Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
From bioavailability scarcity to energy barriers: limitations of anaerobic microbial reductive defluorination
Yi
Ren
 a,
Wenhao
Deng
b and
Mike
Manefield
*a
a,
Wenhao
Deng
b and
Mike
Manefield
*a
aWater Research Centre, School of Civil and Environmental Engineering, Sydney, NSW 2052, Australia. E-mail: manefield@unsw.edu.au
bKey Laboratory of Material Chemistry for Energy Conversion and Storage, Ministry of Education, Hubei Key Laboratory of Bioinorganic Chemistry and Materia Medica, Hubei Key Laboratory of Materials Chemistry and Service Failure, School of Chemistry and Chemical Engineering, Huazhong University of Science and Technology, Wuhan 430074, P. R. China
First published on 10th December 2025
Abstract
Reductive dehalogenases in organohalide-respiring bacteria underpin anaerobic bioremediation of chlorinated pollutants but are rarely effective for reductive defluorination of per- and polyfluoroalkyl substances. However, the physicochemical basis for this selectivity remains unclear. Here, we integrate quantum chemistry and molecular dynamics to evaluate constraints on microbial reductive defluorination. The scarcity of naturally occurring organofluorines has imposed limited evolutionary selective pressure, explaining the absence of robust defluorination pathways. Using quantum mechanical calculations, we show that organofluorines have low bioavailability, due to increasingly unfavorable solvation free energies of fluorinated ethenes in both polar and nonpolar solvents, impeding cellular uptake. Using molecular dynamics simulations, we show that the substrate recognition by reductive dehalogenases is compromised, due to progressively weaker van der Waals energies as chlorines are replaced by fluorines. A tetrafluorinated ligand can form hydrogen bonds with polar residues and is preferentially stabilised in a sub-pocket away from the catalytic site. Using quantum mechanics calculations with a cluster model of the active site, we show that the reductive cleavage of the C–F bond has prohibitively high energy barriers. Together, these results explain the limited anaerobic microbial reductive defluorination of linear per- and polyfluoroalkyl substances and highlight why engineering applications are unlikely to succeed. The workflow provides a screening framework for assessing biodegradability of new organofluorines prior to industrial deployment.
1. Introduction
Reductive dehalogenases (RDases) produced by organohalide-respiring bacteria (ORB) are natural bioremediation tools available in groundwater for organohalogen contamination. ORB are obligate anaerobes, including Dehalococcoides mccartyi, Dehalobacter restrictus and Desulfitobacterium dichloroeliminans, that conserve energy by coupling electron donors (e.g., H2 or organics) to organohalogens serving as terminal electron acceptors.1 RDases are membrane associated, cobalamin (B12)-dependent enzymes that contain a corrinoid cofactor and two iron–sulfur clusters to shuttle reducing equivalents during catalytic reactions.2 One of the RDase crystal structures is PceA (PDB id: 4UQU)3 from anaerobic ORB Sulfurospirillum multivorans. PceA is a dimer with a buried active site encircled by aromatic residues.3 Mechanistically, the cobalamin cofactor mediates reductive C–Cl bond cleavage of tetra-/trichloroethene through a concerted, proton-coupled electron-transfer transition state in which an active-site tyrosine donates the proton and the proximal iron–sulfur cluster supplies the electron.4,5There are numerous RDases from different ORB with various organohalogen substrate specificities. TmrA from Dehalobacter UNSWDHB efficiently catalyzes trichloromethane, a common inhibitor among ORB.6,7 The closely related CfrA and DcrA (95.2% identity in amino acids), from the Dehalobacter strain CF and strain DCA, respectively, target different short-chain chlorinated ethanes.8,9 PceA from Sulfurospirillum multivorans (previously named Dehalospirillum multivorans) can dechlorinate tetra/trichloroethenes.10,11 VcrA from Dehalococcoides sp. strains VS and BAV1 can dechlorinate vinyl chloride.12 But confirmed RDase reactivity toward organofluorinated substrates is rare.
Per- and polyfluoroalkyl substances (PFASs) comprise partially or fully fluorinated alkyl chains bearing carboxylate or sulfonate headgroups.13 Strong C–F bonds and low polarizability confer exceptional thermal and chemical stability,14,15 enabling uses such as aqueous film-forming foams (AFFFs) at firefighting sites.16–18 Widespread deployment has produced pervasive contamination with groundwater as a major sink (μg L−1 levels reported) and associated risks to drinking water supplies.19–23
PFAS persist in part because indigenous microorganisms rarely affect complete biotransformation. Under aerobic conditions, strains including Gordonia sp. NB4-1Y, Dietzia aurantiaca J3, Rhodococcus jostii RHA1 and several Pseudomonas spp. can transform selected fluorotelomer precursors (e.g., 6:2 FTAB/FTS, 4:2–8:2 FTOH and 5:3 FTCA).24–30 Cobalamin-mediated reductive defluorination shows that branched perfluorinated compounds are energetically possible to reduce anaerobically.31 However, biodegradation of PFAS has not been observed in groundwater32,33 where anaerobic and reducing conditions predominate.34 Although it is now trivial to enrich anaerobic dechlorinating bacteria, comparable enrichment of anaerobic defluorinating bacteria has not been achieved. Notably, one dechlorinating enrichment culture reductively defluorinated branched unsaturated per/polyfluoronated compounds (PFMeUPA and FTMeUPA) over 150 days,35 the responsible microorganism was not determined for reductive defluorination. These results suggest the necessary investigation into the physical basis that restricts anaerobic biological reductive defluorination towards per/polyfluorinated compounds,36 which become the focus of the present study.
Various perspectives can be considered regarding the limited microbial reductive defluorination of PFAS. One significant factor is evolutionary limitations, which stem from the scarcity of naturally occurring organofluorinated compounds. PFAS are synthetic, so microorganisms have experienced minimal selective pressure to evolve efficient defluorination pathways.37–40 The bioavailability of pollutants can also be a limiting factor for microbial adaptation towards anaerobic reductive defluorination. The bioavailability is dictated by physical chemistry perspectives,41 and negatively correlated with lipophilicity, and shaped by polarity and solvation energy.42 Fluorination alters electron distribution of ligands and intermolecular interactions, typically increasing hydrophobicity and diminishing polarity. PFAS have positive partition coefficients, indicating lipophilicity,43 but the impacts of various fluorination degrees on bioavailability have not been fully investigated. Ligand polarity and dispersion contacts affect binding free energy within RDase active sites,44 and stronger, more complementary binding generally lowers catalytic barriers by positioning reactive groups optimally.45 But the influence of various fluorination on ligand binding free energy with RDase is unknown. Currently, the only crystalized structure of an anaerobic RDase is PceA,3 providing an opportunity to explore the binding free energy between organofluorinated compounds and the enzyme using molecular dynamics simulations. Another aspect of limited biodegradability is the energy barrier of reductive defluorination.46 For dehalogenation catalyzed by PceA, a concerted transition-state has been shown to lower the energy barrier for perchloroethene reduction,47 with a proton donated by an active-site tyrosine to the ligand.4 However, the energy barrier for linear fluorinated ligands with PceA has not been calculated.
In the present study, for the first time, we present a rigorous theoretical computational investigation into the physicochemical limitations of microbial reductive defluorination of linear organofluorinated compounds. We first discuss evolutionary and environmental context for organofluorine exposure. Next, we use quantum mechanics calculations to quantify ligand polarity and solvation free energies as a function of fluorination degree and quantify the bioavailability of organofluorinated ligands. Subsequently, we apply molecular dynamics simulations to calculate binding free energies of chlorinated versus fluorinated ligands in PceA active sites. Finally, we use the quantum mechanics method to calculate the activation energy barriers for enzymatic C–F bond cleavage. Together, these results explain why anaerobic defluorination of linear PFAS is rarely observed, delineate the narrow chemical space where it might occur, and offer a generalizable screening framework for assessing the biodegradability of new organofluorines prior to industrial application.
