Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
From conformational lability to conformational control in short-peptide crystals
Joanna
Bojarska
 *a,
Martin
Breza
*a,
Martin
Breza
 b,
Sepideh
Jafari
b,
Sepideh
Jafari
 a and
Wojciech M.
Wolf
a
a and
Wojciech M.
Wolf
a
aChemistry Department, Institute of Ecological and Inorganic Chemistry, Technical University of Lodz, 116 Zeromskiego St., Lodz, 90-924, Poland. E-mail: joanna.bojarska@p.lodz.pl
bPhysical Chemistry, Slovak Technical University, Radlinskeho 9, Bratislava, Slovakia
First published on 17th April 2026
Abstract
Short peptides occupy a distinctive position between proteins and traditional small molecules. They combine the functional richness typical of biomolecules with the synthetic accessibility and crystallographic tractability characteristic of small-molecule systems. Their favourable physicochemical properties and exceptional structural diversity make them difficult to replicate with other classes of molecular building blocks, while enabling systematic exploration of sequence–structure relationships in crystalline materials. At the same time, short-peptide crystals are often regarded as conformationally labile, a feature that has traditionally been viewed as an obstacle to predictive peptide crystal engineering. Recent studies, however, suggest that conformational flexibility in short peptides can shift from being treated as a crystallographic complication to being explored as a potentially controllable structural variable. A key challenge is distinguishing genuine crystal-state conformational control from apparent structural changes arising from crystallisation pathways, phase conversion or recrystallisation. In this Highlight we propose a practical framework for identifying when conformational lability translates into experimentally demonstrable conformational control in peptide crystals. We discuss representative model systems, outline experimental strategies required to distinguish crystal-state conformational selection from pathway-dependent effects, and highlight emerging cases in which controlled peptide conformation influences structural organisation and functional behaviour in molecular materials.
Introduction
Short peptides occupy a unique position between proteins and traditional small organic molecules. They combine the structural programmability and functional richness typical of biomolecules with the synthetic accessibility and crystallographic tractability characteristic of small-molecule systems.1–3 This combination makes short peptides particularly useful model systems for peptide crystal engineering. Sequence-level modifications can often be correlated directly with changes in molecular conformation, intermolecular recognition and crystal packing. Compared with longer peptides, short sequences often display favourable physicochemical and pharmacokinetic properties. At the same time, they remain simple enough to allow systematic sequence variation and precise structural analysis.4 Despite their small size, short peptides generate remarkable structural and functional diversity. Even di- and tripeptides can form diverse supramolecular architectures through hydrogen bonding, aromatic interactions and other non-covalent forces.5–7 This versatility has positioned them as attractive platforms for designing functional molecular materials.8–10 At the same time, recent advances in peptide synthesis, biotechnology and bioinformatics are rapidly expanding the accessible sequence space and enabling more systematic exploration of peptide-based systems.11–13 These developments are helping overcome traditional limitations of peptide materials and revealing the full structural and functional potential of short-peptide assemblies. Consequently, interest in short peptides has grown rapidly across fields including supramolecular chemistry, crystal engineering, biomaterials and molecular electronics.2–5A defining feature of short peptides is their intrinsic conformational flexibility. Even very small sequences can adopt multiple backbone geometries depending on sequence, environment and intermolecular interactions. This conformational lability plays a central role in determining packing motifs, polymorphism and phase behaviour in peptide crystals. It also raises a key question for peptide crystal engineering: under what conditions can such flexibility be translated into reproducible conformational control within the crystal lattice? Addressing this question requires distinguishing genuine crystal-state conformational selection from alternative explanations such as phase conversion or recrystallisation. Understanding when conformational flexibility becomes a controllable structural variable may therefore represent an important step toward predictive peptide crystal engineering.
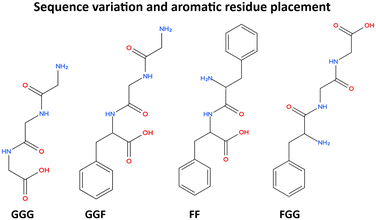
For clarity and accessibility, representative peptide structures discussed throughout this Highlight are illustrated schematically, allowing direct visual comparison of sequence-dependent structural variation. This is particularly relevant for readers from adjacent fields where peptide sequence notation may not be immediately intuitive. Where appropriate, key structural features and representative packing motifs are highlighted to improve clarity.
For the purposes of this Highlight, conformational control is considered most convincing when three criteria are satisfied:
(i) the peptide conformation is reproducibly stabilised within a defined crystal phase,
(ii) the conformational state correlates with specific packing or symmetry outcomes, and
(iii) alternative explanations such as growth effects or recrystallisation can be excluded experimentally. In this context, we use the term conformational control to describe situations in which a specific peptide backbone geometry is stabilised reproducibly by the crystal environment rather than arising from stochastic crystallisation pathways or phase transformations. Recent studies now suggest that in several well-characterised systems conformational lability can be steered to influence crystal form and packing and, in selected cases, measurable functional response.6–10 The broader structural context of conformational variability and polymorphism in peptide and organic crystals has been examined extensively.14–16 The conceptual framework explored in this Highlight is summarised in Scheme 1. Representative short-peptide model systems discussed in this work are shown in Scheme 2.
