Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
TADDOLs and methylcyclohexanones: selectivity, resolution and the kinetics of decomposition
Hana
Bawa
,
Hong
Su
 ,
Stephen
De Doncker
,
Susan A.
Bourne
,
Stephen
De Doncker
,
Susan A.
Bourne
 and
Luigi R.
Nassimbeni
and
Luigi R.
Nassimbeni
 *
*
Centre for Supramolecular Chemistry Research, Department of Chemistry, University of Cape Town, South Africa. E-mail: Luigi.Nassimbeni@uct.ac.za
First published on 3rd February 2026
Abstract
Three chiral TADDOL (α,α,α,α′-tetraaryl-2,2-disubstituted 1,3-dioxolane-4,5-dimethanol) host molecules were employed to yield inclusion compounds with three isomers of methylcyclohexanones as guests. Although 4-methylcyclohexanone was not a chiral compound due to its internal symmetry, the other two isomers were racemates, and TADDOLs acted as potential resolving agents. The selectivity preference of the methylcyclohexanones for each host was established using solution nuclear magnetic resonance (NMR) spectroscopy, and the crystal structure of each host–guest compound was elucidated by single-crystal X-ray diffraction. The packing of the structures was analysed to explain the resulting resolution of the enantiomeric guests. The kinetics of decomposition were investigated for a representative TADDOL host.
Introduction
The resolution of racemic modifications has a long history. It started with Pasteur's spectacular experiment in 1848, when he observed that the double salt of sodium ammonium tartaric acid crystallised with hemihedral facets and these crystals were mirror images. He separated them into distinct batches (triage), dissolved them and noted that when these solutions were viewed in plane polarised light, they exhibited opposite optical rotations. He deduced that the asymmetry (chirality) was not only a feature of the crystals but also of the molecules themselves, which was a brilliant deduction.1 An English translation of Pasteur's paper was published by Gal in 2013.2Since that time, there have been many discoveries and technical applications to the problem of resolution of racemates involving various analytical techniques. These include chiral chromatography, the formation of diastereomeric salts, using enzymes that regulate and catalyse biochemical pathways, and resolution by entrainment. In particular, the work by Vries et al.3 demonstrated that that using a mixture of similar host compounds yields better resolution than employing individual compounds. This method, known as the “Dutch resolution method” has proved valuable in the pharmaceutical industry.
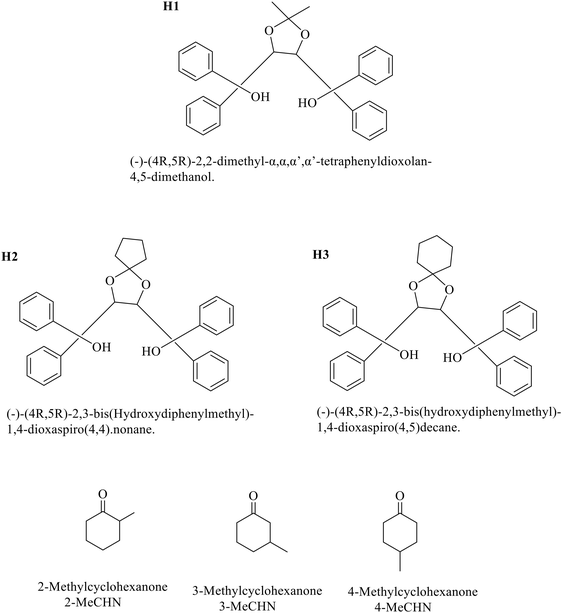
In this work, we present the results of the selectivity and resolution of methylcyclohexanones by three TADDOL chiral host compounds, which are also used as resolving agents for two of the three racemic methylcyclohexanones: 2-methylcyclohexanone (2-MeCHN) and 3-methylcyclohexanone (3-MeCHN), both of which are chiral, and 4-methylcyclohexanone (4-MeCHN), which is not chiral due to its internal symmetry. TADDOL is an acronym for α,α,α,α′-tetraaryl-2,2-disubstituted 1,3-dioxolane-4,5-dimethanol. The three host molecules, labelled H1, H2, and H3, together with the methylcyclohexanone guests, are shown in Scheme 1.
