 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Using light polarisation to unravel the structure of insulin fibril polymorphs
Bethany Caldwella,
Xiaoqi Yea,
Alana Maerivoet b,
Jillian Madine
b,
Jillian Madine b and
Heike Arnolds
b and
Heike Arnolds *a
*a
aDepartment of Chemistry, University of Liverpool, Liverpool, UK, Heike. E-mail: Arnolds@liverpool.ac.uk
bInstitute of Systems, Molecular and Integrative Biology, University of Liverpool, Liverpool, UK
First published on 6th May 2026
Abstract
Amyloid fibrils lie at the core of major human degenerative diseases, such as Alzheimer's, Parkinson's, and diabetes. The structural polymorphism of amyloid fibrils has been linked to disease severity and many polymorph structures of proteins such as Aβ or α-synuclein are now known in detail. Amyloid fibril polymorphism has also been observed for insulin amyloid fibrils, but very little is known about the underlying structural differences. Here, we investigate the structure of insulin amyloid fibril polymorphs created in different solvents by Raman, infrared and fluorescence spectroscopy. We show that insulin polymorphs have different twists of their β-sheets which impacts alignment of backbone carbonyls relative to the fibril axis, changes β-turn structure and disulfide bond conformation, changes alignment of tyrosines and the hydrogen-bonding state of Gln and Glu side chains. These different structures arise because solvent composition changes hydrophobic and hydrogen-bonding interactions between insulin monomers. This work is a first step towards understanding how different environmental conditions create specific insulin fibril structures.
Introduction
Amyloid fibrils were first recognised for their common cross-β structure which is adopted by widely different proteins1 ranging from natively unfolded proteins such as Amyloid-β to globular proteins such as insulin.2 In the cross-β structure, each protein monomer is arranged in a plane and planes stack via backbone hydrogen bonds to form long fibrils. This arrangement on one hand leads to a common fibril backbone structure, but on the other hand allows for a large variety of monomer conformations within each plane, leading to amyloid fibril polymorphism.3 Beyond the backbone hydrogen bonds between monomers, amyloid fibrils are stabilised by ladders of uncharged identical side chains and by exclusion of water when complementary ladders form a steric zipper.4 Structural frustrations in these structures such as steric clashes, charge repulsion and hydrophobicity and polarity of side chains can be relieved by variations in the planar monomer fold which leads to different polymorphs.3 These structural frustrations make amyloid fibril structures sensitive to environmental factors such as solvent composition, pH, ionic strength or presence of ligands. The ensuing wide range of amyloid fibril polymorphs has been linked to disease severity3,5,6 but is also a useful property which can help to create protein structures with defined function.7In order to gain a mechanistic understanding of the link between polymorph structure and amyloid toxicity, the structures of many of the polymorphs of proteins involved in neurodegenerative diseases have been determined by crystallography,8 solid-state nuclear magnetic resonance (NMR)9 or cryo-electron microscopy (cryo-EM).10 For example, for α-synuclein, more than 140 amyloid fibril structures have so far been deposited in the Protein Data Bank.11 This extensive structural knowledge of amyloid fibrils helps to understand the role of mutations in disease and the design of therapeutic interventions.12
Insulin is a small globular hormone with native α-helical structure which regulates glucose metabolism and is used in the treatment of diabetes. Insulin amyloidosis can occur when patients repeatedly inject into the same site.13,14 Insulin is also a frequently used model protein to understand amyloid fibril formation15–17 and many studies have investigated the influence of environmental factors specifically on insulin amyloid polymorphism. The existence of different insulin fibril polymorphs has been confirmed by imaging,18,19 vibrational dichroism,20,21 amide I band shape19 and spectral properties of various external probes.22–24 It has been shown multiple times that different solvent properties such as hydrogen bonding ability, dielectric constant, specific binding of solvent25 or salt molecules26 and protonation state of amino acid side chains20 lead to different insulin fibril polymorphs with clear spectral or morphological differences. Similar to other amyloid diseases, it has been confirmed that insulin polymorph structure changes the cytotoxicity of insulin fibrils.27–30 Unlike other amyloids though, very little is known about the polymorphic landscape of insulin fibrils since only two structures of insulin amyloid fibrils have been determined with atomic level detail using solid-state NMR31 and cryo-EM.32
Here we use Raman and polarized Raman spectroscopy, together with infrared and fluorescence spectroscopy to determine structural differences between insulin fibrils formed in three different solvents at acidic pH. We chose acetic acid and dilute HCl as used in the earliest reports of morphological and spectral differences in insulin fibrils26,33,34 as well as in the recent solid-state NMR study of insulin fibrils.31 These two acids create solutions of insulin monomers and dimers respectively.35 It has been hypothesised that insulin monomers and dimers form structurally different seeds which can grow into different fibril structures.18 We also used acetonitrile as a cosolvent with dilute HCl because this creates a monomeric insulin solution36 without specific binding between solvent molecules and insulin as observed for acetic acid.25 Acetonitrile is a common additive in reverse-phase chromatography of insulin and other proteins37 but has not been used as a cosolvent in insulin aggregation to the best of our knowledge.
Raman spectroscopy, particularly coupled with polarized Raman spectra of aligned protein fibrils,38–40 is emerging as a middle ground between simple diagnostic tools such as the response of amyloid-specific dyes or the appearance of β-sheet peaks in the amide I band in infrared spectroscopy41 and much more demanding techniques such as cryo-EM or solid-state NMR. Raman spectroscopy has two specific advantages over IR spectroscopy in a study of fibril polymorphs: firstly the amide III band is more prominent and its frequency links directly to the β-sheet conformation;39 secondly polarization measurements are straightforward and allow to determine the alignment of several fibril structural features relative to each other.42
Here we show that insulin fibril polymorphs essentially differ in the twist of their β-sheets. This altered Ramachandran angle ψ of the β-sheet changes the alignment of backbone carbonyls relative to the fibril axis, the β-turn structure of the fibril, disulfide bridge conformation and orientation of some side chains. We support these conclusions drawn from Raman spectroscopy by both infrared and intrinsic fluorescence spectroscopy. We attribute the structural changes to the presence of solvent components such as acetonitrile or acetic acid, which modify insulin-water and insulin-insulin interactions.
Experimental
Materials
Human insulin was purchased from Biogems; glacial acetic acid from Amresco; hydrochloric acid S.G. 1.18 (∼37%) from Fisher Scientific; acetonitrile from Alfa Aesar; ThT from Sigma Aldrich; protein solutions are filtered with 0.22 µm PTFE hydrophilic filters from Avantor and aggregated in 2 mL LoBind tubes from Eppendorf.Insulin sample preparation
Human insulin (50 mg) was weighed into a 5 mL volumetric flask and dissolved in 5 mL of either 20% acetic acid (v/v) (AA), 0.02 M hydrochloric acid (HCl), or 0.02 M HCl with 30% acetonitrile (ACN) yielding a final protein concentration of 10 mg mL−1. Insulin solutions were placed in a water bath at 60 °C without agitation and left undisturbed for 9 days. These fibrils were used as seeds to enhance the structural homogeneity of insulin fibrils.43 Seeds were added to native insulin solutions at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 (v/v) ratio prior to aggregation. The seeded mixtures were then subjected to the same quiescent aggregation as the parent sample. We chose the same protein concentration, solvents, incubation temperature and use of seeding as in the NMR study of insulin fibrils to ensure that we obtain fibrils with a similar structure.31 We iteratively repeated seeding for three generations, yielding one unseeded parent generation and three subsequent seeded generations. The final generation fibrils showed the highest degree of alignment and were studied in detail for this work. For these fibrils, centrifugation of samples and measurement of supernatant absorbance at 277 nm showed that 53% of insulin converted into fibrils in acetic acid, 72% in ACN and 88% in HCl.
20 (v/v) ratio prior to aggregation. The seeded mixtures were then subjected to the same quiescent aggregation as the parent sample. We chose the same protein concentration, solvents, incubation temperature and use of seeding as in the NMR study of insulin fibrils to ensure that we obtain fibrils with a similar structure.31 We iteratively repeated seeding for three generations, yielding one unseeded parent generation and three subsequent seeded generations. The final generation fibrils showed the highest degree of alignment and were studied in detail for this work. For these fibrils, centrifugation of samples and measurement of supernatant absorbance at 277 nm showed that 53% of insulin converted into fibrils in acetic acid, 72% in ACN and 88% in HCl.
