 Open Access Article
Open Access ArticleMachine learning for accelerated prediction of size distributions of spherical nanoparticles from small-angle X-ray scattering
Qiaoyu
Guo†
 ,
Fei
Xie†
,
Zhe
Sun
,
Xuechen
Jiao
* and
Wancheng
Yu
,
Fei
Xie†
,
Zhe
Sun
,
Xuechen
Jiao
* and
Wancheng
Yu
 *
*
National Synchrotron Radiation Laboratory, University of Science and Technology of China, Hefei, Anhui 230029, China. E-mail: xjiao@ustc.edu.cn; ywcheng@ustc.edu.cn
First published on 3rd March 2026
Abstract
Small-angle X-ray scattering (SAXS) is widely used for characterizing the particle size distribution (PSD) at the mesoscale. Conventional extraction of PSD from SAXS data typically relies on traditional numerical methods such as Monte Carlo algorithms. However, these approaches often suffer from low computational efficiency and inherent difficulties in resolving complex multimodal distributions, thus limiting their applicability in high-throughput or real-time SAXS data analysis. To overcome these limitations, here we develop a feed-forward neural network (FNN) model for the accurate and efficient PSD analysis. By embedding physical constraints via iterative fine-tuning, the FNN model yields physically plausible predictions and resolves key PSD features accurately, including peak positions, peak widths, and low-abundance subpopulations for both simulated and experimental SAXS data. Validation against synchrotron radiation SAXS measurements and scanning electronic microscopy (SEM) characterization of silica and polystyrene nanoparticles shows strong agreement with PSDs obtained from the Monte Carlo algorithm (McSAS) and direct imaging analysis. Importantly, the FNN model achieves approximately 1800-fold acceleration in computation speed with a processing time of ∼50 ms per one-dimensional scattering curve, far surpassing the conventional McSAS method while maintaining robust predictive performance for both monodisperse and polydisperse systems. This work provides a practical tool for the rapid, high-precision analysis of complex particle systems in materials science and nanotechnology, partially addressing the long-standing challenge of real-time scattering data analysis.
1. Introduction
Particle size distribution (PSD) is a critical physical characteristic of nanoparticles and plays a vital role in their performance across various fields including nanomedicine,1–4 coatings5–7 and lithium-ion batteries.8–10 In the field of nanomedicine, Copelli et al. demonstrated that the PSD of active pharmaceutical ingredients affects the performance of a pharmaceutical drug product significantly.11 Similarly, another study on biodegradable microspheres for pulmonary drug delivery showed that a narrow PSD not only enhances drug delivery efficiency but also facilitates clearer investigation of drug release behavior.12 In the field of coatings, PSD exerts a significant impact on both coating performance and preparation processes. For example, calcium carbonate with a broad PSD induces the formation of a distinct particle-size gradient in the cross-sectional microstructure of coatings, leading to an irregular stress distribution and potentially compromising the coating stability.6 Zhang et al. further revealed that during the Cu coating process of silicon carbide (SiC) nanoparticles, a broad PSD of SiC nanoparticles results in preferential coating of Cu on large particles, whereas small particles are barely fully encapsulated.5 Thus, a narrow PSD of SiC nanoparticles is essential for achieving uniform Cu coatings. In the field of lithium-ion batteries, PSD profoundly influences charge–discharge performance.8 Farkhondeh et al. highlighted that spherical graphite particles with a narrow PSD deliver better electrochemical performance than those with a broad PSD, achieving higher coulombic efficiency during cycling.13 Furthermore, Zhang et al. reported that a uniform PSD helps suppress polarization buildup in the late discharge stage, thereby enhancing cycling stability.14 Chung et al. also showed that electrode materials with a monodisperse PSD yield higher power density at high discharge rates, whereas polydisperse systems favor higher energy density at low C-rates.15 Beyond material performance, PSD also governs flow stability in particulate systems. Kasper et al. observed that in slowly rotating drums containing wet granular matter, both liquid viscosity and particle size dictate the transition from intermittent avalanching to continuous flow.16 Balmforth et al. further linked avalanche amplitude to the initial free-surface angle, which itself depends on the particle size and system geometry.17 Mao et al. noted that while PSD has negligible effects in dry avalanches, the presence of a fluid phase strongly amplifies its influence on the motion of fluid–grain mixtures.18Small-angle X-ray scattering (SAXS) is a well-established technique for determining PSDs in nanoparticle systems.19,20 It exploits scattering resulting from electron density inhomogeneities as an X-ray beam traverses a sample, offering statistical information averaged over a large ensemble of nanoparticles. Extracting PSDs from scattering data typically involves numerical inversion methods. Widely used approaches include the stepwise extracting size distribution calculation (SESDC) method,21,22 the truncated singular value decomposition (TSVD) method,23 and the Monte Carlo method.24–26 The SESDC method segments and linearizes the Guinier curve, subsequently analyzing the distribution characteristics of different particle size intervals stepwise via Guinier approximation. This method is suitable for both monodisperse and dilute polydisperse systems. Unlike the SESDC method, the TSVD method is better suited for polydisperse systems. It conducts singular value decomposition based on particle shape models (e.g., spheres, cylinders and ellipsoid) and builds an approximate matrix using a truncation strategy to retrieve the PSD, effectively suppressing high-frequency noise interference. However, the choice of the truncation parameter directly affects the retention degree of PSD details, which might lead to substantially divergent solutions. The Monte Carlo method employs extensive random sampling to simulate scattering data based on predefined shape models, iteratively tuning parameters until the optimal goodness of fit between simulated and experimental SAXS profiles is achieved. McSAS stands as a leading implementation tool for this approach.25,26 Despite its exceptional efficacy in resolving multimodal and asymmetric PSDs, the method suffers from low computational efficiency, resulting in lengthy processing that restricts its utility in high-throughput SAXS data analysis pipelines. In recent years, as high-throughput experimental techniques have advanced rapidly, the data volume produced by SAXS experiments has surged exponentially, thereby presenting formidable challenges to the real-time retrieval of PSD.27,28