TADDOLs are widely used host compounds, and their structures are well represented in the Cambridge Structural Database (CSD),4 which lists more than 100 compounds of the TADDOL structure and their inclusion compounds. They were described in a review entitled ‘Molecular Recognition in the Solid State’, which explains that they are important because they are derived from naturally occurring tartaric acid.5 TADDOLs are used as separating agents for dialkyl arylphosphine oxides,6 heterocyclic molecules7 and methylpyridines.8
Methylcyclohexanones have also attracted research interest, and their structures have been studied, with CSD yielding a total of 38 structures as guest molecules with a variety of hosts. For example deoxycholic acid (DCA) includes all the methylcyclohexanones and competition experiments showed the selectivity preference was of the order 2-MeCHN > 3-MeCHN > 4-MeCHN, which were confirmed by their structures, NMR and thermal analysis.9 The host 2,2′-(benzene-1,4-diyl-diethynylene)diborneol, a dumb-bell-shaped molecule, resolved both 2-MeCHN and 3-MeCHN as the S conformers.10 An interesting result was reported in which the recrystallization of 3-MeCHN with a chiral host resulted in partial resolution, but recrystallization from an equimolar mixture of this host and a similar but achiral host achieved full resolution of 3-MeCHN. The achiral host acts as a nucleation inhibitor and the enhanced resolution was attributed to kinetic effects.11
Experimental
Materials
TADDOL host compounds, H1, H2, and H3, were synthesised by Fumio Toda,12 and methylcyclohexanones were purchased from Sigma Aldrich and used as received. Inclusion compounds with single guests were prepared by dissolving the host (100 mg, 0.2 mmol) with the appropriate guest isomer (530 mg, 5 mmol) under gentle heating and stirring. Suitable crystals were obtained by slow evaporation of the solution at ambient temperature.Single-crystal x-ray diffraction
In this study, all crystallographic data were collected using a Bruker D8 Venture single-crystal X-ray diffractometer equipped with graphite-monochromatic MoKα radiation (λ = 0.71073 Å), which was generated by applying a Bruker K780 X-ray generator operating at 50 kV and 1.4 mA. Data collection was performed at 100 K.Intensity data were collected using both phi (ϕ) and omega (ω) scan techniques, and data reduction and unit cell refinement were carried out using SAINT-Plus,13 followed by XPREP,14 which uses systematic absence to determine the space group. Absorption corrections of the collected intensities were applied using SADABS.15 The structures were solved using direct methods with SHELXS-97 and refined using full-matrix least-squares methods in SHELXL.16 Hydrogen atoms were refined using riding models. For the O–H and N–H groups, the H atoms were allowed to refine freely. The structure solution and refinement processes were carried out using the X-Seed graphical interface,17 while final molecular graphics and high-resolution images were produced using X-Seed in conjunction with POV-RAY.18
1H nuclear magnetic resonance (NMR) spectrometry
1H NMR spectra were recorded using a Bruker 400 MHz spectrometer with DMSO as the internal standard. The samples were blotted dry, crushed, and dissolved in deuterated d6-DMSO. Appropriate signals were integrated to determine the concentration and relative proportions of the guests.Results and discussion
Selectivity for methylcyclohexanones guests by crystallization
The selectivity of H1, H2 and H3 for the methylcyclohexanones was determined by dissolving a fixed quantity of each host in an equimolar solution of the MeCHN guests. The starting solutions were prepared with an excess of liquid guests to ensure that selectivity was not influenced by concentration once crystals began forming (the molar host![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) guest ratio was 1
guest ratio was 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 49). The ensuing inclusion compound crystals were analysed by NMR (Table 1). The results are plotted on an equilateral triangle, in which the apices represent the pure guest (MeCHNs), Fig. 1. The starting concentration is shown as a black dot representing the mole fraction of each MeCHN guest as 1/3, and their final concentrations are shown by the arrows for H1, H2 and H3. H1 shows a modest preference for 3-MeCHN, while H2 and H3 disfavour 2-MeCHN and tend towards 4-MeCHN.
49). The ensuing inclusion compound crystals were analysed by NMR (Table 1). The results are plotted on an equilateral triangle, in which the apices represent the pure guest (MeCHNs), Fig. 1. The starting concentration is shown as a black dot representing the mole fraction of each MeCHN guest as 1/3, and their final concentrations are shown by the arrows for H1, H2 and H3. H1 shows a modest preference for 3-MeCHN, while H2 and H3 disfavour 2-MeCHN and tend towards 4-MeCHN.
| 2-MeCHN | 3-MeCHN | 4-MeCH | |
|---|---|---|---|
| Start | 33.3% | 33.3% | 33.3% |
| End H1 | 25% | 50% | 25% |
| End H2 | 0% | 25% | 75% |
| End H3 | 0% | 27% | 73% |
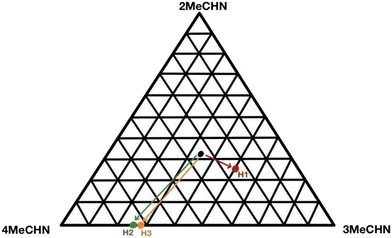 | ||
| Fig. 1 Selectivity of the hosts H1, H2 and H3 for the mixtures of methylcyclohexanones (2-MeCHN, 3-MeCHN, and 4-MeCHN). | ||
Structural results
We attempted to obtain crystals by dissolving each of the three hosts H1, H2 and H3, with each of the methylcyclohexanone guests 2-MeCHN, 3-MeCHN and 4-MeCHN (nine solutions, Table S1). However, we failed to obtain crystals from the solution of H3 with 3-MeCHN, which would have corresponded to structure VIII and thus we determined only eight crystal structures, labelled in roman numbers from I to IX.One of the common features of all these structures is that the conformation of the host molecules places their hydroxyl groups in close proximity, resulting in an intramolecular hydrogen bond (O–H⋯O) and a further hydrogen bond to the guest (O–H⋯O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) of the carbonyl O of the methylcyclohexanone. The details of the hydrogen bonding for all the structures are reported in the SI in Tables S2 and S3.