Fibril alignment
After aggregation, 2 mL aliquots of each sample were transferred into Eppendorf tubes and centrifuged at room temperature at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 g for 30 minutes. The supernatant was discarded, and the pellets were resuspended in their respective aggregation solutions. This centrifugation and washing process was repeated three times to remove unaggregated insulin. The washed fibril samples were then sonicated in an ultrasonic bath (GT SONIC-D Series Digital Ultrasonic Cleaner) at room temperature for 5 minutes to promote fibril shortening and alignment. Fibrils were diluted in the respective aggregation solutions 1
000 g for 30 minutes. The supernatant was discarded, and the pellets were resuspended in their respective aggregation solutions. This centrifugation and washing process was repeated three times to remove unaggregated insulin. The washed fibril samples were then sonicated in an ultrasonic bath (GT SONIC-D Series Digital Ultrasonic Cleaner) at room temperature for 5 minutes to promote fibril shortening and alignment. Fibrils were diluted in the respective aggregation solutions 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 (v/v) and aligned by drop coating deposition Raman.44 In this approach, 20 µL of the fibril solution was deposited onto the surface of a polished Si(111) wafer and allowed to air-dry. As the solvent evaporated this caused a net flow outward from the centre, which carried the fibrils towards the perimeter of the droplet, leading to a self-organisation along the edge, forming a characteristic coffee-ring pattern. Fibrils align with the fibril axis directed tangentially along the droplet perimeter, which is shown by varying the relative orientation between incident laser polarisation and rim direction (Fig. S1). A higher degree of fibril alignment creates a higher birefringence, which is detectable as a brighter area in a cross-polarised reflected light image (Fig. S2).
2 (v/v) and aligned by drop coating deposition Raman.44 In this approach, 20 µL of the fibril solution was deposited onto the surface of a polished Si(111) wafer and allowed to air-dry. As the solvent evaporated this caused a net flow outward from the centre, which carried the fibrils towards the perimeter of the droplet, leading to a self-organisation along the edge, forming a characteristic coffee-ring pattern. Fibrils align with the fibril axis directed tangentially along the droplet perimeter, which is shown by varying the relative orientation between incident laser polarisation and rim direction (Fig. S1). A higher degree of fibril alignment creates a higher birefringence, which is detectable as a brighter area in a cross-polarised reflected light image (Fig. S2).
Fluorescence
All fluorescence measurements were recorded on a Horiba Fluoromax-4 fluorimeter. The excitation slit width was set to 5 nm, while the emission slit width was narrowed to 1 nm for increased resolution, integration time of 0.1 s per point.For intrinsic tyrosine fluorescence, all samples were excited at 280 nm. Native samples in their original solvents at 10 mg mL−1 concentration were measured in a quartz cuvette with excitation along a 2 mm pathlength (absorbance ca. 2) and emission along 10 mm pathlength. The inner filter effect was corrected using absorbance spectra recorded in the same cuvette.45 Intrinsic tyrosine fluorescence of fibril polymorphs was recorded for washed fibrils resuspended in 0.02 M HCl at a concentration of 0.75 mg mL−1 in a quartz cuvette with excitation along the 4 mm pathlength (absorbance ca. 0.3) and emission along the 10 mm pathlength.
For ThT fluorescence, 1 mL of ThT stock solution (8 mg ThT dissolved in 0.02 M HCl) was diluted to 50 mL with 0.02 M HCl. A 3 mL aliquot was transferred into a cuvette for fluorescence measurement. The emission spectra were recorded following excitation at 450 nm over a range of 470 to 700 nm. Subsequently, 100 µL of washed fibrils resuspended in 0.02 M HCl at a concentration of 0.75 mg mL−1 were added, mixed gently, and left to incubate for 2 minutes before a second fluorescence measurement was taken.
Fibril samples used for intrinsic and ThT fluorescence had concentrations that varied by less than 10% (as verified by absorbance at 277 nm) and ThT fluorescence peak heights were therefore not corrected relative to each other.
Transmission electron microscopy (TEM)
TEM images were taken using a FEI 120kV Tecnai G2 Spirit BioTWIN transition electron microscope with a SIS Megaview III camera, to understand the morphologies of insulin fibrils formed under various conditions. The fibril samples were first diluted to 1 mg mL−1 to reduce fibril clustering before being sonicated in an ultrasonic bath (GT SONIC-D Series Digital Ultrasonic Cleaner) at room temperature for 5 minutes to further break down any long fibrils. 5 µL of the fibril solution was then pipetted onto carbon-coated copper grids, blotted and then stained with 2% uranyl acetate for 30 seconds for image contrasting. Fibril widths were analysed using ImageJ.Fourier-transform infrared spectroscopy (FT-IR)
Following aggregation, 20 µL of each insulin fibril sample was drop coated onto PTFE tape and left to dry in a desiccator overnight. Dried samples were then analysed using a Bruker Vertex 70 FT-IR spectrometer equipped with a diamond attenuated total reflection accessory.Raman and polarized Raman spectroscopy
The Renishaw in Via Reflex Raman microscope was employed with a 532 nm laser, 10 s acquisition time, 10% laser power and 10 accumulations. The laser was focussed using a 50× objective from Olympus (NA = 0.5). Spectra shown in Fig. S2 were acquired with a 785 nm laser and 50% laser power and the same acquisition time, accumulations and objective.Small angle X-ray scattering (SAXS)
SAXS measurements were performed in a Nanostar SAXS instrument (Bruker) equipped with a Metal Jet source (Gallium anode). The generated beam is collimated by a three-pinhole setup which mitigates parasitic background and provides a nearly circular high flux beam with approximate diameter of 550 µm at the detector position. The 2D scattering patterns were collected for 1800 seconds by a charge-coupled device (CCD) detector (VÅNTEC 2000) at sample-to-detector distance (SDD) of 107 cm (silver behenate, AgBh, was used as a standard for calibration). This provides the modulus of the transfer moment vector, q = 4sin(θ)λ−1, where 2θ is the scattering angle and the X-ray wavelength λ is 1.3420 Å, q is experimentally accessible from 0.025 to 0.25 Å−1. Glassy carbon was placed between the sample and detector and the total scattering was recorded for 300 s, allowing for the correction of the sample's transmission. Measurements were performed at room temperature and in vacuum, the data was spatially corrected. Azimuthal averaging and background correction were performed in the data reduction step using the software Diffrac.SAXS. Normalization to absolute scale was done using water measured in the exact same conditions as the samples and solvents. For this purpose, reusable quartz capillaries mounted in stainless steel cases were used in all measurements. Concentrations of solutions were determined from the absorbance at 277 nm measured in a 2 mm quartz cuvette (9.1 mg mL−1 for AA, 9.4 mg mL−1 for HCl, 9.8 mg mL−1 for ACN) and then used to scale the scattering intensities for AA and HCl to the same concentrations as ACN.Results and discussion
Native insulin structure
Native insulin consists of two peptide chains linked by two disulfide bonds with a third disulfide bond stabilising a turn in the A chain between two short helices. The central section of the B chain forms a longer helix while the C-terminal section can form an antiparallel β-sheet with a second insulin monomer, as shown in Fig. 1A. | ||
| Fig. 1 Properties of native 10 mg mL−1 insulin solutions in the various solvents used in this study. (A) Insulin dimer structure at pH 2 from pdb 1GUJ.26 The A chains of the dimer are shown in blue and grey, the B chains of the dimer are shown in red and orange. (B) Overlay of SAXS scattering curves. (C) Tyrosine intrinsic fluorescence spectra. Peak heights have been corrected for the inner filter effect. (D) IR spectra of dried native insulin solutions. | ||
For human insulin, 20% acetic acid and 35% ACN at pH 3.6 create monomeric insulin solutions,36,46 while dissolution in dilute HCl at higher insulin concentrations creates dimers.47 However, size measurements of salt-free human insulin solutions containing some Zn2+ ions, at the high concentration of 10 mg mL−1 under our specific solvent conditions are not available and we therefore used small-angle X-ray scattering to determine the oligomer state, with the scattering curves shown in Fig. 1B. Human insulin has the same form factor in 30% ACN+ 0.02 M HCl and in 20% acetic acid, compatible with insulin monomers, while a 0.02 M HCl solution leads to a distinct increase in size, compatible with dimer formation (theoretical SAXS curves for insulin monomer and dimer structures are shown in Fig. S3). At these high concentrations, the slope of I(q) at low scattering vectors q is negative for insulin in dilute HCl and positive for insulin in ACN and acetic acid. The former finding indicates long-range attractive interactions for human insulin in HCl, the latter repulsive interactions for human insulin in ACN or acetic acid in agreement with observations made for bovine insulin.
