Open Access Article
Open Access ArticleProbing histidine tautomers by theoretical X-ray absorption spectroscopy for biological and pathological studies
Hyundong
Kim
a,
Sompriya
Chatterjee
ab,
Myounwoo
Kim
a,
Yeonsig
Nam
*b,
Shaul
Mukamel
 *b and
Jin Yong
Lee
*b and
Jin Yong
Lee
 *a
*a
aDepartment of Chemistry, Sungkyunkwan University, Suwon 16419, Korea. E-mail: jinylee@skku.edu
bDepartment of Chemistry and Physics & Astronomy, University of California, Irvine, CA 92697-2025, USA. E-mail: ysnamc91@gmail.com; smukamel@uci.edu
First published on 6th February 2026
Abstract
Differentiating histidine tautomeric states is critical for understanding their role in neurodegenerative disease pathogenesis. Shifts in tautomeric equilibrium may contribute to driving protein misfolding and aggregation, suggesting potential therapeutic interventions to mitigate disease progression. Using the second-order restricted active space perturbation theory (RASPT2) method, this study proposes X-ray absorption spectroscopy (XAS) as a viable approach for distinguishing histidine tautomeric states. Pristine histidine exhibits distinct N K-edge spectral features depending on its tautomeric form. To improve tautomer discrimination, functionalization of histidine with various electron-donating or withdrawing groups was explored to modify the electronic environment of the imidazole ring. Substituent effects induce shifts of major peaks, enhancing spectral separability. This strategy highlights XAS labeling as a promising tool for differentiating tautomeric structures in complex biomolecules and will provide novel insights into histidine's role in both biological and pathological processes, ultimately informing therapeutic discovery.
Introduction
Histidine is an amino acid integral to various biological processes, contributing to protein structure,1 enzymatic catalysis,2,3 metal ion coordination,4,5 and buffer systems.6 Its imidazole side chain offers versatility, enabling non-covalent interactions such as hydrogen bonding and metal ion coordination, as well as catalysis of acid–base reactions.4,7 A distinctive trait that sets histidine apart from other amino acids is its ability to tautomerize, wherein the imidazole proton relocates between the two ring nitrogens (Nδ and Nε, Fig. 1a). This process enables histidine to create distinct electronic and structural forms to participate in diverse chemical reactions and supramolecular interactions.8 Tautomerism, a feature observed in biomolecules such as nucleic acid bases,9,10 and amino acids,11 influences biochemical processes and has been linked to neurodegenerative diseases like Alzheimer's (AD),12 Parkinson's (PD),13 and prion diseases.14 According to the amyloid hypothesis, a key factor in AD is the aggregation of amyloid beta (Aβ) peptides, derived from the cleavage of amyloid precursor protein (APP) by β- and γ-secretases,15–18 where histidine residues may play a modulatory role.In the Aβ (1–40) sequence, histidine residues at positions 6, 13, and 14 influence aggregation pathways. Replica-exchange molecular dynamics (REMD) simulations indicate that the δ tautomer preferentially stabilizes aggregation-prone conformations.19 This finding motivated the 'histidine tautomerism hypothesis', which postulates that specific tautomeric states of histidine can modulate protein aggregation.20 Beyond Aβ (1–40), this hypothesis extends to Aβ (1–42),21 tau proteins,22 and prion proteins,23 suggesting that histidine tautomerism may represent a unifying mechanism influencing the onset or progression of multiple neurodegenerative diseases. A deeper understanding of how different tautomeric forms, especially the δ tautomer, stabilize or destabilize aggregation-prone conformations could thus guide the design of targeted therapies aimed at altering histidine tautomerization to prevent or slow protein aggregation. This, in turn, motivates approaches that can directly distinguish histidine tautomeric states.
While understanding histidine tautomerism is essential for exploring its role in various biochemical and pathological processes, achieving direct experimental differentiation remains challenging due to the subtle electronic and structural differences between tautomers. Differentiating histidine tautomers can provide critical insights into how tautomeric shifts influence protein structure, aggregation, and function, as seen in neurodegenerative diseases like Alzheimer's and Parkinson's, where these factors are relevant to disease progression and therapeutic design. Current techniques, including low-temperature nuclear magnetic resonance (NMR) and mass spectrometry (MS), have provided valuable information on tautomeric molar ratios24,25 and allowed differentiation through labeling,26 though their resolution of individual tautomeric states remains limited. Notably, 2D IR spectroscopy has been employed to examine the impact of histidine tautomers on Aβ aggregation, providing indirect evidence of how tautomeric shifts influence peptide aggregation.27,28 However, 2D IR spectroscopy primarily reflects changes in secondary or tertiary protein structure, rather than directly probing tautomeric shifts within the histidine imidazole ring. Addressing these limitations will require novel experimental approaches or computational techniques that enable direct and precise differentiation of histidine tautomers.