The following sections expand on these elements by discussing how intrinsic and extrinsic factors contribute to this framework, and how local conformational preferences propagate into packing and functional outcomes.
The following sections discuss how conformational lability in short peptides can, under suitable conditions, translate into experimentally observable conformational control in the solid state. We first discuss intrinsic molecular determinants that bias peptide conformation, followed by extrinsic environmental factors such as hydration and crystallisation conditions. Finally, we summarise experimental and computational approaches used to establish conformational control and highlight emerging examples in which controlled peptide conformation leads directly to measurable structural or functional responses in molecular materials.
The practical question is crystallographic: when does conformational lability become reproducibly translated into conformational control in short-peptide crystals, and how can this be distinguished from pathway-dependent growth, partial phase conversion or recrystallisation into a different phase? Cyclic and strongly constrained systems provide useful context, but the clearest examples still come from a small set of linear, hydration-sensitive and aromatic short-peptide systems. The field is now reaching a stage where a small number of well-characterised model systems enable systematic experimental examination of conformational control.
Molecular and environmental determinants of conformational control
In short-peptide crystals, conformational control rarely has a single origin. It usually reflects an interplay between molecular preorganization and environmental conditions. In the clearest cases, these factors work together strongly enough for local conformational preferences to influence the final crystal structure. To keep the discussion focused, the main narrative centres on systems with the strongest crystallographic support, while adjacent boundary cases are included only where they clarify the mechanistic limits of conformational control. Related studies on diphenylalanine systems and pH-responsive short-peptide assemblies further show how solvent environment, templated crystallisation and assembly conditions can redirect structural outcome in peptide materials.17–20Intrinsic factors
Intrinsic control arises from structural features of the peptide itself. Although it does not eliminate conformational lability, it can restrict the accessible conformational space sufficiently to favour specific geometries. Molecular preorganization is therefore a key factor in any discussion of conformational control. Here, preorganization refers to the tendency of a peptide to favour specific conformational states prior to the establishment of the full packing arrangement.11,12Cyclization provides a clear model for built-in conformational restriction because it limits accessible backbone geometry from the outset. In many cyclic-peptide studies, however, the evidence is stronger for biased self-assembly than for direct crystal-state conformational selection, so these systems are more useful as informative boundary cases rather than as direct prototypes for crystal-state switching in simple linear short peptides. This is largely because the strong preorganization imposed by cyclization already restricts conformational variability, leaving less scope for subtle conformational preferences to redirect crystal packing.11 Unlike cyclic peptides, which typically impose strong preorganization, linear short peptides remain synthetically flexible and sequence-programmable, making them particularly attractive platforms for exploring how local conformational preferences propagate into crystal packing.
Oligomers built from N-methyl-L/D-alanine residues further show that limited bond rotation can be used to program local peptide shape at the level of individual residues, and thus to introduce local structural preference through chain architecture. Their relevance here lies mainly in demonstrating programmable local preorganization rather than direct crystal-state selection in canonical short-peptide crystals.12
Beyond these systems, strong intrinsic conformational bias is also well documented in Aib (α-aminoisobutyric acid)-rich peptides, which stabilise 310- and α-helical geometries. Classic crystallographic studies established this behaviour in Aib-containing helices.21 More recent work and reviews confirm the broader role of non-canonical residues, including Aib, in preorganising peptide structure.22
Weak intramolecular contacts can also favour one local geometry over another, but their crystallographic significance is most convincing when they correlate with a resolved alternative packing arrangement or a conformationally distinct polymorph.13,15,16 In constrained helical peptides, an intramolecular CH⋯π interaction was linked to conformational transitions and conformational polymorphism within a defined temperature range.13 In that system, two stable polymorphs were observed, and density functional theory (DFT) calculations connected their relative stability to a switchable intramolecular CH⋯π interaction over a defined temperature window. Related observations have also been reported for peptide-derived systems containing pseudo-cyclic motifs stabilized by weak intramolecular interactions. Together with peptide-derived crystal families in which small local structural changes redirect supramolecular packing preferences, these studies show how subtle intramolecular preferences can influence local geometry without fully determining the final crystal-state outcome.15,16
Among the intrinsic factors, sequence programming is one of the clearest ways to steer structural outcome. In several well-characterised systems, sequence-dependent changes propagate into packing asymmetry, symmetry breaking and function. In many cases the clearest evidence concerns reproducible changes in packing and symmetry, while the conformational contribution itself is inferred indirectly from the resulting crystal structures.9,10 The strongest present evidence comes from constrained, aromatic and hydration-sensitive systems, including recent piezoelectric tripeptides in which residue placement changes crystal symmetry and electromechanical response. These systems provide some of the clearest recent examples illustrating how sequence-level structural preferences propagate into packing asymmetry and function.6–10
At the structural level, these effects are often reflected in specific hydrogen-bonding motifs and supramolecular synthons, such as head-to-tail chains, β-sheet-like assemblies or helical packing arrangements, which propagate local conformational preferences into long-range crystal organisation. These motifs are typically defined by backbone hydrogen-bond networks and can be correlated with specific backbone conformations such as extended pPII or β-sheet-like arrangements.
