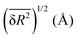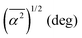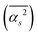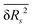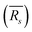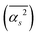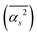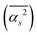Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Perfluorinated alkyl groups induce unexpected hydrophobic hydration structure
Simon
Schulke†
 a,
Elio
Casalini†
a,
Elio
Casalini†
 b,
Jaspreet
Kaur
b,
Jaspreet
Kaur
 a,
Sulejman
Skoko
a,
Sulejman
Skoko
 b,
Gerhard
Schwaab
b,
Gerhard
Schwaab
 a,
Martina
Havenith
a,
Martina
Havenith
 *a and
Ana
Vila Verde
*a and
Ana
Vila Verde
 *b
*b
aFaculty for Chemistry and Biochemistry, Ruhr-University Bochum, Universitätsstraße 150, 44801 Bochum, Germany. E-mail: Martina.Havenith@ruhr-uni-bochum.de
bFaculty of Physics, University of Duisburg-Essen, Lotharstr. 1, 47057 Duisburg, Germany. E-mail: ana.araujo-vila-verde@uni-due.de
First published on 19th March 2026
Abstract
Perfluorination of the terminal methyl group in ethanol gives rise to different thermodynamics of mixing with water. To understand its origin, we probe structure, thermodynamics and collective vibration modes in aqueous solutions of ethanol (EtOH) or 2,2,2-trifluoroethanol (TFE) by using Terahertz (THz) spectroscopy and molecular dynamics simulations. The THz spectra show mainly two features: a mostly entropy-related peak (below 200 cm−1) related to weaker water–water hydrogen bonds, and an enthalpy-related large band above 200 cm−1 related to water–solute hydrogen bonds. The entropic feature is red-shifted for TFE relative to EtOH, consistent with TFE's subpopulation of weaker solvation shell water–water hydrogen bonds found in the simulations. By contrast, the thermodynamics of mixing is dominated by three effects: the higher probability of forming water–water hydrogen bonds in the solvation shell of either solute than in the bulk; the fact that TFE induces a smaller perturbation per water molecule than EtOH, despite perturbing a slightly larger number of water molecules than EtOH, and TFE's weaker solute–water hydrogen bond. The three effects determine the more negative (favourable) enthalpy of mixing and more negative (unfavourable) entropy of mixing of EtOH relative to TFE at low concentrations. The results confirm that hydrophobic solvation of perfluorinated groups is fundamentally different from that of their alkylated equivalents and have implications for the development of models to predict solubility of perfluorinated molecules.
1 Introduction
Organofluorine is absent from almost every natural biosystem, despite the fact that fluorine is the 13th most abundant element on the earth's crust: only approximately 30 natural molecules with C–F bonds are known.1 Fluorine is therefore not considered an essential element for life: it is not required for any known vital cellular function, and in fact, its presence in biological systems can be toxic at elevated levels.2–7 Nevertheless, about 20% of commercial drugs present fluorinated groups,8,9 a prevalence that reflects the ability of fluorine substitutions to dramatically change reactivity and physical–chemical properties of molecules and materials.10 Perfluorination – whereby all alkyl hydrogen atoms of one or more carbons in a molecule are replaced by fluorine – is a widely employed chemical strategy in materials and pharmaceutical sciences.11–14 It is known to enhance the thermal stability and chemical inertness of molecules,15 and increase the hydrophobicity of compounds – as reflected by more positive hydration free energies.16–20 An increased magnitude of the hydrophobic effect leads to a higher tendency to aggregate in aqueous solvents to reduce the solvent-exposed surface area.21 The stronger hydrophobic effect associated with perfluoroalkyl substituents relative to their alkyl counterparts is thought to be at least partially responsible for the higher stability of folded protein conformations22–24 and protein–ligand affinities25,26 in proteins and/or ligands with perfluorinated hydrophobic amino acids. We note, however, that there are examples where this correlation has not been observed, such as molecular force balances in aqueous binary solvents.27 Perfluorination, in contrast to other chemical modifications, such as alkylation and sylilation, preserves the overall molecular topology of the solutes. The similar topology is advantageous e.g. when using perfluorinated hydrophobic amino acids to control the stability of folded protein-based drugs.22–24,28,29That perfluorination leads to more positive hydration free energies has been known for decades.16 Nevertheless, how this increase relates to changes in hydration shell structure and dynamics continues to be actively debated. For alkanes with fewer than four carbon atoms, the hydration free energies are positive and very similar between them. For their perfluorinated counterparts, hydration free energies are consistently less favourable (i.e., more positive) than expected, but clearly increase with the number of carbon atoms—suggesting weaker solute–solvent interactions and an enthalpic cost relative to the alkanes.30 In contrast, early simulation studies using classical force fields concluded that small perfluoroalkanes have more favourable dispersion and electrostatic interactions with water than equivalent alkanes.31 More accurate calculations, using wave function-based symmetry-adapted perturbation theory at the DF-SAPT2+(CCD)δMP2 level of theory and employing the augmented, correlation-consistent, triple-ζ Dunning basis sets (aug-cc-pVTZ) concluded that water molecules indeed interact more favourably with perfluoroalkyl than alkyl groups, when one of the hydrogen atoms points towards the group.32 For larger alkanes, with more than four carbon atoms, the more positive hydration free energies observed in perfluorinated systems have been primarily attributed to their increased molecular volume,31 indicating a predominantly entropic penalty.
Clarifying differences in hydration between alkyl and perfluoroalkyl groups is advantageously done by using small, polar molecules in water as model systems, because their high solubility (in contrast to that of alkanes and perfluoroalkanes) enables a high signal-to-noise ratio. Moreover, studying perfluorinated groups is essential for understanding and predicting the local effects of perfluorination in larger and more complex systems such as drug molecules. Previous works20,32 have investigated perfluorination of C2 to C9 diols and of the terminal methyl group of ethanol (EtOH). Raman MCR spectroscopy and molecular dynamics (MD) simulations were conducted, focusing on the OH stretching region, which probes the local HB network indirectly. These analyses report a less extensive perturbation of the hydrogen bond network by the perfluorinated molecule, in apparent contradiction with hydration free energies, which suggest that perfluorinated molecules perturb their solvation shell more extensively,19,20,32 and also in apparent contradiction with the above-mentioned early simulation studies.31
Direct insight into whether the hydrogen-bond network of water in the Terahertz (THz) region differs between the solvation shell of perfluoroalkyl vs. alkyl substituents is of particular interest because it directly connects to the thermodynamics of hydration.33–35 Majumdar et al.'s initial study,36 using THz-Attenuated Total Reflectance (ATR) spectroscopy to probe the local hydration shell structure and dynamics, suggested that the solvation behaviour of 2,2,2-trifluoroethanol (TFE) is more entropically driven and that the solvation of EtOH is more enthalpically driven. These results are qualitatively in line with excess molar free energies of mixing.37–39 They accompany the spectroscopic analysis with MD simulations, focusing on different populations of hydrogen bonds for the solvation of TFE and EtOH, and conclude that EtOH perturbs the hydrogen bond network of surrounding waters less than TFE due to a greater number of solvation shell defects and weaker heteromolecular interactions in case of TFE solvation. This interpretation is in direct contrast to the above mentioned Raman and MD studies, which indicate that the solvation shell of TFE is less perturbed than that of EtOH.19,20,32
The above summary highlights the challenges surrounding understanding how perfluorination of (alkyl) hydrophobic groups alters the structure and dynamics of solvation shells, and thus the thermodynamics of solvation. It also points to the need of addressing this problem through the simultaneous use of techniques probing structure, dynamics and thermodynamics of solvation to gain comprehensive understanding. We do so by applying a combination of analytical models, classical molecular dynamics (CMD) simulations, ab initio molecular dynamics-density functional theory (AIMD-DFT) simulations, and transmission THz spectroscopy to study EtOH and TFE in aqueous solution. Classical molecular dynamics (CMD) simulations provide detailed statistics on solvation shell structures and input for the analytical Random Network Model to qualitatively quantify solvent–solvent entropy and enthalpy contributions. Simultaneously, comparing the solvation shell structure revealed by CMD and AIMD with known experimental spectroscopic markers for specific thermodynamically relevant solvation shell structures of the whole solvation shell by means of THz spectroscopy, we are able to connect these features to thermodynamic observables. The link between CMD-derived structural statistics and THz spectroscopic signatures was confirmed through density functional theory-based ab initio molecular dynamics (AIMD-DFT) simulations, which were consistent both with measured spectroscopic features and with the structure revealed from CMD. This integrated approach yields a comprehensive picture of solvation shell perturbation by these solutes and elucidates the molecular origins of their differing responses in the two solvent environments.
