 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Precision-engineered polymer topologies: balancing potent antibacterial efficacy with enhanced biocompatibility
Md
Aquib
,
Vinod Kumar
Kannaujiya
and
Cyrille
Boyer
 *
*
Cluster for Advanced Macromolecular Design (CAMD) and Australian Centre for NanoMedicine (ACN), School of Chemical Engineering, UNSW Australia, Sydney, NSW 2052, Australia. E-mail: cboyer@unsw.edu.au
First published on 17th April 2026
Abstract
The escalating threat of antimicrobial resistance demands the development of innovative therapeutic strategies. Host-defense peptides (HDPs) and their synthetic mimics exhibit broad-spectrum antibacterial activity, yet their clinical translation is frequently hindered by cytotoxicity arising from nonspecific interactions with mammalian cell membranes. Recent advances in polymer science, particularly controlled polymerization techniques, have enabled the precise synthesis of well-defined antibacterial polymers and systematic evaluation of structure–activity relationships. This review provides a comprehensive overview of the development and fundamental mechanisms of antibiotic resistance and highlights the evolution of antibacterial polymers from HDP-inspired molecular designs toward precise biofunction control. Key advances in optimizing structural determinants governing antibacterial activity and biocompatibility are discussed, with particular emphasis on compositional parameters and polymer topology, including linear, cyclic, hyperbranched, star, and bottle brush architectures. In addition, emerging approaches based on stimuli-responsive and self-immolative polymers that enable on-demand degradation are presented as effective means to reduce long-term cytotoxicity and environmental accumulation. Concurrently, increasing attention has been directed toward non-amphiphilic polymer systems, which expand the current design landscape by offering alternative pathways to antibacterial activity beyond conventional amphiphilic design principles. Finally, future perspectives involving synergistic and combination therapies, together with data-driven and AI-assisted approaches for pathogen-specific polymer design, are outlined as promising routes to improve selectivity while maintaining or even enhancing antimicrobial efficacy.
Introduction
The increasing prevalence of infections caused by multidrug-resistant (MDR) bacterial and fungal pathogens in recent years represents a serious threat to human health.1–3 This global health concern has been largely driven by the widespread misuse and overuse of antibiotics,4,5 resulting in the consumption of over 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 tons of these drugs annually across both human and veterinary sectors.6–9
000 tons of these drugs annually across both human and veterinary sectors.6–9
To contextualize the global healthcare challenge known as antimicrobial resistance (AMR), approximately 1.14 million deaths in 2021 were directly linked to MDR bacteria, and projections indicate that this number may nearly double to 1.91 million by 2050.10 AMR not only contributes to substantial mortality but also disrupts the global economy, potentially costing the European economy €11.7 billion annually due to prolonged hospitalizations if innovative solutions are not developed.11 Alarmingly, a 2022 report by the World Health Organization concluded that current and pipeline antibiotics are inadequate to address the rapidly evolving AMR crisis.12 Even among the nine new antibiotics approved for “critical” bacterial pathogens, resistance has already emerged in most cases.13,14 This issue is concerning for Gram-negative bacteria, especially MDR strains, which are often linked to life-threatening infections and are notoriously more difficult to treat than Gram-positive bacteria.15–20 These developments collectively reinforce the urgent need for innovative strategies and intensified research into novel antibacterial agents.21
One promising approach to addressing infections caused by Gram-negative and MDR bacteria involves the use of host-defense peptides (HDPs), also known as antimicrobial peptides.22–24 These molecules are key components of the innate immune system in multicellular organisms and serve as natural defenses against invading pathogens.25 HDPs typically consist of amino acid residues that include cationic, hydrophobic, and hydrophilic groups, allowing them to form amphipathic structures.25–27 These structures, characterized by an overall positive charge, primarily exert bactericidal effects by disrupting bacterial membranes.25–27 Membrane disruption by HDPs is generally driven by their cationic and amphipathic nature. The positively charged groups initially interact with the negatively charged bacterial membranes via electrostatic interactions and/or hydrogen bonding. This is followed by the insertion of hydrophobic regions into the membrane, compromising its integrity and ultimately leading to cell lysis and bacterial death.28,29 This broad and non-specific mechanism of action may also reduce the likelihood of resistance development against HDPs.26,30–32