The insulin dimer interface consists of both hydrophobic residues (Val B12, Phe B24 and B25, Leu A13 and B17) and hydrophilic residues with the ability to form hydrogen bonds (Ser B9, Glu B13, Tyr B16 and B26 and Thr B27). Acetic acid is known to solvate hydrophobic residues well48 and can in addition form hydrogen bonds with side chains. Acetonitrile is a common solvent used in chromatography to reduce hydrophobic interactions between proteins and silica.37 Both solvents therefore have the ability to disrupt interactions across the dimer interface and keep insulin monomeric which is reflected in repulsive interactions in SAXS. However, despite very similar SAXS form factors, there are differences in insulin structure in solution. Intrinsic fluorescence of tyrosine in insulin is known to quench upon increased solvent exposure.49 As shown in Fig. 1C, Tyr fluorescence of native insulin solutions increases in the presence of ACN and decreases in the presence of acetic acid relative to HCl. This means ACN reduces Tyr exposure to solvent compared to dilute HCl, while acetic acid increases Tyr exposure to solvent. Native human insulin in ACN studied by NMR was found to adopt a very compact monomer structure, similar to the monomers making up the insulin T6 hexamer36 in keeping with our observation of reduced Tyr exposure to solvent in ACN. Earlier work on bovine insulin found that acetic acid creates a more solvent-permeable insulin structure, perhaps because of its ability to hydrogen bond to insulin through its carboxylic acid groups.25 Fig. 1D shows IR spectra of native insulin in the three solvents. HCl and ACN dissolved insulin shows a clear symmetric COOH stretch at 1730 cm−1 from insulin's 4 Glu side chains which are fully protonated at pH values of 2 and below. Insulin dissolved in acetic acid however does not show this peak, instead it shows higher absorbance at 1570 cm−1 where we normally find the Glu COO− symmetric stretching peak.50 Alongside an additional peak around 1400 cm−1 (the acetate –CH3 deformation) and an enhanced absorbance around 1540 cm−1 (C–O stretch)51 this indicates that acetate ions hydrogen-bond to insulin via the Glu carboxylic acid groups.
The increased solvent permeability probably explains why insulin in acetic acid forms fibrils much earlier than in HCl, while addition of ACN results in the slowest aggregation.
Fibril morphology
TEM images of stained insulin fibrils prepared in 0.02 M HCl, 0.02 M HCl with 30% ACN and in 20% acetic acid are shown in Fig. 2. We will for ease of reading refer to these in the following as HCl, ACN and AA fibrils. Fibrils are several microns long. The uranyl acetate staining generally leads to a flat appearance of imaged fibrils regardless of solvent, only in some cases is a twist clearly discernible (see discussion in next section). The ratio of fibril length to end-to-end distance is 1.02 ± 0.02 for all three fibril types, indicating similar flexibility regardless of solvent. We used representative images of fibrils acquired at different magnifications and measured fibril widths at ca. 1000 positions for each fibril type (Fig. S4–S13). Within error bars, all three fibril types have a width of ca. 10 nm with a standard deviation of around 3 nm. However, it can be seen from the histograms in Fig. 2D that the width distribution for HCl fibrils is broader and has its maximum around 7 nm, while ACN and AA fibrils width distributions have a maximum around 10 nm.The most recent solid-state NMR study of the structure of insulin fibrils, prepared in both dilute HCl and in 20% acetic acid at the same concentration and aggregation temperature as used here,31 found that the two insulin chains rearrange into a U-shape which constitutes a single protofilament with a cross section of 5 × 2.5 nm. The hydrophobic side chains on the exterior of this U-shape are then buried by formation of a tetrameric bundle of protofilaments with a cross section of approximately 10 × 5 nm. A tetramer however is not the only possible quaternary assembly, as shown by molecular modelling of insulin fibrils.52 The different width distribution of HCl fibrils could therefore be caused by units smaller than a tetramer.
Previous atomic force microscopy on the dependence of insulin fibril morphology on solvent conditions reported lower cross-sectional heights for insulin fibrils grown in 25 mM HCl or a pH 2 phosphate buffer, compared to insulin fibrils grown in 20% acetic acid, with the latter widths increasing with NaCl concentration.19,22 Insulin fibril widths are also known to be pH dependent, with an increase for pH values above 2.4 and a reversal of supramolecular chirality.20,22 The change in fibril widths has previously been assigned to self-assembly of a variable number of protofilaments.53 A recent AFM study of insulin fibril morphology highlighted the complexity of fibril assembly by demonstrating the existence of both twisted ribbons and helical fibrils of different widths as well as intertwining fibrils.54
Morphological data however cannot tell us whether there are structural differences at secondary or tertiary structure level or whether we have the same amyloid fold but a different quaternary structure assembly. We will next look at secondary and tertiary structural differences between fibrils as seen by vibrational spectroscopy.
β-Sheet dihedral angles
The secondary structure of proteins can be determined from amide I and amide III regions in Raman spectroscopy, shown for the three fibril types in Fig. 3A. We will compare these and other experimental results in the following to the insulin fibril model derived from NMR data (pdb 8RVT31). We have included a ribbon diagram of the lowest energy NMR structure in Fig. 3B. Each insulin monomer rearranges into a U-shaped conformation with two sections of β-strand connected by a β-turn. A and B chains alternately stack to form the protofibril and the structure is stabilised by the intrachain disulfide bond A6–A11, while A and B chains remain connected by two interchain disulfide bonds A7–B7 and A20–B19. The C-terminal region of the B-chain (residues 21–30) remains unstructured and is not observable in solid-state NMR for both HCl and AA fibrils. In addition, AA fibrils possess an unstructured N-terminal region of the B chain (residues 1–5) which is attributed to repulsion between protonated His B5 sidechains. | ||
| Fig. 3 (A) Raman spectra of third generation fibril polymorphs grown from 0.02 M HCl, 0.02 M HCl + 30% ACN (“ACN”) and 20% acetic acid (“AA”). Spectra have been normalised to the height of the Phe peak at 1005 cm−1. (B) Ribbon diagram of insulin fibrils from pdb 8RVT model 1, disulfide bonds are shown in yellow, A chain in blue, B chain in red, residues B21–B30 are not included in the model.31 | ||
The amide I band of the amyloid β-sheet in Raman is an in-phase coupled vibration of backbone carbonyls and is typically found between 1670 and 1675 cm−1. As Fig. 3A shows, all three polymorphs have a band at ca. 1672.5 cm−1 and only differ by 0.3 cm−1 in peak positions. The position of this band is determined by coupling between carbonyls across different β-strands therefore different polymorphs must have very similar stacking distances in the fibril direction as expected from X-ray diffraction and other structural studies.5 The amide I Raman band has smaller contributions from the out-of-phase β-sheet vibration at ca. 1628 cm−1 and the β-turn at ca. 1660 cm−1, but wavenumber differences between fibrils for these structural elements are again quite small. Spectral differences between polymorphs are best seen in the amide III region which arises from a coupled N-H bending and C–N stretching vibration of the peptide backbone. Within this region, the amide III3 band between 1200 and 1300 cm−1 depends on secondary structure because coupling between the N–H and Cα–H bending motions leads to a dependence on Ramachandran angle ψ.55 This structure dependence has been used to determine ψ distributions for amyloid fibrils from Raman spectroscopy.38 The peak around 1220 cm−1 has been assigned to β-sheet structures; In Fig. 3, HCl fibrils have this peak at 1223.5 cm−1, while ACN and AA fibrils have the peak at 1226.6 and 1227.3 cm−1 respectively. The average β-sheet frequencies recorded in 4 different spots on each sample are 1223.3 ± 0.3 cm−1 for HCl, 1226.6 ± 0.3 cm−1 for ACN and 1227.5 ± 0.5 cm−1for AA fibrils.
