Open Access Article
Open Access ArticleDesign and concise synthesis of halicyclamine-inspired bis-macrocycles targeting DedA membrane proteins
Toshiaki Wayama a,
Riku Horinouchib,
Masayoshi Arai*c,
Hirofumi Ohashid,
Koichi Watashi*d,
Satoshi Yoshidae,
Sota Sato†
a,
Riku Horinouchib,
Masayoshi Arai*c,
Hirofumi Ohashid,
Koichi Watashi*d,
Satoshi Yoshidae,
Sota Sato†
 ef,
Sachiko Tsukamoto
ef,
Sachiko Tsukamoto g and
Hiroki Oguri
g and
Hiroki Oguri *a
*a
aDepartment of Chemistry, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan. E-mail: hirokioguri@g.ecc.u-tokyo.ac.jp
bSchool of Pharmaceutical Sciences, The University of Osaka, 1-6 Yamadaoka, Suita, Osaka 565-0871, Japan
cGraduate School and School of Pharmaceutical Sciences, The University of Osaka, 1-6 Yamadaoka, Suita, Osaka 565-0871, Japan. E-mail: araim@phs.osaka-u.ac.jp
dDepartment of Drug Development, National Institute of Infectious Diseases, Japan Institute for Health Security, 1-23-1 Toyama, Shinjuku-ku, Tokyo 162-8640, Japan. E-mail: watashi.k@jihs.go.jp
eDepartment of Applied Chemistry, School of Engineering, The University of Tokyo, FS CREATION, Mitsui LINK Lab, Kashiwanoha 1, 6-6-2 Kashiwanoha, Kashiwa, Chiba 277-0882, Japan
fInstitute for Molecular Science, 5-1 Higashiyama, Myodaiji, Okazaki, Aichi 444-8787, Japan
gDepartment of Natural Medicines, Graduate School of Pharmaceutical Sciences, Kumamoto University, 5-1 Oehonmachi, Chuo-ku, Kumamoto 862-0973, Japan
First published on 10th April 2026
Abstract
DedA membrane proteins promote the formation of double-membrane vesicles serving as replication platforms for RNA viruses, including SARS-CoV-2. Inspired by the only known inhibitor, halicyclamine A, we developed a concise biomimetic synthesis of halicyclamine-inspired macrocycles, identifying bis-macrocycles that suppress SARS-CoV-2 replication, while exhibiting DedA-associated activity in a bacterial overexpression assay.
Positive-sense single-stranded RNA viruses, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), replicate in unique membrane compartments such as double-membrane vesicles (DMVs) derived from the endoplasmic reticulum and formed specifically in infected cells (Fig. 1).1 Biogenesis of these membranous replication organelles involves both viral and host factors. The host DedA superfamily, such as TMEM41B2 and VMP1,3 which function as lipid scramblases to remodel the lipid bilayer, has been reported to drive the maturation of the membrane structures into DMVs.4,5 These DMVs are formed in the cells infected with positive-sense single-stranded RNA viruses, including not only SARS-CoV-2 but also hepatitis C virus, enteroviruses, and noroviruses.6 While viral proteases and polymerases have been primary targets of antiviral therapeutics, formation of DMVs is expected to be an alternative target of drugs, especially those effective against multiple viruses. However, chemical modulation of host DedA membrane proteins as an antiviral strategy remains largely unexplored.7
 | ||
| Fig. 1 The estimated function of DedA membrane proteins in the replication cycle of SARS-CoV-2 and halicyclamine A (1), a reported inhibitor of bacterial DedA. | ||
To date, the sponge-derived manzamine alkaloid halicyclamine A (1) has been reported as the only molecule that targets DedA membrane proteins (Fig. 1).8 Manzamine alkaloids,9,10 exemplified by halicyclamine A (1), possess characteristic architectures comprising multicyclic bis-nitrogenated cores and macrocyclic alkyl loops, and display diverse biological activities such as antimalarial11 and antituberculosis12 effects. Their sp3-rich macrocyclic scaffolds occupy an underexplored chemical space for the development of next-generation medium-sized molecule therapeutics.13
In this study, we report the concise synthesis of halicyclamine-inspired macrocyclic mimics and demonstrate activity consistent with DedA-associated modulation, thereby supporting their relevance for host-targeted antiviral strategies.
