data Sets as Minimum Spanning Trees, J. Cheminf., 2020, 12(1), 12, DOI:10.1186/s13321-020-0416-x
.
- S. Sosnin, Chemical Space Visual Navigation in the Era of Deep Learning and Big Data, Drug Discovery Today, 2025, 30(7), 104392, DOI:10.1016/j.drudis.2025.104392
.
- K. Castillo-Mendieta, Y. Marrero-Ponce, E. A. Márquez, J. L. García-Giménez, A. Antunes and G. Agüero-Chapin, Mapping the Antibiofilm Peptide Space with Similarity Networks and Curated Negative Sets, ACS Omega, 2025, 10(49), 60457–60476, DOI:10.1021/acsomega.5c07679
.
- B. Zhang, M. Vogt, G. M. Maggiora and J. Bajorath, Design of Chemical Space Networks Using a Tanimoto Similarity Variant Based upon Maximum Common Substructures, J. Comput.-Aided. Mol. Des., 2015, 29(10), 937–950, DOI:10.1007/s10822-015-9872-1
.
- C. Burri and R. Brun, Eflornithine for the Treatment of Human African Trypanosomiasis, Parasitol. Res., 2003, 90(Supp 1), S49–52, DOI:10.1007/s00436-002-0766-5
.
- D. de Llano García, Y. Marrero-Ponce, G. Agüero-Chapin, H. Rodríguez, F. J. Ferri, E. A. Márquez, J. R. Mora, F. Martinez-Rios and Y. Pérez-Castillo, Mapping the Chemical Space of Antiviral Peptides with Half-Space Proximal and Metadata Networks Through Interactive Data Mining, Computers, 2025, 14(10), 423, DOI:10.3390/computers14100423
.
- M. Zwierzyna, M. Vogt, G. M. Maggiora and J. Bajorath, Design and Characterization of Chemical Space Networks for Different Compound Data Sets, J. Comput.-Aided. Mol. Des., 2015, 29(2), 113–125, DOI:10.1007/s10822-014-9821-4
.
- K. Castillo-Mendieta, G. Agüero-Chapin, E. A. Marquez, Y. Perez-Castillo, S. J. Barigye, N. S. Vispo, C. R. García-Jacas and Y. Marrero-Ponce, Peptide Hemolytic Activity Analysis Using Visual Data Mining of Similarity-Based Complex Networks, npj Syst. Biol. Appl., 2024, 10(1), 115, DOI:10.1038/s41540-024-00429-2
.
- G. M. Maggiora and J. Bajorath, Chemical Space Networks: A Powerful New Paradigm for the Description of Chemical Space, J. Comput.-Aided. Mol. Des., 2014, 28(8), 795–802, DOI:10.1007/s10822-014-9760-0
.
- L. Aguilera-Mendoza, S. Ayala-Ruano, F. Martinez-Rios, E. Chavez, C. R. García-Jacas, C. A. Brizuela and Y. Marrero-Ponce, StarPep Toolbox: An Open-Source Software to Assist Chemical Space Analysis of Bioactive Peptides and Their Functions Using Complex Networks, Bioinformatics, 2023, 39(8), btad506, DOI:10.1093/bioinformatics/btad506
.
- S. Ayala-Ruano, Y. Marrero-Ponce, L. Aguilera-Mendoza, N. Pérez, G. Agüero-Chapin, A. Antunes and A. C. Aguilar, Network Science and Group Fusion Similarity-Based Searching to Explore the Chemical Space of Antiparasitic Peptides, ACS Omega, 2022, 7(50), 46012–46036, DOI:10.1021/acsomega.2c03398
.
- K. Castillo-Mendieta, G. Agüero-Chapin, J. R. Mora, N. Pérez, E. Contreras-Torres, J. R. Valdes-Martini, F. Martinez-Rios and Y. Marrero-Ponce, Unraveling the Hemolytic Toxicity Tapestry of Peptides Using Chemical Space Complex Networks, Toxicol. Sci., 2024, 202(2), 236–249, DOI:10.1093/toxsci/kfae115
.
- G. Agüero-Chapin, A. Antunes, J. R. Mora, N. Pérez, E. Contreras-Torres, J. R. Valdes-Martini, F. Martinez-Rios, C. H. Zambrano and Y. Marrero-Ponce, Complex Networks Analyses of Antibiofilm Peptides: An Emerging Tool for Next-Generation Antimicrobials’ Discovery, Antibiotics, 2023, 12(4), 747, DOI:10.3390/antibiotics12040747
.
- R. Kunimoto and J. Bajorath, Combining Similarity Searching and Network Analysis for the Identification of Active Compounds, ACS Omega, 2018, 3(4), 3768–3777, DOI:10.1021/acsomega.8b00344
.
- Y.-C. Lo, S. Senese, C.-M. Li, Q. Hu, Y. Huang, R. Damoiseaux and J. Z. Torres, Large-Scale Chemical Similarity Networks for Target Profiling of Compounds Identified in Cell-Based Chemical Screens, PLoS Comput. Biol., 2015, 11(3), e1004153, DOI:10.1371/journal.pcbi.1004153
.
- Q. Wang, X. Hu, Z. Wei, H. Lu and H. Liu, Reinforcement Learning-Driven Exploration of Peptide Space: Accelerating Generation of Drug-like Peptides, Briefings Bioinf., 2024, 25(5) DOI:10.1093/bib/bbae444
.
- J. L. Medina-Franco, A. L. Chávez-Hernández, E. López-López and F. I. Saldívar-González, Chemical Multiverse: An Expanded View of Chemical Space, Mol. Inf., 2022, 41(11) DOI:10.1002/minf.202200116
.
- H. Eckert and J. Bajorath, Exploring Peptide-Likeness of Active Molecules Using 2D Fingerprint Methods, J. Chem. Inf. Model., 2007, 47(4), 1366–1378, DOI:10.1021/ci700086m
.
- M. Orsi and J. Reymond, Navigating a 1E+60 Chemical Space of Peptide/Peptoid Oligomers, Mol. Inf., 2024, 44(1) DOI:10.1002/minf.202400186
.
- S. Stalmans, N. Bracke, E. Wynendaele, B. Gevaert, K. Peremans, C. Burvenich, I. Polis and B. De Spiegeleer, Cell-Penetrating Peptides Selectively Cross the Blood-Brain Barrier In Vivo, PLoS One, 2015, 10(10), e0139652, DOI:10.1371/journal.pone.0139652
.
- E. Wynendaele, F. Verbeke, S. Stalmans, B. Gevaert, Y. Janssens, C. Van De Wiele, K. Peremans, C. Burvenich and B. De Spiegeleer, Quorum Sensing Peptides Selectively Penetrate the Blood-Brain Barrier, PLoS One, 2015, 10(11), e0142071, DOI:10.1371/journal.pone.0142071
.
- J. J. Naveja and J. L. Medina-Franco, Finding Constellations in Chemical Space Through Core Analysis, Front. Chem., 2019, 7 DOI:10.3389/fchem.2019.00510
.
- S. R. Langdon, N. Brown and J. Blagg, Scaffold Diversity of Exemplified Medicinal Chemistry Space, J. Chem. Inf. Model., 2011, 51(9), 2174–2185, DOI:10.1021/ci2001428
.
- G. W. Bemis and M. A. Murcko, The Properties of Known Drugs. 1. Molecular Frameworks, J. Med. Chem., 1996, 39(15), 2887–2893, DOI:10.1021/jm9602928
.
- A. Capecchi, D. Probst and J.-L. Reymond, One Molecular Fingerprint to Rule Them All: Drugs, Biomolecules, and the Metabolome, J. Cheminf., 2020, 12(1), 43, DOI:10.1186/s13321-020-00445-4
.
- B. C. Doak, B. Over, F. Giordanetto and J. Kihlberg, Oral Druggable Space beyond the Rule of 5: Insights from Drugs and Clinical Candidates, Chem. Biol., 2014, 21(9), 1115–1142, DOI:10.1016/j.chembiol.2014.08.013
.
- A. T. Bockus, C. M. McEwen and R. S. Lokey, Form and Function in Cyclic Peptide Natural Products: A Pharmacokinetic Perspective, Curr. Top. Med. Chem., 2013, 13(7), 821–836, DOI:10.2174/1568026611313070005
.
