Open Access Article
Open Access ArticleCoordination compounds based on a phosphonic diamide-TEMPO diradical and CuII, NdIII, EuIII and TbIII ions: synthesis, magnetic properties and application in the aerobic oxidation of allylic and benzylic alcohols
Yolanda Navarro a,
Stéphane Soriano
a,
Stéphane Soriano c,
Henrique de Castro Silva Jr.
c,
Henrique de Castro Silva Jr. bd,
María José Iglesias
bd,
María José Iglesias a,
Guilherme P. Guedes
a,
Guilherme P. Guedes *b and
Fernando López Ortiz
*b and
Fernando López Ortiz *a
*a
aÁrea de Química Orgánica, Centro de Investigación CIAIMBITAL, Universidad de Almería, Carretera de Sacramento s/n, 04120 Almería, Spain. E-mail: flortiz@ual.es
bUniversidade Federal Fluminense, Instituto de Química, Niterói, Rio de Janeiro, Brazil. E-mail: guilherme_guedes@id.uff.br
cInstituto de Física, Universidade Federal Fluminense, Niterói, Rio de Janeiro, Brazil
dInstituto de Química, Departamento de Química Fundamental, Universidade Federal Rural do Rio de Janeiro, Seropédica, Rio de Janeiro, Brazil
First published on 19th February 2026
Abstract
The synthesis and structure of a novel phenylphosphonic diamide bis-TEMPO (2,2,6,6-tetramethylpiperidine-1-oxyl) diradical (ppdATEMPO) and of four M(hfac)n (hfac = hexafluoroacetylacetonate; M = CuII, NdIII, EuIII or TbIII; n = 2, 3) complexes are reported herein. The X-ray diffraction studies revealed that the compounds consist of five-coordinated [Cu(ppdATEMPO)(hfac)2] and eight-coordinated [Ln(ppdATEMPO)2(hfac)3] (Ln = NdIII, EuIII, TbIII) complexes due to the exclusive binding of the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group of the ligand to the metal ions. Two-dimensional (2D) supramolecular networks are formed through intermolecular N–O⋯H–N(P) hydrogen bonding interactions. In the case of the free diradical, the multilayer supramolecular structure is additionally supported by P
O group of the ligand to the metal ions. Two-dimensional (2D) supramolecular networks are formed through intermolecular N–O⋯H–N(P) hydrogen bonding interactions. In the case of the free diradical, the multilayer supramolecular structure is additionally supported by P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O⋯H–N and N–O⋯H–C inter-chain interactions. A DFT study confirmed the exclusive coordination of ppdATEMPO to all metal cations investigated via the oxygen atom of the P
O⋯H–N and N–O⋯H–C inter-chain interactions. A DFT study confirmed the exclusive coordination of ppdATEMPO to all metal cations investigated via the oxygen atom of the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group. This is attributed to the greater donor ability and steric accessibility of the P
O group. This is attributed to the greater donor ability and steric accessibility of the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group in comparison with the two nitroxide moieties. The magnetic measurements of the complexes indicated the presence of weak intermolecular antiferromagnetic interactions. The behaviour of the copper complex as a catalysts in the aerobic oxidation of alcohols has been also investigated.
O group in comparison with the two nitroxide moieties. The magnetic measurements of the complexes indicated the presence of weak intermolecular antiferromagnetic interactions. The behaviour of the copper complex as a catalysts in the aerobic oxidation of alcohols has been also investigated.
Introduction
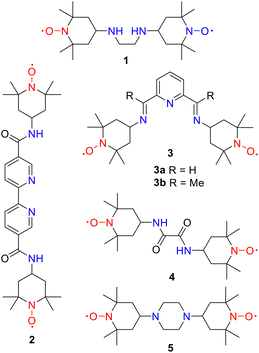
Nitroxyl radicals are a highly valuable class of open-shell compound, with a wide range of applications across various fields, including catalysis,1 molecular biology and medicine,2 energy storage systems3 or materials science.4 Radicals of the TEMPO (2,2,6,6-tetramethylpiperidine-1-oxyl) family play a key role as they can be easily functionalised in the 4-position, thereby expanding the range of applications.5 Stable organic diradicals are currently the focus of great attention. The presence of two unpaired electrons in these molecules endows them with properties that are distinct from, or superior to, those of monoradicals due to their spin-related characteristics.6 These multi-spin systems have numerous applications, including use in organic magnets,6,7 components in spintronic,8 and optical devices,9 building blocks for energy harvesting and storage systems,6b,8b,10 polymerization initiators,11 contrast agents for magnetic resonance imaging,12 polarising agents in DNP-NMR (Dynamic Nuclear Polarization-Nuclear Magnetic Resonance) spectroscopy,13 probes for the investigation of cellular processes,14 and bioactive molecules.6d,15 A large variety of bis(ATEMPO) diradicals have been synthesised by connecting two ATEMPO units. The properties of these molecules can be tuned by adjusting the various structural parameters, like the length, flexibility and functional groups present in the linker.6,16 The heteroatoms present within the linker may function as additional coordination sites for metal cations, leading to the formation of heterospin systems, which exhibit unique architectures and physicochemical properties. The topology and conformation adopted by the diradical will determine the possible spin–spin intra- and intermolecular interactions. Additionally, metal ion complexation can result in altered coupling pathways.17Notwithstanding the favourable outlook, a modest number of complexes of bis(ATEMPO) derivatives with transition metals have been characterised at the molecular level. In most cases, these complexes do not show coordination between the metal ion and the nitroxyl group. Thus, the ligand 1 (Fig. 1), with its ATEMPO units connected by an ethylene bridge, forms a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 complex upon reaction with Mn(hfac)2 (hfac = hexafluoroacetylacetonate) through metal cation binding to the ethylenediamine moiety, which has shown very weak antiferromagnetic interactions (estimated J/kB = −2.5 K).18 Similarly, the diradical 2 obtained by amide formation between two ATEMPOs and [2,2′-bipyridine]-5,5′-dicarbonyl dichloride, results in the formation of a dimeric Cu(II) complex with CuCl2 via metal ion chelation by the bipyridine heterocycle.19 The compound has been shown to be an effective catalyst in the aerobic oxidation of primary alcohols.
1 complex upon reaction with Mn(hfac)2 (hfac = hexafluoroacetylacetonate) through metal cation binding to the ethylenediamine moiety, which has shown very weak antiferromagnetic interactions (estimated J/kB = −2.5 K).18 Similarly, the diradical 2 obtained by amide formation between two ATEMPOs and [2,2′-bipyridine]-5,5′-dicarbonyl dichloride, results in the formation of a dimeric Cu(II) complex with CuCl2 via metal ion chelation by the bipyridine heterocycle.19 The compound has been shown to be an effective catalyst in the aerobic oxidation of primary alcohols.
A higher structural complexity was observed in the complexation of 3a (R = H) with CuBr2. A cluster was formed consisting of four Cu(I) and four Cu(II) ions, and two ligands acting as tridentate coordinating agents by binding through the nitrogen atoms of the pyridinebis(imine) moiety to two Cu(II) ions.20 The mixed Cu(I)/Cu(II) complex exhibited antiferromagnetic behaviour assigned to intermolecular nitroxide⋯nitroxide (2J/kB = −25.6 ± 0.6 K) and intramolecular copper(II)⋯copper(II) interactions (estimated 2J/kB = −85 ± 2 K). By contrast, the methyl derivative 3b reacts with [NiII(CH3CN)5](BF4)2 to give a dicationic Ni(II) complex with a more simple architecture. The Ni(II) ion is N,N,N-tricoordinated to 3b and completes the pseudo-octahedral geometry by binding to three acetonitrile molecules.21 The diradical 4, having a bisamide bridge, is the precursor of a dinuclear Pd(II) complex, in which the metal ions are N,O-chelated by the bisamide linker.22 A number of examples of bis(ATEMPO) complexes have been synthesised by derivatisation of suitably functionalised metal complexes, typically through amide linkages. The metal-complex scaffolds include ferrocene- and cobaltocene-bridges,23 cyclometallated iridium derivatives,15c and boron-containing iron(II) clathrochelates.24
Complexation by metal ion binding to the oxygen atom of the N–O· groups of bis(ATEMPO) diradicals has only been observed for the reaction of ligand 5 with an excess of Mn(hfac)2.18 The value of the Curie constant in the dinuclear complex formed is lower than expected due to the cancellation of one of the spins of each Mn atom with the TEMPO radical. The estimated exchange constant J/kB = −424 K is indicative of the existence of strong antiferromagnetic spin interactions.
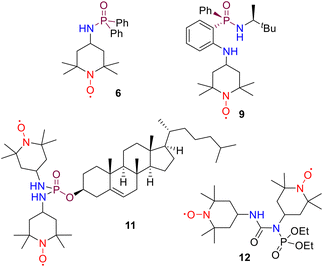
It has been shown that the substituents present in the C-4 position of TEMPO-type nitroxides can have a considerable impact on their physicochemical properties. This phenomenon is associated to the electronic nature of these substituents, despite the absence of π-conjugation between the radical and the substituent in these molecules.15b,25 P-Containing 4-amino-TEMPO (ATEMPO) stable mononitroxyl radicals have proven to be particularly interesting. They can be readily synthesised via P–N bond-forming reactions. To date, numerous compounds have been prepared including, phosphoramidates, thiophosphoramidates, phosphorodiamidates, phosphorodiamidic acids, phosphoric acid triamides, thiophosphoric acid triamides or cyclotriphosphazenes, among others.26 In recent years, our research group has described several examples of phosphinic amide, thiophosphinic amide and selenophosphinic amide27 ATEMPO radicals, and their application in molecular magnetism. Radical 6 (dppnTEMPO) has been used as building block for the construction of coordination compounds of first row metal ions 7 (CuII, CoII, MnII), acting as a mono or bidentate ligand through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and N–O· groups (Fig. 2).28 However, complexes with the rare earth metals GdIII, TbIII and DyIII, 8, exhibited monodimensional polymeric chains, with the ligand functioning as a bridge between paramagnetic sites (Fig. 2).29 A magnetic inequivalence has been identified between lanthanide ions in the TbIII and DyIII complexes. Interestingly, the ATEMPO radical has also been introduced into a P-stereogenic phosphinamide 9. The coordination behaviour of radical 9 with CuII gave rise to a chiral complex 10 with a singular polymeric helicate chain where the radical acts again as a bridge between copper atoms through the P
O and N–O· groups (Fig. 2).28 However, complexes with the rare earth metals GdIII, TbIII and DyIII, 8, exhibited monodimensional polymeric chains, with the ligand functioning as a bridge between paramagnetic sites (Fig. 2).29 A magnetic inequivalence has been identified between lanthanide ions in the TbIII and DyIII complexes. Interestingly, the ATEMPO radical has also been introduced into a P-stereogenic phosphinamide 9. The coordination behaviour of radical 9 with CuII gave rise to a chiral complex 10 with a singular polymeric helicate chain where the radical acts again as a bridge between copper atoms through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and N–O· groups. The enantiopure compounds show weak antiferromagnetic interactions for the ligand, while ferro- and antiferromagnetic properties are present in complex 10 due to the direct coordination of the nitroxide and copper paramagnetic sites.30
O and N–O· groups. The enantiopure compounds show weak antiferromagnetic interactions for the ligand, while ferro- and antiferromagnetic properties are present in complex 10 due to the direct coordination of the nitroxide and copper paramagnetic sites.30
In contrast to prevailing expectations, the number of reported P-containing TEMPO bis(nitroxides) has been found to be significantly lower than anticipated. These are limited to a few examples of bis(4-hydroxy-TEMPO) with phosphonate,31 phosphonoselenoate,32 phosphoramidate33 and pyrophosphate34 functional groups. To the best of our knowledge, only one example of a bis(ATEMPO) diradical with a P-based linker has hitherto been reported. The phosphorodiamidate 11 bearing a cholesteryl substituent has been prepared as a potential spin probe to monitor processes within living systems (Fig. 2).35 A related phosphorus-containing bis(ATEMPO) diradical is the phosphoramidate 12, in which the phosphorus functional group is linked to one of the nitrogen atoms of the urea bridge involving the two ATEMPO units (Fig. 2).36 The molecular structure of these compounds, in addition to their magnetic properties and synthetic applications remain to be explored.
A highly attractive application of nitroxyl radicals involves their use as catalysts in the selective oxidation of primary alcohols to carbonyl compounds. Traditional methods require stoichiometric amounts of oxidants leading to a large amount of toxic wastes.37 The utilisation of molecular oxygen as oxidant, in combination with a minimal amount of a metal catalyst is advantageous due to the formation of benign by-products and the low cost of O2.38 For this purpose, copper is the metal of choice as it is inexpensive and is present in a variety of enzymes such as the galactose oxidase, which catalyses the oxidation of alcohols in vivo.39 The combination of Cu-complexes with TEMPO and O2 is a very active catalytic system for the selective oxidation of primary alcohols.1b,c,40
In this work, we present the synthesis of a novel phosphonic diamide bis(ATEMPO) diradical and its application to the design of magnetic materials by combination of the organic diradical with metal ions. In order to evaluate the differences in the magnetic interactions involving transition-metal complexes and lanthanide-based compounds, we have synthesised four new coordination complexes, incorporating CuII, NdIII, EuIII and TbIII ions. DFT calculations provided support for the exclusive complexation through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group that was observed. The study of the magnetic behaviour of these compounds revealed prevailing weak intermolecular antiferromagnetic exchange interactions. Additionally, the copper complex has been examined as a catalyst in the aerobic oxidation of alcohols.