C) of the carbonyl O of the methylcyclohexanone. The details of the hydrogen bonding for all the structures are reported in the SI in Tables S2 and S3.
The description of these eight crystal structures has been organised by guest species.
Compounds of 2-methylcyclohexanone: I, IV and VII
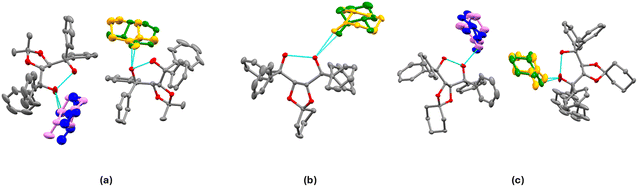
The details of the crystallographic refinement are illustrated in Table 2. Fig. 2 shows the asymmetric units (ASU) for H1·(2-MeCHN) (I), H2·(2-MeCHN) (IV) and H3·(2-MeCHN) (VII).| Structure | I | IV | VII |
|---|---|---|---|
| Structural formula | H1·(2-MeCHN) | H2·(2-MeCHN) | H3·(2-MeCHN) |
| Structural formula | C31H30O4·C7H12O | C33H32O4·C7H12O | C34H34O4·C7H12O |
| Host/guest ratio | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| Molecular mass (g mol−1) | 578.71 | 604.75 | 618.78 |
| Data collection temp (K) | 100(2) | 100(2) | 100(2) |
| Crystal system | Monoclinic | Orthorhombic | Triclinic |
| Space group | P21 | P212121 | P1 |
| a (Å) | 9.448(1) | 9.254(7) | 9.334(1) |
| b (Å) | 9.526(1) | 16.608(1) | 9.951(1) |
| c (Å) | 34.801(4) | 21.476(2) | 18.611(3) |
| α (°) | 90 | 90 | 100.611(4) |
| β (°) | 90.218(4) | 90 | 98.637(4) |
| γ (°) | 90 | 90 | 91.317(4) |
| Volume (Å3) | 3112.0(8) | 3300.5(4) | 1677.5(4) |
| Z | 4 | 4 | 2 |
| D c, calc density (g cm−3) | 1.227 | 1.217 | 1.225 |
| F(000) | 1240 | 1296 | 664 |
| Range θ(°) | 2.156–27.195 | 2.396–28.312 | 2.085–28.491 |
| Reflections collected | 127![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 696 696 |
43![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 859 859 |
57![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 694 694 |
| No data I > 2σ(I) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 886 886 |
7499 | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 728 728 |
| Final R indices [I > 2σ(I)], R1/wR2 | 0.1151/0.3212 | 0.0386/0.0901 | 0.0898/0.1080 |
| R indices (all data), R1/wR2 | 0.1199/0.3244 | 0.0440/0.0933 | 0.2041/0.2155 |
| Goodness of fit on F2 | 1.058 | 1.028 | 1.052 |
| CCDC no | 2479981 | 2479985 | 2479987 |
Sin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ/λ (Å−1) θ/λ (Å−1) |
0.643 | 0.667 | 0.671 |
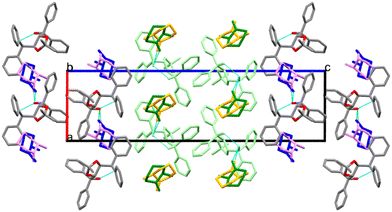
Structure I, H1·(2-MeCHN), crystallizes in space group P21 with Z = 4. There are two host molecules in the asymmetric unit (ASU), Fig. 2a. Both guest enantiomers lie on the same site, giving rise to disorder for which we modelled the site occupancy factors. The guests labelled with the suffix X (62% S) and Y (38% R) are hydrogen bonded to host molecule A. Those labelled W (45% S) and Z (55% R) are hydrogen bonded to host molecule B. They are shown in Fig. 2a using the colour code green/violet for the R-enantiomer and orange/blue for the S-enantiomer. The packing shown in Fig. 3 demonstrates that the guest 2-MeCHN molecules are located in channels along [010] and that the two crystallographically independent hosts pack in layers parallel to [010].
We note that this compound is difficult to crystallize. This structure shows that the host captures 2-MeCHN, but these are enclathrated in approximately equal proportions of R and S enantiomers. The enantiomeric excess (e.e.) of S was only 7%. Thus, the resolution of 2-MeCHN using H1 does not work in practice.