- H. Sunde, K. Ryder, A. E.-D. A. Bekhit and A. Carne, Analysis of Peptides in a Sheep Beta Lactoglobulin Hydrolysate as a Model to Evaluate the Effect of Peptide Amino Acid Sequence on Bioactivity, Food Chem., 2021, 365, 130346, DOI:10.1016/j.foodchem.2021.130346
.
- B. Mishra, J. Lakshmaiah Narayana, T. Lushnikova, Y. Zhang, R. M. Golla, D. Zarena and G. Wang, Sequence Permutation Generates Peptides with Different Antimicrobial and Antibiofilm Activities, Pharmaceuticals, 2020, 13(10), 271, DOI:10.3390/ph13100271
.
- B. Gevaert, S. Stalmans, E. Wynendaele, L. Taevernier, N. Bracke, M. D’;Hondt and B. De Spiegeleer, Exploration of the Medicinal Peptide Space, Protein Pept. Lett., 2016, 23(4), 324–335, DOI:10.2174/0929866523666160215162326
.
- E. C. L. de Oliveira, K. S. da Costa, P. S. Taube, A. H. Lima and C. de S. de S. Junior, Biological Membrane-Penetrating Peptides: Computational Prediction and Applications, Front. Cell. Infect. Microbiol., 2022, 12 DOI:10.3389/fcimb.2022.838259
.
- M. Sánchez-Navarro, M. Teixidó and E. Giralt, Jumping Hurdles: Peptides Able to Overcome Biological Barriers, Acc. Chem. Res., 2017, 50(8), 1847–1854, DOI:10.1021/acs.accounts.7b00204
.
- J. E. Bock, J. Gavenonis and J. A. Kritzer, Getting in Shape: Controlling Peptide Bioactivity and Bioavailability Using Conformational Constraints, ACS Chem. Biol., 2013, 8(3), 488–499, DOI:10.1021/cb300515u
.
- S. Ramazi and J. Zahiri, Post-Translational Modifications in Proteins: Resources, Tools and Prediction Methods, Database, 2021, 2021 DOI:10.1093/database/baab012
.
- J. Miao, M. L. Descoteaux and Y. S. Lin, Structure Prediction of Cyclic Peptides by Molecular Dynamics + Machine Learning, Chem. Sci., 2021, 12(44), 14927–14936, 10.1039/d1sc05562c
.
- P. Petkov, E. Lilkova, N. Ilieva and L. Litov, Self-Association of Antimicrobial Peptides: A Molecular Dynamics Simulation Study on Bombinin, Int. J. Mol. Sci., 2019, 20(21), 5450, DOI:10.3390/ijms20215450
.
- J. R. Allison, Computational Methods for Exploring Protein Conformations, Biochem. Soc. Trans., 2020, 48(4), 1707–1724, DOI:10.1042/BST20200193
.
- G. Siligardi and A. F. Drake, The Importance of Extended Conformations and, in Particular, the P II Conformation for the Molecular Recognition of Peptides, Biopolymers, 1995, 37(4), 281–292, DOI:10.1002/bip.360370406
.
- C. Herrera-León, F. Ramos-Martín, H. El Btaouri, V. Antonietti, P. Sonnet, L. Martiny, F. Zevolini, C. Falciani, C. Sarazin and N. D’Amelio, The Influence of Short Motifs on the Anticancer Activity of HB43 Peptide, Pharmaceutics, 2022, 14(5), 1089, DOI:10.3390/pharmaceutics14051089
.
- M. A. Schmitt, B. Weisblum and S. H. Gellman, Interplay among Folding, Sequence, and Lipophilicity in the Antibacterial and Hemolytic Activities of α/β-Peptides, J. Am. Chem. Soc., 2007, 129(2), 417–428, DOI:10.1021/ja0666553
.
- R. N. Chapman, G. Dimartino and P. S. Arora, A Highly Stable Short α-Helix Constrained by a Main-Chain Hydrogen-Bond Surrogate, J. Am. Chem. Soc., 2004, 126(39), 12252–12253, DOI:10.1021/ja0466659
.
- S. E. Miller, N. R. Kallenbach and P. S. Arora, Reversible α-Helix Formation Controlled by a Hydrogen Bond Surrogate, Tetrahedron, 2012, 68(23), 4434–4437, DOI:10.1016/j.tet.2011.12.068
.
- T. Marcelo Der Torossian, A. F. Silva, F. L. Alves, M. L. Capurro, A. Miranda and O. Vani Xavier, Highly Potential Antiplasmodial Restricted Peptides, Chem. Biol. Drug Des., 2015, 85(2), 163–171, DOI:10.1111/cbdd.12354
.
- M. Der Torossian Torres, A. F. Silva, F. L. Alves, M. L. Capurro, A. Miranda and V. X. Oliveira Junior, The Importance of Ring Size and Position for the Antiplasmodial Activity of Angiotensin II Restricted Analogs, Int. J. Pept. Res. Ther., 2014, 20(3), 277–287, DOI:10.1007/s10989-014-9392-1
.
- F. Milletti, Cell-Penetrating Peptides: Classes, Origin, and Current Landscape, Drug Discovery Today, 2012, 17(15–16), 850–860, DOI:10.1016/j.drudis.2012.03.002
.
- B. Oller-Salvia, M. Sánchez-Navarro, E. Giralt and M. Teixidó, Blood–Brain Barrier Shuttle Peptides: An Emerging Paradigm for Brain Delivery, Chem. Soc. Rev., 2016, 45(17), 4690–4707, 10.1039/C6CS00076B
.
- T. A. Ramelot, J. Palmer, G. T. Montelione and G. Bhardwaj, Cell-Permeable Chameleonic Peptides: Exploiting Conformational Dynamics in de Novo Cyclic Peptide Design, Curr. Opin. Struct. Biol., 2023, 80, 102603, DOI:10.1016/j.sbi.2023.102603
.
- C. D. Payne, B. Franke, M. F. Fisher, F. Hajiaghaalipour, C. E. McAleese, A. Song, C. Eliasson, J. Zhang, A. S. Jayasena, G. Vadlamani, R. J. Clark, R. F. Minchin, J. S. Mylne and K. J. Rosengren, A Chameleonic Macrocyclic Peptide with Drug Delivery Applications, Chem. Sci., 2021, 12(19), 6670–6683, 10.1039/D1SC00692D
.
- D. Lee, J. Choi, M. J. Yang, C.-J. Park and J. Seo, Controlling the Chameleonic Behavior and Membrane Permeability of Cyclosporine Derivatives via Backbone and Side Chain Modifications, J. Med. Chem., 2023, 66(18), 13189–13204, DOI:10.1021/acs.jmedchem.3c01140
.
- S. M. Linker, C. Schellhaas, A. S. Kamenik, M. M. Veldhuizen, F. Waibl, H.-J. Roth, M. Fouché, S. Rodde and S. Riniker, Lessons for Oral Bioavailability: How Conformationally Flexible Cyclic Peptides Enter and Cross Lipid Membranes, J. Med. Chem., 2023, 66(4), 2773–2788, DOI:10.1021/acs.jmedchem.2c01837
.
- P. Ertl, B. Rohde and P. Selzer, Fast Calculation of Molecular Polar Surface Area as a Sum of Fragment-Based Contributions and Its Application to the Prediction of Drug Transport Properties, J. Med. Chem., 2000, 43(20), 3714–3717, DOI:10.1021/jm000942e
.
- P. Matsson, B. C. Doak, B. Over and J. Kihlberg, Cell Permeability beyond the Rule of 5, Adv. Drug Delivery Rev., 2016, 101, 42–61, DOI:10.1016/j.addr.2016.03.013
.
- J. M. Galúcio, E. F. Monteiro, D. A. de Jesus, C. H. Costa, R. C. Siqueira, G. B. dos Santos, J. Lameira and K. S. da Costa, In Silico Identification of Natural Products with Anticancer Activity Using a Chemo-Structural Database of Brazilian Biodiversity, Comput. Biol. Chem., 2019, 83, 107102, DOI:10.1016/j.compbiolchem.2019.107102
.
