Open Access Article
Open Access ArticleProline/sidechain C–H/O interactions stabilize cis-proline
Harrison C.
Oven
,
Himal K.
Ganguly
 and
Neal J.
Zondlo
and
Neal J.
Zondlo
 *
*
Department of Chemistry and Biochemistry, University of Delaware, Newark, DE 19716, USA. E-mail: zondlo@udel.edu
First published on 27th January 2026
Abstract
Bioinformatics analysis was conducted on proteins in the PDB to identify local structures that can stabilize the cis-proline conformation. C–H/O interactions were observed between a sidechain oxygen and Pro C–Hα in the cis-proline conformation at Glu–Pro, Asp–Pro, Gln–Pro, Asn–Pro, Ser–Pro, and Thr–Pro sequences. These C–H/O interactions are apparently most stabilizing at Glu–Pro sequences, which have a substantially higher than average frequency of cis-proline (7.1% of all Glu–Pro amide bonds in the PDB). DFT calculations were conducted to understand the bases and geometries of C–H/O interactions in these sequences. Computationally, these residues all exhibit close C–H/O interactions (substantially below the 2.72 Å sum of the van der Waals radii of H and O), with the closest C–H/O interactions observed with the anionic oxygens of Glu and Asp, and with closer interactions for the anionic residues than the neutral carboxamides Gln and Asn. DFT calculations revealed that C–H/O interactions also stabilize cis-proline at phosphoserine–proline and phosphothreonine–proline sequences, with closer C–H/O interactions in the dianionic forms of phosphorylated residues that predominate at physiological pH. These results also provide an explanation for the observed higher activation barrier for amide bond isomerism at phosphoserine–proline and phosphothreonine–proline sequences. Calculations suggested that C–H/O interactions mediated by these residues could also stabilize non-proline cis amide bonds, which are often functionally important when observed.
Introduction
cis amide bonds exhibit substantial changes in structure compared to trans amide bonds (Fig. 1a), both in peptides and in globular proteins.1,2 In addition, cis–trans isomerism at proline amide bonds is important in the dynamics of proteins.3–6 In isolated peptide sequences, cis amide bonds are higher energy than trans amide bonds, due to both (1) the steric clash between the Cα of adjacent residues that is present in cis amides and (2) n → π* interactions that stabilize the trans amide conformation (Fig. 1).3,7 For proline residues, the Cα/Cα steric clash in cis-proline is partially counteracted by a Cα/Cδ steric clash in trans-proline. Due to these counterbalancing interactions, 5% of all proline residues in the PDB exhibit the cis-proline conformation, with higher frequencies of cis-proline often observed in simple peptides.8–10 In contrast, for non-proline residues, the absence of a similar steric clash in the cis-amide conformation leads to a much lower frequency of cis amide (∼0.03% of all non-proline amide bonds).11Due to the geometric constraints imposed by the cis amide conformation, the vast majority of structures with cis amide bonds are β-turns (type VI), with Cα/Cα distances between the i and i + 3 residues less than 7 Å.12–14 In addition, amide cis–trans isomerism is often the slow step in protein folding, with a timescale (seconds to minutes) that is much longer than that of other protein folding transitions (microseconds to milliseconds).4,15,16 At proline residues, the cis–trans amide interconversion is catalyzed by prolyl isomerases, including cyclophilins, FKBPs, and parvulins.17,18 At non-proline residues, where the energy difference between trans-amide and cis-amide is higher in the unfolded state, and thus where a larger native barrier exists for the trans-to-cis conversion (leading to t1/2 ∼ 1000 s), amide isomerism is mediated by molecular chaperones such as DnaK.19–22
cis amide bonds are highly evolutionarily conserved, likely due to the substantial differences in structure and relative orientations of the protein chains on each side of the amide bond when comparing trans versus cis amides.23 Proteins can exhibit switches in function as a result of proline cis–trans isomerism, with different activities or interaction partners in each amide conformation about a specific proline residue.5,6,24–27
Despite the functional importance of proline cis–trans isomerism, there is still an incomplete understanding of the factors stabilizing or promoting the cis-proline conformation.9,28,29 Within globular proteins, the three-dimensional folded structure certainly plays a significant role in stabilizing the cis-amide conformation. Folding, however, plays a more minor role in intrinsically disordered regions of proteins (IDPs), where the dynamics of proline cis–trans isomerism are often critical.2,30 Moreover, even within globular proteins, there are substantial differences in the frequency of cis-proline depending on the adjacent residues. Proline–proline, aromatic–proline, and proline–aromatic sequences are substantially more likely to adopt the cis-proline conformation, both in globular proteins and within disordered peptides.31–37 The higher frequency of cis-proline in aromatic–proline and proline–aromatic sequences is primarily due to C–H/π interactions, between the aromatic (π) ring and the proline ring and/or C–Hα of the pre-proline residue, which stabilize the cis-proline conformation.38
However, other bases for the stabilization of cis-proline are less well understood. For example, prior bioinformatics analyses have indicated that Glu–Pro sequences in the PDB also exhibit a significantly higher than typical frequency of cis-proline.9,28 Moreover, local sequences can dramatically impact the dynamics of proline cis–trans isomerism. Protein serine/threonine phosphorylation results in a ∼1 kcal mol−1 higher barrier for proline cis–trans isomerism. The substantially slower dynamics (∼5–10-fold lower rates of interconversion) are resolved in vivo by the phosphorylation-dependent prolyl isomerase Pin1.39–44 Pin1 overexpression is associated with cancers due to Pin1 promoting cell cycle progression.45 In contrast, Pin1 is depleted in Alzheimer's disease, and suppression of Pin1 levels in disease models leads to more rapid tau aggregation and neurodegeneration.46 Phosphorylation-dependent proline cis–trans isomerization in the RNA polymerase II C-terminal domain is also central to transcription.47–50 However, the bases for these observations of effects of phosphorylation on proline cis–trans isomerization rates have remained unexplained.
Recently, we demonstrated that cis-proline is stabilized in Ser–Pro sequences via a C–H/O interaction51–58 between the sidechain Ser oxygen and the Pro C–Hα (Fig. 1b).59,60 This work was supported by data from small-molecule X-ray crystallography, solution-state NMR spectroscopy, bioinformatics analysis of the PDB, and DFT calculations on model peptides. In Ser–Pro, the cis-stabilizing effect of C–H/O interactions is counterbalanced by hydrogen-bonding interactions between the Ser hydroxyl and backbone amides that are present in trans-proline. Similarly, based on NMR data and molecular modeling, we recently proposed that Ser/Thr phosphorylation increases the activation barrier at pSer–Pro and pThr–Pro sequences in part via stabilization of cis-proline via C–H/O interactions between a phosphate oxygen and Pro C–Hα.61
C–H/O interactions can be described electrostatically, as the interaction between the partial negative charge (δ−) on an oxygen with the partial positive charge (δ+) on the hydrogen of a polarized C–H bond. The Hα of amino acids exhibit significant δ+ due to bond polarization via the amide nitrogen and carbonyl carbon that are attached to Cα.55,56 This reduced electron density on Hα is reflected in the more downfield chemical shift of protein Hα (∼4–5 ppm) compared to other aliphatic hydrogens in proteins.62
Importantly, however, C–H/O interactions have only a modest dependence on electrostatics,63–65 and are primarily stabilizing via stereoelectronic (molecular orbital-based) effects. Stabilization occurs due to electron delocalization between oxygen lone pair orbitals and the antibonding (σ*) orbital of the C–H bond. Evidence for the predominantly stereoelectronic nature of C–H/O interactions comes both from experiment (due to only partial charges being involved, the underlying electrostatics energies are inherently weak in water, and thus the observation of these interactions being stabilizing in water is inconsistent with a primary electrostatics basis) and from calculations. For example, only minimal differences in calculated interaction energies of the model C–H/O interaction DMF·CHCl3 are observed between ε = 9 and ε = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000.65
000.65
X-ray crystallography is also consistent with a predominantly stereoelectronic nature for C–H/O interactions. H⋯O distances significantly below the 2.72 Å sum of the van der Waals radii of H and O are inconsistent with a purely electrostatics-based interaction, but are consistent with electron delocalization (partial covalency) being important in the interaction.65 C–H/O interactions are also dependent on the electron density on the oxygen, with a more electron-rich oxygen being a better electron donor.66 Thus, calculations on model systems demonstrate a closer H⋯O distance and stronger interaction with oxygens that are part of dianions than monoanions, and with anions compared to neutral oxygens.
Collectively, these results suggested the possibility that protein sidechain oxygens might more generally interact with Pro C–Hα to stabilize the cis-proline conformation via C–H/O interactions. We examine this hypothesis herein, via a combination of bioinformatics analysis of the PDB and DFT calculations. The potential for local proline/sidechain C–H/O interactions will be examined via bioinformatics analysis of structures with cis-proline. The structures identified via bioinformatics will then be analyzed via DFT calculations to determine the potential bases for main chain/sidechain C–H/O interactions to stabilize the cis-proline conformation.
Methods
Bioinformatics
On May 9, 2024, a search of the Protein Data Bank (PDB) was conducted for structures with resolution ≤2.5 Å and with sequence similarities less than 30% using the PISCES server.67 The search yielded 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 599 total structures. Perl scripts were written to extract lists of structures containing Glu–Pro, Gln–Pro, Asp–Pro, Asn–Pro, Ser–Pro, and Thr–Pro sequences. Additional perl scripts were written to individually analyze the lists of X–Pro structures and calculate dihedral angles, specified interatomic distances, amide hydrogen positions, and Hα positions for the examined residues. The data sets were manually refined to exclude structures with nearby broken backbone bonds and structures that contained Pro with positive ϕ dihedral angles. The data sets used for analyses contained 4453 EP structures (4138 with trans-proline and 315 with cis-proline), 3519 QP structures (3354 with trans-proline and 165 with cis-proline), 5322 DP structures (5155 with trans-proline and 167 with cis-proline), 4648 NP structures (4440 with trans-proline and 208 with cis-proline), 5188 SP structures (4895 with trans-proline and 293 with cis-proline), and 5392 TP structures (5188 with trans-proline and 204 with cis-proline). Additional details are in the SI.