2. Computational details
2.1. Selected ligands
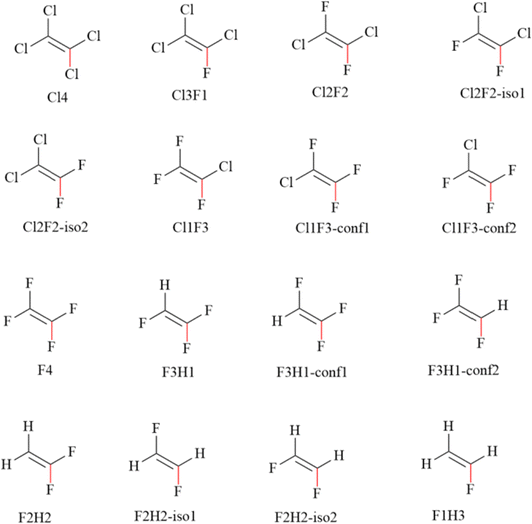
The ligands shown in Chart 1 are halogenated ethenes with varying degrees of chlorination and fluorination. The Cl4 ligand in Chart 1 is the native substrate of PceA,3,48 and is used as the reference for evaluating the energy barriers of reductive defluorination by PceA. Halogenated ethenes with progressively higher levels of fluorination were then selected to assess the influence of fluorine substitution on defluorination. The strengths of the carbon–halogen bonds highlighted in red were probed by calculating the energy barriers for reductive dehalogenation. In addition, the effects of different relative positions of fluorine atoms were examined by including positional isomers. These ligands were used for calculations of defluorination energy barriers, electron distributions, and solvation free energies.2.2. Solvation free energy calculations
We used the following equation49 to calculate the solvation energy of the ligands:| ΔGsolv = Gsoln − Ggas |
ΔGsolv is the solvation free energy, Gsoln is the energy in the solution phase, and Ggas is the energy in the gas phase. Solvation energies were calculated using Gaussian 16.50 For solution-phase energies, ligand geometries were optimized with the M05-2X density functional and 6-31G* basis set51 in combination with the solvation model density (SMD) calculation,49 using water or n-octanol as the solvent, respectively. Dipole moments were obtained from frequency calculations with water as the solvent. The octanol–water partition coefficients (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pow) of the tested ligands were estimated using MarvinSketch, version 23.17.0, 2023, ChemAxon (https://www.chemaxon.com).52 The ChemAxon log
Pow) of the tested ligands were estimated using MarvinSketch, version 23.17.0, 2023, ChemAxon (https://www.chemaxon.com).52 The ChemAxon log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P model is an atom/fragment-based QSPR method derived from the fragmental approach of Viswanadhan,53 in which a molecule is decomposed into predefined atom types and fragments, and the overall log
P model is an atom/fragment-based QSPR method derived from the fragmental approach of Viswanadhan,53 in which a molecule is decomposed into predefined atom types and fragments, and the overall log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P is calculated as the sum of atomic/fragment contributions plus empirically derived correction terms fitted to experimental n-octanol/water partition data. Under the default “ChemAxon” settings, this model and related fragment-based schemes have been shown to reproduce experimental log
P is calculated as the sum of atomic/fragment contributions plus empirically derived correction terms fitted to experimental n-octanol/water partition data. Under the default “ChemAxon” settings, this model and related fragment-based schemes have been shown to reproduce experimental log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values with mean absolute errors of ∼0.2–0.4 log units for drug-like and environmental organic molecules and to perform comparably to or better than more computationally demanding quantum-chemical solvation protocols.52 To understand the influence of fluorine substitution on molecular polarity, electrostatic potential (ESP) maps were calculated using Multiwfn54,55 and are shown in Fig. 2, following the visualization schemes used in previous studies.56–58
P values with mean absolute errors of ∼0.2–0.4 log units for drug-like and environmental organic molecules and to perform comparably to or better than more computationally demanding quantum-chemical solvation protocols.52 To understand the influence of fluorine substitution on molecular polarity, electrostatic potential (ESP) maps were calculated using Multiwfn54,55 and are shown in Fig. 2, following the visualization schemes used in previous studies.56–58
2.3. Molecular dynamics simulation details and binding free energy calculations
To calculate the PceA–ligand binding free energies from molecular dynamics simulations, the enzyme cofactor and ligands needed to be parameterized. Cobalamin, the cofactor of PceA, and the selected ligands were parameterized at the B3LYP/ 6-311G(d,p) level of theory (B3LYP-D3/6-311+g(2d,2p) for the ligands) using Gaussian 09.59 Based on a previous study, the Co atom in cobalamin adopts a +2 oxidation state in the reactant,4 and we therefore assigned the cobalamin cofactor an overall charge of 0 and a doublet spin state.The Sobtop package60 was used to assign the generalized AMBER force field (GAFF)61 atom types for the selected ligands (Chart 1, Cl4, Cl3F1, Cl2F2, Cl1F3 and F4). For the cobalamin cofactor, the cobalt atom was assigned a universal force field (UFF) atom type,62 while GAFF atom types were used for the remaining cobalamin atoms. A similar strategy, which uses UFF for the metal center and GAFF for the surrounding organic framework, has been adopted in previous studies.63,64 Atomic charges were derived using the restrained electrostatic potential (RESP) scheme, as implemented in Multiwfn.54 Detailed force field parameters of the cobalamin and ligands, including atomic names, types, charges, ε and σ values, and bonds, angles and dihedrals of cobalamin and ligands are shown in the SI, Tables S1–S4 and Fig. S1 and S2. The cobalamin cofactor parameters were validated by comparing the Co–N bond distances and the N–Co–N angles of adjacent coordinating nitrogen atoms and the validation results are shown in Fig. S3–S5 and Table S5 of the SI.
The protein structure of PceA (PDB is 4UQU) was simulated with Amber forcefield ff14SB.65 Protonation states at pH 7.0 were assigned with the CHARMM-GUI platform,66 and verified by PROPKA 3. Histidine residues were specified using AMBER residue names, including HID (Nδ-H), HIE (Nε-H), and HIP (+1, both nitrogen atoms protonated). In the final model, the 5 histidine residues are HID 40, HIP 187, HIP 357, HIE 400 and 449HID. Other titratable residues were kept in their canonical protonation states at pH 7 (Asp/Glu deprotonated, Lys/Arg protonated, and Tyr neutral). Water molecule parameters were modelled as TIP3P.67 All the systems were solvated in a cubic box containing 20037 TIP3P water molecules, with a dimension of 89 Å in x, y and z dimensions (default setting in CHARMM-GUI with 10 Å from the edge of the protein). The charge of the systems was neutralised to zero by adding 2 sodium ions.
All simulations and trajectory analyses were performed with GROMACS version 2024.3-gpuvolta.68,69 The starting structures, consisting of PceA with a ligand docked in the active site, were first relaxed by 1000 steps of steepest-descent energy minimization. Equilibration was then carried out in two stages. First, a constant-volume canonical (NVT) equilibration was run at 303.15 K using a V-rescale thermostat70 for 4 nanoseconds (ns) with a time step of 1 fs. This was followed by a constant-pressure isothermal–isobaric (NPT) equilibration at 1 bar using the Parrinello–Rahman barostat,71,72 with isotropic pressure coupling. Next, the NPT equilibration was performed in four stages, during which position restraints were applied to the protein and cofactor. Force constants of 400 and 40 kJ mol−1 nm−2 were initially applied to the backbone and side chains, respectively (as in the NVT stage), and were decreased by 100 and 10 at each subsequent stage. Position restraints on the ligands were also applied, starting at 500 kJ mol−1 nm−2, and reduced by 125 kJ mol−1 nm−2 at each stage. In the final stage of equilibration, all position restraints were removed, and the system was fully relaxed. The Verlet cutoff scheme73 was employed, with a 1.2 nm cutoff for short-range van der Waals interactions. Long-range electrostatics were treated with the particle–mesh Ewald (PME) method.74 All bonds involving hydrogen atoms were constrained using the LINCS algorithm.75
After equilibration, three independent 100 ns production runs were performed with a 2fs time step. The SHAKE algorithm76 was used to constrain all bonds involving hydrogen atoms during production. The molecular dynamics simulations of PceA were validated by comparison with the PceA crystal structure. These validation results are provided in Table S6 and Fig. S6 of the SI. The backbone root-mean-square deviations (RMSDs) for all production simulations are shown in Fig. S7.
The enthalpic components of the binding free energies of the protein–ligand complexes in solvents77,78 were calculated using the molecular mechanics Poisson–Boltzmann surface area (MM-PBSA) approach79 as implemented in the g_mmpbsa tool,80 with the default parameters.
Binding free energies were evaluated with respect to the canonical PceA active site, defined by the binding mode of the native Cl4 ligand. For each ligand, a total of 1000 snapshots were sampled evenly from the trajectory in which the ligand remained associated with this pocket for analysis. For F4, this corresponds to the time window during which it occupies the canonical pocket (0–60 ns) prior to migration into the adjacent sub-pocket. For each system, three independent simulations were performed, and the reported values are averages over these three runs. We did not include an explicit entropic term (−TΔS)81 in the MM/PBSA calculation. Normal-mode or quasi-harmonic entropy estimates for large protein systems are computationally demanding and often poorly converged, which can increase the uncertainty and degrade the reliability of the present calculations.82 The detailed equations used for the MM-PBSA analysis are provided in the SI.
Potential hydrogen bonds between the protein and the fluorinated ligands were quantified using MDAnalysis (version 2.9.0).83 Trajectories were recentred and made whole prior to analysis. The acceptors were the ligand fluorine atoms (F*). Donor hydrogens were all hydrogens on the protein and each H was mapped to its nearest N or O heavy atom within 1.2 Å to define the donor X–H pair. A contact was counted if it satisfied the geometric criteria H⋯F ≤ 3.5 Å and ∠(X–H⋯F) ≥ 120°(with X = N or O). Counts were averaged over the production window (reported as mean ± SD across replicate simulations).
2.4. QM calculations
To compute the energy barriers for reductive defluorination, DFT calculations were performed using a QM cluster model constructed from the X-ray structure of PceA, following the same method of Liao et al.4 It is important to emphasize that reliable transition-state calculations require both a sufficiently large active-site model to capture the substrate environment and an accurate description of the relative energies of the transition states. The enzyme cluster approach used here84 satisfies both criteria and has been successfully applied to multiple systems, including epoxide hydrolases and arylmalonate decarboxylase.85,86All DFT calculations were carried out by using the Gaussian 09 suite of programs.59 The QM cluster model included the truncated corrin ring and seven key residues: Phe38, Thr242, Tyr246, Arg305, Leu306 and Trp376. Geometries were optimized by using the B3LYP-D3 functional87 with the LANL08 basis set for Co,88 and 6-31G(d,p) for all other elements. To avoid artificial expansion or other rearrangements, atoms at the periphery of the cluster were fixed at their crystallographic positions where the truncations were made.89,90 The fixed atoms were selected in the same way as in the study of Liao et al,4 shown in the SI, Fig. S8. Frequency calculations were performed at the same level of theory on the optimized geometries to obtain zero-point vibrational energy (ZPE) corrections and the imaginary frequencies of the transition states, which are reported in Table 2. Cartesian coordinates of all DFT-optimized structures are provided in the XYZ_coordinates.zip file in the Git-hub repository of the present study.