O group that was observed. The study of the magnetic behaviour of these compounds revealed prevailing weak intermolecular antiferromagnetic exchange interactions. Additionally, the copper complex has been examined as a catalyst in the aerobic oxidation of alcohols.
Experimental section
Materials
The synthesis of the diradical was carried out under a dry N2 gas atmosphere using standard Schlenk procedures. THF (99.5%, purchased from Panreac) was distilled from sodium/benzophenone immediately prior to its use. Commercial solvents n-hexane (95%, Panreac) and ethyl acetate (99.8%, Panreac) were distilled prior to their use, n-heptane (HPLC grade, Scharlau) and dichloroethane (99%, Sigma-Aldrich) were used as received. Et3N (99%, Acros Organics) was dried over KOH pellets and then distilled under N2 atmosphere. PhPCl2 (97%, Acros Organics) was distilled prior to use. The [M(hfac)n] building blocks (M = CuII (ABCR), NdIII (ABCR), EuIII (99.9%, Rare Earth Products) and TbIII (99.9%, Rare Earth Products)) were used as received. 4-Amino-TEMPO (97%, ABCR) and the alcohols used for the catalysis (18a: 99%, Acros Organics; 18b: 99%, Acros Organics; 18c: 98%, Sigma-Aldrich; 18d: 99%, ThermoScientific; 18e: 98%, Sigma-Aldrich; 18f: 97%, Acros Organics; 18g: 98%, Sigma-Aldrich; 18h: 99%, Sigma-Aldrich; 18i: 98%, Merck; 18j: 99%, Sigma-Aldrich; 18k: 98%, Sigma-Aldrich; 18l: 98%, Sigma-Aldrich; 18m: 98%, Fluka; 18n: 98%, Sigma-Aldrich; 18o: 98%, Sigma-Aldrich) were also used as received. NMR spectra were obtained on a Bruker Avance 300 (1H, 300.13 MHz, 31P, 121.49 MHz) using a 5 mm BBFO 1H, BB (19F–15N) probe. The spectral references used were internal tetramethylsilane for 1H and external 85% H3PO4 for 31P. Infrared spectra were recorded on a Bruker Alpha FTIR spectrophotometer. High resolution mass spectra were recorded on an Agilent Technologies LC/MSD TOF instrument using electrospray ionization. The melting points were measured on a STUART SMP20 capillary melting point apparatus.X-ray diffraction
X-ray diffraction crystallographic data were collected using a Bruker APEX II CCD area detector at 100 K, using CuKα (λ = 1.54178 Å) or MoKα radiation (λ = 0.71073 Å). Data collection and cell refinement were performed with Bruker APEX241 and Bruker SAINT,42 respectively. Data reduction was done using SAINT. Empirical multiscan absorption correction using equivalent reflections was performed using the SADABS program.43 The structure solution and full matrix least-squares refinements based on F2 were performed using SHELXS44 and SHELXL45 programs, respectively. All atoms except hydrogen were refined anisotropically. The structures were drawn using the Mercury program.46 Details of data collection and structure refinement for compounds 13–17 are summarized in Table S1 (SI). The ORTEP47 view of the asymmetric units are depicted in Fig. S14–S18 (SI). The Cremer–Pople puckering parameters were calculated for compound 13 using PLATON (v-100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 117).48 Compound 14 exhibits large accessible voids, which are occupied by disordered crystallisation solvent molecules. A solvent masking routine implemented in the Olex2 program was employed,49 revealing a total void volume of 436 Å3 and 184 electrons per unit cell, corresponding to 23 electrons per asymmetric unit (Z = 8). This electron count is in good agreement with the presence of half toluene molecule per asymmetric unit (25 e−). Disorder was observed for at least one hfac ligand in 14–16, as well as for several CF3 groups and one nitroxide group in 17. The hfac ligands, CF3 and nitroxide moieties were modelled as disordered over two alternative conformations, except in 14, where one CF3 group required modelling over three alternative orientations. The occupancies of all disorder components were freely refined. Distance restraints (SADI) and equivalent displacement parameter constraints (EADP), together with ISOR or SIMU restraints, were applied to ensure chemically sensible C–F, C–C, C–O, N–O and C–N bond lengths and reasonable displacement parameters. For compounds 15 and 16, the crystal data were refined as inversion twins using the twin law (−1 0 0/0 –1 0/0 0 –1), yielding refined twin fractions of 0.961(5) and 0.039(5) for 15, and 0.815(9) and 0.185(9) for 16.
117).48 Compound 14 exhibits large accessible voids, which are occupied by disordered crystallisation solvent molecules. A solvent masking routine implemented in the Olex2 program was employed,49 revealing a total void volume of 436 Å3 and 184 electrons per unit cell, corresponding to 23 electrons per asymmetric unit (Z = 8). This electron count is in good agreement with the presence of half toluene molecule per asymmetric unit (25 e−). Disorder was observed for at least one hfac ligand in 14–16, as well as for several CF3 groups and one nitroxide group in 17. The hfac ligands, CF3 and nitroxide moieties were modelled as disordered over two alternative conformations, except in 14, where one CF3 group required modelling over three alternative orientations. The occupancies of all disorder components were freely refined. Distance restraints (SADI) and equivalent displacement parameter constraints (EADP), together with ISOR or SIMU restraints, were applied to ensure chemically sensible C–F, C–C, C–O, N–O and C–N bond lengths and reasonable displacement parameters. For compounds 15 and 16, the crystal data were refined as inversion twins using the twin law (−1 0 0/0 –1 0/0 0 –1), yielding refined twin fractions of 0.961(5) and 0.039(5) for 15, and 0.815(9) and 0.185(9) for 16.
Computational details
All quantum-mechanical calculations at the molecular level were performed using the ORCA 6.1.0 package50 with tight convergence criteria set to 1.000 × 10−08 Eh. The molecular structure used for computational calculations was based on a full conformational sampling using the GOAT51 module in ORCA and the lowest conformer was reoptimized at the ωB97M-V/Def2-TZVP52,53 level of theory. Electronic and magnetic properties were investigated using Density Functional Theory (DFT) with an unrestricted Kohn-Sham wavefunction also at the ωB97M-V/Def2-TZVP level. The broken-symmetry methodology was based on the widely used Heisenberg-Dirac-van Vleck Hamiltonian (eqn (1)), and the magnetic coupling constant (Jab) was evaluated using spin projection as proposed by Soda and Yamaguchi (eqn (2)).54,55Natural Bond Orbital calculations used the NBO 7.0 package.56 For each ligand we extracted (i) natural charges to compare donor basicity, (ii) natural spin densities to examine radical character and N/O spin distribution in the TEMPO fragments, (iii) second-order donor–acceptor stabilization energies E(2) for the key lone-pair interactions, and (iv) Natural Localized Molecular Orbitals NLMO-based steric exchange energies.
| Ĥ = −2J·Sa·Sb | (1) |
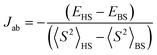 | (2) |
The spin-averaged total steric exchange from the NLMO partition was used as a global measure of steric congestion for ligands 1, 5, 6 and 13.
Magnetic measurements
The measurements were performed using a Quantum Design MPMS-XL instrument. The samples were mounted in gelatine capsules. The working field was applied at approximately 35 K, which indicates that 2 K was attained under field cooling conditions, starting at 35 K. The RSO method was used and consists of repeatedly moving the sample through the gradiometer. The amplitude of the displacement is 4 cm (2 cm above and 2 cm below the center of the gradiometer). The temperature sweep was subdivided into two segments: the initial segment ranged from 2 K to 10 K at a heating rate of 0.5 K min−1. This approach was adopted to ensure the acquisition of a substantial number of points within the area deemed to be of paramount interest. The second measurement was obtained at a faster pace (10–300 K at 3 K min−1) and with a reduced number of points. An additional measurement was carried out using an empty capsule under identical conditions to ascertain the effect of the capsule on the measurement and to enable the implementation of necessary corrections.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). HRMS (ESI) calcd. for [C24H42N4O3P]+: 465.2995, found: 465.2998. The structural assignment was confirmed by adding phenylhydrazine to the NMR sample to give N,N′-bis(1-hydroxy-2,2,6,6-tetramethylpiperidin-4-yl)-P-phenylphosphonic diamide. 1H NMR δ (CDCl3, 300.13 MHz) 7.86 (ddd, 2H, 3JPH 12.0, 3JHH 8.2, 4JHH 1.5 Hz), 7.50 (m, 3H), 3.48 (m, 2H), 2.80 (m, 2H), 1.91 (m, 2H), 1.73 (m, 2H), 1.43 (m, 4H), 1.21 (s, 6H), 1.19 (s, 6H), 1.16 (s, 6H), 1.15 (s, 6H). 31P NMR δ (CDCl3, 121.49 MHz) 18.9 ppm.
O). HRMS (ESI) calcd. for [C24H42N4O3P]+: 465.2995, found: 465.2998. The structural assignment was confirmed by adding phenylhydrazine to the NMR sample to give N,N′-bis(1-hydroxy-2,2,6,6-tetramethylpiperidin-4-yl)-P-phenylphosphonic diamide. 1H NMR δ (CDCl3, 300.13 MHz) 7.86 (ddd, 2H, 3JPH 12.0, 3JHH 8.2, 4JHH 1.5 Hz), 7.50 (m, 3H), 3.48 (m, 2H), 2.80 (m, 2H), 1.91 (m, 2H), 1.73 (m, 2H), 1.43 (m, 4H), 1.21 (s, 6H), 1.19 (s, 6H), 1.16 (s, 6H), 1.15 (s, 6H). 31P NMR δ (CDCl3, 121.49 MHz) 18.9 ppm.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 675 (m). Anal. Calc. for C34H43CuF12N4O7P: C, 43.34%; H, 4.60%; N, 5.95%; found: C, 43.33%; H, 4.90%; N, 6.05% (elemental analysis indicates that no solvent is present in the final structure. The loss of the toluene molecule is attributable to the sample preparation procedure, in which the compound is freeze-dried under vacuum for several days in order to remove any trace of volatile impurities). 15 [Nd(ppdATEMPO)2(hfac)3]: Yield: 66%. Mp: 195–196 °C. IR (ATR, cm−1): 3341 (w, NH), 1651 (s, CO), 1252 (s, CF), 1194 (s, CF), 1140 (s, P
O), 675 (m). Anal. Calc. for C34H43CuF12N4O7P: C, 43.34%; H, 4.60%; N, 5.95%; found: C, 43.33%; H, 4.90%; N, 6.05% (elemental analysis indicates that no solvent is present in the final structure. The loss of the toluene molecule is attributable to the sample preparation procedure, in which the compound is freeze-dried under vacuum for several days in order to remove any trace of volatile impurities). 15 [Nd(ppdATEMPO)2(hfac)3]: Yield: 66%. Mp: 195–196 °C. IR (ATR, cm−1): 3341 (w, NH), 1651 (s, CO), 1252 (s, CF), 1194 (s, CF), 1140 (s, P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 659 (m). 31P NMR δ (CDCl3, 121.49 MHz) 101 ppm. Anal. Calc. for C63H85NdF18N8O12P2: C, 44.65%; H, 5.06%; N, 6.61%; found: C, 45.37%; H, 5.732%; N, 6.39%. 16 [Eu(ppdATEMPO)2(hfac)3]: Yield: 63%. Mp: 145–146 °C. IR (ATR, cm−1): 3344 (w, NH), 1651 (s, CO), 1252 (s, CF), 1194 (s, CF), 1138 (s, P
O), 659 (m). 31P NMR δ (CDCl3, 121.49 MHz) 101 ppm. Anal. Calc. for C63H85NdF18N8O12P2: C, 44.65%; H, 5.06%; N, 6.61%; found: C, 45.37%; H, 5.732%; N, 6.39%. 16 [Eu(ppdATEMPO)2(hfac)3]: Yield: 63%. Mp: 145–146 °C. IR (ATR, cm−1): 3344 (w, NH), 1651 (s, CO), 1252 (s, CF), 1194 (s, CF), 1138 (s, P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 659 (m). 31P NMR δ (CD3CN 121.49 MHz) −120 ppm. Anal. Calc. for C63H85EuF18N8O12P2: C, 44.37%; H, 5.20%; N, 6.57%; found: C, 44.48%; H, 5.184%; N, 6.63%. 17 [Tb(ppdATEMPO)2(hfac)3]: Yield: 80%. Mp: 207–208 °C. IR (ATR, cm−1): 3321 (w, NH), 1646 (s, CO), 1258 (s, CF), 1202 (s, CF), 1148 (s, P
O), 659 (m). 31P NMR δ (CD3CN 121.49 MHz) −120 ppm. Anal. Calc. for C63H85EuF18N8O12P2: C, 44.37%; H, 5.20%; N, 6.57%; found: C, 44.48%; H, 5.184%; N, 6.63%. 17 [Tb(ppdATEMPO)2(hfac)3]: Yield: 80%. Mp: 207–208 °C. IR (ATR, cm−1): 3321 (w, NH), 1646 (s, CO), 1258 (s, CF), 1202 (s, CF), 1148 (s, P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 670 (m). 31P NMR δ (CDCl3, 121.49 MHz) −223 ppm. Anal. Calc. for C63H85TbF18N8O12P2: C, 44.27%; H, 5.01%; N, 6.56%; found: C, 44.17%; H, 5.292%; N, 6.72%.