2 Methods
2.1 Transmission THz spectroscopy
TFE, 99.8% pure, was purchased from Thermo Fisher Scientific (catalog number: 13975, LOT number: A0449707) and used without further purification. EtOH, 99.8% pure was purchased from VWR (catalog number: 20821.321, LOT number: 25C194007) and used without further purification. Ultra pure water with a resistivity of 18.2 MΩ cm was used for sample preparation. The densities of all samples were measured at the corresponding temperature with a density meter (DMA 4500, Anton Paar, Graz, AT), to calculate the exact concentration of each sample from the masses of its components.Broadband THz-Far Infrared Fourier Transform absorption measurements were performed using a Vertex 80v FTIR spectrometer (Bruker, Billerica, MA, USA) with an aperture of 8 mm and equipped with a liquid helium cooled bolometer (Infrared Laboratories, Tucson, AZ, USA) as detector, a Globar as light source and a Mylar-Multilayer beam-splitter. The sample compartment was continuously purged with N2 gas to minimize atmospheric water vapor lines (relative humidity less than 1%), while the interferometer compartment was evacuated. Samples were loaded into a temperature-controlled transmission cell (TFC-S25-3, Harrick Scientific, Pleasantville, NY, USA) via a mini peristaltic pump (Ismatec MS-4/12 Reglo Digital, Zurich, CH). Sample temperatures were set to 20 °C and controlled with a coolant circulator (Huber Ministat 240, Offenburg, DE). The sample layer thickness was fixed by PTFE spacers with a nominal thickness of 13 µm, except for the pure TFE and EtOH measurements where spacers with a nominal thickness of 25 µm, 50 µm, 100 µm, 150 µm, 200 µm were used. The exact layer thicknesses were determined by recording the etalons of the empty cell in the mid-infrared region (3000 cm−1 to 7000 cm−1). Each sample was measured in the frequency range from 30 cm−1 to 680 cm−1 as an average over 128 scans with a resolution of 1 cm−1.
2.2 Ab initio molecular dynamics
We computed the THz spectra of one solute molecule, either EtOH or TFE, in water from AIMD simulations with a DFT functional, following established procedures (see ref. 35 and 40). In all cases, the simulation box included 256 water molecules, which corresponds approximately to two solvation shells around the solutes and little or no bulk-like water, and either 1 or zero solute molecules. For each system (solute molecule in water and water in the bulk), classical MD (CMD) was employed to generate ten structurally distinct configurations, which served as starting points for the subsequent AIMD simulations.All AIMD simulations used the revPBE functional, as it accurately reproduces the oxygen–oxygen radial distribution function of liquid water.41 For each configuration, an initial AIMD equilibration of 100 steps was carried out using a DZVP-MOLOPT-SR-GTH basis set in an NVT ensemble, with a global Nosé–Hoover thermostat applied every 10 steps. All subsequent simulations were performed using the TZV2P-MOLOPT-GTH basis set. This strategy was adopted to mitigate the so-called QM shock that can arise when initializing QM simulations directly from classical MD configurations. Direct equilibration with the TZV2P-MOLOPT-GTH basis set resulted in unstable SCF behavior and convergence failures. In contrast, initiating equilibration with the more robust DZVP-MOLOPT-SR-GTH basis set enabled the electronic structure to stabilize before progressively increasing the basis set flexibility. The change of the basis set was followed by a second equilibration stage to ensure that the system properly adapted to the corresponding level of theory. This equilibration consisted of a 1 ps “massive” thermalization run (2000 steps with a 0.5 fs time step), during which a Nosé–Hoover thermostat was applied every 10 steps to all atoms to bring the system to the target temperature. Finally, a third equilibration stage was carried out through 10 ps global thermalization with a Nosé–Hoover thermostat applied every 100 steps.
The final configuration of the last equilibration stage was used as the starting point for production runs performed in the NVE ensemble, of durations varying between 5 ps and 20 ps and during which the Wannier centers—localized representations of the electronic density—were computed. These were used as input for the software package TRAVIS,42 which calculated the vibrational spectra as the Fourier transform of the dipole moment autocorrelation function, corrected using the harmonic approximation.43 For each system, the ten resulting sets of Wannier center data were averaged to produce the final THz spectra for EtOH, TFE, and bulk water.
The statistical uncertainty of the calculated observables (including the calculated spectra), generically denoted as Y, was estimated as the standard error of the mean (SEM):
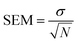 | (1) |
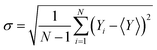 | (2) |
2.3 Classical molecular dynamics
Classical molecular dynamics (CMD) simulations were done using the TIP4P-Ew water model,44 and the force field optimized by Robalo and Vila Verde18 for the solutes. All simulation boxes are rectangular, with dimensions (10 × 4 × 4) nm3, and contain two solute molecules restrained at a minimum distance of ≈2.5 nm from each other—twice the distance at which the solute water radial distribution function converges to 1—to improve sampling while effectively remaining at infinite dilution. Systems were assembled and all simulations were performed using the Gromacs 2020.4 software package.45 The Lennard-Jones interactions were set to zero at 1.4 nm, and long range dispersion corrections were applied to both pressure and energy.46 Long-range electrostatics were treated with the Particle Mesh Ewald (PME)47 scheme, with a 1.4 nm cutoff, a grid spacing of 0.12 nm and a fourth order interpolation. Constraints (LINCS48) were applied to all bonds in the equilibration process and all bonds involving hydrogen atoms in the production phase. The accuracy of the LINCS solver was set with a single iteration and a Taylor expansion order of 4, balancing computational efficiency and numerical stability. For each simulated composition, a steepest descent minimization was performed. Integration of the equation of motion was done using a leapfrog algorithm for equilibration and with the stochastic dynamics (SD) algorithm for production runs, with a 1 fs time step. Systems were equilibrated for 100 ps in the NpT ensemble, employing the Berendsen barostat and thermostat.49 Each production run was 40 ns long, using the default Langevin thermostat with τT = 0.1 ps and T = 298 K, and the Parrinello – Rahman barostat50 with τp = 1 ps and reference pressure of 1 bar with a compressibility factor of 4.5 × 10−5 bar−1, saving the configurations for analysis every 10 ps. Fourteen production runs were performed per solute. Statistics for contacts were collected using in-house tcl scripts running in VMD,51 and molecular images were also obtained with that software package.2.4 Characterizing hydrogen bond networks from simulation
The geometry of O–H⋯O configurations was characterized through probability density distributions, ps(d, α), of the oxygen–oxygen distance, d, and HB angle, defined as α = 180°–(O–Ĥ⋯O) (graphically displayed in the Results section),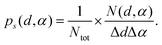 | (3) |
Perturbations of the hydrogen bond network structure for different water subpopulations a and b can then be quantified through pairwise subtractions of the ps(d, α), to obtain:
| Δp(d, α)a–b = pa(d, α) − pb(d, α). | (4) |
| ns(d, α) = ps(d, α) × ntOH,s × ΔdΔα. | (5) |
The magnitude, Δn(d, α), of structural differences between the solvation shells of the two alcohols can then be calculated as:
| Δn(d, α) = nTFE(d, α) − nEtOH(d, α). | (6) |
2.5 Random network model
The random network model (RNM) developed by Sceats and Rice52–54 was employed to analyze CMD simulations of bulk water and of aqueous systems containing either EtOH or TFE approaching infinitely dilute conditions. Briefly, the RNM connects the structure of water to its thermodynamic properties, by describing water as a random network referenced to the structure of ice I (which has a perfectly tetrahedral local geometry) at 0 K.52–54 The model explicitly accounts for the combinatorial entropy arising from the continual formation and disruption of hydrogen bonds, as well as for the mean energetic stabilization associated with each hydrogen bond, among other contributions. Importantly, the model does not constitute a fundamental statistical–mechanical theory, but rather an analytical construct aimed at separating the thermodynamic contributions of water molecules based on the characteristic timescales of different hydrogen-bonding modes. The enthalpy is obtained as the sum of the configurational, lattice vibrational and intramolecular distortion contributions; the entropy consists of the sum of the configurational and lattice vibrational contributions.53,54The RN model as implemented here takes as input only the following descriptors of water structure: the mean distance, ![[R with combining macron]](https://www.rsc.org/images/entities/i_char_0052_0304.gif) , between hydrogen bonded water oxygen atoms, the mean square deviation,
, between hydrogen bonded water oxygen atoms, the mean square deviation, 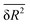 , of this distance, and the mean square value,
, of this distance, and the mean square value, 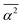 , of the angle α for hydrogen bonded configurations. These values were calculated from the CMD simulations considering that hydrogen bonds exist if the oxygen–oxygen distance is <3 Å and if α < 50°, and then rescaled to minimize force-field-dependency as described in SI Section S2.7.1. The hydrogen bond criteria used when calculating input values for the RN model deviate from those indicated in Section 1.4 because for the RN model we favored identifying first-neighbor water molecules while maximizing entropy–enthalpy cancellation in water of solvation. The values of the unscaled observables are shown in Table 1 for water in the bulk and water in the first solvation shell of TFE or EtOH; the scaled values are shown in Table S1. The solvation shell is defined by the cutoff distances, rcut,solv, of 5.58 Å (EtOH) and 5.72 Å (TFE) between the water oxygen and any of the carbon or oxygen atoms in the solutes. These distances reflect the position of the minima of the radial distribution functions reported in Section S2.6.