- M. Rossi Sebastiano, B. C. Doak, M. Backlund, V. Poongavanam, B. Over, G. Ermondi, G. Caron, P. Matsson and J. Kihlberg, Impact of Dynamically Exposed Polarity on Permeability and Solubility of Chameleonic Drugs beyond the Rule of 5, J. Med. Chem., 2018, 61(9), 4189–4202, DOI:10.1021/acs.jmedchem.8b00347
.
- A. Daina and V. Zoete, A BOILED-Egg To Predict Gastrointestinal Absorption and Brain Penetration of Small Molecules, ChemMedChem, 2016, 1117–1121, DOI:10.1002/cmdc.201600182
.
- C. A. S. Bergström, W. N. Charman and C. J. H. Porter, Computational Prediction of Formulation Strategies for Beyond-Rule-of-5 Compounds, Adv. Drug Delivery Rev., 2016, 101, 6–21, DOI:10.1016/j.addr.2016.02.005
.
- D. F. Veber, S. R. Johnson, H.-Y. Y. Cheng, B. R. Smith, K. W. Ward and K. D. Kopple, Molecular Properties That Influence the Oral Bioavailability of Drug Candidates, J. Med. Chem., 2002, 45(12), 2615–2623, DOI:10.1021/jm020017n
.
- P. Matsson and J. Kihlberg, How Big Is Too Big for Cell Permeability?, J. Med. Chem., 2017, 60(5), 1662–1664, DOI:10.1021/acs.jmedchem.7b00237
.
- C. R. W. Guimarães, A. M. Mathiowetz, M. Shalaeva, G. Goetz and S. Liras, Use of 3D Properties to Characterize Beyond Rule-of-5 Property Space for Passive Permeation, J. Chem. Inf. Model., 2012, 52(4), 882–890, DOI:10.1021/ci300010y
.
- A. Whitty, M. Zhong, L. Viarengo, D. Beglov, D. R. Hall and S. Vajda, Quantifying the Chameleonic Properties of Macrocycles and Other High-Molecular-Weight Drugs, Drug Discovery Today, 2016, 21(5), 712–717, DOI:10.1016/j.drudis.2016.02.005
.
- N. El Tayar, A. E. Mark, P. Vallat, R. M. Brunne, B. Testa and W. F. van Gunsteren, Solvent-Dependent Conformation and Hydrogen-Bonding Capacity of Cyclosporin A: Evidence from Partition Coefficients and Molecular Dynamics Simulations, J. Med. Chem., 1993, 36(24), 3757–3764, DOI:10.1021/jm00076a002
.
- E. Eiríksdóttir, K. Konate, Ü. Langel, G. Divita and S. Deshayes, Secondary Structure of Cell-Penetrating Peptides Controls Membrane Interaction and Insertion, Biochim. Biophys. Acta, Biomembr., 2010, 1798(6), 1119–1128, DOI:10.1016/j.bbamem.2010.03.005
.
- J. S. Appelbaum, J. R. LaRochelle, B. A. Smith, D. M. Balkin, J. M. Holub and A. Schepartz, Arginine Topology Controls Escape of Minimally Cationic Proteins from Early Endosomes to the Cytoplasm, Chem. Biol., 2012, 19(7), 819–830, DOI:10.1016/j.chembiol.2012.05.022
.
- G. H. Bird, E. Mazzola, K. Opoku-Nsiah, M. A. Lammert, M. Godes, D. S. Neuberg and L. D. Walensky, Biophysical Determinants for Cellular Uptake of Hydrocarbon-Stapled Peptide Helices, Nat. Chem. Biol., 2016, 12(10), 845–852, DOI:10.1038/nchembio.2153
.
- H. Yamashita, M. Oba, T. Misawa, M. Tanaka, T. Hattori, M. Naito, M. Kurihara and Y. Demizu, A Helix-Stabilized Cell-Penetrating Peptide as an Intracellular Delivery Tool, ChemBioChem, 2016, 17(2), 137–140, DOI:10.1002/cbic.201500468
.
- T. R. White, C. M. Renzelman, A. C. Rand, T. Rezai, C. M. McEwen, V. M. Gelev, R. A. Turner, R. G. Linington, S. S. F. Leung, A. S. Kalgutkar, J. N. Bauman, Y. Zhang, S. Liras, D. A. Price, A. M. Mathiowetz, M. P. Jacobson and R. S. Lokey, On-Resin N-Methylation of Cyclic Peptides for Discovery of Orally Bioavailable Scaffolds, Nat. Chem. Biol., 2011, 7(11), 810–817, DOI:10.1038/nchembio.664
.
- J. A. Seixas Feio, E. C. L. de Oliveira, C. de S. de Sales, K. S. da Costa and A. H. L. e Lima, Investigating Molecular Descriptors in Cell-Penetrating Peptides Prediction with Deep Learning: Employing N, O, and Hydrophobicity According to the Eisenberg Scale, PLoS One, 2024, 19(6), e0305253, DOI:10.1371/journal.pone.0305253
.
- G. B. Santos, A. Ganesan and F. S. Emery, Oral Administration of Peptide-Based Drugs: Beyond Lipinski's Rule, ChemMedChem, 2016, 11(20), 2245–2251, DOI:10.1002/cmdc.201600288
.
- C. A. Grambow, H. Weir, C. N. Cunningham, T. Biancalani and K. V. Chuang, CREMP: Conformer-Rotamer Ensembles of Macrocyclic Peptides for Machine Learning, Sci. Data, 2024, 11(1), 859, DOI:10.1038/s41597-024-03698-y
.
- J. Wang, Z. Liu, S. Zhao, T. Xu, H. Wang, S. Z. Li and W. Li, Deep Learning Empowers the Discovery of Self-Assembling Peptides with Over 10 Trillion Sequences, Adv. Sci., 2023, 10(31) DOI:10.1002/advs.202301544
.
- S. Sarkar, W. Gu and E. W. Schmidt, Expanding the Chemical Space of Synthetic Cyclic Peptides Using a Promiscuous Macrocyclase from Prenylagaramide Biosynthesis, ACS Catal., 2020, 10(13), 7146–7153, DOI:10.1021/acscatal.0c00623
.
- F. Lovering, Escape from Flatland 2: Complexity and Promiscuity, MedChemComm, 2013, 4(3), 515, 10.1039/c2md20347b
.
- T. T. Wager, X. Hou, P. R. Verhoest and A. Villalobos, Moving beyond Rules: The Development of a Central Nervous System Multiparameter Optimization (CNS MPO) Approach to Enable Alignment of Druglike Properties, ACS Chem. Neurosci., 2010, 1(6), 435–449, DOI:10.1021/cn100008c
.
- F. Lovering, J. Bikker and C. Humblet, Escape from Flatland: Increasing Saturation as an Approach to Improving Clinical Success, J. Med. Chem., 2009, 52(21), 6752–6756, DOI:10.1021/jm901241e
.
- T. Cheng, Y. Zhao, X. Li, F. Lin, Y. Xu, X. Zhang, Y. Li, R. Wang and L. Lai, Computation of Octanol-Water Partition Coefficients by Guiding an Additive Model with Knowledge, J. Chem. Inf. Model., 2007, 47(6), 2140–2148, DOI:10.1021/ci700257y
.
- A. K. Ghose, V. N. Viswanadhan and J. J. Wendoloski, Prediction of Hydrophobic (Lipophilic) Properties of Small Organic Molecules Using Fragmental Methods: An Analysis of ALOGP and CLOGP Methods, J. Phys. Chem. A, 1998, 102(21), 3762–3772, DOI:10.1021/jp980230o
.
- S. J. Thompson, C. K. Hattotuwagama, J. D. Holliday and D. R. Flower, On the Hydrophobicity
of Peptides: Comparing Empirical Predictions of Peptide Log P Values, Bioinformation, 2006, 1(7), 237–241, DOI:10.6026/97320630001237
.
- L. B. Kier, An Index of Molecular Flexibility from Kappa Shape Attributes, Quant. Struct. Relat., 1989, 8(3), 221–224, DOI:10.1002/qsar.19890080307
.