Recent advances in machine learning (ML) have opened new avenues for scattering data analysis. For instance, Molodenskiy et al. developed a feed-forward neural network (FNN) for predicting protein molecular weights and maximum dimensions from SAXS data.29 Zhou et al. designed a compact, lightweight yet efficient network (SEDCNN) for denoising experimental SAXS/wide-angle X-ray scattering diffraction images.30 Zhao et al. proposed a variational autoencoder (VAE) multilayer perceptron (MLP) neural network to establish the comprehensive processing–structure relationship of isotactic polypropylene films.31 Zhao et al. developed SAXSNN to reconstruct the morphology from experimental SAXS patterns directly without modeling by using a physics-aware neural network. By incorporating the basic scattering principle into the network, the trained SAXSNN has suggested to capture the complex mapping between the SAXS patterns in reciprocal space and the corresponding morphologies in real space in an unsupervised way.32 For nanoparticle-related studies, SAXS data analysis relies on the shape models, and the model selection and classification based on scattering data has thus become a research focus attracting extensive attention. Tomaszewski et al. developed a tool named SCAN, which integrates multiple ML algorithms to recommend optimal shape models for nanoparticle characterization.33 This tool takes 11 commonly predefined nanostructure models as the training set, achieving an overall accuracy of 95%–97% in model selection and classification. Monge et al. adopted a convolutional neural network (CNN) for nanoparticle model selection,34 whose training set comprises 75![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 scattering data from 9 nanoparticle models, enabling rapid screening of the optimal form factor for users. In addition to the aforementioned works on shape model classification, the quantification of nanoparticle's PSD also remains a challenge. Li et al. investigated the dual challenges of shape classification and size prediction for nanoparticles. They used simulated SAXS data of four shape models as the training set to develop SAXSNET, a dedicated network for nanomorphology classification that achieves a prediction accuracy of over 96%.35 In parallel, they adopted the random forest and XGboost regression algorithms for PSD predictions. However, further improvement on the prediction accuracy of PSD is needed when this method is applied to experimental SAXS data.
000 scattering data from 9 nanoparticle models, enabling rapid screening of the optimal form factor for users. In addition to the aforementioned works on shape model classification, the quantification of nanoparticle's PSD also remains a challenge. Li et al. investigated the dual challenges of shape classification and size prediction for nanoparticles. They used simulated SAXS data of four shape models as the training set to develop SAXSNET, a dedicated network for nanomorphology classification that achieves a prediction accuracy of over 96%.35 In parallel, they adopted the random forest and XGboost regression algorithms for PSD predictions. However, further improvement on the prediction accuracy of PSD is needed when this method is applied to experimental SAXS data.
Beyond scattering, PSD can also be derived from microscopy images. Kim et al. proposed an automated algorithm that combines computer vision and ML to extract PSD from SEM images.36 This method first adopts a pre-trained CNN for morphological classification, and then performs size measurement via specialized algorithms such as the distance transform for core-only particles and watershed-based segmentation for core–shell structures. Moreover, the physical scale is determined automatically using an efficient and accurate scene text detector (EAST) to locate the scale bar, and the Tesseract algorithm to recognize scale values and their units, thus enabling high-throughput PSD analysis. Zahedi et al. further proposed a fully automated, high-throughput SEM pipeline that integrates deep learning segmentation (U-Net/LinkNet) with automatic scale calibration. Specifically, the scale-bar length is detected via probabilistic Hough transform, while the scale value and unit are extracted using EAST-based text detection combined with an optical character recognition algorithm. This end-to-end pipeline enables the conversion of pixel-level contours to physically calibrated sizes, and direct the PSD output without manual intervention.37 Despite these imaging-based advances, ML methods for directly predicting PSD from scattering data remain underdeveloped.38–40
In summary, traditional SAXS inversion methods are often too slow for high-throughput or real-time analysis, while existing ML approaches for PSD prediction from scattering data still have room for improvement in accuracy and robustness. To address this, we develop an FNN-based model specifically for PSD prediction of spherical nanoparticles. By leveraging the nonlinear mapping capability of neural networks and incorporating physical constraints during training, our approach aims to deliver both high accuracy and computational efficiency, providing a practical tool for real-time, high-throughput SAXS data analysis.