Structure IV, H2·(2-MeCHN), crystallises in space group P212121 with Z = 4, and each host molecule enclathrates one guest molecule. The guest molecule is disordered and shares the same location, as shown in Fig. 2b. This shows the intramolecular H-bond (O⋯O = 2.673(1) Å), and the guests X (orange; S) and Y (green; R) are hydrogen-bonded via (host) O–H⋯O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C(2-MeCHN) = 2.76(1) Å and 2.80(1) Å, respectively. The resolution is poor with 52% R and 48% S, yielding an enantiomeric excess (e.e.) of only 4%.
C(2-MeCHN) = 2.76(1) Å and 2.80(1) Å, respectively. The resolution is poor with 52% R and 48% S, yielding an enantiomeric excess (e.e.) of only 4%.
Structure VII, H3·(2-MeCHN), crystallises in space group P1 with Z = 2, and a host![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) guest ratio of 1
guest ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 host A is partly disordered in one phenyl ring, and both 2-MeCHN guests are disordered and display mixed chirality. The guest H-bonded to host A is labelled X (30% R) and Y (70% S), with O⋯O = 2.79(2) Å and 2.62(1) Å. Host B is similarly H-bonded to guest W (38% S) and Z (62% R) with (host) O⋯O (guest) 2.69(3) Å and 2.76(1) Å (Fig. 2c). Overall, considering both guests, the (e.e.) is 24% R, a poor resolution result.
1 host A is partly disordered in one phenyl ring, and both 2-MeCHN guests are disordered and display mixed chirality. The guest H-bonded to host A is labelled X (30% R) and Y (70% S), with O⋯O = 2.79(2) Å and 2.62(1) Å. Host B is similarly H-bonded to guest W (38% S) and Z (62% R) with (host) O⋯O (guest) 2.69(3) Å and 2.76(1) Å (Fig. 2c). Overall, considering both guests, the (e.e.) is 24% R, a poor resolution result.
The packings of IV and VII are shown in Fig. 4. Guests are located in channels in both cases.
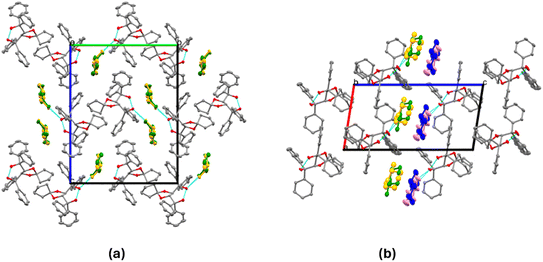 | ||
| Fig. 4 Packing in (a) IV and (b) VII, showing the guests located in channels. The guest 2-MeCHN is disordered, with both enantiomers occupying the same sites and different site occupancy factors. | ||
Compounds of 3-methylcyclohexanone: II, V and VIII
Fig. 5 shows the asymmetric units (ASU) for H1·(3-MeCHN) (II) and H2·(3-MeCHN) (V). Details of the crystallographic refinement are shown in Table 3. Structure VIII, H3·(3-MeCHN), did not yield suitable crystals despite several crystallization attempts lasting over 5 months. Therefore, no structure is reported.| Structure | II | V |
|---|---|---|
| Structural formula | H1·(3-MeCHN) | H2·(3-MeCHN) |
| Structural formula | C31H30O4·C7H12O | C33H32O4·C7H12O |
| Host/guest ratio | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| Molecular mass (g mol−1) | 578.71 | 604.75 |
| Data collection temp (K) | 100(2) | 100(2) |
| Crystal system | Orthorhombic | Orthorhombic |
| Space group | P212121 | P212121 |
| a (Å) | 9.256(1) | 9.655(7) |
| b (Å) | 9.383(1) | 11.226(7) |
| c (Å) | 35.837(5) | 29.717(2) |
| α (°) | 90 | 90 |
| β (°) | 90 | 90 |
| γ (°) | 90 | 90 |
| Volume (Å3) | 3112.3(6) | 3220.6(4) |
| Z | 4 | 4 |
| D c, calc density (g cm−3) | 1.235 | 1.247 |
| F(000) | 1240 | 1296 |
| Range θ(°) | 2.450–28.367 | 1.939–28.336 |
| Reflections collected | 37![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 416 416 |
80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 407 407 |
| No data I > 2σ(I) | 7084 | 7465 |
| Final R indices [I > 2σ(I)], R1/wR2 | 0.0654/0.0714 | 0.0393/0.0436 |
| R indices (all data), R1/wR2 | 0.1703/0.1753 | 0.0979/0.1013 |
| Goodness of fit on F2 | 1.049 | 1.036 |
| CCDC no | 2479982 | 2479986 |
Sin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ/λ (Å−1) θ/λ (Å−1) |
0.668 | 0.668 |
Both II, H1·(3-MeCHN), and V, H2·(3-MeCHN), crystallize in space group P212121 with Z = 4, and a host![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) guest ratio of 1
guest ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 although the unit cell axis lengths differ significantly. The asymmetric units for these structures are shown in Fig. 5. This illustrates both the intramolecular H-bond and the H-bond to the 3-MeCHN guest (details are shown in Tables S2 and S3). The conformation at the 3-methyl carbon is S in II, and R in V, indicating that H1 and H2 can resolve 3-methylcyclohexanone. The guest molecules fill cavities in the crystal structure, as shown in Fig. 6 for II and in Fig. S1 for V.