Extrinsic factors
Extrinsic influences on conformational control in short-peptide crystals arise mainly from hydration state and solvent environment. In several cases, conformational preferences correlate with specific supramolecular synthons, particularly hydrogen-bonded motifs that stabilise distinct backbone geometries and propagate into crystal packing arrangements. These interactions provide a structural link between local conformational bias and long-range packing organisation and can stabilise distinct local geometries or redirect crystallisation toward different structural outcomes.Hydration can stabilize a distinct, crystallographically resolved conformer within the crystal. Solvent effects, by contrast, more commonly redirect packing selection during crystal growth.
Triglycine is the clearest hydration-linked case discussed here, whereas diphenylalanine-derived systems are better viewed as solvent- or pathway-directed outcomes with conformational consequences. In triglycine, the hydrated form adopts an extended peptide backbone geometry (polyproline II, pPII), showing directly that water can stabilize a distinct crystal-state geometry through participation in the hydrogen-bond network.6 The pPII conformation is characterised by typical φ/ψ angles around −75°/145°, providing a distinct structural reference for comparison. The hydrated triglycine structure shown in Fig. 1 illustrates the extended pPII backbone stabilised by a water-mediated hydrogen-bond network within the crystal lattice.
Guo et al.6 identified the hydrated crystal specifically as a dihydrate adopting pPII geometry, providing one of the clearest examples so far of a water-stabilized crystal-state peptide conformation. Subsequent work suggested that humidity can convert a sheet-like form into a hydrated pPII form (Fig. 1), consistent with a hydration-linked structural change.7 However, the available evidence does not yet exclude partial dissolution and recrystallisation.
Although the authors interpret the change as solid-state, the accompanying morphological transformation in scanning electron microscopy (SEM) figures leaves open the possibility of surface dissolution–recrystallisation. For that reason, triglycine is best regarded as a useful experimental reference system rather than as a definitive example of direct crystal-state conformational switching.
Recent humidity-responsive peptide crystals also demonstrate that reversible structural reconfiguration can occur in peptide solids under changes in humidity. These observations suggest that controlled structural adaptation may be achievable in suitably designed peptide materials, although the precise role of conformational switching within a single crystal phase remains to be established. Such systems suggest that water can act not only as a crystallisation variable but also as a structural trigger in peptide solids.6,7 Resolving that point will likely require in situ variable-humidity single-crystal X-ray diffraction or a combination of variable-humidity powder X-ray diffraction (PXRD) with Raman spectroscopy or dynamic vapour sorption, ideally alongside direct monitoring of crystal morphology. Triglycine therefore serves as a useful reference system for studying hydration-linked conformational behaviour in short-peptide crystals.6,7
Solvent effects are illustrated most clearly by diphenylalanine-based systems. Different solvated crystal forms of diphenylalanine (FF) show that the same peptide can access different structural outcomes under different crystallisation conditions,17 while N-capped diphenylalanine demonstrates more directly that solvent composition can redirect crystal growth toward distinct packing states.8 In that system, solvent composition was shown to switch growth between two distinct crystal packings, including monoclinic and orthorhombic forms, providing an unusually clear example of solvent-dependent packing selection at the single-crystal level. These studies show that solvent can redirect crystal growth and favour different packing outcomes. They therefore support solvent-directed structural selection more directly than crystal-state conformational switching. Related work on templated crystallisation and secondary processing further reinforces the need to distinguish conformational selection from kinetic redirection of growth.18
pH is more useful as an emerging rather than an established handle for conformational control in short-peptide crystals. Ma et al. reported pH-dependent packing variation and chirality inversion at the assembly level.19 A correction to that article was published subsequently.20 The result is best regarded as a boundary case at the level of supramolecular assembly rather than as evidence for direct crystal-state conformational control in short-peptide crystals.19
Environmental factors can do more than perturb short-peptide crystallisation. In favourable systems, they can reinforce local conformational preferences strongly enough to redirect structural outcome. More often, however, they act through hydration state, growth pathway or phase selection rather than through direct crystal-state conformational selection. The strongest examples appear when intrinsic preorganization and extrinsic triggers reinforce one another, which helps explain why many well-characterized cases cluster in aromatic and strongly hydrated systems. Simpler non-aromatic linear di- and tripeptides often provide fewer independent packing handles, so weak local conformational preferences are more easily overridden by backbone hydrogen bonding, packing degeneracy and chain entropy.3,6–8,23
Methods for establishing conformational control
Demonstrating conformational control requires experimental evidence that the observed conformation belongs to a well-defined crystal phase rather than to crystallisation history or phase transformation. In practice, this means distinguishing true crystal-state conformational selection from pathway-dependent crystal growth, phase conversion, or surface recrystallisation. Techniques that link molecular conformation directly to crystal packing and phase identity are therefore essential.24–27Single-crystal X-ray diffraction (SCXRD) and powder X-ray diffraction (PXRD) remain the primary structural tools because they provide the most direct view of which conformer is actually present in the crystal lattice.24 When external variables such as humidity influence the system, diffraction experiments performed under controlled environmental conditions become particularly important. For hydration-sensitive peptide crystals, the most informative studies combine variable-humidity diffraction with complementary measurements that track both structural and environmental changes. In practice, variable-humidity PXRD coupled with Raman spectroscopy and dynamic vapour sorption (DVS) provides a powerful experimental combination. Diffraction follows phase evolution, Raman spectroscopy tracks backbone conformation through shifts in the amide I and amide II vibrational bands, and vapour-sorption measurements quantify changes in hydration state. When these measurements are complemented by in situ observation of crystal morphology, artefacts arising from surface dissolution or partial recrystallisation can be reliably excluded.