Although absorption spectroscopy is a powerful technique for probing molecular electronic structure, it remains underutilized in this context. Previous efforts, such as Bermúdez's study using rotational spectroscopy,8 have highlighted the challenges of distinguishing histidine tautomers. Rotational spectroscopy faces limitations due to its limited resolution and its inability to perform the highly localized, element- and site-specific analyses necessary to differentiate closely related tautomeric states. Similarly, UV-visible (UV-vis) spectroscopy, which probes electronic transitions of valence electrons, provides insights into molecular structure but lacks the element- and site-specificity needed to resolve subtle electronic differences at individual atoms. In contrast, X-ray absorption spectroscopy (XAS) offers a significant advantage by exciting core electrons to unoccupied states at much higher energies. By targeting core electron shells, XAS enables highly localized, element-specific, and site-specific analysis, making it a well-suited method for distinguishing histidine's tautomeric states. This capability positions XAS as a promising tool for advancing our understanding of complex biomolecular systems.29 Therefore, by leveraging XAS's site specificity for core electrons, the present study aims to address the challenges in resolving closely related tautomeric forms.
In this study, we employed multi-reference electronic structure theory to investigate the spectroscopic difference between tautomers. We show that tautomers are spectrally distinguishable due to the different core hole relaxation and screening induced by the position of the proton. We further discuss how the functionalized imidazole ring affects the spectroscopic features.
Methods
Electronic structure calculations
The pristine histidine and the derivatives with three tautomeric forms were optimized at the B3LYP/aug-cc-pVDZ level by using OpenMolcas software.30 All optimized geometries are provided in the SI.To study the electronic structure of the histidine tautomers and the derivatives, we employed a post-Hartree–Fock multi-reference wave function approach in the form of state-averaged restricted active space self-consistent field (SA-RASSCF) followed by second-order restricted active space perturbation theory (RASPT2) calculation, an extension of complete active space self-consistent field (CASSCF), and second-order complete active space perturbation theory (CASPT2). These methods can predict core-excited final states with high orbital accuracy, and therefore, they are widely used for X-ray spectroscopic simulations of small molecular systems, particularly when compared to alternative electronic structure approaches.31,32 For SA-RASSCF simulations with 10 roots, we used active space (10, 1/9/0) for the pristine histidine and (12, 1/11/0) for the derivatives, where the number in parentheses refers to the number of electrons in the active space, and the number of active orbitals in the RAS1/RAS2/RAS3 space, respectively. In the active space, π bonding (πRing) and antibonding 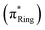 orbitals formed along the imidazole ring are included (Fig. 1b and c). To accurately describe the core electrons, we employed uncontracted basis sets of aug-cc-pCVTZ33,34 on every nitrogen atom and aug-cc-pVDZ on the other atoms. We enforced a single hole in the core orbital to calculate the core-excited state. RASPT2 calculations were done on top of the RASSCF reference by implementing an imaginary shift of 0.2 a.u. to prevent the intruder state problem. Among the 10 calculated roots, only the converged states predominantly characterized by single-electron excitations were selected for analysis, as they are responsible for the near-edge features in the N K-edge spectra. Canonical molecular orbitals obtained from RASPT2 calculations were visualized using an isovalue of 0.3 a.u.
orbitals formed along the imidazole ring are included (Fig. 1b and c). To accurately describe the core electrons, we employed uncontracted basis sets of aug-cc-pCVTZ33,34 on every nitrogen atom and aug-cc-pVDZ on the other atoms. We enforced a single hole in the core orbital to calculate the core-excited state. RASPT2 calculations were done on top of the RASSCF reference by implementing an imaginary shift of 0.2 a.u. to prevent the intruder state problem. Among the 10 calculated roots, only the converged states predominantly characterized by single-electron excitations were selected for analysis, as they are responsible for the near-edge features in the N K-edge spectra. Canonical molecular orbitals obtained from RASPT2 calculations were visualized using an isovalue of 0.3 a.u.
