Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Structure-directing roles of polycarboxylate anions in the formation of Ag(I) coordination polymers with mixed ligands
Yi-Lin Shen,
Chia-Li Liu,
Hsun-Chih Tai,
Zhi-Ling Chen and
Jhy-Der Chen *
*
Department of Chemistry, Chung Yuan Christian University, Chung-Li, Taiwan, R.O.C. E-mail: jdchen@cycu.edu.tw
First published on 28th January 2026
Abstract
Hydrothermal reactions of bis-pyridyl-bis-amide with polycarboxylic acid and Ag(I) salt afforded {[Ag(L1)][HCDDC]}n [L1 = N,N′-di(4-methylpyridyl)oxamide; H2CDDC = 7-carboxy-1,3-dioxo-1,3-dihydrobenzo[de]isochromene-6-carboxylic acid], 1, {[Ag(L1)][1,4-HNDC]·2H2O}n (1,4-H2NDC = 1,4-naphthalenedicarboxylic acid), 2, [Ag(1,2,4,5-H2BTEC)0.5(L1)0.5]n (1,2,4,5-H4BTC = 1,2,4,5-benzenetetracarboxylic acid), 3, {[Ag(L1)(H2O)][1,2,4,5-H3CTC]·H2O}n (1,2,4,5-H4CTC = 1,2,4,5-cyclohexanetetracarboxylic acid), 4, {[Ag(L2)][1,4-HNDC]·H2O}n [L2 = N,N′-di(3-pyridyl)oxamide; 1,4-H2NDC = 1,4-naphthalenedicarboxylic acid], 5, {[Ag2(L1)2][1,2,4-HBTC]·2H2O}n (1,2,4-H3BTC = 1,2,4-benzenetricarboxylic acid), 6, {[Ag2(L1)1.5][(1,3,5-HBTC)(H2O)]·2H2O}n (1,3,5-H3BTC = 1,3,5-benzenetricarboxylic acid), 7, {[Ag(L1)0.5][(1,3,5-H2BTC)]}n, 8, and {[Ag5(1,3,5-HBTC)(1,3,5-BTC)(L1)2]·5H2O}n, 9, which have been structurally identified by using single-crystal X-ray diffraction. Complexes 1, 2, 4, 5 and 6 are 1D zigzag chains and 8 forms a 1D looped chain, whereas 3 features a 3D framework with the fsh topology, 7 reveals a linear tetranuclear molecule and 9 adopts a 3-fold interpenetrated 3D framework with the sra topology. The roles of the polycarboxylate anions in determining the structural diversity of these Ag(I) complexes are evaluated.
Introduction
The rational design and synthesis of novel coordination polymers (CPs) have become an active area of investigation because such that the range of these crystalline functional materials with pre-selected physical and chemical properties can be further expanded.1–11 The varieties of the CPs that can be approached rely on the presence of suitable metal–ligand interactions and supramolecular contacts, i.e., hydrogen bonds and other weak interactions. The properties of CPs are mostly dominated by their structures and compositions and it is well known that the structural type of CPs is determined by several factors such as the identities of metal ions, counter ions and organic linkers.12–15 Therefore, controlling the appropriate factors to construct suitable CPs have become an exciting topic in the crystal engineering of CPs. On the other hand, Ag(I) CPs have attracted great attention due to their structural variability and extensive industrial and medical applications.16–24While many Ag(I) CPs with mixed ligands involving dipyridyl and polycarboxylate ligands have been reported, the coordination of the polycarboxylate ligands to Ag(I) ions remain unjustified, presumably due to the hard base character of the polycarboxylate ligands and the soft acid character of the Ag(I) ion. Short Ag⋯Ag and Ag⋯O distances were observed in the structurally characterized complexes with bridging ligands (ligand-supported) and the attractive phenomenon of Ag⋯Ag interactions was generally accepted after examples of ligand-unsupported ones were reported.25 The Ag⋯Ag distance is generally longer than that of the metallic Ag⋯Ag contact (2.89 Å), but shorter than the van der Waals contact distance (3.40 A°), whereas the Ag⋯O distances are longer than the upper limit of 2.3–2.6 Å for Ag–O bonds,26 but are significantly shorter than the corresponding van der Waals contacts of Ag and O atoms which is 3.24 Å (van der Waals radii for Ag = 1.72 and O = 1.52 Å).27 The Ag⋯O distances to the polycarboxylate anions are thus important in determining the structural topologies of the Ag(I) CPs thus prepared and should be carefully evaluated.
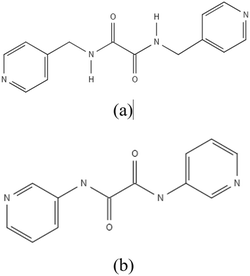
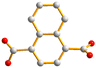
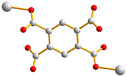
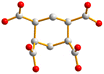
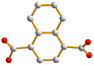
With this background information, we sought to investigate the roles of the polycarboxylate ligands in determining the structural diversity of the bis-pyridyl-bis-amide (bpba)-based Ag(I) CPs. Herein, we report the syntheses and structures of {[Ag(L1)][HCDDC]}n [L1 = N,N′-di(4-methylpyridyl)oxamide; H2CDDC = 7-carboxy-1,3-dioxo-1,3-dihydrobenzo[de]isochromene-6-carboxylic acid], 1, {[Ag(L1)][1,4-HNDC]·2H2O}n (1,4-H2NDC = 1,4-naphthalenedicarboxylic acid), 2, [Ag(1,2,4,5-H2BTEC)0.5(L1)0.5]n (1,2,4,5-H4BTC = 1,2,4,5-benzenetetracarboxylic acid), 3, {[Ag(L1)(H2O)][1,2,4,5-H3CTC]·H2O}n (1,2,4,5-H4CTC = 1,2,4,5-cyclohexanetetracarboxylic acid), 4, {[Ag(L2)][1,4-HNDC]·H2O}n [L2 = N,N′-di(3-pyridyl)oxamide; 1,4-H2NDC = 1,4-naphthalenedicarboxylic acid], 5, {[Ag2(L1)2][1,2,4-HBTC]·2H2O}n (1,2,4-H3BTC = 1,2,4-benzenetricarboxylic acid), 6, {[Ag2(L1)1.5][(1,3,5-HBTC)(H2O)]·2H2O}n (1,3,5-H3BTC = 1,3,5-benzenetricarboxylic acid), 7, {[Ag(L1)0.5][(1,3,5-H2BTC)]}n, 8, and {[Ag5(1,3,5-HBTC)(1,3,5-BTC)(L1)2]·5H2O}n, 9. The roles of the Ag⋯Ag and Ag⋯O interactions in determining the topological structures of these CPs are discussed. Fig. 1 depicts the structures of L1 and L2 and Fig. 2 shows the structures of the polycarboxylic acids, respectively.