2. Methods
2.1 Dataset construction
With the advancement of the scattering technique, its applications in nanoparticle research have expanded remarkably, accompanied by the development of various models for nanoparticle characterization. In this work, a dataset consisting of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 scattering curves was generated using the form factor model for spherical particles, which served as the training data for the FNN model. The scattering intensity I(q, r) can be expressed by the following formula:41
000 scattering curves was generated using the form factor model for spherical particles, which served as the training data for the FNN model. The scattering intensity I(q, r) can be expressed by the following formula:41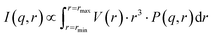 | (1) |
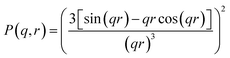 | (2) |
To acquire a sufficiently large and diverse set of scattering intensity data, a series of distinct PSD profiles were generated. For dataset construction, 1 to 4 Gaussian peaks were first generated with stochastic assignment of their means, standard deviations and weights. These peaks were then fitted using a Gaussian distribution function and normalized to a total sum of unity, after which the corresponding simulated scattering curves were calculated. To mimic real experimental conditions, 3% random noise was introduced into the scattering intensity data. The resulting dataset was split into training and validation subsets with a ratio of 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2. The former was used for training the FNN, while the latter served for hyperparameter optimization to alleviate overfitting.
2. The former was used for training the FNN, while the latter served for hyperparameter optimization to alleviate overfitting.
2.2 FNN architecture
In this work, a fully connected FNN was built to infer the PSDs of spherical particles from SAXS data with a standard architecture of an input layer, hidden layers, and an output layer, as shown in Fig. 1(a). The network's input layer takes the scattering curve I as an input of a size 500, and the output layer yields a size distribution V(r) of a size of 500. Two hidden layers are included with 2048 and 1024 neurons, respectively. The ReLU activation function is adopted for the hidden layers, while the sigmoid function is used for the output layer to ensure non-negativity of the output values. For the FNN training, the Adam optimization algorithm was employed with a batch size of 64, a learning rate of 0.001, and a maximum of 300 training epochs. The calculations were performed on a computer with the following hardware specifications: a 12th Gen Intel® Core™ i7-12700K processor with a base speed of 3.60 GHz, 64 GB of RAM, and a 2 TB hard drive, running on a 64-bit version of Windows 11. The root mean squared error (RMSE) was selected as the loss function, which quantifies the deviation between the FNN-predicted PSD and the ground-truth PSD, and its expression is given as follows:42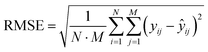 | (3) |
 | ||
| Fig. 1 (a) The FNN architecture trained for predicting the PSD. (b) Plots of the training and validation loss during training. | ||
The generalization capability of the FNN model is enhanced via neural network fine-tuning to improve its consistency with experimental data, where the fine-tuning procedure incorporates physical constraints to iteratively refine the predicted PSD. Taking the initial PSD prediction V(r)pred of the neural network as the starting point, we perform up to 500 optimization iterations with each iteration comprising three key steps: (i) calculating the scattering intensity Icalc from the current V(r)pred, (ii) deriving a correction vector from the ratio of experimental to calculated intensities, and (iii) calculating a correction coefficient for V(r)pred through back-projection, and subsequently updating V(r)pred with non-negativity constraints and normalization. Specifically, any negative values in V(r)pred are clipped to zero, after which the distribution is normalized so that the total volume fraction over the full radius range is equal to 1. This iterative optimization during FNN training improves the PSD inversion accuracy progressively. The key merit of this method lies in its dual advantage. That is, it synergistically combines the efficient data-driven prediction of neural networks with physically interpretable constraints.