Despite their therapeutic potential, HDPs face several limitations that hinder widespread clinical application, including complex and costly synthesis, susceptibility to endogenous and/or exogenous peptidases, and toxicity to host cells.24,33,34 To address these issues, antimicrobial polymers (APs) have emerged as promising synthetic analogs of HDPs. However, achieving high antibacterial potency in APs often comes at the expense of increased cytotoxicity, as excessive cationic charge density or hydrophobic content can disrupt mammalian membranes and cause hemolysis.35–40 Therefore, the rational design of APs requires careful optimization of the balance between bactericidal activity and cytocompatibility to enhance selectivity toward bacterial cells.41–43
Thanks to advances in polymer chemistry—particularly the development of reversible deactivation radical polymerization (RDRP) techniques such as atom transfer radical polymerization (ATRP) and reversible addition–fragmentation chain transfer (RAFT) polymerization, have enabled the scalable synthesis of well-defined polymers that are more resistant to proteolysis and economically viable to produce.24,44–51 These controlled polymerization strategies offer precise control over amphiphilic balance, molecular weight, dispersity, composition, and overall macromolecular topology, thereby allowing fine-tuning of APs to address the limitations associated with both natural HDPs and conventional synthetic polymer analogs.52–58
Among various molecular design parameters, polymer topology has emerged as a particularly influential factor. Non-linear architectures—such as hyperbranched (HP),59 cyclic,60,61 star (SP),56,57 and bottle brush polymers,62,63 often exhibit superior biocompatibility and selective antibacterial activity compared to their linear counterparts.24,59,60,64 Despite the growing research interest in topology-driven polymeric materials, most prior investigations have predominantly focused on linear amphiphilic copolymers.27,46,49,54,64,65 Consequently, systematic studies comparing the effects of distinct polymer topologies on antibacterial efficacy and biocompatibility remain limited. Moreover, several reports have highlighted that the long-term accumulation of synthetic polymers in vivo and their potential release into the environment after administration remain significant concerns.66–68 To mitigate these challenges, stimuli-responsive APs that undergo degradation in response to biological or external stimuli such as enzymatic activity, redox conditions, light, or pH could emerge as a promising strategy,67,69,70 expanding their applications from drug-delivery systems to antimicrobial materials.67,71–73 Upon degradation, these systems yield low-molecular-weight fragments, that can be readily cleared from the body, thereby reducing systemic accumulation and minimizing toxicity.67,71
Consequently, the development of APs has accelerated significantly, leading to the emergence of a wide variety of polymeric systems with diverse structural and functional attributes. Several recent reviews have examined APs from multiple perspectives, including molecular design principles,24,74,75 antimicrobial mechanisms,24,76,77 bacterial uptake and killing pathways,76 antibiofilm strategies,78,79 advanced antibacterial polymeric nanostructures,80 and polypeptide-based systems.77,81–83 Building on these important contributions, the present review provides a more integrated and focused perspective, with a specific emphasis on synthetic APs to enable a targeted and in-depth analysis of the field. In particular, this review encompasses the fundamental mechanisms of antibiotic resistance together with the evolutionary trajectory of antibacterial polymers from HDP-inspired structural design toward precise biofunction control. Key advances in the optimization of structural determinants governing antibacterial efficacy and biocompatibility, including compositional parameters and polymer topology, are critically discussed. In addition, emerging strategies based on stimuli-responsive and self-immolative polymers are highlighted, as these systems enable on-demand degradation and offer a promising route to mitigate long-term cytotoxicity, alongside recent developments in non-amphiphilic polymer systems that exhibit distinct antibacterial mechanisms. Finally, prospective design directions, including synergistic and combination therapeutic strategies as well as data-driven and AI-assisted approaches for pathogen-specific polymer design, are outlined. These emerging paradigms hold significant promise for overcoming the persistent challenge of toxicity while maintaining or even enhancing antimicrobial potency, thereby improving overall therapeutic selectivity.