Mikhonin et al. derived the following relationship between amide III3 peak frequency and Ramachandran angle ψ for unknown hydrogen bonding status of the backbone:55
| ṽ = 1250 cm−1–54 cm−1 sin(ψ + 26°) | (1) |
This results in an average β-sheet Ramachandran angle ψ of 125° for HCl fibrils, 128° for ACN and 129° for AA fibrils. Different ψ angles arise from different twists of the protofilament stack. The insulin fibril structure pdb 8RVT contains 10 different models.31 The two lowest energy models 1 and 2 have average β-sheet ψ angles of 126.8° and 131.8°, which compares well to the amide III3 deduced angles. The β-sheet twist angles derived56 from residues 12 and 18 in the A and B chain are 0.23° and 1.77° which translates to left-handed fibril crossover distances of 740 and 100 nm for models 1 and 2, respectively. We only found two fibrils in the TEM images which showed a periodic variation in staining (Fig. S14). The HCl fibril has an average cross-over distance of 100 ± 10 nm (from six cross-overs) and the AA fibril has an average crossover distance of 66 ± 3 nm (from 8 cross-overs). ACN fibril images had almost no discernible cross-overs and only a single instance of two consecutive cross-overs at a distance of 110 nm. The lower β-sheet ψ angle of HCl fibrils is expected to lead to a longer cross-over distance than for AA or ACN fibrils. This relationship enables protofilaments to twist around one another always interacting via the same region. An early cryo-EM study of insulin fibrils measured a 20–30% variability in crossover distance.53 The variability relates to a subtle change in the underlying β-sheet structure which changes the β-sheet twist. An analysis of this phenomenon is demanding in TEM but easily captured in the amide III3 peak widths, which are 21 cm−1 FWHM for HCl and ACN fibrils and 24 cm−1 FWHM for AA fibrils. These widths translate to ψ ranges of ca. ± 15° which encompasses the flat ribbons we mostly observe as well as 100 nm cross-over distances.
Each fibril type possesses further peaks in the range between 1240 and 1270 cm−1. Analogous to the β-sheet Ramachandran angle ψ distribution, ACN and AA fibrils result in spectra that are broadly similar and distinct from HCl fibril spectra. Due to the sinusoidal relationship between wavenumber and ψ, both α-helices and β-turns can have amide III3 peaks in this wavenumber range. Peak fitting of this region shows that HCl fibrils possess a single peak at 1252 cm−1, while ACN and AA fibrils are best described by double peaks at 1249/1265 cm−1 for ACN and at 1248/1261 cm−1 for AA. All fibrils possess a broad peak around 1280 to 1290 cm−1, which is typical for disordered regions and likely arises from the C-terminal tail of the insulin B-chain, which could not be resolved in solid-state NMR. Given the clear β-sheet amide I peak at 1672.5 cm−1, assignment of the additional amide III3 peaks to α-helices makes no sense. Our prior work on insulin fibrils showed that the β-turn of the fibrils contributes significantly to the infrared spectra, as it consists of 12 out of 51 residues.57 We therefore suggest that the additional peaks arise from differently structured β-turns. We used eqn (1) above to calculate Raman amide III3 spectra from NMR models 1 and 2, included in the supporting information, Fig. S15, which illustrate that the assignment of the additional peaks to different β-turn structures is a reasonable assumption.
Disulfide bond conformation
As Fig. 4 shows, characteristic peaks for S–S stretching vibrations are found between about 500 and 550 cm−1, while C–S stretching peaks are found between ca. 650 and 780 cm−1. The small shoulder at 495 cm−1 is too low for a typical S–S stretching vibration and this region has been assigned to skeletal bending modes previously.58 The two remaining peaks occur around 515 and 530 cm−1 for insulin fibrils, in agreement with prior literature.59,60 The peak near 515 cm−1 belongs to the two interchain disulfide bonds A7–B7 and A20–B19. It changes in height together with the amide I mode in polarised Raman spectra42 (also see SI Fig. S16), which means that the interchain disulfide bonds are oriented parallel to the fibril axis as seen in the NMR structure 8RVT.31 This disulfide bond orientation precludes the alternative insulin fibril structure determined from cryo-EM data (8SBD32). In those fibrils, A and B chains occupy a single plane and then those planes stack to form the fibril, which places the interchain disulfide bonds at a right angle to the fibril axis. This alternative insulin structure could have been created by the 10× lower insulin concentration used.42The peak at ca. 535 cm−1 belongs to the intrachain disulfide bond. It is noticeable that ACN and AA fibrils have a much higher intensity S–S peak than HCl fibrils relative to phenylalanine. The drop in intensity might indicate an n → π* electron donation from sulfur s and p orbitals to backbone carbonyls which could reduce the polarizability of the S–S bond and which is particularly effective for certain dihedral angles.61 Differences in disulfide peak height have been noticed as a distinguishing feature of insulin polymorphs previously.60
The S–S peak frequency relates to the dihedral angles of the CSSC group62 and whether the CCSSCC group is arranged as a staple, hook or a spiral.63 HCl has the lowest frequencies with 512.6 and 531.6 cm−1, AA has 513.5 and 533.3 cm−1, ACN has 514.6 and 534.2 cm−1. The conformations around the C–S bonds are also different, because the C–S stretching frequency depends on which atom is in trans position to sulfur across the adjacent C–C bond,64 although there is substantial spectral overlap between different conformers.65 All polymorphs possess a similarly broad band around 680 cm−1, which is typical for a PH conformer. ACN and AA fibrils in addition have a broad band centred around 740 cm−1 which encompasses both PN and PC conformers, whereas HCl fibrils have two distinct peaks in this region. Extracting the dihedral angles of the NCCS group from pdb 8RVT, we find that model 1 with a low fibril twist has 4 out of the 6 NCCS groups in PC conformation and two in PH conformation, while model 2 with a higher twist has 2 NCCS groups in PC conformation and 4 in PH conformation. To accommodate the twist, the conformation of both the intrachain A–A11 disulfide bond and the A20–B19 disulfide bond changes. The change in disulfide spectra is therefore consistent with the change in β-sheet twist angles observed.
Fibril alignment from polarized Raman spectra
We show polarized Raman spectra of the three fibril polymorphs acquired in 4 polarisation combinations in Fig. 5. Fibrils align tangentially along the rim of a dried drop44 (Fig. S1). Laser polarization parallel to the fibril axis is designated as Z, polarization perpendicular to the fibril axis as X, the first letter denotes the incoming laser polarization direction, the second letter the polarization direction of Raman scattered light. The good overlap between cross-polarized spectra shows that there is no laser-induced damage or artefact created by the polarization optics. The degree of alignment can be quantified with the help of the pseudo-order factor:
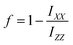 | (2) |
Similarly, both the amide III3 β-sheet and β-turn bands show large differences between fibril types, which qualitatively indicates a different alignment of these structural features. The large discrepancy between amide III3 and amide I β-sheet pseudo-order factors is puzzling at first since the principal axes of amide III and amide I β-sheet tensors are thought to be the same.66 A possible explanation is a difference from nearly cylindrical symmetry of the amide I tensor to non-cylindrical symmetry for amide III tensor, i.e. we would expect these to be of the form shown below:
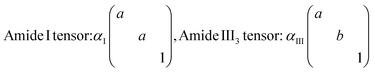 | (3) |
This would explain the high f values for the amide I mode compared to much lower values for the amide III3 mode.67
The derivation of the order parameters 〈P2〉 and 〈P4〉 for the amide I mode, the corresponding Lagrange multipliers and the most probable and preferred orientation distribution functions from polarized Raman spectra has been described in detail before.38,44 The relevant numerical values are found in Table S1 and the preferred orientation distribution functions are shown in Fig. 6.