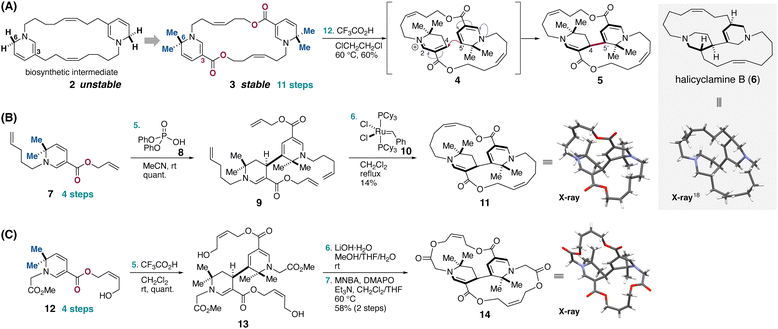
It has been proposed that halicyclamines are biosynthesized via an intramolecular transannular cyclization of a C2-symmetric macrocyclic hypothetical intermediate 2 bearing a pair of 1,6-dihydropyridine (DHP) units.14 Baldwin and co-workers chemically synthesized this highly unstable intermediate 2 and attempted transannular cyclizations; however, disproportionation of the 1,6-DHP units predominated, resulting in rapid decomposition within 1 h (Scheme 1A).15 To overcome this problem, we strategically stabilized the 1,6-DHP moiety while preserving its reactivity toward formation of the bis-nitrogenated bicyclic scaffold. Specifically, 1,6-DHP 3 was designed to incorporate a geminal dimethyl group at C6 to completely suppress hydride transfer from the 1,6-DHP unit, together with an electron-withdrawing ester substituent at C3 to enhance oxidative stability.16 Upon treatment with a Brønsted acid catalyst, macrocyclic C2-symmetric intermediate 3 underwent a biomimetic, regio-controlled transannular cyclization between C4 and C5′ of the two DHP units via an iminium intermediate 4, affording the halicyclamine-type macrocycle 5.17 This compound 5 features a macrocyclic ring of the same size as halicyclamine B (6).18 Despite this success, the previously reported route required 12 synthetic steps in total.
In this study, we report a redesigned synthetic approach that enables rapid and customizable access to bis-macrocyclic halicyclamine analogues exhibiting activity consistent with DedA-associated modulation. To this end, we examined two alternative macrocyclization strategies, beginning with a ring-closing olefin metathesis (RCM)-based approach (Scheme 1B). When 1,6-DHP 7, readily synthesized in four steps, was subjected to a Brønsted acid-catalyzed regio-controlled dimerization between the C4 and C5′ positions, the corresponding halicyclamine-type framework 9 was obtained in quantitative yield. Subsequent RCM using first-generation Grubbs catalyst 10 furnished halicyclamine-type bis-macrocycle 11 in six steps. The modest yield of this transformation is likely attributable to competing metathesis pathways during the two sequential RCM events, leading to undesired macrocycles and related byproducts.15c Single-crystal X-ray diffraction analysis confirmed that 11 adopts a three-dimensional architecture closely resembling that of halicyclamine A (1) and B (6)18 (see Fig. S3–S4 for overlays of these structures). Overall, this approach reduced the total number of steps by approximately half compared with our previous biomimetic route17 (Scheme 1A).
