Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Molecular docking, MM/GBSA, FEP/MD, and DFT/MM MD studies on predicting binding affinity of carbonic anhydrase II inhibitors
Ryoma
Shimizu
,
Yu
Takano
and
Toru
Saito
 *
*
Department of Biomedical Information Sciences, Graduate School of Information Sciences, Hiroshima City University, 3-4-1 Ozuka-Higashi, Asa-Minami-Ku, Hiroshima 731-3194, Japan. E-mail: tsaito@hiroshima-cu.ac.jp; Tel: +81 82 830 1617
First published on 18th March 2026
Abstract
Human carbonic anhydrase II (hCAII) is one of the zinc-containing metalloenzymes that catalyzes various hydration reactions. We report an investigation of whether modern computational tools can predict inhibitory potency of a set of sulfonamides against hCAII. The methods used are molecular docking, molecular dynamics simulations coupled with the free energy perturbation theory (FEP/MD), molecular mechanics combined with the generalized Born and surface area continuum solvation (MM/GBSA), and quantum mechanics/molecular mechanics (QM/MM) metadynamics simulations. A comparison is presented between experimental and computed binding free energy properties. All MD-based approaches demonstrate robust performance with R2 values in the range of 0.89 to 0.99 for the subset of structurally simple sulfonamides, underscoring the importance of accounting for dynamic protein–ligand interactions. With regard to the other subset comprised of structurally more diverse sulfonamides, for which the QM(B3LYP)/MM(CHARMM) metadynamics approach is less affordable, the FEP/MD method yields an R2 value of 0.70. Notably, the R2 value increases to 0.80 after the removal of one outlier (chlorzolamide).
1 Introduction
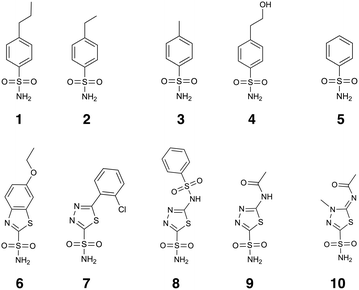
Carbonic anhydrases (CAs) are zinc containing metalloenzymes that catalyze a variety of hydration reactions such as reversible interconversion between CO2 and HCO3−.1–3 Among eight distinct families, only α-CAs are found in higher vertebrates.4 Fifteen α-CA isoforms that are structurally similar but functionally dissimilar have been identified in humans.5 In the active site of human CAs (hCAs), a Zn(II) ion is coordinated to three histidine residues (His94, His96, and His119) and to a water molecule or a hydroxide ion in a distorted tetrahedral manner (Fig. 1a). Different isoforms are responsible for distinct diseases including glaucoma (hCAI, hCAII, hCAIV, and hCAXII) and cancers (hCAIX and hCAXII).3To date, sulfonamides that replace the bound water species and directly bind to the Zn(II) ion have been the most widely used inhibitors,6 and several sulfonamide drugs have long been effective antiglaucoma drugs in clinical use.7,8 However, existing inhibitors face challenges as exemplified by the lack of selectivity. When another isozyme is a target, inhibitors exhibiting poor selectivity may give rise to undesired side effects, thereby resulting in clinical trial failure. As such, emphasis is placed on how to design inhibitors with high selectivity between isoforms.9–11 Because hCAII is abundantly expressed and broadly distributed in human tissues,12 there have been plenty of experimental and computational studies that scrutinize the impact of scaffolds and substituents on binding of sulfonamides to hCAII and hence on their inhibitory activity and selectivity.13–35 X-ray crystallographic studies have provided in-depth insights into how an inhibitor is accommodated in the active site, as well as the hydrophobic and hydrophilic interactions between the inhibitor and nearby amino acid residues (Fig. 1b).20–22,28 For convenience purposes, an inhibitor will also be referred to as a ligand hereafter. Currently, a series of sulfonamide-based analogues show subnanomolar inhibition potency in terms of the equilibrium dissociation constant (KD = koff/kon) and inhibitory constant (Ki) values, where kon (koff) represents the protein–ligand association (dissociation) rate constant.36 Gaspari et al. consistently measured these kinetic constants (kon, koff, and KD) for five benzenesulfonamides 1–5 (Fig. 2) using surface plasmon resonance methodology. In line with the stopped-flow fluorescence experiments,13,14 the authors found that the kon values increase with the type of substituent in the 4-position in the order CH2CH2CH3 (ligand 1) > CH2CH3 (ligand 2) > CH3 (ligand 3) > CH2CH2OH (ligand 4) > H (ligand 5).28
On the computational side, among the available set of modern computational methodologies,37–41 molecular docking simulations, molecular dynamics (MD) based free energy calculations, and machine/deep learning techniques have mainly been exploited,16,18,24,26–30,33–35 compared with computationally demanding first-principles quantum mechanics (QM) methods represented by density functional theory (DFT).31,34 These molecular mechanics (MM) based calculations have a great advantage over QM methods by virtue of much lower computational costs, but at the same time they require fine-tuned force field parameters that can reproduce the geometry of the zinc active site. One of the realistic goals of these approaches is to predict the binding affinity and inhibitory potency of a ligand based on structural and energetic properties and, in turn, to offer guidelines for designing appropriate new ligands. Molecular docking and more expensive MM combined with the generalized Born and surface area continuum solvation (MM/GBSA) simulations39,42,43 have widely been assessed.18,24,26,27,29,30,33,35
Very recently, Taylor and Ho applied the molecular docking simulations using the AutoDock Vina (Vina) package in conjunction with the AutoDock4Zn force field44 and MM/GBSA simulations to a dataset of 32 chemically diverse ligands.33 The authors demonstrated that the former showed a moderately strong correlation with experiment with an R2 coefficient of 0.64, and that the latter had a stronger correlation up to R2 = 0.77 when B3LYP-D3(BJ)45,46 derived electrostatic potential (ESP) charges were used for the ligands. MD simulations coupled with the alchemical free energy perturbation theory (denoted as FEP/MD)37,47 are more robust than the MM/GBSA calculations. Guimarães reported that the MM/GBSA approach yielded an R2 value of 0.71 for 13 monosubstituted benzenesulfonamides.26 But its application appears to be confined to structurally highly similar and simple sulfonamides presumably due to the challenges in setting up and computational cost.16,26
In the present study, we have systematically assessed whether docking scores and (relative) binding free energies computed with molecular docking, QM/MM metadynamics, MM/GBSA, and FEP/MD simulations can be compatible with the experimental binding free energies calculated from KD values (ΔGKD) for structurally highly similar monosubstituted benzenesulfonamides 1–5. We have also investigated how the choice of force field parameter sets affects their results. Kinetic properties represented by kon are related to the activation free energy36 and have been reported to correlate better with experimental drug efficacy compared with KD.48–50 As such, another comparison was carried out between the experimental free energies based on kon values (ΔGkon) and the activation free energy for the binding process of each ligand (ΔG‡) predicted by QM/MM metadynamics simulations. Correlations between the ΔGkon values and computed (relative) binding free energies and between the experimental ΔGKD values and calculated ΔG‡ values were also investigated for comparison. Then, by means of validated and feasible methods, correlation was checked with the ΔG values calculated from Ki values (ΔGKi) for structurally less similar and diverse ethoxzolamide (ligand 6), chlorzolamide (ligand 7), benzolamide (ligand 8), acetazolamide (ligand 9), and methazolamide (ligand 10) as illustrated in Fig. 2.