599 total structures. Perl scripts were written to extract lists of structures containing Glu–Pro, Gln–Pro, Asp–Pro, Asn–Pro, Ser–Pro, and Thr–Pro sequences. Additional perl scripts were written to individually analyze the lists of X–Pro structures and calculate dihedral angles, specified interatomic distances, amide hydrogen positions, and Hα positions for the examined residues. The data sets were manually refined to exclude structures with nearby broken backbone bonds and structures that contained Pro with positive ϕ dihedral angles. The data sets used for analyses contained 4453 EP structures (4138 with trans-proline and 315 with cis-proline), 3519 QP structures (3354 with trans-proline and 165 with cis-proline), 5322 DP structures (5155 with trans-proline and 167 with cis-proline), 4648 NP structures (4440 with trans-proline and 208 with cis-proline), 5188 SP structures (4895 with trans-proline and 293 with cis-proline), and 5392 TP structures (5188 with trans-proline and 204 with cis-proline). Additional details are in the SI.
Computational chemistry
Calculations were conducted with Gaussian 09.68 Initial structures derived from the PDB were subjected to iterative geometry optimization. For all molecules, final optimization was conducted using the M06-2X DFT functional with the 6-311++G(d,p) basis set (including diffuse and polarization functions).69,70 Calculations were conducted in implicit water (IEFPCM) unless stated otherwise.71Rotamer analysis to estimate interaction energies72 (Fig. S15) was conducted on molecules with Glu−, Glu0, or Gln via rotation of the χ2 torsion angle to t (180°) within GaussView, followed by full geometry optimization from this initial structure. The calculated interaction energies were determined via comparison of the calculated electronic energies of the interacting versus non-interacting rotamer. Interaction energies as a function of solvent were determined similarly, via full geometry optimization of the interacting and non-interacting rotamers in the indicated solvent (IEFPCM), vacuum (ε = 1), or condition with artificial dielectric constant (ε = 1000; 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, 100
000, 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000; or 1
000; or 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000), with the calculated relative electronic energies of the interacting versus non-interacting rotamer compared (Table S13). C–H/O interaction distances as a function of solvent and residue identity were also determined (Tables S14 and S15).
000), with the calculated relative electronic energies of the interacting versus non-interacting rotamer compared (Table S13). C–H/O interaction distances as a function of solvent and residue identity were also determined (Tables S14 and S15).
Structures with phosphoserine or phosphothreonine were conducted via bond rotations on models described previously,73 followed by bond rotations within GaussView and subsequent full geometry optimization. Structures at non-proline cis amide bonds were generated via modification of the relevant structures with Pro, followed by full geometry optimization.
Additional geometric details of all computational models, as well as their relative energies and the coordinates for all models, are in the SI.
NBO analysis was conducted using NBO6 as implemented within Gaussian09.74,75 Visualization was conducted in GaussView 5 with isovalues of 0.02. Atoms in Molecules (AIM) analysis was conducted using Multiwfn.76–78
Results
Bioinformatics analysis of X–Pro sequences in the PDB
In order to examine the possibility of C–H/O interactions stabilizing cis-proline, we examined high-resolution structures of X–Pro sequences in the PDB for all amino acids with an oxygen in an aliphatic sidechain (X = Asp, Asn, Glu, Gln, Ser, Thr).67 Post-translational modifications can also add oxygens to protein sidechains (e.g. Cys sulfenic acid (Cys–SOH or Cys–SO−), sulfinate (Cys–SO2−), and sulfonate (Cys–SO3−); phosphoserine or phosphothreonine). However, there is no example of an X-ray crystal structure of an isolated X–cis-proline in the PDB with X = one of these post-translationally modified amino acids. All structures were analyzed for the conformation at the X–Pro amide bond (trans-proline, ω ∼ 180°; or cis-proline, ω ∼ 0°) and the conformations at the main chain (ϕ, ψ) and sidechain (χ1, χ2, and/or χ3, as appropriate) for both residues. In addition, the distance between any sidechain oxygen atom and Pro Hα was measured for all structures. The position of Hα was calculated geometrically from the positions of the heavy atoms attached to Pro Cα (Fig. S1), as the position of Hα is typically not included in PDB files.Glu–Pro sequences exhibited the highest frequency of cis-Pro (7.1% of Glu–Pro sequences) (Table 1), and were observed at greater than the typical 5.3% frequency of cis-Pro at all X–Pro sequences. Ser–Pro sequences also had an elevated frequency of cis-Pro, as we had observed previously.59 In contrast, Gln–Pro and Asn–Pro had modestly lower than average frequencies of cis-Pro, and Thr–Pro and Asp–Pro had substantially lower than average frequencies of cis-Pro (Table 1). These frequencies of cis-proline for each amino acid are consistent with prior analyses that were conducted on fewer structures and with less stringent resolution limits than were used in the current analysis.8,9
| X | Number of structures | % of structures | |||
|---|---|---|---|---|---|
| Total | X–trans-Pro | X–cis-Pro | X–trans-Pro | X–cis-Pro | |
| Glu | 4453 | 4138 | 315 | 92.9 | 7.1 |
| Ser | 5188 | 4893 | 295 | 94.3 | 5.7 |
| Gln | 3519 | 3354 | 165 | 95.3 | 4.7 |
| Asn | 4648 | 4440 | 208 | 95.5 | 4.5 |
| Thr | 5392 | 5188 | 204 | 96.2 | 3.8 |
| Asp | 5322 | 5155 | 167 | 96.9 | 3.1 |
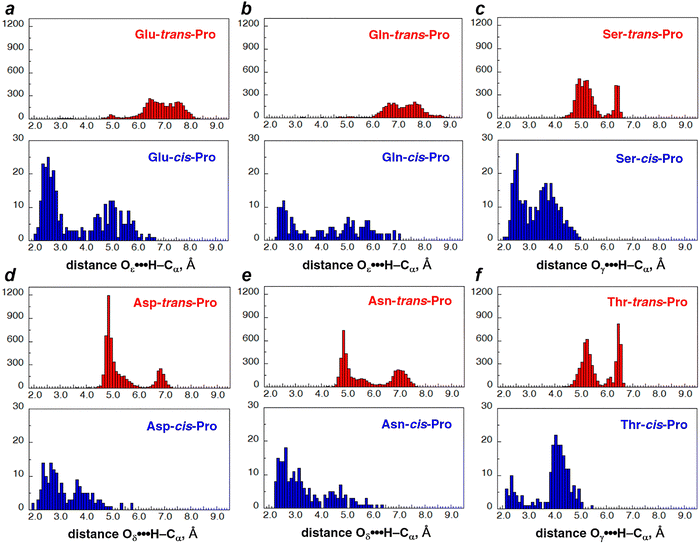
Distances for potential C–H/O interactions were examined for all amino acids in both trans-Pro and cis-Pro (Fig. 2, Table 2, Tables S2 and S3). These data indicated that C–H/O interactions were present between a sidechain oxygen and Pro C–Hα for a significant number of structures with cis-Pro, with potential C–H/O interactions defined initially as O⋯Hα distances < 3.00 Å. Notably, in cis-Pro for all amino acids, a bimodal or trimodal distribution of O⋯Hα distances was observed (Fig. 2), with one distribution of distances distinguished by O⋯Hα distances < 3.00 Å. In contrast, essentially no structures with trans-Pro had O⋯Hα distances < 4 Å (Table S3), indicating that Pro C–Hα/O interactions are not significant for trans-Pro.
| X–cis-Pro | % of X–cis-Pro with Cα–H⋯O distances ≤ | |||
|---|---|---|---|---|
| 2.30 Å | 2.50 Å | 2.75 Å | 3.00 Å | |
| Glu | 5.7 | 20.3 | 38.1 | 47.3 |
| Ser | 3.1 | 16.0 | 31.4 | 38.6 |
| Gln | 0.6 | 12.7 | 26.1 | 34.5 |
| Asn | 3.8 | 17.8 | 36.1 | 46.6 |
| Thr | 4.9 | 12.7 | 16.7 | 18.1 |
| Asp | 7.2 | 21.0 | 39.5 | 52.1 |
Analysis as a function of distance and amino acid type indicated substantial differences in frequencies of potential C–H/O interactions. The highest frequency of close O⋯Hα distances was observed for Glu–cis-Pro and Asp–cis-Pro sequences: ∼50% of these structures had an O⋯Hα distance ≤ 3.0 Å, and 37% of these structures had an O⋯Hα distance ≤ 2.72 Å. Notably, 20% of these structures had an O⋯Hα distance ≤ 2.50 Å, which is well below the sum of the van der Waals radii of H and O, and a distance that would be associated with a particularly favorable C–H/O interaction despite the superficial steric clash at these distances. In contrast, Gln–cis-Pro and Thr–cis-Pro sequences had the lowest frequencies of O⋯Hα distances that would be consistent with a C–H/O interaction. Notably, however, Thr–Pro sequences had the third highest frequency of very close (≤2.30 Å) O⋯Hα distances, suggesting that, while Thr–cis-Pro are less likely to exhibit a C–H/O interaction, they were still capable of exhibiting close C–H/O interactions.