As a complementary strategy to olefin metathesis, we next explored a macrolactonization-based approach for the construction of 14 (Scheme 1C). Brønsted acid-catalyzed dimerization of 1,6-DHP 12, which was likewise readily synthesized in four steps, afforded halicyclamine-type dimer 13 in quantitative yield. Subsequent site-selective hydrolysis of the two methyl ester groups in 13, followed by macrolactonization under 2-methyl-6-nitrobenzoic anhydride (MNBA) and 4-dimethylaminopyridine N-oxide (DMAPO) conditions,19 provided halicyclamine-type bis-macrocycle 14 in seven steps. Single-crystal X-ray diffraction analysis confirmed that 14 similarly reproduces the three-dimensional architecture of halicyclamine A (1) and B (6)18 (Fig. S3–S4). Notably, both alternative synthetic routes (Scheme 1B and C) strategically integrate our biomimetic dimerization platform with site-selective and operationally straightforward macrocyclization methods. These routes are substantially shorter than our previously reported synthesis of halicyclamine analogue 5 and provide rapid access to bis-macrocyclic alkaloidal scaffolds that faithfully recapitulate the natural product architecture while enabling controlled structural diversification.20
With a concise and modular synthetic route established, we next examined whether the synthesized halicyclamine analogues functionally engage DedA membrane proteins in a cellular context. Accordingly, their inhibitory activities were evaluated using an established DedA overexpression assay in Mycobacterium smegmatis and benchmarked against representative members of manzamine alkaloid family (Fig. 2). Following the reported protocol,8 a DedA-overexpressing M. smegmatis strain was prepared, and its responses to each compound were compared with those of the corresponding wild-type strain using an MTT-based viability assay.21 In this assay format, compounds that exhibit reduced inhibitory effects against the DedA-overexpressing strain relative to the wild-type strain are interpreted as functionally engaging DedA membrane proteins, consistent with prior genetic and chemical validation studies.8 Halicyclamine A (1) was not available in sufficient quantity from natural sources; therefore, halicyclamine B (6),18 which is closely related in structure to the synthesized analogues, was selected as the reference compound.22 The natural product halicyclamine B (6) displayed weak growth inhibition against the wild-type strain, with an MIC value of 100 µM, whereas no inhibition was observed against the DedA-overexpressing strain at concentrations up to 100 µM. Although the absolute potency was modest, this differential response indicates a discernible resistance shift associated with DedA overexpression. The concisely synthesized halicyclamine-type macrocycle 11 showed a closely comparable profile, exhibiting an MIC of 100 µM against the wild-type strain and no detectable inhibition of the DedA-overexpressing strain up to 100 µM, resulting in a resistance shift similar in magnitude to that of halicyclamine B (6). In contrast, the halicyclamine analogue 14, assembled via macrolactonization, displayed enhanced inhibitory activity, with MIC values of 30 µM against the wild-type strain and 60 µM against the DedA-overexpressing strain. The larger separation between these values corresponds to a more pronounced resistance shift, exceeding that observed for the natural product 6, and is consistent with enhanced DedA-related activity under these assay conditions. By comparison, macrocycle 15 lacking the central transannular bond17 and the macrocyclization precursors 9 and 13 showed no detectable inhibitory activity against either strain. For reference, manzamine A (16), a representative macrocyclic marine alkaloid,23 was also evaluated and exhibited no detectable inhibitory activity under the same assay conditions. Collectively, these results suggest that DedA-associated growth inhibition in M. smegmatis is preferentially linked to halicyclamine-type bis-macrocyclic scaffolds and appears to depend on both the bis-nitrogenated core framework and the bis-macrocyclic architecture.
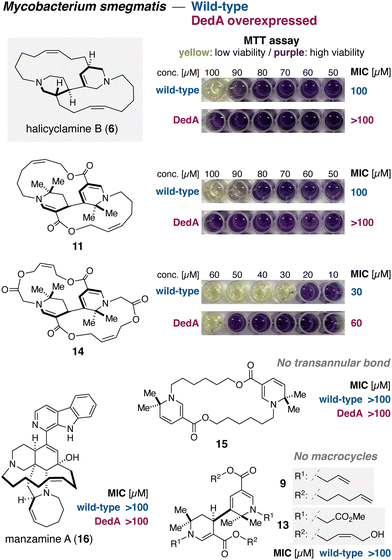 | ||
| Fig. 2 Evaluation of inhibitory activity toward wild-type and DedA-overexpressed Mycobacterium smegmatis. | ||
To gain initial insight into whether the designed halicyclamine-type analogues translate into functional antiviral activity, the anti-SARS-CoV-2 replication activities of the synthetic analogues were evaluated in a cell-based SARS-CoV-2 infection assay (Fig. 3). The synthetic bis-macrocyclic analogues 11 and 14 showed antiviral potency, with IC50 values of 1.7 and 1.4 µM, respectively. In contrast, both macrocycle 15, which lacks the central transannular bond,17 and macrocyclization precursor 13 displayed only weak replication-inhibitory activity (IC50 > 10 µM) under the same conditions. This preliminary structure–activity relationship suggests that the bis-macrocyclic architecture, particularly the presence of the central transannular bond, substantially enhances antiviral potency. Collectively, these findings highlight halicyclamine-inspired bis-macrocyclic scaffolds as promising lead structures for antiviral drug discovery.
In summary, bis-macrocyclic analogues 11 and 14 were newly designed on the basis of halicyclamine A (1) and rapidly synthesized via a biomimetic dimerization strategy. DedA-associated activity of these analogues was supported by the differential responses observed between wild-type and DedA-overexpressing Mycobacterium smegmatis strains. In line with this DedA-associated activity, these halicyclamine-inspired analogues 11 and 14 potently suppressed SARS-CoV-2 replication in a cell-based infection assay.