1 although the unit cell axis lengths differ significantly. The asymmetric units for these structures are shown in Fig. 5. This illustrates both the intramolecular H-bond and the H-bond to the 3-MeCHN guest (details are shown in Tables S2 and S3). The conformation at the 3-methyl carbon is S in II, and R in V, indicating that H1 and H2 can resolve 3-methylcyclohexanone. The guest molecules fill cavities in the crystal structure, as shown in Fig. 6 for II and in Fig. S1 for V.
Compounds of 4-methylcyclohexanone: III, VI and IX
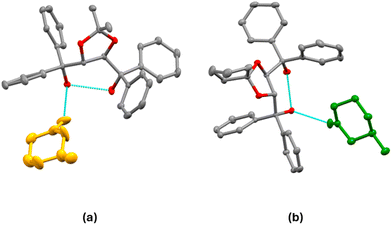
Fig. 7 shows the asymmetric units (ASU) for H1·(4-MeCHN) (III), H2·(4-MeCHN) (VI) and H3·(4-MeCHN) (IX). Details of the crystallographic refinement are presented in Table 4.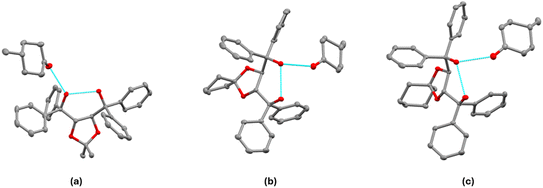 | ||
| Fig. 7 Asymmetric units of (a) III, (b) VI, and (c) IX. All displacement ellipsoids are drawn at 50% probability. | ||
| Structure | III | VI | IX |
|---|---|---|---|
| Structural formula | H1·(4-MeCHN) | H2·(4-MeCHN) | H3·(4-MeCHN) |
| Structural formula | C31H30O4·C7H12O | C33H32O4·C7H12O | C34H34O4·C7H12O |
| Host/guest ratio | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| Molecular mass (g mol−1) | 578.71 | 604.75 | 618.78 |
| Data collection temp (K) | 100(2) | 100(2) | 100(2) |
| Crystal system | Orthorhombic | Orthorhombic | Orthorhombic |
| Space group | P212121 | P212121 | P212121 |
| a (Å) | 9.324(2) | 9.181(2) | 9.324(2) |
| b (Å) | 12.457(2) | 15.953(4) | 15.978(4) |
| c (Å) | 26.951(7) | 22.039(6) | 22.244(6) |
| α (°) | 90 | 90 | 90 |
| β (°) | 90 | 90 | 90 |
| γ (°) | 90 | 90 | 90 |
| Volume (Å3) | 3130.3(12) | 3228.0(1) | 3314.0(14) |
| Z | 4 | 4 | 4 |
| D c, calc density (g cm−3) | 1.228 | 1.244 | 1.240 |
| F(000) | 1240 | 1296 | 1328 |
| Range θ(°) | 2.311–28.361 | 2.403–28.384 | 2.368–28.337 |
| Reflections collected | 47![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 934 934 |
81![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 428 428 |
91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 602 602 |
| No data I > 2σ(I) | 6885 | 7220 | 7856 |
| Final R indices [I > 2σ(I)], R1/wR2 | 0.0391/0.0486 | 0.038/0.084 | 0.0321/0.0345 |
| R indices (all data), R1/wR2 | 0.0890/0.0941 | 0.047/0.089 | 0.0793/0.0808 |
| Goodness of fit on F2 | 1.041 | 1.029 | 1.032 |
| CCDC no | 2479983 | 2479997 | 2480004 |
Sin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ/λ (Å−1) θ/λ (Å−1) |
0.668 | 0.668 | 0.668 |
All three structures with 4-MeCHN crystallize in the space group P212121 with Z = 4 and a host![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) guest ratio of 1
guest ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and have similar unit cell dimensions. The asymmetric units for these structures are shown in Fig. 7, showing both the intramolecular H-bond and the H-bond to the 4-MeCHN guest (details are shown in Tables S2 and S3). 4-MeCHN is achiral and is included here for comparison purposes. The methyl group on the cyclohexanone is equatorial in all three structures.