, of the angle α for hydrogen bonded configurations. These values were calculated from the CMD simulations considering that hydrogen bonds exist if the oxygen–oxygen distance is <3 Å and if α < 50°, and then rescaled to minimize force-field-dependency as described in SI Section S2.7.1. The hydrogen bond criteria used when calculating input values for the RN model deviate from those indicated in Section 1.4 because for the RN model we favored identifying first-neighbor water molecules while maximizing entropy–enthalpy cancellation in water of solvation. The values of the unscaled observables are shown in Table 1 for water in the bulk and water in the first solvation shell of TFE or EtOH; the scaled values are shown in Table S1. The solvation shell is defined by the cutoff distances, rcut,solv, of 5.58 Å (EtOH) and 5.72 Å (TFE) between the water oxygen and any of the carbon or oxygen atoms in the solutes. These distances reflect the position of the minima of the radial distribution functions reported in Section S2.6.
![[R with combining macron]](https://www.rsc.org/images/entities/i_char_0052_0304.gif) ,
, 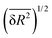 for water in the bulk, and water in the first solvation shell of the indicated solutes, obtained from CMD simulations
for water in the bulk, and water in the first solvation shell of the indicated solutes, obtained from CMD simulations
For solvation shells beyond the first, water is substantially less perturbed relative to bulk water, rendering the difference of the ![[R with combining macron]](https://www.rsc.org/images/entities/i_char_0052_0304.gif) ,
, 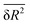 and
and 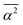 observables between those solvation shells and water in the bulk comparable to their statistical uncertainty. That high uncertainty combined with the larger number of water molecules in solvation shells beyond the first introduce significant statistical noise in the thermodynamic observables calculated using the RN model. For this reason, we restrict our analysis using the RN model to the first solvation shell.
observables between those solvation shells and water in the bulk comparable to their statistical uncertainty. That high uncertainty combined with the larger number of water molecules in solvation shells beyond the first introduce significant statistical noise in the thermodynamic observables calculated using the RN model. For this reason, we restrict our analysis using the RN model to the first solvation shell.
The entropic and enthalpic components to the thermodynamics of hydration of the first solvation shell of each solute was calculated relative to water in the bulk as:
| ΔB11 = N11(B11 − B0) | (7) |
3 Results and discussion
3.1 Different thermodynamics of solvation between TFE and EtOH
Differences in solvation behaviour between EtOH and TFE can be advantageously quantified by looking at the thermodynamic excess mixing properties, which can be measured accurately. The excess properties refer to the non-ideal part of mixing and can be calculated in general by the following equation for a certain property y: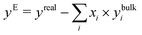 | (8) |
Here, yE is the excess molar property of the mixture, yreal is the real, measurable molar property of the mixture, xi are the mole-fractions of the components of the mixture and ybulki is the measurable molar property y of the pure component i. This expression gives a direct estimate of how interactions between components of a mixture change the overall properties. Fig. 1 shows selected excess mixing thermodynamic properties for aqueous mixtures of EtOH and TFE.
 | ||
| Fig. 1 Comparison of selected thermodynamic properties of EtOH (black; ΔGE and ΔHE from ref. 38 and 55 with −TΔSE calculated from the two, ϕEV from ref. 56 and 57) and TFE (green; ΔGE and ΔHE from ref. 37 with −TΔSE calculated from the two, ϕEV from ref. 58) in water as function of their mole fraction. (i) Excess Gibbs free energy of mixing ΔGE. (ii) Excess enthalpy of mixing ΔHE. (iii) Excess entropic contribution of mixing (−TΔS)E. (iv) Excess apparent molar volume ϕEV of mixing. | ||
The excess Gibbs free energy of mixing in Fig. 1(i) shows a more unfavourable mixing for TFE compared to EtOH but in general the two substances share a similar behaviour. This is in line with the more hydrophobic nature of TFE compared to EtOH, as indicated by its less favourable solvation free energy.30 However, looking at the enthalpic and entropic contributions to the overall Gibbs free energy of mixing, significant differences between the two substances arise. The excess mixing enthalpy in Fig. 1(ii) is always negative for EtOH, while for TFE it is only negative for low concentrations and even then it is always less negative than for EtOH. This difference is surprising because TFE is still fully miscible with water, and points towards weaker and even unfavourable TFE-water interactions. The entropic contribution shown in Fig. 1(iii), however, shows that entropy of solvation is substantially unfavourable across the whole mixing range for EtOH, whereas for TFE it is less unfavourable, and in fact becomes favourable in solutions with high TFE concentration. Lastly, the excess apparent molar volume in Fig. 1(iv) shows smaller deviations from the ideal behaviour for TFE, which points to less pertubation of the system by TFE compared to EtOH and less strong interactions between water and TFE compared to water and EtOH.
3.2 THz spectroscopy
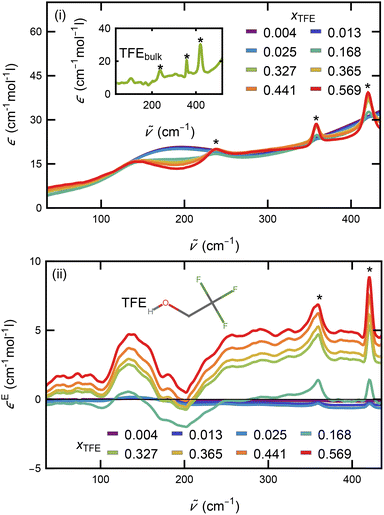
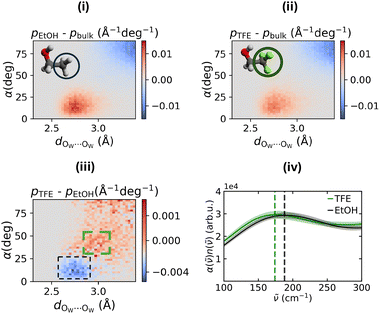
In order to have spectroscopic results comparable to these thermodynamic properties, we measured TFE-water mixtures across the whole mixing range (Fig. 2(i)) and EtOH-water mixtures in the first fifth of the mixing range (Fig. S1(i)), as well as spectra of the pure components (insets in Fig. 2(i) and Fig. S1(i)), in the THz range from 30 cm−1 to 680 cm−1. From these measurements we calculated the excess molar extinction coefficient,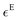 , by subtracting the mole-fraction-weighted bulk molar extinction coefficients of TFE or EtOH and of water from the mixture, in line with eqn (8):
, by subtracting the mole-fraction-weighted bulk molar extinction coefficients of TFE or EtOH and of water from the mixture, in line with eqn (8):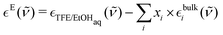 | (9) |
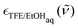 being the frequency-dependent (
being the frequency-dependent (![[small nu, Greek, tilde]](https://www.rsc.org/images/entities/i_char_e0e1.gif) ) molar extinction coefficient of the aqueous alcohol mixture,
) molar extinction coefficient of the aqueous alcohol mixture, 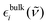 being the frequency-dependent molar extinction coefficient of bulk substance i and xi being the mole fraction of that substance, where i is water, TFE or EtOH. This procedure eliminates contributions to the spectra arising from either bulk water, bulk TFE or bulk EtOH interactions, and leaves only the differences relative to the bulk phases due to interactions between water and TFE or water and EtOH, as well as altered water–water, TFE–TFE and EtOH–EtOH interactions resulting from mixing, effectively probing the local solvation environment.
being the frequency-dependent molar extinction coefficient of bulk substance i and xi being the mole fraction of that substance, where i is water, TFE or EtOH. This procedure eliminates contributions to the spectra arising from either bulk water, bulk TFE or bulk EtOH interactions, and leaves only the differences relative to the bulk phases due to interactions between water and TFE or water and EtOH, as well as altered water–water, TFE–TFE and EtOH–EtOH interactions resulting from mixing, effectively probing the local solvation environment.
The excess molar extinction coefficients of TFE solutions at selected mole-fractions of TFE are shown in Fig. 2(ii) in the frequency range from 35 cm−1 to 435 cm−1. Several absorption features can be seen. The overall intensity initially drops and starts to increase for xTFE > 0.168. The broad absorption at 140 cm−1 continuously increases in intensity with increasing TFE concentration. This peak resembles the cavity-wrap water feature identified for all alcohols from methanol to pentanol from our prior work developing the concept of THz-Calorimetry,33–35,59 and the simulation results presented below confirm this assignment also for TFE. A more detailed explanation of THz-Calorimetry and an analysis and comparison with EtOH is given below.