Host DedA membrane proteins remain largely unexplored as targets of antiviral drug development. The present findings therefore suggest that these natural product-inspired analogues represent promising starting points for further investigation. Although DedA-associated activity was evaluated in this study using an established M. smegmatis-based assay, DedA homologues in humans and mycobacteria are expected to differ in sequence and structure. Moreover, SARS-CoV-2 replication is a complex process involving multiple host DedA-related homologues and viral proteins,5 and the activity responses observed in the M. smegmatis-based assay may therefore not directly correspond to antiviral activity against SARS-CoV-2. Future work will focus on assessing inhibitory activity against human DedA homologues to clarify the translational potential of this strategy for the development of new host-directed antiviral agents.
T. W. and H. Oguri designed the project. T. W. performed the organic synthesis. H. Ohashi and K. W. assessed antiviral activity against SARS-CoV-2, while T. W., R. H., and M. A. conducted assays using Mycobacterium smegmatis. T. W., S. Y., and S. S. performed X-ray crystallographic analysis. S. T. provided the authentic samples of natural products. T. W. and H. Oguri wrote the manuscript with input from all authors.
Conflicts of interest
There are no conflicts to declare.Data availability
The data underlying this study are available in the published article and its supporting information (SI). See DOI: https://doi.org/10.1039/d6cc01052k.CCDC 2519925 (11) and 2519926 (14) contain the supplementary crystallographic data for this paper.24a,b
Acknowledgements
This work was partially supported by JSPS KAKENHI (JP22H00346 and JP22H05127), the Astellas Foundation for Research on Metabolic Disorders, the Naito Foundation, and the Asahi Glass Foundation (H. Oguri). Additional support was provided by JSPS KAKENHI (JP25K21782) and JST MIRAI program (JPMJMI22G1) (K.W.) as well as JSPS KAKENHI (JP24K18457) (H. Ohashi). T.W. is a Research Fellow of JSPS DC2 and gratefully acknowledges financial support from JSPS KAKENHI (JP24KJ0618) to complete this research work. T. W. also acknowledges the Program for Leading Graduate Schools, World-leading Innovative Graduate Study Program for Materials Research, Industry and Technology (MERIT-WINGS). We were granted access to the X-ray diffraction instruments at KEK Photon Factory BL17A beamline (2023G604). This work was further inspired by the international and interdisciplinary environments of the JSPS Asian CORE Program, “Asian Chemical Biology Initiative” as well as the JSPS A3 Foresight Program. Figures were created using BioRender. We are grateful to Dr Nobuto Kaneko (UTokyo) for his advice on the DFT calculations.References
- For reviews, see: (a) A. R. Fehr and S. Perlman, Methods Mol. Biol., 2015, 1282, 1 CrossRef CAS PubMed; (b) P. V’kovski, A. Kratzel, S. Steiner, H. Stalder and V. Thiel, Nat. Rev. Microbiol., 2021, 19, 155 CrossRef PubMed; (c) H. Yang and Z. Rao, Nat. Rev. Microbiol., 2021, 19, 685 CrossRef CAS PubMed; (d) V. C. Soares, I. B. G. Moreira and S. S. G. S. Dias, Viruses, 2025, 17, 1064 CrossRef CAS PubMed.
- (a) F. Moretti, P. Bergman, S. Dodgson, D. Marcellin, I. Claerr, J. M. Goodwin, R. DeJesus, Z. Kang, C. Antczak, D. Begue, D. Bonenfant, A. Graff, C. Genoud, J. S. Reece-Hoyes, C. Russ, Z. Yang, G. R. Hoffman, M. Mueller, L. O. Murphy, R. J. Xavier and B. Nyfeler, EMBO Rep., 2018, 19, e45889 CrossRef PubMed; (b) K. Morita, Y. Hama, T. Izume, N. Tamura, T. Ueno, Y. Yamashita, Y. Sakamaki, K. Mimura, H. Morishita, W. Shihoya, O. Nureki, H. Mano and N. Mizushima, J. Cell Biol., 2018, 217, 3817 CrossRef CAS PubMed; (c) C. J. Shoemaker, T. Q. Huang, N. R. Weir, N. J. Polyakov, S. W. Schultz and V. Denic, PLoS Biol., 2019, 17, e2007044 CrossRef CAS PubMed; (d) J. D. Trimarco, B. E. Heaton, R. R. Chaparian, R. A. Binder, G. C. Gray, V. D. Menachery and N. S. Heaton, PLoS Pathog., 2021, 17, e1009599 CrossRef CAS PubMed.