2.3 Model performance evaluation
To systematically evaluate the prediction performance of the proposed FNN, 3000 sets of simulated test data were generated for each type of PSD with 1 to 4 peaks. The mean relative error (MRE) was employed to quantify the difference between the FNN-predicted PSD and the ground-truth one,43 and its expression is given as follows: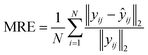 | (4) |
The coefficient of determination (R2) is a core metric for evaluating the prediction performance of ML regression models.44 Accordingly, this work also adopts this metric to quantify the degree of agreement between the predicted and ground-truth PSDs. R2 is expressed as follows:
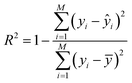 | (5) |
For the experimental validation, scattering data of two spherical nanoparticle samples were collected at the resonant soft X-ray scattering beamline of the Hefei light source BL05U-B. This beamline features prominent advantages including high photon flux, a small divergence angle, tunable energy, and an adjustable sample-to-detector distance. One sample is a silica (SiO2) spherical nanoparticle with a nominal radius of 250 nm, and the other is a polystyrene spherical nanoparticle with a nominal radius of 150 nm, both purchased from Nantong Zhichuan Microsphere Biotechnology Co., Ltd, China. Both samples were prepared by drop-casting onto Si3N4 windows and then air-drying. Si3N4 windows have an inner frame dimension of 1.5 mm × 1.5 mm, an outer frame dimension of 5 mm × 5 mm, and a window thickness of 100 nm. The CCD detector used in the experiments is Greateyes Lotte-i 4k4k, which has a detection area of 61.68 mm × 61.44 mm, a pixel size of 15 µm × 15 µm, an operating temperature of −80 °C, and a working vacuum of 1 × 10−4 Pa. The measurement parameters for the SiO2 sample were set as follows: an X-ray energy of 525 eV, an exposure time of 1 s, and a sample-to-detector distance of 170 mm. For the polystyrene sample, the parameters were set as follows: an X-ray energy of 410 eV, an exposure time of 10 s, and a sample-to-detector distance of 172 mm. In the experiments, the nanoparticle samples were mounted on a dedicated sample holder for X-ray scattering measurements. Prior to formal scattering measurements, dark backgrounds of the detector were collected and subsequently subtracted from the raw sample scattering data. All experimental scattering data were processed using ScatterX software developed by Xie et al. to obtain the final one-dimensional scattering curves.45 For complementary morphological characterization, scanning electron microscopy (SEM) was performed to directly visualize the spherical nanoparticles and verify their actual PSDs.
3. Results and discussion
To validate the performance of the trained FNN in predicting PSDs of both monodisperse and polydisperse spherical nanoparticles, we systematically evaluated test datasets containing 1 to 4 peaks, each consisting of 3000 simulated samples.As shown in Fig. 2, the MREs of the FNN-predicted PSDs are below 0.3 across all distributions, while the mean of R2 exceeds 0.93. Specifically, MRE and R2 values across test datasets with different peak numbers are comparable, and the prediction robustness keeps stable as the peak number increases. These results demonstrate the excellent mapping capability of the developed FNN model for both unimodal and multimodal PSDs.
To evaluate the prediction performance of the FNN intuitively, representative samples were randomly selected from test datasets containing 1 to 4 peaks. The predicted and ground-truth PSDs along with their corresponding scattering intensity profiles are compared, as shown in Fig. 3. The FNN model achieves high-fidelity PSD predictions across all test cases. For the one-peak case (Fig. 3(a)), the predicted curve accurately captures both the peak position and the full width at half maximum (FWHM) of the ground-truth distribution with a MRE of 0.1664 and an R2 value of 0.9613. For the two-peak case (Fig. 3(b)), the model correctly resolves the peak separation and relative intensity ratio without peak position misassignment, yielding a MRE of 0.2070 and an R2 value of 0.9318. Fig. 3(c) and (d) show that, for the three- and four-peak cases, FNN-based predictions also show good agreement with the ground-truth for all critical parameters including peak positions, intensity ratios, and FWHM. Notably, even those peaks with relatively lower content are accurately resolved in terms of position and distribution width, with a MRE of 0.2359 and an R2 value of 0.9094 for the three-peak case and a MRE of 0.2014 and an R2 value of 0.9503 for the four-peak case. The corresponding scattering intensity curves (Fig. 3(e)–(h)) further verify the consistency between the FNN-predicted and ground-truth profiles. Across all cases, the MRE between the FNN-predicted and ground-truth scattering curves is below 0.04 with R2 values all exceeding 0.999. This suggests an excellent match for both monodisperse and polydisperse systems. The stable performance confirms the reliability of the FNN in deconvoluting complex scattering signals and accurately reconstructing multi-peak PSDs.