1 and have similar unit cell dimensions. The asymmetric units for these structures are shown in Fig. 7, showing both the intramolecular H-bond and the H-bond to the 4-MeCHN guest (details are shown in Tables S2 and S3). 4-MeCHN is achiral and is included here for comparison purposes. The methyl group on the cyclohexanone is equatorial in all three structures.
Comparison of related structures
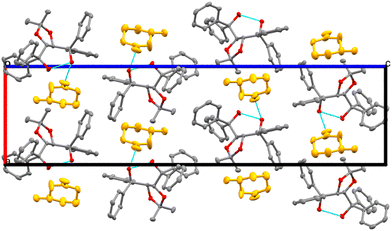
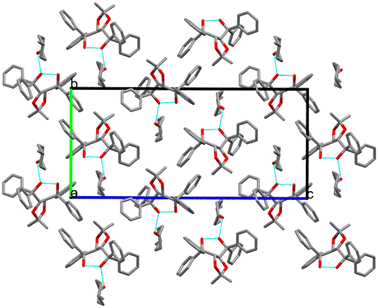
The packing is similar for structures II, III, IV, V, VI and IX, with the guests located in channels running parallel to [100], as illustrated by the example in Fig. 8. We compared the guest orientation in the series of inclusion compounds with H2 (IV, V and VI). In structure IV, the 2-MeCHN guests are unresolved and are located in channels parallel to [100]. Their respective site occupancies are 52% (R) and 48% (S) (Table S3). Hydrogen bonds, (host)-O–H⋯O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C-(guest), are similar, with O⋯O distances of 2.76(1) Å (R) and 2.80(1) Å (S). The mean plane of the 2-MeCHN guests is perpendicular to [001].
C-(guest), are similar, with O⋯O distances of 2.76(1) Å (R) and 2.80(1) Å (S). The mean plane of the 2-MeCHN guests is perpendicular to [001].
Structure V has 3-MeCHN guests lying in channels parallel to [100] but oriented with their mean planes parallel to [110]. (Host)-O–H⋯O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C-(guest) hydrogen bonds have O⋯O distances of 2.77(2) Å.
C-(guest) hydrogen bonds have O⋯O distances of 2.77(2) Å.
Although the channels in structure VI are also parallel to [100], the 4-MeCHN guests are oriented with their mean planes perpendicular to [001] and an O⋯O hydrogen bond distance of 2.76(2) Å.
We observe that none of the structures of the inclusion compounds with H1, H2, and H3 crystallised with 2-MeCHN were resolved. The crystals obtained from these solutions were the slowest to appear. We also note that in structure II, H1·(3-MeCHN), host H1 resolved 3-MeCHN as ‘S’, but that in structure V, H2·(3-MeCHN), resolved 3-MeCHN as ‘R’. In these hosts, the groups bonded at position 2 of the dioxolane ring are quite different in bulk (dimethyl versus cyclic pentomic in H1 and H2, respectively). Analysis of the non-bonded interactions showed that the cyclic pentomic ring has a close contact of 3.93(1) Å with the chiral carbon (C) atom of 3-MeCHN, but this has no equivalent with H1. Although 4-MeCHN is achiral, it is included in this study for comparison; the structures of all three inclusion compounds with this guest are isostructural.
Kinetics of decomposition
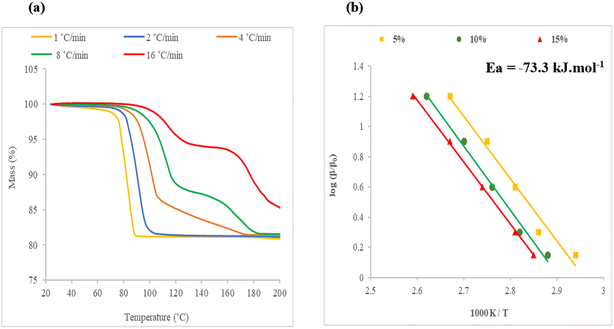
The inclusion compounds derived by the combination of H2 with each of the methylcyclohexanones were subjected to thermal analysis at predetermined heating rates. The results were analysed by applying the method of Flynn, Wall and Ozawa.19–21 The decomposition showed different curves representing the extent of reaction (α), corresponding to the loss of the guest molecule.We note that the TG curves, showing the decomposition of the inclusion compounds, display a two-step reaction, which is particularly pronounced in structure VI, H2·(4-MeCHN). In order to ensure consistency in the comparison of activation energies obtained, we analysed only the early decomposition reaction in each case. The results are shown in Fig. 9–11. A small decomposition step at the beginning of some TG curves is attributed to the surface solvent.