To better mimic aqueous experimental conditions, the polarizable continuum model (PCM) as the conductor model (C-PCM) implemented in OpenMolcas was used during geometric optimizations and RASSCF/PT2 calculations.35 As Y. Nishimoto reported that the combination of CASPT2 and PCM can sufficiently predict a compound's spectroscopic properties,36 the solvent effect was considered by incorporating nonequilibrium C-PCM in the RASSCF calculations.37 For the case of RASPT2 calculations, reaction field effects were added to the one-electron Hamiltonian. Transition dipole moments (TDMs) were computed from RASSCF wavefunctions by using the restricted active space state interaction (RASSI) module implemented in the OpenMolcas package.38
X-ray absorption spectroscopy calculations
The intensity of the X-ray absorption spectra was obtained from the imaginary part of the linear optical susceptibility χ(1) with respect to the absorption frequency ω using the following equation:39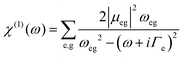 | (1) |
Results and discussion
XAS of pristine histidine
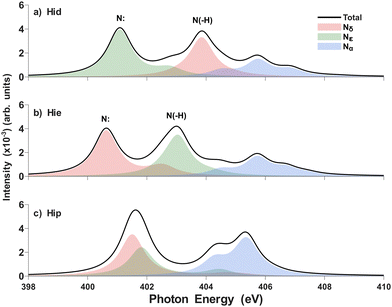
Fig. 1a shows the three major tautomeric forms (δ tautomer and ε tautomer) of zwitterionic or protonated (π) histidine alongside their protonation sites (Nδ and Nε), providing a structural framework for analyzing tautomeric shifts. The backbone nitrogen (Nα), which has a positive charge, also generates a consistent K-edge peak with respect to tautomeric states that is expected to be similar to the peak for ammonium in water.42 In the molecular orbital diagram for the δ tautomer, three nitrogen 1s orbitals are ordered by energy above the oxygen 1s orbitals, highlighting the distinct electronic environments of the nitrogen atoms (Fig. 1b). The ε tautomer shows a similar diagram with the inverted energy level of Nδ and Nε. Since histidine has an aromatic imidazole ring, the π* orbitals play an important role as arrival orbitals for the transition of N 1s orbitals.The theoretical XAS for pristine histidine (Fig. 2) shows three dominant peaks associated with Nδ, Nε, and Nα that are related to  orbitals. Two nitrogen atoms of the imidazole ring for the case of zwitterionic histidine can be regarded as unprotonated pyridinic (Nε for δ tautomer and Nδ for ε tautomer, N:) and protonated pyrrolic (N(–H)) nitrogen, respectively, and will produce well-resolved peaks.43 Initially, we focused on distinguishing the δ and ε tautomeric states of the pristine histidine, as these are the most prevalent forms under physiological conditions. In the calculated spectra, there are two major peaks originating from the imidazole ring's
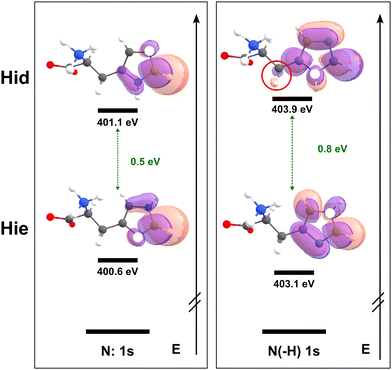
orbitals. Two nitrogen atoms of the imidazole ring for the case of zwitterionic histidine can be regarded as unprotonated pyridinic (Nε for δ tautomer and Nδ for ε tautomer, N:) and protonated pyrrolic (N(–H)) nitrogen, respectively, and will produce well-resolved peaks.43 Initially, we focused on distinguishing the δ and ε tautomeric states of the pristine histidine, as these are the most prevalent forms under physiological conditions. In the calculated spectra, there are two major peaks originating from the imidazole ring's 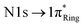 transitions (red and green). The δ tautomer's Nε peak at 401.1 eV is separated from the ε tautomer's Nδ peak at 400.6 eV. Likewise, the δ tautomer's Nδ peak (403.9 eV) is separated from the ε tautomer's Nε peak (403.1 eV). These distinct peaks show the possibility of two tautomers being spectroscopically distinguishable when probed by high spectral resolution X-ray spectroscopy.44 To find out the origin of this difference, we investigated the characteristics of molecular orbitals involved in the peaks (Fig. 3). The peaks from unprotonated pyridinic nitrogens are observed at lower photon energies and associated with excitations into a final state orbital with partial
transitions (red and green). The δ tautomer's Nε peak at 401.1 eV is separated from the ε tautomer's Nδ peak at 400.6 eV. Likewise, the δ tautomer's Nδ peak (403.9 eV) is separated from the ε tautomer's Nε peak (403.1 eV). These distinct peaks show the possibility of two tautomers being spectroscopically distinguishable when probed by high spectral resolution X-ray spectroscopy.44 To find out the origin of this difference, we investigated the characteristics of molecular orbitals involved in the peaks (Fig. 3). The peaks from unprotonated pyridinic nitrogens are observed at lower photon energies and associated with excitations into a final state orbital with partial  character (see left panel of Fig. 3). The higher ones are from protonated pyrrolic nitrogens to
character (see left panel of Fig. 3). The higher ones are from protonated pyrrolic nitrogens to  orbitals. We found that the later
orbitals. We found that the later  orbital can interact with the σ orbital on the beta carbon (Cβ, see Fig. 1a), which connects the imidazole ring and the backbone. This shows that the inequivalence between two imidazole nitrogens gives rise to distinct features in core electron excitation. The unprotonated pyridinic N K-edge features exhibit only minor shifts between tautomers, making a detailed qualitative interpretation difficult because arrival state orbitals show similar