- M. Randić and M. Pompe, The Variable Molecular Descriptors Based on Distance Related Matrices, J. Chem. Inf. Comput. Sci., 2001, 41(3), 575–581, DOI:10.1021/ci0001029
.
- F. R. Burden, A Chemically Intuitive Molecular Index Based on the Eigenvalues of a Modified Adjacency Matrix, Quant. Struct. Relat., 1997, 16(4), 309–314, DOI:10.1002/qsar.19970160406
.
- I. Gutman, B. Furtula and V. Katanić, Randić Index and Information, AKCE Int. J. Graphs Comb., 2018, 15(3), 307–312, DOI:10.1016/j.akcej.2017.09.006
.
-
D. Sabirov, A. Zimina and I. Shepelevich, Complexity of Molecular Ensembles with Basak's Indices: Applying Structural Information Content, 2025, pp. 113–121 DOI:10.1007/978-3-031-67841-7_6
.
- P. M. Andersson, M. Sjöström, S. Wold and T. Lundstedt, Comparison between Physicochemical and Calculated Molecular Descriptors, J. Chemom., 2000, 14(5–6), 629–642, DOI:10.1002/1099-128X(200009/12)14:5/6
![[double bond splayed right]](https://www.rsc.org/images/entities/char_e00a.gif) 629::AID-CEM606
629::AID-CEM606![[double bond splayed left]](https://www.rsc.org/images/entities/char_e009.gif) 3.0.CO;2-M
3.0.CO;2-M .
- J. Singh, B. Shaik, V. K. Agrawal and P. V. Khadikar, SAR Studies on β-Cell KATP Channel Openers, Interdiscip. Sci.: Comput. Life Sci., 2012, 4(3), 215–222, DOI:10.1007/s12539-012-0135-8
.
- V. Consonni, R. Todeschini and M. Pavan, Structure/Response Correlations and Similarity/Diversity Analysis by GETAWAY Descriptors. 1. Theory of the Novel 3D Molecular Descriptors, J. Chem. Inf. Comput. Sci., 2002, 42(3), 682–692, DOI:10.1021/ci015504a
.
- O. Devinyak, D. Havrylyuk and R. Lesyk, 3D-MoRSE Descriptors Explained, J. Mol. Graphics Modell., 2014, 54, 194–203, DOI:10.1016/j.jmgm.2014.10.006
.
- R. Todeschini and P. Gramatica, The Whim Theory: New 3D Molecular Descriptors for Qsar in Environmental Modelling, SAR QSAR Environ. Res., 1997, 7(1–4), 89–115, DOI:10.1080/10629369708039126
.
- P. Gramatica, WHIM Descriptors of Shape, QSAR Comb. Sci., 2006, 25(4), 327–332, DOI:10.1002/qsar.200510159
.
- O. Méndez-Lucio and J. L. Medina-Franco, The Many Roles of Molecular Complexity in Drug Discovery, Drug Discovery Today, 2017, 22(1), 120–126, DOI:10.1016/j.drudis.2016.08.009
.
- M. Paradís-Bas, J. Tulla-Puche and F. Albericio, The Road to the Synthesis of “Difficult Peptides, Chem. Soc. Rev., 2016, 45(3), 631–654, 10.1039/C5CS00680E
.
- P. A. Clemons, N. E. Bodycombe, H. A. Carrinski, J. A. Wilson, A. F. Shamji, B. K. Wagner, A. N. Koehler and S. L. Schreiber, Small Molecules of Different Origins Have Distinct Distributions of Structural Complexity That Correlate with Protein-Binding Profiles, Proc. Natl. Acad. Sci. U. S. A., 2010, 107(44), 18787–18792, DOI:10.1073/pnas.1012741107
.
- D. Eisenberg, W. Wilcox and A. D. McLachlan, Hydrophobicity and Amphiphilicity in Protein Structure, J. Cell. Biochem., 1986, 31(1), 11–17, DOI:10.1002/jcb.240310103
.
- W. M. Meylan and P. H. Howard, Atom/Fragment Contribution Method for Estimating Octanol–Water Partition Coefficients, J. Pharm. Sci., 1995, 84(1), 83–92, DOI:10.1002/jps.2600840120
.
- M. Oeller, R. J. D. Kang, H. L. Bolt, A. L. Gomes dos Santos, A. L. Weinmann, A. Nikitidis, P. Zlatoidsky, W. Su, W. Czechtizky, L. De Maria, P. Sormanni and M. Vendruscolo, Sequence-Based Prediction of the Intrinsic Solubility of Peptides Containing Non-Natural Amino Acids, Nat. Commun., 2023, 14(1), 7475, DOI:10.1038/s41467-023-42940-w
.
- H. Xiong, B. L. Buckwalter, H. M. Shieh and M. H. Hecht, Periodicity of Polar and Nonpolar Amino Acids Is the Major Determinant of Secondary Structure in Self-Assembling Oligomeric Peptides, Proc. Natl. Acad. Sci. U. S. A., 1995, 92(14), 6349–6353, DOI:10.1073/pnas.92.14.6349
.
- A. E. Kister and I. Gelfand, Finding of Residues Crucial for Supersecondary Structure Formation, Proc. Natl. Acad. Sci. U. S. A., 2009, 106(45), 18996–19000, DOI:10.1073/pnas.0909714106
.
- A. Rajput, A. K. Gupta and M. Kumar, Prediction and Analysis of Quorum Sensing Peptides Based on Sequence Features, PLoS One, 2015, 10(3), e0120066, DOI:10.1371/journal.pone.0120066
.
- W. Chen, H. Ding, P. Feng, H. Lin and K. C. Chou, IACP: A Sequence-Based Tool for Identifying Anticancer Peptides, Oncotarget, 2016, 7(13), 16895–16909, DOI:10.18632/oncotarget.7815
.
- P. Bhadra, J. Yan, J. Li, S. Fong and S. W. I. Siu, AmPEP: Sequence-Based Prediction of Antimicrobial Peptides Using Distribution Patterns of Amino Acid Properties and Random Forest, Sci. Rep., 2018, 8(1), 1697, DOI:10.1038/s41598-018-19752-w
.
- K.-Y. Huang, Y.-J. Tseng, H.-J. Kao, C.-H. Chen, H.-H. Yang and S.-L. Weng, Identification of Subtypes of Anticancer Peptides Based on Sequential Features and Physicochemical Properties, Sci. Rep., 2021, 11(1), 13594, DOI:10.1038/s41598-021-93124-9
.
- W. R. Pearson and D. J. Lipman, Improved Tools for Biological Sequence Comparison, Proc. Natl. Acad. Sci. U. S. A., 1988, 85(8), 2444–2448, DOI:10.1073/pnas.85.8.2444
.
- T. Fox, M. Bieler, P. Haebel, R. Ochoa, S. Peters and A. Weber, BILN: A Human-Readable Line Notation for Complex Peptides, J. Chem. Inf. Model., 2022, 62(17), 3942–3947, DOI:10.1021/acs.jcim.2c00703
.
- T. Zhang, H. Li, H. Xi, R. V. Stanton and S. H. Rotstein, HELM: A Hierarchical Notation Language for Complex Biomolecule Structure Representation, J. Chem. Inf. Model., 2012, 52(10), 2796–2806, DOI:10.1021/ci3001925
.
- V. Erckes, M. Abderrahmane, M. Jusot, C. Steuer and R. Ochoa, Peptide Cheminformatics Tools: Making Computational Tasks Accessible in Peptide Drug Discovery, Drug Discovery Today, 2026, 31(2), 104612, DOI:10.1016/j.drudis.2026.104612
.
- Y. Su, T. Doherty, A. J. Waring, P. Ruchala and M. Hong, Roles of Arginine and Lysine Residues in the Translocation of a Cell-Penetrating Peptide From 13C, 31P, And 19F Solid-State NMR, Biochemistry, 2009, 48(21), 4587–4595, DOI:10.1021/bi900080d
.
- A. Gräslund, F. Madani, S. Lindberg, Ü. Langel and S. Futaki, Mechanisms of Cellular Uptake of Cell-Penetrating Peptides, J. Biophys., 2011, 2011, 1–10, DOI:10.1155/2011/414729
.