O), 670 (m). 31P NMR δ (CDCl3, 121.49 MHz) −223 ppm. Anal. Calc. for C63H85TbF18N8O12P2: C, 44.27%; H, 5.01%; N, 6.56%; found: C, 44.17%; H, 5.292%; N, 6.72%.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 for compounds 19, 23 and 27; 1
1 for compounds 19, 23 and 27; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 for 20, 29 and 30; and 1
2 for 20, 29 and 30; and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 for 26) as the eluent mixture.
4 for 26) as the eluent mixture.Results and discussion
Synthesis and structural characterization
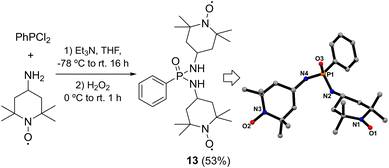
The synthesis of the new phosphonic amide-TEMPO diradical 13 has been carried out by extension of the methodology developed by our research group for the preparation of radical 6.28 The reaction consisted of the nucleophilic substitution of dichlorophenylphosphine with 2 equivalents of 4-amino-TEMPO in the presence of Et3N at −78 °C, followed by oxidation with H2O2 (Scheme 1). After purification through crystallisation in ethyl acetate, compound 13 was obtained as red crystals suitable for X-ray diffraction in 53% yield. The displacement of the two chlorides of PhPCl2 by two ATEMPO units in 13 is corroborated by the quasi-molecular ion of m/z 465.2998 observed in the HRMS (ESI) mass spectrum, as compared with the calculated value obtained for [C24H42N4O3P]+ of m/z 465.2995. The 1H-NMR spectrum of 13 showed the characteristic broad signals for one phenyl ring and two C4-heterosubstituted TEMPO units (Fig. S1).With regard to the 31P NMR spectrum, the relatively broad signal observed at δP 23.8 ppm is consistent with the expected chemical shift range for a phosphonic diamide linkage (Fig. S2).57 These assignments were confirmed through the measurement of the 1H- and 31P NMR spectra of the bis(hydroxylamine) obtained by reduction of 13 upon treatment with phenylhydrazine (Fig. S3 and S4).
The reaction of diradical 13 with equimolar amounts of M(hfac)n (M = CuII, NdIII, EuIII and TbIII) in hot n-heptane afforded complexes 14–17 in good to excellent yields (50% for 14, 63% for 15, 66% for 16 and 80% for 17) (Scheme 2).
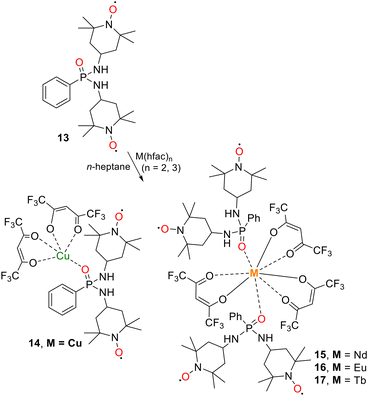 | ||
| Scheme 2 Synthesis and structural representation of complexes 14 [Cu(ppdATEMPO)(hfac)2] and 15–17 [Ln(ppdATEMPO)2(hfac)3], Ln = Nd(III), Eu(III), Tb(III). | ||
Regarding the NMR spectroscopic characterization of the complexes, only the 31P{1H} NMR spectra of the lanthanide derivatives 15–17 provided useful information, despite their broadness (Fig. S5). The observed δP were 101 ppm (half height linewidth W½ = 427 Hz) for 15, −120 ppm (W½ = 1952 Hz) for 16 and −223 ppm (W½ = 636 Hz) for 17. These values correspond to isotropic shifts of +77, −144 and −247 ppm, respectively. For the Cu(II) complex 14 no 31P{1H} NMR signal could be detected within the chemical shift range of 1400 to −800 ppm. The addition of phenylhydrazine to the NMR samples of 15–17 resulted in the loss of the 31P signals due to decomposition. The 1H NMR spectra of all complexes exhibited a high degree of signal broadening, which precluded the possibility of performing a structural analysis.
Crystal structures
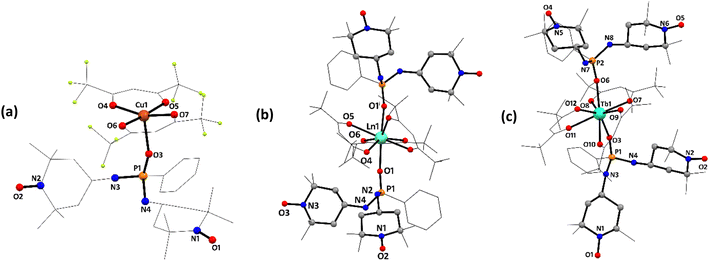
Single crystal X-ray diffraction studies were performed on compounds 13–17. Their molecular structures are shown in Scheme 1 and Fig. 3–5. ORTEP diagrams illustrating the refined structures of these compounds can be found in Fig. S14–S18 (SI). Selected geometrical parameters found in 13–17 are listed in Tables 1, 2 and S2. Table S1 details a selection of the experimental data and refinement parameters derived from compounds 13–17 (SI).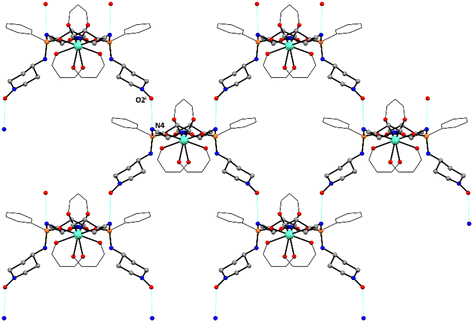 | ||
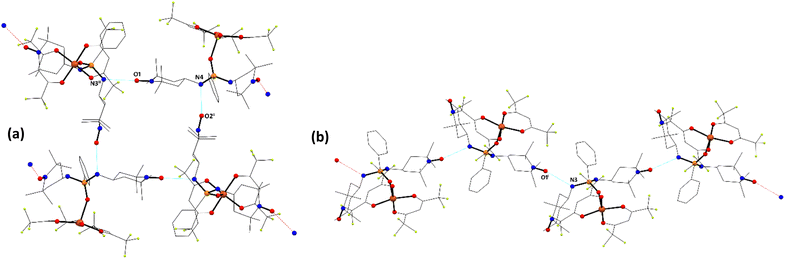
| Fig. 5 Crystal packing of compound 15 and 16 showing intermolecular hydrogen bonds (cyan dotted lines). Hydrogen atoms, parts of 13 and hfac ligands have been omitted for clarity. | ||
| P1–O3 | 1.4814(1) | N1–O1 | 1.291(1) |
| P1–N2 | 1.620(1) | N3–O2 | 1.291(2) |
| P1–N4 | 1.650(1) | N2–C5 | 1.467(2) |
| P1–C19 | 1.802(1) | N4–C14 | 1.481(2) |
| O3–P1–N2 | 111.67(6) | N4–P1–C19 | 102.16(6) |
| O3–P1–N4 | 117.64(6) | P1–N4–C14 | 119.0(1) |
| O3–P1–C19 | 110.88(6) | O1–N1–C1 | 115.8(1) |
| C19–P1–N2 | 111.01(7) | O1–N1–C7 | 115.9(1) |
| N2–P1–N4 | 102.90(6) | O2–N3–C10 | 115.6(1) |
| P1–N2–C5 | 125.8(1) | O2–N3–C16 | 116.0(1) |
| Cu1–O3 | 2.118(2) | P1–N3 | 1.626(3) | |
| Cu1–O4 | 1.965(2) | P1–N4 | 1.641(4) | |
| Cu1–O5a | 1.950(13) | P1–C19 | 1.796(4) | |
| Cu1–O6a | 1.945(4) | N1–O1 | 1.289(4) | |
| Cu1–O7a | 1.96(4) | N2–O2 | 1.288(4) | |
| P1–O3 | 1.488(3) | |||
| O3–P1–N3 | 118.7(2) | N3–P1–N4 | 103.0(2) | |
| O3–P1–C19 | 110.2(2) | N4–P1–C19 | 114.2(2) | |
| O3–P1–N4 | 107.3(2) | P1–O3–Cu1 | 151.6(2) | |
| N3–P1–C19 | 103.0(2) | |||
| Hydrogen-bond geometry | ||||
|---|---|---|---|---|
| Symmetry codes: (i) x, −y, z + 1/2; (ii) –x + 1/2, y − 1/2, −z + 1/2.a Average bond length resulting from full or partial positional disorder of the hfac ligand over two sites. | ||||
| D–H⋯A | D–H | H⋯A | D⋯A | D–H⋯A |
| N3–H3⋯O1i | 0.83(4) | 2.11(4) | 2.883(3) | 155(3) |
| N4–H4⋯O2ii | 0.78(4) | 2.21(2) | 2.962(3) | 163(4) |
Compound 13 crystallises in the monoclinic P21/c group and its molecular structure is shown in Scheme 1 and Fig. S14. The phosphorus atom adopts a distorted tetrahedral geometry with angles varying between 102.16(6) and 117.64(6)° (Table 1). The P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond distance is 1.481(1) Å, in agreement with other observed in related systems (Table 1).58 The nitrogen atoms bonded to the P
O bond distance is 1.481(1) Å, in agreement with other observed in related systems (Table 1).58 The nitrogen atoms bonded to the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group deviate slightly from a planar configuration (sum of bond-angles for N1/N2 about 1.1°/11.4° less than the ideal 360° of a sp2 configuration).
O group deviate slightly from a planar configuration (sum of bond-angles for N1/N2 about 1.1°/11.4° less than the ideal 360° of a sp2 configuration).
The piperidine rings of the TEMPO moieties are in the expected chair configuration, with N–O· distances of 1.291(2) Å, similar to other nitroxide fragments described in the literature.27,30,59 The conformations of the six-membered rings N1/C1/C4–C7 and N2/C10/C13–C16 were confirmed by Cremer–Pople puckering analysis. The puckering amplitudes are Q = 0.4930(15) and 0.5079(16) Å, with corresponding θ values of 160.96(16)° and 156.98(17)°, and φ values of 351.4(5)° and 3.9(5)°, respectively. The θ values are close to 180°, consistent with slightly distorted chair conformations, while the φ values indicate minor deviations from idealized chair geometry.60
The N–O· groups of the ATEMPO units are arranged quasi-perpendicularly leading to an intramolecular O⋯O distance between the radical moieties of 10.012(2) Å. The P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group and the N–H of the phosphonic amide linkage antiparallel to it (torsion angle O3–P1–N2–H2 of 172(1)°) are involved in intermolecular hydrogen bonds between neighbour molecules of moderate strength according to Jeffrey's classification (N2–H2⋯O3i, 2.782(2) Å, i = 3/2 − x, y − 1/2, 1/2 − z).61
O group and the N–H of the phosphonic amide linkage antiparallel to it (torsion angle O3–P1–N2–H2 of 172(1)°) are involved in intermolecular hydrogen bonds between neighbour molecules of moderate strength according to Jeffrey's classification (N2–H2⋯O3i, 2.782(2) Å, i = 3/2 − x, y − 1/2, 1/2 − z).61
This feature, characteristic of phosphoryl compounds containing an N–H group,62 gives rise to a linear polymeric chain with a quasi-planar arrangement of the O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) P–N–H moieties of the molecules and an alternating right/left orientation with respect to the P
P–N–H moieties of the molecules and an alternating right/left orientation with respect to the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O axis of the ATEMPO unit acting as a hydrogen donor. Furthermore, the second N–H group in the phosphonic amide molecule is involved in a short contact with the oxygen atom of the nitroxyl group of an adjacent molecule (N4–H4⋯O1ii, 2.66(2) Å, ii = x − 1/2, 1/2 − y, z − 1/2).
O axis of the ATEMPO unit acting as a hydrogen donor. Furthermore, the second N–H group in the phosphonic amide molecule is involved in a short contact with the oxygen atom of the nitroxyl group of an adjacent molecule (N4–H4⋯O1ii, 2.66(2) Å, ii = x − 1/2, 1/2 − y, z − 1/2).