To validate the performance of the FNN model on experimental data, synchrotron radiation X-ray scattering and SEM measurements were performed on SiO2 and polystyrene samples. As displayed in Fig. 4(a)and (b), SEM analysis reveals that the SiO2 sample consists primarily of uniformly sized particles with a tiny fraction of smaller nanoparticles and double-particle aggregates, while the polystyrene sample is primarily composed of monodisperse spherical particles with a minor fraction of smaller particles. Experimental scattering data for the two samples were collected, and the PSD inversion was carried out using the FNN and McSAS methods, respectively, with the corresponding scattering curves generated for each approach. Notably, the structural parameters for the McSAS and FNN methods were identical, with both methods using a spherical shape model and a particle size range of 1–500 nm. For McSAS, the minimum uncertainty estimate was set to 1%, and the convergence criterion was set to 1, ensuring that the simulation results achieved sufficient convergence. A comparison with experimental scattering data confirms close agreement between the scattering profiles obtained from FNN and McSAS and the measured curves for both samples, as presented in Fig. 4(c) and (d). Specifically, both methods exhibit strong consistency with the experimental data, whether in terms of the peak intensity, overall trend, and detailed fluctuations of the scattering curves. This indicates that the FNN and McSAS methods can accurately reproduce the global features and key details embedded in the scattering signals. Regarding the fitting accuracy of scattering curves, the FNN method achieves R2 values of 0.9952 and 0.9973 for the SiO2 and polystyrene samples, while the corresponding values for the McSAS method are 0.9714 and 0.9845, as shown in Table 1. Consistently, the FNN yields MRE values of 0.06528 for SiO2 and 0.04638 for polystyrene, which are lower than those of McSAS (0.147 for SiO2 and 0.1033 for polystyrene). These results confirm that both methods achieve good reconstruction accuracy with FNN delivering higher R2 and lower MRE, demonstrating its superior performance. Notably, while achieving comparable reconstruction accuracy, the FNN completes processing in approximately 50 ms per sample, in contrast to the 15 minutes required by McSAS per sample, representing an approximately 1800-fold improvement in computational efficiency.
| Sample | Method | R 2 | MRE |
|---|---|---|---|
| SiO2 | FNN | 0.9952 | 0.06528 |
| McSAS | 0.9714 | 0.147 | |
| Polystyrene | FNN | 0.9973 | 0.04638 |
| McSAS | 0.9845 | 0.1033 |
Further comparison of the PSD results obtained by the FNN and McSAS methods for both samples shows good agreement, particularly in the peak positions of the main and secondary distribution peaks. As seen in Fig. 5(a) and (b), the PSD of SiO2 exhibits a multi-peak feature. The dominant peak lies around 260 nm ranging from 250 to 270 nm, and a secondary peak appears in the range 300–330 nm. A smaller distribution is observed between 175 and 200 nm. Trace amounts of particles also present around 130–140 nm and 50–60 nm. Similarly, the PSD of polystyrene in Fig. 5(d) and (e) also displays multiple peaks. The strongest peak is centered at 160 nm (155–165 nm), with additional populations in the ranges of 180–210 nm, 220–250 nm, and 110–140 nm together with a trace distribution between 60 and 90 nm. For the PSD analysis, the MRE between the FNN-predicted and McSAS-retrieved is 0.5363 and 0.754 for the SiO2 and polystyrene samples, respectively. These results confirm that the FNN and McSAS methods are highly consistent in identifying the positions and relative intensities of the multi-peak features, supporting the reliability of the FNN model in analyzing complex multi-modal PSDs.
 | ||
| Fig. 5 PSDs of (a)–(c) SiO2 and (d)–(f) polystyrene obtained from scattering data using (a) and (d) FNN, (b) and (e) McSAS, and from (c) and (f) SEM image analysis. | ||
Notably, the radius distributions of spherical particles were also obtained by analyzing SEM images of SiO2 and polystyrene samples in this work. As shown in Fig. 5(c)–(f), the main radii of SiO2 and polystyrene are 260 nm and 160 nm, respectively. This suggests that the PSDs derived from SEM are consistent with those obtained from scattering measurements in terms of the main radius sizes. However, PSDs derived from SEM fail to capture the distribution features in other radius ranges revealed by the FNN and McSAS. This discrepancy fundamentally arises from the inherent statistical advantages of the scattering technique over SEM, particularly their capacity to perform ensemble averaging over large nanoparticle populations. Nevertheless, the SEM results strongly corroborate the accuracy of the FNN-predicted PSDs.
To assess the performance of the FNN model under varying conditions, its robustness and efficiency were systematically evaluated by considering factors such as noise levels, hidden layer sizes, and peak width. The results indicate that noise levels had minimal impact on the model's computational efficiency (see Fig. S1 and S2 and Table S1 of the SI). The effect of hidden layer size on the computation time and accuracy was also assessed, as presented in Tables S2 and S3 of the SI. The computational time was found to be unaffected by the size of the hidden layers and the number of peaks in the PSD, remaining steady at around 50 ms per sample. The model achieved the best performance with two hidden layers consisting of 2048 and 1024 neurons. Additionally, the accuracy of the FNN model was not influenced by changes in peak width, as demonstrated in Fig. S3–S5 of the SI. The performance validation of the FNN model on both simulated and experimental scattering data exhibits its feasibility and versatility for the accurate and efficient PSD extraction of both monodisperse and polydisperse systems. These results further underscore the model's robustness and adaptability across different types of data and system complexities. Unlike traditional methods, the FNN synergizes machine learning's capacity for high-dimensional nonlinear mapping with physics-informed optimization, employing neural network training for pattern recognition and physics-constrained parameter fine-tuning. This dual strategy enables efficient, high-precision SAXS analysis of intricate PSDs while ensuring physical plausibility. The computational efficiency of the FNN makes it especially valuable for scenarios requiring large-scale scattering data processing. In real time, high-frequency monitoring or high-throughput batch analysis, its millisecond-level inversion speed per curve can overcome the efficiency bottleneck of traditional methods, offering a new technical method for the rapid characterization of complex particle systems.