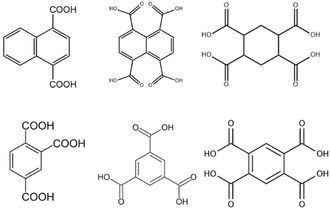 | ||
| Fig. 2 From left to right, first row: structures of 1,4-H2NDC, 1,4,5,8-H4NTC and 1,2,4,5-H4CTC. Second row: 1,2,4-H3BTC, 1,3,5-H3BTC and 1,2,4,5-H4BTC. | ||
Experimental details
General procedures
IR spectra (KBr disk) were obtained from a JASCO FT/IR-4200 FT-IR spectrometer. Elemental analyses were performed on an Elementar vario EL III analyzer. Powder X-ray diffraction was carried out on a Bruker D2 PHASER diffractometer with a CuKα (λα = 1.54 Å) radiation.Materials
The reagents 1,4-H2NDC, 1,2,4-H3BTC, and 1,3,5-H3BTC were purchased from Alfa Aesar and 1,2,4,5-H4CTC and 1,4,5,8-H4NTC from NOVA. Silver acetate was obtained from ACROS. The ligands N,N′-di(4-methylpyridyl)oxamide (L1) and N,N′-di(3-pyridyl)oxamide (L2) were prepared according to published procedures.28X-ray crystallography
The diffraction data for complexes 1–9 were collected on a Bruker AXS SMART APEX II CCD diffractometer, which was equipped with a graphite-monochromated Mo Kα (λα = 0.71073 Å) radiation. Data reduction was carried out by standard methods with the use of well-established computational procedures.29 The structure factors were obtained after Lorentz and polarization corrections. An empirical absorption correction based on “multi-scan” was applied to the data. The positions of some of the heavier atoms were located by the direct or Patterson method. The remaining atoms were found in a series of alternating difference Fourier maps and least-squares refinements, while the hydrogen atoms except those of the water molecules were added by using the HADD command in SHELXTL 6.1012.30 Table 1 lists the crystal data for 1–9. ORTEP drawings for the crystal structures are given in the SI (Fig. S1–S9).| CP/complex | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 |
|---|---|---|---|---|---|---|---|---|---|
| a R1 = ∑‖Fo∣ – ∣Fc‖/∑∣Fo∣.b wR2 = [∑w(Fo2 – Fc2)2/∑w(Fo2)2]1/2. w = 1/[σ2(Fo2) + (ap)2 + (bp)], p = [max (Fo2 or 0) + 2(Fc2)]/3. a = 0.0391, b = 3.1802 for 1; a = 0.0478, b = 0.39 for 2; a = 0.0193, b = 7.5031 for 3; a = 0.0407, b = 1.4676 for 4; a = 0.0256, b = 1.0625 for 5; a = 0.0538, b = 0 for 6; a = 0.0376, b = 8.4310 for 7; a = 0.0353, b = 0.5672 for 8; a = 0.0428, b = 4.4862 for 9.c Quality-of-fit = [∑w(∣Fo2∣ – ∣Fc2∣)2/(Nobserved – Nparameters)]1/2. | |||||||||
| Formula | C28H19AgN4O9 | C26H25AgN4O8 | C12H9AgN2O5 | C24H29AgN4O12 | C24H19AgN4O7 | C37H36Ag2N8O12 | C30H31Ag2N6O12 | C16H12AgN2O7 | C46H45Ag5N8O21 |
| Formula weight | 663.34 | 629.37 | 369.08 | 673.38 | 583.30 | 1000.48 | 883.35 | 452.15 | 1585.25 |
| Temperature, K | 296(2) | 293(2) | 293(2) | 293(2) | 296(2) | 293(2) | 293(2) | 293(2) | 293(2) |
| Crystal system | Triclinic | Triclinic | Monoclinic | Monoclinic | Monoclinic | Triclinic | Triclinic | Triclinic | Triclinic |
| Space group | P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) |
P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) |
P21/n | P21/c | P21/n | P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) |
P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) |
P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) |
P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) |
| a, Å | 9.4392(8) | 9.096(2) | 12.779(2) | 11.277(2) | 8.5629(16) | 9.4624(19) | 10.8579(15) | 8.6861(19) | 9.627(6) |
| b, Å | 10.4331(9) | 9.557(2) | 7.1294(14) | 11.398(2) | 25.199(5) | 11.7312(19) | 12.1551(15) | 8.914(2) | 10.094(7) |
| c, Å | 14.5148(13) | 15.651(4) | 13.218(2) | 21.089(4) | 10.864(2) | 19.160(3) | 13.0789(16) | 11.391(3) | 31.171(18) |
| α, ° | 76.969(2) | 76.424(8) | 90 | 90 | 90 | 95.359(6) | 78.820(4) | 84.881(7) | 85.529(16) |
| β, ° | 85.423(2) | 80.583(8) | 93.331(4) | 91.707(5) | 93.170(6) | 95.325(7) | 85.334(5) | 70.317(5) | 85.355(17) |
| γ, ° | 66.140(2) | 84.483(8) | 90 | 90 | 90 | 106.993 | 78.078(5) | 70.793(5) | 63.937(16) |
| V, Å3 | 1273.48(19) | 1302.3(6) | 1202.2(4) | 2709.5(9) | 2340.6(8) | 2008.9(6) | 1655.3(4) | 784.0(3) | 2709(3) |
| Z | 2 | 2 | 4 | 4 | 4 | 2 | 2 | 2 | 2 |
| Dcalc, mg m−3 | 1.730 | 1.605 | 2.039 | 1.651 | 1.655 | 1.654 | 1.772 | 1.915 | 1.943 |
| F(000) | 668 | 640 | 728 | 1376 | 1176 | 1008 | 886 | 450 | 1560 |
| μ(Mo Kα), mm−1 | 0.858 | 0.831 | 1.699 | 0.815 | 0.915 | 1.047 | 1.256 | 1.331 | 1.860 |
| Range (2θ) for data collection, ° | 2.88–51.98 | 2.70–52.00 | 4.31–56.77 | 3.61–56.88 | 3.23–52.00 | 2.15–52.00 | 3.17–52.00 | 3.79–56.74 | 2.62–56.45 |
| Independent reflections | 5009 [R(int) = 0.0790] | 5111 [R(int) = 0.0837] | 2981 [R(int) = 0.0383] | 6787 [R(int) = 0.0272] | 4616 [R(int) = 0.0805] | 7913 [R(int) = 0.1393] | 6483 [R(int) = 0.0708] | 3905 [R(int) = 0.0455] | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 236 [R(int) = 0.0249] 236 [R(int) = 0.0249] |
| Data/restraints/parameters | 5009/0/386 | 5111/0/352 | 2981/0/182 | 6787/0/370 | 4616/0/325 | 7913/0/542 | 6483/0/451 | 3905/0/236 | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 236/0/752 236/0/752 |
| Quality-of-fit indicatorc | 1.010 | 1.052 | 1.163 | 1.048 | 1.020 | 1.003 | 1.053 | 1.071 | 1.072 |