2 Results and discussion
2.1 Molecular docking
We examined the Vina, AutoDock4 (AD4), and genetic optimization for ligand docking (GOLD) approaches. Note that the AutoDock4Zn force field was commonly used for Vina and AD4. When the anionic form of each ligand was docked into the binding pocket, non-bonded interactions stabilized the binding of the N atom of the sulfonamide group to the Zn(II) ion. Concerning ligand 1, the predicted best pose reproduced its X-ray crystal structure (4YXU, see also Fig. S1)28 regardless of the docking technique used. Table 1 collects the relationship between the experimental ΔGKD and ΔGkon values and the ligand binding affinities in terms of the binding energies (ΔEVina and ΔEAD4) and GOLD fitness scores (GFSs) for 1–5.| Ligand | ΔGKDab | ΔGkonab | ΔGKiac | ΔEVinaa | ΔEAD4a | GFS | ΔΔGad | ΔGad |
|---|---|---|---|---|---|---|---|---|
a In kcal mol−1.
b From ref. 28. The formulae ΔGKD = RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(KD) and ΔGkon= −RT ln(KD) and ΔGkon= −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(kon) were used with R = 8.314 J (K mol)−1 and T = 300 K.
c From ref. 19. The formula ΔGKi = RT ln(kon) were used with R = 8.314 J (K mol)−1 and T = 300 K.
c From ref. 19. The formula ΔGKi = RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(Ki) was used with R = 8.314 J (K mol)−1 and T = 300 K.
d Using the covalent bond (non-bonded) model. Data are presented as mean ± standard deviation. ln(Ki) was used with R = 8.314 J (K mol)−1 and T = 300 K.
d Using the covalent bond (non-bonded) model. Data are presented as mean ± standard deviation.
|
||||||||
| 1 | −9.91 | −9.45 | N.A. | −8.75 | −8.84 | 56.28 | 0.0 (0.0) | −19.5 ± 2.5 (−13.0 ± 1.1) |
| 2 | −9.47 | −9.15 | N.A. | −8.60 | −8.66 | 55.56 | 1.6 ± 0.5 (2.1 ± 0.8) | −17.9 ± 3.2 (−11.6 ± 1.9) |
| 3 | −8.89 | −8.65 | N.A. | −8.72 | −8.74 | 53.07 | 2.2 ± 0.4 (2.1 ± 0.5) | −16.8 ± 0.8 (−11.1 ± 0.4) |
| 4 | −8.74 | −8.51 | N.A. | −8.50 | −8.67 | 55.69 | 2.1 ± 0.1 (2.4 ± 0.9) | −16.4 ± 2.6 (−10.4 ± 2.0) |
| 5 | −8.16 | −8.43 | N.A. | −8.43 | −8.46 | 51.75 | 2.8 ± 0.3 (3.5 ± 0.5) | −15.1 ± 1.1 (−12.1 ± 4.3) |
| 6 | N.A. | N.A. | −12.57 | −8.79 | −8.83 | 60.61 | 0.0 | −23.1 ± 1.9 |
| 7 | N.A. | N.A. | −11.95 | −9.40 | −9.53 | 60.38 | −1.6 ± 2.8 | −25.5 ± 1.5 |
| 8 | N.A. | N.A. | −11.20 | −9.47 | −9.79 | 67.88 | 1.9 ± 0.6 | −28.9 ± 1.5 |
| 9 | N.A. | N.A. | −10.98 | −8.63 | −8.71 | 55.36 | 4.9 ± 1.0 | −17.9 ± 1.2 |
| 10 | N.A. | N.A. | −10.89 | −8.99 | −9.00 | 58.57 | 4.7 ± 1.8 | −13.2 ± 0.4 |
As expected, these values both appear to correlate more with ΔGKD than with ΔGkon judged by R2 (Fig. S2a–f). Vina and AD4 yielded slightly different results due to different scoring functions, which have been discussed elsewhere.51,52 The magnitudes of the ΔEVina values decrease in the order 1 > 3 > 2 > 4 > 5, whereas the ΔEAD4 values follow the order 1 > 3 > 4 > 2 > 5. Despite the underestimation of the binding energy of ligand 2, ΔEVina and ΔEAD4 are moderately correlated with the ΔGKD values for 1–5, with R2 values of 0.61 and 0.71 (see Fig. S2a and b). On the other hand, the GOLD fitness scores decrease in the order 1 > 4 > 2 > 3 > 5 and, except for 4, the behavior is consistent with the experimentally observed order 1 > 2 > 3 > 4 > 5. The results to some extent support the fact that the magnitude of ligand binding energy increases with increasing alkyl chain length, with an R2 coefficient of 0.65 (Fig. S2c). However, it is clearly seen that the evaluation of the score tends to prioritize similarity in size over difference in hydrophobicity, thereby leading 1 and 4 to exhibit much the same scores (56.28 vs. 55.69).
Table 1 also summarizes the relationship between the experimental ΔGKi values and the calculated results (ΔEVina, ΔEAD4, and GFSs) for 6–10. In contrast to the case of ligands 1–5, all of the molecular docking methods cannot be applicable to ligands 6–10, which have more diverse and unsystematic molecular structures. The calculated results obtained with Vina, AD4, and GOLD demonstrate no correlation with the ΔGKi values, yielding R2 values in the range of 0.00 to 0.02 (Fig. S2g–i). Ligand 8 is predicted to be too stable, whereas ligand 6 appears to be less stable than it should be based on the experimental ΔGKi values. Strictly speaking, the KD and Ki values are not entirely equivalent but could be quite similar.53–55 Because these values are converted to kcal mol−1 using the equation: ΔGX = RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) X (X = KD and Ki), it may be more feasible to aggregate and analyze data for all the ligands. Fig. 3a–c illustrate the overall correlations between the experimental and calculated data for 1–10, with weak-to-moderate R2 values of 0.44, 0.37 and 0.51 for Vina, AD4, and GOLD, respectively.
X (X = KD and Ki), it may be more feasible to aggregate and analyze data for all the ligands. Fig. 3a–c illustrate the overall correlations between the experimental and calculated data for 1–10, with weak-to-moderate R2 values of 0.44, 0.37 and 0.51 for Vina, AD4, and GOLD, respectively.