For searching transition state geometry, we started from the reactant minimum, we performed two-dimensional relaxed energy scans along the C–F (breaking bond) and H–C (forming bond from Tyr246) distances. At each grid point, these two distances were constrained to fixed values, while all other internal coordinates in the cluster were optimized and selected peripheral atoms were kept fixed at their crystallographic positions to define the cluster boundary. We chose relaxed scans so that the active-site environment (Tyr246, Arg305, cobalt coordination sphere, and nearby residues) can reorganize optimally along the reaction path, whereas a rigid scan would artificially freeze the environment and distort the barrier.
The approximate transition state region was identified as the highest energy point along the minimum-energy path between reactant-like and product-like regions of this 2D surface. This structure was then used as the initial guess for an unconstrained transition state optimization (with only the boundary atoms fixed, same as in the reactant). The resulting stationary point typically exhibits a single dominant imaginary frequency that is localized on the reacting centre and involves concerted C–F bond cleavage and H–C bond formation, together with other small imaginary modes confined to peripheral groups near the fixed boundary. We verified this by visual inspection of the vibrational normal modes (GaussView animations) and therefore identify the structure with the dominant reaction-coordinate imaginary mode as the chemically relevant transition state for each ligand.
Final energies of all intermediates and transition states were obtained from single-point calculations on the optimized geometries using a larger basis set, namely LANL08 for Co and 6-311+G(2d,2p) for all other atoms. To account for solvation effects from the protein environment, single-point calculations were also performed on the optimized structures at the same level of theory with the SMD solvation model.91 The dielectric constant (ε) was set to 4, consistent with the value used previously by Liao et al.4 The calculation of the energy difference between the unprotonated Tyr246, CoII and the protonated Tyr246, CoI can be found in the SI. Based on the calculated activation barriers, reaction kinetics were estimated using the Eyring equation.92–95
3. Results and discussion
3.1. Scarcity of naturally occurring organofluorine compounds limits bioavailability
The biodegradability of newly synthesized pollutants is subjected to the duration and extent of microbial exposure, which in turn depends on the compounds’ natural environmental abundance.37,39 Multiple polyhalogenated organochlorines (e.g., tetrachloroethene and trichloroethene) are produced naturally by volcanic activity and marine algae, and trichloromethane and dichloromethane have been detected in volcanic gases,96 indicating their presence of organochlorines on Earth well before the emergence of life (3.8–3.9 billion years ago).97 Consequently, RDases likely have an ancient evolutionary origin, having evolved to dechlorinate naturally occurring organohalogens,36,98–100 long before industrial-scale synthesis of organohalogen compounds began in the 1940s.101 This prior exposure plausibly primed RDases with broad substrate specificity to adapt to anthropogenic chlorinated pollutants, particularly when their molecular structures match or resemble natural organohalogen analogues.37,39By contrast, microbial reductive defluorination remains limited, consistent with the relative scarcity of naturally occurring organofluorines. This limitation is likely attributable to the relative scarcity of naturally occurring organofluorinated compounds. The cosmogenic formation of fluorine requires stringent astrophysical nucleosynthetic pathways,102–104 yielding far lower cosmic and, therefore, terrestrial abundance than other halogens and nearby elements such as C, N, O, and Ne.104 On Earth, fluorine is predominantly sequestered in crustal and mantle minerals, where its high electronegativity promotes strong ionic bonding with silicate minerals limiting its distribution in the biosphere.104 Volcanic eruptions are among the few natural processes that release hydrogen fluoride and trace amounts of organic fluorinated compounds into the environment.105 Although the mass production of PFAS compounds began around the same time as organochlorine solvents in the 1940s and 1950s,106 their natural analogues are rare, and sustained microbial exposure has likely been insufficient for the evolution of efficient anaerobic defluorination pathways.
This scarcity imposes an evolutionary disadvantage for microorganisms to develop robust defluorination metabolic pathways. A notable exception is fluoroacetate dehalogenase, which defluorinates fluoroacetate, which is a monofluorinated compound structurally similar to common metabolic intermediates, but the activity is largely confined to singly fluorinated substrates.107,108 In contrast, the heavily substituted, highly fluorinated architectures of industrial PFAS exhibit minimal resemblance to natural metabolites, further reducing recognition by existing enzyme repertoires and hindering adaptation.
3.2. Polarity of organofluorines limits bioavailability
Bioavailability governs the exposure of organisms to pollutants in water, soil, and sediments,109 and therefore strongly influences biodegradation. When contaminants partition out of the aqueous phase by adsorbing onto hydrophobic surfaces, accumulating at water and non-aqueous phase liquid (NAPL) interfaces or forming separate NAPL phases,110 the diffusive mass transfer to microbes and extracellular enzymes diminishes, limiting biodegradation. Molecular polarity controls hydrophobicity and thus bioavailability,41 because hydrophobic pollutants desorb and repartition slowly from solids or NAPLs into water.111,112 The octanol–water partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pow) is a standard proxy for hydrophobicity that helps to estimate bioavailability at NAPL–water interfaces,113 reflecting the balance of electrostatic and dispersion interactions that is strongly modulated by molecular polarity, especially dipole–dipole contributions.114
Pow) is a standard proxy for hydrophobicity that helps to estimate bioavailability at NAPL–water interfaces,113 reflecting the balance of electrostatic and dispersion interactions that is strongly modulated by molecular polarity, especially dipole–dipole contributions.114
Low-polarity pollutants can occupy both hydrophobic and certain hydrophilic micropores via a dispersion force.115 Water (polarity 1.85 Debye) preferentially saturates the most polar sites, leaving low-polarity organics to compete for nonpolar domains.116 At AFFFs contaminated sites, PFAS frequently persist in hydrophobic micropores, where small pore sizes and tortuous pathways hinder desorption,116 interactions between perfluoroalkyl tails and nonpolar matrix regions further reduce mobility and bioavailability.117
Therefore, information about solvation energy in both the polar solvent and the nonpolar solvent can be obtained from quantum mechanics calculations, from which the bioavailability of the ligands can be inferred.118 We sought a system where we could examine the influence the fluorination on polarity and solvation energy to determine their bioavailability, and 12 organohalogenated ligands (Chart 1) with varying degrees of chlorination and fluorination were calculated for solvation free energy (ΔGsolv) in octanol and water, and the dipole moments were also calculated to serve as a quantitative measure of molecular polarity. These results can provide insights for the influence of chlorination and fluorination degree as well as the relative positions of fluorine substitutions on the molecular polarity and bioavailability.
As shown in Table 1, we observed that with the increasing degree of fluorination and decreasing chlorination yields higher (i.e., less favourable) ΔGsolv in both octanol and water, indicating reduced affinity for water and, even more unexpectedly, for octanol. For example, Cl4 is the most favorable octanol (−2.705 kcal mol−1) and moderately unfavorable in water (+1.889 kcal mol−1), whereas F4 is unfavourable in both (1.633 kcal mol−1 in octanol and 4.011 kcal mol−1 in water), and lower bioavailability. Ligands containing only fluorine substitutions (from F4 to F1H3) generally exhibit a larger dipole and lower log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pow, indicating a greater tendency to partition into polar solvents, indicating a diminished octanol preference and (depending on isomer) somewhat improved water interaction relative to fully perfluorinated, highly symmetric structures.
Pow, indicating a greater tendency to partition into polar solvents, indicating a diminished octanol preference and (depending on isomer) somewhat improved water interaction relative to fully perfluorinated, highly symmetric structures.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pow) were calculated according to the methods described in Section 2
Pow) were calculated according to the methods described in Section 2
| Ligands | ΔGsolv octanol (kcal mol−1) | ΔGsolv water (kcal mol−1) | Dipole moment in water (Debye) | log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pow estimated by MARVIN Pow estimated by MARVIN |
|---|---|---|---|---|
| Cl4 | −2.705 | 1.889 | 0.000 | 2.522 |
| Cl3F1 | −1.700 | 2.374 | 0.590 | 2.230 |
| Cl2F2 | −0.569 | 2.982 | 0.000 | 1.938 |
| Cl2F2-iso1 | −0.567 | 2.971 | 0.924 | 1.938 |
| Cl2F2-iso2 | −0.778 | 2.709 | 0.426 | 1.938 |
| Cl1F3 | 0.461 | 3.418 | 0.450 | 1.646 |
| F4 | 1.633 | 4.011 | 0.000 | 1.354 |
| F3H1 | −0.024 | 1.974 | 1.712 | 1.210 |
| F2H2 | −1.850 | −0.300 | 3.101 | 1.240 |
| F2H2-iso1 | 0.131 | 2.056 | 1.632 | 1.240 |
| F2H2-iso2 | −1.110 | 0.655 | 0.000 | 1.240 |
| F1H3 | −1.118 | 0.471 | 1.819 | 1.097 |
We observed that the dipole moment of ligands in water can be influenced by the fluorination degree or varying positions of fluorine substitution, which can impact the ligands and ΔGsolv in water. Therefore, the relationship between the calculated dipole moment and ΔGsolv in water is identified for the 12 ligands calculated by fitting against the dipole moment and ΔGsolv (Fig. 1). These data showed a general trend, the ΔGsolv in water decreases (more favorable) as the dipole moment increases, reflecting the dominant role of electrostatic and dipole–dipole interactions in polar solvents,119 and therefore higher bioavailability. Symmetry and substitution pattern control these properties: highly symmetric molecules (Cl4, Cl2F2, F4, and F2H2-iso2) have near-zero dipoles due to vector cancellation and therefore show higher (less favorable). In contrast, asymmetric isomers (Cl3F1, Cl2F2-iso1, and F2H2) possess larger dipoles and correspondingly more favorable hydration.
