 Open Access Article
Open Access ArticlePyMESpec: a Python toolbox for automated modulation excitation spectroscopic data analysis and transient experiments
Alfred
Worrad
ab,
Quentin
Kim
ab,
Sagar
Sourav
cd,
Himaghna
Bhattacharjee
ab and
Dionisios G.
Vlachos
 *abc
*abc
aDepartment of Chemical and Biomolecular Engineering, University of Delaware, Newark, DE 19716, USA. E-mail: vlachos@udel.edu
bCenter for Catalytic Science and Technology, University of Delaware, Newark, DE 19716, USA
cDelaware Energy Institute, University of Delaware, 221 Academy St., Newark, DE 19716, USA
dDepartment of Chemical Engineering, Indian Institute of Technology Madras, Chennai, Tamil Nadu 600036, India
First published on 14th January 2026
Abstract
Recent advances in laboratory automation have created a growing need for efficient and scalable tools to process experimental data without human intervention. Of great interest to the catalysis community is the analysis of time-resolved spectroscopic data, which can capture transients and identify highly active minority species. One such technique is modulation excitation spectroscopy (MES). Modulation produces large datasets unsuitable for manual processing. We introduce the python modulation excitation spectroscopy (PyMESpec) toolkit, an open-source library for analyzing MES experiments. PyMESpec offers a fast and flexible baseline correction, phase-sensitive detection (PSD), chemometric deconvolution, and automated reaction rate extraction. PyMESpec can be used through a command-line interface with configuration files or via a graphical user interface (GUI). This facilitates high-throughput, reproducible processing of spectroscopic datasets in automated and adaptive experimentation. PyMESpec applies to large datasets from spectroscopies and transient experiments in general. We demonstrate it using spectra from diffuse reflectance infrared Fourier transform spectroscopy (DRIFTS) over CeO2 and modulation excitation ultraviolet-visible (ME UV-vis) and near-ambient pressure x-ray photoemission spectroscopy (NAP-XPS) on vanadia/titania catalyst for oxidative propane dehydrogenation.
1. Introduction
Automated laboratory experiments, often referred to as the self-driving lab, are increasingly sought after as they enable efficient use of equipment and higher data throughput handling compared to individual researchers manually handling data and tasks. They improve productivity, accelerate discovery, and reduce human error in routine tasks.1–4 Automation also brings significant challenges, including the planning of sequential experiments to explore the parameter space efficiently, data processing, and data interpretation.3,5 Software toolkits, such as NEXTorch,6 have been developed to design experiments and guide catalyst optimization using Bayesian optimization. While human in the loop is essential for high-level decision-making, creating data analysis pipelines for self-driving labs is a necessity, but often complex, potentially non-uniform, and situationally dependent.One challenge in heterogeneous catalysis research is that the reactive sites often exist in small populations that are hard to distinguish from less catalytically active, abundant species.7,8 Advanced spectroscopic techniques and data analysis tools can discern subtle signals, and with transient perturbations, identify catalytically active sites and elucidate reaction mechanisms in situ or under operando conditions.
Modulation excitation spectroscopy (MES) is a powerful and sensitive technique that enhances the signal-to-noise ratio, enabling kinetic and mechanistic analysis.9–11 Through periodic modulations, it allows phase-sensitive detection to isolate dynamic spectral responses from static background signals and noise. Temperature, pressure, pH, isotopic labels, electrical potential, and light flux, are often modulated8 and data is collected using a host of spectroscopic techniques, such as ATR,7,12 DRIFTS,13–16 Raman,17 XRD,18 and UV-vis.15,17 Time-resolved techniques,9 such as temporal analysis of products (TAP)19 and steady-state isotopic transient kinetic analysis (SSITKA),20 also leverage controlled transients to extract mechanistic insights by generating large datasets.