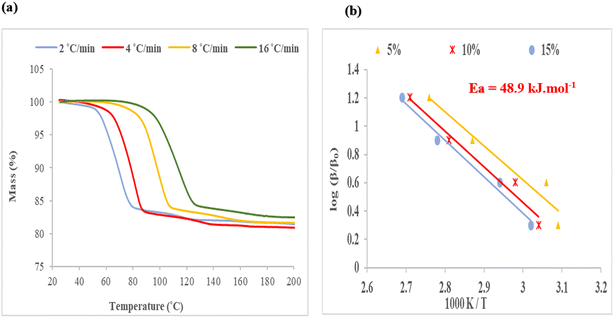
An example is shown in Fig. 9(a) for the decomposition of structure IV, H2·(2-MeCHN), in which the heating rates, β, were changed geometrically as β+ 2, 4, 8, 16 °C min−1, and the plot of log (β/β0) versus 1000 K/T is shown in Fig. 9(b). The slope of these curves yielded an average energy of activation of 48.9 kJ mol−1.
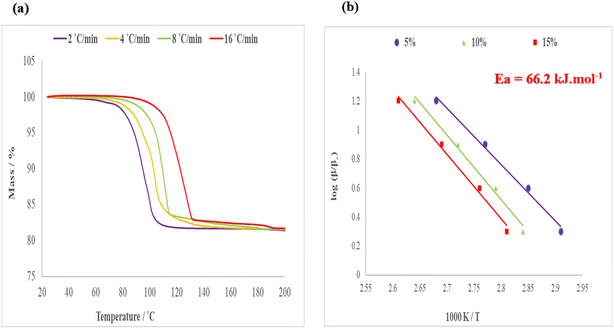
The same procedure was followed for the decomposition of structure V, H2·(3-MeCHN), where the decay curves are shown in Fig. 10(a) and the corresponding semi-logarithmic plot is depicted in Fig. 10(b). In this instance, the activation energy of decomposition has an average of 66.2 kJ mol−1.
The experiment of the decomposition of structure VI, H2·(4-MeCHN) showed an uncommon result, and it was noted that only at heating rates of 1 °C min−1 and 2 °C min−1 did the decomposition reaction display a stoichiometry with an H![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) G ratio of 1
G ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. At higher heating rates, the decomposition shows a more complex mechanism of guest loss. We therefore analysed only the beginning of the reaction, namely at a mass loss of 5%, 10% and 15%. The decomposition traces are shown in Fig. 11(a), and the corresponding semi-logarithmic plot in Fig. 11(b), which gave an activation energy of the starting reaction of 73.3 kJ mol−1.
1. At higher heating rates, the decomposition shows a more complex mechanism of guest loss. We therefore analysed only the beginning of the reaction, namely at a mass loss of 5%, 10% and 15%. The decomposition traces are shown in Fig. 11(a), and the corresponding semi-logarithmic plot in Fig. 11(b), which gave an activation energy of the starting reaction of 73.3 kJ mol−1.
We note that all the structures with the guest (2-MeCHN) are disfavoured in comparison with (3-MeCHN) and (4-MeCHN) (see Fig. 1). It is noteworthy that the calculated densities of the structures with (2-MeCHN) are lower, possibly resulting from the disorder of this guest that occurs in these structures. The activation energy for the H2·(2-MeCHN) decomposition reaction is 48.9 kJ mol−1, which is lower than that of H2·(3-MeCHN) and H2·(4-MeCHN) with the average values of 66.2 kJ mol−1 and 73.3 kJ mol−1, respectively. This indicates that a poorly packed structure is likely to present a lower activation energy of decomposition. We also carried out the same analysis on H1·(3-MeCHN) for comparison. The activation energy averages 73.1 kJ mol−1 (Fig. S4), which is similar to that of H2·(4-MeCHN). We note that Fig. 1 shows that H1 preferentially includes 3-MeCHN while H2 prefers 4-MeCHN, so the activation energies are consistent with the selectivity results.
Conclusion
Three similar chiral TADDOL host molecules were employed to form inclusion compounds with the three isomers of methylcyclohexanones. The selectivity, as measured from the solution NMR, showed that 2-MeCHN was disfavoured relative to 3-MeCHN and 4-MeCHN. The structure of each inclusion compound was elucidated by single-crystal X-ray diffraction. Nine host–guest solutions were prepared, but one failed to yield suitable crystals [H3·(3-MeCHN)]. Each host was exposed to racemic 2-MeCHN and 3-MeCHN to test for possible enantiomeric resolution. The result of the guest resolution was that 2-MeCHN was not resolved by any of the hosts, while H1 and H2 were capable of resolving 3-MeCHN, a result consistent with the selectivity experiments. All the structures with 4-MeCHN as a guest showed the methyl group of the latter in the equatorial position. The inclusion compounds with H2 were subjected to thermal decomposition at various heating temperatures, which allowed for an estimation of the activation energies of the decomposition reactions. The activation energy for the loss of 2-MeCHN is significantly lower than that for either 3-MeCHN or 4-MeCHN, which is again consistent with the selectivity experiments.Author contributions
HB: investigation, analysis, writing – original draft; HS and SDD: data collection, analysis; SAB: supervision, analysis, writing – review & editing; LRN: conceptualisation, supervision, writing – review & editing.Conflicts of interest
The authors declare no competing interests.Data availability
The data supporting this article have been included in the article or as part of the supplementary information (SI). Supplementary information: crystallographic data, thermogravimetry and NMR spectra. See DOI: https://doi.org/10.1039/d5ce01022e.CCDC 2479981–2479983, 2479985–2479987, 2479997 and 2480004 contain the supplementary crystallographic data for this paper.22a–h
Acknowledgements
We thank the National Research Foundation (South Africa), the University of Cape Town, for research grants and the Libyan Government for the financial support.References
- L. Pasteur, Mémoire sur la relation qui peut exister entre la forme cristalline et la composition chimique. et sur la cause de la polarisation rotatoire, C. R. Séances Acad. Sci., 1848, 26, 535–538 Search PubMed.