orbital can interact with the σ orbital on the beta carbon (Cβ, see Fig. 1a), which connects the imidazole ring and the backbone. This shows that the inequivalence between two imidazole nitrogens gives rise to distinct features in core electron excitation. The unprotonated pyridinic N K-edge features exhibit only minor shifts between tautomers, making a detailed qualitative interpretation difficult because arrival state orbitals show similar  orbitals including no interaction between the backbone. In contrast, the pyrrolic N K-edge peaks display systematic shifts, reflecting the relevance to the excited electron's density. The major origin of the core excitation energy can be rationalized in terms of two competing effects. First, direct core hole relaxation by the valence electron density due to the reduced screening of the nucleus, which serves as an attractive energy that induces stabilization of the core level energy (N 1s energy in this study), resulting in a blue-shift in the final spectra from increasing transition energy. Second, polarization/screening arising from interaction between the excited electron and the valence electrons effectively reduces the core–hole potential and induces stabilization of the final core-excited state, resulting in a red-shift in the spectra.45 Direct core hole relaxation is not the primary cause, because the core-level energy associated with the pyrrolic site is nearly unchanged between tautomers. Therefore, the polarization/screening effect–i.e. differences in the excited electron's density (and resulting electron–hole interaction) must be dominant in this case. In the ε tautomer, the excited electron is delocalized over the imidazole ring while retaining substantial density near the core hole. This enhances the electron–hole interaction and the polarization/screening of the valence electrons toward the core hole, resulting in stabilization of the final state. In the δ tautomer, σ(C–C) or σ(C–H) on Cβ can interact with the
orbitals including no interaction between the backbone. In contrast, the pyrrolic N K-edge peaks display systematic shifts, reflecting the relevance to the excited electron's density. The major origin of the core excitation energy can be rationalized in terms of two competing effects. First, direct core hole relaxation by the valence electron density due to the reduced screening of the nucleus, which serves as an attractive energy that induces stabilization of the core level energy (N 1s energy in this study), resulting in a blue-shift in the final spectra from increasing transition energy. Second, polarization/screening arising from interaction between the excited electron and the valence electrons effectively reduces the core–hole potential and induces stabilization of the final core-excited state, resulting in a red-shift in the spectra.45 Direct core hole relaxation is not the primary cause, because the core-level energy associated with the pyrrolic site is nearly unchanged between tautomers. Therefore, the polarization/screening effect–i.e. differences in the excited electron's density (and resulting electron–hole interaction) must be dominant in this case. In the ε tautomer, the excited electron is delocalized over the imidazole ring while retaining substantial density near the core hole. This enhances the electron–hole interaction and the polarization/screening of the valence electrons toward the core hole, resulting in stabilization of the final state. In the δ tautomer, σ(C–C) or σ(C–H) on Cβ can interact with the  orbital, resulting in hyperconjugation. This means delocalization of π* electrons to the backbone, reducing electron density around the core hole (nitrogen) and further weakening screening. The pyrrolic peak therefore blueshifts in the δ tautomer. These features serve as reliable spectroscopic markers for distinguishing histidine tautomers.
orbital, resulting in hyperconjugation. This means delocalization of π* electrons to the backbone, reducing electron density around the core hole (nitrogen) and further weakening screening. The pyrrolic peak therefore blueshifts in the δ tautomer. These features serve as reliable spectroscopic markers for distinguishing histidine tautomers.
The backbone amine's signals (blue shades in Fig. 2a and b) are mainly attributed to σ*(Nα–H) with a relatively diffuse character. These peaks remain consistent and distant from the imidazole nitrogen peaks, making them negligible. For the case of the π histidine, the Nδ and Nε peaks that come from 1s to the final state exhibit partial  character and largely overlap to produce one intense peak. Nα produces a few distinct peaks that come from σ*(Nα–H) (Fig. S9). This implies that the protonated form can be easily distinguished from the zwitterionic tautomer.
character and largely overlap to produce one intense peak. Nα produces a few distinct peaks that come from σ*(Nα–H) (Fig. S9). This implies that the protonated form can be easily distinguished from the zwitterionic tautomer.