Besides the aforementioned interchain interaction, there exists a short contact between the NO of the second ATEMPO unit in the phosphonic amide molecule and the meta CH of a nearby P–Ph moiety (C21–H21⋯O2ii distance of 2.531 Å).63 This network of intermolecular hydrogen bonds and C–H⋯O short contacts, together with other weak C⋯H and H⋯H interactions (average distance of 2.851 Å and 2.361 Å, respectively),64 generates a supramolecular multilayer structure. The shortest intrachain distance between the paramagnetic sites is 8.608(2) Å, while the closest separation among them is 4.981(2) Å. To the best of our knowledge, this is the first molecular characterisation of a phosphonic amide ATEMPO diradical.
In all complexes, coordination of the ligand to the metal ions takes place exclusively through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group. All attempts to coordinate the nitroxide moieties by increasing the amount of the metal or temperature and reaction times were unsuccessful.
O group. All attempts to coordinate the nitroxide moieties by increasing the amount of the metal or temperature and reaction times were unsuccessful.
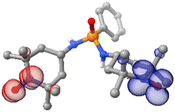
Compound 14 crystallises (method B) in the monoclinic C2/c space group and its molecular structure is shown in Fig. 3a and S15. The most representative distances and bond angles are recorded in Table 2. The asymmetric unit comprises one radical molecule 13 coordinated to a Cu(hfac)2 moiety through a phosphonic oxygen atom. The metal ion is pentacoordinated by four oxygen atoms from the hfac ligands (O4, O5, O6, and O7) and by the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O oxygen atom of ligand 13 (O3). In this coordination environment, the oxygen atoms of the hfac ligands define the basal plane, whereas the phosphine oxide oxygen atom occupies the apical position, resulting in a distorted square-pyramidal geometry. The calculated τ5 parameter is 0.18, consistent with this assignment (τ5 = 0 corresponds to an ideal square-pyramidal geometry). Both hfac ligands exhibit positional disorder; for simplicity, the corresponding Cu–O bond lengths are reported as average values in Table 2, and only one disorder component is depicted in Fig. 3a.
O oxygen atom of ligand 13 (O3). In this coordination environment, the oxygen atoms of the hfac ligands define the basal plane, whereas the phosphine oxide oxygen atom occupies the apical position, resulting in a distorted square-pyramidal geometry. The calculated τ5 parameter is 0.18, consistent with this assignment (τ5 = 0 corresponds to an ideal square-pyramidal geometry). Both hfac ligands exhibit positional disorder; for simplicity, the corresponding Cu–O bond lengths are reported as average values in Table 2, and only one disorder component is depicted in Fig. 3a.
Bond distances in the base range from 1.94(1)–1.965(2) Å and are slightly longer for Cu–O3 (2.118(2) Å), in agreement to other related systems described in the literature.65 P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond distance is 1.488(2) Å, very similar to the observed for the ligand (1.481(1) Å), indicating that coordination to the metal ion does not affect remarkably the P
O bond distance is 1.488(2) Å, very similar to the observed for the ligand (1.481(1) Å), indicating that coordination to the metal ion does not affect remarkably the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond. In this complex, both nitroxide moieties remain uncoordinated and their bond distances are similar to the free ligand and other TEMPO-derived compounds.66 The arrangement of the ATEMPOs is analogous to that found in the ligand, with a slight increase in the distance between them (distance O⋯O, 10.991(4) Å). A network of O⋯H–N hydrogen bonds (Fig. 4a, average bond distance 2.16 Å) and other C⋯F (3.080 Å), H⋯H (2.243 Å) and F⋯F (average bond distances 2.868 Å) weak interactions stabilizes a supramolecular zigzag chain (Fig. 4b).
O bond. In this complex, both nitroxide moieties remain uncoordinated and their bond distances are similar to the free ligand and other TEMPO-derived compounds.66 The arrangement of the ATEMPOs is analogous to that found in the ligand, with a slight increase in the distance between them (distance O⋯O, 10.991(4) Å). A network of O⋯H–N hydrogen bonds (Fig. 4a, average bond distance 2.16 Å) and other C⋯F (3.080 Å), H⋯H (2.243 Å) and F⋯F (average bond distances 2.868 Å) weak interactions stabilizes a supramolecular zigzag chain (Fig. 4b).
The shortest distance between paramagnetic sites is 6.320(4) Å, found for interchain nitroxide interactions, followed by 7.391(3) Å for Cu⋯O–N, and 14.595(1) Å between copper atoms. Compound 14 is the first example of a monophosphonic diamide dinitroxide copper(II) complex. Joesten and co-workers reported the crystal structure of a bis(phosphonic diamide)copper(II) perchlorate in which the Cu(II) ion is chelated by the two P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups of the ligand.67 Recently, manganese(II) complexes of the type [MnX2L2], where X = Cl, Br, I, and L is N,N,N′,N′-tetramethyl-P-indol-1-ylphosphonic diamide, have been described.68
O groups of the ligand.67 Recently, manganese(II) complexes of the type [MnX2L2], where X = Cl, Br, I, and L is N,N,N′,N′-tetramethyl-P-indol-1-ylphosphonic diamide, have been described.68
Compounds 15 and 16 crystallise in the Aba2 space group, while compound 17 crystallises in the monoclinic P21/n. The most representative distances and bond angles are recorded in Table S2. The three complexes presented the same molecular structure (Fig. 3b and c), which consists of an eight-coordinated Ln(III) atom bonded to six oxygen atoms from three hfac moieties and two units of 13 through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group arranged in a practically linear fashion. In compound 15, the Nd atom adopts a distorted D4d (square antiprism) geometry, while in complex 17 the geometry around the metal ion is defined as a biaugmented trigonal prism J50 (C2v).69 The Ln⋯Ohfac bond distances range from 2.39(6)–2.534(16) Å, 2.395(3)–2.461(10) Å, and 2.357(2)–2.441(2) Å in 15–17, respectively, being in the low-range limit compared to similar compounds.28,64,70 In all compounds, the P
O group arranged in a practically linear fashion. In compound 15, the Nd atom adopts a distorted D4d (square antiprism) geometry, while in complex 17 the geometry around the metal ion is defined as a biaugmented trigonal prism J50 (C2v).69 The Ln⋯Ohfac bond distances range from 2.39(6)–2.534(16) Å, 2.395(3)–2.461(10) Å, and 2.357(2)–2.441(2) Å in 15–17, respectively, being in the low-range limit compared to similar compounds.28,64,70 In all compounds, the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond distances are slightly longer than the free ligand due to coordination to the metal ion.
O bond distances are slightly longer than the free ligand due to coordination to the metal ion.
The Ln–O–P bond angles are nearly identical for 15 and 16, with values of 156.4(2)° and 155.84(14)°, respectively, while the analogous bond angles in terbium derivative 17 are markedly larger, reaching 161.96(13)° and 165.36(14)°. The shortest intermolecular Ln⋯Ln distances are 13.1274(2) Å for 15, 13.1083(2) Å for 16 and 9.9890(7) Å in compound 17. The crystal packing in compounds 15–17 is stabilized by a series of short contacts, including C(sp2)–H⋯O, C(sp3)–H⋯C(sp2), and F⋯F interactions, as well as by a hydrogen-bonding network involving nitroxide moieties and NH groups from the phosphonic amide functionalities (Fig. 5). The N–O⋯H–N hydrogen bond distances range from 2.01(2) to 2.16(8) Å (see Table S2).
A limited number of complexes of the phosphonic diamides L = tBuPO(NHiPr)2 and (Ad)PO(NHiPr)2 (Ad = adamantyl) with lanthanide ions of formula [L2Ln(H2O)5][X]3·L2 (Sol), where Ln = Nd(III),71 Dy(III), Er(III),72 Tb(III), Ho(III);73 X = Cl, Br, I; and Sol = H2O, CH2Cl2, have been recently reported. All complexes contain a seven-coordinated metal cation, with the two phosphonic diamide ligands coordinated to the Ln(III) ion through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group, positioned in a diaxial geometry (OP–Ln–OP bond angles in the range of 172.19° to 177.66°).
O group, positioned in a diaxial geometry (OP–Ln–OP bond angles in the range of 172.19° to 177.66°).
Interestingly, these compounds behaved as single-ion magnets (SIM). The axial arrangement of the phosphorus-based ligand and the symmetry around the Ln(III) ions were identified as the primary contributors to their magnetic properties, with a minor modulation attributed to secondary coordination sphere anions.74 In addition, [L3Ln(NO3)3] complexes of tBuPO(NHiPr)2 with nine-coordinated Ln = Ce(III) and La(III) ions were also described.75
Ligand coordination analysis
As previously mentioned, the phosphinic amide TEMPO radical 6 (Fig. 2) is capable of acting as either a mono- or bidentate ligand through metal coordination involving the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and N–O· groups. However, the results obtained demonstrate that the tridentate phenylphosphonic diamide bis-TEMPO 13 binds exclusively through the P
O and N–O· groups. However, the results obtained demonstrate that the tridentate phenylphosphonic diamide bis-TEMPO 13 binds exclusively through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group, even in the presence of an excess of metal cation. This behavior mirrors that observed in the case of the tetradentate ligands 1 and 5 (Fig. 1), in which metal coordination occurs through the nitrogen atoms of the linker between the TEMPO radicals or the nitroxide donor sites, respectively. In order to understand the role played by the substituent on the C4 position of TEMPO radicals on their coordination properties,15b,25 a DFT study of the different coordination modes of ligands 1, 5, 6 and 13 was undertaken.
O group, even in the presence of an excess of metal cation. This behavior mirrors that observed in the case of the tetradentate ligands 1 and 5 (Fig. 1), in which metal coordination occurs through the nitrogen atoms of the linker between the TEMPO radicals or the nitroxide donor sites, respectively. In order to understand the role played by the substituent on the C4 position of TEMPO radicals on their coordination properties,15b,25 a DFT study of the different coordination modes of ligands 1, 5, 6 and 13 was undertaken.
Ligand 1 provides the electronic baseline. The bridging amine nitrogens are the strongest donors, with natural charges around −0.60 and lone pairs that are not significantly involved in hyperconjugative delocalization. The two TEMPO moieties show the usual radical signature: moderate negative charge at oxygen (≈−0.42), substantial spin shared between N and O (Table S3), and O-lone-pair donor–acceptor stabilization energies of ≈7–8 kcal mol−1 (Table S4c). Thus the nitroxide groups in ligand 1 are electronically good donors. They do not coordinate because the corresponding approach vector is sterically expensive: the TEMPO ring shields the oxygen along the N–O axis, and the NLMO-resolved steric analysis shows a dense network of local Pauli repulsions between the nitroxide lone pairs/bonds and the adjacent backbone. Several pairwise disjoint steric terms reach 5–12 kcal mol−1 for motions that bring a metal toward O(TEMPO), and the local dEi values in the nitroxide region are high (≈55–60 kcal mol−1). In contrast, approach vectors toward the amine lone pairs are less congested, so ligand 1 naturally binds through its NH donors even though its global steric budget is lower than that of ligands 5 and 6.
The ligand 5 provides a useful extension of the analysis because it contains two nitroxide units, as in ligand 1, but incorporates a piperazine ring with two closed-shell nitrogen donors. Although these piperazine nitrogens carry significant negative charge (≈−0.44), the NBO results show that their lone pairs are strongly delocalized within the local σ-framework, as reflected in the relatively large donor–acceptor stabilization energies associated with these sites (≈28 kcal mol−1; Table S4b). These values are the highest among all potential donors in ligand 5 and indicate that the piperazine lone pairs are electronically committed to internal hyperconjugation rather than external coordination. The NLMO-resolved steric analysis further shows that the local steric exchange around the piperazine nitrogens is the most congested region of the molecule, with spin-averaged steric contributions of ≈85 kcal mol−1 (Table S5), nearly a factor of two greater than those associated with approach vectors toward the nitroxide oxygens. By contrast, the nitroxide groups retain the same electronic features observed in ligand 1, with moderate negative charge at the oxygens (≈−0.42), substantial N/O spin sharing, and donor–acceptor energies in the usual 7–8 kcal mol−1 range (Table S4c). Importantly, the steric cost for approaching either nitroxide oxygens in ligand 5 is markedly lower (≈38 kcal mol−1; Table S5) than that of the piperazine nitrogens. As a result, ligand 5 binds exclusively through its two nitroxide oxygens: the piperazine nitrogen atoms, despite their formal basicity, are both electronically less available and sterically inaccessible for coordination.