Protein sidechain conformations exhibiting C–H/O interactions in cis-proline
The distributions of sidechain rotamers were examined for each amino acid.79 Comparison of the rotamer distributions in trans-Pro versus in cis-Pro (Table 3) indicated several trends. Glu and Gln residues exhibited a significantly higher frequency of the g−χ1 rotamer in Glx–cis-Pro, while Gln (but not Glu) exhibited a significantly higher frequency of the g+χ2 rotamer. For Asp and Asn, the major χ1 rotamer was t, for both trans-Pro and cis-Pro. Most notably, Ser and Thr exhibited a substantially higher fraction of the t χ1 rotamer for cis-Pro than trans-Pro; the t rotamer was previously59 identified to be associated with C–H/O interactions in Ser–cis-Pro.| Rotamer | Glu | Gln | Asp | Asn | Ser | Thr | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| χ 1 | χ 2 | χ 1 | χ 2 | χ 1 | χ 1 | χ 1 | χ 1 | |||
| trans | g − | −60 | 62.4 | 53.6 | 65.9 | 35.7 | 19.1 | 33.0 | 23.6 | 40.5 |
| g + | +60 | 9.2 | 31.5 | 9.6 | 13.4 | 6.8 | 7.6 | 39.3 | 52.4 | |
| t | ±180 | 28.4 | 14.8 | 24.5 | 50.8 | 74.0 | 59.4 | 37.1 | 7.1 | |
| cis | g − | −60 | 71.7 | 41.9 | 75.2 | 10.9 | 29.3 | 32.7 | 28.7 | 33.8 |
| g + | +60 | 5.7 | 18.1 | 5.5 | 45.5 | 7.8 | 3.4 | 15.7 | 45.1 | |
| t | ±180 | 22.5 | 40.0 | 19.4 | 43.6 | 62.9 | 63.9 | 55.6 | 21.1 | |
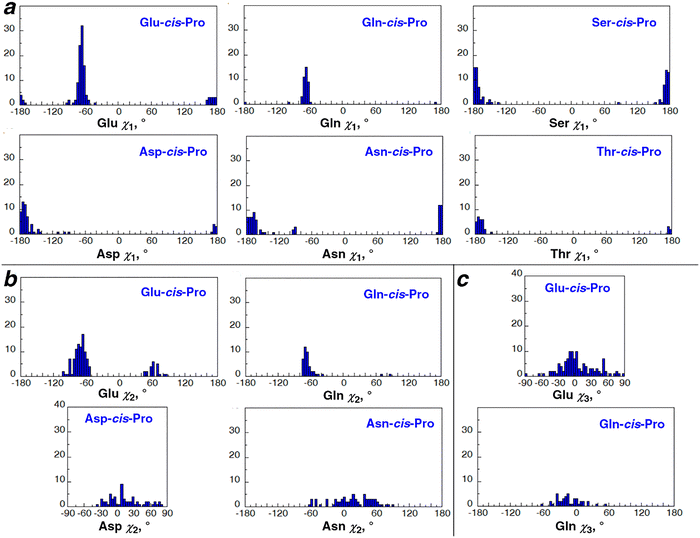
In order to identify the sidechain conformations associated with potential C–H/O interactions, all structures with O⋯Hα distances ≤ 2.72 Å were analyzed separately (Fig. 3, Fig. S3–S6). These results indicated strong conformational preferences when a C–H/O interaction was present. Glu and Gln exhibited C–H/O interactions predominantly with the g−χ1 rotamer, with the t χ1 rotamer a minor conformation for C–H/O interactions with Glu (Fig. 3a). In contrast, for Asp and Asn, C–H/O interactions were associated predominantly with the t χ1 rotamer. Similarly, for Ser and Thr, C–H/O interactions were observed almost exclusively with the t χ1 rotamer.
Glu and Gln also showed a strong preference for the g−χ2 rotamer for C–H/O interactions, with Glu also exhibiting a small population with the g+χ2 rotamer (Fig. 3b, Fig. S4). The Asp and Asn χ2 rotamers (Fig. 3b, Fig. S5), and the equivalent Glu and Gln χ3 rotamers (Fig. 3c, Fig. S4), showed a broader distribution between −60° and +60°.
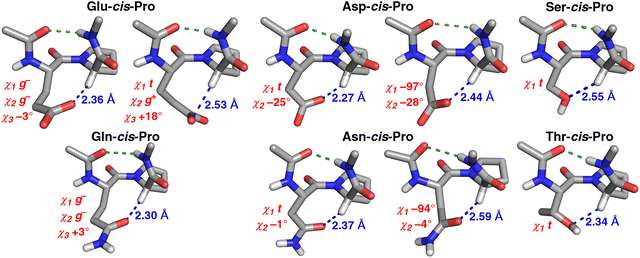
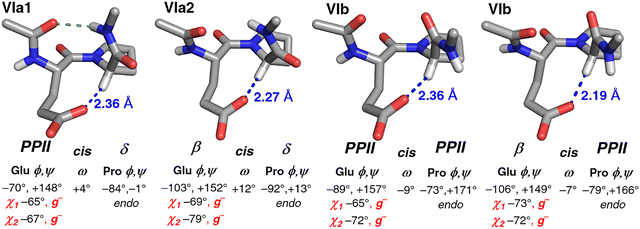
Individual structures from the PDB were analyzed for all of these identified combinations of conformations with potential C–H/O interactions (Fig. 4). All structures exhibited clear evidence of C–H/O interactions, including good interaction geometries. Analysis of these structures indicated two different interaction modes for Glu and Gln: a more common one where the Glx χ1 and χ2 rotamers were both g−, and a less common one where the χ1 rotamer was t and the χ2 rotamer was g+. These results are consistent with analysis of sidechain torsion angle distributions in C–H/O interactions (Fig. 3). Similarly, Asp and Asn each had two different combinations of χ1 and χ2 rotamers, (t, 0 ± 90°) and (∼−100°, ∼−30°). Ser and Thr exhibited C–H/O interactions through the t χ1 rotamer. Close C–H/O interactions were observed in all type VI β-turn subtypes12–14 (VIa1 [PcisD], VIa2 [BcisD], and VIb [PcisP and BcisP]) (Fig. 5, Fig. S8, Tables S10 and S11). For Glu, Gln, and Ser, C–H/O interactions were relatively overrepresented in type VIa β-turns (Pro in the δ conformation), while for Asp, Asn, and Thr, C–H/O interactions were substantially more likely in BcisP type VIb β-turns (type VIb: Pro in the PPII/β conformations) (seen by comparing conformation frequencies in Tables S9 versus S11).
All residues examined exhibited evidence of frequent and close C–H/O interactions with cis-Pro (Table 2 and Fig. 4), but substantially different frequencies of cis-Pro (Table 1). For example, Glu and Asp have essentially identical likelihoods of C–H/O interactions in cis-Pro (found in ∼40%–50% of all cis-Pro structures), but very different frequencies of cis-Pro. Asp has been previously shown to frequently interact via hydrogen bonds with the local protein backbone.80 Indeed, numerous examples were found of Asp–trans-Pro structures with hydrogen bonds to the pre-Asp (i − 1), Asp (i), and post-proline (i + 2 and i + 3) amide hydrogens (Fig. 6). These sidechain–main chain hydrogen bonds inherently compete with other conformations, and could relatively promote the trans-Pro conformation in Asp–Pro due to the greater strength of hydrogen bonds, despite favorable C–H/O interactions that are observed with Asp–cis-Pro.
 | ||
| Fig. 6 Asp–trans-Pro structures in the PDB stabilized via local hydrogen bonds between main-chain amide hydrogens and the Asp carboxylate. (left to right) pdb 5ik4, pdb 5w2f, pdb 6a9w, and pdb 3cov. | ||
DFT calculations on C–H/O interactions in Ac–X–cis-Pro–NHMe peptides
In order to better understand the nature of C–H/O interactions in stabilizing specific conformations in cis-Pro, DFT calculations were conducted on model Ac–X–cis-Pro–NHMe peptides, X = Glu, Gln, Asp, Asn, Ser, Thr. Quantum mechanics-based calculations were used instead of force field-based calculations due to the inherently quantum mechanical nature of C–H/O interactions, and the absence of full parametrization of these interactions in common force fields. Indeed, by typical molecular mechanics-based approaches, the close approach of the Pro Hα to the sidechain oxygen would be treated as a repulsive interaction, due to H⋯O distances that are below the sum of the van der Waals radii of these atoms. These systematic biases against close approaches of atoms in protein structure determination software could also impact the refinement of protein crystal structures, and thus the apparent distances observed in the PDB.53While C–H/O interactions were observed in all type VI β-turn subtypes (Fig. 5, Table S11), calculations were conducted on those structures in type VIa1 (PcisD) β-turns (Tables S5 and S6). Unlike other type VI β-turn subtypes, type VIa1 β-turns exhibit an i/i + 3 C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Oi⋯Hi+3–N main-chain/main-chain hydrogen bond, which restrains the geometry and provides interaction partners for these main-chain hydrogen-bonding groups, simplifying the calculations and analysis.12–14 Geometry optimization calculations were conducted using structures from the PDB (Fig. 4) as initial models, examining all different combinations of sidechain torsion angles that exhibited C–H/O interactions.68–71,81
Oi⋯Hi+3–N main-chain/main-chain hydrogen bond, which restrains the geometry and provides interaction partners for these main-chain hydrogen-bonding groups, simplifying the calculations and analysis.12–14 Geometry optimization calculations were conducted using structures from the PDB (Fig. 4) as initial models, examining all different combinations of sidechain torsion angles that exhibited C–H/O interactions.68–71,81
As was seen in the PDB analysis, Glu and Gln exhibited two distinct combinations of conformations with C–H/O interactions, with (χ1, χ2) = (g−, g−) or (t, g+) (Fig. 7). Closer H⋯O interactions were observed in the (g−, g−) sidechain rotamer pair for both residues, and these (g−, g−) structures were also lower in energy than the (t, g+) structures by 1.3–1.5 kcal mol−1 in these calculations. The (g−, g−) structures exhibited a more classical oxygen-based type of C–H/O interaction, with hydrogen bond-like geometries.
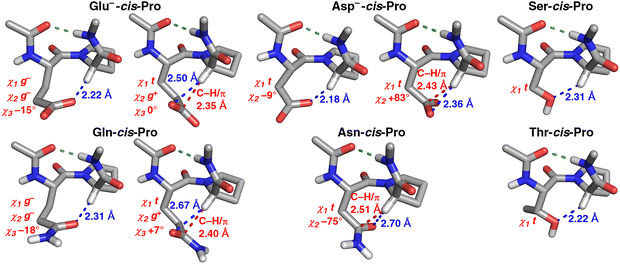 | ||
Fig. 7 C–H/O interactions with Pro C–Hα in geometry-optimized structures with X–cis-Pro amide bonds in type VIa1 β-turns. Structures from the PDB (Fig. 4) were modified to Ac–X–cis-Pro–NHMe and then subjected to geometry optimization to examine the conformations at pre-proline residues which can stabilize the cis-Pro conformation via C–H/O interactions (Hα⋯O distances, blue). Glu, Gln, Asp, and Asn all have one interaction mode where the nearest interaction is a C–H/π interaction between the amide or carboxylate π orbitals and Pro C–Hα (red). C–H/π interactions were defined as CC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O⋯H distances at or below the sum of the van der Waals radii of C and H (≤2.90 Å). Conformations at Asp–Pro with Asp χ1 = −100°, χ2 = −30° and Asn–Pro with Asn χ1 = −90°, χ2 = −30° were observed with C–H/O interactions with Pro C–Hα in the PDB but optimized to alternative structures with intraresidue C–H/O interactions, and thus were not included. Additional geometric information is in Fig. S10. O⋯H distances at or below the sum of the van der Waals radii of C and H (≤2.90 Å). Conformations at Asp–Pro with Asp χ1 = −100°, χ2 = −30° and Asn–Pro with Asn χ1 = −90°, χ2 = −30° were observed with C–H/O interactions with Pro C–Hα in the PDB but optimized to alternative structures with intraresidue C–H/O interactions, and thus were not included. Additional geometric information is in Fig. S10. | ||
In contrast, the (t, g+) structures exhibited longer O⋯H distances, but had a geometry that is more typical of cation/π or C–H/π interactions,82–84 with the Pro Hα interacting closely with the π face of the carboxylate or carboxamide (a geometry also observed in the PDB, Table S7). Indeed, these structures had very close CC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O⋯Hα distances (2.35–2.40 Å), well below the 2.90 Å sum of the van der Waals radii of C and H. C–H/O interactions with carbonyls may be mediated via either of the oxygen lone pairs (Os or Op) or via the π molecular orbital serving as the electron donor that interacts with the C–H σ* orbital.65 The calculations indicate that these distinct interaction modes may be alternatively observed in different conformations of the Glu or Gln sidechains.