XAS of histidine derivatives
Even though the histidine itself shows a distinct spectrum by its tautomeric form, we tried to simulate XAS for the histidine derivatives to study the improvement of tautomer discrimination because about 0.5 eV of the difference sometimes becomes not distinguishable due to peak shifting or broadening by the adjacent metal ion, molecular vibration, or hydrogen bonding.46,47 We modified the electronic environment of nitrogen atoms in the imidazole ring by functionalizing the delta carbon (Cδ) that is bonded to Nε, as this site directly influences the electronic properties of Nδ and Nε. According to experimental studies,48–50 substitution of a functional group into the imidazole ring is one of the possible ways to achieve this.51,52 Modifying Cδ can cause asymmetric changes to Nδ and Nε whereas modifying Cε, which is bonded to Nδ, causes symmetrical changes to them, failing to improve spectral resolution. We selected three groups—methyl (–CH3) amino (–NH2) and fluoro (–F)—to study their influence on XAS, especially in the imidazole ring (Fig. 1a).For the methyl-substituted histidine (Fig. 4), for the pyridinic N peaks, the δ tautomer's Nε peak (400.6 eV, green) and ε tautomer's Nδ peak (400.2 eV, red) are observed whereas for the pyrrolic N peaks, the δ tautomer's Nδ peak (403.0 eV, red) and ε tautomer's Nε peak (403.3 eV) are produced. In the case of amino-histidine, it shows similar patterns to the case of methyl-substituted histidine, which are shown in the SI (Fig. S7). These results show no improvement in distinguishing between tautomers.
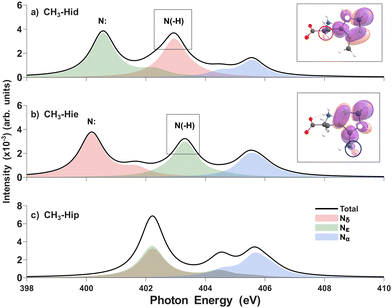 | ||
| Fig. 4 Theoretical N K-edge X-ray absorption spectra of methyl-substituted δ (a) and ε tautomer (b) and π histidine (c) by RASPT2 in aqueous solution, respectively. The main pyrrolic peaks’ corresponding isosurfaces for the excited electron's density are shown (black boxes). The white dots indicate the sites of excitation, which represent core holes. For other details, see the caption of Fig. 2. | ||
As the methyl group is a σ-donating group, it induces a few changes in the XAS spectra. Unprotonated pyridinic N peaks are only slightly shifted, whereas for protonated pyrrolic N peaks, the shift direction differs with the population of excited electrons in Fig. 4 (black boxes). Pyrrolic N peaks of substituted histidines show a hyperconjugation path along not only the backbone but also the substituent. In the δ tautomer, the excited electron shows predominant delocalization along the backbone carbon by hyperconjugation. In the ε tautomer, the excited electron density delocalizes along the substituent. This implies that there are two competitive hyperconjugation paths introduced by substitution. Our results show that delocalization to the methyl group rather than the backbone increases the excitation energy. As hyperconjugation exists between the imidazole ring and the methyl group, the  electron can delocalize toward the methyl group. Although the methyl group is a stronger σ-donor than the backbone, it does not provide an efficient screening channel.53,54 This will weaken the polarization/screening effect, which leads to a blueshift in the ε tautomer. The amine-substituted histidine also shows similar character (Fig. S11).
electron can delocalize toward the methyl group. Although the methyl group is a stronger σ-donor than the backbone, it does not provide an efficient screening channel.53,54 This will weaken the polarization/screening effect, which leads to a blueshift in the ε tautomer. The amine-substituted histidine also shows similar character (Fig. S11).
Besides, in fluoro-substituted histidine (Fig. 4), the pyridinic N peaks (400.9 eV for δ tautomer and 400.8 eV for ε tautomer) and pyrrolic N peak for the δ tautomer (402.6 eV) undergo a red-shift, whereas the pyrrolic N peak for the ε tautomer (404.6 eV) exhibits a blue-shift like previous cases, but the magnitude of the shift is much higher in the ε tautomer's Nε peak. As fluorine is regarded as a strong σ acceptor, it can stabilize the core hole energy level in both the δ and ε tautomers alike. In terms of the polarization/screening, since fluorine shows high electronegativity and low polarizability, more delocalization of excited electrons toward the fluorine means large reduction of the screening effect. As the ε tautomer shows more electron density around the fluorine, this induces a dramatic blueshift of the pyrrolic peak with respect to the δ tautomer. In other words, a larger charge displacement from the imidazole ring to the fluorine in the ε tautomer causes the signature blue-shift.55 Our Mulliken charge analysis of the initial and final state rationalizes this phenomenon (Table S3).
This substitution study reveals that inducing changes in the electronic environment along the imidazole ring causes shifts of existing N K-edge peaks; in particular, some substitutions can enhance tautomer discrimination. The magnitude of interaction between the imidazole ring and the backbone/substituent governs the polarization/screening effect; therefore, fluorine, a strong σ-acceptor, induces a noticeable difference between the δ and ε tautomers (Fig. 5).