These techniques generate large datasets of hundreds to thousands of spectra. Processing such datasets manually is time-consuming and prone to subjective bias and inconsistent interpretation. Automated data analysis software, capable of processing large datasets, could overcome this challenge and provide real-time feedback for adaptive experimentation and expedited exploration. Orange-spectroscopy, an extension of the orange data mining suite,21 can perform preprocessing and visualization of spectroscopic datasets but is not tailored for phase-sensitive detection and kinetic rate extraction in MES experiments.
In this work, we introduce the python modulation excitation spectroscopy (PyMESpec), a python-based package for automating data analysis of MES and transient spectroscopic experiments. PyMESpec facilitates the processing of large-scale spectroscopic datasets by systematic baseline correction, noise filtering, phase-sensitive analysis, chemometric deconvolution, and kinetic rate extraction with minimal user oversight. It can increase throughput and reproducibility, enabling autonomous transient experiments in self-driving laboratories. We demonstrate its use with experimental data of CeO2 diffuse reflectance infrared Fourier transform spectroscopy (DRIFTS) and modulation excitation ultraviolet-visible (ME UV-vis) and near-ambient pressure X-ray photoemission spectroscopy (NAP-XPS) on vanadia/titania catalyst for oxidative propane dehydrogenation.
2. Methods
2.1. Case study tutorials
To demonstrate the capabilities of the PyMESpec suite, we present two experimental datasets involving CeO2 diffuse reflectance infrared Fourier transform spectroscopy (DRIFTS) and modulation excitation ultraviolet-visible (ME UV-vis) and near-ambient pressure X-ray photoemission spectroscopy (NAP-XPS) on vanadia/titania catalyst for oxidative propane dehydrogenation. These examples demonstrate the toolbox's ability to process large, time-resolved spectroscopic data to extract chemical insights via automated workflows. A flow diagram showcasing a standard subset of possible workflows is shown in Fig. S1. All data processing performed within this work can be found as part of a series of Jupyter notebook tutorials on the Github page https://github.com/worradal/PyMESpec under the src/example_files/jupyter_notebook_tutorials/folder. PyMESpec natively supports CSV files across all package operations; however, custom data processing classes can be implemented to handle any file type (if converting to CSV is undesirable). Example screenshots showcasing the GUI layout can be found in the SI, Fig. S3 and S4.ME-UV-vis spectra were collected for the propane (C3) oxidative dehydrogenation (ODH) reaction over this catalyst. An in-house-built low-void volume environmental cell,22 an Avantes ULS2048CL-EVO UV-vis spectrometer equipped with an AvaLight-XE light source, and an adjustable UV/VIS/NIR high-temperature probe were used for data collection. BaSO4 was used as a reference for the background spectrum of the sample absorbance spectra. The 4V/TiO2 sample was calcined in 10% O2/Ar for 30 min at 400 °C. Following this, the sample was conditioned in the C3 ODH reaction mixture (C3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) O2
O2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ar, 1
Ar, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8) for 1 h to achieve a stable structure for the reaction. Following this, O2 was switched periodically with He with a full cycle period of 60 s, where O2 was admitted into the environmental cell for 30 s and replaced by He for 30 s. The data was collected at 100 ms per spectrum with averaging over 10 spectra (1 s per spectrum), leading to 60 spectra per cycle. Spectra were collected for a total of 10 periods.
8) for 1 h to achieve a stable structure for the reaction. Following this, O2 was switched periodically with He with a full cycle period of 60 s, where O2 was admitted into the environmental cell for 30 s and replaced by He for 30 s. The data was collected at 100 ms per spectrum with averaging over 10 spectra (1 s per spectrum), leading to 60 spectra per cycle. Spectra were collected for a total of 10 periods.