O⋯Hα distances (2.35–2.40 Å), well below the 2.90 Å sum of the van der Waals radii of C and H. C–H/O interactions with carbonyls may be mediated via either of the oxygen lone pairs (Os or Op) or via the π molecular orbital serving as the electron donor that interacts with the C–H σ* orbital.65 The calculations indicate that these distinct interaction modes may be alternatively observed in different conformations of the Glu or Gln sidechains.
Notably, closer C–H/O interactions were observed with Glu than with Gln. These results are consistent with bioinformatics data (Table 2), which indicated a substantially greater likelihood of close C–H/O interactions for Glu than Gln. The closer interactions are likely due to the greater electron density on the anionic carboxylate oxygen of Glu than on the formally neutral carboxamide oxygen of Gln, along with the higher energy of the filled orbitals of the more electron-rich carboxylate than the less electron-rich carboxamide. Consistent with this interpretation, geometry optimization calculations on the neutral (acidic) form of Glu (Glu0) demonstrated a significantly longer O⋯Hα distance for the neutral acid compared to the anionic carboxylate (Fig. 8). The neutral Glu0 also exhibited longer C–H/O interaction distances than the Gln carboxamide (2.52 Å versus 2.31 Å), consistent with the more electron-rich nature (and higher-energy π molecular orbitals) of the amide compared to the acid.
Two interaction geometries were identified as stable energy minima for Asp, (χ1, χ2) = (t, −10°) or (t, +80°), while one was identified for Asn (χ1, χ2) = (t, −75°) (Fig. 7). The (t, −10°) geometry for Asp exhibited the closest O⋯Hα distance of any structure in these calculations, as well as a classical hydrogen bond-like geometry, consistent with a very favorable C–H/O interaction. However, notably, this structure was also quite strained in the ω torsion angle (ω = +20°), which imposes an inherent torsional energy cost that would result in destabilization due to reduced electron delocalization in that amide bond. The Asp (χ1, χ2) = (t, +80°) structure and the Asn (χ1, χ2) = (t, −75°) structure had significantly longer O⋯Hα distances, but both exhibited a favorable C–H/π interaction geometry, with close C⋯Hα distances, similar to that observed in the (t, g+) rotamer of Glu and Gln. As was the case with Glu versus Gln, in structures with comparable geometries, closer interaction distances were observed for Asp than Asn, consistent with closer C–H/O interactions with more electron-rich anionic carboxylate electron donors than with neutral amide carbonyls.
C–H/O interactions in the Ser and Thr structures were mediated via the t rotamer, with O⋯Hα distances that were substantially below 2.72 Å (Fig. 7). Interestingly, a closer interaction was observed with Thr than Ser. These results are consistent with bioinformatics data indicating that, while Thr had the lowest overall frequency of C–H/O interactions in cis-Pro (<20% of Thr–cis-Pro had a C–H/O interaction, Table 2), Thr–cis-Pro C–H/O interactions that were present were particularly likely to be close (≤2.30 Å).
Calculations were conducted on Ser–cis-Pro with two different χ2 torsion angles, which represent two different positions of the serine hydroxyl hydrogen and are not directly determinable in the PDB. Notably, the O⋯Hα distance depended on the χ2 torsion angle (Fig. 9), and also depended on the Ser ionization state (Fig. S13), with a closer distance for the atypical anionic Ser. While anionic Ser is strongly thermodynamically disfavored, these results suggest that hydrogen bonding to the sidechain hydroxyl might modulate electron density on oxygen and impact the strength and geometry of C–H/O interactions in cis-Pro.
Calculations on the energetics and inherent bases of C–H/O interactions stabilizing cis-Pro
In order to further understand the underlying basis or bases for C–H/O interactions that stabilize cis-Pro, natural bond orbital (NBO) calculations74,75 were conducted on models derived from all X–cis-Pro sequences. NBO calculations convert the wavefunction into localized molecular orbitals, which can be used to approximate the component molecular orbitals that contribute to stabilization via electron delocalization.In Glu–cis-Pro and Gln–cis-Pro structures with (χ1, χ2) = (g−, g−), NBO calculations indicated substantial stabilization due to interaction of the Pro C–H antibonding (σ*) orbital with two oxygen lone pairs of the carboxylate of Glu or the carbonyl of Gln (Fig. 10a). In contrast, in Glu–cis-Pro and Gln–cis-Pro structures with (χ1, χ2) = (t, g+), interactions with the oxygen lone pairs were greatly reduced (Fig. 10b). However, in this case, NBO calculations demonstrated greater interaction of the Pro C–H σ* orbital with the π molecular orbitals, consistent with the hypothesis above that these structures could be interpreted as being stabilized primarily via a C–H/π interaction mode. Full molecular orbital calculations, which properly address the global (rather than localized) nature of molecular orbitals, clearly demonstrate electron delocalization between the Glx carboxylate or carboxamide and the Pro C–Hα, with through-space electron delocalization stabilizing these conformations (Fig. S14).
The wavefunctions from these calculated structures were also examined via Atoms in Molecules (AIM) analysis.76–78 All structures exhibited bond critical points (BCPs) between sidechain atoms and Pro Hα (Fig. S16). These results are consistent with NBO calculations indicating through-space electron delocalization and partial covalency in these C–H/O interactions. The highest electron densities at the BCPs were for Glu− in the (g−, g−) conformation, for Thr, and for Asp in the (t, −10°) conformation. These results are consistent with these conformations exhibiting the closest C–H/O interactions (Fig. 7). Notably, these electron densities were also similar to those observed in the model acetylene–water C–H/O interaction (Fig. S16).85
In most structures, the BCP was located on an approximately linear path between the sidechain oxygen and Pro Hα. However, for either Glu− or Gln in the (t, g+) conformation, and for Asn in the (t, −75°) conformation, the BCP was on paths between the Pro Hα and the carbonyl carbon (Fig. S16). For Asp in the (t, +80°), a curved path was observed. The results for these conformations are consistent with our analysis above, that these C–H/O interaction modes are more properly described as C–H/π interactions, with electron delocalization between the conjugated π systems and σ* of Pro C–Hα.
In order to understand the energetics and the underlying primary bases for C–H/O interactions that stabilize cis-Pro, calculations were conducted comparing the structures identified above with structures in which one sidechain torsion angle was rotated away from the C–H/O interaction, with subsequent full geometry optimization (Fig. S15).72 For structures with Glu and Gln, the structures resulting from rotating the χ2 torsion angle to t (180°) and subsequent geometry optimization placed the sidechain away from the backbone, allowing an estimation of the C–H/O interaction energy via the energy differences between the interacting and non-interacting rotamers. In contrast, in structures with Asp, Asn, Ser, and Thr, the resultant rotated structures exhibited a new hydrogen bond with the backbone, consistent with the high frequency of sidechain–backbone interactions observed for these amino acids, and precluding application of this approach.
For Glu, the (χ1, χ2) = (g−, g−) rotamer exhibited an interaction energy of −3.0 kcal mol−1 in implicit water using this approach of comparing energies of different rotamers (Table S12). In contrast, the Glu (χ1, χ2) = (t, g+) rotamer had an interaction energy of −1.8 kcal mol−1, with this less favorable interaction energy consistent with its less frequent observation in the PDB. For Gln, the comparable energies were less favorable than those with Glu ([g−, g−] = −2.4 kcal mol−1, [t, g+] = −1.4 kcal mol−1), consistent with stronger C–H/O interactions with Glu than with Gln. Finally, the (g−, g−) rotamer exhibited the weakest interaction energy (−2.0 kcal mol−1) for neutral Glu0, consistent with the less electron-rich nature of a carboxylic acid (Glu0) compared to a carboxamide (Gln) and the longer O⋯Hα distances observed in geometry optimization calculations (Fig. 8). Two important caveats to these rotamer-based interaction energies are (1) that they do not address inherent differences in the energies of the conformations, though these are expected to be relatively small at χ2; and (2) more importantly, that they do not fully address differences in solvation energies (e.g., differences in competitive hydrogen bonds at the oxygen or between water molecules) and other solvation (e.g., organized solvation at Hα by water).
The interacting and non-interacting conformer pairs of Glu, Gln, and Glu0 were used to explore the roles of electrostatics versus stereoelectronic effects in the energetics of C–H/O interactions. CM5 calculations86 of charge on the interacting oxygens and Pro Hα atoms in these structures indicate relatively modest partial charges (Glu −0.49, Gln −0.39, Glu0 carbonyl O −0.34; Pro Hα +0.10 to +0.12). These partial charges would be expected to be associated with minimal favorable energetics via electrostatics in water for these structures with C–H/O interactions. Moreover, considering the very close interactions of the sidechain carbonyl carbon with Pro Hα in the (t, g+) rotamers (C⋯Hα 2.35–2.40 Å, well below the 2.90 Å sum of van der Waals radii of these atoms), the CM5 charges on the carbonyl carbon (+0.16 in Glu and +0.26 in Gln) and Pro Hα (+0.10 to +0.12) suggest that the very close approach of these atoms in a C–H/π interaction mode should be unfavorable due to electrostatic repulsion between positively charged C and H atoms.