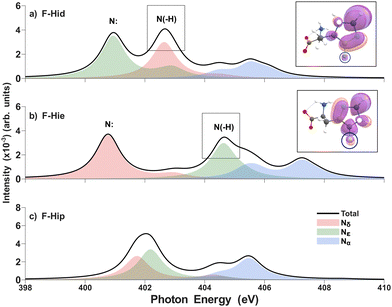 | ||
| Fig. 5 Theoretical N K-edge X-ray absorption spectra of the fluoro-substituted δ (a) and ε tautomer (b) and π histidine (c) by RASPT2 in aqueous solution. The main pyrrolic peaks’ corresponding isosurfaces for the excited electron's density are shown (black boxes). The white dots indicate the sites of excitation, which represent core holes. For other details, see the caption of Fig. 2. | ||
Conclusions
In summary, we propose N K-edge X-ray absorption spectroscopy (XAS) for distinguishing histidine tautomeric forms. Our computational analysis shows that the XAS spectra of pristine histidine show spectral features that differentiate its tautomeric states, originating from the different electronic structure between the pyrrolic or pyridinic nitrogen of the imidazole ring. While substitutions perturb electronic structures, particularly in the imidazole ring, when considering the polarization/screening effect, fluorine induces a pronounced shift of the pyrrolic N peak and makes it easier to distinguish the δ and ε tautomers. This approach opens new possibilities for using XAS to study biomolecules, particularly in biological and pathological mechanisms where histidine tautomerism plays a crucial role, such as enzymatic catalysis and protein aggregation. By enhancing our understanding of histidine's tautomeric states under diverse physiologically relevant conditions, this method could inform therapeutic strategies and advance drug development for neurodegenerative diseases.Conflicts of interest
There are no conflicts to declare.Data availability
The datasets used in this article are available in the supplementary information (SI). Supplementary information: background signal, optimized geometries, X-ray absorption spectra in the gas phase, X-ray absorption spectra-major peaks and molecular orbital transitions for the X-ray absorption spectra. See DOI: https://doi.org/10.1039/d5cp01475a.Acknowledgements
This work was supported by the National Research Foundation (NRF) grant funded by the Korean government (2021R1A2C2095412) and SKKU Academic Research Support Program (Samsung Research Fund), Sungkyunkwan University, 2025.Notes and references
- O. O. Olubiyi and B. Strodel, J. Phys. Chem. B, 2012, 116, 3280–3291 CrossRef CAS PubMed.
- J. Rebek, Struct. Chem., 1990, 1, 129–131 Search PubMed.
- M. E. Brosnan and J. T. Brosnan, J. Nutr., 2020, 150(Suppl. 1), 2570S–2575S CrossRef PubMed.
- R. J. Sundberg and R. B. Martin, Chem. Rev., 1974, 74, 471–517 CrossRef CAS.
- D. P. Smith, D. G. Smith, C. C. Curtain, J. F. Boas, J. R. Pilbrow, G. D. Ciccotosto, T. Lau, D. J. Tew, K. Perez, J. D. Wade, A. I. Bush, S. C. Drew, F. Separovic, C. L. Masters, R. Cappai and K. J. Barnham, J. Biol. Chem., 2006, 281, 15145–15154 Search PubMed.
- N. C. Stellwagen, C. Gelfi and P. G. Righetti, J. Chromatogr. A, 1999, 838, 179–189 CrossRef CAS PubMed.
- R. N. V. Krishna Deepak and R. Sankararamakrishnan, Biochemistry, 2016, 55, 3774–3783 CrossRef CAS PubMed.
- C. Bermúdez, S. Mata, C. Cabezas and J. L. Alonso, Angew. Chem., Int. Ed., 2014, 53, 11015–11018 CrossRef PubMed.
- M. Mons, I. Dimicoli, F. Piuzzi, B. Tardivel and M. Elhanine, J. Phys. Chem. A, 2002, 106, 5088–5094 Search PubMed.
- M. K. Shukla and J. Leszczynski, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2013, 3, 637–649 CAS.
- S.-P. Lu and A. H. Lewin, Tetrahedron, 1998, 54, 15097–15104 CrossRef CAS.
- D. Yanagisawa, N. Shirai, T. Amatsubo, H. Taguchi, K. Hirao, M. Urushitani, S. Morikawa, T. Inubushi, M. Kato, F. Kato, K. Morino, H. Kimura, I. Nakano, C. Yoshida, T. Okada, M. Sano, Y. Wada, K. Wada, A. Yamamoto and I. Tooyama, Biomaterials, 2010, 31, 4179–4185 CrossRef CAS PubMed.
- N. T. Tzvetkov, H.-G. Stammler and L. Antonov, J. Mol. Struct., 2017, 1149, 273–281 CrossRef CAS.
- K. Beyreuther and C. L. Masters, Brain Pathol., 1991, 1, 241–251 Search PubMed.