Beyond 250 cm−1 the spectra are quite flat except for two sharp absorption modes at 360 cm−1 and 420 cm−1. These features are already present for the lowest concentration of TFE in water and are also present in pure TFE with an additional absorption at 236 cm−1 (see inset in Fig. 2(i); these features are marked with asterisks). We therefore assign these absorption modes to intramolecular TFE vibrations, supported by Raman measurements of pure liquid TFE.60 Although we subtracted a scaled bulk TFE molar extinction (eqn (9)), these modes are still visible, indicating that the different environment in aqueous vs. bulk TFE led to changes in the intensity of these absorptions. That these peaks do not show a concentration-dependent frequency shift suggests that the amount of TFE aggregation is not significant. These results cannot exclude that transient TFE aggregates form but are not detected by the techniques used here; separate studies are necessary to investigate this possibility.
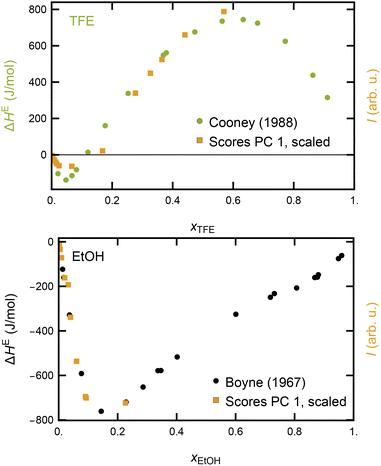 | ||
| Fig. 3 Qualitative comparison between TFE mixing enthalpy (from ref. 37) and scaled (×1289) results from the PCA (upper panel) and between EtOH mixing enthalpy (from ref. 38) and scaled (×1404) results from the PCA (lower panel). The frame label on the right is kept in the same color as the data points of the scores and labeled “I (arb. u.)” to point out that the scores do not share the same unit as the thermodynamic dataset. | ||
The hydrophilic (also called bound-water) contribution arises from water molecules that directly hydrogen bond to the solute. This interaction mainly contributes enthalpically to the solvation process. Formation of a water–solute hydrogen bond modifies the librational mode of the water network, which is located at 680 cm−1 in bulk water. In non-fluorinated alcohols, this solute–water hydrogen bond leads to a stiffening of the librational motion of solvation water molecules, meaning a loss of softer librations and an increase in stiffer librations.33,35 Since we look at the difference to bulk solvent and solute, this stiffening in librational motion should result in a negative contribution at lower frequencies and a positive contribution at higher frequencies, as depicted in the inset in Fig. 4 and from here on called bound-water signature. Our previous studies revealed that this bound-water signature was present in all aqueous solutions of various non-fluorinated alcohols investigated.
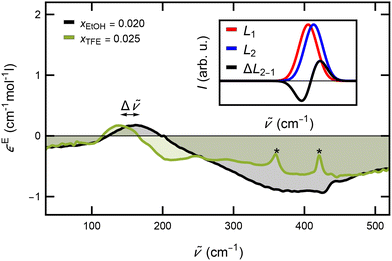 | ||
Fig. 4 Excess molar extinction coefficients of aqueous TFE (green) and EtOH (black) solutions at similar mole-fractions. A redshift (marked by Δ![[small nu, Greek, tilde]](https://www.rsc.org/images/entities/i_char_e0e1.gif) ) by 30 cm−1 of the absorption mode at low frequencies is present for TFE compared to EtOH, indicating a weakened HB network around TFE. Sharp absorption modes at 360 cm−1 and 420 cm−1, only present for TFE and marked by asterisks, can be assigned to intramolecular vibrations of TFE (see Fig. 2 and ref. 60 and 64). The inset explains the origin of the typical bound-water lineshape resulting from a frequency shift of the librational mode. The difference between a reference Gaussian (in red, L1) and a blue-shifted one (in blue, L2) results in a lineshape that has a negative contribution at lower frequencies and a positive contribution at higher frequencies (ΔL2-1). ) by 30 cm−1 of the absorption mode at low frequencies is present for TFE compared to EtOH, indicating a weakened HB network around TFE. Sharp absorption modes at 360 cm−1 and 420 cm−1, only present for TFE and marked by asterisks, can be assigned to intramolecular vibrations of TFE (see Fig. 2 and ref. 60 and 64). The inset explains the origin of the typical bound-water lineshape resulting from a frequency shift of the librational mode. The difference between a reference Gaussian (in red, L1) and a blue-shifted one (in blue, L2) results in a lineshape that has a negative contribution at lower frequencies and a positive contribution at higher frequencies (ΔL2-1). | ||
In short, the THz results show that our excess molar extinction spectra connect to macroscopic excess mixing thermodynamics measurements. The lower integrated excess molar extinction of TFE is in line with its more ideal mixing behaviour from apparent molar volume measurements. The collective motions of the HB network of water probed via THz spectroscopy further reveal an overall looser HB network of water around TFE. The lack of a clear bound-water signature but a distinct presence of a hydrophobic-hydration water mode for TFE compared to EtOH spectroscopically explain the stronger entropic solvation signature of TFE relative to EtOH. In what follows, we connect the spectroscopic features detected experimentally to water structure and thermodynamics of hydration with the help of molecular simulations and the Random Network model.
3.3 AIMD and classical MD capture solvation shell structure
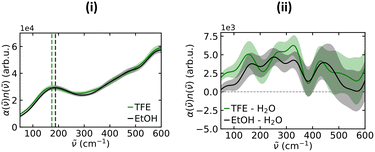
Reliable studies of solvation shell structure using simulations require both a good quality description of the system and good sampling. AIMD simulations offer a direct way of assessing quality through computational predictions of the vibrational spectra in the THz region, which can be compared with experiments. Here, we computed the THz spectra for one molecule of EtOH or TFE in water, as well as for water in the bulk, using AIMD.Fig. 5 presents the THz spectra calculated from the AIMD simulations. The maximum near 200 cm−1 is its most prominent feature and corresponds to the equivalent feature observed in experiment for water and aqueous solutions of alcohols.33,66 It is slightly red-shifted for TFE compared to EtOH, which is in agreement with the experimental results shown in Fig. 4, indicating that the computational model provides an accurate description of the solvation shells for both solutes.
To further assess the quality of the AIMD simulations we calculate the probability of finding a water dangling OH—namely, an OH group with no HB with other water molecules or solutes and that points towards the hydrophobic part of the solute—within the first solvation shell. It is 0.058 ± 0.007 for TFE and 0.016 ± 0.002 for EtOH, indicating a 3.625 fold increase for TFE. This result is in good agreement with Raman-MCR results, which clearly indicate that dangling OH groups, identifiable by a peak at ≈3660 cm−1, are substantially more abundant near TFE than near EtOH.32
Given that the main feature of the THz spectra computed in these simulations is in good agreement with experimental measurements and that these spectra reflect a water structure (shown in SI Section S2.3) consistent with those obtained from CMD simulations (shown in Fig. 7 and in SI Section S2.4), we are confident that the solvation structures sampled through classical MD—benefiting from higher statistical sampling and significantly lower computational cost compared to AIMD—can reliably capture the distinct behaviours of the solvation shells surrounding these two solutes. We base our remaining analysis and discussion on results obtained using classical simulations only.
3.4 Structure and spectroscopic response of solvation shells
We characterise the hydrogen bond network in the simulated systems through the geometry of O–H⋯O configuration, illustrated in Fig. 6. Specifically, we calculate joint probability density distributions, ps(d, α), of the O to O distance, d, and HB angle, α, where s identifies an aqueous system with a solute (EtOH or TFE) or water in the bulk (see eqn (3)). Within this framework, we identify a hydrogen bond when d < 3.2 Å and α < 45°. Fig. 6 shows pTFE(d, α), which describes the water structure in the first hydration shell of TFE, considering only distances and angles between water molecules. The distribution peaks at d ≈ 2.8 Å and α ≈ 15°, typical for the water hydrogen bond network.67 | ||
| Fig. 8 Difference, Δn, between the solvation shell of TFE and EtOH (eqn (6)) considering (i) all the oxygen atoms in the solutes and their first solvation shells, (ii) water OH relative to the alcohol O of the solutes, (iii) alcohol OH group relative to the water oxygen, (iv) water–water within the solvation shell cutoff. Results obtained with classical MD simulations. The dashed lines indicate the cutoffs used to identify hydrogen bonded configurations. | ||
For any (d, α) combination, Δntot is positive, reflecting the fact that the radius of TFE's solvation shell is larger by approximately 0.14 Å, i.e. it encompasses a larger number of water molecules. Nevertheless, the results in Fig. 7(i) and (ii) indicate that each OH⋯H structure in that solvation shell is less perturbed relative to the bulk than for EtOH. Prior Raman-MCR data showed that the OH-stretch region is substantially more intense (after normalizing by concentration) for EtOH(aq) than for TFE(aq).68 Likewise, the unsigned integral of the THz spectra in Fig. 4, which estimates the magnitude of the perturbation of solvation shells relative to bulk, is also larger for EtOH(aq). A higher intensity can arise from a larger number of water molecules that are weakly perturbed or from a smaller number of water molecules experiencing a larger perturbation. The present simulation results clarify that the lower Raman-MCR intensity and lower unsigned integral of the THz spectrum associated with the solvation shell of TFE can be explained by water–water hydrogen bonds experiencing a smaller perturbation relative to the bulk than EtOH.