In many peptide systems the crystals are too small for conventional X-ray diffraction. In such cases, microcrystal electron diffraction (MicroED) and related three-dimensional electron-diffraction methods have become particularly valuable. These techniques allow structural determination of peptide nanocrystals and short amyloid assemblies, in several cases even by ab initio phasing and at sub-Å resolution.25,26 They therefore extend crystallographic analysis to systems that would otherwise remain structurally inaccessible.
Complementary spectroscopic approaches can help resolve structural questions that diffraction alone cannot address. Solid-state nuclear magnetic resonance spectroscopy (solid-state NMR), particularly high-dimensional proton-detected experiments, provides direct insight into conformational heterogeneity and the possible coexistence of multiple backbone geometries within a single crystalline phase.27 Such measurements are especially valuable when subtle conformational differences do not produce clearly distinguishable diffraction patterns.
Computational methods increasingly complement experimental approaches to peptide crystal engineering.
Molecular dynamics (MD) simulations and density functional theory (DFT) calculations remain valuable tools for evaluating the stability of competing conformers and interaction motifs within crystal environments.28,29
More recently, machine-learning interatomic potentials have begun to accelerate exploration of conformational and lattice-energy landscapes in molecular crystals. Neural-network potentials such as atoms-in-molecules neural network (AIMNet2) and message passing atomic cluster expansion (MACE) enable rapid sampling of peptide conformational space and efficient ranking of candidate crystal packings.30,31
The growing availability of community datasets for molecular crystals, including the Open Molecular Crystals 2025 (OMC25) dataset, is further enabling the development and benchmarking of machine-learning models for crystal structure prediction and stability analysis.32
Together, these approaches are beginning to enable computational exploration of peptide conformational landscapes prior to experimental crystallisation.
Artificial-intelligence (AI) tools developed for biomolecular structure prediction can also provide guidance on sequence-dependent local conformational preferences. However, although methods such as AlphaFold3 are highly effective at predicting sequence-level structures, they do not model crystal packing or intermolecular interactions in the solid state and therefore cannot replace explicit crystallographic analysis.33
In practice, the most convincing demonstrations of conformational control combine diffraction under controlled environmental conditions with spectroscopic or nuclear magnetic resonance validation and targeted computational analysis. Only such integrated approaches allow conformational lability to be distinguished from growth history or phase transformation and thus make it possible to demonstrate reproducible conformational control in short-peptide crystal engineering.
From conformational control to function in short-peptide crystals
Recent tripeptide crystals now provide the clearest evidence that local structural preferences can propagate indirectly into function through packing asymmetry and symmetry breaking. Across these studies, residue placement repeatedly alters crystal symmetry and the resulting electromechanical response, making piezoelectric short-peptide crystals one of the clearest currently available illustrations linking local structural preference with functional response.9,10Tan and co-workers showed that extending glycylglycine (GG) to the tripeptides phenylalanylglycylglycine (FGG) and glycylglycylphenylalanine (GGF) converts a centrosymmetric crystal system into non-centrosymmetric ones, giving rise to measurable piezoelectric response (Fig. 2).9 Sequence-dependent symmetry modulation in short peptides provides a clear example of how local structural variation propagates into crystal packing and resulting functional behaviour.
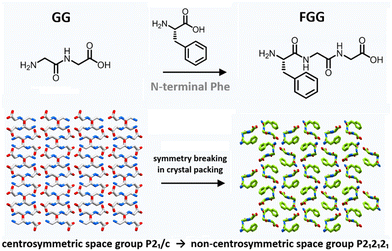 | ||
| Fig. 2 Sequence-induced symmetry modulation in short-peptide crystals. Incorporation of L-phenylalanine at the N-terminus converts centrosymmetric GG crystals (P21/c) into non-centrosymmetric FGG (P212121), illustrating symmetry breaking in crystal packing, adapted from ref. 9. | ||
In the broader context of organic and bio-derived piezoelectrics, these responses are comparable to those reported for established organic materials such as poly(vinylidene fluoride) (PVDF), which typically exhibits piezoelectric coefficients of ∼20–30 pC N−1. Similar magnitudes are observed in natural piezoelectric biomaterials including collagen and nanostructured cellulose, where reported responses generally fall in the ∼10–50 pC N−1 range depending on structural organisation.34,35
Peptide-based hydrogen-bonded organic frameworks and related peptide-derived crystalline materials provide a broader structural context for these comparisons.38
Yang and co-workers extended this picture by showing that sequence programming in a hydrophobic tripeptide series with central phenylalanine (Phe) can further increase packing asymmetry and electromechanical response. In that series, the central Phe residue gave the strongest response, consistent with the largest packing asymmetry identified from structural analysis; for the best-performing system, an effective piezoelectric coefficient measured by piezoresponse force microscopy (PFM) reached 24.0 pC N−1 and the open-circuit voltage reached 2.57 V at 50 N.10
Overall, the studies of Tan et al. (2025) and Yang et al. (2026) provide some of the clearest recent experimental examples showing that precise sequence modification in short peptides can convert centrosymmetric crystal packing into non-centrosymmetric structures and thereby enable measurable piezoelectric response.9,10
These observations provide one of the clearest experimental links so far between sequence-level structural programming and functional response in peptide crystals, highlighting short-peptide crystals as promising platforms for sequence-programmed molecular electromechanical materials.