- (a) N. J. Dusetti, Y. Jiang, M. I. Vaccaro, R. Tomasini, S. A. Azizi, E. L. Calvo, A. Ropolo, F. Fiedler, G. V. Mallo, J.-C. Dagorn and J. L. Iovanna, Biochem. Biophys. Res. Commun., 2002, 290, 641 CrossRef CAS PubMed; (b) A. Ropolo, D. Grasso, R. Pardo, M. L. Sacchetti, C. Archange, A. L. Re, M. Seux, J. Nowak, C. D. Gonzalez, J. L. Iovanna and M. I. Vaccaro, J. Biol. Chem., 2007, 282, 37124 CrossRef CAS PubMed; (c) Y. Tian, Z. Li, W. Hu, H. Ren, E. Tian, Y. U. Zhao, Q. Lu, X. Huang, P. Yang, X. Li, X. Wang, A. L. Kovács, L. Yu and H. Zhang, Cell, 2010, 141, 1042 CrossRef CAS PubMed; (d) E. Itakura and N. Mizushima, J. Cell Biol., 2011, 192, 17 CrossRef CAS PubMed; (e) C. Kishi-Itakura, I. Koyama-Honda, E. Itakura and N. Mizushima, J. Cell Sci., 2014, 127, 4089 CrossRef CAS PubMed.
- (a) A. Ghanbarpour, D. P. Valverde, T. J. Melia and K. M. Reinisch, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2101562118 CrossRef CAS PubMed; (b) D. Huang, B. Xu, L. U. Liu, L. Wu, Y. Zhu, A. Ghanbarpour, Y. Wang, F.-J. Chen, J. Lyu, Y. Hu, Y. Kang, W. Zhou, Y. Wang, X. Ding, W. Li, Z. Jiang, Z. Chen, X. Zhang, H. Zhou, J. Z. Li, C. Guo, W. Zheng, X. Zhang, P. Li, T. Melia, K. Reinisch and X.-W. Chen, Cell Metab., 2021, 33, 1655 CrossRef CAS PubMed; (c) Y. E. Li, Y. Wang, X. Du, T. Zhang, H. Y. Mak, S. E. Hancock, H. McEwen, E. Pandzic, R. M. Whan, Y. C. Aw, I. E. Lukmantara, Y. Yuan, Z. Dong, A. Don, N. Turner, S. Qi and H. Yang, J. Cell Biol., 2021, 220, e202103105 CrossRef CAS PubMed; (d) K. M. Reinisch, X.-W. Chen and T. J. Melia, Contact, 2021, 4, 25152564211024494 CrossRef PubMed; (e) T. Zhang, Y. E. Li, Y. Yuan, X. Du, Y. Wang, X. Dong, H. Yang and S. Qi, Autophagy, 2021, 17, 2048 CrossRef CAS PubMed.
- For review, see: Y. Hama, H. Morishita and N. Mizushima, EMBO Rep., 2022, 23, e53894 CrossRef CAS PubMed.
- (a) J. Baggen, L. Persoons, E. Vanstreels, S. Jansen, D. V. Looveren, B. Boeckx, V. Geudens, J. D. Man, D. Jochmans, J. Wauters, E. Wauters, B. M. Vanaudenaerde, D. Lambrechts, J. Neyts, K. Dallmeier, H. J. Thibaut, M. Jacquemyn, P. Maes and D. Daelemans, Nat. Genet., 2021, 53, 435 CrossRef CAS PubMed; (b) H. H. Hoffmann, W. M. Schneider, K. Rozen-Gagnon, L. A. Miles, F. Schuster, B. Razooky, E. Jacobson, X. Wu, S. Yi, C. M. Rudin, R. M. MacDonald, L. K. McMullan, J. T. Poirier and C. M. Rice, Cell, 2021, 184, 133 CrossRef CAS PubMed.
- For reviews, see: (a) W. Cui, Y. Duan, Y. Gao, W. Wang and H. Yang, Structure, 2024, 32, 1301–1320 CrossRef CAS PubMed; (b) T. Kronenberger, S. A. Laufer and T. Pillaiyar, Drug Discovery Today, 2023, 28, 103579 CrossRef CAS PubMed.