| Final R indices [I > 2σ(I)]a,b | R1 = 0.0592, wR2 = 0.1095 | R1 = 0.0440, wR2 = 0.1037 | R1 = 0.0415, wR2 = 0.1077 | R1 = 0.0306, wR2 = 0.0819 | R1 = 0.0367, wR2 = 0.0618 | R1 = 0.0596, wR2 = 0.1241 | R1 = 0.0591, wR2 = 0.1201 | R1 = 0.0305, wR2 = 0.0767 | R1 = 0.0364, wR2 =0.0949 |
| R indices (all data) | R1 = 0.1253, wR2 = 0.1325 | R1 = 0.0669, wR2 = 0.1150 | R1 = 0.0441, wR2 = 0.1087 | R1 = 0.0353, wR2 = 0.0849 | R1 = 0.0701, wR2 = 0.0699 | R1 = 0.1473, wR2 = 0.1557 | R1 = 0.0974, wR2 = 0.1446 | R1 = 0.0344, wR2 = 0.0792 | R1 = 0.0432, wR2 = 0.0984 |
Results and discussion
Structure of {[Ag(L1)][HCCDC]}n, 1
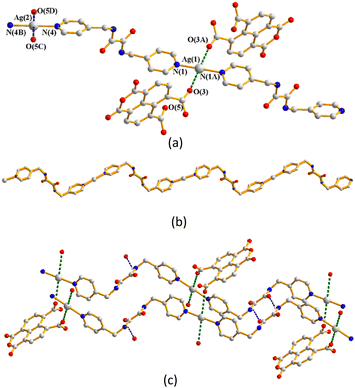
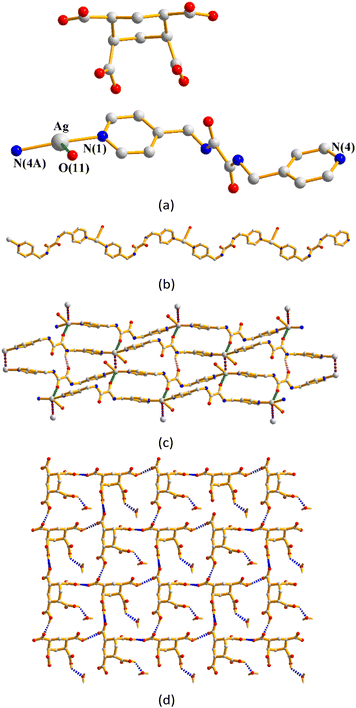
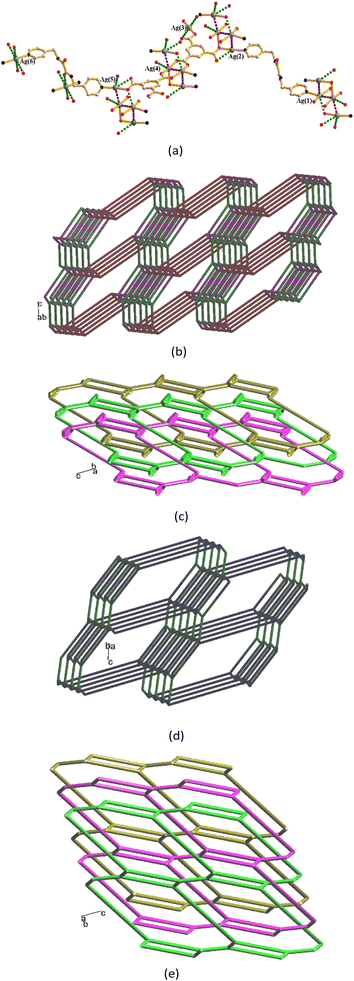
Single-crystal X-ray diffraction analysis shows that complex 1 crystallizes in the triclinic space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) . The asymmetric unit consists of two halves of a Ag(I) cation, one L1 ligand and one HCDDC− anion (H2CDDC = 7-carboxy-1,3-dioxo-1,3-dihydrobenzo[de]isochromene-6-carboxylate). The two independent Ag(I) cations are two-coordinated by two pyridyl nitrogen atoms from two L1 ligands [Ag–N = 2.149(4) and 2.114(5) Å], resulting in linear geometries (∠N–Ag–N = 180°) (Fig. 3(a)). The Ag(I) cations are linked together by L1 ligands to afford 1D zigzag chains (Fig. 3(b)), and the chains are further interlinked through Ag⋯O interactions [2.895(3) and 3.216(4) Å] to the HCDDC− anions to form pairs of double chains (Fig. 3(c)). The double chains are also supported by the self-complementary N–H⋯O hydrogen bonds [O⋯H = 2.151(4) Å; ∠N–H⋯O = 137.8(3)°] to the amide oxygen atoms of the adjacent chains. Transformation of the organic compound from 1,4,5,8-H4NTC to HCDDC− is observed.
. The asymmetric unit consists of two halves of a Ag(I) cation, one L1 ligand and one HCDDC− anion (H2CDDC = 7-carboxy-1,3-dioxo-1,3-dihydrobenzo[de]isochromene-6-carboxylate). The two independent Ag(I) cations are two-coordinated by two pyridyl nitrogen atoms from two L1 ligands [Ag–N = 2.149(4) and 2.114(5) Å], resulting in linear geometries (∠N–Ag–N = 180°) (Fig. 3(a)). The Ag(I) cations are linked together by L1 ligands to afford 1D zigzag chains (Fig. 3(b)), and the chains are further interlinked through Ag⋯O interactions [2.895(3) and 3.216(4) Å] to the HCDDC− anions to form pairs of double chains (Fig. 3(c)). The double chains are also supported by the self-complementary N–H⋯O hydrogen bonds [O⋯H = 2.151(4) Å; ∠N–H⋯O = 137.8(3)°] to the amide oxygen atoms of the adjacent chains. Transformation of the organic compound from 1,4,5,8-H4NTC to HCDDC− is observed.
Structure of {[Ag(L1)][1,4-HNDC]·2H2O}n, 2
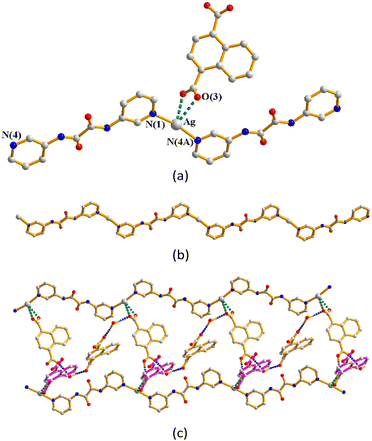
Complex 2 crystallizes in the triclinic space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) and each asymmetric unit consists of one Ag(I) cation, one L1 ligand, one 1,4-HNDC− anion and two co-crystallized water molecules. The Ag(I) cations are two-coordinated by two pyridyl nitrogen atoms from two L1 ligands [Ag–N = 2.176(3) and 2.182(3) Å], resulting in a distorted linear geometry [∠N–Ag–N = 163.94(12)°] (Fig. 4(a)). The Ag(I) cations are linked together by L1 ligands to afford a 1D zigzag chain (Fig. 4(b)), which is further interlinked by an adjacent chain through the Ag⋯Ag [3.4832(8) Å] and Ag⋯O interactions [2.616(3) and 2.662(4) Å] to the 1,4-HNDC− anions to form a double chain (Fig. 4(c)). The double chains are further extended through the O–H⋯O [O⋯H = 1.651(3) Å; ∠O–H⋯O = 164.0(2)°] hydrogen bonds between two adjacent 1,4-HNDC− anions to form a 2D layer. The 2D layers are supported by the O–H⋯O hydrogen bonds from the water molecules to the carboxylate oxygen atoms [O⋯H = 2.132(3) Å; ∠O–H⋯O = 170.3(2)°], the amide oxygen atoms [O⋯H = 1.972(3) Å; ∠O–H⋯O = 169.5(2)°] and water oxygen atoms [O⋯H = 1.984(3) and 2.051(3) Å; ∠O–H⋯ O = 170.0(2) and 151.8(2)°].
and each asymmetric unit consists of one Ag(I) cation, one L1 ligand, one 1,4-HNDC− anion and two co-crystallized water molecules. The Ag(I) cations are two-coordinated by two pyridyl nitrogen atoms from two L1 ligands [Ag–N = 2.176(3) and 2.182(3) Å], resulting in a distorted linear geometry [∠N–Ag–N = 163.94(12)°] (Fig. 4(a)). The Ag(I) cations are linked together by L1 ligands to afford a 1D zigzag chain (Fig. 4(b)), which is further interlinked by an adjacent chain through the Ag⋯Ag [3.4832(8) Å] and Ag⋯O interactions [2.616(3) and 2.662(4) Å] to the 1,4-HNDC− anions to form a double chain (Fig. 4(c)). The double chains are further extended through the O–H⋯O [O⋯H = 1.651(3) Å; ∠O–H⋯O = 164.0(2)°] hydrogen bonds between two adjacent 1,4-HNDC− anions to form a 2D layer. The 2D layers are supported by the O–H⋯O hydrogen bonds from the water molecules to the carboxylate oxygen atoms [O⋯H = 2.132(3) Å; ∠O–H⋯O = 170.3(2)°], the amide oxygen atoms [O⋯H = 1.972(3) Å; ∠O–H⋯O = 169.5(2)°] and water oxygen atoms [O⋯H = 1.984(3) and 2.051(3) Å; ∠O–H⋯ O = 170.0(2) and 151.8(2)°].