Geometry optimization calculations were conducted on both the interacting and non-interacting structures as a function of solvent dielectric constant (ε), examining the interaction distance and the energy difference between interacting and non-interacting rotamers in vacuum (gas phase) (ε = 1), hexane (ε = 1.9), CHCl3 (ε = 4.7), CH2Cl2 (ε = 8.9), acetone (ε = 20.5), acetonitrile (ε = 36), DMSO (ε = 47), H2O (ε = 78), and artificial solvent conditions with ε = 1000, ε = 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, ε = 100
000, ε = 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, and ε = 1
000, and ε = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000. These latter high-dielectric-constant conditions will effectively fully screen out purely electrostatics-based interactions, since electrostatics interaction energy (Eelectrostatics) scales as Eelectrostatics ∼ 1/ε.
000. These latter high-dielectric-constant conditions will effectively fully screen out purely electrostatics-based interactions, since electrostatics interaction energy (Eelectrostatics) scales as Eelectrostatics ∼ 1/ε.
For Glu–cis-Pro structures with (χ1, χ2) = (g−, g−), the closest O⋯Hα distance (2.05 Å) and most favorable interaction energy (−5.7 kcal mol−1) were observed in vacuum, consistent with electrostatics being an important component of C–H/O interactions in vacuum, as well as the inherent destabilization of charge without counterbalancing opposite charges. In contrast, in all solvent conditions with ε ≥ 4.7 (CHCl3 and all more polar conditions), the interaction distances (2.22 Å ± 0.02 Å) and interaction energies (−2.7 to −3.1 kcal mol−1) were similar (Tables S13 and S14). Notably, among these conditions, the most favorable interaction energies were actually with ε = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, conditions that functionally screen out all electrostatics interactions. These results indicate that C–H/O interactions are highly favorable even under conditions with large dielectric constants, suggesting that outside of the most non-polar conditions (vacuum or hexanes), electrostatics play a minimal role in the strength of these C–H/O interactions. Moreover, calculations with neutral oxygen sources (Gln or Glu0), in the (χ1, χ2) = (g−, g−) rotamer, indicate that the O⋯Hα interaction distances (2.305 Å ± 0.01 Å for Gln, 2.38 Å ± 0.01 Å for Glu0) and interaction energies (−2.3 ± 0.2 kcal mol−1 for Gln, −1.96 ± 0.13 kcal mol−1 for Glu0) have minimal solvent dependence. For example, the O⋯Hα distance for Gln was nearly identical in vacuum (ε = 1) (2.313 Å) and in ε = 1
000, conditions that functionally screen out all electrostatics interactions. These results indicate that C–H/O interactions are highly favorable even under conditions with large dielectric constants, suggesting that outside of the most non-polar conditions (vacuum or hexanes), electrostatics play a minimal role in the strength of these C–H/O interactions. Moreover, calculations with neutral oxygen sources (Gln or Glu0), in the (χ1, χ2) = (g−, g−) rotamer, indicate that the O⋯Hα interaction distances (2.305 Å ± 0.01 Å for Gln, 2.38 Å ± 0.01 Å for Glu0) and interaction energies (−2.3 ± 0.2 kcal mol−1 for Gln, −1.96 ± 0.13 kcal mol−1 for Glu0) have minimal solvent dependence. For example, the O⋯Hα distance for Gln was nearly identical in vacuum (ε = 1) (2.313 Å) and in ε = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 (2.315 Å), as was also the case for Glu0 (2.382 Å versus 2.387 Å).
000 (2.315 Å), as was also the case for Glu0 (2.382 Å versus 2.387 Å).
In structures of Glu− or Gln with (χ1, χ2) = (t, g+), the interaction energies and geometries similarly showed minimal dependence on solvent dielectric constant in all conditions other than calculations in vacuum or hexane for Glu−. The close Cπ⋯Hα interaction distances (2.36 Å ± 0.03 Å for Glu−, 2.40 Å ± 0.01 Å for Gln; both of these are ∼0.5 Å below the 2.90 Å sum of the van der Waals radii of C and H) also exhibit minimal dependence on the solvent dielectric constant. These results provide further evidence that C–H/O interactions and C–H/π interactions are fundamentally stereoelectronic in nature, and that the molecular orbital basis of these interactions renders them functionally solvent-independent for their inherent interaction energies.
C–H/O interactions in pSer–cis-Pro and pThr–cis-Pro
Ser/Thr phosphorylation leads to an increased barrier for proline cis–trans isomerization at pSer–Pro and pThr–Pro sequences.39–41 Recently, we proposed the possibility that this increased barrier to isomerization was due to the combination of (a) stabilization of the ground state of trans-Pro via a phosphate–amide hydrogen bond73 and (b) stabilization of the ground state of cis-Pro via a phosphate–Pro C–Hα C–H/O interaction,60,61 with neither of these interactions present and/or impacting the energy of the transition state of isomerization. In order explore this hypothesis, we conducted calculations on pSer–cis-Pro and pThr–cis-Pro structures, in order to examine potential C–H/O interactions with the phosphate that could stabilize cis-Pro (Fig. 11). Calculations were conducted in the physiologically most important dianionic ionization state, in the monoanionic state that can be a significant population in solution, and in the non-physiologically relevant neutral form (Fig. 11).We identified two different pairs of sidechain conformations that exhibit phosphate–proline C–H/O interactions, (χ1, χ2) = (g−, −90°) or (t, +95°). For both pSer and pThr, closer Hα⋯O distances were observed in the (g−, −90°) rotamer pair. Closer Hα⋯O distances were also observed in the dianionic form than in the monoanionic form, with both closer than with the neutral phosphate. Due to multiple Lewis basic oxygens in the phosphate, C–H/O interactions were observed to both the Pro C–Hα and to the pSer/pThr C–Hα. Notably, even in structures with the neutral phosphate, close C–H/O interactions were observed, at Pro C–Hα and/or at the pSer/pThr Hα. In addition, AIM analysis demonstrated BCPs between a phosphate oxygen and Pro Hα in all of these structures (Fig. S18). These results are consistent with the hypothesis that C–H/O interactions can stabilize cis-proline when preceded by pSer or pThr, and that cis-proline-stabilizing C–H/O interactions contribute to the higher activation barrier for proline cis–trans isomerization at these sites in proteins.
C–H/O interactions in cis-non-proline amide bonds
0.03% of non-proline amide bonds are in the cis-amide conformation.11 Because cis-non-proline amide bonds are inherently ∼5 kcal mol−1 higher in energy than those of trans-non-proline,3,20 their observation in proteins requires stabilization of the cis amide by local and/or global structures. The low frequencies of cis-non-proline amide bonds preclude bioinformatics analysis as conducted above.87 However, human phospholipase D exhibits a particularly close C–H/O interaction between Gly C–Hα and the Asp carboxylate, which we propose to stabilize the cis-Gly conformation (Fig. 12).88 We had previously observed59 that C–H/O interactions stabilize a Ser–cis-Arg conformation that is widely conserved in bacterial xanthine–guanine phosphoribosyltransferases.89,90 While not supported by the power of statistics, these structures suggest that C–H/O interactions can be one component stabilizing cis-non-proline conformations.In order to further explore this possibility, we conducted DFT calculations on Ac–X–cis-Ala–NHMe structures, X = Glu, Gln, Asp, Asn, Ser, Thr, pSer, and pThr. All residues exhibited C–H/O interactions between a sidechain oxygen and Ala C–Hα, with Hα⋯O distances and interaction geometries similar to those observed with Pro (Fig. 12b and Fig. S19, S20). Calculations were also conducted on Ac–Glu–cis-Z–NHMe structures, Z = Ala, Gly, Ser, Val, tert-leucine (Tle), in order to examine the dependence of the interaction on the residue identity at the cis amide. All structures exhibited close C–H/O interactions, with the shortest Hα⋯O distances for Ser and the longest for Val (Fig. 12b, c, Fig. S21, Table S16).
In order to test the ability to electronically tune the strength of C–H/O interactions via inductive effects, Ac–Glu–cis-Z–NHMe structures with Z = fluorinated amino acids β-F-Ala, β,β-F2-Ala, β,β,β-F3-Ala, and α-F-Gly were also examined computationally. Consistent with structure being driven by favorable C–H/O interactions, introduction of fluorines led to closer Hα⋯O distances,64,91 with the structure with trifluoroalanine exhibiting a 1.99 Å Hα⋯O distance, comparable to that of typical hydrogen bonds (Fig. 12d, e, Fig. S22, Table S17).
Discussion
cis-Proline amide bonds are associated with substantial changes in protein structure, dynamics, and function.1–6 However, there is an incomplete understanding of local structural effects that stabilize cis-proline. We previously identified that cis-proline in Ser–Pro sequences is stabilized by a C–H/O interaction between the sidechain Ser oxygen and the proline C–Hα.59Herein, we have more broadly examined the possibility of C–H/O interactions stabilizing cis-proline amide bonds via sidechain oxygen lone pairs interacting with Pro C–Hα bonds. Bioinformatics analysis of cis-proline amide bonds at Glu–Pro, Gln–Pro, Asp–Pro, Asn–Pro, Ser–Pro, and Thr–Pro sequences indicate high frequencies of structures with O⋯Hα distances at or below the 2.72 Å sum of the van der Waals radii of O and H, with a significant number of structures with O⋯Hα distances < 2.30 Å.
Analysis of structures with C–H/O interactions, combined with DFT calculations on these structures, identified two interaction modes of the sidechain with cis-proline for Glu, Gln, Asp, and Asn: one with an oxygen lone pair-directed C–H/O interaction ([χ1, χ2] = [g−, g−] for Glu and Gln, [χ1, χ2] = [t, ∼0°] for Asp, Asn), and one with a C–H/π-type interaction geometry of the Pro C–Hα with the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O π molecular orbitals ([χ1, χ2] = [t, g+] for Glu and Gln, = [t, ∼−80°] for Asp, and = [t, ∼+80°] for Asn). C–H/O interactions with Ser and Thr are mediated via the t χ1 rotamer.
O π molecular orbitals ([χ1, χ2] = [t, g+] for Glu and Gln, = [t, ∼−80°] for Asp, and = [t, ∼+80°] for Asn). C–H/O interactions with Ser and Thr are mediated via the t χ1 rotamer.
These C–H/O interactions were further investigated computationally, in order to understand the bases for these interactions. In water, the primary basis of interaction strength is not an electrostatics interaction between the partial negative change on the oxygen and the partial positive charge on the polarized hydrogen. Instead, C–H/O interactions are fundamentally driven by through-space electron delocalization, a molecular orbitals-based effect that leads to orbital mixing between electron-rich oxygen lone pair orbitals (ns or np) on the sidechain oxygen or the π orbitals on the carbonyl and the proline C–H antibonding orbital (σ*), as nO → σ*C–H or πC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O → σ*C–H interactions. These quantum-mechanical effects on structure are currently not well addressed by typical molecular mechanics/force field-based approaches.