- J. Gal, Translation from French, Pasteur L Memoir on the relationship that can exist between crystalline form and chemical composition and on the cause of optical rotation. Citation for chemical breakthrough awards: choosing Pasteur's award-winning publication, Bull. Hist. Chem., 2013, 38, 7–12 CrossRef CAS.
- T. Vries, H. Wynberg, E. van Echten, J. Koek, W. ten Hoeve, R. M. Kellogg, Q. B. Broxterman, A. Minnaard, B. Kaptein, S. van der Sluis, L. Hulshof and J. Kooistra, Angew. Chem., Int. Ed., 1998, 37, 2349–2354 Search PubMed.
- C. R. Groom, I. J. Bruno, M. P. Lightfoot and S. C. Ward, Acta Crystallogr., Sect. B: Struct. Sci., Cryst. Eng. Mater., 2016, 72, 171–179 CrossRef CAS PubMed.
- F. Toda, in Advances in Supramolecular Chemistry, ed. W. G. Gokel, JAI Press, Greenwich, CT, USA, 1992, vol. 2, pp. 141–191 Search PubMed.
- B. Varga, R. Herbay, G. Székely, T. Holczbauer, J. Madarász, B. Mátravölgyi, E. Fogassy, G. Keglevich and P. Bagi, Eur. J. Org. Chem., 2020, 2020, 1840–1852 CrossRef CAS.
- D. L. Recchia, B. Barton and E. C. Hosten, J. Inclusion Phenom. Macrocyclic Chem., 2025, 105, 291–296 Search PubMed.
- B. Barton, E. C. Hosten and D. V. Jooste, Tetrahedron, 2017, 18, 2662–2673 Search PubMed.
- J. S. B. Boudiombo, H. Su, N. Ravenscroft, S. A. Bourne and L. R. Nassimbeni, Cryst. Growth Des., 2019, 19, 3962–3968 CrossRef.
- E. Batisai, L. R. Nassimbeni and E. Weber, CrystEngComm, 2015, 17, 4205–4209 RSC.
- L. R. Nassimbeni, H. Su and T. L. Curtin, Chem. Commun., 2012, 48, 8526–8528 RSC.
- F. Toda, Topics in Current Chemistry, in Molecular Inclusion and Molecular Recognition - Clathrates I, ed. E. Weber, Springer-Verlag, Berlin, 1987, ch. 3, vol. 140 Search PubMed.
- SAINT-Plus, Madison, Wisconsin, USA, 2004 Search PubMed.
- XPREP, Madison, Wisconsin, USA, 2003 Search PubMed.
- G. M. Sheldrick, SADABS Program for Empirical Absorption Correction of Area Detector Data, Gottingen, 1996 Search PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. C: Struct. Chem., 2015, 71, 3–8 Search PubMed.
- L. J. Barbour, J. Appl. Crystallogr., 2020, 53, 1141–1146 Search PubMed.
- POV-Ray, Persistence of Vision Pty Ltd., Williamstown, Victoria, Australia, 2004 Search PubMed.
- J. H. Flynn and L. A. Wall, J. Polym. Sci., Part B: Polym. Phys., 1996, 4, 323–328 CrossRef.
- T. Ozawa, Bull. Chem. Soc. Jpn., 1965, 38, 1881–1886 CrossRef CAS.
- N. Koga, J. Therm. Anal. Calorim., 2013, 113, 1527–1541 CrossRef CAS.
- (a) CCDC 2479981: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7mdz; (b) CCDC 2479982: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7mf0; (c) CCDC 2479983: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7mg1; (d) CCDC 2479985: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7mj3; (e) CCDC 2479986: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7mk4; (f) CCDC 2479987: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7ml5; (g) CCDC 2479997: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7mxh; (h) CCDC 2480004: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p7n4r.
| This journal is © The Royal Society of Chemistry 2026 |