2.2 QM/MM metadynamics
We hypothesized that the experimental kinetic constants (kon and KD) are comparable to activation and reaction free energies (ΔG‡ and ΔG) that can describe the kinetics of the ligand binding process. For each ligand, we relied on a binding process in which a proton is first transferred from the NH2 group of the neutral sulfonamide to the Zn-bound OH−, generating a transient complex where both the H2O and the anionic sulfonamide are bound to the Zn(II) ion (see also Fig. 4).28,56,57 The concomitant dissociation of the H2O molecule leads to the final ligand-bound complex observed in the crystal structure. For all ligands tested (1–5), we find that the nitrogen of the sulfonamide group is bound to the Zn(II) ion. In fact, the Zn⋯N and the Zn⋯O(proximal) distances of 2.00 ± 0.08 and 3.07 ± 0.22 Å taken from the last 2.5 ps of a randomly selected trajectory for 1 agree well with those of 1.93 and 2.99 Å observed in the 4YXU structure (Fig. S3).28 This N atom and the other distal O atom of the O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S
S![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group properly form hydrogen bonds with the OH and NH group of Thr199 (Fig. 1b and Fig. S4). The same trend can be seen in the other structures complexed with ligands 2–5.
O group properly form hydrogen bonds with the OH and NH group of Thr199 (Fig. 1b and Fig. S4). The same trend can be seen in the other structures complexed with ligands 2–5.
 | ||
| Fig. 4 Collective variables (interatomic distances d1 and d2) that describe the binding process of a sulfonamide to the zinc active center. | ||
First, we discuss the validity of the QM method and the size of the QM region. It should be emphasized that the binding mode of the sulfonamide moiety must be calculated at the DFT level. Even the use of time-tested semiempirical methods58 failed to reproduce the correct binding mode where the N atom of the NH2 group is bound to the Zn(II) ion. The Zn-bound N atom of the NH2 group in the starting ligand 1 was irreversibly switched to one of the O atoms belonging to the O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S
S![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group. The active site both before and after the ligand binding process forms non-covalent interactions with surrounding amino acid residues such as Leu198 and Thr199, depending on the ligand size (Fig. 1b). We believe that non-covalent interactions between the QM and MM regions can sufficiently be described by the MM part of the QM/MM Hamiltonian, and thus the unbiased minimal QM region introduced in Section 3.3 was used. Indeed, the CH⋯π and hydrogen-bonding interactions were well described throughout our QM(B3LYP)/MM(CHARMM)45,59 MD simulations (see the text below).
O group. The active site both before and after the ligand binding process forms non-covalent interactions with surrounding amino acid residues such as Leu198 and Thr199, depending on the ligand size (Fig. 1b). We believe that non-covalent interactions between the QM and MM regions can sufficiently be described by the MM part of the QM/MM Hamiltonian, and thus the unbiased minimal QM region introduced in Section 3.3 was used. Indeed, the CH⋯π and hydrogen-bonding interactions were well described throughout our QM(B3LYP)/MM(CHARMM)45,59 MD simulations (see the text below).
Next, let us turn our attention to the ligand-binding reactions. Fig. 3d displays the relationship between the average ΔGkon and ΔG‡ values calculated by the QM/MM metadynamics simulations for 1–5. The absolute ΔG‡ values decrease in the order 1 > 2 > 3–5 > 4. As listed in Table 2, ligand 1 requires the lowest activation barrier (ΔG‡ = 2.9 ± 0.5 kcal mol−1). In addition to its benzene ring which forms the CH⋯π interactions with Val135 and Leu198, the propyl group can make contacts with hydrophobic residues such as Phe131 and Pro20224,60 (Movie S1). We assume that these interactions contribute to lowering the ΔG‡ value, and thus that the decrease in hydrophobic contacts in the 4-position by substitution with ethyl (2), methyl (3), hydroxyethyl (4) and hydrogen (5) is likely to provide higher average ΔG‡ values. From the inspection of trajectories, ligand 4 does not benefit from hydrophobic interactions but rather forms hydrogen-bonding interactions with a solvent water molecule analogous to the 4YYT structure28 (Movie S2). Additional hydrophobic interactions can be recognized between ligand 5 and Val121, but such interactions are not observed in the 4YX4 structure28 (Movie S3). This discrepancy may explain why the binding of 5 is more favorable than that of 4, with average ΔG‡ values of 6.8 and 8.1 kcal mol−1. Nevertheless, the calculated R2 value 0.92 ensures that the kon value can be predicted more or less quantitatively based on the calculated ΔG‡ values. Meanwhile, these ΔG‡ values correlate less with the KD values, with an R2 value of 0.73 (Fig. S5a).
| Ligand | ΔGKDa | ΔGkona | ΔG‡ | ΔG |
|---|---|---|---|---|
a From ref. 28. The formulae ΔGKD = RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(KD) and ΔGkon = −RT ln(KD) and ΔGkon = −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(kon) were used with R = 8.314 J (K mol)−1 and T = 300 K. ln(kon) were used with R = 8.314 J (K mol)−1 and T = 300 K.
|
||||
| 1 | −9.91 | −9.45 | 2.9 ± 0.5 | −1.9 ± 0.5 |
| 2 | −9.47 | −9.15 | 4.6 ± 1.8 | −3.5 ± 3.9 |
| 3 | −8.89 | −8.65 | 6.8 ± 0.4 | 1.9 ± 1.5 |
| 4 | −8.74 | −8.51 | 8.1 ± 1.9 | 4.8 ± 3.4 |
| 5 | −8.16 | −8.43 | 6.8 ± 1.1 | 1.7 ± 0.4 |
The reaction free energies of the binding process (ΔG) for 1–5 are found to be less correlated with the experimental ΔGKD (R2 = 0.51) and ΔGkon (R2 = 0.70) values as shown in Fig. 3e and S5b. After passing through the transition state, other than the results of 1, some simulations provided a transient complex where the generated H2O continues to be weakly bound to the Zn(II) ion forming hydrogen bonds with Glu106 and Thr199. We suggest that the existence of these two product states gave rise to the inconsistency in the calculated ΔG values with large uncertainties listed in Table 2. Taking the results of ligand 2 as an example, the computed one-dimensional potentials of mean force (1D-PMFs) in Fig. S6 indicate that the excessive sampling of a water-coordinated complex led to a significantly higher endergonicity (ΔG = −8.6 kcal mol−1), whereas the adequate sampling of a water-free complex corresponding to the X-ray structure provided a ΔG value of 0.7 kcal mol−1. Our overall assessment is that the computed activation free energies and reaction free energies highlight substantial (R2 = 0.92) and moderate (R2 = 0.70) correlations with the experimental ΔGkon values, respectively. It is suggested that the use of different collective variables (CVs), a different DFT functional, and/or an unbiased and larger QM region can improve accuracy in predicting reaction free energies and correlation with the experimental ΔGKD values, which will be investigated elsewhere. In view of the computational cost that requires several months of effort and the expectation that the ΔG‡ values would be unlikely to have a strong correlation with the Ki values, we decided against applying the B3LYP/CHARMM metadynamics simulations to ligands 6–10, some of which are larger in size.