O → σ*C–H interactions. These quantum-mechanical effects on structure are currently not well addressed by typical molecular mechanics/force field-based approaches.
More broadly, because the interaction strengths of stereoelectronic effects are inherently less dependent on solvent than are electrostatics interactions, they have the potential to impact protein structures and interactions in diverse contexts and in ways that might be underappreciated.64,65 Considered only in electrostatics terms, C–H/O interactions should be insignificant in water, given the very modest partial charges present on (for example) protein Hα (∼+0.1). Moreover, at the sidechain oxygen atoms, a C–H/O interaction is weaker than a hydrogen bond of those oxygen atoms to water.92 However, the C–H/O interaction also obviates the need for water solvation at Pro Hα, allowing the released water molecules to engage in more favorable water–water hydrogen bonds. C–H/O interactions also convert hydrogen bonds of water with the sidechain oxygen to water–water hydrogen bonds that are likely similar in strength. In addition, by releasing water molecules at both sites, the C–H/O interaction also has potential advantages in translational entropy. Notably, in membrane environments, C–H/O interactions can occur without a water desolvation energy cost.93,94 Globally, these questions can be properly addressed in future QM/MM simulations that explicitly include the quantum mechanical nature both of C–H/O interactions and of hydrogen bonds to water.95,96
Glu–Pro sequences have previously been identified to have a significantly higher than average frequency of the cis-Pro conformation.8,9 However, no basis for this higher frequency of cis-Pro has been identified. The data herein suggest that C–H/O interactions between the Glu carboxylate and Pro Hα specifically stabilize the cis-Pro conformation relative to the trans-Pro conformation, leading to an increased population of cis-Pro at Glu–Pro sequences. C–H/O interactions also impact the conformational ensemble present in structures with cis-Pro. More broadly, C–H/O interactions are one of a series of competing sidechain–mainchain interactions that impact conformations in both trans-Pro and cis-Pro.
Intrinsically disordered proteins (IDPs) and unfolded states of proteins exhibit mixtures of trans-Pro and cis-Pro at proline sites.2,4,10,16 In contrast, in folded proteins, typically a single amide conformation is present at each proline, due to the constraints of tertiary structure and the differences in local structure in trans-Pro versus cis-Pro.23 The work herein provides insights into transient structures that can be present when the cis-Pro conformation is present in IDPs, as well as local sequence elements that can promote cis-Pro.
DFT calculations also demonstrated that cis-proline in phosphoserine–proline and phosphothreonine–proline sequences can be stabilized by favorable C–H/O interactions mediated by the electron-rich phosphate group. This stabilization of the cis-proline conformation by C–H/O interactions (relative to the transition state for isomerization, where these interactions are not present) provides an explanation for the higher barrier and the significantly slower rate of proline cis–trans isomerization at these sites in proteins.40
The strength of the C–H/O interactions at pSer–Pro and pThr–Pro sites, as well as at Glu–Pro and Asp-Pro sites, is impacted by ionization state (charge).66,97 Phosphorylated amino acids typically exist primarily in the dianionic forms, but significant populations of the monoanionic forms can also be present or predominant.73,98,99 Notably, local environment impacts the pKa values of ionizable groups.100,101 The results herein provide a context by which changes in local environment could impact proline cis–trans isomerism at these sites via changes in sidechain ionization state.
Slower proline cis–trans isomerization at pSer–Pro and pThr–Pro sites is a central element both in transcription (via cis–trans isomerization in the Pol II CTD) and in cell cycle progression and cell division (via numerous proteins).39,47–49 It has also been implicated in misfolding of the tau protein in Alzheimer's disease. The phosphorylation-dependent prolyl isomerase Pin1/Ess1 is critical in these processes.41,42,44,45 The identified relevant Ser/Thr phosphorylation sites in these proteins are all in IDPs and/or in intrinsically disordered regions (IDRs) of proteins. In addition, other prolyl isomerases have been identified to impact the phase separation behavior and aggregation of IDPs and IDRs.44,102–104 The results herein suggest that phosphate–Pro Hα C–H/O interactions are important to the dynamics of these proline cis–trans isomerization events. Moreover, these results suggest that transient protein structures or protein–protein interactions that engage with sidechain phosphates or with other sidechain oxygens could impact the structures and dynamics of the conformational ensemble, and thereby change protein function.
Finally, all encoded amino acids have a C–Hα equivalent to that in proline. We demonstrate computationally that sidechain–main chain C–H/O interactions are also capable of stabilizing cis amide bonds at non-proline residues. Collectively, these results suggest that C–H/O interactions between a sidechain oxygen and the main-chain C–Hα at the subsequent residue can stabilize a cis amide bond both at proline and at non-proline residues, and that the interaction strength is dependent on the identity of both residues.
Conflicts of interest
There are no conflicts to declare.Data availability
Summaries of key data are included in the main manuscript. Additional data are in the supplementary information (SI), including additional bioinformatics analysis and the coordinates for all structures determined computationally. Supplementary information is available. See DOI: https://doi.org/10.1039/d5cp03423j.Acknowledgements
We thank NSF (CHE-2004110 and BIO-1616490) and the DOD CDMRP PRARP program (AZ140115) for funding. Instrumentation support was provided by NIH (GM110758) and NSF (CHE-1229234).References
- M. W. MacArthur and J. M. Thornton, Influence of proline residues on protein conformation, J. Mol. Biol., 1991, 218, 397–412 CrossRef CAS PubMed.
- B. Mateos, C. Conrad-Billroth, M. Schiavina, A. Beier, G. Kontaxis, R. Konrat, I. C. Felli and R. Pierattelli, The Ambivalent Role of Proline Residues in an Intrinsically Disordered Protein: From Disorder Promoters to Compaction Facilitators, J. Mol. Biol., 2020, 432, 3093–3111 Search PubMed.
- G. Fischer, Chemical aspects of peptide bond isomerisation, Chem. Soc. Rev., 2000, 29, 119–127 RSC.
- J. F. Brandts, H. R. Halvorson and M. Brennan, Consideration of the possibility that the slow step in protein denaturation reactions is due to cis–trans isomerization of proline residues, Biochemistry, 1975, 14, 4953–4963 CrossRef CAS PubMed.
- A. H. Andreotti, Native State Proline Isomerization: An Intrinsic Molecular Switch, Biochemistry, 2003, 42, 9515–9524 Search PubMed.
- B. Eckert, A. Martin, J. Balbach and F. X. Schmid, Prolyl isomerization as a moleular timer in phage infection, Nat. Struct. Mol. Biol., 2005, 12, 619–623 CrossRef CAS PubMed.
- R. W. Newberry and R. T. Raines, The n → pi* interaction, Acc. Chem. Res., 2017, 50, 1838–1846 CrossRef CAS PubMed.
- D. E. Stewart, A. Sarkar and J. E. Wampler, Occurrence and role of cis peptide-bonds in protein structures, J. Mol. Biol., 1990, 214, 253–260 Search PubMed.
- D. Pal and P. Chakrabarti, Cis Peptide Bonds in Proteins: Residues Involved, their Conformations, Interactions and Locations, J. Mol. Biol., 1999, 294, 271–288 CrossRef CAS PubMed.
- T. R. Alderson, J. H. Lee, C. Charlier, J. F. Ying and A. Bax, Propensity for cis-Proline Formation in Unfolded Proteins, ChemBioChem, 2018, 19, 37–42 CrossRef PubMed.
- A. Jabs, M. S. Weiss and R. Hilgenfeld, Non-proline cis peptide bonds in proteins, J. Mol. Biol., 1999, 286, 291–304 Search PubMed.
- C. M. Wilmot and J. M. Thornton, Beta-Turns and Their Distortions – a Proposed New Nomenclature, Protein Eng., 1990, 3, 479–493 Search PubMed.
- E. G. Hutchinson and J. M. Thornton, A revised set of potentials for beta-turn formation in proteins, Protein Sci., 1994, 3, 2207–2216 CrossRef CAS PubMed.
- M. Shapovalov, S. Vucetic and R. L. Dunbrack, A new clustering and nomenclature for beta turns derived from high-resolution protein structures, PLoS Comput. Biol., 2019, 15, e1006844 CrossRef PubMed.
- C. Grathwohl and K. Wüthrich, NMR studies of the rates of proline cis–trans isomerization in oligopeptides, Biopolymers, 1981, 20, 2623–2633 CrossRef CAS.
- W. J. Wedemeyer, E. Welker and H. A. Scheraga, Proline cis–trans isomerization and protein folding, Biochemistry, 2002, 41, 14637–14644 CrossRef CAS PubMed.
- G. Fischer, Peptidyl-prolyl cis/trans isomerases and their effectors, Angew. Chem., Int. Ed. Engl., 1994, 33, 1415–1436 CrossRef.
- F. X. Schmid, Prolyl isomerases, Adv. Protein Chem., 2002, 59, 243–282 Search PubMed.
- C. Odefey, L. M. Mayr and F. X. Schmid, Non-prolyl cis–trans peptide bond isomerization as a rate-determining step in protein unfolding and refolding, J. Mol. Biol., 1995, 245, 69–78 Search PubMed.
- G. Scherer, M. L. Kramer, M. Schutkowski, U. Reimer and G. Fischer, Barriers to rotation of secondary amide peptide bonds, J. Am. Chem. Soc., 1998, 120, 5568–5574 Search PubMed.
- G. Pappenberger, H. Aygun, J. W. Engels, U. Reimer, G. Fischer and T. Kiefhaber, Nonprolyl cis peptide bonds in unfolded proteins cause complex folding kinetics, Nat. Struct. Biol., 2001, 8, 452–458 Search PubMed.
- C. Schiene-Fischer, J. Habazettl, F. X. Schmid and G. Fischer, The hsp70 chaperone DnaK is a secondary amide peptide bond cis–trans isomerase, Nat. Struct. Biol., 2002, 9, 419–424 CrossRef CAS PubMed.
- S. Lorenzen, B. Peters, A. Goede, R. Preissner and C. Frommel, Conservation of cis prolyl bonds in proteins during evolution, Proteins: Struct., Funct., Bioinf., 2005, 58, 589–595 CrossRef CAS PubMed.