Introducing a phosphoryl group in ligand ppdATEMPO (13) reorganizes the donor hierarchy. The phosphoryl oxygen becomes the strongest basic site, with natural charge near −1.10 and the largest donor-acceptor stabilization energies in the system (≈17–18 kcal mol−1; Table S4a), arising from efficient lone-pair donation from the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O oxygen into the adjacent P–N and P–C framework. The amide nitrogens are electronically deactivated toward external binding by N → P π-donation, consistent with their charges (≈ −1.00) and P–N delocalization. In contrast, the nitroxide groups in 13 remain essentially indistinguishable from those in 1, 5 and 6: their charges, spin densities and lone-pair E(2) values exhibit the same 7–8 kcal mol−1 interval (Tables S3 and S4c). The absence of nitroxide coordination in 13 is therefore not an electronic effect.
O oxygen into the adjacent P–N and P–C framework. The amide nitrogens are electronically deactivated toward external binding by N → P π-donation, consistent with their charges (≈ −1.00) and P–N delocalization. In contrast, the nitroxide groups in 13 remain essentially indistinguishable from those in 1, 5 and 6: their charges, spin densities and lone-pair E(2) values exhibit the same 7–8 kcal mol−1 interval (Tables S3 and S4c). The absence of nitroxide coordination in 13 is therefore not an electronic effect.
Steric accessibility becomes decisive once two TEMPO units flank the phosphorus center. The spin-averaged total steric exchange rises from ≈1629 kcal mol−1 in ligand 1 to ≈1747 kcal mol−1 in ligand 6 and ≈2035 kcal mol−1 in ligand 13 (Table S6), reflecting the progressive increase in steric congestion. In 13, the two TEMPO units create a rigid steric enclosure: the phosphoryl oxygen points along a relatively open vector, whereas both nitroxide oxygens lie in bulky TEMPO rings. The local steric terms around the nitroxide sites are larger than in 6, and the incremental steric penalty required to place a metal ion in the nitroxide coordination region is prohibitive. As a result, only the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O site is sterically accessible for binding.
O site is sterically accessible for binding.
The comparison with ligand 6 clarifies the role of the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O anchor. Ligand 6 carries a single TEMPO group and presents a less crowded environment around phosphorus. Coordination at the phosphoryl oxygen is favored because it is both, the electronically dominant donor, and lies along an accessible approach vector. Once the metal ion is bound at the P
O anchor. Ligand 6 carries a single TEMPO group and presents a less crowded environment around phosphorus. Coordination at the phosphoryl oxygen is favored because it is both, the electronically dominant donor, and lies along an accessible approach vector. Once the metal ion is bound at the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O site, only a modest additional displacement is required to bring the nitroxide oxygen into a geometry where interaction becomes feasible. The incremental steric penalty associated with this secondary motion is relatively small: the key pairwise steric exchanges along the nitroxide arm in 6 are on the order of 1–3 kcal mol−1, and the local dEi values, although characteristic of a TEMPO fragment, are markedly lower than those encountered when attempting first-contact coordination at the nitroxide of ligand 1. As a result, ligand 6 can engage the metal ion at both the phosphoryl and nitroxide sites because the metal ion approaches the TEMPO unit from a pre-anchored, low-steric pathway rather than attempting to enter the TEMPO region directly.
O site, only a modest additional displacement is required to bring the nitroxide oxygen into a geometry where interaction becomes feasible. The incremental steric penalty associated with this secondary motion is relatively small: the key pairwise steric exchanges along the nitroxide arm in 6 are on the order of 1–3 kcal mol−1, and the local dEi values, although characteristic of a TEMPO fragment, are markedly lower than those encountered when attempting first-contact coordination at the nitroxide of ligand 1. As a result, ligand 6 can engage the metal ion at both the phosphoryl and nitroxide sites because the metal ion approaches the TEMPO unit from a pre-anchored, low-steric pathway rather than attempting to enter the TEMPO region directly.
Broken-symmetry calculations further support the independence of the TEMPO units. The weak antiferromagnetic coupling observed for ligand 13 (J ≈ −0.35 cm−1) and the large spatial separation between the two N–O units (>8 Å) are consistent with electronically intact radicals that remain magnetically isolated (Fig. 6). The coordination through P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O does not perturb the electronic structure of the nitroxide moieties, reinforcing the conclusion that their non-coordination in 13 is steric rather than electronic.
O does not perturb the electronic structure of the nitroxide moieties, reinforcing the conclusion that their non-coordination in 13 is steric rather than electronic.
Magnetic properties
Lanthanide-based monometallic complexes hold considerable potential as nanomagnetic materials due to the strong magnetic anisotropy of the lanthanide ions.76 The crystal-field splitting and coordination geometry have proven to be pivotal in determining their magnetic behaviour. It has been demonstrated that these effects combine favourably in complexes of bulky phosphonic diamides with rare-earth ions leading to promising high-performance Ln(III)-SIMs.72–76The magnetic properties of compounds 14–17 were investigated between 2 K and 300 K. Fig. 7 represents the thermal dependence of χMT, the product of the molar magnetic susceptibility by the temperature, for all compounds. At room temperature, the values of χMT are 1.2 (14), 3.0 (15), 2.8 (16) and 12.5 (17) cm3 mol−1 K.
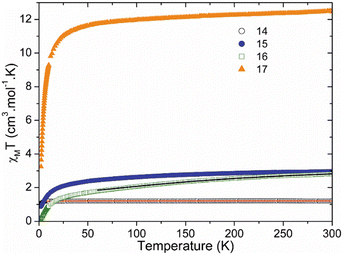 | ||
| Fig. 7 Thermal dependence of the χMT product for compounds 14–17. The solid lines (red and black) represent the best fits (see SI text for details). | ||
For compound 14, this value is close to the expected one for three uncoupled S = 1/2 (1.13 cm3 mol−1 K). On lowering temperature, χMT remains nearly constant down to 8 K, below which it decreases to 1.07 cm3 mol−1 K, indicating the presence of antiferromagnetic interactions. The reciprocal susceptibility, plotted against temperature, can be modeled by the Curie–Weiss law (Fig. S13), leading to C = 1.21 cm3 mol−1 K and θ = −0.23 K, the respective Curie and Weiss constants. The antiferromagnetic interactions in compound 14 are very weak. Analysis of the crystal structure shows that the radicals are relatively far apart from each other and from the metal ion. Because there is no direct coordination between the CuII ion and the ATEMPO radicals through the N–O moiety, and because the piperidinyl ring does not provide an efficient pathway for magnetic exchange, the magnetic interactions between the organic radicals and the metal center are expected to be negligible.77 This expectation is corroborated by the ab initio calculations (see above). Therefore, in order to reproduce the magnetic data, we only consider intermolecular interactions through a mean field approximation.
The best fit parameters for 14 are grad = 2 (fixed), gCu = 2.230(5) and zJ′ = −0.18(1) cm−1. As expected from the structure, the intermolecular interaction is weak and antiferromagnetic. This interaction may be carried out through the network of hydrogen bonds involving the TEMPO oxygen atoms and the amide group of neighboring molecules (N3–H3⋯O1i and N4–H4⋯O2ii).
For compounds 15 and 17, the χMT values at room temperature are close to those expected for four uncoupled radicals and Ln(III) free ions (3.1 and 13.3 cm3 mol−1 K, respectively). In contrast, for the Eu derivative 16, this value is significantly higher due to the thermal population of excited states of Eu3+ ion at room temperature.78 Upon cooling, compounds 15–17 display a similar trend, with χMT gradually decreasing and exhibiting a more pronounced drop at the lowest temperatures. Similarly to compound 14, this behavior at low-temperature likely arises from antiferromagnetic intermolecular interactions. Furthermore, for 15 and 17, the gradual decrease of χMT is attributed to the depopulation of the MJ sublevels of the lanthanide free ion ground state. In the case of compound 16, the continuous decrease results from the depopulation of the low-lying spin–orbit coupled states.79 Accordingly, the magnetic data of compound 16 were analyzed in the high temperature region (>60 K), taking into account the contributions from four uncoupled radicals and from the spin-orbit coupling of the europium ion (see SI). The best fit, shown in Fig. 7, was obtained using grad = 2 (fixed) and λ = 360(2) cm−1, in good agreement with the literature.79
Catalysis
The development of efficient oxidation systems by using low hazard and economic catalysts and oxidants under mild conditions, has turned into an important aim for catalytic-process design. Cu-TEMPO mixtures have emerged as one of the most effective catalysts for selective alcohol oxidation. However, there is still numerous challenges in the search of practicality. This includes mild reaction conditions, low pressures of O2, low catalyst loading, low temperatures, and the avoidance of toxic additives.1b,c,40,80Taking into account the above considerations, we envisioned the use of the copper-diradical complex 14 for the aerobic oxidation of alcohols. For that purpose, the reaction of 4-nitrobenzyl alcohol 18a was used as the model to optimize the reaction conditions (Table 3). Unless otherwise noted, all reactions were carried out under an air atmosphere. The completion of the reaction was established by TLC.
| Entry | Catalyst | Solvent | T (°C) | Time (h) | Yielda |
|---|---|---|---|---|---|
| a NMR yield determined through integration of the 1H NMR spectrum of the reaction crude.b ACN: acetonitrile.c DCE: dichloroethane.d Isolated yield.e Under an O2 atmosphere. | |||||
| 1 | 14 | ACNb | 25 | 72 | 32 |
| 2 | 14 | DCEc | 25 | 22 | 91(90)d |
| 3 | 14 | DCE | 25 | 72 | 100 |
| 4 | 14 | DCE | 50 | 7 | 81 |
| 5 | 14 | DCE | 50 | 30 | 100 |
| 6e | 14 | DCE | 25 | 8 | 69 |
| 7 | 13 + Cu(hfac)2 | DCE | 25 | 22 | 90 |
| 8 | 13 | DCE | 25 | 22 | 0 |
| 9 | Cu(hfac)2 | DCE | 25 | 22 | 0 |
First, reaction of 18a in the presence of 10% of 14 in acetonitrile, afforded benzaldehyde 19 in only 32% yield after 72 hours, probably due to the moderate solubility of the catalyst (entry 1). Change of the solvent to DCE was crucial, increasing the yield of the reaction to 91% in 22 h (entry 2). However, the reaction did not reach completion until 72 h (entry 3). Increasing the temperature to 50 °C did not lead to a remarkable improvement, as shown by TLC. After 7 h, yield was 81% (entry 4). The reaction was completed within 30 h (entry 5). Similar results were observed by increasing the concentration of oxygen with an O2 balloon, affording aldehyde 19 in 69% yield after 8 h (entry 6). To our delight, the reaction can be carried out by the in situ formation of 14 from diradical 13 and Cu(hfac)2, increasing the atom efficiency of the transformation. In these conditions, 19 was formed in an excellent 90% yield after 22 h (entry 7). As expected, control experiments using only the TEMPO diradical (entry 8) or the copper salt (entry 9) did not promote the oxidation of benzyl alcohol 18a. In all cases, no over oxidation to carboxylic acids was observed.
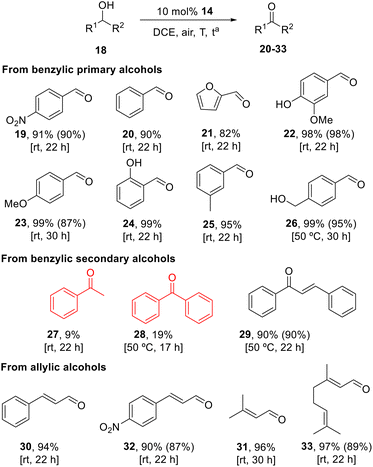
Based on the results in Table 3, entry 2 was selected as the optimal conditions to explore the scope of the reaction (Scheme 3). The oxidation of benzyl alcohol to give 20 took place after 22 h in 90% yield. The yield of 21 obtained in the reaction of furfuryl alcohol was marginally lower (82%). The most favourable outcomes were observed when electron-donating substituted phenyl groups such as methyl, hydroxy and methoxy substituents in ortho, meta and para positions, were utilised. The expected benzaldehydes 22–25 were formed in excellent yields of up to 99% at room temperature. The double oxidation of diol 1,4-phenylenedimethanol necessitated an increase in the temperature to 50 °C. In the case of the secondary benzyl alcohols 1-phenylethan-1-ol and diphenylmetanol, substandard results were obtained. The respective ketones were produced in yields lower than 20%, even after the reaction temperature was increased to 50 °C. The exception to this observation is phenylcinnamyl alcohol, in which the hydroxyl group occupies both an allylic and benzylic position. The oxidation process at 50 °C for a duration of 22 hours resulted in the synthesis of (E)-chalcone 29 in 90% yield.
We were also pleased to observe that this protocol may be used in the oxidation of allylic alcohols. The application of the standard oxidation procedure to cinnamyl alcohol, 4-nitrocinnamyl alcohol, prenol and geraniol, gave rise to the respective aldehydes 30–33 in excellent yields. Besides this fact, isomerization of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond during the course of the reactions was very limited, indicating that the oxidation of allylic primary alcohols using 14 as catalyst is very selective.
C bond during the course of the reactions was very limited, indicating that the oxidation of allylic primary alcohols using 14 as catalyst is very selective.