- T. Liu, Y. Liu, H. Y. Kao and D. Pei, Membrane Permeable Cyclic Peptidyl Inhibitors against Human Peptidylprolyl Isomer Ase Pin1, J. Med. Chem., 2010, 53(6), 2494–2501, DOI:10.1021/jm901778v
.
- J. J. Cronican, D. B. Thompson, K. T. Beier, B. R. McNaughton, C. L. Cepko and D. R. Liu, Potent Delivery of Functional Proteins into Mammalian Cells in Vitro and in Vivo Using a Supercharged Protein, ACS Chem. Biol., 2010, 5(8), 747–752, DOI:10.1021/cb1001153
.
- B. Manavalan, S. Subramaniyam, T. H. Shin, M. O. Kim and G. Lee, Machine-Learning-Based Prediction of Cell-Penetrating Peptides and Their Uptake Efficiency with Improved Accuracy, J. Proteome Res., 2018, 17(8), 2715–2726, DOI:10.1021/acs.jproteome.8b00148
.
- R. M. Bernardes-Loch, G. de Oliveira Almeida, I. T. Brasiliano, W. Meira Jr, D. E. V. Pires, M. C. Baracat-Pereira and S. de Azevedo Silveira, PerseuCPP: A Machine Learning Strategy to Predict Cell-Penetrating Peptides and Their Uptake Efficiency, Bioinf. Adv., 2024, 5(1) DOI:10.1093/bioadv/vbaf213
.
- P. Charoenkwan, P. Chumnanpuen, N. Schaduangrat and W. Shoombuatong, Stack-AVP: A Stacked Ensemble Predictor Based on Multi-View Information for Fast and Accurate Discovery of Antiviral Peptides, J. Mol. Biol., 2025, 437(6), 168853, DOI:10.1016/j.jmb.2024.168853
.
- R. Pugliese and F. Gelain, Peptidic Biomaterials: From Self-Assembling to Regenerative Medicine, Trends Biotechnol., 2017, 35(2), 145–158, DOI:10.1016/j.tibtech.2016.09.004
.
- H. Hosseinkhani, P.-D. Hong and D.-S. Yu, Self-Assembled Proteins and Peptides for Regenerative Medicine, Chem. Rev., 2013, 113(7), 4837–4861, DOI:10.1021/cr300131h
.
- B. He, X. Yuan, A. Zhou, H. Zhang and D. Jiang, Designer Functionalised Self-Assembling Peptide Nanofibre Scaffolds for Cartilage Tissue Engineering, Expert Rev. Mol. Med., 2014, 16, e12, DOI:10.1017/erm.2014.13
.
- W. Sun, D. A. Gregory and X. Zhao, Designed Peptide Amphiphiles as Scaffolds for Tissue Engineering, Adv. Colloid Interface Sci., 2023, 314, 102866, DOI:10.1016/j.cis.2023.102866
.
- N. Ni, Y. Hu, H. Ren, C. Luo, P. Li, J.-B. Wan and H. Su, Self-Assembling Peptide Nanofiber Scaffolds Enhance Dopaminergic Differentiation of Mouse Pluripotent Stem Cells in 3-Dimensional Culture, PLoS One, 2013, 8(12), e84504, DOI:10.1371/journal.pone.0084504
.
- E. Gazit, A Possible Role for Π-stacking in the Self-assembly of Amyloid Fibrils, FASEB J., 2002, 16(1), 77–83, DOI:10.1096/fj.01-0442hyp
.
- N. P. King, J. B. Bale, W. Sheffler, D. E. McNamara, S. Gonen, T. Gonen, T. O. Yeates and D. Baker, Accurate Design of Co-Assembling Multi-Component Protein Nanomaterials, Nature, 2014, 510(7503), 103–108, DOI:10.1038/nature13404
.
- F. Fontana and F. Gelain, Probing Mechanical Properties and Failure Mechanisms of Fibrils of Self-Assembling Peptides, Nanoscale Adv., 2020, 2(1), 190–198, 10.1039/C9NA00621D
.
- P. Chakraborty, Y. Tang, T. Yamamoto, Y. Yao, T. Guterman, S. Zilberzwige-Tal, N. Adadi, W. Ji, T. Dvir, A. Ramamoorthy, G. Wei and E. Gazit, Unusual Two-Step Assembly of a Minimalistic Dipeptide-Based Functional Hypergelator, Adv. Mater., 2020, 32(9) DOI:10.1002/adma.201906043
.
- N. R. Lee, C. J. Bowerman and B. L. Nilsson, Effects of Varied Sequence Pattern on the Self-Assembly of Amphipathic Peptides, Biomacromolecules, 2013, 14(9), 3267–3277, DOI:10.1021/bm400876s
.
- S. Zhang, Lipid-like Self-Assembling Peptides, Acc. Chem. Res., 2012, 45(12), 2142–2150, DOI:10.1021/ar300034v
.
- S. C. Yuan, J. A. Lewis, H. Sai, S. J. Weigand, L. C. Palmer and S. I. Stupp, Peptide Sequence Determines Structural Sensitivity to Supramolecular Polymerization Pathways and Bioactivity, J. Am. Chem. Soc., 2022, 144(36), 16512–16523, DOI:10.1021/jacs.2c05759
.
- P. Charoenkwan, P. Chumnanpuen, N. Schaduangrat, C. Oh, B. Manavalan and W. Shoombuatong, PSRQSP: An Effective Approach for the Interpretable Prediction of Quorum Sensing Peptide Using Propensity Score Representation Learning, Comput. Biol. Med., 2023, 158, 106784, DOI:10.1016/j.compbiomed.2023.106784
.
- P. Pandey, V. Patel, N. V. George and S. S. Mallajosyula, KELM-CPPpred: Kernel Extreme Learning Machine Based Prediction Model for Cell-Penetrating Peptides, J. Proteome Res., 2018, 17(9), 3214–3222, DOI:10.1021/acs.jproteome.8b00322
.
- L. Yao, P. Xie, J. Guan, C.-R. Chung, W. Zhang, J. Deng, Y. Huang, Y.-C. Chiang and T.-Y. Lee, ACP-CapsPred: An Explainable Computational Framework for Identification and Functional Prediction of Anticancer Peptides Based on Capsule Network, Briefings Bioinf., 2024, 25(5) DOI:10.1093/bib/bbae460
.
- Y. Zuo, Y. Lv, Z. Wei, L. Yang, G. Li and G. Fan, IDPF-PseRAAAC: A Web-Server for Identifying the Defensin Peptide Family and Subfamily Using Pseudo Reduced Amino Acid Alphabet Composition, PLoS One, 2015, 10(12), e0145541, DOI:10.1371/journal.pone.0145541
.
- P. J. A. Cock, T. Antao, J. T. Chang, B. A. Chapman, C. J. Cox, A. Dalke, I. Friedberg, T. Hamelryck, F. Kauff, B. Wilczynski and M. J. L. De Hoon, Biopython: Freely Available Python Tools for Computational Molecular Biology and Bioinformatics, Bioinformatics, 2009, 25(11), 1422–1423, DOI:10.1093/bioinformatics/btp163
.
- J. Dong, Z. J. Yao, L. Zhang, F. Luo, Q. Lin, A. P. Lu, A. F. Chen and D. S. Cao, PyBioMed: A Python Library for Various Molecular Representations of Chemicals, Proteins and DNAs and Their Interactions, J. Cheminf., 2018, 10(1), 16, DOI:10.1186/s13321-018-0270-2
.
- R. Ochoa and P. Cossio, PepFun: Open Source Protocols for Peptide-Related Computational Analysis, Molecules, 2021, 26(6), 1664, DOI:10.3390/molecules26061664
.
- D. Osorio, P. Rondón-Villarreal and R. Torres, Peptides: A Package for Data Mining of Antimicrobial Peptides, R J., 2015, 7(1), 4, DOI:10.32614/RJ-2015-001
.
- R. Ochoa and K. Deibler, PepFuNN: Novo Nordisk Open-Source Toolkit to Enable Peptide in Silico Analysis, J. Pept. Sci., 2025, 31(2) DOI:10.1002/psc.3666
.
- S. S. Sahu and G. Panda, A Novel Feature Representation Method Based on Chou's Pseudo Amino Acid Composition for Protein Structural Class Prediction, Comput. Biol. Chem., 2010, 34(5–6), 320–327, DOI:10.1016/j.compbiolchem.2010.09.002
.