Next, electrostatic potential distributions of the ligands can be obtained with the density functional theory calculation, from which the spatial distribution of partial charges across the molecular surface can be visualized in Fig. 2, thereby offering a qualitative representation of polarity. The charge anisotropy can be visualized (Fig. 2a–f) from the chlorine-substituted ligands display pronounced positive charged regions (σ-holes) on the Cl atom, which enhance specific electrostatic interactions with polar environments.120–122 In contrast, the fluorine atom with strong electronegativity and small atom radius can effectively neutralize the Sigma-hole effect on F (Fig. 2b–f), contributing to the immiscibility of fluorinated compounds with water. This occurs because water–water hydrogen bonding interactions are stronger than water–fluorine interactions.15 Fluorine can amplify the σ-hole on adjacent Cl120,121 (Fig. 2b–d and f), explaining why some mixed Cl/F structures retain localized positive potential yet fully fluorinated analogs do not. Consequently, despite similarly low dipoles, Cl4 hydrates more favorably than F4, consistent with the larger atom size of Cl and higher polarizability that strengthen dispersion interactions with both solvents.
The cis isomers Cl2F2-iso1 and F2H2 yield the largest dipole moments with their respective series (0.924 and 3.101 Debye, respectively) (Fig. 2d and i), and show more favorable hydration. F2H2 exhibits the most favorable ΔGsolv in water (−0.3 kcal mol−1) and favorable solvation in octanol (−1.850 kcal mol−1). Exceptions (e.g., F2H2-iso1) highlight that small changes in substitution can invert solvent preferences. Introducing hydrogen increases polarity and broadens electrostatic potential distributions (Fig. 2h–l), enabling stronger interactions with water and therefore higher bioavailability.123
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pow of ligands decrease with increasing fluorination (Table 1, Cl4 to F4), while ΔGsolv in both octanol and water generally increases, disfavoring bulk partitioning into either phase. Together, these effects promote the accumulation of the ligands at interfaces (air–water and NAPL–water boundaries).113,124,125 Interfacial localization reduces effective concentrations in the aqueous phase and slows diffusive supply to microbes, thereby lowering bioavailability and constraining biodegradation.
Pow of ligands decrease with increasing fluorination (Table 1, Cl4 to F4), while ΔGsolv in both octanol and water generally increases, disfavoring bulk partitioning into either phase. Together, these effects promote the accumulation of the ligands at interfaces (air–water and NAPL–water boundaries).113,124,125 Interfacial localization reduces effective concentrations in the aqueous phase and slows diffusive supply to microbes, thereby lowering bioavailability and constraining biodegradation.
3.3. Binding affinity of ligands affects organofluorine biodegradability
RDases catalyze reductive elimination of halogens from aliphatic and aromatic organohalides.126 PceA, obtained from the anaerobic microorganism Sulfurospirillum multivorans,3 is a practical model for analyzing ligand recognition and specificity. Information about PceA substrate specificity can be obtained from molecular dynamics simulations and binding free energy calculations, from which information about influence of fluorine substitution on ligand binding free energy can be inferred.Therefore, a series of ligands with varying degrees of chlorination/fluorination (Chart 1, Cl4, Cl3F1, Cl2F2, Cl1F3 and F4) were selected for binding free energy calculations. These ligands were selected to test halogen substitution effects while retaining close similarity to known PceA substrates.127,128 The active site of PceA forms an aromatic cage dominated by tyrosine, tryptophan, and phenylalanine side chains, favoring hydrophobic/dispersion contacts.129 To validate the molecular dynamics simulation with the crystal structure of PceA (chain-B of PDB id: 4UR0), we examined the simulation results of the protein structure with the native ligand (trichloroethene). We observed that, across three independent replicates, all non-hydrogen atom RMSDs for the protein and the active site remained below resolution of the crystal structure (1.65 Å), and key residues maintained native orientations (Table S6 and Fig. S6), indicating that the present simulation is in good agreement with the atomic positions from the crystallographic coordinates, and the results are reproducible.
To assess the influence of fluorine atoms on ligand binding, and to determine whether fluorinated ligands can engage in polar interactions with residues in the PceA active site, we examined the ligand binding positions of Cl4 and F4 with respect to the cobalamin and residues surrounding the active site (Fig. 3). Cl4 sits deeply in the pocket with one Cl oriented toward the cobalamin cobalt and extensive van der Waals contacts to Phe38, Tyr246, Trp96, and Trp376, with Phe38 acting as a hydrophobic “gate” that helps retain the ligand (Fig. 3a). Although the F4 ligand was initially placed in the same canonical pocket and orientation as Cl4, it remains associated with this site for approximately the first 60 ns of the trajectory, after which it gradually leaves the canonical pocket and migrates into an adjacent sub-pocket, a small binding pocket that is similarly formed by multiple aromatic residues, including Phe57, Tyr61, Trp96, Tyr102, and Tyr382. This observation indicates that the fully fluorinated ligand is poorly accommodated in the native PceA binding environment and instead preferentially occupies an alternative, rather than the catalytically relevant site (Fig. 3b). The ligand F4 can form hydrogen bonds with polar residues (Fig. 3c). Prior work on fluoroacetate dehalogenase from Rhodopseudomonas shows that polar/halide-binding residues (Ser/Thr/Arg) can stabilize fluoride and enhance defluorination; engineering increased polarity in the pocket improved activity.130 These comparisons suggest that polar contacts may assist defluorination, but in the hydrophobic active site PceA, the polar interaction cannot compensate for reduced dispersion with F-substituted substrates.
As the degrees of fluorination and chlorination modulate ligand polarity, they can substantially influence the thermodynamic binding profile within the enzyme's active site.131,132 To assess the effect of fluorine substitution on enzyme–ligand interactions, Binding free energies were evaluated with respect to the canonical PceA active site pocket (Fig. 4), using snapshots in which each ligand is associated with this site. For F4, this corresponds to the portion of the trajectory during which it resides in the canonical site (the first 60 ns), prior to its migration into the adjacent sub-pocket. This definition allows a direct comparison of how progressively fluorinated ligands interact with the native Cl4 binding environment, even though F4 does not remain stably bound there. The calculated binding free energies shows a monotonic loss of affinity with increasing fluorination (Fig. 4a), from −15.19 ± 0.22 kcal mol−1 for Cl4 to −7 ± 0.41 kcal mol−1 for F4, indicating that higher degrees of fluorination result in less favourable binding. This trend suggests that PceA exhibits strong substrate specificity toward chlorinated ethenes.
To further analyze these interactions, the binding free energy was decomposed into four components: van der Waals energy, electrostatic energy, polar solvation energy, and SASA (non-polar solvation energy).80 van der Waals interactions are the dominant favorable contribution (Fig. 4b), which weakens from −19.62 ± 0.27 kcal mol−1 (Cl4) to −9.44 ± 0.05 kcal mol−1 (F4), consistent with the halogen van der Waals radius order I > Br > Cl ≫ F.133 The smaller atom of F lowers molecular polarizability and reduces contact area in a hydrophobic pocket.134 The non-polar solvation energies contribution changes only modestly and becomes less favorable with increasing fluorination (shifting from −2.09 ± 0.03 kcal mol−1 for Cl4 to −1.51 ± 0.02 kcal mol−1 for F4). Electrostatic interaction energies are small and show no clear correlation with halogen substitution pattern. Polar solvation energies are positive and oppose ligand binding, which decrease from 6.50 ± 0.15 kcal mol−1 (Cl4) to 3.97 ± 0.26 kcal mol−1 (F4), reflecting a smaller desolvation penalty for the unfavourable solvated fluorinated ligands. This Polar solvation energy term differs conceptually from the standalone ΔGsolv in Table 1; here, it measures the electrostatic cost of moving the ligand from water into the nonpolar pocket during binding.81 Overall, the loss of van der Waals stabilization with fluorination outweighs the slightly lower desolvation penalty, yielding weaker net binding.
These results align with the literature showing stronger binding for larger (iodinated ligands vs. brominated analogs135 and chlorinated aromatic ligands vs. fluorinated analogs136) more polarizable halogens in hydrophobic binding pockets. In PceA, dispersion dominates, so atomic size and polarizability govern affinity. Consequently, highly fluorinated substrates bind weakly and are less stably accommodated in the active site, helping to explain their poor biodegradability by RDases.
3.4. Reductive defluorination energy barriers limit the bio-defluorination
The activation energy barrier can limit the proceeding of a chemical reaction. While PceA efficiently lowers the barrier for reductive dechlorination,10 its activity toward defluorination is constrained by the intrinsically higher activation energy for C–F cleavage, which is the strongest single bond in organic chemistry.108,137–139To calculate the energy barrier of reductive defluorination by PceA, the QM cluster models for transition-state searches were built from representative configurations in which each ligand occupies a catalytically competent pose in the canonical active site pocket (i.e., the same site as Cl4), irrespective of its equilibrium residence time in that pose. Thus, the calculated energy barrier reported below probe the intrinsic reactivity of each ligand toward C–X bond cleavage in the native PceA environment, complementing the MD-based analysis of binding preferences.
We calculated reductive defluorination with PceA, using the concerted proton-coupled electron transfer (PCET) mechanism established for dechlorination.4 Briefly, an electron reduces Co(II) to Co(I), Tyr246 is protonated, ligand binding generates a concerted transition-state, Tyr246 donates H+ to the substrate carbon, and halide departure occurs simultaneously with electron transfer (Fig. 5). Due to the lack of experimental data of energy profile associated with reductive dechlorination or defluorination with PceA, to validate our cluster model, we optimized the reactant state and transition-state of the cluster model with Cl4 as the ligand, and the calculated energy barrier of the Cl4 dechlorination is (12.69 kcal mol−1) in good agreement with the literature (12.5 kcal mol−1).4 Therefore, this cluster model will be used for calculating the energy barrier of reductive dechlorination of other organofluorinated ligands.
 | ||
| Fig. 5 PceA proton coupled electron transfer (PCET) dehalogenation mechanism.4 The reaction mechanism involves the initial coordination of the chlorinated substrate to the Co(II) center, followed by PCET that reduces Co(II) to Co(I), and protonates Tyr246. Then, the concerted transition-state forms where Tyr246 donates H+ to the substrate carbon as the C–Cl bond cleaves and generates a transient Co(III)–substrate complex. Chloride departs, H substitutes, and the cofactor returns to Co(II), completing the catalytic cycle. | ||
We present the optimized cluster model of PceA with Cl3F1 in a transition-state (Fig. 6a).