Structure of [Ag(1,2,4,5-H2BTEC)0.5(L1)0.5]n, 3
The structure of complex 3 was solved in the monoclinic space group P21/n. The asymmetric unit consists of one Ag(I) cation, a half of an L1 ligand, and a half of a 1,2,4,5-H2BTC2− ligand. Each of the Ag(I) cations is two-coordinated by one pyridyl nitrogen atom from the L1 ligands [Ag–N = 2.178(4) Å] and one oxygen atom from the 1,2,4,5-H2BTC2− ligand [Ag–O = 2.218(3) Å], resulting in a distorted linear geometry [∠N–Ag–O = 164.46(14)°] (Fig. 5(a)). The Ag(I) cations are also coordinated by two amide oxygen atoms from two L1 ligands [Ag⋯O = 2.536(3) and 2.597(3) Å]. If the two oxygen atoms are considered to be involved in the coordination environment, the Ag(I) cation adopts a drastically distorted tetrahedral geometry. Moreover, adjacent Ag(I) ions are linked by Ag⋯Ag [3.4763(7) Å] to form a dinuclear unit (Fig. 5(b)). Considering the two Ag⋯O interactions with distances of 2.536(3) and 2.597(3) Å as coordinated covalent bonds, and regarding the L1 ligand as 6-connected nodes and Ag(I) ions as 4-connected nodes, with the 1,2,4,5-H2BTC2− ligand as linkers, the structure can be simplified as a 4,6-connected 3D net with the (43·63)2(46·66·83)-fsh topology (standard representation) (Fig. 5(c)). Moreover, if the L1 ligands are considered as 4-connected nodes, and the dinuclear Ag(I) units as 6-connected nodes, with the 1,2,4,5-H2BTC2− ligand as linkers, the structure can be further simplified as a 4,6-connected 3D net with the (44·610·8)(44·62)-fsc topology (cluster representation) (Fig. 5(d)), determined by using ToposPro.31Structure of {[Ag(L1)(H2O)][1,2,4,5-H3CTC]·H2O}n, 4
Crystals of complex 4 conform to the monoclinic space group P21/n and the asymmetric unit comprises one Ag(I) cation, one L1 ligand, one 1,2,4,5-H3CTC− ligand and two water molecules. One of the two water molecules interacts significantly with the Ag(I) cation [Ag⋯O = 2.584(2) Å]. The Ag(I) cation is two-coordinated by two pyridyl nitrogen atoms from two L1 ligands [Ag–N = 2.1395(16) and 2.1519(16) Å], resulting in a distorted linear geometry (Fig. 6(a)). If the Ag⋯O distance of 2.584(2) Å to the oxygen atom of the water molecule is regarded as a coordinate covalent bond, the Ag(I) cation may adopt a distorted triangular geometry. The Ag(I) cations are linked by the L1 ligands to form 1D zigzag chains (Fig. 6(b)). The 1D chains are supported by the Ag⋯Ag interactions [3.2002(5) Å] and N–H⋯O [O⋯H = 2.033(2); ∠N–H⋯O = 151.3(1)°] hydrogen bonds to form double chains (Fig. 6(c)), which are also supported by the Ag⋯O interactions [Ag⋯O = 2.687(2) Å] to the amide oxygen atoms of the other pair (Fig. 6(d)).Structure of {[Ag(L2)][1,4-HNDC]·H2O}n, 5
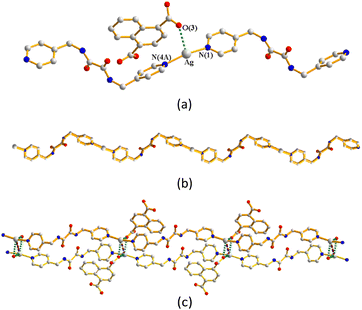
Complex 5 crystallizes in the monoclinic space group P21/n. The asymmetric unit consists of one Ag(I) cation, one L2 ligand, one 1,4-HNDC− ligand and one cocrystallized water molecule. The Ag(I) cation is two-coordinated by two pyridyl nitrogen atoms from two L2 ligands [Ag–N = 2.182(3) and 2.189(3) Å], resulting in a distorted linear geometry, while the anionic 1,4-HNDC− ligand interacts with the Ag(I) cation through the Ag⋯O interaction [Ag–O = 2.635(2) and 2.754(2) Å] (Fig. 7(a)). The Ag(I) cations are linked by the L2 ligands to afford 1D zigzag chains (Fig. 7(b)). Moreover, the 1D zigzag chains are supported by extensive O–H⋯O hydrogen bonds, originating from the carboxylate hydrogen atoms to the carboxylate oxygen atoms [O⋯H = 1.695(2) Å; ∠O–H⋯O = 169.1(2)°] and from the water hydrogen atoms to the carboxylate oxygen atoms [O⋯H = 1.937(3) and 2.034(2) Å; ∠O–H⋯O = 171.8(2) and 169.1(2)°] (Fig. 7(c)).Structure of {[Ag2(L1)2][1,2,4-HBTC]·2H2O}n, 6
Crystals of complex 6 conform to the triclinic space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) . The asymmetric unit consists of two Ag(I) cations, two L1 ligands, one 1,2,4-HBTC2− ligand and two cocrystallized water molecules. The Ag(I) metal centers are two-coordinated by one nitrogen atom from two L1 ligands [Ag–N = 2.146(5) Å], resulting in a distorted linear geometry (Fig. 8(a)). The Ag(I) cations are linked by the L1 ligands to afford 1D zigzag chains (Fig. 8(b)). Moreover, the 1D zigzag chains are supported by the Ag⋯Ag interactions [Ag⋯Ag = 3.1054(10) and 3.5257(10) Å] and Ag⋯O interactions to the amide oxygen atoms [Ag⋯O = 2.862(4) and 3.096(4) Å], carboxylate oxygen atoms [Ag⋯O = 2.862(4) and 2.968(5) Å] and to the water oxygen atoms [Ag⋯O = 2.930(7), 2.884(5) and 3.049(5) Å]. Self-complementary N–H⋯O hydrogen bonds [O⋯H = 2.138(4) and 1.992(6) Å; ∠N–H⋯O = 149.9(4) and 147.8(4)°], N–H⋯O hydrogen bonds to the carboxylate oxygen atoms [O⋯H = 2.150(6) and 1.961(6) Å; ∠N–H⋯O = 162.1(4) and 146.5(4)°] and O–H⋯O hydrogen bonds [O⋯H = 2.096(6) Å; ∠O–H⋯O = 136.0(4)°] from the water molecules to the carboxylate oxygen atoms are also observed (Fig. 8(c)).