Near ambient pressure X-ray photoelectron spectroscopy (NAP-XPS) spectra were collected at Lehigh University using a SPECS DeviSim NAP reactor cell coupled with a PHOIBOS 150 NAP electron energy analyzer equipped with an Al Kα source. 4V/TiO2 was heated in 1 mbar of O2 to 400 °C before cooling down to 130 °C and pulling vacuum. 1 mbar of ethanol was then introduced to the chamber for 140 min. Spectra of the O 1s and V 2p XPS region (540–505 eV) were taken every 14 minutes during the reduction by ethanol. The NAP-XPS data was analyzed using CasaXPS23 and used to compute pure component UV-vis spectra for V5+, V4+ and V3+ species, for chemometric deconvolution. A tutorial for obtaining the pure component UV-vis spectra from known fractions in XPS can be found in the Jupyter notebook tutorial “computing_pure_components_tutorial.ipynb.” Additional details on the matching of XPS and UV-vis can be found in Sections S1, S2, and in Fig. S2 of the SI.
2.2. MES data structure
We first outline the structure of the data. An experimental variable, such as a reactant concentration, is periodically modulated, commonly via a step waveform, and the system's response is monitored using a spectroscopic technique sensitive to chemical or structural changes. The time-resolved spectroscopic data is a signal I(t,ν), where t is the time of collection and ν is the spectral coordinate within each spectrum (e.g., wavenumber, frequency, or energy). The I(ti,νj) data forms a matrix where each row corresponds to a spectrum collected at time ti, and each column corresponds to the spectral coordinate νj, with each element containing the signal I(ti,νj); a pictorial representation is shown in Fig. 1. This data structure allows automated analysis in PyMESpec. | ||
| Fig. 1 Data structure I(t,ν) of MES experiments, with each row representing a spectrum collected at a point in time, and each column the temporal progression for a select spectral coordinate value. | ||
3. Results and discussion
3.1. Baseline correction
Nearly all spectroscopic techniques suffer from background fluctuations due to environmental, instrumental, or sample effects.24 Our library includes a comprehensive baseline correction module, allowing the use of standard correction methods in the form of linear or polynomial baseline subtraction for simple baseline drifts. We also implement asymmetric reweighted penalized least squares (arPLS)25 to effectively preserve true spectral features, even for more complex baseline corrections. Correction methods are tunable through PyMESpec package scripting, with simplified versions available in GUI. The baseline correction module is modular and extensible. Customized correction methods can be implemented by defining classes that specify how the correction is applied to a 1D spectrum.Baseline correction schemes can be applied to individual spectra, full I(t,ν) matrices, or select ranges of the spectral coordinate. We showcase the arPLS method, which corrects for baseline drift in a set of CeO2 FTIR spectra at different temperatures in Fig. 2A. In the original FTIR data, a temperature-dependent baseline drift is present that we wish to remove. We highlight the extracted baseline in relation to the original and baseline corrected version of a single spectrum in Fig. 2B for clear emphasis on the change in the spectrum. Upon applying arPLS baseline correction, the change in the hydroxyl region (∼3000–3800 cm−1) is more clearly visible and is less subject to thermal noise. For convenience and reproducibility, the exact code used for this baseline correction can be found in the example tutorial Jupyter notebook “CeO2_DRIFTS_tutorial.ipynb” on the GitHub page, which walks the user through the impact that adjusting parameters has on the shape of the baseline. For an example tutorial on how to define a custom baseline correction class and implemented it within a workflow, we would like to direct the reader to the tutorial located in “CeO2_DRIFTS_custom_baseline_tutorial.ipynb.”
3.2. MES phase-sensitive detection (PSD)
The first major MES-specific feature of the PyMESpec package is PSD, which isolates signals that oscillate at a chosen modulation frequency at different phase offsets from the experimental modulation. PSD can be performed either on an average modulation cycle or the full experimental time series, depending on the signal quality and analysis objectives. The average-cycle approach, which is a common practice but not a strict requirement, involves averaging individual modulation periods to form a representative cycle. The conventional formulation of PSD7 is provided in eqn (1). | (1) |
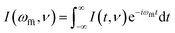 | (2) |
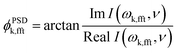 | (3) |
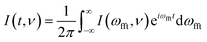 | (4) |
Since PSD commonly uses k = 1 for the demodulation index (the fundamental frequency), we employ this for our vanadia on titania case study tutorial. The supported 4V/TiO2 MES experiment conducted at 400 °C switching on and off O2, while keeping propane flow constant, is shown in Fig. 3. For a direct comparison between the time index (Fig. 3A) and the phase angle (Fig. 3B) in this example, use 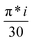 , where i is the time index.