2.3 Assessment of force field parameters for the active site
To describe the geometry of the zinc active site within a framework of MM force field parameters, two models namely non-bonded (NB) and covalent bond (CB) models61,62 have mainly been adopted. The NB model treats metal coordination geometries through Coulombic and Lennard-Jones (LJ) contributions only. In addition to these non-bonded interactions, the CB model delineates bonds, angles, and torsions explicitly, and equilibrium parameters and force constants are usually obtained from QM calculations.The NB model together with the default LJ parameters gave an incorrect zinc coordination geometry, as with the aforementioned results from semi-empirical methods. The resulting Zn⋯N and Zn⋯O distances are 4.39 ± 0.55 and 2.00 ± 0.06 Å. According to Lu and Voth, the LJ parameters optimized for hCAII–sulfonamide complexes yielded a Zn⋯N distance of 1.99 Å at the cost of a longer Zn⋯O separation of 4.11 Å.18 On the other hand, these distances obtained with the CB model are found to be 1.96 ± 0.05 and 2.67 ± 0.14 Å (Fig. S7). Details of our CB model will be described in 3.2, and derived force field parameters are given in the SI. Taking into account the fact that similar structures were obtained at the QM/MM and MM levels, we perceive the agreement in the protein–ligand binding site as sufficient to justify the feasibility of the CB model on the prediction of hCAII–sulfonamide complexes.
2.4 FEP/MD
As mentioned above, the ligand-binding process to hCAII may involve some chemical transformations, and more importantly, this makes it hard to calculate the absolute binding free energy based on cost-efficient classical MD simulations. Alternatively, binding free energy differences between two ligands (referred to as ΔΔG) are often evaluated using alchemical free energy perturbation MD (FEP/MD) simulations. In the FEP/MD procedure, the 4-propylphenyl group of 1 was changed to another substituent, such as phenyl (5), while the sulfonamide group remained unchanged. The average ΔΔG values for 2–5 with respect to 1 were computed by the FEP/MD simulations with both the NB and CB models as summarized in Table 1, and then were compared with the ΔGKD and ΔGkon values (Fig. 3f and S5c). We see that our computed results are able to derive the structure–activity relationship of the 4-position of benzenesulfonamides. The predicted ΔΔG values increase in the order 1 < 2 < 3–4 < 5 regardless of the chosen model. The CB model slightly demonstrates superior performance over the NB model in terms of R2 (0.89 vs. 0.84). This is true for when the computed ΔΔG values are compared with ΔGkon values (0.88 vs. 0.75) as shown in Fig. S5c. What surprised us is that the inaccurate zinc coordination geometry of the starting ligand 1 predicted by the NB model does not substantially reduce the accuracy for the prediction of ΔΔG, presumably because the FEP-generated binding modes of 2–5 may also have incorrect coordination geometries as the starting complex and allowed for cancellation of errors. That said, comparison of the results from these models strongly suggests that one must choose the CB model. We expect that the FEP/MD simulations based on the CB model are likely to be transferable to the exploration of the relationship between Ki and ΔΔG values for 6–10.The ΔΔG values for 7–10 with respect to 6 (and not to 1) were evaluated with only the CB model. Unlike the molecular docking calculations mentioned above and the MM/GBSA simulations described below, it is impossible to aggregate and analyze data for all ligands 1–10 as a whole. The calculated average values increase in the order 7 < 6 < 8 < 9–10 and show a moderate correlation with the experimental data with an R2 value of 0.70 (Fig. 3g). Although ligands 7–10 share the same thiadiazole scaffold, ligand 7 is the only outlier with large uncertainties and is recognized as the main cause that may lead to a significant deterioration in the R2 value. The removal of ligand 7 actually yields an R2 value of 0.80. Moreover, we infer from visual inspection of trajectories that 7 tends to form a halogen bond63 with nearby residues such as Gln92, which is not experimentally observed in hCAII but in the hCAIX mimic.19 If ligand 7 was to be involved in the formation of a halogen bond like in the case of the hCAIX mimic, it would cause active-site conformational changes and exhibit a two-fold increase in inhibitory activity. Considering this assumption, ΔGKi is expected to be −12.30 kcal mol−1, and therefore, the R2 value is increased to 0.82.
2.5 MM/GBSA
It has been argued that MM/GBSA calculations only need a fraction of the computational cost of FEP/MD simulations and can deal with structurally dissimilar ligands.26,33 We began by carrying out the MM/GBSA simulations to evaluate the binding free energies (ΔG) for structurally similar ligands 1–5. The results reveal that the choice of which force field model to use for the zinc active site is crucial for the correlation with the experimental binding free energies. The use of the NB model did not display a satisfying correlation with either ΔGKD or ΔGkon. Unlike the FEP/MD simulations, the MM/GBSA calculations collected internal energies on zinc coordination geometries that may vary depending on ligands, thereby resulting in the largest standard deviation of 4.3 kcal mol−1 observed for the smallest ligand 5 (Table 1) and much smaller R2 values of 0.22 for ΔGKD and 0.43 for ΔGkon (Fig. S5d and e). The CB model yielded larger absolute average ΔG values for 1–5 than the NB model. The large standard deviations among 10 independent runs are found to be 2.5, 3.2, and 2.6 kcal mol−1 for 1, 2, and 4, possibly reflecting the notion that these ligands have relatively long and flexible substituents. Nevertheless, reproducibility can be seen in the order of the absolute average ΔG values, that is, 1 > 2 > 3 > 4 > 5, and thus we find an excellent correlation between the experimental ΔGKD (ΔGkon) and calculated ΔGkon values predicted utilizing the CB model, with an R2 value of 0.99 (0.93). This ensures that the removal of the ligand conformational entropy term is proven to be valid for structurally similar ligands. To our surprise, the MM/GBSA procedure outperforms more computationally demanding QM(B3LYP)/MM(CHARMM) metadynamics and FEP/MD calculations, and we applied the approach with the CB model to the prediction of binding free energies for a set of structurally more diverse and dissimilar ligands 6–10.Although the reproducibility of the ΔG values for 6–10 seems higher than that for 1–5 as judged by the smaller standard deviations listed in Table 1, the ΔG values for 7 and 8 tend to be overestimated compared to that for 6. Consequently, the collected results demonstrate a poor correlation with the experimental data (R2 = 0.21, Fig. S5f). We consider that the reason for this significant drop may be their more diverse orientations, as evidenced by superpositions of all five protein–ligand complexes in crystal structures.19 The sulfonamide cores evidently possess similar orientations by virtue of the hydrogen bonds with Thr199, while the orientations of thiadiazole rings and substituent moieties differ from each other. Thus, our results indicate that the neglect of entropic terms does not work with the benefits of cancellation of errors unless the effects of the conformational entropy are similar in magnitude. Taking all data together (ligands 1–10), the overall R2 value is shown to be 0.40 (Fig. 3h). It should be noted that additional calculations, in which CGenFF charges were replaced with DFT-calculated ESP charges, did not improve the results of the present study.