 | ||
Fig. 6 (A) Preferred orientation distribution functions of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angle relative to the fibril axis. (B) Models 1 and 2 of 8RVT showing how the fibril twist influences the carbonyl directions. O angle relative to the fibril axis. (B) Models 1 and 2 of 8RVT showing how the fibril twist influences the carbonyl directions. | ||
As expected, the β-sheet carbonyls are mainly aligned along the fibril axis, however, there are distinct differences between fibrils. The most probable C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angle is lowest for the HCl fibril polymorph at 14° and increases to 15.6° for ACN fibrils and 18° for AA fibrils. This is similar to two earlier studies,42,44 which derived most probable angles of 21 respectively 12° for insulin fibrils prepared in HCl under similar aggregation conditions to ours. The difference of 4° between the most probable C
O angle is lowest for the HCl fibril polymorph at 14° and increases to 15.6° for ACN fibrils and 18° for AA fibrils. This is similar to two earlier studies,42,44 which derived most probable angles of 21 respectively 12° for insulin fibrils prepared in HCl under similar aggregation conditions to ours. The difference of 4° between the most probable C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angles of HCl and AA fibrils derived from the amide I mode alignment matches well to the β-sheet ψ angle difference of 4° derived from amide III3 frequencies. Both HCl and ACN fibrils possess a 90° peak in their preferred orientation distribution functions and this has been interpreted in other work as a fraction of fibrils without net orientation.38
O angles of HCl and AA fibrils derived from the amide I mode alignment matches well to the β-sheet ψ angle difference of 4° derived from amide III3 frequencies. Both HCl and ACN fibrils possess a 90° peak in their preferred orientation distribution functions and this has been interpreted in other work as a fraction of fibrils without net orientation.38
Changes in amino acid side chains
Fig. 7A shows IR spectra of fibril polymorphs with amide I, II and III regions. The IR amide III β-sheet peaks show the same pattern of lowest β-sheet frequency for HCl fibrils, while the close overlap between amide I and II peaks confirm the earlier statement from Raman spectroscopy that the coupling between carbonyls in different β-strands is very similar for different fibril types. Fig. 7B shows the same IR spectra after resolution enhancement in the amide II and amide I regions to illustrate changes in the narrower side chain peaks which overlap the amide group vibrations. Here we can see that some side chain bands change in absorbance or shift in frequency. Bands reducing in intensity for ACN fibrils are seen at 1660 cm−1 and 1715 cm−1. The former can be assigned to Gln side chains, while the latter belongs to hydrogen-bonded COOH groups found in both Glu side chains and C-termini at this low pH. A more detailed interpretation is hampered by the fact that there are 3 Gln and 4 Glu residues in different positions, some in the core of the U-shaped fibril model and some on the outside.Moreover, IR spectra show peak shifts/absorbance changes of some tyrosine modes for HCl fibrils relative to ACN and AA fibrils, which could be interpreted as an increased amount of tyrosine side chains which are either deprotonated or experience a strong hydrogen bond. This can be probed independently by intrinsic tyrosine fluorescence. For this measurement, we resuspended all fibrils into the same 0.02 M HCl solution to be able to detect the tyrosine environment without the influence of ACN or acetic acid. As Fig. 7C shows, the intrinsic tyrosine fluorescence is highest for HCl fibrils and lowest for acetic acid fibrils. In the common interpretation of this ratio, this means that tyrosines are least exposed to water for HCl fibrils and most exposed to water for AA fibrils.49 However, there is no emission near 350 nm, which would be expected if Tyr had deprotonated to form Tyr-O−. A better interpretation of the Tyr spectral changes seen in IR then is that some of the 4 Tyr sidechains in HCl fibrils are hydrogen bonding to other side chains, which would explain why the Tyr pseudo-order factor was highest for HCl fibrils. Again, a more detailed interpretation is not possible because the 4 tyrosines are found in different positions within the A and B chains.
Fig. 7D also includes results from the thioflavin T (ThT) standard fluorescence test for amyloid fibril formation.41 The degree of binding of the amyloid dye ThT is different for the different fibrils, as indicated by both different fluorescence intensities and peak wavelengths in Fig. 7D. Particularly the shift in peak position has been linked to the existence of different insulin polymorphs,22 however ThT fluorescence is only an indirect probe of insulin fibril polymorphism which does not explain the origin of structural differences.
Overall, the vibrational spectra show that the three solvents produce fibrils with very similar secondary structure (as deduced from amide I peak positions), but small differences in tertiary structure as deduced from changes in β-sheet carbonyl orientation distribution, disulfide and side chain spectra.
Influence of aggregation solvents
We finally consider the influence of the aggregation solvent. Previous studies have discussed whether the oligomer state of native insulin influences the fibril structure and fibril morphology,29,34,68 even though the broader assumption is that insulin oligomers first dissociate to form monomers which then unfold to form a partially-folded intermediate, before a nucleus or later protofilaments and fibrils can form.26,69,70 How then is it possible that insulin dimers (present in an HCl solution) lead to a different fibril structure than insulin monomers (present in acetic acid or acetonitrile solutions) if an unfolded monomer is the seed for fibril formation?A possible explanation for this is the difference in insulin hydration in the different solvents which has been speculated to influence its fibril self-assembly before.71 For example, the hydration shell of bovine insulin monomers in acetic acid has been found to be distinctly thinner than the hydration shell of bovine insulin dimers in hydrochloric acid.72 This could be related to the ability of acetic acid to hydrogen bond to insulin (and therefore to displace water molecules) as shown through our IR spectra for human insulin and in earlier work for bovine insulin.25 Co-solvents such as ethanol or the acetic acid and acetonitrile used here also change the dielectric constant and hydrogen bonding ability of the solvent, which is known to increase repulsive interactions between bovine insulin molecules and create monomeric insulin solutions.25,71,73,74 ACN has not been used as a solvent in insulin aggregation to the best of our knowledge, but its ability to solvate amino acids as well as the protein lysozyme has been studied75 and it has been used to modify peptide self-assembly.76,77 These studies have confirmed that peptide hydrogen bonding is strengthened in the presence of ACN, while hydrophobic interactions are weakened due to solvation of hydrophobic amino acid side chains by ACN, again displacing water molecules. Even though both co-solvents displace water molecules around different side chains, the overall reduced hydration shell could be supportive for a distinct fibril structure.
We can also consider solvent influence from the point of view of relieving structural frustrations in the planar monomer fold.3 The most important structural frustrations in the insulin fibril structure arise from three glutamic acid residues in positions A4, A17 and B1331 which form part of the mostly hydrophobic fibril core. With acetic acid forming hydrogen bonds to glutamic acid and ACN weakening hydrophobic interactions between the adjacent hydrophobic residues, both co-solvents could lead to a slightly different packing of the fibril core, which then results in a different β-sheet twist and disulfide conformation.
In addition, we need to consider an interplay between tertiary and quaternary structure of insulin fibrils. For example, if acetic acid and acetonitrile facilitate self-assembly of the U-shaped insulin protofibrils into a tetrameric bundle, then this tetramer could be stabilised by an increased twist of the fibril backbone. In principle any solvent change that facilitates self-assembly of protofibrils into tetramers, e.g. through a reduced charge repulsion could have the same effect. This would provide an overarching explanation for the different fibril morphologies reported in the literature.
Conclusions
Insulin fibril polymorphism has been widely observed but very few links have been made between observed morphology and fibril structure. Drawing together information deduced from polarized Raman spectroscopy as well as infrared and fluorescence spectroscopy we have shown that structural differences consist of changes in β-sheet twists, and β-turn structure, which impact disulfide bond conformation, alignment of tyrosines and hydrogen-bonding of Gln and Glu side chains. These different structures arise because solvent components acetonitrile and acetic acid are able to change hydrophobic and hydrogen-bonding interactions between insulin monomers. This work shows the great potential of polarized Raman spectroscopy to start linking environmental conditions to specific insulin fibril structures.Author contributions
BC: investigation, formal analysis, writing – original draft; XY: methodology, investigation, formal analysis, conceptualisation, writing – original draft; AM: investigation, writing – review &editing; JM: conceptualisation, supervision, writing – review&editing; HA: conceptualisation, visualisation, investigation and formal analysis, supervision, writing – review&editing.Conflicts of interest
There are no conflicts of interest to declare.Data availability
Data will be made available on request.Supplementary information: additional Raman spectra, simulation and analysis, TEM analysis, SAXS simulation. See DOI: https://doi.org/10.1039/d5cp04579g.