, where i is the time index.
The static sections of the spectra are removed, isolating only the contributions from active species. The region above 800 nm in the original signal, dominated by noise, is also smoothed considerably. The phase-sensitive plot indicates that regions around 400 nm and 600 nm appear ∼180 degrees out of phase with each other. This is consistent with the regions ∼425 nm and ∼550–650 nm being associated with the oxidized (V5+) and reduced (V4+ and V3+) species, respectively. This out-of-phase behavior was severely obscured in the original spectra, whereas upon PSD, the reduction and reoxidation cycles between the oxidized and reduced species are clearly seen. For additional considerations on the kinetic analysis of MES-PSD, we refer the readers to the comprehensive works by Bravo and co-workers.14,27 A tutorial Jupyter notebook is available for the analysis of V/TiO2, with the PSD section in the first part of the “V_TiO2_tutorial.ipynb.” We note that, for other non-MES applications, general noise filtering methods utilizing Savitzky-Golay28 and low-pass filtering are also implemented.
3.3. Chemometric analysis
Another prominent feature of our software package is the chemometric analysis module. Under the assumption that linear mixing applies, we can quantify the relative contributions of chemical species or catalytic sites. This is achieved using a library of reference spectra for each pure component. At each time step, the measured spectrum is deconvoluted using non-negative linear least squares (NNLS)29 to ensure physically meaningful (i.e., non-negative) contributions from each reference. The deconvolution solves the optimization problem given in eqn (5):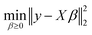 | (5) |
 | (6) |
![[small sigma, Greek, circumflex]](https://www.rsc.org/images/entities/i_char_e111.gif) 2 is the estimated error of the residuals, and the ii refers to the i-th diagonal element of the covariance matrix (XTX)−1. To enforce the non-negativity constraint on the resulting confidence intervals, we bound the lower limit at zero. Additionally, when interpreting results in terms of fractional species contributions, we cap the upper bound at 1 to reflect a closed composition space. This chemometric approach is applied to the same 4V/TiO2 system used in the PSD analysis. The results are illustrated in Fig. 4.
2 is the estimated error of the residuals, and the ii refers to the i-th diagonal element of the covariance matrix (XTX)−1. To enforce the non-negativity constraint on the resulting confidence intervals, we bound the lower limit at zero. Additionally, when interpreting results in terms of fractional species contributions, we cap the upper bound at 1 to reflect a closed composition space. This chemometric approach is applied to the same 4V/TiO2 system used in the PSD analysis. The results are illustrated in Fig. 4.
Through chemometrics analysis, we obtain fractions for vanadium in each oxidation state, allowing us to track the working state of the catalyst over time. In this example, the fraction of V5+ decreases as it reduces to V4+ and V3+ species during propane flow in the absence of O2. When O2 is switched on, the reduced species quickly reoxidize to V5+. The system does not completely reoxidize to 100% V5+, as there exists a baseline of reduced vanadium species under O2 and propane flow. In previous computational work, we provided evidence that a small set of highly active V5+ isolated species may form30 that reduce faster than most other V5+ species, which may explain why the working state of the catalyst is not being fully oxidized. The chemometrics analysis is available in the tutorial notebook “V_TiO2_tutorial.ipynb.”