3 Computational details
3.1 Molecular docking simulations
All molecular docking calculations were performed using Vina (version 1.2.7),64 AD4 (version 4.2.5),65 and GOLD with the Hermes visualizer tool (version 2024.2.0).66 The AutoDock4Zn force field44 was employed in conjunction with Vina and AD4. The X-ray crystallographic structure of hCAII complexed with ligand 1 was taken from Protein Data Bank (PDB entry 4YXU).28 Hydrogen atoms were added, after the removal of 1 and crystal water molecules. The atomic charges of deprotonated ligands 1–10 were assigned using the ChelpG method67 preceded by geometry optimizations at the B3LYP/def2-SVP level of theory with the ORCA program.45,68,69 With regard to the GOLD docking program, each ligand was docked into the active site of the receptor with a 10 Å sphere centered at the Zn(II) ion. The best docking pose for each compound was chosen based on the GOLD fitness score. For AD4, on the other hand, the grid size was set to 26 × 26 × 18 points with a spacing of 0.375 Å centered at the coordinate that corresponds to the center of mass of benzene in the 4XYU structure. The generated affinity maps were used for the subsequent Vina and AD4 simulations. Guided by a previous benchmark study,52 default parameters were used, except that the exhaustiveness value and energy_range were set to 400 and 100 for Vina and that the number of docking runs (ga_run) and the number of energy evaluations (ga_num_evals) were set to 100 and 2.5 × 106 for AD4, respectively. The best docking pose for each ligand was identified on the basis of the binding energy.3.2 System setup
The hCAII structures in complex with ligands 1–10 were constructed referring to the X-ray crystallographic data.19,28 Hydrogen atoms were added with the CHARMM-GUI input generator.70 The force field parameters of the neutral form of ligands 1–5 for QM/MM metadynamics simulations71 were generated by the CGenFF method.72 Likewise, those of the anionic form of ligands 1–10 were also parameterized for FEP/MD and MM-GBSA simulations. In the NB model, the CHARMM LJ parameters were used for Zn, while the CHARMM36 force field parameters59 and TIP3P water models73 were applied to the amino acid residues and water molecules. On the basis of the CB model, the force field parameters for the active site containing the Zn(II) ion, the sulfonamide core, and three histidine residues were parameterized using the antechamber module and metal center parameter builder (MCPB) method implemented in AmberTools24.61,74 The equilibrium parameters (bond lengths and angles) and force constants were obtained at the B3LYP/6-31G* level of theory45,75 by using the Gaussian16 program.76 The generated parameters were converted to CHARMM-formatted topology and parameter files provided in the SI.Subsequently, each system was immersed in a 70 Å × 70 Å × 70 Å cube box of water molecules using the visual molecular dynamics (VMD) program.77 In accordance with our previous studies,78,79 the following conditions were used for classical MD simulations unless otherwise noted. All classic MD runs were performed in the NPT ensemble using the NAMD3 program80,81 under periodic boundary conditions at 300 K. Long-range electrostatic interactions were treated by the particle mesh Ewald method82 with a tolerance of 10−6. The electrostatic and van der Waals interactions were cut off at 14 Å with a switching distance of 12 Å.
3.3 QM/MM metadynamics
Each neutral ligand was manually placed slightly away from the Zn(II)–OH− species, maintaining the same conformation as that of the anionic form binding to the Zn(II) ion. The ligand, Zn(II) ion, Zn-bound OH−, and the imidazole rings of His94, His96, and His119 were considered as the QM region and were treated by first-principles RIJCOSX-B3LYP/def2-SVP calculations.45,68,83,84 For each ligand, the total charge and spin multiplicities in the QM region were set to 1 and 1, while the MM subsystem was described by the CHARMM36 force field parameters. The system was minimized over 2000 steps, followed by a 10 ns classical MD simulation with a time step of 2 fs. During the MD simulation, the QM atoms were kept fixed along with our previous studies.78,79 Subsequently, QM/MM MD equilibration was run for 2 ps with a time step of 0.5 fs using ORCA and NAMD3 together with the NAMD–ORCA interface,85 which manages data communication between QM and MM results computed with these programs. Electrostatic interactions between QM and MM atoms were dealt with the electrostatic embedding scheme, and the QM–MM boundary atoms were capped by link (hydrogen) atoms with the charge shift method.To determine free energy landscape, we performed three independent B3LYP/CHARMM metadynamics simulations. The collective variables (CVs) were set to drive the reactions (Fig. 4). The first CV (d1) corresponds to the dissociation of OH ligand and the second one (d2) represents the binding of the N atom of the sulfonamide group to Zn. 1D-PMFs for the ligand-binding mechanisms were investigated in the NVT ensemble at 300 K. The time step, hill weight, hill width, and hill frequency were set to 0.5 fs, 0.50 kcal mol−1, 2.5 Å, and 50 fs, respectively. The simulation time was set up to 100 ps. Upon the formation of H2O that would move away from the reaction site, three independent runs generated consistent 1D-PMFs, and the simulations were considered to be converged.
3.4 FEP/MD
The binding free energy differences between ligand L1 to ligand L2 can be expressed as ΔΔG = ΔGL2 − ΔGL1. When the involvement of chemical reactions in the protein–ligand process interferes with the direct calculation of ΔGL1 and ΔGL2per se, the use of a thermodynamic cycle allows us to regard ΔΔG as equivalent to ΔGB − ΔGA as shown in Fig. 5. The subscripts A and B represent alchemical transformation from L1 to L2 in the solvent and in the solvated protein, respectively. To this end, alchemical FEP/MD simulations introduce unphysical intermediate states that connect L1 and L2 with a coupling parameter λ. The λ value for the initial and final states are often set to 0 and 1.Here, ligand 1 was chosen as the starting ligand corresponding to L1, and relative binding free energies of 2–5 (ΔΔG) were calculated by FEP/MD simulations implemented in the NAMD3 program. To compute the ΔGA term, 1 was solvated in a 30 Å × 30 Å × 30 Å cube box of water molecules with the VMD program. The solvated protein structure complexed with 1 for the calculation of ΔGB was prepared in the way described above. For both the initial systems, the energy minimization was employed for 10 ps, followed by three independent equilibration simulations for 5 ns with a time step of 1.0 fs. In what follows, we took the sulfonamide group as the core and performed three independent FEP/MD simulations, each of which used 21λ windows for the forward transformation (from λ = 0 to 1 in steps of 0.05) and backward transformation (from λ = 1 to 0 in steps of 0.05). The equilibrium and production run times were set to 10 ps and 40 ps for each λ value. Likewise, the ΔΔG values for 7–10 were also calculated with reference to 6.