4. Conclusions
In this work, we developed an FNN model for the accurate and efficient prediction of PSD from scattering data, demonstrating distinct advantages over traditional methods. The FNN model achieves high-precision PSD characterization for both monodisperse and polydisperse systems. For simulated systems containing 1–4 peaks, the FNN model maintains a mean relative error (MRE) below 0.5 while accurately resolving critical features, including peak positions, full width at half maximum (FWHM), relative intensities, and even low-abundance minor peaks. Experimental validation using SiO2 and polystyrene nanoparticles confirms that the FNN-predicted PSDs closely match those obtained by the conventional McSAS method. Furthermore, SEM analysis independently verifies key peak positions (260 nm for SiO2 and 160 nm for polystyrene), corroborating the reliability of the FNN model for PSD analysis. Remarkably, the FNN model exhibits unprecedented computational efficiency. It infers the PSD from a single scattering curve in approximately 50 ms, which is roughly 1800 times faster than the McSAS method (15 minutes per curve). This breakthrough addresses the long-standing efficiency bottleneck of traditional inversion approach and enables real-time processing in scenarios such as in situ and high-throughput experiments. The method effectively synergizes the advantages of ML with physical plausibility by integrating physical constraints through iterative fine-tuning, ensuring that the predicted PSDs adhere to the fundamental physics of scattering while preserving the strong capability of ML to map high-dimensional nonlinear relationships. Additionally, by leveraging the statistical advantage of scattering measurements, the FNN model can identify minor size populations that are undetectable by SEM, thus enabling more comprehensive structural characterization. In summary, the FNN model with its exceptional accuracy, speed, and physical consistency offers a novel technical approach for the rapid, high-precision analysis of complex particle systems in materials science and nanotechnology.Conflicts of interest
The authors declare no competing interests.Data availability
The data that support the findings of this study are available upon request.The source code is openly available in Science Data Bank at https://www.scidb.cn/s/b6NVvm.
Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5cp04166j.
Acknowledgements
We thank the resonant soft X-ray scattering beamline (Jinhua) BL05U-B of Hefei Light Source, jointly constructed by the Zhejiang Institute of Optoelectronics and the National Synchrotron Radiation Laboratory of the University of Science and Technology of China. This work was supported by the project of the Anhui Provincial Natural Science Foundation (2308085MA19) and the National Key Research and Development Program of China (2023YFA1609800).References
- R. J. Hintz and K. C. Johnson, The Effect of Particle-Size Distribution on Dissolution Rate and Oral Absorption, Int. J. Pharm., 1989, 51, 9–17 CrossRef CAS.
- S. Gimondi, H. Ferreira, R. L. Reis and N. M. Neves, Size-Dependent Polymeric Nanoparticle Distribution in a Static versus Dynamic Microfluidic Blood Vessel Model: Implications for Nanoparticle-Based Drug Delivery, ACS Appl. Nano Mater., 2023, 6, 7364–7374 CrossRef CAS PubMed.
- R. Liu, S. S. Huang, Y. H. Wan, G. H. Ma and Z. G. Su, Preparation of insulin-loaded PLA/PLGA microcapsules by a novel membrane emulsification method and its release in vitro, Colloids Surf., B, 2006, 51, 30–38 CrossRef CAS PubMed.
- H. Bagheri, H. Hashemipour and S. Ghader, Population balance modeling: application in nanoparticle formation through rapid expansion of supercritical solution, Comput. Part. Mech., 2019, 6, 721–737 CrossRef.
- R. Zhang, L. Gao, L. Yu and J. K. Guo, Influence of particle size-distribution on the phase homogeneity between SiC particles and Cu coating, J. Inorg. Mater., 2002, 17, 1311–1314 CAS.
- Y. Wu and L. F. Francis, Effect of particle size distribution on stress development and microstructure of particulate coatings, J. Coat. Technol. Res., 2017, 14, 455–465 CrossRef CAS.
- L. D. D. Haddad, R. R. Neves, P. V. de Oliveira, W. J. dos Santos, A. N. de Carvalho and W. J. dos Santos, Influence of particle shape and size distribution on coating mortar properties, J. Mater. Res. Technol., 2020, 9, 9299–9314 CrossRef.
- M. Farkhondeh and C. Delacourt, Mathematical Modeling of Commercial LiFePO4 Electrodes Based on Variable Solid-State Diffusivity, J. Electrochem. Soc., 2012, 159, A177–A192 CrossRef CAS.
- S. T. Taleghani, B. Marcos, K. Zaghib and G. Lantagne, A Study on the Effect of Porosity and Particles Size Distribution on Li-Ion Battery Performance, J. Electrochem. Soc., 2017, 164, E3179–E3189 CrossRef CAS.