3.4. Automated rate analysis
Another key feature of our software package is the automated extraction of kinetic rates. Rate analysis can be performed either on a cycle-averaged pulse, which enhances the signal-to-noise ratio, or on individual pulses, enabling the user to track trends over multiple pulses, such as in catalyst deactivation. Numerous commercial instrument packages include basic rate estimation tools; they are typically designed for small datasets, require user interaction to select data points, or are not easily incorporated into modular workflows. Our implementation is focused on large datasets requiring minimal human intervention. Additionally, our package can compute rates directly from the raw spectra or from the species concentrations obtained from the chemometric analysis shown in the previous section.Regardless of the type of data selected, for rate extraction on a dataset containing multiple pulses, it will first identify the maxima and minima of the I(t,ν) signal (or concentration) using SciPy's find_peaks method.31 The maxima and minima are then used as the start and end points for each kinetic window. To extract rates from each kinetic window, a kinetic equation chosen by the user is fit to the data, with an exponential fit being the default. Users can implement their own fitting function beyond the exponential function, most commonly used for adsorption, desorption, or redox processes. Optionally, for increased accuracy, the initial peak detection can be followed by adaptive interval refinement, in which the fitting boundaries are adjusted iteratively to minimize the root mean square error (RMSE) of the fitted model.
While the PyMESpec comes with a built-in peak detection scheme, the framework is modular and can accommodate custom peak detection algorithms, including machine learning approaches such as convolutional neural networks (CNNs), for handling complex or low-signal-to-noise-ratio systems. We apply the automated rate calculation to the 4V/TiO2 system to extract the redox rates for the V5+ species in Fig. 5.
 | ||
| Fig. 5 Automated peak detection and rate calculation for each pulse in the MES experiment based on the V5+ species as calculated from the populations computed in the chemometrics section (Fig. 4). (A) Fit pulsed rates from absolute maxima and minima before adaptive peak finding, and (B) sections and calculated rates from adaptive peak finding. | ||
This automat ed rate detection scheme can compute the reduction and reoxidation rates based on the chemometric deconvolutions for the 4V/TiO2 system. Our adaptive fitting scheme drastically improves the rate-fitting and detection of regions with a change in conditions. The reoxidation process is faster than the reduction process by approximately an order of magnitude. This information is unattainable in steady state experiments, further illustrating the importance of transient experiments. While assigning the start and end points directly to the times when a modulation occurs may seem trivial, this is not feasible in sequential reactions (A → B → C) when the delay of intermediate species is not explicitly known. This analysis is available in the tutorial notebook “V_TiO2_tutorial.ipynb.”
4. Conclusions
High-throughput data analysis workflows are required for spectral analysis of highly complex transient spectral techniques. We have outlined PyMESpec, a flexible data analysis software package and a pipeline that requires minimal human oversight. In its most complete form, it takes raw transient spectra, employs sophisticated baseline correction algorithms to get comparable spectra, which, using phase-sensitive detection, can filter for signals of interest. Additionally, deconvolutions are applied to obtain active species balances based on reference spectra enabling the calculations of reaction rates. These individual components can be flexibly utilized for individual spectrum purposes as well.Author contributions
A. W. wrote the PyMESpec software. Q. K. and S. S. conducted experiments. H. B. helped with initial code development. D. G. V. provided supervision. A. W. wrote the manuscript with input from S. S. and D. G. V. All authors contributed to the analysis and discussion of the results.Conflicts of interest
There are no conflicts of interest to declare.Data availability
PyMESpec is freely available on its GitHub project page (https://github.com/worradal/PyMESpec) along with detailed user documentation, Jupyter notebook tutorials for the python package, and example configuration files for the GUI and command line implementation. Details can be found on the GitHub page.Supplementary information is available. See DOI: https://doi.org/10.1039/d5cp04322k.
Acknowledgements
This work was supported by the U.S. Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences, under Award Number DE-SC0024085. The authors would like to thank Dr Ryan Thorpe for his assistance with XPS experiments performed at the Institute for Functional Materials and Devices at Lehigh University and for his help with the analysis. We acknowledge the use of Visual Studio Code for the expedited writing of unit tests and documentation.References
- C. Scheurer and K. Reuter, Nat. Catal., 2025, 8, 13–19 CrossRef CAS.
- Y. Xie, K. Sattari, C. Zhang and J. Lin, Prog. Mater. Sci., 2023, 132, 101043 CrossRef.