3.5 MM/GBSA
The MM/GBSA simulation was carried out using MolAICal86 based on the NAMD trajectories. Each protein–ligand (PL) complex19 prepared in 3.2 was first minimized for 20 ps and equilibrated for 2.5 ns using NAMD3. Snapshots were taken from each of the 10 independent 2.5-ns-long trajectories at 10 ps intervals, which amounted to 250 snapshots. Then, both the protein-only (P) and ligand-only (L) structures were extracted from the PL complex by removing either the ligand or protein from each snapshot. The binding free energy ΔG can be expressed as| ΔG = 〈GPL − GP − GL〉PL | (1) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 〉PL represents an ensemble average over a single trajectory of the PL complex.87,88 Each free energy (G) is calculated from
〉PL represents an ensemble average over a single trajectory of the PL complex.87,88 Each free energy (G) is calculated from| G = ΔEint + ΔEelec + ΔEvdW + ΔGGB + ΔGSA | (2) |
4 Conclusions
In the present study, we have performed molecular docking, FEP/MD, MM/GBSA, and QM(B3LYP)/MM(CHARMM) metadynamics simulations to evaluate the binding affinity of hCAII inhibitors. All classical MD-based methods are found to excellently reproduce the experimental equilibrium dissociation constants for the subset of structurally simple sulfonamides, yielding R2 values of 0.89 and 0.99 for FEP/MD and MM/GBSA, respectively. These results highlight the importance of dynamic interactions between ligands and the zinc active site, and claim that the use of the CB model is necessary to reproduce correct zinc–ligand coordination geometries at the MM level. The ΔG‡ values computed with the B3LYP/CHARMM metadynamics simulations also demonstrate a substantial correlation with the experimental association rate constants (R2 = 0.92). Tests on the other subset consisting of structurally more diverse sulfonamides indicate that MM/GBSA shows poor performance compared with more robust FEP/MD in terms of R2 (0.21 vs. 0.70). Evidently, FEP/MD simulations are much more efficient than the B3LYP/CHARMM simulations that require several weeks of effort. One shortcoming of FEP/MD is that it is likely to overestimate the binding affinity of a chlorine-containing ligand, leading ligand 7 (chlorzolamide) to be the only outlier. The R2 value is restored to 0.80 and 0.82, when 7 is excluded or the Ki value from the hCAIX mimic is instead used for the experimental data. Even without these assumptions, we anticipate that the FEP/MD approach will be of use as a reference tool for predicting binding free energies of a variety of hCAII inhibitors.Author contributions
Ryoma Shimizu: formal analysis, investigation, methodology, validation, visualization, and writing – review and editing. Yu Takano: funding acquisition, resources, supervision, validation, and writing – review and editing. Toru Saito: conceptualization, funding acquisition, resources, supervision, validation, writing – original draft, and writing – review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Fig. S1, overlay of the best docked pose (green) with respect to the X-ray crystallographic structure (magenta) obtained with (a) AutoDock Vina, (b) AutoDock4, and (c) GOLD simulations; Fig. S2, comparison of binding free energies (ΔGkon, ΔGKD, ΔGKi) between the experimental and calculated results obtained by (a), (d) and (g) AutoDock Vina, (b), (e) and (h) AutoDock4, and (c), (f) and (i) GOLD calculations; Fig. S3, (a) representative snapshot of the zinc active site and (b) evolutions of the Zn⋯O and Zn⋯N distances during a 100 ps QM/MM metadynamics simulation; Fig. S4, (a) representative snapshot of the zinc active site and Thr199 and (b) evolutions of the N(Thr199)⋯O(sulfonamide) and O(Thr199)⋯N(sulfonamide) distances during a 100 ps QM/MM metadynamics simulation; Fig. S5, comparison of binding free energies (ΔGkon, ΔGKD, ΔGKi) between the experimental and calculated results obtained by (a) and (b) QM(B3LYP)/MM(CHARMM) metadynamics for 1–5, (c) FEP/MD for 1–5, (d) and (e) MM/GBSA for 1–5, and (f) MM/GBSA for 6–10; Fig. S6, one-dimensional potentials of mean force (1D-PMF) from the reactant (R) to product (P) state via transition state (TS) for the formation of (a) water-free and (b) water-coordinate complexes for ligand 2, together with their representative snapshots in P taken from the 100-ps-long QM(B3LYP)/MM(CHARMM) metadynamics simulations; Fig. S7, (a) representative snapshot of the zinc active site taken from the 5-ns-long equilibration and (b) evolutions of the Zn⋯O and Zn⋯N distances with the NB model. (c) and (d) those obtained with the CB model; topology and parameter files for the zinc active site and sulfonamide core (PDF). Movie S1: interactions between 1 and amino acid residues during the binding process predicted by QM/MM metadynamics simulations. Movie S2: interactions between 4 and amino acid residues during the binding process predicted by QM/MM metadynamics simulations. Movie S3: interactions between 5 and amino acid residues during the binding process predicted by QM/MM metadynamics simulations. See DOI: https://doi.org/10.1039/d5cp04452a.Acknowledgements
This work was supported by the Grant-in-Aid for Scientific Research (C) (No. 22K06164 and 24K08357) from the Japan Society for the Promotion of Science, and by the Grant-in-Aid for Transformative Research Areas (A) “System biosynthesis based on accumulation prediction, and creation of biological reactions” (No. 25H01591) and “Progressive condensed matter physics inspired by hyper-ordered structures” (No. 20H05883) from the Ministry of Education, Culture, Sports, Science and Technology (MEXT).Notes and references
- C. T. Supuran, A. Scozzafava and A. Casini, Med. Res. Rev., 2003, 23, 146–189 CrossRef CAS PubMed.
- V. Alterio, A. Di Fiore, K. D'Ambrosio, C. T. Supuran and G. De Simone, Chem. Rev., 2012, 112, 4421–4468 CrossRef CAS PubMed.
- N. Naeem, A. Sadiq, G. A. Othman, H. M. Yassin and E. U. Mughal, RSC Adv., 2024, 14, 35769–35970 RSC.
- C. T. Supuran, Expert Opin. Drug Discovery, 2017, 12, 61–88 CrossRef CAS.
- C. Supuran, Clin. Sci., 2021, 135, 1233–1249 CrossRef CAS PubMed.
- T. Mann and D. Keilin, Nature, 1940, 146, 164–165 CrossRef CAS.
- H. A. Quigley, Lancet, 2011, 377, 1367–1377 CrossRef PubMed.
- E. Masini, F. Carta, A. Scozzafava and C. T. Supuran, Expert Opin. Ther. Pat., 2013, 23, 705–716 CrossRef CAS PubMed.
- G. La Regina, A. Coluccia, V. Famiglini, S. Pelliccia, L. Monti, D. Vullo, E. Nuti, V. Alterio, G. De Simone, S. M. Monti, P. Pan, S. Parkkila, C. T. Supuran, A. Rossello and R. Silvestri, J. Med. Chem., 2015, 58, 8564–8572 CrossRef CAS PubMed.
- A. Bonardi, A. Nocentini, S. Bua, J. Combs, C. Lomelino, J. Andring, L. Lucarini, S. Sgambellone, E. Masini, R. McKenna, P. Gratteri and C. T. Supuran, J. Med. Chem., 2020, 63, 7422–7444 CrossRef CAS PubMed.
- P. Gopinath and M. K. Kathiravan, RSC Adv., 2021, 11, 38079–38093 RSC.
- A. Nocentini, W. A. Donald and C. T. Supuran, Carbonic Anhydrases, 2019, 151–185 CrossRef CAS.
- P. W. Taylor, R. W. King and A. S. V. Burgen, Biochemistry, 1970, 9, 2638–2645 CrossRef CAS PubMed.
- R. W. King and A. S. V. Burgen, Proc. R. Soc. London, Ser. B, 1976, 193, 107–125 CAS.
- A. E. Eriksson, T. A. Jones and A. Liljas, Proteins: Struct., Funct., Bioinf., 1988, 4, 274–282 CrossRef CAS PubMed.
- K. A. Rossi, K. M. Merz Jr., G. M. Smith and J. J. Baldwin, J. Med. Chem., 1995, 38, 2061–2069 CrossRef CAS PubMed.