- S. C. R. Lummis, D. L. Beene, L. W. Lee, H. A. Lester, R. W. Broadhurst and D. A. Dougherty, Cis–trans isomerization at a proline opens the pore of a neurotransmitter-gated channel, Nature, 2005, 438, 248–252 Search PubMed.
- K. P. Lu, G. Finn, T. H. Lee and L. K. Nicholson, Prolyl cis–trans isomerization as a molecular timer, Nat. Chem. Biol., 2007, 3, 619–629 CrossRef CAS PubMed.
- P. A. M. Schmidpeter, J. R. Koch and F. X. Schmid, Control of protein function by prolyl isomerization, Biochim. Biophys. Acta, Gen. Subj., 2015, 1850, 1973–1982 Search PubMed.
- K. S. Mineev, M. A. Chernykh, V. V. Motov, D. A. Prudnikova, D. M. Pavlenko, A. I. Kuzmenkov, S. Peigneur, J. Tytgat and A. A. Vassilevski, A scorpion toxin affecting sodium channels shows double cis–trans isomerism, FEBS Lett., 2023, 597, 2358–2368 Search PubMed.
- R. Bhattacharyya and P. Chakrabarti, Stereospecific Interactions of Proline Residues in Protein Structures and Complexes, J. Mol. Biol., 2003, 331, 925–940 Search PubMed.
- B. Wathen and Z. C. Jia, Local and nonlocal environments around cis peptides, J. Proteome Res., 2008, 7, 145–153 Search PubMed.
- S. D. Hanes, Prolyl isomerases in gene transcription, Biochim. Biophys. Acta, Gen. Subj., 2015, 1850, 2017–2034 Search PubMed.
- H. J. Dyson, M. Rance, R. A. Houghten, R. A. Lerner and P. E. Wright, Folding of Immunogenic Peptide-Fragments of Proteins in Water Solution: 1. Sequence Requirements for the Formation of a Reverse Turn, J. Mol. Biol., 1988, 201, 161–200 Search PubMed.
- U. Reimer, G. Scherer, M. Drewello, S. Kruber, M. Schutkowski and G. Fischer, Side-chain effects on peptidyl-prolyl cis/trans isomerization, J. Mol. Biol., 1998, 279, 449–460 Search PubMed.
- W.-J. Wu and D. P. Raleigh, Local Control of Peptide Conformation: Stabilization of cis Proline Peptide Bonds by Aromatic Proline Interactions, Biopolymers, 1998, 45, 381–394 Search PubMed.
- F. Nardi, J. Kemmink, M. Sattler and R. C. Wade, The cisproline(i-1)-aromatic(i) interaction: Folding of the Ala-cisPro-Tyr peptide characterized by NMR and theoretical approaches, J. Biomol. NMR, 2000, 17, 63–77 CrossRef CAS PubMed.
- L. Halab and W. D. Lubell, Effect of sequence on peptide geometry in 5-tert-butylprolyl type VI beta-turn mimics, J. Am. Chem. Soc., 2002, 124, 2474–2484 CrossRef CAS PubMed.
- K. M. Thomas, D. Naduthambi and N. J. Zondlo, Electronic control of amide cis–trans isomerism via the aromatic-prolyl interaction, J. Am. Chem. Soc., 2006, 128, 2216–2217 CrossRef CAS PubMed.
- A. M. Brown and N. J. Zondlo, A Propensity Scale for Type II Polyproline Helices (PPII): Aromatic Amino Acids in Proline-Rich Sequences Strongly Disfavor PPII Due to Proline-Aromatic Interactions, Biochemistry, 2012, 51, 5041–5051 Search PubMed.
- N. J. Zondlo, Aromatic-Proline Interactions: Electronically Tunable CH/pi Interactions, Acc. Chem. Res., 2013, 46, 1039–1049 CrossRef CAS PubMed.
- M. B. Yaffe, M. Schutkowski, M. H. Shen, X. Z. Zhou, P. T. Stukenberg, J. U. Rahfeld, J. Xu, J. Kuang, M. W. Kirschner, G. Fischer, L. C. Cantley and K. P. Lu, Sequence-specific and phosphorylation-dependent proline isomerization: A potential mitotic regulatory mechanism, Science, 1997, 278, 1957–1960 CrossRef CAS PubMed.
- M. Schutkowski, A. Bernhardt, X. Z. Zhou, M. H. Shen, U. Reimer, J. U. Rahfeld, K. P. Lu and G. Fischer, Role of phosphorylation in determining the backbone dynamics of the serine/threonine-proline motif and Pin1 substrate recognition, Biochemistry, 1998, 37, 5566–5575 Search PubMed.
- M. Weiwad, A. Werner, P. Rucknagel, A. Schierhorn, G. Kullertz and G. Fischer, Catalysis of proline-directed protein phosphorylation by peptidyl-prolyl cis/trans isomerases, J. Mol. Biol., 2004, 339, 635–646 CrossRef CAS PubMed.
- Y. C. Liou, X. Z. Zhou and K. P. Lu, Prolyl isomerase Pin1 as a molecular switch to determine the fate of phosphoproteins, Trends Biochem. Sci., 2011, 36, 501–514 Search PubMed.
- K. Nakamura, A. Greenwood, L. Binder, E. H. Bigio, S. Denial, L. Nicholson, X. Z. Zhou and K. P. Lu, Proline Isomer-Specific Antibodies Reveal the Early Pathogenic Tau Conformation in Alzheimer's Disease, Cell, 2012, 149, 232–244 Search PubMed.
- S. N. Zhuang, P. Chakraborty and M. Zweckstetter, Regulation of tau by peptidyl-prolyl isomerases, Curr. Opin. Struct. Biol., 2024, 84, 102799 CrossRef PubMed.
- Z. M. Lu and T. Hunter, Prolyl isomerase Pin1 in cancer, Cell Res., 2014, 24, 1033–1049 CrossRef CAS PubMed.
- L. Wang, Y. Zhou, D. M. Chen and T. H. Lee, Peptidyl-Prolyl Cis/Trans Isomerase Pin1 and Alzheimer's Disease, Front. Cell Dev. Biol., 2020, 8, 355 CrossRef PubMed.
- A. R. Bataille, C. Jeronimo, P. E. Jacques, L. Laramee, M. E. Fortin, A. Forest, M. Bergeron, S. D. Hanes and F. Robert, A Universal RNA Polymerase II CTD Cycle Is Orchestrated by Complex Interplays between Kinase, Phosphatase, and Isomerase Enzymes along Genes, Mol. Cell, 2012, 45, 158–170 Search PubMed.
- M. M. Zhang, X. J. Wang, X. Chen, M. E. Bowman, Y. H. Luo, J. P. Noel, A. D. Ellington, F. A. Etzkorn and Y. Zhang, Structural and Kinetic Analysis of Prolyl-isomerization/Phosphorylation Cross-Talk in the CTD Code, ACS Chem. Biol., 2012, 7, 1462–1470 Search PubMed.
- E. B. Gibbs, F. Y. Lu, B. Portz, M. J. Fisher, B. P. Medellin, T. N. Laremore, Y. J. Zhang, D. S. Gilmour and S. A. Showalter, Phosphorylation induces sequence-specific conformational switches in the RNA polymerase II C-terminal domain, Nat. Commun., 2017, 8, article 15233 CrossRef PubMed.
- D. Eick and M. Geyer, The RNA Polymerase II Carboxy-Terminal Domain (CTD) Code, Chem. Rev., 2013, 113, 8456–8490 Search PubMed.
- D. J. Sutor, Evidence for Existence of C–H⋯O Hydrogen Bonds in Crystals, J. Chem. Soc., 1963, 1105–1110 RSC.
- S. Krimm, Hydrogen Bonding of C–H⋯O
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C in Proteins, Science, 1967, 158, 530–531 CrossRef CAS PubMed.
C in Proteins, Science, 1967, 158, 530–531 CrossRef CAS PubMed. - Z. S. Derewenda, L. Lee and U. Derewenda, The Occurence of C–H⋯O Hydrogen Bonds in Proteins, J. Mol. Biol., 1995, 252, 248–262 Search PubMed.
- G. R. Desiraju, The C–H⋯O Hydrogen Bond: Structural Implications and Supramolecular Design, Acc. Chem. Res., 1996, 29, 441–449 Search PubMed.
- P. Chakrabarti and S. Chakrabarti, C–H⋯O hydrogen bond involving proline residues in alpha-helices, J. Mol. Biol., 1998, 284, 867–873 Search PubMed.
- Y. L. Gu, T. Kar and S. Scheiner, Fundamental properties of the CH⋯O interaction: Is it a true hydrogen bond?, J. Am. Chem. Soc., 1999, 121, 9411–9422 Search PubMed.
- C. R. Jones, P. K. Baruah, A. L. Thompson, S. Scheiner and M. D. Smith, Can a C–H⋯O Interaction Be a Determinant of Conformation?, J. Am. Chem. Soc., 2012, 134, 12064–12071 CrossRef CAS PubMed.
- S. Horowitz and R. C. Trievel, Carbon-Oxygen Hydrogen Bonding in Biological Structure and Function, J. Biol. Chem., 2012, 287, 41576–41582 CrossRef CAS PubMed.
- H. C. Oven, G. P. A. Yap and N. J. Zondlo, Helical twists and b-turns in structures at serine-proline sequences: stabilization of cis-proline and type VI b-turns via C–H/O interactions, Proteins, 2024, 92, 1190–1205 CrossRef CAS PubMed.
- H. K. Ganguly, B. A. Ludwig, C. M. Tressler, M. R. Bhatt, A. K. Pandey, C. M. Quinn, S. Bai, G. P. A. Yap and N. J. Zondlo, 4,4-Difluoroproline as a Unique 19F NMR Probe of Proline Conformation, Biochemistry, 2024, 63, 1131–1146 CrossRef CAS PubMed.
- H. K. Ganguly, M. B. Elbaum, N. J. Daniecki and N. J. Zondlo, Serine-404 Phosphorylation and the R406W Modification in Tau Stabilize the cis-Proline Amide Bond, via Phosphoserine-Proline C–H/O and Proline-Aromatic C–H/π Interactions, ChemRxiv, 2023, preprint DOI:10.26434/chemrxiv-2023-w152b.
- G. Merutka, H. J. Dyson and P. E. Wright, Random Coil H-1 Chemical-Shifts Obtained as a Function of Temperature and Trifluoroethanol Concentration for the Peptide Series Ggxgg, J. Biomol. NMR, 1995, 5, 14–24 Search PubMed.