Acknowledgements
We thank Dr Pedro Oseliero-Filho at the Materials Innovation Factory Liverpool for help with acquisition and analysis of SAXS data. Xiaoqi Ye gratefully acknowledges financial support from the China Scholarship Council (Grant No. 201906870026). We acknowledge the Biomedical EM Unit Shared Research Facility at the University of Liverpool.References
- M. Sunde; L. C. Serpell; M. Bartlam; P. E. Fraser; M. B. Pepys and C. C. F. Blake Common Core Structure of Amyloid Fibrils by Synchrotron X-Ray Diffraction.
- F. Chiti and C. M. Dobson, Protein Misfolding, Functional Amyloid, and Human Disease, Annu. Rev. Biochem., 2006, 75(1), 333–366, DOI:10.1146/annurev.biochem.75.101304.123901.
- M. R. Sawaya, M. P. Hughes, J. A. Rodriguez, R. Riek and D. S. Eisenberg, The Expanding Amyloid Family: Structure, Stability, Function, and Pathogenesis, Cell, 2021, 184(19), 4857–4873, DOI:10.1016/j.cell.2021.08.013.
- M. R. Sawaya, S. Sambashivan, R. Nelson, M. I. Ivanova, S. A. Sievers, M. I. Apostol, M. J. Thompson, M. Balbirnie, J. J. W. Wiltzius, H. T. McFarlane, A. Ø. Madsen, C. Riekel and D. Eisenberg, Atomic Structures of Amyloid Cross-β Spines Reveal Varied Steric Zippers, Nature, 2007, 447(7143), 453–457, DOI:10.1038/nature05695.
- R. Gallardo, N. A. Ranson and S. E. Radford, Amyloid Structures: Much More than Just a Cross-β Fold, Curr. Opin. Struct. Biol., 2020, 60, 7–16, DOI:10.1016/j.sbi.2019.09.001.
- D. Li and C. Liu, Conformational Strains of Pathogenic Amyloid Proteins in Neurodegenerative Diseases, Nat. Rev. Neurosci., 2022, 23(9), 523–534, DOI:10.1038/s41583-022-00603-7.
- X. Wang, S. Zhang, J. Zhang, Y. Wang, X. Jiang, Y. Tao, D. Li, C. Zhong and C. Liu, Rational Design of Functional Amyloid Fibrillar Assemblies, Chem. Soc. Rev., 2023, 52(14), 4603–4631, 10.1039/D2CS00756H.
- L. C. Serpell; P. E. Fraser and M. Sunde X-Ray Fiber Diffraction of Amyloid Fibrils, in Methods in Enzymology, Elsevier, 1999, vol. 309, pp. 526–536 DOI:10.1016/S0076-6879(99)09036-9.
- R. Tycko, The Evolving Role of Solid State Nuclear Magnetic Resonance Methods in Studies of Amyloid Fibrils, Curr. Opin. Struct. Biol., 2025, 92, 103043, DOI:10.1016/j.sbi.2025.103043.
- S. H. W. Scheres, B. Ryskeldi-Falcon and M. Goedert, Molecular Pathology of Neurodegenerative Diseases by Cryo-EM of Amyloids, Nature, 2023, 621(7980), 701–710, DOI:10.1038/s41586-023-06437-2.
- Searching RCSB Protein Data Bank for 3D Structures Similar to Pdb 6A6B, a Cryo-EM Fibril Structure of Human Wild-Type α-Syn (1–140), 2025.
- D. Li and L. Cong, Molecular Rules Governing the Structural Polymorphism of Amyloid Fibrils in Neurodegenerative Diseases, Structure, 2023, 31, 1335–1347, DOI:10.1016/j.str.2023.08.006.
- K. Iwaya, T. Zako, J. Fukunaga, K. M. Sörgjerd, K. Ogata, K. Kogure, H. Kosano, M. Noritake, M. Maeda, Y. Ando, Y. Katsura and T. Nagase, Toxicity of Insulin-Derived Amyloidosis: A Case Report, BMC Endocr. Disord., 2019, 19(1), 61, DOI:10.1186/s12902-019-0385-0.
- T. Nagase, Y. Katsura, Y. Iwaki, K. Nemoto, H. Sekine, K. Miwa, T. Oh-i, K. Kou, K. Iwaya, M. Noritake and T. Matsuoka, The Insulin Ball, The Lancet, 2009, 373(9658), 184, DOI:10.1016/S0140-6736(09)60041-6.
- M. H. A. Fagihi and S. Bhattacharjee, Amyloid Fibrillation of Insulin: Amelioration Strategies and Implications for Translation, ACS Pharmacol. Transl. Sci., 2022, 5(11), 1050–1061, DOI:10.1021/acsptsci.2c00174.
- A. Das, M. Shah and I. Saraogi, Molecular Aspects of Insulin Aggregation and Various Therapeutic Interventions, ACS Bio Med. Chem. Au, 2022, 2, 205–221, DOI:10.1021/acsbiomedchemau.1c00054.
- M. Akbarian, R. Yousefi, F. Farjadian and V. N. Uversky, Insulin Fibrillation: Toward Strategies for Attenuating the Process, Chem. Commun., 2020, 56(77), 11354–11373, 10.1039/d0cc05171c.
- O. M. Selivanova and O. V. Galzitskaya, Structural Polymorphism and Possible Pathways of Amyloid Fibril Formation on the Example of Insulin Protein, Biochemistry, 2012, 77(11), 1237–1247, DOI:10.1134/s0006297912110028.
- M. Ziaunys, K. Mikalauskaite, A. Sakalauskas and V. Smirnovas, Study of Insulin Aggregation and Fibril Structure under Different Environmental Conditions, IJMS, 2024, 25(17), 9406, DOI:10.3390/ijms25179406.
- D. Kurouski, R. A. Lombardi, R. K. Dukor, I. K. Lednev and L. A. Nafie, Direct Observation and PH Control of Reversed Supramolecular Chirality in Insulin Fibrils by Vibrational Circular Dichroism, Chem. Commun., 2010, 46(38), 7154, 10.1039/c0cc02423f.
- M. Krupová, J. Kessler and P. Bouř, Polymorphism of Amyloid Fibrils Induced by Catalytic Seeding: A Vibrational Circular Dichroism Study, Chem. Phys. Chem., 2021, 22(1), 83–91, DOI:10.1002/cphc.202000797.
- M. Ziaunys, A. Sakalauskas and V. Smirnovas, Identifying Insulin Fibril Conformational Differences by Thioflavin-T Binding Characteristics, Biomacromolecules, 2020, 21(12), 4989–4997, DOI:10.1021/acs.biomac.0c01178.
- T. Hiramatsu, N. Yamamoto, S. Ha, Y. Masuda, M. Yasuda, M. Ishigaki, K. Yuzu, Y. Ozaki and E. Chatani, Iodine Staining as a Useful Probe for Distinguishing Insulin Amyloid Polymorphs, Sci. Rep., 2020, 10(1), 16741, DOI:10.1038/s41598-020-73460-y.
- W. Dzwolak and M. Pecul, Chiral Bias of Amyloid Fibrils Revealed by the Twisted Conformation of Thioflavin T: An Induced Circular Dichroism/DFT Study, FEBS Lett., 2005, 579(29), 6601–6603, DOI:10.1016/j.febslet.2005.10.048.
- W. Dzwolak, R. Ravindra and R. Winter, Hydration and Structure - the Two Sides of the Insulin Aggregation Process, Phys. Chem. Chem. Phys., 2004, 6(8), 1938–1943, 10.1039/b314086e.