- J. Gao, S. Qiao and G. M. Whitesides, J. Med. Chem., 1995, 38, 2292–2301 CrossRef CAS PubMed.
- D. Lu and G. A. Voth, Proteins: Struct., Funct., Bioinf., 1998, 33, 119–134 CrossRef CAS.
- C. Genis, K. H. Sippel, N. Case, W. Cao, B. S. Avvaru, L. J. Tartaglia, L. Govindasamy, C. Tu, M. Agbandje-McKenna, D. N. Silverman, C. J. Rosser and R. McKenna, Biochemistry, 2009, 48, 1322–1331 CrossRef CAS PubMed.
- B. S. Avvaru, J. M. Wagner, A. Maresca, A. Scozzafava, A. H. Robbins, C. T. Supuran and R. McKenna, Bioorg. Med. Chem. Lett., 2010, 20, 4376–4381 CrossRef CAS PubMed.
- F. Pacchiano, M. Aggarwal, B. S. Avvaru, A. H. Robbins, A. Scozzafava, R. McKenna and C. T. Supuran, Chem. Commun., 2010, 46, 8371–8373 RSC.
- N. Hen, M. Bialer, B. Yagen, A. Maresca, M. Aggarwal, A. H. Robbins, R. McKenna, A. Scozzafava and C. T. Supuran, J. Med. Chem., 2011, 54, 3977–3981 CrossRef CAS PubMed.
- J. Mecinović, P. W. Snyder, K. A. Mirica, S. Bai, E. T. Mack, R. L. Kwant, D. T. Moustakas, A. Héroux and G. M. Whitesides, J. Am. Chem. Soc., 2011, 133, 14017–14026 CrossRef PubMed.
- P. W. Snyder, J. Mecinović, D. T. Moustakas, S. W. Thomas, M. Harder, E. T. Mack, M. R. Lockett, A. Héroux, W. Sherman and G. M. Whitesides, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 17889–17894 CrossRef CAS PubMed.
- M. Lopez, H. Vu, C. K. Wang, M. G. Wolf, G. Groenhof, A. Innocenti, C. T. Supuran and S.-A. Poulsen, J. Am. Chem. Soc., 2011, 133, 18452–18462 CrossRef CAS PubMed.
- C. R. W. Guimarães, J. Chem. Theory Comput., 2011, 7, 2296–2306 CrossRef PubMed.
- M. Schmid, E. S. Nogueira, F. W. Monnard, T. R. Ward and M. Meuwly, Chem. Sci., 2012, 3, 690–700 RSC.
- R. Gaspari, C. Rechlin, A. Heine, G. Bottegoni, W. Rocchia, D. Schwarz, J. Bomke, H.-D. Gerber, G. Klebe and A. Cavalli, J. Med. Chem., 2016, 59, 4245–4256 CrossRef CAS PubMed.
- V. Chahal, S. Nirwan and R. Kakkar, Biophys. Chem., 2020, 265, 106439 CrossRef CAS PubMed.
- A. Tinivella, L. Pinzi and G. Rastelli, J. Cheminf., 2021, 13, 18 CAS.
- Y. Jiang, C. T. Supuran and J. Ho, J. Phys. Chem. A, 2022, 126, 9207–9217 CrossRef CAS PubMed.
- C. B. O'Herin, Y. W. Moriuchi, T. A. Bemis, A. J. Kohlbrand, M. D. Burkart and S. M. Cohen, J. Med. Chem., 2023, 66, 2789–2803 CrossRef PubMed.
- M. Taylor and J. Ho, J. Comput.-Aided Mol. Des., 2023, 37, 167–182 CrossRef CAS PubMed.
- M. Taylor, H. Mun and J. Ho, J. Phys. Chem. B, 2025, 129, 1475–1485 CrossRef CAS PubMed.
- L. T. Nguyen, M. Attarroshan, A. J. Cutright, S. L. Stokes, J. P. Emerson and S. R. Gwaltney, Polyhedron, 2025, 280, 117683 CrossRef CAS.
- S. Decherchi and A. Cavalli, Chem. Rev., 2020, 120, 12788–12833 CrossRef CAS PubMed.
- W. L. Jorgensen and L. L. Thomas, J. Chem. Theory Comput., 2008, 4, 869–876 CrossRef CAS PubMed.
- H. M. Senn and W. Thiel, Angew. Chem., Int. Ed., 2009, 48, 1198–1229 CrossRef CAS PubMed.
- E. Wang, H. Sun, J. Wang, Z. Wang, H. Liu, J. Z. H. Zhang and T. Hou, Chem. Rev., 2019, 119, 9478–9508 CrossRef CAS PubMed.
- J. Mieres-Perez, Y. Almeida-Hernandez, W. Sander and E. Sanchez-Garcia, Chem. Rev., 2025, 125, 7023–7056 CrossRef CAS PubMed.
- A. Ghidini, E. Serra and A. Cavalli, Acc. Chem. Res., 2025, 58, 3137–3145 CrossRef CAS PubMed.
- I. Massova and P. A. Kollman, Perspect. Drug Discovery Des., 2000, 18, 113–135 CrossRef CAS.
- S. Genheden and U. Ryde, Expert Opin. Drug Discovery, 2015, 10, 449–461 CrossRef CAS PubMed.
- D. Santos-Martins, S. Forli, M. J. A. Ramos and A. J. Olson, J. Chem. Inf. Model., 2014, 54, 2371–2379 CrossRef CAS PubMed.
- A. D. Becke, J. Chem. Phys., 1993, 98, 5648–5652 CrossRef CAS.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- R. W. Zwanzig, J. Chem. Phys., 1954, 22, 1420–1426 CrossRef CAS.
- N. J. Bruce, G. K. Ganotra, D. B. Kokh, S. K. Sadiq and R. C. Wade, Curr. Opin. Struct. Biol., 2018, 49, 1–10 CrossRef CAS PubMed.
- N. Ahalawat and J. Mondal, J. Phys. Chem. Lett., 2021, 12, 633–641 CrossRef CAS PubMed.
- E. Maximova, E. B. Postnikov, A. I. Lavrova, V. Farafonov and D. Nerukh, J. Phys. Chem. Lett., 2021, 12, 10631–10636 CrossRef CAS PubMed.
- P. Chaskar, V. Zoete and U. F. Röhrig, J. Chem. Inf. Model., 2017, 57, 73–84 CrossRef CAS PubMed.
- N. T. Nguyen, T. H. Nguyen, T. N. H. Pham, N. T. Huy, M. V. Bay, M. Q. Pham, P. C. Nam, V. V. Vu and S. T. Ngo, J. Chem. Inf. Model., 2020, 60, 204–211 CrossRef CAS PubMed.
- A. L. Klinger, D. F. McComsey, V. Smith-Swintosky, R. P. Shank and B. E. Maryanoff, J. Med. Chem., 2006, 49, 3496–3500 CrossRef CAS PubMed.
- Y. Yu, L. M. Sternicki, D. H. Hilko, R. J. Jarrott, G. Di Trapani, K. F. Tonissen and S.-A. Poulsen, J. Med. Chem., 2024, 67, 15862–15872 CrossRef CAS PubMed.