- K.-J. Park and M. Kanehisa, Prediction of Protein Subcellular Locations by Support Vector Machines Using Compositions of Amino Acids and Amino Acid Pairs, Bioinformatics, 2003, 19(13), 1656–1663, DOI:10.1093/bioinformatics/btg222
.
- P. Agrawal, D. Bhagat, M. Mahalwal, N. Sharma and G. P. S. Raghava, AntiCP 2.0: An Updated Model for Predicting Anticancer Peptides, Briefings Bioinf., 2021, 22(3) DOI:10.1093/bib/bbaa153
.
-
L. Li, Dimension Reduction for High-Dimensional Data, 2010, pp. 417–434 DOI:10.1007/978-1-60761-580-4_14
.
- B. Rao, C. Zhou, G. Zhang, R. Su and L. Wei, ACPred-Fuse: Fusing Multi-View Information Improves the Prediction of Anticancer Peptides, Briefings Bioinf., 2020, 21(5), 1846–1855, DOI:10.1093/bib/bbz088
.
- Y. Zuo, Y. Li, Y. Chen, G. Li, Z. Yan and L. Yang, PseKRAAC: A Flexible Web Server for Generating Pseudo K-Tuple Reduced Amino Acids Composition, Bioinformatics, 2017, 33(1), 122–124, DOI:10.1093/bioinformatics/btw564
.
- K. Chen, L. Kurgan and M. Rahbari, Prediction of Protein Crystallization Using Collocation of Amino Acid Pairs, Biochem. Biophys. Res. Commun., 2007, 355(3), 764–769, DOI:10.1016/j.bbrc.2007.02.040
.
- R. Dai, W. Zhang, W. Tang, E. Wynendaele, Q. Zhu, Y. Bin, B. De Spiegeleer and J. Xia, BBPpred: Sequence-Based Prediction of Blood-Brain Barrier Peptides with Feature Representation Learning and Logistic Regression, J. Chem. Inf. Model., 2021, 61(1), 525–534, DOI:10.1021/acs.jcim.0c01115
.
- L. Wei, C. Zhou, R. Su and Q. Zou, PEPred-Suite: Improved and Robust Prediction of Therapeutic Peptides Using Adaptive Feature Representation Learning, Bioinformatics, 2019, 35(21), 4272–4280, DOI:10.1093/bioinformatics/btz246
.
- S. Kawashima, P. Pokarowski, M. Pokarowska, A. Kolinski, T. Katayama and M. Kanehisa, AAindex: Amino Acid Index Database, Progress Report 2008, Nucleic Acids Res., 2007, 36(Database), D202–D205, DOI:10.1093/nar/gkm998
.
- Z. Dosztányi, V. Csizmók, P. Tompa and I. Simon, The Pairwise Energy Content Estimated from Amino Acid Composition Discriminates between Folded and Intrinsically Unstructured Proteins, J. Mol. Biol., 2005, 347(4), 827–839, DOI:10.1016/j.jmb.2005.01.071
.
- M. Sandberg, L. Eriksson, J. Jonsson, M. Sjöström and S. Wold, New Chemical Descriptors Relevant for the Design of Biologically Active Peptides. A Multivariate Characterization of 87 Amino Acids, J. Med. Chem., 1998, 41(14), 2481–2491, DOI:10.1021/jm9700575
.
- S. Henikoff and J. G. Henikoff, Amino Acid Substitution Matrices from Protein Blocks, Proc. Natl. Acad. Sci. U. S. A., 1992, 89(22), 10915–10919, DOI:10.1073/pnas.89.22.10915
.
- E. Wynendaele, N. Debunne, Y. Janssens, A. De Spiegeleer, F. Verbeke, L. Tack, S. Van Welden, E. Goossens, D. Knappe, R. Hoffmann, C. Van De Wiele, D. Laukens, P. Van Eenoo, L. Vereecke, F. Van Immerseel, O. De Wever and B. De Spiegeleer, The Quorum Sensing Peptide EntF* Promotes Colorectal Cancer Metastasis in Mice: A New Factor in the Host-Microbiome Interaction, BMC Biol., 2022, 20(1), 151, DOI:10.1186/s12915-022-01317-z
.
- J. Xu, F. Li, C. Li, X. Guo, C. Landersdorfer, H.-H. Shen, A. Y. Peleg, J. Li, S. Imoto, J. Yao, T. Akutsu and J. Song, IAMPCN: A Deep-Learning Approach for Identifying Antimicrobial Peptides and Their Functional Activities, Briefings Bioinf., 2023, 24(4) DOI:10.1093/bib/bbad240
.
- J. Guan, L. Yao, C.-R. Chung, Y.-C. Chiang and T.-Y. Lee, StackTHPred: Identifying Tumor-Homing Peptides through GBDT-Based Feature Selection with Stacking Ensemble Architecture, Int. J. Mol. Sci., 2023, 24(12), 10348, DOI:10.3390/ijms241210348
.
- C. Chen, L.-X. Chen, X.-Y. Zou and P.-X. Cai, Predicting Protein Structural Class Based on Multi-Features Fusion, J. Theor. Biol., 2008, 253(2), 388–392, DOI:10.1016/j.jtbi.2008.03.009
.
- Q. Dai, Y. Li, X. Liu, Y. Yao, Y. Cao and P. He, Comparison Study on Statistical Features of Predicted Secondary Structures for Protein Structural Class Prediction: From Content to Position, BMC Bioinf., 2013, 14(1), 152, DOI:10.1186/1471-2105-14-152
.
- S. Zhang, S. Ding and T. Wang, High-Accuracy Prediction of Protein Structural Class for Low-Similarity Sequences Based on Predicted Secondary Structure, Biochimie, 2011, 93(4), 710–714, DOI:10.1016/j.biochi.2011.01.001
.
- R. R. Sokal, N. L. Oden and B. A. Thomson, Local Spatial Autocorrelation in Biological Variables, Biol. J. Linn. Soc., 1998, 65(1), 41–62, DOI:10.1111/j.1095-8312.1998.tb00350.x
.
- N. Xiao, D.-S. Cao, M.-F. Zhu and Q.-S. Xu, Protr/ProtrWeb: R Package and Web Server for Generating Various Numerical Representation Schemes of Protein Sequences, Bioinformatics, 2015, 31(11), 1857–1859, DOI:10.1093/bioinformatics/btv042
.
- E. Bizzotto, G. Zampieri, L. Treu, P. Filannino, R. Di Cagno and S. Campanaro, Classification of Bioactive Peptides: A Systematic Benchmark of Models and Encodings, Comput. Struct. Biotechnol. J., 2024, 23, 2442–2452, DOI:10.1016/j.csbj.2024.05.040
.
- K. C. Chou, Prediction of Protein Cellular Attributes Using Pseudo-Amino Acid Composition, Proteins:Struct., Funct., Bioinf., 2001, 43(3), 246–255, DOI:10.1002/prot.1035
.
- K.-C. Chou, Prediction of Protein Subcellular Locations by Incorporating Quasi-Sequence-Order Effect, Biochem. Biophys. Res. Commun., 2000, 278(2), 477–483, DOI:10.1006/bbrc.2000.3815
.
- J. Kyte and R. F. Doolittle, A Simple Method for Displaying the Hydropathic Character of a Protein, J. Mol. Biol., 1982, 157(1), 105–132, DOI:10.1016/0022-2836(82)90515-0
.
- M. Vihinen, E. Torkkila and P. Riikonen, Accuracy of Protein Flexibility Predictions, Proteins: Struct., Funct., Bioinf., 1994, 19(2), 141–149, DOI:10.1002/prot.340190207
.
- J. G. Henikoff and S. Henikoff, Using Substitution Probabilities to Improve Position-Specific Scoring Matrices, Bioinformatics, 1996, 12(2), 135–143, DOI:10.1093/bioinformatics/12.2.135
.
- R. Grantham, Amino Acid Difference Formula to Help Explain Protein Evolution, Science, 1974, 185(4154), 862–864, DOI:10.1126/science.185.4154.862
.