. The asymmetric unit consists of two Ag(I) cations, two L1 ligands, one 1,2,4-HBTC2− ligand and two cocrystallized water molecules. The Ag(I) metal centers are two-coordinated by one nitrogen atom from two L1 ligands [Ag–N = 2.146(5) Å], resulting in a distorted linear geometry (Fig. 8(a)). The Ag(I) cations are linked by the L1 ligands to afford 1D zigzag chains (Fig. 8(b)). Moreover, the 1D zigzag chains are supported by the Ag⋯Ag interactions [Ag⋯Ag = 3.1054(10) and 3.5257(10) Å] and Ag⋯O interactions to the amide oxygen atoms [Ag⋯O = 2.862(4) and 3.096(4) Å], carboxylate oxygen atoms [Ag⋯O = 2.862(4) and 2.968(5) Å] and to the water oxygen atoms [Ag⋯O = 2.930(7), 2.884(5) and 3.049(5) Å]. Self-complementary N–H⋯O hydrogen bonds [O⋯H = 2.138(4) and 1.992(6) Å; ∠N–H⋯O = 149.9(4) and 147.8(4)°], N–H⋯O hydrogen bonds to the carboxylate oxygen atoms [O⋯H = 2.150(6) and 1.961(6) Å; ∠N–H⋯O = 162.1(4) and 146.5(4)°] and O–H⋯O hydrogen bonds [O⋯H = 2.096(6) Å; ∠O–H⋯O = 136.0(4)°] from the water molecules to the carboxylate oxygen atoms are also observed (Fig. 8(c)).
Structure of {[Ag2(L1)1.5][(1,3,5-HBTC)(H2O)]·2H2O}n, 7
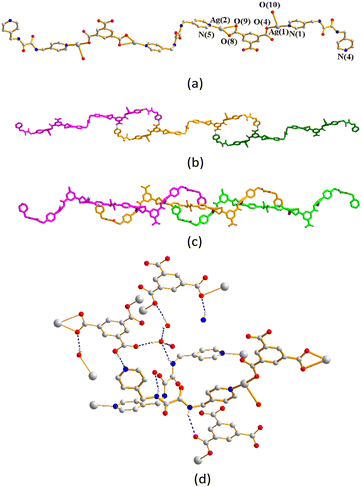
The structure of complex 7 was solved in the triclinic space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) and each asymmetric unit comprises two Ag(I) cations, one and a half L1 ligands, one 1,3,5-HBTC2− ligand, one coordinated water molecule and two co-crystallized water molecule. The Ag(1) atom is linked by two oxygen atoms from the 1,3,5-HBTC2− ligand [Ag–O = 2.176(4) Å] and water molecule [Ag–O = 2.534(5) Å] and one pyridyl nitrogen atom from the L1 ligand [Ag–N =2.175(6) Å]. Considering the Ag–O distance of 2.534(5) Å as a coordinate covalent bond, the Ag(1) atom adopts a distorted T-shaped geometry, whereas the Ag(2) metal center is three-coordinated by two oxygen atoms from the 1,3,5-HBTC2− ligand [Ag–O = 2.326(6) and 2.550(6) Å] and one pyridyl nitrogen atom from the L1 ligand [Ag–N =2.191(6) Å], resulting in a distorted triangular geometry. The Ag(I) ions are linked together by the 1,3,5-HBTC2− and L1 ligands to afford a linear tetranuclear molecule (Fig. 9(a)), which are linked by the O–H⋯N [H⋯N = 1.775(5) Å; ∠O–H⋯N = 166.1(3)°] hydrogen bonds from the carboxylate hydrogen atom to N(4) that terminates the extension of the tetranuclear molecule. Moreover, the tetranuclear molecules are interweaved through the Ag⋯Ag interactions [Ag⋯Ag = 3.0190(8) Å] to form a 1D chain (Fig. 9(b)). The interweaved chains (Fig. 9(c)) are also supported by the N–H⋯O hydrogen bonds from the amine hydrogen atoms to the carboxylate oxygen atoms [H⋯O = 2.362(4) and 2.234(5) Å; ∠N–H⋯O = 131.3(3) and 155.1(4)°] and coordinated water molecules [H⋯O = 2.164(5) Å; ∠N–H⋯O = 152.9(4)°], as well as the O–H⋯O hydrogen bonds from the co-crystallized water molecules to the amide oxygen atoms [O⋯H = 1.984(5) Å; ∠O–H⋯O = 169.3(4)°] and the carboxylate oxygen atoms [O⋯H = 1.942(4), 2.033(4) and 2.026(5) Å; ∠O–H⋯O = 169.5(4), 145.1(4) and 138.3(4)°]. O–H⋯O hydrogen bonds between the co-crystallized water molecules are also observed [O⋯H = 2.100(5) Å; ∠O–H⋯O = 148.4(4)°] (Fig. 9(d)).
and each asymmetric unit comprises two Ag(I) cations, one and a half L1 ligands, one 1,3,5-HBTC2− ligand, one coordinated water molecule and two co-crystallized water molecule. The Ag(1) atom is linked by two oxygen atoms from the 1,3,5-HBTC2− ligand [Ag–O = 2.176(4) Å] and water molecule [Ag–O = 2.534(5) Å] and one pyridyl nitrogen atom from the L1 ligand [Ag–N =2.175(6) Å]. Considering the Ag–O distance of 2.534(5) Å as a coordinate covalent bond, the Ag(1) atom adopts a distorted T-shaped geometry, whereas the Ag(2) metal center is three-coordinated by two oxygen atoms from the 1,3,5-HBTC2− ligand [Ag–O = 2.326(6) and 2.550(6) Å] and one pyridyl nitrogen atom from the L1 ligand [Ag–N =2.191(6) Å], resulting in a distorted triangular geometry. The Ag(I) ions are linked together by the 1,3,5-HBTC2− and L1 ligands to afford a linear tetranuclear molecule (Fig. 9(a)), which are linked by the O–H⋯N [H⋯N = 1.775(5) Å; ∠O–H⋯N = 166.1(3)°] hydrogen bonds from the carboxylate hydrogen atom to N(4) that terminates the extension of the tetranuclear molecule. Moreover, the tetranuclear molecules are interweaved through the Ag⋯Ag interactions [Ag⋯Ag = 3.0190(8) Å] to form a 1D chain (Fig. 9(b)). The interweaved chains (Fig. 9(c)) are also supported by the N–H⋯O hydrogen bonds from the amine hydrogen atoms to the carboxylate oxygen atoms [H⋯O = 2.362(4) and 2.234(5) Å; ∠N–H⋯O = 131.3(3) and 155.1(4)°] and coordinated water molecules [H⋯O = 2.164(5) Å; ∠N–H⋯O = 152.9(4)°], as well as the O–H⋯O hydrogen bonds from the co-crystallized water molecules to the amide oxygen atoms [O⋯H = 1.984(5) Å; ∠O–H⋯O = 169.3(4)°] and the carboxylate oxygen atoms [O⋯H = 1.942(4), 2.033(4) and 2.026(5) Å; ∠O–H⋯O = 169.5(4), 145.1(4) and 138.3(4)°]. O–H⋯O hydrogen bonds between the co-crystallized water molecules are also observed [O⋯H = 2.100(5) Å; ∠O–H⋯O = 148.4(4)°] (Fig. 9(d)).