More standardized comparison across crystal orientation, tensor assignment, sample history and measurement geometry should now help define these structure–property trends more rigorously.
These cases also broaden how variation among short-peptide forms can be viewed. In some systems, structural flexibility may open access to more than one functional state. Dong and co-workers reported a soft porous crystal based on helical self-assembly of a cysteine-derived dipeptide. This system reversibly switched between guest-filled and guest-free phases and showed pentane/hexane selectivity greater than 20 under ambient conditions.36
Structural flexibility can therefore contribute to phase transitions, sorption behaviour and mechanical actuation in peptide-based crystals.
Such systems remain relatively few in number and structurally less well defined than the best hydration- and symmetry-related examples.
The clearest functional examples still cluster around piezoelectricity and adaptive porosity, while adjacent peptide-based crystalline systems suggest a broader, but less securely resolved, landscape.37–41
A particularly clear recent illustration comes from peptide-based crystalline solids that reversibly switch between layered and hexagonally packed architectures in response to humidity while maintaining crystal integrity in the solid state, thereby coupling adaptive packing with tunable mechanical behaviour.42
Thus, these studies indicate that short-peptide crystals can link local conformational preference with crystallographic outcome and, in a growing number of systems, measurable functional behaviour.
Representative systems are summarized in Table 1, with emphasis on how local structural preference is introduced, how it propagates into crystallographic outcome and, in selected cases, how it supports measurable response. Several systems are included as adjacent boundary cases not because they provide direct evidence for crystal-state conformational control, but because they clarify where supramolecular redirection, guest response or assembly-level effects begin to overlap with the present problem.
| System | Main determinant/trigger | Local conformational/structural feature | Crystallographic outcome | Functional relevance | Level of evidence | Ref. |
|---|---|---|---|---|---|---|
| a For triglycine, the hydration-linked structural change is well supported, but the pathway remains under debate between crystal-state transformation and dissolution–recrystallisation. | ||||||
| FGG/GGF | Sequence change (Phe placement) | Sequence-dependent local structural preference linked to non-centrosymmetric packing | Symmetry breaking | Piezoelectric response | Strong | 9 |
| Hydrophobic tripeptide series with central Phe | Sequence programming/aromatic residue placement | Sequence-dependent local structural preference | Increased non-centrosymmetric packing asymmetry | Enhanced piezoelectric response | Strong | 10 |
| Triglycine (GGG) | Hydration/humidity | Hydrated pPII conformation versus sheet-like form | Hydration-linked structural change | Humidity-responsive structural changea | Moderatea | 6, 7 |
| N-capped diphenylalanine | Solvent composition/growth conditions | Local structural preference during crystal growth | Packing transition during crystal growth | Not established | Moderate | 8 |
| Diphenylalanine (FF) solvated forms | Solvent/crystallisation conditions | Solvent-dependent structural variation | Multiple crystal outcomes | Not established | Boundary | 17 |
| Constrained helical peptides | Intramolecular C–H⋯π contact; temperature | Conformational polymorphism | Distinct polymorphs | Not established | Boundary | 13 |
| Short peptide assemblies | pH | Assembly-level packing variation/chirality inversion | Assembly-state structural redirection | Not established at direct crystal-state conformer level | Boundary | 19 |
| Layered dipeptide crystals | CO2/guest stimulus | Guest-dependent structural reorganization | Symmetry breaking | Stimulus-responsive behaviour | Boundary | 39 |
| Cysteine-derived dipeptide | Helical self-assembly/guest exchange | Flexible crystal framework | Reversible guest-filled/guest-free phases | Selective pentane/hexane adsorption | Boundary | 36 |
Conclusions and outlook
Recent work has begun to change how conformational flexibility in short-peptide crystals is viewed. What was long considered mainly a crystallographic complication can, in certain systems, act as a structural variable that influences crystal packing and symmetry. A small but growing number of well-characterised examples shows that local conformational preferences may propagate into supramolecular organisation and, in favourable cases, measurable functional behaviour.At the same time, such behaviour appears only in specific classes of systems. The clearest cases involve short peptides that combine intrinsic conformational bias with external triggers such as hydration or aromatic interactions. In these systems, relatively small differences in backbone geometry can be amplified through packing interactions. Conformational control therefore appears to be conditional rather than universal.
A central challenge remains the reliable identification of genuine crystal-state conformational selection. Apparent structural changes may arise from crystallisation pathways, partial phase conversion or surface recrystallisation. Careful experimental design is therefore essential. Variable-humidity or variable-temperature diffraction experiments, combined with spectroscopic or sorption measurements, will likely play an important role in clarifying these mechanisms.
Short linear peptides provide a particularly useful experimental platform for such studies. Their sequences can be modified systematically while the resulting crystal structures remain relatively simple. This allows direct comparison of how small sequence changes influence molecular conformation, crystal packing and, in favourable cases, physical response.