In this transition-state, the key residue Tyr246 donates a proton to the carbon atom with the leaving fluorine atom, which is simultaneously coordinated to the cobalt center (Fig. 6a). For each ligand, the transition state region was first located by a 2D relaxed scan along the C–F and H–C coordinates, and the highest energy point along the minimum-energy path was then used as the initial guess for a full transition state optimization (Fig. 6b). In a representative case (Cl3F1), the transition-state occurs with a C–F distance of 1.65 Å, a H–C distance of 1.84 Å, a F–Co distance of 2.11 Å and a H (from Arg305)–O (from Tyr246) distance of 1.90 Å (Fig. 6c).
The reductive defluorination energy barriers of all calculated ligands greatly exceed the Cl4 dechlorination barrier (Fig. 7a–d), consistent with previous work by Liao et al.,139 in which the energy barrier for defluorination catalyzed by the dehalogenase NpRdhA was reported as 36.6 kcal mol−1, compared to 16.6 kcal mol−1 for dechlorination. Among mixed Cl/F series (Fig. 7a and c), the lowest barriers were obtained for Cl2F2-iso1 and Cl1F3 (∼34 kcal mol−1), still more than 2 times higher than dechlorination. Substituent arrangement can influence the energy barrier, by placing Cl on the reacting carbon and aligning a neighboring F on the same face modestly lowers the barrier, likely by polarizing the C–F bond and stabilizing the concerted transition-state. Fully F/H-substituted ligands (Fig. 7b and d) display even higher barriers (often >40 kcal mol−1), with one exception (F1H3, 22.96 kcal mol−1). This trend can be attributed to the presence of hydrogen atoms, which are less electronegative and exert a weaker electron-withdrawing effect than chlorine. As a result, the C–F bonds in these molecules are less polarized, making them more resistant to cleavage and thereby increasing the energy barrier.
 | ||
| Fig. 7 Energy barrier of PceA-catalyzed defluorination calculation and dechlorination. (a) The energy profile for defluorination of Cl4 to F4 and (b) the energy profile for defluorination of Cl4 and F3H1 to F1H3. (c) and (d) Reaction coordinate profiles (React, Int and TS) for the same sets with Cl4 dechlorination (12.69 kcal mol−1) as the reference (−0.45 V). Energies from a DFT cluster model with single point calculation (Co LANL08; others 6-311+G(2d,2p)) and SMD (ε = 4) solvation. These energies can be approximated as the free energies in solution, the same as in the case of the approach adopted previously by Liao et al.4 | ||
Increasing fluorination strengthens C–F bonds and increases energy barriers. Fluoroacetate defluorination by fluoroacetate dehalogenase increases from 11.2 to 24.4 kcal mol−1.115 Each added F increases the positive charge at the α-carbon and the negative charge on F, enhancing Coulombic stabilization, while the negative hyperconjugation further reinforces the bond.15 Together, these electrostatic and orbital effects elevate bond dissociation energy barriers for highly fluorinated substrates.
To evaluate how fluorination affects PceA defluorination barriers, Table 2 reports key transition-state distances (H–C, C–F, and F–Co) for each ligand, and Fig. S9 (SI) correlates these distances with activation energies.
| Cl4 | Cl3F1 | Cl2F2 | Cl2F2 iso1 | Cl2F2 iso2 | Cl1F3 | Cl1F3-conf1 | Cl1F3-conf2 | |
|---|---|---|---|---|---|---|---|---|
| H–C distance (Å) | 2.32 | 1.84 | 1.80 | 1.84 | 1.73 | 1.79 | 1.69 | 1.72 |
| C–X distance (Å) | 1.96 | 1.65 | 1.62 | 1.66 | 1.65 | 1.65 | 1.62 | 1.65 |
| X–Co distance (Å) | 2.54 | 2.12 | 2.15 | 2.12 | 2.09 | 2.14 | 2.12 | 2.10 |
| C–X–Co angle (degree) | 173.61 | 163.17 | 160.38 | 170.59 | 165.50 | 167.52 | 162.80 | 167.05 |
| Imaginary frequency (i cm−1) | −135.02 | −418.75 | −429.25 | −414.52 | −399.08 | −429.59 | −411.45 | −394.23 |
| F4 | F3H1 | F3H1-conf1 | F3H1-conf2 | F2H2 | F2H2-iso1 | F2H2-iso2 | F1H3 | |
|---|---|---|---|---|---|---|---|---|
| H–C distance (Å) | 1.68 | 1.78 | 1.73 | 1.89 | 1.78 | 1.94 | 1.88 | 1.73 |
| C–X distance (Å) | 1.63 | 1.71 | 1.73 | 1.73 | 1.80 | 1.88 | 1.79 | 1.94 |
| X–Co distance (Å) | 2.12 | 2.06 | 2.06 | 2.07 | 2.01 | 1.99 | 2.03 | 1.98 |
| C–X–Co angle (degree) | 164.95 | 164.43 | 165.06 | 166.73 | 164.29 | 168.08 | 165.38 | 164.67 |
| Imaginary frequency (i cm−1) | −412.24 | −434.35 | −414.69 | −393.22 | −432.51 | −306.89 | −406.61 | −399.40 |
From results in Table 2, when comparing the geometry of optimized transition states between reductive dechlorination and defluorination, we observed shorter H–C, C–F, and F–Co distances and smaller C–F–Co angles, indicating tighter, more bent transition-states, which involve later proton transfer and later bond cleavage than that of the C–Cl bond. Previous study links shorter halogen–Co to higher barriers,139 which are less thermodynamically favorable. Across our ligand set, however, correlations between the barrier height and individual geometric descriptors are weak (Fig. S9). The transition-state could occur later with longer C–F distance, even with lower energy barrier, for example, Cl1F3-conf2 (37.47 kcal mol−1, H–C = 1.72 Å) versus Cl3F1 (39.62 kcal mol−1, H–C = 1.84 Å) shows that a less-advanced proton transfer can coincide with a higher barrier. Thus, transition-state geometry reflects a multifactor balance (dispersion, electrostatics, and local polarity) rather than a single controlling distance.
With the established differences in activation energy barriers between reductive dechlorination and defluorination, we estimated the relative reaction kinetics by using the Eyring equation.92–95 Derived from transition state theory, the Eyring equation describes a linear free-energy relationship, whereby reactions with lower activation energies are thermodynamically more favorable, proceed at faster rates. Conversely, less thermodynamically favorable reactions are slower.140 Applying this framework, reaction rate constants were estimated based on the activation energies calculated in the present study, as summarized in Table 3. These values can provide quantitative insights into the substantial kinetic disadvantage associated with enzymatic defluorination relative to dechlorination.
| Cl4 | Cl3F1 | Cl2F2 | Cl2F2 iso1 | Cl2F2 iso2 | Cl1F3 | Cl1F3-conf1 | Cl1F3-conf2 | |
|---|---|---|---|---|---|---|---|---|
| Energy barrier (kcal mol−1) | 12.69177 | 34.53463 | 34.28687 | 32.65786 | 41.97199 | 29.30322 | 45.89149 | 41.23589 |
| Relative difference in reaction kinetics (mol s−1) | 3.55 × 103 | 8.17 × 1015 times slower | 5.4 × 1015 times slower | 3.51 × 1014 times slower | 2.14 × 1021 times slower | 1.26 × 1012 times slower | 1.53 × 1024 times slower | 6.23 × 1020 times slower |
| F4 | F3H1 | F3H1-conf1 | F3H1-conf2 | F2H2 | F2H2-iso1 | F2H2-iso2 | F1H3 | |
|---|---|---|---|---|---|---|---|---|
| Energy barrier (kcal mol−1) | 34.31208 | 50.42838 | 51.70852 | 50.68764 | 55.65925 | 39.65456 | 46.75546 | 43.22858 |
| Relative difference in reaction kinetics (mol s−1) | 5.62 × 1015 times slower | 3.09 × 1027 times slower | 2.65 × 1028 times slower | 4.78 × 1027 times slower | 2.01 × 1031 times slower | 4.4 × 1019 times slower | 6.54 × 1024 times slower | 1.76 × 1022 times slower |
All defluorination reactions are vastly slower, ranging from the smallest difference of 1.26 × 1012 times slower (Cl1F3) to the highest difference of 2.01 × 1031 times (F2H2). These kinetics render anaerobic enzymatic reductive defluorination of linear PFAS impractical for engineered remediation, which can be attributed, in part, to the absence of finely tuned polar/halide-stabilizing residues needed to stabilize fluorinated substrates and the nascent fluoride in the PCET transition state. The binding free energy weakens with increasing fluorination as van der Waals contacts diminish due to the smaller atom radius of fluorine (Section 3.3). In contrast, fluoroacetate dehalogenase (FAcD) uses Arg111 and Arg114 to hydrogen bond the substrate carboxylate and Asp110 as a nucleophile, and the released F− is further stabilized with a hydrogen bond by His155, Trp156, and Tyr219.46,107,141–144 Haloacid dehalogenase (DeHa2) likewise employs polar residues Arg/Ser/Thr to orient the substrate, cleave C–F, and expel fluoride.145 Without such features, PceA cannot effectively stabilize the C–F PCET transition state, making defluorination thermodynamically and kinetically unfavorable. These constraints indicate that anaerobic RDase-based biodefluorination of linear PFAS is not practicable at engineering scales.