- S. Scheiner, T. Kar and Y. L. Gu, Strength of the (CH)-H-alpha center dot center dot O hydrogen bond of amino acid residues, J. Biol. Chem., 2001, 276, 9832–9837 CrossRef CAS PubMed.
- S. Scheiner and T. Kar, Effect of solvent upon CH center dot center dot center dot O hydrogen bonds with implications for protein folding, J. Phys. Chem. B, 2005, 109, 3681–3689 CrossRef CAS PubMed.
- N. J. Daniecki, M. R. Bhatt, G. P. A. Yap and N. J. Zondlo, Proline C–H Bonds as Loci for Proline Assembly via C–H/O Interactions, ChemBioChem, 2022, 23, e202200409 CrossRef CAS PubMed.
- B. Nepal and S. Scheiner, Anionic CH center dot center dot center dot X-Hydrogen Bonds: Origin of Their Strength, Geometry, and Other Properties, Chem. – Eur. J., 2015, 21, 1474–1481 CrossRef CAS PubMed.
- G. Wang and R. L. Dunbrack, PISCES: a protein sequence culling server, Bioinformatics, 2003, 19, 1589–1591 Search PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. J. A. Montgomery, J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, E. Brothers, K. N. Kudin, V. N. Staroverov, T. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, O. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian 09, Revision D.01, Gaussian, Inc., Wallingford, CT, 2013 Search PubMed.
- Y. Zhao and D. G. Truhlar, The M06 suite of density functionals for main group thermochemistry, thermochemical kinetics, noncovalent interactions, excited states, and transition elements: two new functionals and systematic testing of four M06-class functionals and 12 other functionals, Theor. Chem. Acc., 2008, 120, 215–241 Search PubMed.
- K. Raghavachari, J. S. Binkley, R. Seeger and J. A. Pople, Self-Consistent Molecular Orbital Methods. 20. Basis sets for correlated wave functions, J. Chem. Phys., 1980, 72, 650–654 Search PubMed.
- J. Tomasi, B. Mennucci and E. Cances, The IEF version of the PCM solvation method: an overview of a new method addressed to study molecular solutes at the QM ab initio level, J. Mol. Struct. THEOCHEM, 1999, 464, 211–226 Search PubMed.
- H. K. Ganguly, M. B. Elbaum and N. J. Zondlo, Proline-Aromatic Sequences Stabilize Turns via C–H/π interactions in both cis-Proline and trans-Proline, Biochemistry, 2025, 64, 2848–2866 CrossRef CAS PubMed.
- A. K. Pandey, H. K. Ganguly, S. K. Sinha, K. E. Daniels, G. P. A. Yap, S. Patel and N. J. Zondlo, An Inherent Structural Difference Between Serine and Threonine Phosphorylation: Phosphothreonine Prefers an Ordered, Compact, Cyclic Conformation, ACS Chem. Biol., 2023, 18, 1938–1958 Search PubMed.
- C. R. Glendening, C. R. Landis and F. Weinhold, Natural bond orbital methods, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2012, 2, 1–42 Search PubMed.
- E. D. Glendening, C. R. Landis and F. Weinhold, NBO 6.0: Natural bond orbital analysis program, J. Comput. Chem., 2013, 34, 1429–1437 Search PubMed.
- R. F. W. Bader, Atoms in Molecules, Acc. Chem. Res., 1985, 18, 9–15 Search PubMed.
- T. Lu and F. W. Chen, Multiwfn: A multifunctional wavefunction analyzer, J. Comput. Chem., 2012, 33, 580–592 CrossRef CAS PubMed.
- T. Lu, A comprehensive electron wavefunction analysis toolbox for chemists, Multiwfn, J. Chem. Phys., 2024, 161, 082503 Search PubMed.
- S. C. Lovell, J. M. Word, J. S. Richardson and D. C. Richardson, The Penultimate Rotamer Library, Proteins, 2000, 40, 389–408 Search PubMed.
- M. Vijayakumar, H. Qian and H. X. Zhou, Hydrogen bonds between short polar side chains and peptide backbone: Prevalence in proteins and effects on helix-forming propensities, Proteins, 1999, 34, 497–507 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery, J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16, Revision C.01, Gaussian, Inc., Wallingford, CT, 2019 Search PubMed.
- J. C. Ma and D. A. Dougherty, The Cation–π Interaction, Chem. Rev., 1997, 97, 1303–1324 Search PubMed.
- M. Nishio, CH/pi hydrogen bonds in crystals, CrystEngComm, 2004, 6, 130–158 RSC.
- M. Nishio, Y. Umezawa, J. Fantini, M. S. Weiss and P. Chakrabarti, CH-pi hydrogen bonds in biological macromolecules, Phys. Chem. Chem. Phys., 2014, 16, 12648–12683 RSC.
- S. Emamian, T. Lu, H. Kruse and H. Emamian, Exploring Nature and Predicting Strength of Hydrogen Bonds: A Correlation Analysis Between Atoms-in-Molecules Descriptors, Binding Energies, and Energy Components of Symmetry-Adapted Perturbation Theory, J. Comput. Chem., 2019, 40, 2868–2881 Search PubMed.
- A. V. Marenich, S. V. Jerome, C. J. Cramer and D. G. Truhlar, Charge Model 5: An Extension of Hirshfeld Population Analysis for the Accurate Description of Molecular Interactions in Gaseous and Condensed Phases, J. Chem. Theory Comput., 2012, 8, 527–541 Search PubMed.
- K. P. Exarchos, T. P. Exarchos, G. Rigas, C. Papaloukas and D. I. Fotiadis, Extraction of consensus protein patterns in regions containing non-proline cis peptide bonds and their functional assessment, BMC Bioinf., 2011, 12, article 142 CrossRef PubMed.
- P. Magotti, I. Bauer, M. Igarashi, M. Babgoli, R. Marotta, D. Piomelli and G. Garau, Structure of Human N-Acylphosphatidylethanolamine-Hydrolyzing Phospholipase D: Regulation of Fatty Acid Ethanolamide Biosynthesis by Bile Acids, Structure, 2015, 23, 598–604 CrossRef CAS PubMed.
- S. Vos, J. deJersey and J. L. Martin, Crystal structure of Escherichia coli xanthine phosphoribosyltransferase, Biochemistry, 1997, 36, 4125–4134 Search PubMed.
- S. Vos, R. J. Parry, M. R. Burns, J. de Jersey and J. L. Martin, Structures of free and complexed forms of Escherichia coli xanthine-guanine phosphoribosyltransferase, J. Mol. Biol., 1998, 282, 875–889 CrossRef CAS PubMed.
- C. D. Sessler, M. Rahm, S. Becker, J. M. Goldberg, F. Wang and S. J. Lippard, CF2H, a Hydrogen Bond Donor, J. Am. Chem. Soc., 2017, 139, 9325–9332 CrossRef CAS PubMed.
- S. Scheiner, Weak H-bonds. Comparisons of CH center dot center dot center dot O to NH center dot center dot center dot O in proteins and PH center dot center dot center dot N to direct P center dot center dot center dot N interactions, Phys. Chem. Chem. Phys., 2011, 13, 13860–13872 RSC.
- A. Senes, I. Ubarretxena-Belandia and D. M. Engelman, The C alpha-H center dot center dot center dot O hydrogen bond: A determinant of stability and specificity in transmembrane helix interactions, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 9056–9061 Search PubMed.
- A. Senes, D. E. Engel and W. F. DeGrado, Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs, Curr. Opin. Struct. Biol., 2004, 14, 465–479 Search PubMed.
- A. Warshel, Computer Modeling of Chemical Reactions in Enzymes and Solutions, Wiley, 1997 Search PubMed.
- A. Prah, M. Purg, J. Stare, R. Vianello and J. Mavri, How Monoamine Oxidase A Decomposes Serotonin: An Empirical Valence Bond Simulation of the Reactive Step, J. Phys. Chem. B, 2020, 124, 8259–8265 CrossRef CAS PubMed.
- B. Nepal and S. Scheiner, Effect of Ionic Charge on the CH center dot center dot center dot pi Hydrogen Bond, J. Phys. Chem. A, 2014, 118, 9575–9587 CrossRef CAS PubMed.
- E. A. Bienkiewicz and K. J. Lumb, Random-coil chemical shifts of phosphorylated amino acids, J. Biomol. NMR, 1999, 15, 203–206 CrossRef CAS PubMed.
- R. Hendus-Altenburger, C. B. Fernandes, K. Bugge, M. B. A. Kunze, W. Boomsma and B. B. Kragelund, Random coil chemical shifts for serine, threonine and tyrosine phosphorylation over a broad pH range, J. Biomol. NMR, 2019, 73, 713–725 Search PubMed.
- D. L. Z. Caetano, R. Metzler, A. G. Cherstvy and S. J. de Carvalho, Adsorption of lysozyme into a charged confining pore, Phys. Chem. Chem. Phys., 2021, 23, 27195–27206 RSC.
- F. A. Africo, A. G. Cherstvy and S. J. de Carvalho, Critical adsorption of polyelectrolytes onto highly oppositely charged surfaces: Effects of charge renormalization, J. Chem. Phys., 2024, 161, 194905 CrossRef CAS PubMed.
- J. D. Baker, L. B. Shelton, D. L. Zheng, F. Favretto, B. A. Nordhues, A. Darling, L. E. Sullivan, Z. Y. Sun, P. K. Solanki, M. D. Martin, A. Suntharalingam, J. J. Sabbagh, S. Becker, E. Mandelkow, V. N. Uversky, M. Zweckstetter, C. A. Dickey, J. Koren and L. J. Blair, Human cyclophilin 40 unravels neurotoxic amyloids, PLoS Biol., 2017, 15, 2001336 CrossRef PubMed.
- M. Babu, F. Favretto and M. Zweckstetter, Peptidyl Prolyl Isomerase A Modulates the Liquid-Liquid Phase Separation of Proline-Rich IDPs, J. Am. Chem. Soc., 2022, 144, 16157–16163 Search PubMed.
- G. L. Parra, E. J. Sohn, X. Xu and D. S. Libich, The Spliceosomal Peptidyl Prolyl Isomerase Like 1 Interacts with the Low-Complexity Domain of the RNA Binding Protein EWS Modulating Its Phase Separation Behavior, Biochemistry, 2025, 64, 3173–3177 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2026 |