- R. K. Singh, N. G. Chamachi, S. Chakrabarty and A. Mukherjee, J. Phys. Chem. B, 2017, 121, 550–564 CrossRef CAS PubMed.
- J. A. Hardy and D. Allsop, Trends Pharmacol. Sci., 1991, 12, 383–388 CrossRef CAS PubMed.
- D. J. Selkoe, Neuron, 1991, 6, 487–498 CrossRef CAS PubMed.
- J. A. Hardy and G. A. Higgins, Science, 1992, 256, 184–185 CrossRef CAS PubMed.
- H. Shi, B. Kang and J. Y. Lee, J. Phys. Chem. B, 2016, 120, 11405–11411 CrossRef CAS PubMed.
- X. Xing, W. Zhao, D. Hu, B. Kang, H. Shi, J. Y. Lee and H. Ai, ACS Chem. Neurosci., 2019, 10, 2602–2608 Search PubMed.
- H. Shi and J. Y. Lee, ACS Chem. Neurosci., 2017, 8, 669–675 CrossRef CAS PubMed.
- H. Li, N. Li, Y. Tang and J. Y. Lee, ACS Chem. Neurosci., 2021, 12, 1983–1988 Search PubMed.
- S. Chatterjee, A. Salimi and J. Y. Lee, ACS Chem. Neurosci., 2021, 12, 3203–3213 Search PubMed.
- M. Tanokura, Biochim. Biophys. Acta, 1983, 742, 576–585 CrossRef CAS PubMed.
- W. F. Reynolds, I. R. Peak, M. H. Freedman and J. R. Lyerla, J. Am. Chem. Soc., 1973, 95, 328–331 CrossRef CAS PubMed.
- X. Pan, Z. J. Kirsch and R. W. Vachet, Anal. Chem., 2022, 94, 1003–1010 CrossRef CAS PubMed.
- Y. Nam, M. Kalathingal, S. Saito and J. Y. Lee, Biophys. J., 2020, 119, 831–842 CrossRef CAS PubMed.
- S. Chatterjee, Y. Nam, A. Salimi and J. Y. Lee, Phys. Chem. Chem. Phys., 2022, 24, 18691–18702 RSC.
- W. Malzer, C. Schlesiger and B. Kanngießer, Spectrochim. Acta, Part B, 2021, 177, 106101 CrossRef CAS.
- G. L. Manni, I. F. Galván, A. Alavi, F. Aleotti, F. Aquilante, J. Autschbach, D. Avagliano, A. Baiardi, J. J. Bao, S. Battaglia, L. Birnoschi, A. Blanco-González, S. I. Bokarev, R. Broer, R. Cacciari, P. B. Calio, R. K. Carlson, R. C. Couto, L. Cerdán, L. F. Chibotaru, N. F. Chilton, J. R. Church, I. Conti, S. Coriani, J. Cuéllar-Zuquin, R. E. Daoud, N. Dattani, P. Decleva, C. de Graaf, M. G. Delcey, L. De Vico, W. Dobrautz, S. S. Dong, R. Feng, N. Ferré, M. Filatov (Gulak), L. Gagliardi, M. Garavelli, L. González, Y. Guan, M. Guo, M. R. Hennefarth, M. R. Hermes, C. E. Hoyer, M. Huix-Rotllant, V. K. Jaiswal, A. Kaiser, D. S. Kaliakin, M. Khamesian, D. S. King, V. Kochetov, M. Krośnicki, A. A. Kumaar, E. D. Larsson, S. Lehtola, M.-B. Lepetit, H. Lischka, P. L. Ríos, M. Lundberg, D. Ma, S. Mai, P. Marquetand, I. C. D. Merritt, F. Montorsi, M. Mörchen, A. Nenov, V. H. A. Nguyen, Y. Nishimoto, M. S. Oakley, M. Olivucci, M. Oppel, D. Padula, R. Pandharkar, Q. M. Phung, F. Plasser, G. Raggi, E. Rebolini, M. Reiher, I. Rivalta, D. Roca-Sanjuán, T. Romig, A. A. Safari, A. Sánchez-Mansilla, A. M. Sand, I. Schapiro, T. R. Scott, J. Segarra-Martí, F. Segatta, D.-C. Sergentu, P. Sharma, R. Shepard, Y. Shu, J. K. Staab, T. P. Straatsma, L. K. Sørensen, B. N. C. Tenorio, D. G. Truhlar, L. Ungur, M. Vacher, V. Veryazov, T. A. Voß, O. Weser, D. Wu, X. Yang, D. Yarkony, C. Zhou, J. P. Zobel and R. Lindh, J. Chem. Theory Comput., 2023, 19, 6933–6991 CrossRef PubMed.
- F. Montorsi, F. Segatta, A. Nenov, S. Mukamel and M. Garavelli, J. Chem. Theory Comput., 2022, 18, 1003–1016 CrossRef CAS PubMed.