The toxicity of released fluoride ions may also hinder microbial adaptation toward reductive defluorination, in which the released F− can induce oxidative stress and perturb redox balance, necessitating efficient fluoride export mechanisms.36,135,146 Functional fluoride efflux transporters are identified to be essential for the reductive defluorination of branched polyfluorocarboxylic acids by Acetobacterium spp.136 Nonetheless, the microbial reductive defluorination of PFAS with a linear structure remains a significant challenge. Practical strategies include pretreatment to generate more labile fragments, specialized consortia, and enzyme engineering to introduce polar/halide-binding residues.35,136 The computational workflow here can serve as a pre-market biodegradability screen for new compounds, informing regulatory assessments and helping prevent persistent pollutants from entering commerce.
4. Conclusions
The present study critically evaluates the challenges associated with microbial reductive dehalogenation of per- and polyfluoroalkyl substances (PFASs), with a specific focus on the reductive dehalogenase enzyme PceA. It begins by addressing the geochemical scarcity of fluorine on Earth, which has limited microbial evolutionary exposure to organofluorinated compounds and impeded the development of effective enzymatic degradation pathways. The physicochemical properties of PFAS, particularly the low polarity, the high hydrophobicity, and the strong C–F bonds, were shown to significantly reduce bioavailability and hinder enzymatic recognition. The degree of fluorination and relative position of fluorine substitution can influence the molecular charge distribution. The degree of fluorination can also influence the van der Waals interaction within the hydrophobic active site of PceA, which can be crucial in substrate recognition, and ultimately lead to poor accommodation fluorinated ligands, due to unfavourable van der Waals interactions and non-polar solvation energies. A tetrafluorinated ligand is preferentially accommodated by a sub-pocket away from the catalytic site. Although polar residues may partially stabilize fluorine atoms through electrostatic interactions or hydrogen bonding, this effect is insufficient to overcome the intrinsic energetic constraints. Quantum chemical calculations further demonstrated that PFAS compounds exhibit significantly higher activation barriers for C–F bond cleavage compared to C–Cl bond cleavage, resulting in markedly slower reaction kinetics. Taken together, these findings from a theoretical and mechanistic perspective indicate that the exploitation of organohalide-respiring bacteria for the anaerobic reductive defluorination of linear PFAS is not feasible on an engineering scale. This study underscores the urgent need for alternative technologies that are better suited to mitigate the environmental and health impacts of these persistent and recalcitrant pollutants.Author contributions
The manuscript was written through contributions of all authors. All authors gave approval to the final version of the manuscript. M. M. conceived and designed research and edited this manuscript. Y. R. undertook the computation works and wrote the original draft. W. D. instructed and reviewed all the density functional theory calculations. All authors read and approved the manuscript.Conflicts of interest
The authors declare no competing financial interest.Data availability
The data underlying this study for validation, and reproducing the results are openly available and free of charge in the GitHub repository: https://github.com/feammox/Theoretical_limitation_PFAS_degradation.git. The data include the: solvation free energy ΔGsolv calculations: the Gaussian 16 package output log files, which can be used to generate the input calculation details and obtain the outputs energy results, he molecular dynamics simulation by Gromacs/2024.3-gpuvolta: the molecular mechanics parameters files, including topology files of all ligands, cobalamin cofactor, iron–sulfur cluster, and PceA protein forcefield parameters, which can be used to generate the input files for molecular dynamics simulation by Gromacs/2024.3-gpuvolta. the output files of binding free energy calculation by using g_mmpbsa, the quantum mechanics calculation for the energy barrier of reductive defluorination: the frequency calculation results, including the Cartesian coordinates of each system and energy calculation results. The example input file for large basis set and solvation effect calculations can be found in the Cl4 folder. The software availability: the Gaussian software can be purchased from https://gaussian.com/, the Gromacs software can be downloaded from https://www.gromacs.org/ and the crystal structure of PceA (PDB id: 4UQU) can be downloaded from https://www.rcsb.org/.Supplementary information (SI): the detailed force field parameters of cobalamin and ligands, including atomic names, types, charges, ε and σ, bonds, angles and dihedrals of cobalamin and ligands, validation results of cobalamin cofactor parameters, validation of PceA molecular dynamics simulation, the cluster model of PceA for QM calculation, with the fixed atoms indicated and the Cartesian coordinates of all density functional theory of optimized cluster structures. See DOI: https://doi.org/10.1039/d5cp03866a.
Acknowledgements
Y. R. is supported by the Research Training Program (RTP) Scholarship, provided by the Australian Government. We acknowledge the discussions with the computational chemist Junming Ho from the School of Chemistry, University of New South Wales, Sydney Australia, who provided critical reading and valuable suggestions for this manuscript. We acknowledge the funding from the Australian Research Council and Tetratech Coffey for LP170100116. We also acknowledge the computation resources from UNSW HPC scheme 10.26190/PMN5-7J50.References
- J. Koenig, M. Lee and M. Manefield, Rev. Environ. Sci. Biotechnol., 2014, 14, 49–71 CrossRef.
- B. Kräutler, W. Fieber, S. Ostermann, M. Fasching, K. H. Ongania, K. Gruber, C. Kratky, C. Mikl, A. Siebert and G. Diekert, Helv. Chim. Acta, 2003, 86, 3698–3716 CrossRef.
- M. Bommer, C. Kunze, J. Fesseler, T. Schubert, G. Diekert and H. Dobbek, Science, 2014, 346, 455–458 CrossRef CAS PubMed.
- R. Z. Liao, S. L. Chen and P. E. Siegbahn, Chemistry, 2016, 22, 12391–12399 CrossRef CAS PubMed.
- X. Zhang, Z. Wang, Z. Li, S. Shaik and B. Wang, ACS Catal., 2023, 13, 1173–1185 CrossRef CAS.
- Y. K. Wong, S. I. Holland, H. Ertan, M. Manefield and M. Lee, Environ. Microbiol., 2016, 18, 3092–3105 CrossRef CAS PubMed.
- M. Lee, A. Low, O. Zemb, J. Koenig, A. Michaelsen and M. Manefield, Environ. Microbiol., 2012, 14, 883–894 CrossRef CAS PubMed.
- S. Tang, P. H. Wang, S. A. Higgins, F. E. Loffler and E. A. Edwards, Front. Microbiol., 2016, 7, 100 Search PubMed.
- S. Tang and E. A. Edwards, Philos. Trans. R. Soc. London, Ser. B Philos. Trans.: Biol. Sci., 2013, 368, 20120318 CrossRef PubMed.
- A. Neumann, G. Wohlfarth and G. Diekert, J. Bacteriol., 1998, 180, 4140–4145 CrossRef CAS PubMed.
- A. Neumann, A. Siebert, T. Trescher, S. Reinhardt, G. Wohlfarth and G. Diekert, Arch. Microbiol., 2002, 177, 420–426 CrossRef CAS PubMed.
- P. J. McMurdie, S. F. Behrens, J. A. Muller, J. Goke, K. M. Ritalahti, R. Wagner, E. Goltsman, A. Lapidus, S. Holmes, F. E. Loffler and A. M. Spormann, PLoS Genet., 2009, 5, e1000714 CrossRef PubMed.
- C. F. Kwiatkowski, D. Q. Andrews, L. S. Birnbaum, T. A. Bruton, J. C. DeWitt, D. R. U. Knappe, M. V. Maffini, M. F. Miller, K. E. Pelch, A. Reade, A. Soehl, X. Trier, M. Venier, C. C. Wagner, Z. Wang and A. Blum, Environ. Sci. Technol. Lett., 2020, 7, 532–543 CrossRef CAS PubMed.
- A. v Bondi, J. Phys. Chem., 1964, 68, 441–451 CrossRef CAS.
- D. M. Lemal, J. Org. Chem., 2004, 69, 1–11 CrossRef CAS PubMed.
- C. A. Moody and J. A. Field, Environ. Sci. Technol., 2000, 34, 3864–3870 CrossRef CAS.
- E. Kissa, Fluorinated surfactants and repellents, CRC Press, 2001 Search PubMed.
- R. C. Buck, J. Franklin, U. Berger, J. M. Conder, I. T. Cousins, P. De Voogt, A. A. Jensen, K. Kannan, S. A. Mabury and S. P. van Leeuwen, Integr. Environ. Assess. Manage., 2011, 7, 513–541 CrossRef CAS PubMed.
- J. L. Sims, K. M. Stroski, S. Kim, G. Killeen, R. Ehalt, M. F. Simcik and B. W. Brooks, Sci. Total Environ., 2022, 816, 151535 CrossRef CAS PubMed.
- B. Lopez, P. Ollivier, A. Togola, N. Baran and J.-P. Ghestem, Sci. Total Environ., 2015, 518, 562–573 CrossRef PubMed.
- W. J. Backe, T. C. Day and J. A. Field, Environ. Sci. Technol., 2013, 47, 5226–5234 CrossRef CAS PubMed.
- G. Munoz, P. Labadie, F. Botta, F. Lestremau, B. Lopez, E. Geneste, P. Pardon, M. H. Devier and H. Budzinski, Sci. Total Environ., 2017, 607–608, 243–252 Search PubMed.
- E. Hepburn, C. Madden, D. Szabo, T. L. Coggan, B. Clarke and M. Currell, Environ. Pollut., 2019, 248, 101–113 CrossRef CAS PubMed.
- D. M. J. Shaw, G. Munoz, E. M. Bottos, S. V. Duy, S. Sauve, J. Liu and J. D. Van Hamme, Sci. Total Environ., 2019, 647, 690–698 CrossRef CAS PubMed.