- V. Paketurytė-Latvė, A. Smirnov, E. Manakova, L. Baranauskiene, V. Petrauskas, A. Zubrienė, J. Matulienė, V. Dudutienė, E. Čapkauskaitė, A. Zakšauskas, J. Leitans, S. Gražulis, K. Tars and D. Matulis, IUCrJ, 2024, 11, 556–569 CrossRef PubMed.
- P. W. Taylor, R. W. King and A. S. V. Burgen, Biochemistry, 1970, 9, 3894–3902 CrossRef CAS PubMed.
- L. L. Kiefer, S. A. Paterno and C. A. Fierke, J. Am. Chem. Soc., 1995, 117, 6831–6837 CrossRef CAS.
- C. Bannwarth, S. Ehlert and S. Grimme, J. Chem. Theory Comput., 2019, 15, 1652–1671 CrossRef CAS PubMed.
- J. Huang and A. D. MacKerell Jr, J. Comput. Chem., 2013, 34, 2135–2145 CrossRef CAS PubMed.
- P. A. Boriack, D. W. Christianson, J. Kingery-Wood and G. M. Whitesides, J. Med. Chem., 1995, 38, 2286–2291 CrossRef CAS PubMed.
- P. Li and K. M. Merz Jr., J. Chem. Inf. Model., 2016, 56, 599–604 CrossRef CAS PubMed.
- T. K. Piskorz, B. Lee, S. Zhan and F. Duarte, J. Chem. Theory Comput., 2024, 20, 9060–9071 CrossRef CAS PubMed.
- G. Cavallo, P. Metrangolo, R. Milani, T. Pilati, A. Priimagi, G. Resnati and G. Terraneo, Chem. Rev., 2016, 116, 2478–2601 CrossRef CAS PubMed.
- J. Eberhardt, D. Santos-Martins, A. F. Tillack and S. Forli, J. Chem. Inf. Model., 2021, 61, 3891–3898 CrossRef CAS PubMed.
- G. M. Morris, R. Huey, W. Lindstrom, M. F. Sanner, R. K. Belew, D. S. Goodsell and A. J. Olson, J. Comput. Chem., 2009, 30, 2785–2791 CrossRef CAS PubMed.
- G. Jones, P. Willett, R. C. Glen, A. R. Leach and R. Taylor, J. Mol. Biol., 1997, 267, 727–748 CrossRef CAS PubMed.
- C. M. Breneman and K. B. Wiberg, J. Comput. Chem., 1990, 11, 361–373 CrossRef CAS.
- F. Weigend and R. Ahlrichs, Phys. Chem. Chem. Phys., 2005, 7, 3297–3305 RSC.
- F. Neese, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2012, 2, 73–78 CAS.
- J. Lee, X. Cheng, J. M. Swails, M. S. Yeom, P. K. Eastman, J. A. Lemkul, S. Wei, J. Buckner, J. C. Jeong and Y. Qi, et al. , J. Chem. Theory Comput., 2016, 12, 405–413 CrossRef CAS PubMed.
- G. Bussi, A. Laio and M. Parrinello, Phys. Rev. Lett., 2006, 96, 090601 CrossRef PubMed.
- K. Vanommeslaeghe, E. Hatcher, C. Acharya, S. Kundu, S. Zhong, J. Shim, E. Darian, O. Guvench, P. Lopes, I. Vorobyov and A. D. Mackerell Jr., J. Comput. Chem., 2010, 31, 671–690 CrossRef CAS PubMed.
- W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. W. Impey and M. L. Klein, J. Chem. Phys., 1983, 79, 926–935 CrossRef CAS.
- D. A. Case, H. M. Aktulga, K. Belfon, D. S. Cerutti, G. A. Cisneros, V. W. D. Cruzeiro, N. Forouzesh, T. J. Giese, A. W. Götz, H. Gohlke, S. Izadi, K. Kasavajhala, M. C. Kaymak, E. King, T. Kurtzman, T.-S. Lee, P. Li, J. Liu, T. Luchko, R. Luo, M. Manathunga, M. R. Machado, H. M. Nguyen, K. A. O’Hearn, A. V. Onufriev, F. Pan, S. Pantano, R. Qi, A. Rahnamoun, A. Risheh, S. Schott-Verdugo, A. Shajan, J. Swails, J. Wang, H. Wei, X. Wu, Y. Wu, S. Zhang, S. Zhao, Q. Zhu, T. E. I. Cheatham, D. R. Roe, A. Roitberg, C. Simmerling, D. M. York, M. C. Nagan and K. M. Merz Jr., J. Chem. Inf. Model., 2023, 63, 6183–6191 CrossRef CAS PubMed.
- P. C. Hariharan and J. A. Pople, Theor. Chim. Acta, 1973, 28, 213–222 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery, Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16 Revision B.01, Gaussian Inc., Wallingford CT, 2016 Search PubMed.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graphics, 1996, 14, 33–38 CrossRef CAS PubMed.
- T. Saito and Y. Takano, J. Phys. Chem. B, 2022, 126, 2087–2097 CrossRef CAS PubMed.
- S. Suenaga, Y. Takano and T. Saito, Molecules, 2023, 28, 2697 CrossRef CAS PubMed.
- M. T. Nelson, W. Humphrey, A. Gursoy, A. Dalke, L. V. Kalé, R. D. Skeel and K. Schulten, Int. J. High Perform. Comput. Appl., 1996, 10, 251–268 Search PubMed.
- J. C. Phillips, D. J. Hardy, J. D. C. Maia, J. E. Stone, J. A. V. Ribeiro, R. C. Bernardi, R. Buch, G. Fiorin, J. Hénin, W. Jiang, R. McGreevy, M. C. R. Melo, B. K. Radak, R. D. Skeel, A. Singharoy, Y. Wang, B. Roux, A. Aksimentiev, Z. Luthey-Schulten, L. V. Kalé, K. Schulten, C. Chipot and E. Tajkhorshid, J. Chem. Phys., 2020, 153, 044130 CrossRef CAS PubMed.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- F. Neese, J. Comput. Chem., 2003, 24, 1740–1747 CrossRef CAS PubMed.
- F. Neese, F. Wennmohs, A. Hansen and U. Becker, Chem. Phys., 2009, 356, 98–109 CrossRef CAS.
- M. C. R. Melo, R. C. Bernardi, T. Rudack, M. Scheurer, C. Riplinger, J. C. Phillips, J. D. Maia, G. B. Rocha, J. V. Ribeiro, J. E. Stone, F. Neese, K. Schulten and Z. Luthey-Schulten, Nat. Methods, 2018, 15, 351–354 CrossRef CAS PubMed.
- Q. Bai, S. Tan, T. Xu, H. Liu, J. Huang and X. Yao, Briefings Bioinf., 2020, 22, bbaa161 CrossRef PubMed.
- S. Genheden, P. Mikulskis, L. Hu, J. Kongsted, P. Söderhjelm and U. Ryde, J. Am. Chem. Soc., 2011, 133, 13081–13092 CrossRef CAS PubMed.
- S. Genheden, I. Nilsson and U. Ryde, J. Chem. Inf. Model., 2011, 51, 947–958 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2026 |