In our view, further progress will depend on combining systematic peptide series with complementary experimental and computational approaches. Emerging diffraction methods capable of resolving very small crystals, together with solid-state spectroscopies and improved computational models, are beginning to expand what can be analysed in these systems.
If these approaches continue to develop, short-peptide crystals may become valuable model systems for understanding how sequence-encoded structural preferences influence crystal packing and emergent properties in molecular materials. In our view, establishing such relationships will be essential for determining when conformational flexibility can be used as a controllable structural element rather than remaining a source of crystallographic complexity. Improved integration of structural visualisation with crystallographic analysis will further enhance accessibility and interpretation of peptide crystal systems.
Author contributions
J. B. developed and expanded the conceptual framework of conformational control in short-peptide crystals, wrote the manuscript, led the revision in response to peer review, supervised the project, and served as the corresponding author. W. M. W. initiated the original concept related to conformational lability of short peptides and contributed to the revision of the manuscript. M. B. contributed to theoretical calculations and related analysis. S. J. contributed to the initial drafting of sections related to molecular docking and molecular dynamics simulations. All authors reviewed and approved the final version of the manuscript. This work was completed while S. J. was a Doctoral Candidate at the Interdisciplinary Doctoral School, Lodz University of Technology, Poland.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
AI tools were used solely for language polishing, phrasing suggestions and minor text restructuring. The authors have reviewed, edited and fully approve the final manuscript. The authors thank the editor for the invitation to contribute this Highlight Review.References
- A. J. Cruz-Cabeza and J. Bernstein, Conformational polymorphism, Chem. Rev., 2014, 114(4), 2170–2191, DOI:10.1021/cr400249d.
- V. Apostolopoulos, J. Bojarska, T.-T. Chai, S. Elnagdy, K. Kaczmarek, J. Matsoukas, R. New, K. Parang, O. Paredes Lopez, H. Parhiz, C. O. Perera, M. Pickholz, M. Remko, M. Saviano, M. Skwarczynski, Y. Tang, W. M. Wolf, T. Yoshiya, J. Zabrocki, P. Zielenkiewicz, M. AlKhazindar, V. Barriga, K. Kelaidonis, E. Mousavinezhad Sarasia and I. Toth, A global review on short peptides: frontiers and perspectives, Molecules, 2021, 26(2), 430, DOI:10.3390/molecules26020430.
- K. Kaygisiz, D. Sementa, V. Athiyarath, X. Chen and R. V. Ulijn, Context dependence in assembly code for supramolecular peptide materials and systems, Nat. Rev. Mater., 2025, 10(6), 449–472, DOI:10.1038/s41578-025-00782-6.
- S. Routray, M. K. Baroi, P. Das and D. Das, The journey of short peptides: from molecules to materials, Chem. Commun., 2026, 62(22), 6004–6028, 10.1039/D5CC05199A.
- K. Tao, H. Wu, L. Adler-Abramovich, J. Zhang, X. Fan, Y. Wang, Y. Zhang, S. A. M. Tofail, D. Mei, J. Li and E. Gazit, Aromatic short peptide architectonics: assembly and engineering, Prog. Mater. Sci., 2024, 142, 101240, DOI:10.1016/j.pmatsci.2024.101240.
- M. Guo, I. Rosbottom, L. Zhou, C. W. Yong, L. Zhou, Q. Yin, I. T. Todorov, E. Errington and J. Y. Y. Heng, Triglycine (GGG) adopts a polyproline II (pPII) conformation in its hydrated crystal form: revealing the role of water in peptide crystallization, J. Phys. Chem. Lett., 2021, 12(34), 8416–8422, DOI:10.1021/acs.jpclett.1c01622.
- Z. Zhu, J. Yang, H. Zhao, Y. Wu, T. Bai, H. Zhang, Y. Song, T. Ji, J. Hu and Y. Gao, Humidity-responsive self-assembly of short peptides with super-flexibility, CrystEngComm, 2024, 26, 1667–1670, 10.1039/D4CE00031E.
- Y. Dan, Z. A. Arnon, Y. Tang, M. Kravikass, Y. Zhou, R. Misra, O. S. Tiwari, L. J. W. Shimon, R. Beck, E. Gazit, G. Wei and L. Adler-Abramovich, Effect of solvent-induced packing transitions on N-capped diphenylalanine peptide crystal growth, Nat. Commun., 2025, 16, 6106, DOI:10.1038/s41467-025-61331-x.
- M.-L. Tan, Y. Luo, S. Liu, Y. Huo, J. Zhao, X. Xu, J. Song and W. Ji, Structural modulation in short peptide crystals from centrosymmetric to noncentrosymmetric achieving piezoelectricity, ACS Appl. Mater. Interfaces, 2025, 17, 59135–59147, DOI:10.1021/acsami.5c15632.
- X. Yang, S. Liu, H. Lu, Y. Wang, H. Zhang, W. Ji and H. Wang, Peptide sequence programmed piezoelectric response by supramolecular self-assembly, Adv. Sci., 2026, 13(12), e15237, DOI:10.1002/advs.202515237.