- N. B. Ch, A. Paramane and P. R. Randive, Influence of anode particle size distribution on internal short-circuit behaviour of lithium-ion battery, Ionics, 2025, 31, 4053–4072 CrossRef CAS.
- D. Copelli, A. Cavecchi, C. Merusi and R. Leardi, Multivariate evaluation of the effect of the particle size distribution of an Active Pharmaceutical Ingredient on the performance of a pharmaceutical drug product: a real-case study, Chemom. Intell. Lab. Syst., 2018, 178, 1–10 CrossRef CAS.
- R. Zhao, J. Xu and B. Guo, Preparation and Evaluation of Biodegradable Microspheres with Narrow Size Distribution for Pulmonary Delivery, Indian J. Pharm. Sci., 2017, 79, 930–938 CAS.
- L. Bläubaum, F. Röder, C. Nowak, H. S. Chan, A. Kwade and U. Krewer, Impact of Particle Size Distribution on Performance of Lithium-Ion Batteries, ChemElectroChem, 2020, 7, 4755–4766 CrossRef.
- J. Zhang, J. S. Qiao, K. N. Sun and Z. H. Wang, Balancing particle properties for practical lithium-ion batteries, Particuology, 2022, 61, 18–29 CrossRef CAS.
- D. W. Chung, P. R. Shearing, N. P. Brandon, S. J. Harris and R. E. García, Particle Size Polydispersity in Li-Ion Batteries, J. Electrochem. Soc., 2014, 161, A422–A430 CrossRef CAS.
- J. H. Kasper, V. Magnanimo, S. D. M. de Jong, A. Beek and A. Jarray, Effect of viscosity on the avalanche dynamics and flow transition of wet granular matter, Particuology, 2021, 59, 64–75 CrossRef CAS.
- N. J. Balmforth and J. N. McElwaine, From episodic avalanching to continuous flow in a granular drum, Granular Matter, 2018, 20, 52 CrossRef.
- W. Mao, Y. Wang, P. Yang, Y. Huang and H. Zheng, Dynamics of granular debris flows against slit dams based on the CFD–DEM method: effect of grain size distribution and ambient environments, Acta Geotech., 2023, 18, 5811–5838 CrossRef.
- M. Doktorova, N. Kucerka, J. J. Kinnun, J. J. Pan, D. Marquardt, H. L. Scott, R. M. Venable, R. W. Pastor, S. R. Wassall, J. Katsaras and F. A. Heberle, Molecular Structure of Sphingomyelin in Fluid Phase Bilayers Determined by the Joint Analysis of Small-Angle Neutron and X-ray Scattering Data, J. Phys. Chem. B, 2020, 124, 5186–5200 CrossRef CAS PubMed.
- L. Caselli, L. Conti, I. De Santis and D. Berti, Small-angle X-ray and neutron scattering applied to lipid-based nanoparticles: Recent advancements across different length scales, Adv. Colloid Interface Sci., 2024, 327, 103156 CrossRef CAS PubMed.
- R. C. Chen, Z. H. Li and J. H. He, Determination of upper limit of Guinier approximation on dilute polydisperse system in SAXS, Nucl. Instrum. Methods Phys. Res., Sect. B, 2024, 552, 165377 CrossRef CAS.
- R. C. Chen and J. H. He, Stepwise Extracting Size Distribution Calculation (SESDC) Method for SAXS Dilute Polydisperse Spherical Systems, Adv. Theory Simul., 2025, 2500323 CrossRef CAS.
- X. J. Zhu, J. Shen, W. Liu, X. M. Sun and Y. J. Wang, Nonnegative least-squares truncated singular value decomposition to particle size distribution inversion from dynamic light scattering data, Appl. Opt., 2010, 49, 6591–6596 CrossRef PubMed.
- I. Breßler, B. Pauw and A. Thünemann, McSAS: A package for extracting quantitative form-free distributions, 2014, p. 1900 Search PubMed.
- I. Bressler, B. R. Pauw and A. F. Thünemann, McSAS: software for the retrieval of model parameter distributions from scattering patterns, J. Appl. Crystallogr., 2015, 48, 962–969 CrossRef CAS PubMed.
- B. R. Pauw, C. Kastner and A. F. Thunemann, Nanoparticle size distribution quantification: results of a small-angle X-ray scattering inter-laboratory comparison, J. Appl. Crystallogr., 2017, 50, 1280–1288 CrossRef CAS PubMed.
- R. Fini, M. Magnani, C. V. Santilli and S. H. Pulcinelli, Tuning the Formation and Growth of Platinum Nanoparticles Using Surfactant: SAXS Study of the Aggregative Growth Mechanism, ACS Appl. Mater. Interfaces, 2025, 17, 41237–41248 CrossRef CAS PubMed.