Fig. 8(ii)–(iv) show the contributions of water-to-alcohol, alcohol-to-water and water–water populations to Fig. 8(i). Clearly, the dominant contribution comes from water–water hydrogen bonds, shown in Fig. 8(iv), as expected. Nevertheless, the results show that replacing CH3 by CF3 substantially modifies direct alcohol–water hydrogen bonds, as shown in Fig. 8(ii) and (iii). These results are consistent with 1H NMR measurements and simulations, which confirm that the TFE hydroxy hydrogen has a higher hydrogen bonding ability than that of EtOH, whereas the TFE oxygen has a much lower hydrogen bonding ability than that of EtOH.65
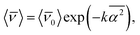 | (10) |
![[small nu, Greek, macron]](https://www.rsc.org/images/entities/i_char_e0ce.gif) 0〉 being the average frequency in a given reference state, k a positive constant and
0〉 being the average frequency in a given reference state, k a positive constant and 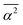 the mean square hydrogen bond angle.52–54 In our CMD simulations,
the mean square hydrogen bond angle.52–54 In our CMD simulations, 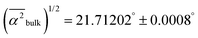 for water in the bulk, whereas for water-to-alcohol hydrogen bonds with TFE it is larger
for water in the bulk, whereas for water-to-alcohol hydrogen bonds with TFE it is larger 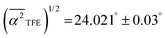 and for EtOH it is smaller
and for EtOH it is smaller 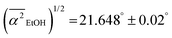 . These results confirm that the average hydrogen bond angle associated water-to-EtOH hydrogen bonds is smaller than bulk, and thus should lead to a blue-shift in the libration frequency and to a bound-water signature, consistent with the experimentally measured THz spectra. By contrast, the weaker water-to-alcohol hydrogen bonds in case of TFE give rise to a broader distribution in their hydrogen bond angle and should lead to a red-shift in the frequency of this libration mode. The red-shift, combined with the lower abundance of these hydrogen bonds, is consistent with the absence of the bound-water signature in the THz spectra of TFE(aq).
. These results confirm that the average hydrogen bond angle associated water-to-EtOH hydrogen bonds is smaller than bulk, and thus should lead to a blue-shift in the libration frequency and to a bound-water signature, consistent with the experimentally measured THz spectra. By contrast, the weaker water-to-alcohol hydrogen bonds in case of TFE give rise to a broader distribution in their hydrogen bond angle and should lead to a red-shift in the frequency of this libration mode. The red-shift, combined with the lower abundance of these hydrogen bonds, is consistent with the absence of the bound-water signature in the THz spectra of TFE(aq).
3.5 Thermodynamics of solvation, and its connection to structure and spectroscopy
Briefly, the RN model assumes that the thermodynamics of liquid water can be described by the mean position of its water oxygen atoms, which function as nodes connected by a random hydrogen-bond network (also called lattice), together with the thermal fluctuations of the network; further details are given in the Methods section. The model successfully describes the thermodynamics of solid and liquid water in a wide range of temperatures52–54 as well as the thermodynamics of solvation of both non-polar and ionic solutes.70,71Within this framework, the solvent–solvent free energy is expressed as
| Gtot,vv = Htot,vv − TStot,vv | (11) |
The Random Network Model results for TFE and EtOH, summarized in Table 2, clearly indicate that they perturb their solvation shells qualitatively similarly: both pay an entropic cost (negative ΔSvv,11), compensated by an enthalpic gain (negative ΔHvv,11). Nevertheless, there are substantial quantitative differences between them.
| ΔSvv,11 (J mol−1 K−1) | ΔHvv,11 (J mol−1) | |
|---|---|---|
| TFE | −2.30 ± 0.05 | −795 ± 45 |
| EtOH | −6.80 ± 0.05 | −1922 ± 40 |
| ΔΔSvv,11 | ΔΔHvv,11 | |
| −4.50 ± 0.07 | −1127 ± 60 |
EtOH perturbs its solvation shell substantially more than TFE relative to water in the bulk, as indicated by its larger absolute values of ΔSvv,11 and ΔHvv,11. This difference does not directly reflect alcohol-water hydrogen bonds, which are not considered in the RN model, but arises solely from differences in the water–water network of their solvation shells. As demonstrated in Fig. 7(i)–(iii), EtOH induces a larger perturbation per water molecule in its solvation shell than TFE. This effect dominates over the slightly larger number of water molecules in TFE's first solvation shell relative to EtOH's (Table 2, caption). As discussed in the previous section, these results are consistent with the higher spectroscopic signal in the THz and Raman region associated with the solvation shell of EtOH vs. TFE. It also agrees with the experimental thermodynamics of solvation data shown in Fig. 1(ii) and (iii) in the dilute regime, indicating that changes in solvent–solvent interactions upon solvation contribute substantially towards the experimentally-measured excess solvation properties (which include the solute–solvent contribution).
By construction, the RN model decomposes the liquid water enthalpy and entropy into different terms. The model relies on the assumption that there is a separation of time scales for different water motions: libration (450 cm−1 to 900 cm−1) and hindered translation (60 cm−1 to 200 cm−1) are fast and occur within a quasi-static network of hydrogen bonded molecules, whereas diffusion and relaxation processes related to it occur more slowly. As a result, the energy and entropy of the random network consist of configurational terms associated with the average position of the molecules, and vibrational terms comprising an intramolecular term plus “lattice modes” relative to a static configuration of the random network.53 The “lattice modes” can be librational, OO translational and hydrogen bond bending modes.
The main terms responsible for the different solvent–solvent thermodynamics of solvation between TFE and EtOH are summarized in Table 3, together with the structural parameter on which they depend; all terms are shown in SI Section S2.7.2. These results clearly indicate that differences in thermodynamics of solvation between EtOH and TFE can be traced back to differences in the mean square hydrogen bond angle 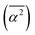 in their solvation shells. This angle is narrowest for EtOH (Table S1), leading to a more favourable configurational enthalpy associated with angle bending in the solvation shell of EtOH than in the solvation shell of TFE or for water in the bulk. Simultaneously, this angle also makes the lattice libration enthalpy, the configurational entropy associated with angle bending, and the entropies of lattice hydrogen bond bending, libration and OO translation less favourable for EtOH than for TFE or for water in the bulk.
in their solvation shells. This angle is narrowest for EtOH (Table S1), leading to a more favourable configurational enthalpy associated with angle bending in the solvation shell of EtOH than in the solvation shell of TFE or for water in the bulk. Simultaneously, this angle also makes the lattice libration enthalpy, the configurational entropy associated with angle bending, and the entropies of lattice hydrogen bond bending, libration and OO translation less favourable for EtOH than for TFE or for water in the bulk.
Additionally, the average distance between water oxygen atoms is smallest for the solvation shell of EtOH (Table S1). This difference makes both the configurational enthalpy associated with OO stretching and the vibrational intramolecular zero-point distortion contribution more favourable for the solvation shell of EtOH than for TFE, but has little effect on solvation entropy (Table 3). Nevertheless, this difference in solvation shell structure between TFE and EtOH has a much smaller impact on the thermodynamics of solvation than the changes in hydrogen bond angle.
The average OO distance between hydrogen bonded water molecules in the solvation shell of TFE and of EtOH is smaller than for water in the bulk (Table 1 and Table S1). This difference suggests that hydrogen bonds are stronger in the solvation shells than in water in the bulk. How can this finding be reconciled with the red-shifted OO stretching band in the solvation shells of TFE and EtOH relative to water in the bulk, seen in the THz spectra (Fig. 4 and 5, 140 cm−1 and 165 cm−1)? This apparent contradiction originates from the fact that the solvation shell subpopulations with stretched hydrogen bonds, which give rise to the red-shifted OO stretching bands, are small and contribute little to the average values listed in Table S1 and Table 1. Moreover, we calculated the average OO distances between hydrogen bonded water molecules separately for the cavity-wrap and for the HBhyd2bulk, subpopulations near CF3 and CH3, as shown in Table 4. We found that both subpopulations near CF3 exhibit larger OO distances compared to bulk water, consistent with an overall weakening of hydrogen bonding and a red shift in the hydrogen-bond stretching region. In contrast, for EtOH, only the HBhyd2bulk population has larger OO distance than bulk, with the cavity-wrap water showing slightly shorter OO distances than bulk. These results are robust to the choice of rcut to define the wrap-water and HBhyd2bulk populations (Fig. S9).