- V. Mendez, S. Holland, S. Bhardwaj, J. McDonald, S. Khan, D. O'Carroll, R. Pickford, S. Richards, C. O'Farrell, N. Coleman, M. Lee and M. J. Manefield, Sci. Total Environ., 2022, 829, 154587 CrossRef CAS PubMed.
- S. Bhardwaj, M. Lee, D. O'Carroll, J. McDonald, K. Osborne, S. Khan, R. Pickford, N. Coleman, C. O’Farrell and S. Richards, J. Hazard. Mater., 2024, 478, 135510 CrossRef CAS PubMed.
- S.-H. Yang, Y. Shi, M. Strynar and K.-H. Chu, J. Hazard. Mater., 2022, 423, 127052 CrossRef CAS PubMed.
- J. Liu, L. S. Lee, L. F. Nies, C. H. Nakatsu and R. F. Turco, Environ. Sci. Technol., 2007, 41, 8024–8030 CrossRef CAS PubMed.
- M. H. Kim, N. Wang and K. H. Chu, Appl. Microbiol. Biotechnol., 2014, 98, 1831–1840 CrossRef CAS PubMed.
- M. H. Kim, N. Wang, T. McDonald and K. H. Chu, Biotechnol. Bioeng., 2012, 109, 3041–3048 CrossRef CAS PubMed.
- F. Torres, V. Ochoa-Herrera, P. Blowers and R. Sierra-Alvarez, Chemosphere, 2009, 76, 1143–1149 CrossRef CAS PubMed.
- R. McGregor and Y. Zhao, Biorem. J., 2021, 31, 7–17 Search PubMed.
- B. Xu, S. Liu, J. L. Zhou, C. Zheng, J. Weifeng, B. Chen, T. Zhang and W. Qiu, J. Hazard. Mater., 2021, 412, 125159 Search PubMed.
- T. H. Christensen, P. L. Bjerg, S. A. Banwart, R. Jakobsen, G. Heron and H.-J. Albrechtsen, J. Contam. Hydrol., 2000, 45, 165–241 CrossRef CAS.
- Y. Yu, K. Zhang, Z. Li, C. Ren, J. Chen, Y. H. Lin, J. Liu and Y. Men, Environ. Sci. Technol., 2020, 54, 14393–14402 CrossRef CAS PubMed.
- L. P. Wackett, Microb. Biotechnol., 2021 DOI:10.1111/1751-7915.13928.
- S. D. Copley, Curr. Opin. Struct. Biol., 2021, 69, 41–49 CrossRef CAS PubMed.
- S. D. Copley, Phys. Biol., 2020 DOI:10.1088/1478-3975/ab8697.
- S. D. Copley, Nat. Chem. Biol., 2009, 5, 559–566 Search PubMed.
- Y. Ren and M. Manefield, Appl. Microbiol. Biotechnol., 2025, 109, 1–10 Search PubMed.
- M. Manefield, M. Lee and J. Koenig, in Microbial ecology of extreme environments, Springer, 2017, pp. 201–213 Search PubMed.
- P. Heinrich and T. Braunbeck, Comp. Biochem. Physiol., Part C:Toxicol. Pharmacol., 2019, 221, 59–67 CAS.
- E. R. Christensen, Y. Wang, J. Huo and A. Li, J. Environ. Chem, Eng., 2022, 10, 107201 CrossRef CAS.
- S. Genheden and U. Ryde, Expert. Opin. Drug Discovery, 2015, 10, 449–461 Search PubMed.
- J. Kua, Y. Zhang and J. A. McCammon, J. Am. Chem. Soc., 2002, 124, 8260–8267 CrossRef CAS PubMed.
- Y. Yue, J. Chen, L. Bao, J. Wang, Y. Li and Q. Zhang, Chemosphere, 2020, 254, 126803 CrossRef CAS PubMed.
- A. Williams, Chem. Soc. Rev., 1994, 23, 93–100 RSC.
- C. Kunze, M. Bommer, W. R. Hagen, M. Uksa, H. Dobbek, T. Schubert and G. Diekert, Nat. Commun., 2017, 8, 15858 CrossRef CAS PubMed.
- J. Ho, A. Klamt and M. L. Coote, J. Phys. Chem. A, 2010, 114, 13442–13444 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, F. Williams, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16 Rev. C.01, Gaussian Inc., Wallingford CT, 2016 Search PubMed.
- Y. Zhao, N. E. Schultz and D. G. Truhlar, J. Chem. Theory Comput., 2006, 2, 364–382 CrossRef PubMed.
- V. Kundi and J. Ho, J. Phys. Chem. B, 2019, 123, 6810–6822 CrossRef CAS PubMed.
- V. N. Viswanadhan, A. K. Ghose, G. R. Revankar and R. K. Robins, J. Chem. Inf. Comput. Sci., 1989, 29, 163–172 CrossRef CAS.
- T. Lu and F. Chen, J. Comput. Chem., 2012, 33, 580–592 CrossRef CAS PubMed.
- J. Zhang and T. Lu, Phys. Chem. Chem.Phys., 2021, 23, 20323–20328 RSC.
- T. Lu and F. Chen, J. Mol. Graphics Modell., 2012, 38, 314–323 CrossRef CAS PubMed.
- S. Manzetti and T. Lu, J. Phys. Org. Chem., 2013, 26, 473–483 CrossRef CAS.
- T. Lu and S. Manzetti, Struct. Chem., 2014, 25, 1521–1533 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. A. Jr., J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, E. Brothers, K. N. Kudin, V. N. Staroverov, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, Ö. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian 09 Revision A.2, Gaussian Inc., Wallingford CT, 2009 Search PubMed.
- T. Lu and F. Chen, J. Comput. Chem., 2012, 33, 580–592 CrossRef CAS PubMed.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, J. Comput. Chem., 2004, 25, 1157–1174 CrossRef CAS PubMed.
- A. K. Rappé, C. J. Casewit, K. Colwell, W. A. Goddard III and W. M. Skiff, J. Am. Chem. Soc., 1992, 114, 10024–10035 CrossRef.
- N. Keot and M. Sarma, J. Phys. Chem. A, 2024, 128, 8346–8359 CrossRef CAS PubMed.
- D. Li, T. Sun, T. Ma, W. Zhang, Q. Sun, M. Cheng, Z. Zha, W. Xie and Z. Tao, Adv. Funct. Mater., 2024 DOI:10.1002/adfm.202405145.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, J. Chem. Theory Comput., 2015, 11, 3696–3713 CrossRef CAS PubMed.
- S. Jo, T. Kim, V. G. Iyer and W. Im, J. Comput. Chem., 2008, 29, 1859–1865 CrossRef CAS PubMed.
- W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. Impey and M. Klein, J. Chem. Phys., 1983, 79, 926–935 Search PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1, 19–25 CrossRef.
- M. Abraham, A. Alekseenko, V. Basov, C. Bergh, E. Briand, A. Brown, M. Doijade, G. Fiorin, S. Fleischmann, S. Gorelov, G. Gouaillardet, A. Grey, M. E. Irrgang, F. Jalalypour, J. Jordan, C. Kutzner, J. A. Lemkul, M. Lundborg, P. Merz, V. Miletic, D. Morozov, J. Nabet, S. Pall, A. Pasquadibisceglie, M. Pellegrino, H. Santuz, R. Schulz, T. Shugaeva, A. Shvetsov, A. Villa, S. Wingbermuehle, B. Hess and E. Lindahl, Gromacs, 2024 Search PubMed.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 014101 Search PubMed.
- M. Parrinello and A. Rahman, J. Appl. Phys., 1981, 52, 7182–7190 CrossRef CAS.
- Q. Ke, X. Gong, S. Liao, C. Duan and L. Li, J. Mol. Liq., 2022, 365, 120116 CrossRef CAS.
- H. Grubmüller, H. Heller, A. Windemuth and K. Schulten, Mol. Simul., 1991, 6, 121–142 CrossRef.
- H. G. Petersen, J. Chem. Phys., 1995, 103, 3668–3679 CrossRef CAS.
- B. Hess, H. Bekker, H. J. Berendsen and J. G. Fraaije, J. Comput. Chem., 1997, 18, 1463–1472 CrossRef CAS.
- J.-P. Ryckaert, G. Ciccotti and H. J. Berendsen, J. Comput. Phys., 1977, 23, 327–341 CrossRef CAS.
- P. A. Kollman, I. Massova, C. Reyes, B. Kuhn, S. Huo, L. Chong, M. Lee, T. Lee, Y. Duan and W. Wang, Acc. Chem. Res., 2000, 33, 889–897 CrossRef CAS PubMed.
- M. K. Gilson and B. Honig, Proteins: Struct., Funct., Bioinf., 1988, 4, 7–18 CrossRef CAS PubMed.
- N. Homeyer and H. Gohlke, Mol. Inform., 2012, 31, 114–122 CrossRef CAS PubMed.
- R. Kumari, R. Kumar, Open Source Drug Discovery and A. Lynn, J. Chem. Inf. Model., 2014, 54, 1951–1962 CrossRef CAS PubMed.
- J. B. Chaires, Annu. Rev. Biophys., 2008, 37, 135–151 Search PubMed.
- C. Wang, D. Greene, L. Xiao, R. Qi and R. Luo, Front. Mol. Biosci., 2017, 4, 87 Search PubMed.
- N. Michaud-Agrawal, E. J. Denning, T. B. Woolf and O. Beckstein, J. Comput. Chem., 2011, 32, 2319–2327 CrossRef CAS PubMed.
- F. Himo, J. Am. Chem. Soc., 2017, 139, 6780–6786 CrossRef CAS PubMed.
- M. E. Lind and F. Himo, Angew. Chem., Int. Ed., 2013, 52, 4563–4567 CrossRef CAS PubMed.