The aerobic oxidation of alcohols catalysed by 14 shows comparable performance compared to analogous procedures using catalytic systems based on mononuclear Cu(II) complexes as catalyst and mediated by TEMPO. For example, the square-pyramidal Cu(II) complexes with tridentate NNS, N(NH)S and NNO ligands reported by Bagh and coworkers,81 and those contain tetradentate N2O2 bis(phenol) diamine ligands in the presence of Cs2CO3,82 show similar results. Moreover, the performance of 14 is superior to that of the (Cu(tpy-TEMPO)Cl2·2DMF) complex (tpy = 4-(2,2′:6′,2″-terpyridin-4′-yloxy)) in combination with DBU (DBU = 1,8-diazabicyclo[5.4.0]undec-7-ene).83
However, it requires longer reaction times than some of the most efficient Cu(I) and Cu(II)-(bpy)-TEMPO (bpy = 2,2′-bipyridine) catalytic systems. The oxidation of allylic and benzylic alcohols using the catalysts developed by Stahl84 (Cu(I)(OTf)/bipy/TEMPO with NMI as a base in acetonitrile) (OTf = trifluoromethanesulfonate, NMI = N-methylimidazole) and Sheldon85 (Cu(II)(OTf)2/bipy/TEMPO with KOtBu as a base in acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water 2
water 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) are typically completed in less than 5 hours of reaction. It is important to note, that complex 14 contains the metal cation and the nitroxyl radical as ligand and can be readily prepared following a straightforward protocol using commercial reagents. Furthermore, the use of supplementary reagents such as a base (NMI, DBU, etc.) is unnecessary.
1) are typically completed in less than 5 hours of reaction. It is important to note, that complex 14 contains the metal cation and the nitroxyl radical as ligand and can be readily prepared following a straightforward protocol using commercial reagents. Furthermore, the use of supplementary reagents such as a base (NMI, DBU, etc.) is unnecessary.
Conclusions
The synthesis and molecular characterisation of the first example of a new phosphonic diamide-TEMPO diradical (ppdATEMPO), and of its coordination compounds with CuII, NdIII, EuIII and TbIII ions, have been described. The solid-state structure of the diradical was found to exhibit a quasi-perpendicular arrangement of the ATEMPO units and an architecture showing a 3D network formed via P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O⋯H–N, N–O⋯H–N(P) and N–O⋯H–C inter-chain hydrogen bonding. The X-ray diffraction analyses of the metal complexes demonstrated distinctive structural features. In contrast to the structurally related phosphinic amide TEMPO mononitroxyl derivative, the diradical acts as a monodentate ligand exclusively through the P
O⋯H–N, N–O⋯H–N(P) and N–O⋯H–C inter-chain hydrogen bonding. The X-ray diffraction analyses of the metal complexes demonstrated distinctive structural features. In contrast to the structurally related phosphinic amide TEMPO mononitroxyl derivative, the diradical acts as a monodentate ligand exclusively through the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group in all cases. This results in five-(CuII) and eight-coordinated (LnIII) mononuclear complexes, which show discrete molecules organised in 2D supramolecular layers or chains through intermolecular interactions. The different coordination modes of ppdATEMPO (13) vs. dppnTEMPO (6) reflect how donor strength and steric accessibility interplay around the key donor sites. The magnetic measurements revealed weak antiferromagnetic interactions in complex 14, which were rationalized as intermolecular in nature and are likely present in all compounds. In addition, the copper(II) complex 14 proved to be a good catalyst in the aerobic oxidation of a wide scope of allylic and benzylic primary alcohols. The expected aldehydes were formed in excellent yields and selectivities in mild reaction conditions under air. The formation of ketones in high yields was only possible when the hydroxyl group underwent doubly activation, with occupation of both an allylic and a benzyl position.
O group in all cases. This results in five-(CuII) and eight-coordinated (LnIII) mononuclear complexes, which show discrete molecules organised in 2D supramolecular layers or chains through intermolecular interactions. The different coordination modes of ppdATEMPO (13) vs. dppnTEMPO (6) reflect how donor strength and steric accessibility interplay around the key donor sites. The magnetic measurements revealed weak antiferromagnetic interactions in complex 14, which were rationalized as intermolecular in nature and are likely present in all compounds. In addition, the copper(II) complex 14 proved to be a good catalyst in the aerobic oxidation of a wide scope of allylic and benzylic primary alcohols. The expected aldehydes were formed in excellent yields and selectivities in mild reaction conditions under air. The formation of ketones in high yields was only possible when the hydroxyl group underwent doubly activation, with occupation of both an allylic and a benzyl position.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI).Supplementary information is available. See DOI: https://doi.org/10.1039/d5ce01150g.
CCDC 2501504–2501508 (13–17) contain the supplementary crystallographic data for this paper.86a–e
Acknowledgements
Financial support from Ministerio de Economía y Competitividad (MINECO), including the European Regional Development Fund (ERDF) program (projects CTQ2014-57157-P, CTQ2016-75671-P), and the Research and Transfer Plan of the University of Almeria, funded by “Consejería de Universidad, Investigación e Innovación de la Junta de Andalucía” (ERDF 2021-2027. Objective RSO1.1. Programme: 54.A; project PPUENTE2020/007) is gratefully acknowledged. GPG acknowledges the financial support from Fundação Carlos Chagas de Amparo à Pesquisa do Estado do Rio de Janeiro – FAPERJ (project numbers E-26/202.720/2018 and E-26/201.314/2022) and Conselho Nacional de Desenvolvimento Científico e Tecnológico – CNPq (project numbers 304671/2020-7 and 307135/2023-3). Theoretical calculations were performed using the Lobo Carneiro supercomputer from Núcleo Avançado de Computação de Alto Desemprenho (NACAD), under Project ID a20006. The authors would also like to thank the National Laboratory for Scientific Computing (LNCC/MCTI, Brazil) for providing HPC resources of the SDumont supercomputer, which have contributed to the research results reported within this paper. URL: https://sdumont.lncc.br. We are grateful to the Low Temperature and Magnetometry Service of the Institute of Materials Science of Barcelona (ICMAB) for carrying out the magnetism measurements.Notes and references
- (a) H. A. Beejapur, Q. Zhang, K. Hu, L. Zhu, J. Wang and Z. Ye, ACS Catal., 2019, 9, 2777 CrossRef CAS; (b) S. Nagasawa, Y. Sasano and Y. Iwabuchi, Heterocycles, 2022, 105, 61 CrossRef CAS PubMed; (c) D. Leifert and A. Studer, Chem. Rev., 2023, 123, 10302 CrossRef CAS PubMed.
- (a) F. Hyodo, H. Eto, T. Naganuma, N. Koyasu, A. E. Elhelaly, Y. Noda, H. Kato, M. Murata, T. Akahoshi, M. Hashizume, H. Utsumi and M. Matsuo, Antioxid. Redox Signaling, 2022, 36, 172 CrossRef CAS PubMed; (b) K. Gwozdzinski, A. Pieniazek and L. Gwozdzinski, Molecules, 2025, 30, 2159 CrossRef CAS PubMed.
- (a) Y. Xie, K. Zhang, Y. Yamauchi and Z. Jia, Mater. Horiz., 2021, 8, 803 RSC; (b) A. A. Vereshchagin, A. Y. Kalnin, A. I. Volkov, D. A. Lukyanov and O. V. Levin, Energies, 2022, 15, 2699 CrossRef CAS; (c) C. M. Davis, C. E. Boronski, T. Yang, T. Liu and Z. Liang, Batteries, 2023, 9, 504 CrossRef CAS.
- (a) G. Audran, E. G. Bagryanskaya, S. R. A. Marque and P. Postnikov, Polymer, 2020, 12, 1481 CrossRef CAS PubMed; (b) H. R. Lamontagne and B. H. Lessard, ACS Appl. Polym. Mater., 2020, 2, 5327 CrossRef CAS.
- (a) Z. Zhou and L. Liu, Curr. Org. Chem., 2014, 18, 459 CrossRef CAS; (b) J. A. Zarling, V. E. Brunt, A. K. Vallerga, W. Li, A. Tao, D. A. Zarling and C. T. Minson, Front. Genet., 2015, 6, 325 Search PubMed; (c) B. Bognár, G. Úr, C. Sár, O. H. Hankovszky, K. Hideg and T. Kálai, Curr. Org. Chem., 2019, 23, 480 CrossRef; (d) B. Chen, E. Pentzer, S. Mitchell, N. Sinclair and B. Gurkan, Mol. Syst. Des. Eng., 2020, 5, 1147 RSC; (e) Z. Wang, Q. Wang, X. Liao, H. Han, R. Sun and M. Xie, Dyes Pigm., 2024, 228, 112236 CrossRef CAS; (f) B. Dutta, N. Adhikari, R. P. Borah, B. Bhattacharyya and D. Sarma, J. Mol. Struct., 2025, 1322, 140198 CrossRef CAS.
- (a) M. Abe, Chem. Rev., 2013, 113, 7011 CrossRef CAS PubMed; (b) J. J. Dressler and M. M. Haley, J. Phys. Org. Chem., 2020, 33, e4114 CrossRef CAS; (c) Z. Yang Shu and A. Rajca, Chem. Rev., 2023, 123, 11954 CrossRef PubMed; (d) Y. Zhu, Z. Zhu, S. Wang, Q. Peng and A. Abdurahman, Angew. Chem., Int. Ed., 2025, 64, e202423470 CrossRef CAS PubMed.
- (a) D. Cho, K. C. Ko and J. Y. Lee, Int. J. Quantum Chem., 2016, 116, 578 CrossRef CAS; (b) V. Lloveras, E. Badetti, K. Wurst, V. Chechik, J. Veciana and J. Vidal-Gancedo, Chem. – Eur. J., 2016, 22, 1805 CrossRef CAS PubMed; (c) J. Tuček, K. Holá, A. B. Bourlinos, P. Błoński, A. Bakandritsos, J. Ugolotti, M. Dubecký, F. Karlický, V. Ranc, K. Čépe, M. Otyepka and R. Zbořil, Nat. Commun., 2017, 8, 14525 CrossRef PubMed; (d) S. Yu, Y. Song, Y. Bu and X. Song, New J. Chem., 2023, 47, 19243 RSC; (e) Q. Jiang, L. Wang, H. Wei, Y. Peng, G. Xu, Z. Li, P. Liu, Z. Hu, W. Niu, Y. Chen, H. Tang, W. Zeng and G. Li, Angew. Chem., Int. Ed., 2025, 64, e202422994 CrossRef CAS PubMed.
- (a) Z. Zeng, X. Shi, C. Chi, J. T. López-Navarrete, J. Casado and J. Wu, Chem. Soc. Rev., 2015, 44, 6578 RSC; (b) X. Hu, W. Wang, D. Wang and Y. Zheng, J. Mater. Chem. C, 2018, 6, 11232 RSC; (c) Y. Lang, Y. Hei, T. Yu, Y. Yang, M. Chen, J. Zhang, F.-G. Zhao, Y. Zheng and X. Liu, Chem. – Eur. J., 2024, 30, e202302943 CrossRef PubMed; (d) R. Casares, S. Rodríguez-González, Á. Martínez-Pinel, I. R. Márquez, M. T. González, C. Díaz, F. Martín, J. M. Cuerva, E. Leary and A. Millán, J. Am. Chem. Soc., 2024, 146, 29977 CrossRef CAS PubMed.
- (a) D. Bhattacharya, S. Shil, A. Misra, L. Bytautas and D. J. Klein, Int. J. Quantum Chem., 2015, 115, 1561 CrossRef CAS; (b) R. Bano, M. Asghar, K. Ayub, T. Mahmood, J. Iqbal, S. Tabassum, R. Zakaria and M. A. Gilani, Front. Mater., 2021, 8, 783239 CrossRef; (c) Z. X. Chen, Y. Li and F. Huang, Chem, 2021, 7, 288 CrossRef CAS; (d) Y. R. Poh, D. Morozov, N. P. Kazmierczak, R. G. Hadt, G. Groenhof and J. Yuen-Zhou, J. Am. Chem. Soc., 2024, 146, 15549 CrossRef CAS PubMed.
- (a) K. Fujikura, H. Akutsu, J.-I. Yamada, S. Nakatsuji and M. Satoh, Synth. Met., 2015, 208, 17 CrossRef CAS; (b) K. Kim and J.-K. Kim, Korean J. Chem. Eng., 2016, 33, 858 Search PubMed; (c) J. Sun, E. Zhao, J. Liang, H. Li, S. Zhao, G. Wang, X. Gu and B. Z. Tang, Adv. Mater., 2022, 34, 2108048 Search PubMed; (d) S. Fang, J. Huang, R. Tao, Q. Wei, X. Ding, S. Yajima, Z. Chen, W. Zhu, C. Liu, Y. Li, N. Yin, L. Song, Y. Liu, G. Shi, H. Wu, Y. Gao, X. Wen, Q. Chen, Q. Shen, Y. Li, Z. Liu, Y. Li and W. Ma, Adv. Mater., 2023, 35, 2212184 Search PubMed.