![[R with combining macron]](https://www.rsc.org/images/entities/i_char_0052_0304.gif) , between neighbouring water oxygen atoms (for which oxygen–oxygen distance <3.5 Å and α < 40°) for the wrap- and HBhyd2bulk populations near CH3 and of CF3. The wrap water population consists of hydrogen bond donors and acceptors within the cutoff distance, rcut < 3.66 Å, between water oxygen and the carbon atom of these groups; the HBhyd2bulk consists of hydrogen bonds between wrap-water molecules and water molecules beyond rcut. Differences to the bulk values Δ
, between neighbouring water oxygen atoms (for which oxygen–oxygen distance <3.5 Å and α < 40°) for the wrap- and HBhyd2bulk populations near CH3 and of CF3. The wrap water population consists of hydrogen bond donors and acceptors within the cutoff distance, rcut < 3.66 Å, between water oxygen and the carbon atom of these groups; the HBhyd2bulk consists of hydrogen bonds between wrap-water molecules and water molecules beyond rcut. Differences to the bulk values Δ![[R with combining macron]](https://www.rsc.org/images/entities/i_char_0052_0304.gif) are shown to facilitate comparisons
are shown to facilitate comparisons
| Population |
![[R with combining macron]](https://www.rsc.org/images/entities/i_char_0052_0304.gif) (Å) (Å) |
Δ![[R with combining macron]](https://www.rsc.org/images/entities/i_char_0052_0304.gif) (Å) (Å) |
|---|---|---|
| Bulk | 2.85113 | — |
| ±0.0001 | — | |
| EtOH | 2.84473 | -0.00640 |
| wrap | ±0.0006 | ±0.0006 |
| TFE | 2.85775 | 0.00662 |
| wrap | ±0.0009 | ±0.0009 |
| EtOH | 2.85558 | 0.00445 |
| HBhyd2bulk | ±0.0001 | ±0.0001 |
| TFE | 2.86168 | 0.01055 |
| HBhyd2bulk | ±0.0002 | ±0.0002 |
4 Conclusions
We have used THz spectroscopy, classical and ab initio molecular dynamics simulations and the Random Network Model of water developed by Sceats et al.,52–54 to investigate the structure, dynamics and thermodynamics of hydration of methyl and perfluoromethyl groups, using TFE and EtOH in water as model systems.The higher hydrophobicity of TFE relative to EtOH reflects partially TFE's lower number of water-to-alcohol hydrogen bonds, and the fact that they are also weaker than water-to-EtOH hydrogen bonds, as demonstrated by AIMD and CMD simulations (Fig. 8). Both aspects are visible in the water libration part of the experimental THz spectra (Fig. 4, for ![[small nu, Greek, macron]](https://www.rsc.org/images/entities/i_char_e0ce.gif) > 200 cm−1), with the excess THz spectra of EtOH(aq) showing the bound-water feature that is expected when a libration mode shifts to higher frequencies relative to water in the bulk. Higher libration frequencies are expected when the average hydrogen bond angle is smaller.54 By contrast, this signature is not visible in the excess THz spectra of TFE(aq).
> 200 cm−1), with the excess THz spectra of EtOH(aq) showing the bound-water feature that is expected when a libration mode shifts to higher frequencies relative to water in the bulk. Higher libration frequencies are expected when the average hydrogen bond angle is smaller.54 By contrast, this signature is not visible in the excess THz spectra of TFE(aq).
A body of evidence has demonstrated CF3 groups are more hydrophobic than CH319,25,28,30 due to their larger volume,31 and thus also contribute to the higher hydrophobicity of TFE relative to EtOH. Our results add to this picture, clarifying that the different water perturbation induced by CF3 groups relative to CH3 cannot be explained by their different volume. Instead, their different perturbation signals that alkyl and perfluoroalkyl groups have different hydrophobic hydration. Specifically, CF3 groups weaken the hydrogen bonds in their hydrophobic solvation shell more than CH3. As a result, the hydrogen bond stretching mode found around 165 cm−1 and associated with hydrophobic solvation of alkyl substituents33 is substantially red shifted, to 140 cm−1, for TFE(aq) relative to EtOH(aq). These results add to prior work by some of us, that indicated that the solvation shell of perfluorinated groups is fundamentally different from that of alkyl groups because perfluorinated groups induce substantially more dangling water OH groups than the corresponding alkyl groups due to stabilizing electrostatic and dispersion interactions stemming from the polarity of the CF bond.19,20,32 The substantially more frequent dangling OH groups near perfluorinated groups possibly contribute to the unusual weakening (larger OO distance) of the hydrogen bond network near them.
Despite the fundamental differences in hydration between CF3 and CH3, the larger volume of CF3 and the different water-to-alcohol hydrogen bond propensities of EtOH vs. TFE, TFE perturbs its water of solvation far less than EtOH as clearly visible in our results: the solvent–solvent enthalpy of hydration of EtOH is substantially more favourable (more negative) than for TFE (Table 2), in part because the probability that a water–water hydrogen bond forms in EtOH's solvation shell is larger than in the bulk and than near TFE (Fig. 7), whereas the solvent–solvent entropy of solvation is (necessarily) more unfavourable for EtOH. The smaller overall perturbation is spectroscopically detectable also in the smaller unsigned integrals of the excess THz spectra of TFE(aq) and EtOH(aq) and in excess Raman spectra.32
These results have interesting implications for materials and drug design, as the replacement of alkyl groups by their perfluorinated counterparts will increase hydrophobicity while decreasing the structural perturbation per water molecule in their solvation shells. Moreover, they are relevant for studying the mechanisms by which water mixtures of either TFE or ethanol stabilize helical conformations in proteins.73–77 Simulation results indicate that, for TFE, this effect correlates with non-specific interactions between TFE and the protein, and that the TFE-carbonyl hydrogen bond destabilizes the helical structure contrary to expectations.75 Intermolecular Nuclear Overhauser effect measurements and molecular dynamics simulations suggest these contributions are also at play for EtOH.76 Less understood is the possible contribution of alcohol-induced perturbation of solvent structure to the effect of either alcohol on conformational stability of proteins. Our results suggest that this contribution might be distinct for TFE relative to EtOH because of their different impact in solvent structure. The lower hydrogen-bonding ability of TFE's hydroxy group suggests that it might have a lower ability than EtOH to hydrogen bond to the carbonyl groups in the protein backbone, consistent with TFE's stronger effect on protein stability. Our results point to the need of comparative studies between the two alcohols to mechanistically understand changes in protein conformational stability as a function of alcohol concentration in the binary aqueous solvent.
Conflicts of interest
The authors have no conflict of interest to report.Data availability
Data supporting this article have been included as part of the supplementary information (SI). Replication data (simulation and experiment) is available free of charge at https://doi.org/10.17877/RESOLV-2025-MI644MUC, TUDOdata. Supplementary information: further computational and experimental details. See DOI: https://doi.org/10.1039/d5cp04598c.Acknowledgements
Funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany's Excellence Strategy – EXC 2033 – 390677874 – RESOLV. The authors gratefully acknowledge the Gauss Centre for Super-computing e.V. (https://www.gauss-centre.eu) for providing computing time on the national supercomputer HPE Apollo Hawk at the High Performance Computing Center Stuttgart (HLRS) under the grant numbers THySoF/44298 and HySoF/44310.References
- G. W. Gribble, The handbook of environmental chemistry, Springer-Verlag, 2002, ch. 5, vol. 3, pp. 121–136 Search PubMed.
- J. M. T. Thompson and G. Sandford, Organofluorine chemistry, Philos. Trans. R. Soc., A, 2000, 358, 455–471 CrossRef.
- F. H. Nielsen, Micronutrients in parenteral nutrition: boron, silicon, and fluoride, Gastroenterology, 2009, 137, S55–S60 CrossRef CAS PubMed.
- N. R. Johnston and S. A. Strobel, Principles of fluoride toxicity and the cellular response: a review, Arch. Toxicol., 2020, 94, 1051–1069 CrossRef CAS PubMed.
- M. Wei, Y. Ye, M. M. Ali, Y. Chamba, J. Tang and P. Shang, Effect of fluoride on cytotoxicity involved in mitochondrial dysfunction: a review of mechanism, Front. Vet. Sci., 2022, 9, 850771 CrossRef PubMed.
- S. Guth, S. Hüser, A. Roth, G. Degen, P. Diel, K. Edlund, G. Eisenbrand, K.-H. Engel, B. Epe, T. Grune, V. Heinz, T. Henle, H.-U. Humpf, H. Jäger, H.-G. Joost, S. E. Kulling, A. Lampen, A. Mally, R. Marchan, D. Marko, E. Mühle, M. A. Nitsche, E. Röhrdanz, R. Stadler, C. van Thriel, S. Vieths, R. F. Vogel, E. Wascher, C. Watzl, U. Nöthlings and J. G. Hengstler, Toxicity of fluoride: critical evaluation of evidence for human developmental neurotoxicity in epidemiological studies, animal experiments and in vitro analyses, Arch. Toxicol., 2020, 94, 1375–1415 CrossRef CAS PubMed.