- M. Arai, L. Liu, T. Fujimoto, A. Setiawan and M. Kobayashi, Mar. Drugs, 2011, 9, 984 CrossRef CAS PubMed.
- (a) R. Sakai, T. Higa, C. W. Jefford and G. Bernardinelli, J. Am. Chem. Soc., 1986, 108, 6404 CrossRef CAS; (b) J. I. Kobayashi, M. Tsuda, N. Kawasaki, K. Matsumoto and T. Adachi, Tetrahedron Lett., 1994, 35, 4383 CrossRef CAS; (c) M. Jaspars, V. Pasupathy and P. Crews, J. Org. Chem., 1994, 59, 3253 CrossRef CAS.
- For reviews, see: (a) E. Magnier and Y. Langlois, Tetrahedron, 1998, 54, 6201 CrossRef CAS; (b) J.-F. Hu, M. T. Hamann, R. Hill and M. Kelly, Alkaloids, 2003, 60, 207 CAS; (c) R. Duval and E. Poupon, Biomimetic Organic Synthesis, Wiley-VCH, 2011 Search PubMed; (d) T. Kubota, S. Kurimoto and J. Kobayashi, Alkaloids, 2020, 84, 1 CAS.
- (a) K. K. H. Ang, M. J. Holmes, T. Higa, M. T. Hamann and U. A. K. Kara, Antimicrob. Agents Chemother., 2000, 44, 1645 CrossRef CAS PubMed; (b) K. V. Rao, B. D. Santarsiero, A. D. Mesecar, R. F. Schinazi, B. L. Tekwani and M. T. Hamann, J. Nat. Prod., 2003, 66, 823 CrossRef CAS PubMed; (c) J. D. Winkler, A. T. Londregan and M. T. Hamann, Org. Lett., 2006, 8, 2591 CrossRef CAS PubMed.
- M. Arai, M. Sobou, C. Vilchéze, A. Baughn, H. Hashizume, P. Pruksakorn, S. Ishida, M. Matsumoto, W. R. Jacobs and M. Kobayashi, Bioorg. Med. Chem., 2008, 16, 6732 CrossRef CAS PubMed.
- (a) K. A. El Sayed, M. Kelly, U. A. K. Kara, K. K. H. Ang, I. Katsuyama, D. C. Dunbar, A. A. Khan and M. T. Hamann, J. Am. Chem. Soc., 2001, 123, 1804 CrossRef CAS PubMed; (b) S. Matsunaga, Y. Miyata, R. W. M. Van Soest and N. Fusetani, J. Nat. Prod., 2004, 67, 1758 CrossRef CAS PubMed; (c) D. G. Jimenez, V. Poongavanam and J. Kihlberg, J. Med. Chem., 2023, 66, 5377 CrossRef PubMed; (d) F. Lovering, J. Bikker and C. Humblet, J. Med. Chem., 2009, 52, 6752 CrossRef CAS PubMed.
- (a) A. Kaiser, X. Billot, A. Gateau-Olesker, C. Marazano and B. C. Das, J. Am. Chem. Soc., 1998, 120, 8026 CrossRef CAS; (b) J.-C. Wypych, T. M. Nguyen, P. Nuhant, M. Bénéchie and C. Marazano, Angew. Chem., Int. Ed., 2008, 47, 5418 CrossRef CAS PubMed; (c) I. Sinigaglia, T. M. Nguyen, J.-C. Wypych, B. Delpech and C. Marazano, Chem. – Eur. J., 2010, 16, 3594 CrossRef CAS PubMed; (d) Y. Tang, L. Zhu and R. Hong, Tetrahedron Chem, 2022, 3, 100025 CrossRef CAS.