- J. L. Medina-Franco, N. Sánchez-Cruz, E. López-López and B. I. Díaz-Eufracio, Progress on Open Chemoinformatic Tools for Expanding and Exploring the Chemical Space, J. Comput.-Aided. Mol. Des., 2022, 36(5), 341–354, DOI:10.1007/s10822-021-00399-1
.
- E. Wynendaele, B. Gevaert, S. Stalmans, F. Verbeke and B. De Spiegeleer, Exploring the Chemical Space of Quorum Sensing Peptides, Pept. Sci., 2015, 104(5), 544–551, DOI:10.1002/bip.22649
.
- M. Orsi, H. Personne, E. Bonvin, T. Paschoud, B. Olcay, X. Hu, S. Javor and J.-L. Reymond, Chemical Space for Peptide-Based Antimicrobials, Chimia, 2024, 78(10), 648–653, DOI:10.2533/chimia.2024.648
.
- E. Wynendaele, A. Bronselaer, J. Nielandt, M. D’Hondt, S. Stalmans, N. Bracke, F. Verbeke, C. Van De Wiele, G. De Tré and B. De Spiegeleer, Quorumpeps Database: Chemical Space, Microbial Origin and Functionality of Quorum Sensing Peptides, Nucleic Acids Res., 2013, 41(D1), D655–D659, DOI:10.1093/nar/gks1137
.
- A. Capecchi, A. Zhang and J.-L. Reymond, Populating Chemical Space with Peptides Using a Genetic Algorithm, J. Chem. Inf. Model., 2020, 60(1), 121–132, DOI:10.1021/acs.jcim.9b01014
.
- N. Sánchez-Cruz and J. L. Medina-Franco, Statistical-Based Database Fingerprint: Chemical Space Dependent Representation of Compound Databases, J. Cheminf., 2018, 10(1), 55, DOI:10.1186/s13321-018-0311-x
.
- P. Willett, Similarity-Based Virtual Screening Using 2D Fingerprints, Drug Discovery Today, 2006, 1046–1053, DOI:10.1016/j.drudis.2006.10.005
.
- D. Boldini, D. Ballabio, V. Consonni, R. Todeschini, F. Grisoni and S. A. Sieber, Effectiveness of Molecular Fingerprints for Exploring the Chemical Space of Natural Products, J. Cheminf., 2024, 16(1), 35, DOI:10.1186/s13321-024-00830-3
.
- J. L. Durant, B. A. Leland, D. R. Henry and J. G. Nourse, Reoptimization of MDL Keys for Use in Drug Discovery, J. Chem. Inf. Comput. Sci., 2002, 42(6), 1273–1280, DOI:10.1021/ci010132r
.
- R. E. Carhart, D. H. Smith and R. Venkataraghavan, Atom Pairs as Molecular Features in Structure–Activity Studies: Definition and Applications, J. Chem. Inf. Comput. Sci., 1985, 25(2), 64–73, DOI:10.1021/ci00046a002
.
- S. D. Axen, X.-P. Huang, E. L. Cáceres, L. Gendelev, B. L. Roth and M. J. Keiser, A Simple Representation of Three-Dimensional Molecular Structure, J. Med. Chem., 2017, 60(17), 7393–7409, DOI:10.1021/acs.jmedchem.7b00696
.
- D. Rogers and M. Hahn, Extended-Connectivity Fingerprints, J. Chem. Inf. Model., 2010, 50(5), 742–754, DOI:10.1021/ci100050t
.
- S. Riniker and G. A. Landrum, Open-Source Platform to Benchmark Fingerprints for Ligand-Based Virtual Screening, J. Cheminf., 2013, 5(1), 26, DOI:10.1186/1758-2946-5-26
.
- I. Di Bonaventura, X. Jin, R. Visini, D. Probst, S. Javor, B.-H. Gan, G. Michaud, A. Natalello, S. M. Doglia, T. Köhler, C. van Delden, A. Stocker, T. Darbre and J.-L. Reymond, Chemical Space Guided Discovery of Antimicrobial Bridged Bicyclic Peptides against Pseudomonas Aeruginosa and Its Biofilms, Chem. Sci., 2017, 8(10), 6784–6798, 10.1039/C7SC01314K
.
- M. Orsi and J.-L. Reymond, One Chiral Fingerprint to Find Them All, J. Cheminf., 2024, 16(1), 53, DOI:10.1186/s13321-024-00849-6
.
- J. Adamczyk and P. Ludynia, Scikit-Fingerprints: Easy and Efficient Computation of Molecular Fingerprints in Python, SoftwareX, 2024, 28, 101944, DOI:10.1016/j.softx.2024.101944
.
- M. Banck, T. Vandermeersch, N. M. O’Boyle, G. R. Hutchison, C. Morley and C. A. James, Open Babel: An Open Chemical Toolbox, J. Cheminf., 2011, 3(1), 33, DOI:10.1186/1758-2946-3-33
.
- C. Manelfi, V. Tazzari, F. Lunghini, C. Cerchia, A. Fava, A. Pedretti, P. F. W. Stouten, G. Vistoli and A. R. Beccari, “DompeKeys”: A Set of Novel Substructure-Based Descriptors for Efficient Chemical Space Mapping, Development and Structural Interpretation of Machine Learning Models, and Indexing of Large Databases, J. Cheminf., 2024, 16(1), 21, DOI:10.1186/s13321-024-00813-4
.
- A. Bender, H. Y. Mussa, R. C. Glen and S. Reiling, Molecular Similarity Searching Using Atom Environments, Information-Based Feature Selection, and a Naïve Bayesian Classifier, J. Chem. Inf. Comput. Sci., 2004, 44(1), 170–178, DOI:10.1021/ci034207y
.
- A. Rives, J. Meier, T. Sercu, S. Goyal, Z. Lin, J. Liu, D. Guo, M. Ott, C. L. Zitnick, J. Ma and R. Fergus, Biological Structure and Function Emerge from Scaling Unsupervised Learning to 250 Million Protein Sequences, Proc. Natl. Acad. Sci. U. S. A., 2021, 118(15) DOI:10.1073/pnas.2016239118
.
- S. Jaeger, S. Fulle and S. Turk, Mol2vec: Unsupervised Machine Learning Approach with Chemical Intuition, J. Chem. Inf. Model., 2018, 58(1), 27–35, DOI:10.1021/acs.jcim.7b00616
.
- A. Elnaggar, M. Heinzinger, C. Dallago, G. Rehawi, Y. Wang, L. Jones, T. Gibbs, T. Feher, C. Angerer, M. Steinegger, D. Bhowmik and B. Rost, ProtTrans: Toward Understanding the Language of Life Through Self-Supervised Learning, IEEE Trans. Pattern Anal. Mach. Intell., 2022, 44(10), 7112–7127, DOI:10.1109/TPAMI.2021.3095381
.
- K. Yang, K. Swanson, W. Jin, C. Coley, P. Eiden, H. Gao, A. Guzman-Perez, T. Hopper, B. Kelley, M. Mathea, A. Palmer, V. Settels, T. Jaakkola, K. Jensen and R. Barzilay, Analyzing Learned Molecular Representations for Property Prediction, J. Chem. Inf. Model., 2019, 59(8), 3370–3388, DOI:10.1021/acs.jcim.9b00237
.
- R. Gómez-Bombarelli, J. N. Wei, D. Duvenaud, J. M. Hernández-Lobato, B. Sánchez-Lengeling, D. Sheberla, J. Aguilera-Iparraguirre, T. D. Hirzel, R. P. Adams and A. Aspuru-Guzik, Automatic Chemical Design Using a Data-Driven Continuous Representation of Molecules, ACS Cent. Sci., 2018, 4(2), 268–276, DOI:10.1021/acscentsci.7b00572
.
- R. Özçelik, H. Brinkmann, E. Criscuolo and F. Grisoni, Generative Deep Learning for de Novo Drug Design—A Chemical Space Odyssey, J. Chem. Inf. Model., 2025, 65(14), 7352–7372, DOI:10.1021/acs.jcim.5c00641
.
-
G. E. Hinton and S. T. Roweis, Stochastic Neighbor Embedding, Neural Information Processing Systems, 2002 Search PubMed
.