Crystal structure of {[Ag(L1)0.5][(1,3,5-H2BTC)]}n, 8
The structure of complex 8 was solved in the triclinic space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) and each asymmetric unit comprises one Ag(I) cation, a half of an L1 ligand and one 1,3,5-H2BTC− ligand. The Ag(I) cation is three-coordinated by one oxygen atom from the 1,3,5-H2BTC− ligand [Ag–O = 2.164(1) Å], one pyridyl nitrogen atom from the L1 ligand [Ag–N =2.182(1) Å], and one amide oxygen atom of the L1 ligand [Ag–O = 2.584(2) Å], resulting in a distorted T-shaped geometry (Fig. 10(a)). Moreover, the structure is further extended through the Ag⋯Ag [Ag⋯Ag = 3.4378(8) Å] interactions and O–H⋯O hydrogen bonds originating from the carboxylate hydrogen atoms to the carboxylate oxygen atoms [O⋯H = 1.804(2) and 1.8388(14) Å; ∠O–H⋯O = 156.6(1)° and 174.8(1)°] (Fig. 10(b)). If the L1 ligands are considered as four-coordinated nodes, and the Ag(I) ions as two coordinated nodes, the structure of complex 8 can be regarded as a 2,4-connected 1D net with the (42)(4)2-2,4C4 topology (Fig. 10(c)).
and each asymmetric unit comprises one Ag(I) cation, a half of an L1 ligand and one 1,3,5-H2BTC− ligand. The Ag(I) cation is three-coordinated by one oxygen atom from the 1,3,5-H2BTC− ligand [Ag–O = 2.164(1) Å], one pyridyl nitrogen atom from the L1 ligand [Ag–N =2.182(1) Å], and one amide oxygen atom of the L1 ligand [Ag–O = 2.584(2) Å], resulting in a distorted T-shaped geometry (Fig. 10(a)). Moreover, the structure is further extended through the Ag⋯Ag [Ag⋯Ag = 3.4378(8) Å] interactions and O–H⋯O hydrogen bonds originating from the carboxylate hydrogen atoms to the carboxylate oxygen atoms [O⋯H = 1.804(2) and 1.8388(14) Å; ∠O–H⋯O = 156.6(1)° and 174.8(1)°] (Fig. 10(b)). If the L1 ligands are considered as four-coordinated nodes, and the Ag(I) ions as two coordinated nodes, the structure of complex 8 can be regarded as a 2,4-connected 1D net with the (42)(4)2-2,4C4 topology (Fig. 10(c)).
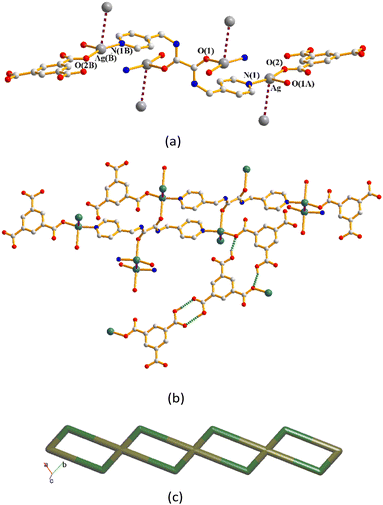 | ||
| Fig. 10 (a) A drawing showing the coordination environments of the Ag(I) cations of 8. (b) A drawing showing the Ag⋯Ag interactions and O–H⋯O hydrogen bonds. (c) A drawing showing the 2,4C4 topology. | ||
Structure of {[Ag5(1,3,5-HBTC)(1,3,5-BTC)(L1)2]·5H2O}n, 9
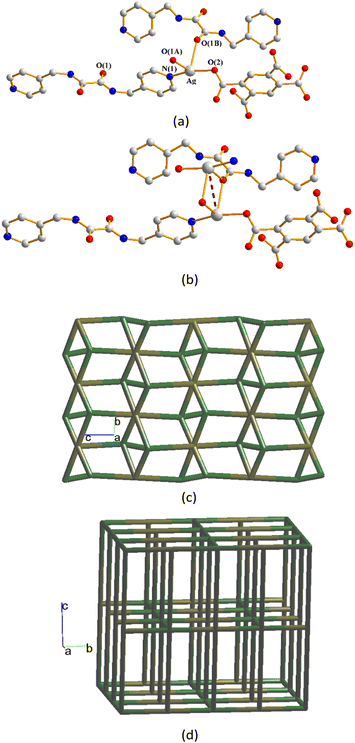
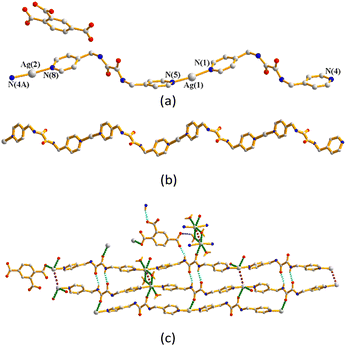
The structure of complex 9 was solved in the triclinic space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) and each asymmetric unit comprises four and two halves of Ag(I) cations, two L1 ligands, one 1,3,5-HBTC2− ligand, one 1,3,5-BTC3− ligand and five co-crystallized water molecules. While the Ag(1) and Ag(4) atoms form the distorted T-shaped geometries and the Ag(2) and Ag(5) atoms form the bent ones, Ag(3) and Ag(6) adopt linear geometries (Fig. 11(a)). The Ag(1) cation is three-coordinated by two oxygen atoms from the 1,3,5-HBTC2− [Ag–O = 2.320(3) Å] and 1,3,5-BTC3− ligands [Ag–O = 2.328(3) Å] and one pyridyl nitrogen atom from the L1 ligand [Ag–N = 2.317(3) Å]. The Ag(2) cation is two-coordinated by one oxygen atom from the 1,3,5-BTC3− [Ag–O = 2.136(3) Å] ligand and one pyridyl nitrogen atom from the L1 ligand [Ag–N = 2.138(3) Å]. The Ag(3) cation is two-coordinated by two symmetry-related oxygen atoms from the 1,3,5-BTC3− [Ag–O = 2.190(3) Å] ligand. The Ag(4) cation is three-coordinated by three oxygen atoms from one 1,3,5-HBTC2− ligand [Ag–O = 2.171(3) Å] and two 1,3,5-BTC3− ligands [Ag–O = 2.182(3) and 2.574(3) Å]. The Ag(5) cation is two-coordinated by one oxygen atom from the 1,3,5-HBTC2− [Ag–O = 2.182(3) Å] ligand and one pyridyl nitrogen atom from the L1 ligand [Ag–N = 2.169(3) Å]. The Ag(6) cation is two-coordinated by two symmetry-related nitrogen atoms from two L1 ligands [Ag–N = 2.227(3) Å].
and each asymmetric unit comprises four and two halves of Ag(I) cations, two L1 ligands, one 1,3,5-HBTC2− ligand, one 1,3,5-BTC3− ligand and five co-crystallized water molecules. While the Ag(1) and Ag(4) atoms form the distorted T-shaped geometries and the Ag(2) and Ag(5) atoms form the bent ones, Ag(3) and Ag(6) adopt linear geometries (Fig. 11(a)). The Ag(1) cation is three-coordinated by two oxygen atoms from the 1,3,5-HBTC2− [Ag–O = 2.320(3) Å] and 1,3,5-BTC3− ligands [Ag–O = 2.328(3) Å] and one pyridyl nitrogen atom from the L1 ligand [Ag–N = 2.317(3) Å]. The Ag(2) cation is two-coordinated by one oxygen atom from the 1,3,5-BTC3− [Ag–O = 2.136(3) Å] ligand and one pyridyl nitrogen atom from the L1 ligand [Ag–N = 2.138(3) Å]. The Ag(3) cation is two-coordinated by two symmetry-related oxygen atoms from the 1,3,5-BTC3− [Ag–O = 2.190(3) Å] ligand. The Ag(4) cation is three-coordinated by three oxygen atoms from one 1,3,5-HBTC2− ligand [Ag–O = 2.171(3) Å] and two 1,3,5-BTC3− ligands [Ag–O = 2.182(3) and 2.574(3) Å]. The Ag(5) cation is two-coordinated by one oxygen atom from the 1,3,5-HBTC2− [Ag–O = 2.182(3) Å] ligand and one pyridyl nitrogen atom from the L1 ligand [Ag–N = 2.169(3) Å]. The Ag(6) cation is two-coordinated by two symmetry-related nitrogen atoms from two L1 ligands [Ag–N = 2.227(3) Å].