- (a) J. E. Baldwin and R. C. Whitehead, Tetrahedron Lett., 1992, 33, 2059 CrossRef CAS; (b) J. E. Baldwin, T. D. W. Claridge, A. J. Culshaw, F. A. Heupel, V. Lee, D. R. Spring, R. C. Whitehead, R. J. Boughtflower, I. M. Mutton and R. J. Upton, Angew. Chem., Int. Ed., 1998, 37, 2661 CrossRef CAS; (c) J. E. Baldwin, T. D. W. Claridge, A. J. Culshaw, F. A. Heupel, V. Lee, D. R. Spring and R. C. Whitehead, Chem. – Eur. J., 1999, 5, 3154 CrossRef; (d) A. T. Piwko, B. G. Miller and J. M. Smith, Nat. Prod. Rep., 2023, 40, 964 RSC; (e) A. Leblond, A. Nguyen, C. Alcover, K. Leblanc, J.-F. Gallard, D. Joseph, E. Poupon and M. A. Beniddir, Org. Lett., 2024, 26, 2163 CrossRef CAS PubMed; (f) S. I. Gómez, E. I. O. Nava, A. Dadawalla, D. Ashong, G. Chen and Q.-H. Chen, Molecules, 2025, 30, 2907 CrossRef PubMed.
- (a) T. Wayama, Y. Arai and H. Oguri, J. Org. Chem., 2022, 87, 5938 CrossRef CAS PubMed; (b) T. Wayama and H. Oguri, Org. Lett., 2026, 28, 982 CrossRef CAS PubMed.
- T. Wayama and H. Oguri, Org. Lett., 2023, 25, 3596 CrossRef CAS PubMed.
- B. Harrison, S. Talapatra, E. Lobkovsky, J. Clardy and P. Crews, Tetrahedron Lett., 1996, 37, 9151 CrossRef CAS.
- I. Shiina, H. Fukui and A. Sasaki, Nat. Protoc., 2007, 2, 2312 CrossRef CAS PubMed.
- Selected examples for synthetic studies, see: (a) G. A. Molander and F. Cadoret, Tetrahedron Lett., 2011, 52, 2199 CrossRef CAS PubMed; (b) A. I. Lansakara, S. V. S. Mariappan and F. C. Pigge, J. Org. Chem., 2016, 81, 10266 CrossRef CAS PubMed; (c) M. G. Banwell, M. J. Coster, N. L. Hungerford, M. J. Garson, S. Su, A. C. Kotzed and M. H. G. Munro, Org. Biomol. Chem., 2012, 10, 154 RSC; (d) A. G. Dalling, G. Späth and A. Fürstner, Angew. Chem., Int. Ed., 2022, 61, e202209651 CrossRef CAS PubMed; (e) B. J. Smith and G. A. Sulikowski, Angew. Chem., Int. Ed., 2010, 49, 1599 CrossRef CAS PubMed; (f) B. J. Smith, T. Qu, M. Mulder, M. J. Noetzel, C. W. Lindsley and G. A. Sulikowski, Tetrahedron, 2010, 66, 4805 CrossRef CAS; (g) Z. D. Firestone, T. A. Grigolo, F. G. Pernichelle and J. M. Smith, J. Am. Chem. Soc., 2026, 148, 6814 CrossRef CAS PubMed.
- (a) T. Mosmann, J. Immunol. Methods, 1983, 65, 55 CrossRef CAS PubMed; (b) G. Abate, A. Aseffa, A. Selassie, S. Goshu, B. Fekade, D. WoldeMeskal and H. Miörner, J. Clin. Microbiol., 2004, 42, 871 CrossRef CAS PubMed.
- H. Kato, A. H. El-Desoky, Y. Takeishi, T. Nehira, E. D. Angkouw, R. E. P. Mangindaan, N. J. de Voogd and S. Tsukamoto, Bioorg. Med. Chem. Lett., 2019, 29, 8 CrossRef CAS PubMed.
- (a) K. Eguchi, Y. Fujiwara, A. Hayashida, H. Horlad, H. Kato, H. Rotinsulu, F. Losung, R. E. P. Mangindaan, N. J. de Voogd, M. Takeya and S. Tsukamoto, Bioorg. Med. Chem., 2013, 21, 3831 CrossRef CAS PubMed; (b) J. R. Palem, G. R. Bedadala, K. A. El Sayed and S.-C. V. Hsia, Planta Med., 2011, 77, 46 CrossRef CAS PubMed; (c) M. Yousaf, N. L. Hammond, J. Peng, S. Wahyuono, K. A. McIntosh, W. N. Charman, A. M. S. Mayer and M. T. Hamann, J. Med. Chem., 2004, 47, 3512 CrossRef CAS PubMed.
- (a) CCDC 2519925: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2ql5xf; (b) CCDC 2519926: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2ql5yg.
Footnote |
| † Present address: JEOL Ltd., 3-1-2 Musashino, Akishima, Tokyo 196-8558, Japan. |
| This journal is © The Royal Society of Chemistry 2026 |