- C. Marquet, M. Heinzinger, T. Olenyi, C. Dallago, K. Erckert, M. Bernhofer, D. Nechaev and B. Rost, Embeddings from Protein Language Models Predict Conservation and Variant Effects, Hum. Genet., 2022, 141(10), 1629–1647, DOI:10.1007/s00439-021-02411-y
.
- S. Renaud and R. A. Mansbach, Latent Spaces for Antimicrobial Peptide Design, Digital Discovery, 2023, 2(2), 441–458, 10.1039/D2DD00091A
.
- S. Gelman, B. Johnson, C. R. Freschlin, A. Sharma, S. D’Costa, J. Peters, A. Gitter and P. A. Romero, Biophysics-Based Protein Language Models for Protein Engineering, Nat. Methods, 2025, 22(9), 1868–1879, DOI:10.1038/s41592-025-02776-2
.
- J. H. Jensen, T. Hoeg-Jensen and S. B. Padkjær, Building a BioChemformatics Database, J. Chem. Inf. Model., 2008, 48(12), 2404–2413, DOI:10.1021/ci800128b
.
- F. Mastrolorito, N. Gambacorta, F. Ciriaco, F. Cutropia, M. V. Togo, V. Belgiovine, A. R. Tondo, D. Trisciuzzi, A. Monaco, R. Bellotti, C. D. Altomare, O. Nicolotti and N. Amoroso, Chemical Space Networks Enhance Toxicity Recognition via Graph Embedding, J. Chem. Inf. Model., 2025, 65(4), 1850–1861, DOI:10.1021/acs.jcim.4c02140
.
- P. Schwaller, T. Laino, T. Gaudin, P. Bolgar, C. A. Hunter, C. Bekas and A. A. Lee, Molecular Transformer: A Model for Uncertainty-Calibrated Chemical Reaction Prediction, ACS Cent. Sci., 2019, 5(9), 1572–1583, DOI:10.1021/acscentsci.9b00576
.
- D. Harding-Larsen, J. Funk, N. G. Madsen, H. Gharabli, C. G. Acevedo-Rocha, S. Mazurenko and D. H. Welner, Protein Representations: Encoding Biological Information for Machine Learning in Biocatalysis, Biotechnol. Adv., 2024, 77, 108459, DOI:10.1016/j.biotechadv.2024.108459
.
- N. E. Siedhoff, A.-M. Illig, U. Schwaneberg and M. D. Davari, PyPEF—An Integrated Framework for Data-Driven Protein Engineering, J. Chem. Inf. Model., 2021, 61(7), 3463–3476, DOI:10.1021/acs.jcim.1c00099
.
- X. Zheng and Y. Tomiura, A BERT-Based Pretraining Model for Extracting Molecular Structural Information from a SMILES Sequence, J. Cheminf., 2024, 16(1), 71, DOI:10.1186/s13321-024-00848-7
.
- T. Ochiai, T. Inukai, M. Akiyama, K. Furui, M. Ohue, N. Matsumori, S. Inuki, M. Uesugi, T. Sunazuka, K. Kikuchi, H. Kakeya and Y. Sakakibara, Variational Autoencoder-Based Chemical Latent Space for Large Molecular Structures with 3D Complexity, Commun. Chem., 2023, 6(1), 249, DOI:10.1038/s42004-023-01054-6
.
- J. van Eck, D. Gogishvili, W. Silva and S. Abeln, PLM-EXplain: Divide and Conquer the Protein Embedding Space, Bioinformatics, 2026, 42(1) DOI:10.1093/bioinformatics/btaf631
.
- M. Leclercq and A. Droit, Protein Language Models: Applications and Perspectives, J. Proteome Res., 2026, 25(2), 507–524, DOI:10.1021/acs.jproteome.5c00506
.
- N. Brandes, D. Ofer, Y. Peleg, N. Rappoport and M. Linial, ProteinBERT: A Universal Deep-Learning Model of Protein Sequence and Function, Bioinformatics, 2022, 38(8), 2102–2110, DOI:10.1093/bioinformatics/btac020
.
- L. Pantolini, G. Studer, J. Pereira, J. Durairaj, G. Tauriello and T. Schwede, Embedding-Based Alignment: Combining Protein Language Models with Dynamic Programming Alignment to Detect Structural Similarities in the Twilight-Zone, Bioinformatics, 2024, 40(1) DOI:10.1093/bioinformatics/btad786
.
- B. E. Suzek, H. Huang, P. McGarvey, R. Mazumder and C. H. Wu, UniRef: Comprehensive and Non-Redundant UniProt Reference Clusters, Bioinformatics, 2007, 23(10), 1282–1288, DOI:10.1093/bioinformatics/btm098
.
- A. L. Feller and C. O. Wilke, Peptide-Aware Chemical Language Model Successfully Predicts Membrane Diffusion of Cyclic Peptides, J. Chem. Inf. Model., 2025, 65(2), 571–579, DOI:10.1021/acs.jcim.4c01441
.
- Z. Du, X. Ding, Y. Xu and Y. Li, UniDL4BioPep: A Universal Deep Learning Architecture for Binary Classification in Peptide Bioactivity, Briefings Bioinf., 2023, 24(3) DOI:10.1093/bib/bbad135
.
- W. Dee, LMPred: Predicting Antimicrobial Peptides Using Pre-Trained Language Models and Deep Learning, Bioinf. Adv., 2022, 2(1) DOI:10.1093/bioadv/vbac021
.
- S. Badrinarayanan, C. Guntuboina, P. Mollaei and A. Barati Farimani, Multi-Peptide: Multimodality Leveraged Language-Graph Learning of Peptide Properties, J. Chem. Inf. Model., 2025, 65(1), 83–91, DOI:10.1021/acs.jcim.4c01443
.
- B. De Spiegeleer, F. Verbeke, M. D’Hondt, A. Hendrix, C. Van De Wiele, C. Burvenich, K. Peremans, O. De Wever, M. Bracke and E. Wynendaele, The Quorum Sensing Peptides PhrG, CSP and EDF Promote Angiogenesis and Invasion of Breast Cancer Cells In Vitro, PLoS One, 2015, 10(3), e0119471, DOI:10.1371/journal.pone.0119471
.
- A. De Spiegeleer, A. Descamps, S. Govindarajan, J. Coudenys, K. Van der borght, H. Hirmz, N. Van Den Noortgate, D. Elewaut, B. De Spiegeleer and E. Wynendaele, Bacterial Quorum-Sensing Peptides as Immune Modulators Present in Systemic Circulation, Biomolecules, 2023, 13(2), 296, DOI:10.3390/biom13020296
.
- A. De Spiegeleer, A. Descamps, E. Wynendaele, P. Naumovski, L. Crombez, M. Planas, L. Feliu, D. Knappe, V. Mouly, A. Bigot, R. Bielza, R. Hoffmann, N. Van Den Noortgate, D. Elewaut and B. De Spiegeleer, Streptococcal Quorum Sensing Peptide CSP-7 Contributes to Muscle Inflammation and Wasting, Biochim. Biophys. Acta, Mol. Basis Dis., 2024, 1870(4), 167094, DOI:10.1016/j.bbadis.2024.167094
.
- M. Cavaco, P. Fraga, J. Valle, R. D. M. Silva, L. Gano, J. D. G. Correia, D. Andreu, M. A. R. B. Castanho and V. Neves, Molecular Determinants for Brain Targeting by Peptides: A Meta-Analysis Approach with Experimental Validation, Fluids Barriers CNS, 2024, 21(1), 45, DOI:10.1186/s12987-024-00545-5
.
- M. Dichiara, B. Amata, R. Turnaturi, A. Marrazzo and E. Amata, Tuning Properties for Blood-Brain Barrier Permeation: A Statistics-Based Analysis, ACS Chem. Neurosci., 2020, 11(1), 34–44, DOI:10.1021/acschemneuro.9b00541
.
- L. T. Nguyen, E. F. Haney and H. J. Vogel, The Expanding Scope of Antimicrobial Peptide Structures and Their Modes of Action, Trends Biotechnol., 2011, 29(9), 464–472, DOI:10.1016/j.tibtech.2011.05.001
.
| This journal is © the Owner Societies 2026 |