The Ag(I) cations are interlinked through the Ag⋯Ag [Ag⋯Ag = 3.3661(22), 2.8959(16) and 3.0012(23) Å] and Ag⋯O interactions [Ag⋯O = 2.970(5), 2.749(4), 2.639(9), 2.910(5) and 2.963(4) Å] to form linear tetranuclear geometries, while the other Ag⋯O interactions [Ag⋯O = 2.799(4), 2.766(3) and 2.976(3) Å] that support the structure are also observed. Considering the Ag(1) and Ag(4) cations and 1,3,5-HBTC2− ligands as 3-connected nodes and 1,3,5-BTC3− ligands as five-connected nodes, with the other Ag(I) cations and L1 ligands as linkers, the structure of 9 can be simplified as a 3,3,3,5-connected 3D net with the (4·102)(4·6·8)(42·63·83·102)(42·6) topology (standard representation) (Fig. 11(b)), showing 3-fold interpenetration (Fig. 11(c)). Moreover, if the 1,3,5-BTC3− ligands and the bridged dinuclear Ag(I) units are considered as 4-connected nodes, the structure can be further simplified as a 4,4-connected 3D net with the (42·63·8)-sra topology (cluster representation) (Fig. 11(d)), revealing the 3-fold interpenetration (Fig. 11(e)).
Ag⋯Ag and Ag⋯O distances and structural types
Table 2 lists the Ag⋯Ag and Ag⋯O distances in complexes 1–9, which demonstrate that these distances are subjected to the changes of the polycarboxylate anions. If the Ag⋯O distance for the coordinate covalent bond is set to be less than 2.6 Å, only complexes 3, 4, 7, 8 and 9 show Ag–O coordinate covalent bonds, while the others adopt Ag⋯O interactions. On the basis of the coordinate covalent bonds involving Ag–N and Ag–O, complexes 1, 2, 4, 5 and 6 form 1D zigzag chains, whereas complex 3 is a 3D framework with the fsc topology, 7 shows interweaved tetranuclear molecules, 8 displays a 1D chain with the 2,4C4 topology and 9 reveals a 3-fold interpenetrated 3D net with the sra topology. It is noted that the Ag⋯Ag distances are also subjected to the changes of the polycarboxylate ligands. The Ag⋯Ag interactions of these complexes except 3 are supported by the polycarboxylate ligands through the weak Ag⋯O interactions and hydrogen bonds, instead of the Ag–O coordinate covalent bonds, which can most probably be regarded as “ligand-unsupported”. The Ag⋯Ag interaction of 3 is thus “ligand-supported”.| Complex | Ag⋯Ag | Ag⋯O |
|---|---|---|
| P: to the polycarboxylate oxygen atom; A: to the amide oxygen atom; W: to the water oxygen atom. | ||
| 1 | 2.895(3) and 3.216(4) (P) | |
| 2 | 3.4832(8) | 2.616(3) and 2.662(4) (P) |
| 3 | 3.4763(7) | 2.536(3) and 2.597(3) (A); 2.218(3) (P) |
| 4 | 3.2002(5) | 2.584(2) (W) |
| 5 | 2.635(2) and 2.754(2) (P) | |
| 6 | 3.1054(10) and 3.5257(10) | 2.862(4) and 3.096(4) (A); 2.862(4) and 2.968(5) (P); 2.930(7), 2.884(5) and 3.049(5) (W) |
| 7 | 3.0190(8) | 2.176(4), 2.326(6) and 2.550(6) (P); 2.534(5) (W) |
| 8 | 3.4378(8) | 2.164(1) (P); 2.584(2) (A) |
| 9 | 3.3661(22), 2.8959(16) and 3.0012(23) | 2.320(3), 2.328(3), 2.136(3), 2.190(3), 2.171(3), 2.182(3) 2.574(3), 2.910(5), 2.970(5), 2.749(4) and 2.976(3) (P); 2.639(9), 2.963(4) and 2.799(4) (W); 2.766(3) (A) |
Ligand conformations and bonding modes
The descriptor for assigning the ligand conformation is summarized pointwise: (1) based on the relative orientation of two C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (or N–H) groups, on the same or opposite direction, the ligand can adopt cis or trans conformation. (2) Due to the relative orientation of the nitrogen atom on the pyridyl ring and the carbonyl group of amide, three more orientations, anti–anti, syn–anti and syn–syn, are possible for the ligand. Accordingly, the ligand conformations of L1 and L2 in complexes 1–9 are summarized in Table 3, in which complex 9 shows two independent L1 ligands with the same ligand conformation. While the L1 ligands in complexes 3 and 8 bridge six and four Ag(I) cations, respectively, through the pyridyl nitrogen and amide oxygen atoms, the ligands in the other complexes bridge two cations through the two pyridyl nitrogen atoms, considering the Ag⋯O distances less than 2.6 Å as the coordinate covalent bonds. Moreover, Table 4 gives the bonding modes of the polycarboxylate ligands. While those in complexes 1, 2, 4, 5 and 6 show no Ag–O coordinate covalent bond, the others coordinate with one to five Ag(I) cations. Noticeably, the anions of 1,3,5-H3BTC in 7–9 coordinate at least with one Ag(I) cation, probably indicating their better basicity than the other polycarboxylate ligands toward the Ag(I) cations.
O (or N–H) groups, on the same or opposite direction, the ligand can adopt cis or trans conformation. (2) Due to the relative orientation of the nitrogen atom on the pyridyl ring and the carbonyl group of amide, three more orientations, anti–anti, syn–anti and syn–syn, are possible for the ligand. Accordingly, the ligand conformations of L1 and L2 in complexes 1–9 are summarized in Table 3, in which complex 9 shows two independent L1 ligands with the same ligand conformation. While the L1 ligands in complexes 3 and 8 bridge six and four Ag(I) cations, respectively, through the pyridyl nitrogen and amide oxygen atoms, the ligands in the other complexes bridge two cations through the two pyridyl nitrogen atoms, considering the Ag⋯O distances less than 2.6 Å as the coordinate covalent bonds. Moreover, Table 4 gives the bonding modes of the polycarboxylate ligands. While those in complexes 1, 2, 4, 5 and 6 show no Ag–O coordinate covalent bond, the others coordinate with one to five Ag(I) cations. Noticeably, the anions of 1,3,5-H3BTC in 7–9 coordinate at least with one Ag(I) cation, probably indicating their better basicity than the other polycarboxylate ligands toward the Ag(I) cations.