- (a) G. Audran, E. G. Bagryanskaya, S. R. A. Marque and P. Postnikov, Polymer, 2020, 12, 1481 CrossRef CAS PubMed; (b) H. R. Lamontagne and B. H. Lessard, ACS Appl. Polym. Mater., 2020, 2, 5327 CrossRef CAS.
- (a) K.-I. Matsumoto, J. B. Mitchell and M. C. Krishna, Molecules, 2021, 26, 1614 Search PubMed; (b) K.-I. Matsumoto, I. Nakanishi, Z. Zhelev, R. Bakalova and I. Aoki, Antioxid. Redox Signaling, 2022, 36, 95 CrossRef CAS PubMed; (c) T. Luo, B. Wang, R. Chen, Q. Qi, R. Wu, S. Xie, H. Chen, J. Han, D. Wu and S. Cao, J. Mater. Chem. B, 2025, 13, 372 RSC.
- (a) D. J. Kubicki, G. Casano, M. Schwarzwälder, S. Abel, C. Sauvée, K. Ganesan, M. Yulikov, A. J. Rossini, G. Jeschke, C. Copéret, A. Lesage, P. Tordo, O. Ouari and L. Emsley, Chem. Sci., 2016, 7, 550 RSC; (b) E. V. Zaytseva and D. G. Mazhukin, Molecules, 2021, 26, 677 CrossRef CAS PubMed; (c) B. E. Ackermann, B. J. Lim, N. Elathram, S. Narayanan and G. T. Debelouchina, ChemBioChem, 2022, 23, e202200577 Search PubMed; (d) W. Y. Chow, G. De Paëpe and S. Hediger, Chem. Rev., 2022, 122, 9795 CrossRef CAS PubMed; (e) D. Dai, Y. Liu, X. He and J. Mao, Magn. Reson. Lett., 2025, 5, 200178 CrossRef CAS PubMed.
- (a) E. A. Legenzov, S. J. Sims, N. D. A. Dirda, G. M. Rosen and J. P. Y. Kao, Biochemistry, 2015, 54, 6973 CrossRef CAS PubMed; (b) P. K. Pachpatil, S. V. Kanojia, A. Ghosh, A. G. Majumdar, A. Wadawale, M. Mohapatra, B. S. Patro, T. K. Ghanty and D. Goswami, Sens. Actuators, B, 2022, 370, 132474 CrossRef CAS.
- (a) E. Damiani, R. Castagna, P. Astolfi and L. Greci, Free Radical Res., 2005, 39, 325 CrossRef CAS PubMed; (b) M. Kavala, V. Brezová, Ĺ. Švorc, Z. Vihonská, P. Olejníková, J. Moncol, J. Kožíšek, P. Herich and P. Szolcsányi, Org. Biomol. Chem., 2014, 12, 4491 RSC; (c) V. Venkatesh, R. Berrocal-Martin, C. J. Wedge, I. Romero-Canelón, C. Sánchez-Cano, J.-I. Song, J. P. C. Coverdale, P. Zhang, G. J. Clarkson, A. Habtemariam, S. W. Magennis, R. J. Deeth and P. J. Sadler, Chem. Sci., 2017, 8, 8271 RSC; (d) P. Poprac, P. Poliak, M. Kavala, Z. Barbieriková, M. Zalibera, M. Fronc, Ľ. Švorc, Z. Vihonská, P. Olejníková, K. Lušpai, V. Lukeš, V. Brezová and P. Szolcsányi, ChemPlusChem, 2017, 82, 1326 CrossRef CAS PubMed; (e) A. D. Verderosa, J. Harris, R. Dhouib, M. Totsika and K. E. Fairfull-Smith, Med. Chem. Commun., 2019, 10, 699 RSC.
- D. Cho, C. K. Ko and J. Y. Lee, J. Phys. Chem. A, 2014, 118, 5112 CrossRef CAS PubMed.
- G. Gagnaire, A. Jeunet and J.-L. Pierre, Tetrahedron Lett., 1989, 30, 6507 CrossRef CAS.
- H. Kinoshita, H. Akutsu, J. Yamada and S. Nakatsuji, Inorg. Chim. Acta, 2008, 361, 4159 CrossRef CAS.
- X. Lu, N. Wu, B. Zhang and K. Deng, J. Coord. Chem., 2017, 70, 475 CrossRef CAS.
- T. Katayama, T. Ishida and T. Nogami, Inorg. Chim. Acta, 2002, 329, 31 CrossRef CAS.
- N. Cosquer, E. Lefebvre, B. Douziech, S. Houille, F. Michaud, C. J. Gómez-García and F. Conan, Inorg. Chim. Acta, 2018, 479, 1 Search PubMed.
- C. Sommer, W. Ponikwar, W. Ponikwar, H. Nöth and W. Beck, Z. Anorg. Allg. Chem., 2003, 629, 2291 Search PubMed.
- S. Yi, B. Captain, M. F. Ottaviani and A. E. Kaifer, Langmuir, 2011, 27, 5624 CrossRef CAS PubMed.
- A. B. Burdukov, E. G. Boguslavskii, V. A. Reznikov, N. V. Pervukhina, M. A. Vershinin, Y. Z. Voloshin, O. A. Varzatskii and Y. N. Bubnov, Russ. Chem. Bull., 2005, 54, 1125 CrossRef CAS.
- (a) H. Moons, E. Goovaerts, V. P. Gubskaya, I. A. Nuretdinov, C. Corvaja and L. Franco, Phys. Chem. Chem. Phys., 2011, 13, 3942 RSC; (b) M. Kavala, R. Boča, Ĺ. Dlháň, V. Brezová, M. Breza, J. Kožíšek, M. Fronc, P. Herich, Ĺ. Švorc and P. Szolcsányi, J. Org. Chem., 2013, 78, 6558 CrossRef CAS PubMed; (c) H. Fan, J. Zhang, M. Ravivarma, H. Li, B. Hu, J. Lei, Y. Feng, S. Xiong, C. He, J. Gong, T. Gao and J. Song, ACS Appl. Mater. Interfaces, 2020, 12, 43568 CrossRef CAS PubMed; (d) R. Azuma, T. Yamasaki, M. C. Emoto, H. Sato-Akaba, K. Sano, M. Munekane, H. G. Fujii and T. Mukai, Free Radical Biol. Med., 2023, 194, 114 CrossRef CAS PubMed; (e) A. Shiroudi, M. Śmiechowski, J. Czub and M. A. Abdel-Rahman, Sci. Rep., 2024, 14, 8434 CrossRef CAS PubMed.
- (a) A. B. Shapiro, L. S. Bogach, V. M. Chumakov, A. A. Kropacheva, V. I. Suskina and É. G. Rozantsev, Russ. Chem. Bull., 1975, 24, 1959 CrossRef; (b) F.-P. Tsui, F. A. Robey, T. W. Engle, S. M. Ludeman and G. Zon, J. Med. Chem., 1982, 25, 1106 CrossRef CAS PubMed; (c) G. Sosnovsky, N. U. M. Rao and S. W. Li, J. Med. Chem., 1986, 29, 2225 CrossRef CAS PubMed; (d) L. D. Protsenko, I. A. Avrutskaya, T. N. Dneprova, E. Yu. Kodintseva, S. M. Andrianova and P. Y. Sologub, Pharm. Chem. J., 1988, 22, 527 CrossRef; (e) G. Sosnovsky, N. U. M. Rao, S. W. Li and H. M. Swartz, J. Org. Chem., 1989, 54, 3667 Search PubMed; (f) S. Nagahara, A. Murakami and K. Makino, Nucleosides Nucleotides, 1992, 11, 889 Search PubMed; (g) B. Huras, J. Zakrzewski and M. Krawczyk, Lett. Org. Chem., 2010, 7, 545 CrossRef CAS.
- S. G. Reis, M. A. del Águila-Sánchez, G. P. Guedes, Y. Navarro, R. A. Allão-Cassaro, G. B. Ferreira, S. Calancea, F. López-Ortiz and M. G. F. Vaz, Polyhedron, 2018, 144, 166 CrossRef.
- S. G. Reis, M. A. del Águila-Sánchez, G. P. Guedes, G. B. Ferreira, M. A. Novak, N. L. Speziali, F. López-Ortiz and M. G. F. Vaz, Dalton Trans., 2014, 43, 14889 RSC.
- S. G. Reis, M. Briganti, S. Soriano, G. P. Guedes, S. Calancea, C. Tiseanu, M. A. Novak, M. A. del Águila-Sánchez, F. Totti, F. López-Ortiz, M. Andruh and M. G. F. Vaz, Inorg. Chem., 2016, 55, 11676 CrossRef CAS PubMed.
- Y. Navarro, G. P. Guedes, M. A. del Águila-Sánchez, M. J. Iglesias, F. Lloret and F. López-Ortiz, Dalton Trans., 2021, 50, 2585 RSC.
- (a) V. P. Gubskaya, L. Sh. Berezhnaya, V. V. Yanilkin, V. I. Morozov, N. V. Nastapova, Yu. Ya. Efremov and I. A. Nuretdinov, Russ. Chem. Bull., 2005, 54, 1642 CrossRef CAS; (b) K. Rizvanov, G. M. Fazleeva, V. P. Gubskaya and I. A. Nuretdinov, J. Anal. Chem., 2011, 66, 1441 CrossRef.
- B. Huras, J. Zakrzewski and M. Krawczyk, Heteroat. Chem., 2011, 22, 137 CrossRef CAS.
- G. Sosnovsky and M. Konieczny, Z. Naturforsch., B, 1977, 32, 87 CrossRef.
- G. Sosnovsky and M. Konieczny, Z. Naturforsch., B, 1977, 32, 321 CrossRef.
- É. G. Rozantsev, V. I. Suskina, Y. A. Ivanov and B. I. Kaspruk, Russ. Chem. Bull., 1973, 22, 1281 Search PubMed.
- (a) V. D. Seń, E. V. Kapustina and V. A. Golubev, Russ. Chem. Bull., 1983, 32, 1906 CrossRef; (b) V. V. Tkachev, V. D. Seń and L. O. Atovmyan, Russ. Chem. Bull., 1987, 36, 16906 Search PubMed.
- S. Caron, R. W. Dugger, S. G. Ruggeri, J. A. Ragan and D. H. B. Ripin, Chem. Rev., 2006, 106, 2943 CrossRef CAS PubMed.
- (a) D. Wang, A. B. Weinstein, P. B. White and S. S. Shannon, Chem. Rev., 2018, 118, 2636 Search PubMed; (b) S. Najafishirtari, K. Friedel Ortega, M. Douthwaite, S. Pattisson, G. J. Hutchings, C. J. Bondue, K. Tschulik, D. Waffel, B. Peng, M. Deitermann, G. W. Busser, M. Muhler and M. Behrens, Chem. – Eur. J., 2021, 27, 16809 CrossRef CAS PubMed; (c) Y. Zhao, Z. Du, B. Guo, X. Shen, S. Li, T. Wang and C. Liang, Chem. – Asian J., 2022, 17, E202200224 CrossRef CAS PubMed; (d) L. Zhao, O. Akdim, X. Huang, K. Wang, M. Douthwaite, S. Pattisson, R. J. Lewis, R. Lin, B. Yao, D. J. Morgan, G. Shaw, Q. He, D. Bethell, S. McIntosh, C. J. Kiely and G. J. Hutchings, ACS Catal., 2023, 13, 2892 CrossRef CAS PubMed; (e) Z. Shariatinia and Z. Karimzadeh, Coord. Chem. Rev., 2025, 526, 216372 CrossRef CAS; (f) C. Bersani, D. Rodríguez-Padrón, D. Ballesteros, E. Rodríguez-Castellón, A. Perosa and M. Selva, ChemSusChem, 2025, 18, e20240088 CrossRef PubMed.
- (a) I. W. C. E. Arends, P. Gamez and R. A. Sheldon, Adv. Inorg. Chem., 2006, 58, 235 CrossRef CAS; (b) B. Xu, J.-P. Lumb and B. A. Arndtsen, Angew. Chem., Int. Ed., 2015, 54, 4208 Search PubMed; (c) E. Safaei, H. Bahrami, A. Pevec, B. Kozlevčar and Z. Jagličić, J. Mol. Struct., 2017, 1133, 526 CrossRef CAS; (d) B. K. Kundu, R. Ranjan, A. Mukherjee, S. M. Mobin and S. Mukhopadhyay, J. Inorg. Biochem., 2019, 195, 164 Search PubMed; (e) C.-W. Chiang, Y.-H. Chou, C.-H. Chou, H.-C. Chen, J.-L. Chen, L.-C. Hsu, W.-H. Huang, H.-L. Li and Y.-H. Liu, Eur. J. Inorg. Chem., 2024, 27, e202300516 CrossRef CAS.