- L. Bäuml and R. de Vivie-Riedle, J. Phys. Chem. B, 2025, 129, 2159–2167 CrossRef PubMed.
- T. H. Dunning, J. Chem. Phys., 1989, 90, 1007–1023 CrossRef CAS.
- D. E. Woon and T. H. Dunning, J. Chem. Phys., 1995, 103, 4572–4585 CrossRef CAS.
- Y. Nishimoto, J. Chem. Theory Comput., 2025, 21, 730–746 CrossRef CAS PubMed.
- A. Bhattacharjee, M. Chavarot-Kerlidou, J. L. Dempsey, H. B. Gray, E. Fujita, J. T. Muckerman, M. Fontecave, V. Artero, G. M. Arantes and M. J. Field, ChemPhysChem, 2014, 15, 2951–2958 CrossRef CAS PubMed.
- P. A. Malmqvist, B. O. Roos and B. Schimmelpfennig, Chem. Phys. Lett., 2002, 357, 230–240 CrossRef CAS.
- B. Mennucci, J. Tomasi, R. Cammi, J. R. Cheeseman, M. J. Frisch, F. J. Devlin, S. Gabriel and P. J. Stephens, J. Phys. Chem. A, 2002, 106, 6102–6113 CrossRef CAS.
- S. Mukamel, Principles of nonlinear optical spectroscopy, Oxford University Press, New York, 1995 Search PubMed.
- V. V. da Cruz, S. Eckert and A. Föhlisch, Phys. Chem. Chem. Phys., 2021, 23, 1835–1848 RSC.
- C. Nicolas and C. Miron, J. Electron Spectrosc. Relat. Phenom., 2012, 185, 267–272 CrossRef CAS.
- M. Ekimova, M. Kubin, M. Ochmann, J. Ludwig, N. Huse, P. Wernet, M. Odelius and E. T. J. Nibbering, J. Phys. Chem. B, 2018, 122, 7737–7746 CrossRef CAS PubMed.
- F. Meyer, M. Blum, A. Benkert, D. Hauschild, Y. L. Jeyachandran, R. G. Wilks, W. Yang, M. Bär, F. Reinert, C. Heske, M. Zharnikov and L. Weinhardt, Phys. Chem. Chem. Phys., 2018, 20, 8302–8310 Search PubMed.
- J. W. Smith and R. J. Saykally, Chem. Rev., 2017, 117, 13909–13934 Search PubMed.
- T. Fransson, S. Coriani, O. Christiansen and P. Norman, J. Chem. Phys., 2013, 138, 124311 CrossRef PubMed.
- P. Frank, M. Benfatto, B. Hedman and K. O. Hodgson, Inorg. Chem., 2012, 51, 2086–2096 Search PubMed.
- R. Yamamura, T. Suenaga, M. Oura, T. Tokushima and O. Takahashi, Chem. Phys. Lett., 2020, 738, 136895 CrossRef CAS.
- J. M. Joo, B. B. Touré and D. Sames, J. Org. Chem., 2010, 75, 4911–4920 CrossRef CAS PubMed.
- A. Mahindra, N. Bagra and R. Jain, J. Org. Chem., 2013, 78, 10954–10959 CrossRef CAS PubMed.
- K. Sharma, K. K. Sharma, A. Mahindra, N. Sehra, N. Bagra, S. Aaghaz, R. Parmar, G. K. Rathod and R. Jain, Med. Res. Rev., 2023, 43, 775–828 CrossRef CAS PubMed.
- M. Wanko, J. Houmøller, K. Støchkel, M.-B. S. Kirketerp, M. Å. Peterson, M. B. Nielsen, S. B. Nielsen and A. Rubio, Phys. Chem. Chem. Phys., 2012, 14, 12905–12911 Search PubMed.
- S. Herbers, P. Buschmann, J. Wang, K. G. Lengsfeld, K. P. R. Nair and J.-U. Grabow, Phys. Chem. Chem. Phys., 2020, 22, 11490–11497 Search PubMed.
- G. Comas-Vilà and P. Salvador, Phys. Chem. Chem. Phys., 2025, 27, 10482–10491 Search PubMed.
- P. Bolognesi, V. Carravetta, L. Sementa, G. Barcaro, S. Monti, P. M. Mishra, A. Cartoni, M. C. Castrovilli, J. Chiarinelli, S. Tosic, B. P. Marinkovic, R. Richter and L. Avaldi, Front. Chem., 2019, 7, 151 Search PubMed.
- C. D. Pemmaraju, R. Copping, S. Wang, M. Janousch, S. J. Teat, T. Tyliszcak, A. Canning, D. K. Shuh and D. Prendergast, Inorg. Chem., 2014, 53, 11415–11425 Search PubMed.
| This journal is © the Owner Societies 2026 |