- M. E. S. Lind and F. Himo, ACS Catal., 2016, 6, 8145–8155 Search PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 Search PubMed.
- L. E. Roy, P. J. Hay and R. L. Martin, J. Chem. Theory Comput., 2008, 4, 1029–1031 Search PubMed.
- M. Prejanò, T. Marino and N. Russo, Front. Chem., 2018, 6, 249 Search PubMed.
- M. R. Blomberg, T. Borowski, F. Himo, R.-Z. Liao and P. E. Siegbahn, Chem. Rev., 2014, 114, 3601–3658 Search PubMed.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, J. Phys. Chem. B, 2009, 113, 6378–6396 CrossRef CAS PubMed.
- H. Eyring, Trans. Faraday Soc., 1938, 34, 41–48 RSC.
- H. Eyring, Chem. Rev., 1935, 17, 65–77 CrossRef CAS.
- H. Eyring, J. Chem. Phys., 1935, 3, 107–115 CrossRef CAS.
- J. J. Rooney, J. Mol. Catal. A: Chem., 1995, 96, L1–L3 CrossRef CAS.
- G. W. Gribble, Prog. Chem. Org. Nat. Prod., 1996, 1–423 CAS.
- N. Lahav, S. Nir and A. C. Elitzur, Prog. Biophys. Biophys. Chem., 2001, 75, 75–120 CrossRef CAS PubMed.
- B. E. Jugder, H. Ertan, S. Bohl, M. Lee, C. P. Marquis and M. Manefield, Front. Microbiol., 2016, 7, 249 Search PubMed.
- R. E. Richardson, Curr. Opin. Biotechnol, 2013, 24, 498–505 CrossRef CAS PubMed.
- L. A. Hug, F. Maphosa, D. Leys, F. E. Loffler, H. Smidt, E. A. Edwards and L. Adrian, Philos. Trans. R. Soc. London, Ser. B Philos. Trans.: Biol. Sci., 2013, 368, 20120322 CrossRef PubMed.
- P. L. McCarty, In situ remediation of chlorinated solvent plumes, 2010, pp. 1–28 Search PubMed.
- S. Woosley and W. Haxton, Nature, 1988, 334, 45–47 CrossRef CAS.
- H. Jönsson and N. Ryde, in Modern Synthesis Processes and Reactivity of Fluorinated Compounds, Elsevier, 2017, pp. 1–6 Search PubMed.
- K. T. Koga and E. F. Rose-Koga, C. R. Chim, 2018, 21, 749–756 CrossRef CAS.
- G. W. Gribble, in Organofluorines, 2002, ch. 5, pp. 121–136 DOI:10.1007/10721878_5.
- L. G. T. Gaines, Am. J. Ind. Med., 2023, 66, 353–378 CrossRef CAS PubMed.
- Y. Yue, J. Fan, G. Xin, Q. Huang, J. B. Wang, Y. Li, Q. Zhang and W. Wang, Environ. Sci. Technol., 2021 DOI:10.1021/acs.est.0c08811.
- Y. Li, R. Zhang, L. Du, Q. Zhang and W. Wang, Catal. Sci. Technol., 2016, 6, 73–80 RSC.
- D. Sijm, R. Kraaij and A. Belfroid, Environ. Pollut., 2000, 108, 113–119 CrossRef CAS PubMed.
- M. L. Brusseau, Water Res., 2021, 190, 116778 CrossRef CAS PubMed.
- M. Garcia-Junco, C. Gomez-Lahoz, J.-L. Niqui-Arroyo and J.-J. Ortega-Calvo, Environ. Sci. Technol., 2003, 37, 2988–2996 CrossRef CAS PubMed.
- S. Ghoshal, A. Ramaswami and R. G. Luthy, Environ. Sci. Technol., 1996, 30, 1282–1291 CrossRef CAS.
- H. Sharifan, M. Bagheri, D. Wang, J. G. Burken, C. P. Higgins, Y. Liang, J. Liu, C. E. Schaefer and J. Blotevogel, Sci. Total Environ., 2021, 771, 145427 CrossRef CAS PubMed.
- S. Suganthi, V. Kannappan and V. Sathyanarayanamoorthi, DJ J. Eng. Chem. Fuel, 2016, 1, 1–14 CrossRef.
- D. D. Do, Adsorption analysis: Equilibria and kinetics (with cd containing computer MATLAB programs), World Scientific, 1998 Search PubMed.
- H. Cheng, E. Hu and Y. Hu, J. Contam. Hydrol., 2012, 129–130, 80–90 CrossRef CAS PubMed.
- C. J. Newell, D. T. Adamson, P. R. Kulkarni, B. N. Nzeribe, J. A. Connor, J. Popovic and H. F. Stroo, Groundwater Monit. Rem., 2021, 41, 76–89 CrossRef.
- G. D. R. Matos, D. Y. Kyu, H. H. Loeffler, J. D. Chodera, M. R. Shirts and D. L. Mobley, J. Chem. Eng. Data, 2017, 62, 1559–1569 CrossRef PubMed.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, J. Chem. Theory Comput., 2008, 4, 877–887 CrossRef CAS PubMed.
- T. Clark, M. Hennemann, J. S. Murray and P. Politzer, J. Mol. Model., 2007, 13, 291–296 CrossRef CAS PubMed.
- M. A. Ibrahim, J. Comput. Chem., 2011, 32, 2564–2574 CrossRef CAS PubMed.
- J. Y. C. Lim and P. D. Beer, Chem, 2018, 4, 731–783 CAS.
- J. Biernacka, K. Betlejewska-Kielak, E. Kłosińska-Szmurło, F. A. Pluciński and A. P. Mazurek, Acta Pol. Pharm., 2013, 70, 877–882 CAS.
- M. L. Brusseau, N. Yan, S. Van Glubt, Y. Wang, W. Chen, Y. Lyu, B. Dungan, K. C. Carroll and F. O. Holguin, Water Res., 2019, 148, 41–50 CrossRef CAS PubMed.
- M. L. Brusseau, Sci. Total Environ., 2018, 613, 176–185 Search PubMed.
- B. E. Jugder, H. Ertan, M. Lee, M. Manefield and C. P. Marquis, Trends Biotechnol., 2015, 33, 595–610 Search PubMed.
- A. Neumann, H. Scholz-Muramatsu and G. Diekert, Arch. Microbiol., 1994, 162, 295–301 Search PubMed.
- A. Neumann, G. Wohlfarth and G. Diekert, J. Biol. Chem., 1996, 271, 16515–16519 CrossRef CAS PubMed.
- K. I. Albanese and M. L. Waters, Chem. Sci., 2021, 12, 8900–8908 Search PubMed.
- H. Zhang, S. Tian, Y. Yue, M. Li, W. Tong, G. Xu, B. Chen, M. Ma, Y. Li and J.-B. Wang, ACS Catal., 2020, 10, 3143–3151 CrossRef CAS.
- H. Gomez-Velasco, A. Rojo-Dominguez and E. Garcia-Hernandez, Biophys. Chem., 2020, 257, 106315 Search PubMed.
- P. Setny, R. Baron and J. A. McCammon, J. Chem. Theory Comput., 2010, 6, 2866–2871 CrossRef CAS PubMed.
- J. Klimeš and A. Michaelides, J. Chem. Phys., 2012, 137, 120901 CrossRef PubMed.
- J. M. Myslinski, J. E. DeLorbe, J. H. Clements and S. F. Martin, J. Am. Chem. Soc., 2011, 133, 18518–18521 CrossRef CAS PubMed.
- V. Ochoa-Herrera, Q. Banihani, G. León, C. Khatri, J. A. Field and R. Sierra-Alvarez, Water Res., 2009, 43, 3177–3186 CrossRef CAS PubMed.
- Y. Yu, F. Xu, W. Zhao, C. Thoma, S. Che, J. E. Richman, B. Jin, Y. Zhu, Y. Xing, L. Wackett and Y. Men, Sci. Adv., 2024, 10, eado2957 Search PubMed.
- Y. Wang and A. Liu, Chem. Soc. Rev., 2020, 49, 4906–4925 Search PubMed.
- Y. Li, Y. Yue, H. Zhang, Z. Yang, H. Wang, S. Tian, J. B. Wang, Q. Zhang and W. Wang, Environ. Int., 2019, 131, 104999 Search PubMed.
- R.-Z. Liao, S.-L. Chen and P. E. M. Siegbahn, ACS Catal., 2015, 5, 7350–7358 CrossRef CAS.
- I. H. Williams, Chem. Soc. Rev., 1993, 22, 277–283 Search PubMed.
- T. Nakayama, T. Kamachi, K. Jitsumori, R. Omi, K. Hirotsu, N. Esaki, T. Kurihara and K. Yoshizawa, Chemistry, 2012, 18, 8392–8402 Search PubMed.
- T. Kamachi, T. Nakayama, O. Shitamichi, K. Jitsumori, T. Kurihara, N. Esaki and K. Yoshizawa, Chemistry, 2009, 15, 7394–7403 Search PubMed.
- M. L. Baron, C. M. Bothroyd, G. I. Rogers, A. Staffa and I. D. Rae, Phytochemistry, 1987, 26, 2293–2295 Search PubMed.
- B. J. Leong, J. S. Folz, U. Bathe, D. G. Clark, O. Fiehn and A. D. Hanson, Phytochemistry, 2022, 202, 113356 Search PubMed.
- I. S. Ridder, H. J. Rozeboom, K. H. Kalk, D. B. Janssen and B. W. Dijkstra, J. Biol. Chem., 1997, 272, 33015–33022 Search PubMed.
- L. P. Wackett, Microb. Biotechnol., 2024, 17, e14463 CrossRef PubMed.
| This journal is © the Owner Societies 2026 |