- A. Di Giorgi, N. La Maida, O. Taoussi, S. Pichini, F. P. Busardò, A. Tini and A. Di Trana, Analysis of perfluoroalkyl substances (PFAS) in conventional and unconventional matrices: clinical outcomes, J. Pharm. Biomed. Anal. Open, 2023, 1, 100002 CrossRef.
- S. Purser, P. R. Moore, S. Swallow and V. Gouverneur, Fluorine in medicinal chemistry, Chem. Soc. Rev., 2008, 37, 320–330 RSC.
- M. Salwiczek, E. K. Nyakatura, U. I. M. Gerling, S. Ye and B. Koksch, Fluorinated amino acids: compatibility with native protein structures and effects on protein-protein interactions, Chem. Soc. Rev., 2012, 41, 2135–2171 RSC.
- B. E. Smart, Fluorine substituent effects (on bioactivity), J. Fluorine Chem., 2001, 109, 3–11 CrossRef CAS.
- E. Henary, S. Casa, T. L. Dost, J. C. Sloop and M. Henary, The role of small molecules containing fluorine atoms in medicine and imaging applications, Pharmaceuticals, 2024, 17, 281 CrossRef CAS PubMed.
- J. Wang, M. Sánchez-Roselló, J. Aceña, C. del Pozo, A. E. Sorochinsky, S. Fustero, V. A. Soloshonok and H. Liu, Fluorine in pharmaceutical industry: fluorinecontaining drugs introduced to the market in the last decade (2001–2011), Chem. Rev., 2014, 114, 2432–2506 CrossRef CAS PubMed.
- R. Dams and K. Hintzer, in Industrial aspects of fluorinated oligomers and polymers, ed. B. Ameduri and H. Sawada, RSC, 2016, ch. 1, vol. 2, pp. 1–31 Search PubMed.
- R. Maynard, Proc. of the CORROSION, 2019, pp. NACE-2019–12903 Search PubMed.
- B. J. Place and J. A. Field, Identification of novel fluorochemicals in aqueous film-forming foams used by the US military, Environ. Sci. Technol., 2012, 46, 7120–7127 CrossRef CAS PubMed.
- E. Wilhelm, R. Battino and R. J. Wilcock, Lowpressure solubility of gases in liquid water, Chem. Rev., 1977, 77, 219–262 CrossRef CAS.
- O. N. Tretinnikov and Y. Ikada, Dynamic wetting and contact angle hysteresis of polymer surfaces studied with the modified wilhelmy balance method, Langmuir, 1994, 10, 1606–1614 CrossRef CAS.
- J. R. Robalo, S. Huhmann, B. Koksch and A. Vila Verde, The multiple origins of the hydrophobicity of fluorinated apolar amino acids, Chem, 2017, 3, 881–897 CAS.
- J. R. Robalo, D. M. de Oliveira, P. Imhof, D. Ben-Amotz and A. Vila Verde, Quantifying how step-wise fluorination tunes local solute hydrophobicity, hydration shell thermodynamics and the quantum mechanical contributions of solute–water interactions, Phys. Chem. Chem. Phys., 2020, 22, 22997–23008 RSC.
- J. R. Robalo, D. Mendes de Oliveira, D. Ben-Amotz and A. Vila Verde, Influence of methylene fluorination and chain length on the hydration shell structure and thermodynamics of linear diols, J. Phys. Chem. B, 2021, 125, 13552–13564 CrossRef CAS PubMed.
- K. A. Dill, Dominant forces in protein folding, Biochemistry, 1990, 29, 7133–7155 CrossRef CAS PubMed.
- E. N. G. Marsh, Fluorinated proteins: from design and synthesis to structure and stability, Acc. Chem. Res., 2014, 47, 2878–2886 CrossRef CAS PubMed.
- A. Langhans, M. Krummhaar, C. Roth and B. Koksch, Catalytically competent fluorinated barnase variants, Synlett, 2024, 1042–1046 CAS.
- B. C. Buer, J. L. Meagher, J. A. Stuckey and E. N. G. Marsh, Structural basis for the enhanced stability of highly fluorinated proteins, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 4810–4815 CrossRef CAS PubMed.
- L. Wehrhan, J. Leppkes, N. Dimos, B. Loll, B. Koksch and B. G. Keller, Water Network in the Binding Pocket of Fluorinated BPTI–Trypsin Complexes-Insights from Simulation and Experiment, J. Phys. Chem. B, 2022, 126, 9985–9999 CrossRef CAS PubMed.
- J. Mecinović, P. W. Snyder, K. A. Mirica, S. Bai, E. T. Mack, R. L. Kwant, D. T. Moustakas, A. Héroux and G. M. Whitesides, Fluoroalkyl and alkyl chains have similar hydrophobicities in binding to the “hydrophobic wall” of carbonic anhydrase, J. Am. Chem. Soc., 2011, 133, 14017–14026 CrossRef PubMed.
- C. Adam, L. Yang and S. L. Cockroft, Partitioning solvophobic and dispersion forces in alkyl and perfluoroalkyl cohesion, Angew. Chem., Int. Ed., 2015, 54, 1164–1167 CrossRef CAS PubMed.
- B. C. Buer and E. N. G. Marsh, Fluorine: a new element in protein design, Protein Sci., 2012, 21, 453–462 CrossRef CAS PubMed.
- S. Huhmann, E. K. Nyakatura, H. Erdbrink, U. I. M. Gerling, C. Czekelius and B. Koksch, Effects of single substitutions with hexafluoroleucine and trifluorovaline on the hydrophobic core formation of a heterodimeric coiled coil, J. Fluorine Chem., 2015, 175, 32–35 CrossRef CAS.
- A. V. Marenich, C. P. Kelly, J. D. Thompson, G. D. Hawkins, C. C. Chambers, D. J. Giesen, P. Winget, C. J. Cramer and D. G. Truhlar, Minnesota solvation database – version, 2012 Search PubMed.
- V. H. Dalvi and P. J. Rossky, Molecular origins of fluorocarbon hydrophobicity, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 13603–13607 CrossRef CAS PubMed.
- J. R. Robalo, L. M. Streacker, D. Mendes de Oliveira, P. Imhof, D. Ben-Amotz and A. Vila Verde, Hydrophobic but water-friendly: favorable water–perfluoromethyl interactions promote hydration shell defects, J. Am. Chem. Soc., 2019, 141, 15856–15868 CrossRef CAS PubMed.
- V. Conti Nibali, S. Pezzotti, F. Sebastiani, D. R. Galimberti, G. Schwaab, M. Heyden, M.-P. Gaigeot and M. Havenith, Wrapping up hydrophobic hydration: locality matters, J. Phys. Chem. Lett., 2020, 11, 4809–4816 CrossRef CAS PubMed.
- S. Pezzotti, W. Chen, F. Novelli, X. Yu, C. Hoberg and M. Havenith, Terahertz calorimetry spotlights the role of water in biological processes, Nat. Rev. Chem., 2025, 9, 481–494 CrossRef CAS PubMed.
- S. Pezzotti, F. Sebastiani, E. P. van Dam, S. Ramos, V. Conti Nibali, G. Schwaab and M. Havenith, Spectroscopic fingerprints of cavity formation and solute insertion as a measure of hydration entropic loss and enthalpic gain, Angew. Chem., Int. Ed., 2022, 61, e202203893 CrossRef CAS PubMed.
- B. B. Majumdar, P. Pyne, R. K. Mitra and D. Das Mahanta, Impact of hydrophobicity on local solvation structures and its connection with the global solubilization thermodynamics of amphiphilic molecules, Phys. Chem. Chem. Phys., 2023, 25, 27161–27169 RSC.
- A. Cooney and K. W. Morcom, Thermodynamic behaviour of mixtures containing fluoroalcohols I. (water + 2,2,2-trifluoroethanol), J. Chem. Thermodyn., 1988, 20, 735–741 CrossRef CAS.
- J. A. Boyne and A. G. Williamson, Enthalpies of mixture of ethanol and water at 25 °C, J. Chem. Eng. Data, 1967, 12, 318 CrossRef CAS.
- J. Larkin, Thermodynamic properties of aqueous nonelectrolyte mixtures I. Excess enthalpy for water + ethanol at 298.15 to 383.15 K, J. Chem. Thermodyn., 1975, 7, 137–148 CrossRef CAS.
- M. Thomas, M. Brehm, R. Fligg, P. Vöhringer and B. Kirchner, Computing vibrational spectra from ab initio molecular dynamics, Phys. Chem. Chem. Phys., 2013, 15, 6608–6622 RSC.