- K. Hu, W. Xiong, C. Sun, C. Wang, J. Li, F. Yin, Y. Jiang, M.-R. Zhang, Z. Li, X. Wang and Z. Li, Self-assembly of constrained cyclic peptides controlled by ring size, CCS Chem., 2020, 2(1), 42–51, DOI:10.31635/ccschem.020.201900047.
- J. Morimoto, M. Yokomine, Y. Shiratori, T. Ueda, T. Nakamuro, K. Takaba, S. Maki-Yonekura, K. Umezawa, K. Miyanishi, Y. Fukuda, T. Watanabe, M. Suga, A. Inayoshi, T. Yoshida, W. Mizukami, K. Takeuchi, K. Yonekura, E. Nakamura and S. Sando, Bottom-up design of peptide shapes in water using oligomers of N-methyl-L/D-alanine, Chem. Sci., 2025, 16(23), 10512–10522, 10.1039/D5SC01483B.
- J. Sun, Z.-Y. Tian, J. Liu, C. Wan, C. Dai, Z. Liu, Y. Xing, Y. Wu, Z. Hou, W. Han, F. Yin, Y. Ye and Z. Li, Intramolecular CH⋯π attraction mediated conformational polymorphism of constrained helical peptides, Chem. Sci., 2024, 15, 14264–14272, 10.1039/D4SC02545H.
- A. Nangia, Conformational polymorphism in organic crystals, Acc. Chem. Res., 2008, 41(5), 595–604, DOI:10.1021/ar700203k.
- J. Bojarska, M. Breza, P. Borowiecki, I. D. Madura, K. Kaczmarek, Z. M. Ziora and W. M. Wolf, An experimental and computational investigation of the cyclopentene-containing peptide-derived compounds: focus on pseudo-cyclic motifs via intramolecular interactions, R. Soc. Open Sci., 2024, 11, 240962, DOI:10.1098/rsos.240962.
- J. Bojarska, M. Breza, M. Remko, P. Borowiecki, A. Fruziński, I. D. Madura, K. Kaczmarek, Z. Leśnikowski, A. Kraj, P. Zielenkiewicz and W. M. Wolf, Supramolecular synthon hierarchy in cyclopropyl-containing peptide-derived compounds, CrystEngComm, 2022, 24, 8372–8389, 10.1039/D2CE01231F.
- Z. Chaker, P. Chervy, Y. Boulard, S. Bressanelli, P. Retailleau, M. Paternostre and T. Charpentier, Systematic method for the exploration, representation, and classification of the diphenylalanine solvatomorphic space, J. Phys. Chem. B, 2021, 125(33), 9454–9466, DOI:10.1021/acs.jpcb.1c04203.
- V. Verma, I. Bade, V. Karde and J. Y. Y. Heng, Experimental elucidation of templated crystallization and secondary processing of peptides, Pharmaceutics, 2023, 15(4), 1288, DOI:10.3390/pharmaceutics15041288.
- X. Ma, K. Qi, X. Ju, Y. Sun, H. Yang, Y. Ke, J. Zhang, Y. Zhao, H. Xu and J. Wang, pH-dependent packing mode variations and chirality inversion in short peptide self-assembly, Angew. Chem., Int. Ed., 2025, 64(37), e202511407, DOI:10.1002/anie.202511407.
- X. Ma, K. Qi, X. Ju, Y. Sun, H. Yang, Y. Ke, J. Zhang, Y. Zhao, H. Xu and J. Wang, Correction to ‘pH-dependent packing mode variations and chirality inversion in short peptide self-assembly’, Angew. Chem., Int. Ed., 2026, 65(11), e8533589, DOI:10.1002/anie.8533589.
- I. L. Karle and P. Balaram, Structural characteristics of alpha-helical peptide molecules containing Aib residues, Biochemistry, 1990, 29, 6747–6756, DOI:10.1021/bi00481a001.
- T. G. Castro, M. Melle-Franco, C. E. A. Sousa, A. Cavaco-Paulo and J. C. Marcos, Non-Canonical Amino Acids as Building Blocks for Peptidomimetics: Structure, Function, and Applications, Biomolecules, 2023, 13, 981, DOI:10.3390/biom13060981.
- K. Qi, H. Qi, M. Wang, X. Ma, Y. Wang, Q. Yao, W. Liu, Y. Zhao, J. Wang, Y. Wang, W. Qi, J. Zhang, J. R. Lu and H. Xu, Chiral inversion induced by aromatic interactions in short peptide assembly, Nat. Commun., 2024, 15, 6186, DOI:10.1038/s41467-024-50448-0.
- P. Sacchi, M. Lusi, A. J. Cruz-Cabeza, E. Nauha and J. Bernstein, Same or different - that is the question: identification of crystal forms from crystal structure data, CrystEngComm, 2020, 22(43), 7170–7185, 10.1039/D0CE00724B.
- L. S. Richards, M. D. Flores, C. Millán, C. Glynn, C.-T. Zee, M. R. Sawaya, M. Gallagher-Jones, R. J. Borges, I. Usón and J. A. Rodriguez, Fragment-based ab initio phasing of peptidic nanocrystals by MicroED, ACS Bio Med Chem Au, 2023, 3(2), 201–210, DOI:10.1021/acsbiomedchemau.2c00082.