Powder X-ray analysis
In order to check the phase purity of the products, powder X-ray diffraction (PXRD) experiments were carried out for complexes 1–9. As shown in Fig. S10–S18, the peak positions of the experimental and simulation PXRD patterns are in agreement with each other.Conclusions
Eight bpba-based Ag(I) CPs and one tetranuclear molecule containing the polycarboxylate ligands have been successfully obtained. Their structural types are susceptible to the changes of the polycarboxylate ligands and reaction conditions. Complexes 7–9 represent unique examples that diverse structures of Ag(I) CPs constructed from L1 and 1,3,5-H3BTC can be obtained by careful evaluation of their metal to ligand ratios. While complexes 3 and 7–9 show Ag–O coordinate covalent bonds, giving 3 a 3D framework with the fsh topology, 7 a linear tetranuclear molecule, 8 a 1D looped chain, and 9 a 3-fold interpenetrated 3D framework with the sra topology, respectively, the other complexes reveal Ag⋯O interactions, adopting 1D zigzag chains. The geometries of the Ag(I) cations and the structural types of these complexes are presumably subjected to the determination of a Ag–O bond, which is set to be less than 2.6 Å in this report. The identities of the polycarboxylate ligands are important in determining the strengths of the “ligand-unsupported” and “ligand-supported” Ag⋯Ag interactions in complexes 2–4 and 6–9 as well.Author contributions
Investigation, Y.-L. S. and C.-L. L.; data curation, H.-C. T. and Z.-L. C.; review and supervision, J.-D. C. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the SI.Supplementary information: ORTEP drawings (Fig. S1–S9). PXRD patterns (Fig. S10–S18). See DOI: https://doi.org/10.1039/d5ce01151e.
CCDC 2512227–2512235 (1–9) contain the supplementary crystallographic data for this paper.32a–i
Acknowledgements
We are grateful to the National Science and Technology Council of the Republic of China for support.References
- W. P. Lustig, S. Mukherjee, N. D. Rudd, A. V. Desai, J. Li and S. K. Ghosh, Chem. Soc. Rev., 2017, 46, 3242–3285 RSC.
- S. R. Batten, S. M. Neville and D. R. Turner, Coordination Polymers Design, Analysis and Application, The Royal Society of Chemistry, London, UK, 2009 Search PubMed.
- S. Mondal and P. Dastidar, Cryst. Growth Des., 2020, 20, 7411–7420 CrossRef CAS.
- V. Chandrasekhar, C. Mohapatra, R. Banerjee and A. Mallick, Inorg. Chem., 2013, 52, 3579–3581 Search PubMed.
- J. Yu, Y. Cheng, X. Zhang, L. Zhou, Z. Song, A. Nezamzadeh-Ejhieh and Y. Huang, J. Environ. Chem. Eng., 2025, 13, 116870 Search PubMed.
- P. Yan, Z. Chen, X. Li, F. Liang, Y. Tan, Y. Lin, K. Yang, C. Xiao, J. Wu and D. Ma, J. Solid State Chem., 2024, 330, 124461 CrossRef CAS.
- F. Liang, D. Ma, L. Qin, Q. Yu, J. Chen, R. Liang, C. Zhong, H. Liao and Z. Peng, Dalton Trans., 2024, 53, 10070–10074 RSC.
- Y. Zhang, H. Tan, J. Zhu, L. Duan, Y. Ding, F. Liang, Y. Li, X. Peng, R. Jiang, J. Yu, J. Fan, Y. Chen, R. Chen and D. Ma, Molecules, 2024, 29, 5903 CrossRef CAS PubMed.
- S. Li, J. Sun, G. Liu, S. Zhang, Z. Zhang and X. Wang, Chin. Chem. Lett., 2024, 35, 109148 CrossRef CAS.
- Y. Ma, L. Yang, X. Bai and K. Wang, Spectrochim. Acta, Part A, 2025, 341, 126471 Search PubMed.
- D. Ma, T. Liang, J. Zheng, G. Chen, Y. Ye, A. Nezamzadeh-Ejhieh, L. Lu, Z. Song and Y. Huang, React. Funct. Polym., 2026, 218, 106520 Search PubMed.
- V. Lakshmanan, Y.-T. Lai, X.-K. Yang, M. Govindaraj, C.-H. Lin and J.-D. Chen, Polymer, 2021, 13, 3018 CAS.
- W.-T. Lee, T.-T. Liao and J.-D. Chen, Int. J. Mol. Sci., 2022, 23, 3603 CrossRef CAS PubMed.
- V. Lakshmanan, C.-Y. Lee, Y.-W. Tseng, Y.-H. Liu, C.-H. Lin and J.-D. Chen, CrystEngComm, 2022, 24, 6076–6086 RSC.
- J. L. Sague, M. Meuwly and K. M. Fromm, CrystEngComm, 2008, 10, 1542–1549 RSC.
- Y.-C. He, X.-H. Li, Y.-M. Ruan, Q. Xu, S.-X. Li, H.-X. Mao, F.-H. Zhao and M.-T. Li, Inorg. Chim. Acta, 2023, 557, 121710 CrossRef CAS.
- Y.-H. Li, D. Sun, G.-G. Luo, F.-J. Liu, H.-J. Hao, Y.-M. Wen and Y. Zhao, J. Mol. Struct., 2011, 1000, 85–91 CrossRef CAS.
- Y. Li a, C. Xiao, S. Li, Q. Chen, B. Li, Q. Liao and J. Niu, J. Solid State Chem., 2013, 200, 251–257 CrossRef.
- V. Moodley, L. Mthethwa, M. N. Pillay, B. Omondi and W. E. van Zyl, Polyhedron, 2015, 99, 87–95 CrossRef CAS.
- O. Z. Yeşilel, G. Günay, C. Darcan, M. S. Soylu, S. Keskin and S. W. Ng, CrystEngComm, 2012, 14, 2817–2825 RSC.
- A. Stephenson and M. D. Ward, RSC Adv., 2012, 2, 10844–10853 RSC.
- H.-Y. Shi, Y.-B. Dong, Y.-Y. Liu and J.-F. Ma, CrystEngComm, 2014, 16, 5110–5120 RSC.
- Z. Ma, X. Song, Z. Li, Y. Ren, J. Wang and Y. Liang, Dalton Trans., 2024, 53, 3797–3807 RSC.
- S. Kintzel, K. Eckhardt, J. Getzschmann, V. Bon, J. Grothe and S. Kaskel, Eur. J. Inorg. Chem., 2020, 3167–3173 CrossRef CAS.
- K. Singh, J. R. Long and P. Stavropoulos, J. Am. Chem. Soc., 1997, 119, 2942–2943 CrossRef CAS.
- S. Muthu, J. H. K. Yip and J. J. Vittal, J. Chem. Soc., Dalton Trans., 2002, 4561–4568 RSC.
- A. Bondi, J. Phys. Chem., 1964, 68, 441–451 CrossRef CAS.
- K. B. Thapa and J.-D. Chen, CrystEngComm, 2015, 17, 4611–4626 RSC.
- Bruker AXS, APEX2, V2008.6, SADABS V2008/1, SAINT V7.60A, SHELXTL V6.14, Bruker AXS Inc., Madison, WI, USA, 2008 Search PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A:Found. Crystallogr., 2008, 64, 112–122 CrossRef CAS PubMed.
- V. A. Blatov, A. P. Shevchenko and D. M. Proserpio, Cryst. Growth Des., 2014, 14, 3576–3586 Search PubMed.
- (a) CCDC 2512227: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5lv; (b) CCDC 2512228: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5mw; (c) CCDC 2512229: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5nx; (d) CCDC 2512230: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5py; (e) CCDC 2512231: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5qz; (f) CCDC 2512232: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5r0; (g) CCDC 2512233: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5s1; (h) CCDC 2512234: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5t2; (i) CCDC 2512235: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2qb5v3.
| This journal is © The Royal Society of Chemistry 2026 |