- K. Forster-Tonigold and A. Groß, Dispersion corrected RPBE studies of liquid water, J. Chem. Phys., 2014, 141, 064501 CrossRef PubMed.
- M. Brehm, M. Thomas, S. Gehrke and B. Kirchner, TRAVIS—A free analyzer for trajectories from molecular simulation, J. Chem. Phys., 2020, 152, 164105 CrossRef CAS PubMed.
- R. Ramírez, T. López-Ciudad, P. Kumar and D. Marx, Quantum corrections to classical timecorrelation functions: hydrogen bonding and anharmonic floppy modes, J. Chem. Phys., 2004, 121, 3973–3983 CrossRef PubMed.
- H. W. Horn, W. C. Swope, J. W. Pitera, J. D. Madura, T. J. Dick, G. L. Hura and T. Head-Gordon, Development of an improved four-site water model for biomolecular simulations: TIP4P-Ew, J. Chem. Phys., 2004, 120, 9665 CrossRef CAS PubMed.
- H. Berendsen, D. van der Spoel and R. van Drunen, GROMACS: a message-passing parallel molecular dynamics implementation, Comput. Phys. Commun., 1995, 91, 43–56 CrossRef CAS.
- M. M. Ghahremanpour, J. Tirado-Rives and W. L. Jorgensen, Refinement of the optimized potentials for liquid simulations force field for thermodynamics and dynamics of liquid alkanes, J. Phys. Chem. B, 2022, 126, 5896–5907 CrossRef CAS PubMed.
- U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, A smooth particle mesh Ewald method, J. Chem. Phys., 1995, 103, 8577–8593 CrossRef CAS.
- B. Hess, H. Bekker, H. J. C. Berendsen and J. G. E. M. Fraaije, LINCS: a linear constraint solver for molecular simulations, J. Comput. Chem., 1997, 18, 1463–1472 CrossRef CAS.
- A. Lemak and N. Balabaev, On the Berendsen thermostat, Mol. Simul., 1994, 13, 177–187 CrossRef CAS.
- M. Parrinello and A. Rahman, Polymorphic transitions in single crystals: a new molecular dynamics method, J. Appl. Phys., 1981, 52, 7182–7190 CrossRef CAS.
- W. Humphrey, A. Dalke and K. Schulten, VMD – Visual molecular dynamics, J. Mol. Graphics, 1996, 14, 33–38 CrossRef CAS PubMed.
- M. G. Sceats, M. Stavola and S. A. Rice, A zeroth order random network model of liquid water, J. Chem. Phys., 1979, 70, 3927–3938 CrossRef CAS.
- M. G. Sceats and S. A. Rice, The enthalpy and heat capacity of liquid water and the ice polymorphs from a random network model, J. Chem. Phys., 1980, 72, 3248–3259 CrossRef CAS.
- M. G. Sceats and S. A. Rice, The entropy of liquid water from the random network model, J. Chem. Phys., 1980, 72, 3260–3262 CrossRef CAS.
- R. F. Lama and B. C.-Y. Lu, Excess thermodynamic properties of aqueous alcohol solutions, J. Chem. Eng. Data, 1965, 10, 216–219 CrossRef CAS.
- B. González, N. Calvar, E. Gómez and Á. Domínguez, Density, Dynamic Viscosity, and Derived Properties of Binary Mixtures of Methanol or Ethanol with Water, Ethyl Acetate, and Methyl Acetate at T = (293.15, 298.15, and 303.15) K, J. Chem. Thermodyn., 2007, 39, 1578–1588 CrossRef.
- R. R. Pawar, S. S. Kale and A. S. Kale, Density, Excess Molar Volumes of Water-Ethanol Binary Mixtures at Various Temperatures, Int. J. Sci. Res. Sci. Technol., 2017, 3, 96–100 Search PubMed.
- T. Minamihonoki, H. Ogawa, H. Nomura and S. Murakami, Thermodynamic properties of binary mixtures of 2,2,2-trifluoroethanol with water or alkanols at T = 298.15 K, Thermochim. Acta, 2007, 459, 80–86 CrossRef CAS.
- F. Böhm, G. Schwaab and M. Havenith, Mapping hydration water around alcohol chains by THz calorimetry, Angew. Chem., Int. Ed., 2017, 56, 9981–9985 CrossRef PubMed.
- M. Perttilä, Vibrational spectra and normal coordinate analysis of 2,2,2-trichloroethanol and 2,2,2-trifluoroethanol, Spectrochim. Acta, Part A, 1979, 35, 585–592 CrossRef.
- V. Sharma, F. Böhm, G. Schwaab and M. Havenith, The low frequency motions of solvated Mn(II) and Ni(II) ions and their halide complexes, Phys. Chem. Chem. Phys., 2014, 16, 25101–25110 RSC.
- D. Decka, G. Schwaab and M. Havenith, A THz/FTIR fingerprint of the solvated proton: evidence for Eigen structure and Zundel dynamics, Phys. Chem. Chem. Phys., 2015, 17, 11898–11907 RSC.
- F. Böhm, V. Sharma, G. Schwaab and M. Havenith, The low frequency modes of solvated ions and ion pairs in aqueous electrolyte solutions: Iron(II) and Iron(III) Chloride, Phys. Chem. Chem. Phys., 2015, 17, 19582–19591 RSC.
- A. J. Barnes, H. E. Hallam and D. Jones, Vapour phase infrared studies of alcohols – I. Intramolecular interactions and self-association, Proc. R. Soc. London, Ser. A, 1973, 335, 97–111 CAS.
- M. Matsugami, R. Yamamoto, T. Kumai, M. Tanaka, T. Umecky and T. Takamuku, Hydrogen bonding in ethanol–water and trifluoroethanol–water mixtures studied by NMR and molecular dynamics simulation, J. Mol. Liq., 2016, 217, 3–11 CrossRef CAS.
- M. Heyden, J. Sun, S. Funkner, G. Mathias, H. Forbert, M. Havenith and D. Marx, Dissecting the THz spectrum of liquid water from first principles via correlations in time and space, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 12068–12073 CrossRef CAS PubMed.
- E. Casalini, A. M. Stingel, B. Moussa, M. Personeni, P. B. Petersen and A. Vila Verde, Hydrophilic–hydrophobic double layers around amphiphilic solutes in mixed solvents, JACS Au, 2025, 5, 2992–2999 CrossRef PubMed.
- J. R. Robalo and A. Vila Verde, Unexpected Trends in the Hydrophobicity of Fluorinated Amino Acids Reflect Competing Changes in Polarity and Conformation, Phys. Chem. Chem. Phys., 2019, 21, 2029–2038 RSC.
- C. H. Rochester and J. R. Symonds, Thermodynamic studies of fluoroalcohols. Part 1.—Vapour pressures and enthalpies of vaporization, J. Chem. Soc., Faraday Trans. 1, 1973, 69, 1267–1273 RSC.
- B. Madan and K. Sharp, Heat capacity changes accompanying hydrophobic and ionic solvation: a Monte Carlo and random network model study, J. Phys. Chem., 1996, 100, 7713–7721 CrossRef CAS.
- K. A. Sharp and B. Madan, Hydrophobic effect, water structure, and heat capacity changes, J. Phys. Chem. B, 1997, 101, 4343–4348 CrossRef CAS.
- A. Ben-Naim, Hydrophobic interaction and structural changes in the solvent, Biopolymers, 1975, 14, 1337–1355 CrossRef CAS.
- P. Luo and R. L. Baldwin, Mechanism of helix induction by trifluoroethanol: a framework for extrapolating the helix-forming properties of peptides from trifluoroethanol/water mixtures back to water, Biochemistry, 1997, 36, 8413–8421 CrossRef CAS PubMed.
- D. Roccatano, G. Colombo, M. Fioroni and A. E. Mark, Mechanism by which 2,2,2-trifluoroethanol/water mixtures stabilize secondarystructure formation in peptides: a molecular dynamics study, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 12179–12184 CrossRef CAS PubMed.
- A. F. Pereira, V. Piccoli and L. Martínez, Trifluoroethanol direct interactions with protein backbones destabilize-helices, J. Mol. Liq., 2022, 365, 120209 CrossRef CAS.
- J. T. Gerig, Investigation of ethanol–peptide and water–peptide interactions through intermolecular nuclear Overhauser effects and molecular dynamics simulations, J. Phys. Chem. B, 2013, 117, 4880–4892 CrossRef CAS PubMed.
- M. Kinoshita, Y. Okamoto and F. Hirata, Peptide conformations in alcohol and water: analyses by the Reference Interaction Site Model theory, J. Am. Chem. Soc., 2000, 122, 2773–2779 CrossRef CAS.
Footnote |
| † These authors have equally contributed to the manuscript. |
| This journal is © the Owner Societies 2026 |