- J. L. Whittingham, D. J. Scott, K. Chance, A. Wilson, J. Finch, J. Brange and G. G. Dodson, Insulin at PH 2: Structural Analysis of the Conditions Promoting Insulin Fibre Formation, J. Mol. Biol., 2002, 318(2), 479–490, DOI:10.1016/s0022-2836(02)00021-9.
- K. Yuzu, N. Yamamoto, M. Noji, M. So, Y. Goto, T. Iwasaki, M. Tsubaki and E. Chatani, Multistep Changes in Amyloid Structure Induced by Cross-Seeding on a Rugged Energy Landscape, Biophys. J., 2021, 120(2), 284–295, DOI:10.1016/j.bpj.2020.12.005.
- E. Bystrenova, Z. Bednarikova, M. Barbalinardo, F. Valle, Z. Gazova and F. Biscarini, Insulin Amyloid Structures and Their Influence on Neural Cells, Colloids Surf., B, 2018, 161, 177–182, DOI:10.1016/j.colsurfb.2017.10.054.
- H. Yoshihara, J. Saito, A. Tanabe, T. Amada, T. Asakura, K. Kitagawa and S. Asada, Characterization of Novel Insulin Fibrils That Show Strong Cytotoxicity Under Physiological PH, J. Pharm. Sci., 2016, 105(4), 1419–1426, DOI:10.1016/j.xphs.2016.01.025.
- W. Mori, K. Yuzu, N. Lobsiger, H. Nishioka, H. Sato, T. Nagase, K. Iwaya, M. Lindgren and T. Zako, Degradation of Insulin Amyloid by Antibiotic Minocycline and Formation of Toxic Intermediates, Sci. Rep., 2021, 11(1), 6857, DOI:10.1038/s41598-021-86001-y.
- S. Suladze, R. Sarkar, N. Rodina, K. Bokvist, M. Krewinkel, D. Scheps, N. Nagel, B. Bardiaux and B. Reif, Atomic Resolution Structure of Full-Length Human Insulin Fibrils, Proc. Natl. Acad. Sci. U. S. A., 2024, 121(23), e2401458121, DOI:10.1073/pnas.2401458121.
- L. Wang, C. E. Hall, E. Uchikawa, D. Chen, E. Choi, X. Zhang and X. Bai, Structural Basis of Insulin Fibrillation, Sci. Adv., 2023, 9(37), eadi1057, DOI:10.1126/sciadv.adi1057.
- L. Nielsen, S. Frokjaer, J. F. Carpenter and J. Brange, Studies of the Structure of Insulin Fibrils by Fourier Transform Infrared (FTIR) Spectroscopy and Electron Microscopy, J. Pharm. Sci., 2001, 90(1), 29–37, DOI:10.1002/1520-6017(200101)90:1<29::aid-jps4>3.0.co;2-4.
- T. Sneideris, D. Darguzis, A. Botyriute, M. Grigaliunas, R. Winter and V. Smirnovas, PH-Driven Polymorphism of Insulin Amyloid-Like Fibrils, PLoS One, 2015, 10(8), e0136602, DOI:10.1371/journal.pone.0136602.
- A. K. Attri, C. Fernandez and A. P. Minton, PH-Dependent Self-Association of Zinc-Free Insulin Characterized by Concentration-Gradient Static Light Scattering, Biophys. Chem., 2010, 148(1–3), 28–33, DOI:10.1016/j.bpc.2010.02.002.
- W. Bocian, J. Sitkowski, E. Bednarek, A. Tarnowska, R. Kawecki and L. Kozerski, Structure of Human Insulin Monomer in Water/Acetonitrile Solution, J. Biomol. NMR, 2008, 40(1), 55–64, DOI:10.1007/s10858-007-9206-2.
- A. M. Katti, K. Kmiotek, J. Z. Geng and P. Goel, A Direct Approach to Insulin Isotherm Analysis in Reversed Phase Chromatography, Chromatographia, 2009, 70(1–2), 31–36, DOI:10.1365/s10337-009-1131-y.
- M. Harper, U. Nudurupati, R. J. Workman, T. I. Lakoba, N. Perez, D. Nelson, Y. Ou and D. Punihaole, Toward Determining Amyloid Fibril Structures Using Experimental Constraints from Raman Spectroscopy, J. Chem. Phys., 2023, 159(22), 225101, DOI:10.1063/5.0177437.
- J. D. Flynn and J. C. Lee, Raman Fingerprints of Amyloid Structures, Chem. Commun., 2018, 54(51), 6983–6986, 10.1039/C8CC03217C.
- J. D. Flynn, R. P. McGlinchey, R. L. Walker and J. C. Lee, Structural Features of α-Synuclein Amyloid Fibrils Revealed by Raman Spectroscopy, J. Biol. Chem., 2018, 293(3), 767–776, DOI:10.1074/jbc.M117.812388.
- M. Nilsson, Techniques to Study Amyloid Fibril Formation in Vitro, Methods, 2004, 34(1), 151–160, DOI:10.1016/j.ymeth.2004.03.012.
- V. Sereda, M. R. Sawaya and I. K. Lednev, Structural Organization of Insulin Fibrils Based on Polarized Raman Spectroscopy: Evaluation of Existing Models, J. Am. Chem. Soc., 2015, 137(35), 11312–11320, DOI:10.1021/jacs.5b07535.
- W. Surmacz-Chwedoruk, V. Babenko, R. Dec, P. Szymczak and W. Dzwolak, The Emergence of Superstructural Order in Insulin Amyloid Fibrils upon Multiple Rounds of Self-Seeding, Sci. Rep., 2016, 6, 32022, DOI:10.1038/srep32022.
- V. Sereda and I. K. Lednev, Polarized Raman Spectroscopy of Aligned Insulin Fibrils, J. Raman Spectrosc., 2014, 45(8), 665–671, DOI:10.1002/jrs.4523.
- J. R. Lakowicz and G. Weber, Quenching of Protein Fluorescence by Oxygen. Detection of Structural Fluctuations in Proteins on the Nanosecond Time Scale, Biochemistry, 1973, 12(21), 4171–4179, DOI:10.1021/bi00745a021.
- V. N. Uversky, L. N. Garriques, I. S. Millett, S. Frokjaer, J. Brange, S. Doniach and A. L. Fink, Prediction of the Association State of Insulin Using Spectral Parameters, J. Pharm. Sci., 2003, 92(4), 847–858, DOI:10.1002/jps.10355.
- E. J. Nettleton, P. Tito, M. Sunde, M. Bouchard, C. M. Dobson and C. V. Robinson, Characterization of the Oligomeric States of Insulin in Self-Assembly and Amyloid Fibril Formation by Mass Spectrometry, Biophys. J., 2000, 79(2), 1053–1065, DOI:10.1016/s0006-3495(00)76359-4.
- A. Kamada, M. Rodriguez-Garcia, F. S. Ruggeri, Y. Shen, A. Levin and T. P. J. Knowles, Controlled Self-Assembly of Plant Proteins into High-Performance Multifunctional Nanostructured Films, Nat. Commun., 2021, 12(1), 3529, DOI:10.1038/s41467-021-23813-6.
- I. B. Bekard and D. E. Dunstan, Tyrosine Autofluorescence as a Measure of Bovine Insulin Fibrillation, Biophys. J., 2009, 97(9), 2521–2531, DOI:10.1016/j.bpj.2009.07.064.
- A. Barth, The Infrared Absorption of Amino Acid Side Chains, Prog. Biophys. Mol. Biol., 2000, 74(3–5), 141–173, DOI:10.1016/S0079-6107(00)00021-3.
- K. Ito and H. J. Bernstein, The Vibrational Spectra of the Formate, Acetate, and Oxalate Ions, Can. J. Chem., 1956, 34(2), 170–178, DOI:10.1139/v56-021.
- J. H. Choi, B. C. H. May, H. Wille and F. E. Cohen, Molecular Modeling of the Misfolded Insulin Subunit and Amyloid Fibril, Biophys. J., 2009, 97(12), 3187–3195, DOI:10.1016/j.bpj.2009.09.042.