- (a) L. Marais and A. J. Swarts, Catalysts, 2019, 9, 395 Search PubMed; (b) T. F. S. Silva and L. M. D. R. S. Martins, Molecules, 2020, 25, 748 CrossRef CAS PubMed; (c) Z. Maa, K. T. Mahmudov, V. A. Aliyeva, A. V. Gurbanov and A. J. L. Pombeiro, Coord. Chem. Rev., 2020, 423, 213482 CrossRef; (d) P. Chandra, J. Mol. Struct., 2023, 1273, 134249 CrossRef CAS; (e) Y. Wang, C. Luo, M. Cai and B. Huang, Catal. Lett., 2024, 154, 1431 CrossRef CAS; (f) Z. Huang, W. Zhong, T. Ge, C. Lu and Y. He, J. Catal., 2025, 441, 115858 Search PubMed.
- Bruker, APEX2 v2014.5–0, Bruker AXS Inc., Madison, Wisconsin, USA, 2007 Search PubMed.
- Bruker, SAINT, Bruker AXS Inc., Madison, Wisconsin, USA, 2007 Search PubMed.
- Bruker, SADABS, Bruker AXS Inc., Madison, Wisconsin, USA, 2001 Search PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A: Found. Crystallogr., 2008, 64, 112 CrossRef CAS PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A: Found. Adv., 2015, 71, 3 Search PubMed.
- C. F. Macrae, P. R. Edgington, P. McCabe, E. Pidcock, G. P. Shields, R. Taylor, M. Towler and J. van de Streek, J. Appl. Crystallogr., 2006, 39, 453 CrossRef CAS.
- L. J. Farrugia, J. Appl. Crystallogr., 2012, 45, 849 Search PubMed.
- A. L. Spek, Acta Crystallogr., Sect. D: Biol. Crystallogr., 2009, 65, 148 Search PubMed.
- O. V. Dolomanov, L. J. Bourhis, R. J. Gildea, J. A. K. Howard and H. Puschmann, J. Appl. Crystallogr., 2009, 42, 339 CrossRef CAS.
- F. Neese, WIRES Comput. Molec. Sci., 2025, 15, e70019 CrossRef.
- B. de Souza, Angew. Chem., Int. Ed., 2025, 64, e202500393 CrossRef CAS PubMed.
- F. Weigend and R. Ahlrichs, Phys. Chem. Chem. Phys., 2005, 7, 3297 RSC.
- N. Mardirossian and M. Head-Gordon, J. Chem. Phys., 2016, 144, 214110 Search PubMed.
- T. Soda, Y. Kitagawa, T. Onishi, Y. Takano, Y. Shigeta, H. Nagao, Y. Yoshioka and K. Yamaguchi, Chem. Phys. Lett., 2000, 319, 223 CrossRef CAS.
- K. Yamaguchi, Y. Takahara and T. Fueno, in Applied Quantum Chemistry, Springer Netherlands, Dordrecht, 1986, pp. 155–184 Search PubMed.
- E. D. Glendening, C. R. Landis and F. Weinhold, J. Comput. Chem., 2019, 40, 2234 CrossRef CAS PubMed.
- (a) G. Ruiz-Gómez, A. Francesch, M. J. Iglesias, F. López-Ortiz, C. Cuevas and M. Serrano-Ruiz, Org. Lett., 2008, 10, 3981 CrossRef PubMed; (b) N. Kumari, A. Jagadeesh, P. Galav, A. Kundun and B. Chakraborty, J. Org. Chem., 2024, 89, 15851 CrossRef CAS PubMed.
- (a) J. Fawcett, M. J. P. Harger and R. Sreedharan-Menon, J. Chem. Soc., Chem. Commun., 1992, 227 RSC; (b) R. Murugavel and R. Pothiraja, New J. Chem., 2003, 27, 968 RSC; (c) S. Parsons, T. Barrett, P. Bailey and P. A. Wood, CCDC 247827, CSD Commun., 2004, DOI:10.5517/cc89wfv; (d) G. Gangadhararao and K. C. Kumara Swamy, Tetrahedron, 2014, 70, 2643 CrossRef CAS.
- (a) R. N. Shibaeva, J. Struct. Chem., 1975, 16, 318 CrossRef; (b) D. Bordeaux and J. Lajzérowicz, Acta Crystallogr., Sect. B, 1977, 33, 1837 CrossRef; (c) H. Fujiwara, E. Fujiwara and H. Kobayashi, Mol. Cryst. Liq. Cryst., 2002, 380, 269 CrossRef CAS; (d) F. Türkyilmaz, G. Kehr, J. Li, C. G. Daniliuc, M. Tesch, A. Studer and G. Erker, Angew. Chem., Int. Ed., 2016, 55, 1470 CrossRef PubMed; (e) M. J. Percino, M. Cerón, G. Soriano-Moro, J. A. Pacheco, M. E. Castro, V. M. Chapela, J. Bonilla-Cruz and E. Saldivar-Guerra, J. Mol. Struct., 2016, 1103, 254 CrossRef; (f) G. Ionita, A. M. Madalan, A. M. Ariciu, A. Medvedovici and P. Ionita, New J. Chem., 2016, 40, 503 RSC.
- D. Cremer and J. A. Pople, J. Am. Chem. Soc., 1975, 97, 1354 CrossRef CAS.
- (a) G. J. Jeffrey, An Introduction to Hydrogen Bonding, Oxford University Press, New York, 1997 Search PubMed; (b) D. Herschlag and M. M. Pinney, Biochemistry, 2018, 57, 3338 Search PubMed.
- F. Hamzehee, M. Pourayoubi, M. Nečasb and D. Choquesillo-Lazarte, Acta Crystallogr., Sect. C: Struct. Chem., 2017, 73, 287 CrossRef CAS PubMed.
- (a) G. R. Desiraju, Acc. Chem. Res., 1996, 29, 441 CrossRef CAS PubMed; (b) Z. S. Derewenda, Int. J. Mol. Sci., 2023, 24, 13165 CrossRef CAS PubMed.
- (a) I. Mata, I. Alkorta, E. Molins and E. Espinosa, Chem. – Eur. J., 2010, 16, 2442 CrossRef CAS PubMed; (b) O. Loveday and J. Echeverría, Cryst. Growth Des., 2021, 21, 5961 CrossRef CAS.
- (a) G. P. Guedes, R. G. Zorzanelli, N. M. Comerlato, N. L. Speziali, S. Santos-Jr and M. G. F. Vaz, Inorg. Chem. Commun., 2012, 23, 59 CrossRef CAS; (b) H. Kinoshita, H. Akutsu, J.-I. Yamada and S. Nakatsuji, Mendeleev Commun., 2006, 305 CrossRef CAS; (c) Y. Navarro, G. P. Guedes, J. Cano, P. Ocón, M. J. Iglesias, F. Lloret and F. López-Ortiz, Dalton Trans., 2020, 49, 6280 RSC.
- F. Iwasaki, J. H. Yoshikawa, H. Yamamoto, K. Takada, E. Kannari, M. Yasui, T. Ishida and T. Nogami, Acta Crystallogr., Sect. B: Struct. Sci., 1999, 55, 1057 CrossRef PubMed.
- P. T. Miller, P. G. Lenhert and M. D. Joesten, Inorg. Chem., 1972, 11, 2221 CrossRef CAS.
- M. Bortoluzzi, V. Ferraro and J. Castro, Dalton Trans., 2021, 50, 3132 RSC.
- M. Llunell, D. J. Casanova, P. Alemany and S. Alvarez, SHAPE 2.1, Universitat de Barcelona, Spain, 2013 Search PubMed.
- (a) E. M. Fatila, A. C. Maahs, E. E. Hetherington, B. J. Cooper, R. E. Cooper, N. N. Daanen, M. Jennings, S. E. Skrabalak and K. E. Preuss, Dalton Trans., 2018, 47, 16232 RSC; (b) D. Y. Chappidi, M. N. Gordon, H. M. Ashberry, J. Huang, B. M. Labedis, R. E. Cooper, B. J. Cooper, V. Carta, S. E. Skrabalak, K. R. Dunbar and E. M. Fatila, Inorg. Chem., 2022, 61, 12197 CrossRef CAS PubMed; (c) S. Naseri, M. Mirzakhani, C. Besnard, L. Guénée, L. Briant, H. Nozary and C. Piguet, Chem. – Eur. J., 2023, 29, e202202727 CrossRef CAS PubMed.
- S. K. Gupta, T. Rajeshkumar, G. Rajaraman and R. Murugavel, Chem. Commun., 2016, 52, 7168 RSC.
- (a) S. K. Gupta, T. Rajeshkumar, G. Rajaraman and R. Murugavel, Chem. Sci., 2016, 7, 5181 RSC; (b) A. Ghatak, G. Bhatt, R. Rana, S. K. Gupta, F. Meyer, G. Rajaraman and R. Murugavel, Chem. – Asian J., 2025, 20, e202401477 CrossRef CAS PubMed.
- S. K. Gupta, T. Rajeshkumar, G. Rajaraman and R. Murugavel, Dalton Trans., 2018, 47, 357 RSC.
- S. K. Gupta, S. Dey, T. Rajeshkumar, G. Rajaraman and R. Murugavel, Chem. – Eur. J., 2022, 28, e20210358 Search PubMed.
- S. K. Gupta, S. Shanmugan, T. Rajeshkumar, A. Borah, M. Damjanović, M. Schulze, W. Wernsdorfer, G. Rajaraman and R. Murugavel, Dalton Trans., 2019, 48, 15928 RSC.
- (a) P. Zhang, L. Zhang, S. Xue, S.-Y. Lin and J. Tang, Chin. Sci. Bull., 2012, 57, 2517 CrossRef CAS; (b) K. Bernot, Eur. J. Inorg. Chem., 2023, 26, e202300336 CrossRef CAS.
- A. S. Florencio, R. A. Allão, M. G. F. Vaz and J. W. D. M. Carneiro, Inorg. Chem. Commun., 2012, 24, 67 CrossRef CAS.
- (a) M. Andruh, E. Bakalbassis, O. Kahn, J. C. Trombe and P. Porchers, Inorg. Chem., 1993, 32, 1616 CrossRef CAS; (b) O. Kahn, Molecular Magnetism, VCH Publishers, Inc., New York, 1993 Search PubMed.
- E. S. Areas, B. P. Rodrigues, A. C. C. do Nascimento, H. C. Silva Jr, G. B. Ferreira, F. S. Miranda, F. Garcia, S. H. Safeer, S. Soriano and G. P. Guedes, J. Braz. Chem. Soc., 2024, 35, e20230160 Search PubMed.
- (a) L. Chimilouski, W. H. Slominski, A. I. Tillmann, D. Will, A. M. dos Santos, G. Farias, E. Martendal, K. P. Naidek and F. R. Xavier, Molecules, 2024, 29, 2634 CrossRef CAS PubMed; (b) M. G. B. Rodrigues, A. L. Pesquero de Melo, M. D. Coutinho-Neto and C. A. Angelucci, ChemCatChem, 2025, 17, e202401444 CrossRef CAS; (c) W. Yang, F. Xu and G. Shi, Catal. Lett., 2025, 155, 10 CrossRef CAS; (d) S. Zhang, W. Huang, E. Fayad, D. N. Binjawhar, K. P. Rakesh and H.-L. Qin, Org. Biomol. Chem., 2025, 23, 8128 Search PubMed.
- N. Ch. Jana, S. Behera, S. K. R. Maharana, R. R. Behera and B. Bagh, Catal. Sci. Technol., 2023, 13, 5422 Search PubMed.
- E. Safaeia, L. Hajikhanmirzaei, B. Karimi, A. Wojtczak, P. Cotič and Y.-I. Lee, Polyhedron, 2016, 106, 153 CrossRef.
- X. Lu, N. Wu, B. Zhang and K. Deng, J. Coord. Chem., 2017, 70, 475 CrossRef CAS.
- (a) J. M. Hoover and S. S. Stahl, J. Am. Chem. Soc., 2011, 133, 16901 CrossRef CAS PubMed; (b) J. M. Hoover and S. S. Stahl, Org. Synth., 2013, 90, 240 CrossRef CAS.
- P. Gamez, I. W. C. E. Arends, J. Reedijk and R. A. Sheldon, Chem. Commun., 2003, 2414 RSC.
- (a) CCDC 2501504: Experimental Crystal Structure Determination, 2026, DOI:10.25505/fiz.icsd.cc2pz0pd; (b) CCDC 2501505: Experimental Crystal Structure Determination, 2026, DOI:10.25505/fiz.icsd.cc2pz0qf; (c) CCDC 2501506: Experimental Crystal Structure Determination, 2026, DOI:10.25505/fiz.icsd.cc2pz0rg; (d) CCDC 2501507: Experimental Crystal Structure Determination, 2026, DOI:10.25505/fiz.icsd.cc2pz0sh; (e) CCDC 2501508: Experimental Crystal Structure Determination, 2026, DOI:10.25505/fiz.icsd.cc2pz0tj.
| This journal is © The Royal Society of Chemistry 2026 |