- M. Seifrid, R. Pollice, A. Aguilar-Granda, Z. Morgan Chan, K. Hotta, C. T. Ser, J. Vestfrid, T. C. Wu and A. Aspuru-Guzik, Acc. Chem. Res., 2022, 55, 2454–2466 CrossRef CAS PubMed.
- Y. Shi, P. L. Prieto, T. Zepel, S. Grunert and J. E. Hein, Acc. Chem. Res., 2021, 54, 546–555 CrossRef CAS PubMed.
- A. Trunschke, Catal. Sci. Technol., 2022, 12, 3650–3669 RSC.
- Y. Wang, T.-Y. Chen and D. G. Vlachos, J. Chem. Inf. Model., 2021, 61, 5312–5319 CrossRef CAS PubMed.
- A. Urakawa, T. Bürgi and A. Baiker, Chem. Eng. Sci., 2008, 63, 4902–4909 CrossRef CAS.
- C. Hess, ACS Catal., 2025, 15, 8005–8013 CrossRef.
- P. Müller and I. Hermans, Ind. Eng. Chem. Res., 2017, 56, 1123–1136 CrossRef.
- D. Baurecht and U. P. Fringeli, Rev. Sci. Instrum., 2001, 72, 3782–3792 CrossRef CAS.
- M. Chen, N. Maeda, A. Baiker and J. Huang, ACS Catal., 2012, 2, 2007–2013 CrossRef CAS.
- A. Urakawa, R. Wirz, T. Bürgi and A. Baiker, J. Phys. Chem. B, 2003, 107, 13061–13068 CrossRef CAS.
- J. Weyel, L. Schumacher, M. Ziemba, M. Pfeiffer and C. Hess, Acc. Chem. Res., 2024, 57, 2643–2652 CrossRef CAS PubMed.
- P. D. Srinivasan, B. S. Patil, H. Zhu and J. J. Bravo-Suárez, React. Chem. Eng., 2019, 4, 862–883 RSC.
- K. Yu, M. Scarpelli, S. Sourav, A. Worrad, J. A. Boscoboinik, L. Ma, S. N. Ehrlich, N. Marinkovic, W. Zheng and D. G. Vlachos, Catal. Sci. Technol., 2024, 14, 4948–4957 RSC.
- I. C. t Have, J. J. G. Kromwijk, M. Monai, D. Ferri, E. B. Sterk, F. Meirer and B. M. Weckhuysen, Nat. Commun., 2022, 13, 324 CrossRef CAS PubMed.
- H. Hoyer and C. Hess, J. Phys. Chem. C, 2024, 128, 21377–21386 CrossRef CAS.
- D. Ferri, M. A. Newton, M. Di Michiel, G. L. Chiarello, S. Yoon, Y. Lu and J. Andrieux, Angew. Chem., Int. Ed., 2014, 53, 8890–8894 CrossRef CAS PubMed.
- K. Morgan, N. Maguire, R. Fushimi, J. T. Gleaves, A. Goguet, M. P. Harold, E. V. Kondratenko, U. Menon, Y. Schuurman and G. S. Yablonsky, Catal. Sci. Technol., 2017, 7, 2416–2439 RSC.
- C. Ledesma, J. Yang, D. Chen and A. Holmen, ACS Catal., 2014, 4, 4527–4547 CrossRef CAS.
- J. Demšar, T. Curk, A. Erjavec, C. Gorup, T. Hočevar, M. Milutinovič, M. Možina, M. Polajnar, M. Toplak, A. Starič, M. Štajdohar, L. Umek, L. Žagar, J. Žbontar, M. Žitnik and B. Zupan, J. Mach. Learn. Res., 2013, 14, 2349–2353 Search PubMed.
- B. S. Patil, P. D. Srinivasan, E. Atchison, H. Zhu and J. J. Bravo-Suárez, React. Chem. Eng., 2019, 4, 667–678 RSC.
- N. Fairley, V. Fernandez, M. Richard-Plouet, C. Guillot-Deudon, J. Walton, E. Smith, D. Flahaut, M. Greiner, M. Biesinger, S. Tougaard, D. Morgan and J. Baltrusaitis, Appl. Surf. Sci. Adv., 2021, 5, 100112 CrossRef.