- L. Samperisi, X. Zou and Z. Huang, Three-dimensional electron diffraction: a powerful structural characterization technique for crystal engineering, CrystEngComm, 2022, 24, 2719–2728, 10.1039/D2CE00051B.
- E. Burakova, S. K. Vasa and R. Linser, Characterization of conformational heterogeneity via higher-dimensionality, proton-detected solid-state NMR, J. Biomol. NMR, 2022, 76, 197–212, DOI:10.1007/s10858-022-00405-0.
- A. N. Hosseini and D. van der Spoel, Simulations of amyloid-forming peptides in the crystal state, Protein J., 2023, 42, 192–204, DOI:10.1007/s10930-023-10119-3.
- T. K. Piskorz, L. Perez-Chirinos, B. Qiao and I. R. Sasselli, Tips and tricks in the modeling of supramolecular peptide assemblies, ACS Omega, 2024, 9(29), 31254–31273, DOI:10.1021/acsomega.4c02628.
- K. S. Nayal, D. Orellana, G. He, D. S. Levine, Y. Yang, L. Barroso-Luque, T. J. Martinez, M. Ceriotti, C. Greenwell and G. M. Day, Efficient Molecular Crystal Structure Prediction and Stability Assessment with AIMNet2 Neural Network Potentials, Cryst. Growth Des., 2025, 25(21), 9092–9106, DOI:10.1021/acs.cgd.5c01001.
- F. Della Pia, B. X. Shi, V. Kapil, A. Zen, D. Alfè and A. Michaelides, Accurate and efficient machine learning interatomic potentials for finite temperature modelling of molecular crystals, Chem. Sci., 2025, 16, 11419–11433, 10.1039/D5SC01325A.
- V. Gharakhanyan, L. Barroso-Luque, Y. Yang, M. Shuaibi, K. Michel, D. S. Levine, M. Dzamba, X. Fu, M. Gao, X. Liu and H. Ni, et al., Open Molecular Crystals 2025 (OMC25) dataset and models, Sci. Data, 2026, 13, 354, DOI:10.1038/s41597-026-06628-2.
- J. Abramson, J. Adler and J. Dunger, et al., Accurate structure prediction of biomolecular interactions with AlphaFold 3, Nature, 2024, 630, 493–500, DOI:10.1038/s41586-024-07487-w.
- N. Ahbab, S. Naz, T.-B. Xu and S. Zhang, A comprehensive review of piezoelectric PVDF polymer fabrications and characteristics, Micromachines, 2025, 16(4), 386, DOI:10.3390/mi16040386.
- R. Wang, C. R. Bowen and Y. Yang, Natural piezoelectric biomaterials: a biocompatible and sustainable building block for biomedical devices, ACS Nano, 2022, 16(11), 16841–16857, DOI:10.1021/acsnano.2c08164.
- Y. Dong, H. Lu, Z. Fang, C. Zhang and H. Cao, Soft porous crystal based on helical self-assembly of dipeptides for selective adsorption, Mater. Today Chem., 2025, 45, 102667, DOI:10.1016/j.mtchem.2025.102667.
- D. F. Brightwell, G. Truccolo, K. Samanta, E. J. Fenn, S. J. Holder, H. J. Shepherd, C. S. Hawes and A. Palma, A reversibly porous supramolecular peptide framework, Chem. – Eur. J., 2022, 28(66), e202202368, DOI:10.1002/chem.202202368.
- T. Vijayakanth, S. Dasgupta, P. Ganatra, S. Rencus-Lazar, A. V. Desai, S. Nandi, R. Jain, S. Bera, A. I. Nguyen, E. Gazit and R. Misra, Peptide hydrogen-bonded organic frameworks, Chem. Soc. Rev., 2024, 53, 3640–3655, 10.1039/D3CS00648D.
- X. Li, Q. Li, A. Wu and J. Li, CO2 induces symmetry breaking in layered dipeptide crystals, Angew. Chem., Int. Ed., 2023, 62(1), e202214184, DOI:10.1002/anie.202214184.
- H. Yuan, Y. Tang, E. Naranjo, P. A. Cazade, Y. Yao, V. Thangavel, R. Yang, S. Rencus-Lazar, D. Thompson, L. J. W. Shimon, G. Wei, X. Chen and E. Gazit, Discrete actuation of water-responsive crystalline metal-peptide frameworks, Angew. Chem., Int. Ed., 2025, 64(43), e202513629, DOI:10.1002/anie.202513629.
- Y. Zhang, T. Pan, S. Guerin, J. Zhang, Y. Tang, Z. Qiu, Q. Li, J. Zhang, F. Wei, J. Wang, R. Yang, D. Mei, D. Thompson, G. Wei, H. Xu and K. Tao, Underlying polymorphism: superhelical crystallization induces architectural and functional diversity, Small, 2026, 22(6), e10542, DOI:10.1002/smll.202510542.
- V. Athiyarath, C. J. Murphy, R. V. Ulijn and X. Chen, et al., Water-Mediated Reconfigurable Topology and Mechanics in Porous Peptide Materials, Matter, 2026, 102669, DOI:10.1016/j.matt.2026.102669 , available online 11 March 2026.
| This journal is © The Royal Society of Chemistry 2026 |