- J. L. Jimenez, E. J. Nettleton, M. Bouchard, C. V. Robinson, C. M. Dobson and H. R. Saibil, The Protofilament Structure of Insulin Amyloid Fibrils, Proc. Natl. Acad. Sci. U. S. A., 2002, 99(14), 9196–9201, DOI:10.1073/pnas.142459399.
- J. Zhou, S. Assenza, M. Tatli, J. Tian, I. M. Ilie, E. L. Starostin, A. Caflisch, T. P. J. Knowles, G. Dietler, F. S. Ruggeri, H. Stahlberg, S. K. Sekatskii and R. Mezzenga, Hierarchical Protofilament Intertwining Rules the Formation of Mixed-Curvature Amyloid Polymorphs, Adv. Sci., 2024, 11(32), 2402740, DOI:10.1002/advs.202402740.
- A. V. Mikhonin, S. V. Bykov, N. S. Myshakina and S. A. Asher, Peptide Secondary Structure Folding Reaction Coordinate: Correlation between UV Raman Amide III Frequency, Ψ Ramachandran Angle, and Hydrogen Bonding, J. Phys. Chem. B, 2006, 110(4), 1928–1943, DOI:10.1021/jp054593h.
- X. Periole, T. Huber, A. Bonito-Oliva, K. C. Aberg, P. C. A. van der Wel, T. P. Sakmar and S. J. Marrink, Energetics Underlying Twist Polymorphisms in Amyloid Fibrils, J. Phys. Chem. B, 2018, 122(3), 1081–1091, DOI:10.1021/acs.jpcb.7b10233.
- X. Ye, J. Madine and H. Arnolds, The Influence of Membrane Curvature on Insulin Aggregation, Colloids Surf., B, 2025, 255, 114914, DOI:10.1016/j.colsurfb.2025.114914.
- S. Mangialardo, F. Piccirilli, A. Perucchi, P. Dore and P. Postorino, Raman Analysis of Insulin Denaturation Induced by High-Pressure and Thermal Treatments: Raman Analysis of Insulin Denaturation Induced by High-Pressure and Thermal Treatments, J. Raman Spectrosc., 2012, 43(6), 692–700, DOI:10.1002/jrs.3097.
- D. Kurouski, J. Washington, M. Ozbil, R. Prabhakar, A. Shekhtman and I. K. Lednev, Disulfide Bridges Remain Intact While Native Insulin Converts into Amyloid Fibrils, PLoS One, 2012, 7(6), e36989, DOI:10.1371/journal.pone.0036989.
- M. Ishigaki, K. Morimoto, E. Chatani and Y. Ozaki, Exploration of Insulin Amyloid Polymorphism Using Raman Spectroscopy and Imaging, Biophys. J., 2020, 118(12), 2997–3007, DOI:10.1016/j.bpj.2020.04.031.
- H. R. Kilgore and R. T. Raines, N →π* Interactions Modulate the Properties of Cysteine Residues and Disulfide Bonds in Proteins, J. Am. Chem. Soc., 2018, 140(50), 17606–17611, DOI:10.1021/jacs.8b09701.
- H. E. Van Wart and H. A. Scheraga, Agreement with the Disulfide Stretching Frequency-Conformation Correlation of Sugeta, Go, and Miyazawa, Proc. Natl. Acad. Sci. U. S. A., 1986, 83(10), 3064–3067, DOI:10.1073/pnas.83.10.3064.
- B. Hernández, F. Pflüger, E. López-Tobar, S. G. Kruglik, J. V. Garcia-Ramos, S. Sanchez-Cortes and M. Ghomi, Disulfide Linkage Raman Markers: A Reconsideration Attempt: Disulfide Raman Markers, J. Raman Spectrosc., 2014, 45(8), 657–664, DOI:10.1002/jrs.4521.
- L. G. Tensmeyer and J. E. Shields, Raman Spectra of Crystalline 4Zn, 2Zn, and Na Insulin, Proc.SPIE, 1990, 1336, 222–234, DOI:10.1117/12.22913.
- W. Qian and S. Krimm, Vibrational Studies of the Disulfide Group in Proteins. Part V. Correlation of SS Stretch Frequencies with the CCSS Dihedral Angle in Known Protein Disulfide Bridges, Biopolymers, 1992, 32(4), 321–326, DOI:10.1002/bip.360320405.
- M. Tsuboi, J. M. Benevides and G. J. Thomas, Jr., Raman Tensors and Their Application in Structural Studies of Biological Systems, Proc. Jpn. Acad., Ser. B, 2009, 85(3), 83–97, DOI:10.2183/pjab.85.83.
- Z. Zhang and H. F. Gleeson, Understanding Liquid Crystal Order Parameters Deduced from Different Vibrations in Polarised Raman Spectroscopy, Liq. Cryst., 2019, 46(2), 219–233, DOI:10.1080/02678292.2018.1485980.
- A. Ahmad, I. S. Millett, S. Doniach, V. N. Uversky and A. L. Fink, Partially Folded Intermediates in Insulin Fibrillation, Biochemistry, 2003, 42(39), 11404–11416, DOI:10.1021/bi034868o.
- L. Nielsen, S. Frokjaer, J. Brange, V. N. Uversky and A. L. Fink, Probing the Mechanism of Insulin Fibril Formation with Insulin Mutants, Biochemistry, 2001, 40(28), 8397–8409, DOI:10.1021/bi0105983.
- Q. X. Hua and M. A. Weiss, Mechanism of Insulin Fibrillation - The Structure of Insulin under Amyloidogenic Conditions Resembles a Protein-Folding Intermediate, J. Biol. Chem., 2004, 279(20), 21449–21460, DOI:10.1074/jbc.M314141200.
- W. Dzwolak, S. Grudzielanek, V. Smirnovas, R. Ravindra, C. Nicolini, R. Jansen, A. Loksztejn, S. Porowski and R. Winter, Ethanol-Perturbed Amyloidogenic Self-Assembly of Insulin: Looking for Origins of Amyloid Strains, Biochemistry, 2005, 44(25), 8948–8958, DOI:10.1021/bi050281t.
- P. Wang, X. Wang, L. Liu, H. Zhao, W. Qi and M. He, The Hydration Shell of Monomeric and Dimeric Insulin Studied by Terahertz Time-Domain Spectroscopy, Biophys. J., 2019, 117(3), 533–541, DOI:10.1016/j.bpj.2019.06.028.
- N. Javid, K. Vogtt, C. Krywka, M. Tolan and R. Winter, Capturing the Interaction Potential of Amyloidogenic Proteins, Phys. Rev. Lett., 2007, 99(2), 028101, DOI:10.1103/PhysRevLett.99.028101.
- V. Vetri, F. Piccirilli, J. Krausser, G. Buscarino, U. Lapinska, B. Vestergaard, A. Zaccone and V. Fodera, Ethanol Controls the Self-Assembly and Mesoscopic Properties of Human Insulin Amyloid Spherulites, J. Phys. Chem. B, 2018, 122(12), 3101–3112, DOI:10.1021/acs.jpcb.8b01779.
- K. Gekko, E. Ohmae, K. Kameyama and T. Takagi, Acetonitrile-Protein Interactions: Amino Acid Solubility and Preferential Solvation, Biochim. Biophys. Acta, Protein Struct. Mol. Enzymol., 1998, 1387(1–2), 195–205, DOI:10.1016/S0167-4838(98)00121-6.
- L. Deng, Y. Zhao, P. Zhou, H. Xu and Y. Wang, Modulation of Intra- and Inter-Sheet Interactions in Short Peptide Self-Assembly by Acetonitrile in Aqueous Solution, Chin. Phys. B, 2016, 25(12), 128704, DOI:10.1088/1674-1056/25/12/128704.
- Y. Zhao, L. Deng, J. Wang, H. Xu and J. R. Lu, Solvent Controlled Structural Transition of KI 4 K Self-Assemblies: From Nanotubes to Nanofibrils, Langmuir, 2015, 31(47), 12975–12983, DOI:10.1021/acs.langmuir.5b02303.
| This journal is © the Owner Societies 2026 |