- C. R. Mittermayr, H. W. Tan and S. D. Brown, Appl. Spectrosc., 2001, 55, 827–833 CrossRef CAS.
- S.-J. Baek, A. Park, Y.-J. Ahn and J. Choo, Analyst, 2015, 140, 250–257 RSC.
- C. R. Harris, K. J. Millman, S. J. van der Walt, R. Gommers, P. Virtanen, D. Cournapeau, E. Wieser, J. Taylor, S. Berg, N. J. Smith, R. Kern, M. Picus, S. Hoyer, M. H. van Kerkwijk, M. Brett, A. Haldane, J. F. del Río, M. Wiebe, P. Peterson, P. Gérard-Marchant, K. Sheppard, T. Reddy, W. Weckesser, H. Abbasi, C. Gohlke and T. E. Oliphant, Nature, 2020, 585, 357–362 CrossRef CAS PubMed.
- A. Torres-Velasco, B. S. Patil, H. Zhu, Y. Qi, S. G. Podkolzin and J. J. Bravo-Suárez, J. Catal., 2024, 434, 115508 CrossRef CAS.
- A. Savitzky and M. J. E. Golay, Anal. Chem., 1964, 36, 1627–1639 CrossRef CAS.
- C. L. Lawson and R. J. Hanson, Solving Least Squares Problems, Society for Industrial and Applied Mathematics, 1995.
- A. Worrad, S. A. Khan, D. G. Vlachos and S. Caratzoulas, J. Phys. Chem. C, 2024, 128, 18933–18941 CrossRef CAS.
- P. Virtanen, R. Gommers, T. E. Oliphant, M. Haberland, T. Reddy, D. Cournapeau, E. Burovski, P. Peterson, W. Weckesser, J. Bright, S. J. van der Walt, M. Brett, J. Wilson, K. J. Millman, N. Mayorov, A. R. J. Nelson, E. Jones, R. Kern, E. Larson, C. J. Carey, İ. Polat, Y. Feng, E. W. Moore, J. VanderPlas, D. Laxalde, J. Perktold, R. Cimrman, I. Henriksen, E. A. Quintero, C. R. Harris, A. M. Archibald, A. H. Ribeiro, F. Pedregosa, P. van Mulbregt, A. Vijaykumar, A. P. Bardelli, A. Rothberg, A. Hilboll, A. Kloeckner, A. Scopatz, A. Lee, A. Rokem, C. N. Woods, C. Fulton, C. Masson, C. Häggström, C. Fitzgerald, D. A. Nicholson, D. R. Hagen, D. V. Pasechnik, E. Olivetti, E. Martin, E. Wieser, F. Silva, F. Lenders, F. Wilhelm, G. Young, G. A. Price, G.-L. Ingold, G. E. Allen, G. R. Lee, H. Audren, I. Probst, J. P. Dietrich, J. Silterra, J. T. Webber, J. Slavič, J. Nothman, J. Buchner, J. Kulick, J. L. Schönberger, J. V. de Miranda Cardoso, J. Reimer, J. Harrington, J. L. C. Rodríguez, J. Nunez-Iglesias, J. Kuczynski, K. Tritz, M. Thoma, M. Newville, M. Kümmerer, M. Bolingbroke, M. Tartre, M. Pak, N. J. Smith, N. Nowaczyk, N. Shebanov, O. Pavlyk, P. A. Brodtkorb, P. Lee, R. T. McGibbon, R. Feldbauer, S. Lewis, S. Tygier, S. Sievert, S. Vigna, S. Peterson, S. More, T. Pudlik, T. Oshima, T. J. Pingel, T. P. Robitaille, T. Spura, T. R. Jones, T. Cera, T. Leslie, T. Zito, T. Krauss, U. Upadhyay, Y. O. Halchenko, Y. Vázquez-Baeza and C. SciPy, Nat. Methods, 2020, 17, 261–272 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2026 |