- L. Q. Huang, J. G. Mai, Q. H. Zhu, Z. Guo, S. Y. Qin, P. L. Yang, X. X. Li, Y. C. Shi, X. T. Wang, Q. N. Wang, N. Li, C. Xie and H. G. Liu, Reversible rearrangement of magnetic nanoparticles in solution studied using time-resolved SAXS method, J. Synchrotron Radiat., 2019, 26, 1294–1301 CrossRef CAS PubMed.
- D. S. Molodenskiy, D. Svergun and A. G. Kikhney, Artificial neural networks for solution scattering data analysis, Structure, 2022, 30, 900–908 CrossRef CAS PubMed.
- Z. Z. Zhou, C. Li, X. X. Bi, C. L. Zhang, Y. K. Huang, J. Zhuang, W. Q. Hua, Z. Dong, L. A. Zhao, Y. Zhang and Y. H. Dong, A machine learning model for textured X-ray scattering and diffraction image denoising, npj Comput. Mater., 2023, 58, 5496 Search PubMed.
- C. Zhao, W. Yu and L. Li, Visualization of small-angle X-ray scattering datasets and processing-structure mapping of isotactic polypropylene films by machine learning, Mater. Des., 2023, 228, 111828 CrossRef CAS.
- C. Zhao, S. Sun, X. Han, J. Zhu, W. Yu and L. Li, Morphology reconstruction from experimental small-angle x-ray scattering patterns by physics-aware neural network, APL Mach. Learn., 2025, 3, 016109 CrossRef.
- P. Tomaszewski, S. Yu, M. Borg and J. Ronnols, Machine Learning-Assisted Analysis of Small Angle X-ray Scattering, 2021 Swedish Workshop on Data Science (SweDS), 2021, pp. 1–6 Search PubMed.
- N. Monge, M. R. Amini and A. Deschamps, Influence of device configuration and noise on a machine learning predictor for the selection of nanoparticle small-angle X-ray scattering models, Acta Crystallogr., Sect. A:Found. Adv., 2024, 80, 405–413 CrossRef CAS PubMed.
- Y. K. Li, L. Y. Liu, X. N. Zhao, S. M. Zhou, X. H. Wu, Y. C. Lai, Z. J. Chen, J. Z. Chen and X. Q. Xing, Deep learning-assisted characterization of nanoparticle growth processes: unveiling SAXS structure evolution, Radiat. Detect. Technol. Methods, 2024, 8, 1712–1728 CrossRef.
- H. Kim, J. Han and T. Y.-J. Han, Machine vision-driven automatic recognition of particle size and morphology in SEM images, Nanoscale, 2020, 12, 19461–19469 RSC.
- R. Zahedi, H. Bagheri, F. Ghasemian, M. Ghazvini and S. Y. Ziaei, Nano-particles size measurement based on semantic segmentation via convolution neural network, Measurement, 2025, 240, 115513 CrossRef.
- M. Frei and F. E. Kruis, Image-based size analysis of agglomerated and partially sintered particles via convolutional neural networks, Powder Technol., 2020, 360, 324–336 CrossRef CAS.
- J. Bals and M. Epple, Deep learning for automated size and shape analysis of nanoparticles in scanning electron microscopy, RSC Adv., 2023, 13, 2795–2802 RSC.
- Y. Wu, M. Lin and S. Rohani, Particle characterization with on-line imaging and neural network image analysis, Chem. Eng. Res. Des., 2020, 157, 114–125 CrossRef CAS.
- J. Bolze, D. Beckers and V. Kogan, Size Distribution Determination of Nanoparticles and Nanosized Pores by Small-Angle X-Ray Scattering (SAXS) on a Multi-Purpose X-Ray Diffractometer, 2012, pp. 1–11 Search PubMed.
- J. G. Ethier, D. J. Audus, D. C. Ryan and R. A. Vaia, Integrating theory with machine learning for predicting polymer solution phase behavior, Giant, 2023, 15, 100171 CrossRef CAS.
- J. Zhou, Z. L. Su, S. Hosseini, Q. Tian, Y. J. Lu, H. Luo, X. Q. Xu, C. P. Chen and J. D. Huang, Decision tree models for the estimation of geo-polymer concrete compressive strength, Math. Biosci. Eng., 2024, 21, 1413–1444 Search PubMed.
- A. Teimouri, A. Challapalli, J. Konlan and G. Q. Li, Machine learning assisted design and optimization of plate-lattice structures with superior specific recovery force, Giant, 2024, 18, 100282 CrossRef.
- F. Xie, M. Xie, B. Song, Q. Guo and X. Jiao, ScatterX: A software for fast processing of high-throughput small-angle scattering data, Chin. Phys. B, 2024, 33, 120101 CrossRef.
Footnote |
| † Q. Guo and F. Xie contributed equally to this work. |
| This journal is © the Owner Societies 2026 |