Development and mechanisms of antibiotic resistance in bacteria
Over the past decades, bacterial infectious diseases caused by multidrug-resistant (MDR) pathogens have posed a significant threat to global public health, remaining a major cause of mortality and economic burden despite extensive research efforts and substantial financial investment.19,84,85 While resistance development in bacteria is a natural evolutionary process, the accelerated emergence of MDR strains is largely driven by human activities. The overuse and misuse of antibiotics in both healthcare and agriculture, coupled with inadequate sanitation, limited access to clean water, and insufficient infection control measures, have created environments conducive to the rapid spread of resistant bacteria.86–90 There are pathogenic MDR bacteria that have acquired resistance to multiple antimicrobial categories, extensively drug-resistant (XDR) bacteria that are susceptible to only one or two antimicrobial drug categories, and pandrug-resistant (PDR) bacteria that are resistant to all clinically available drugs.91,92Although new antimicrobial agents continue to be developed and incorporated into medical treatment (Fig. 1),93 most represent only minor structural modifications of existing antibiotics. As a result, they tend to be ineffective against XDR and PDR strains because they share the same molecular targets and mechanisms of action as their predecessors.93,94 Antibiotics typically act by inhibiting key bacterial processes including: (I) cell wall synthesis, (II) DNA/RNA synthesis (III) protein synthesis, and (IV) folic acid metabolism (Fig. 2).94,95 However, bacteria have evolved a wide array of resistance mechanisms, rendering nearly all existing antibiotics increasingly ineffective. The complexity of the mechanism of action, potency, and efficacy factors such as duration of action or concentration determines whether a novel antibiotic can develop resistance.94
 | ||
| Fig. 1 Timeline of antibiotic development and the subsequent emergence of drug-resistant bacteria. Adapted from ref. 84 under the terms of the CC BY 4.0 license, American Chemical Society, copyright 2024, and modified to include recent updates. | ||
Bacteria can develop resistance to antibiotics through multiple pathways or mechanisms, broadly classified into three main types: intrinsic, acquired, and adaptive resistance.19,96–98 Intrinsic resistance refers to the innate insensitivity of certain bacterial species to specific antibiotics, typically due to inherent structural or functional characteristics.99 This type of resistance is often encoded by chromosomal genes and remains relatively stable over time. Acquired resistance describes a situation in which susceptible bacteria gain the ability to resist antibiotics by altering their metabolic pathway.98,99 This occurs through gene mutations or through horizontal gene transfer, including transformation, transduction, or conjugation.99 Finally, adaptive resistance is a transient, non-heritable state that allows bacteria to tolerate antibiotics under environmental stress, such as exposure to sub-lethal antibiotic concentrations or nutrient limitations.97 This form of resistance dissipates once the stressor is removed. Despite differences in origin and persistence, all resistance types generally act through a set of core molecular mechanisms.19,100–102 These are (A) impermeability of antibiotics (exclusive to Gram-negative specie), (B) efflux pump-out, (C) altered target site, (D) target overproduction, (E) enzymatic inactivation or modification, and (F) bypass pathway (Fig. 2).84,98,100
In addition to these mechanisms, the role of biofilms in AMR is both complex and clinically significant. Bacteria within biofilms can be 10- to 1000-fold more resistant to antibiotics than their planktonic counterparts.103 This enhanced resistance is largely attributed to the extracellular polymeric substance, a protective three-dimensional matrix that acts as a physical barrier to antimicrobial agents. Moreover, the presence of persister cells and the intrinsic heterogeneity within biofilm communities further promote bacterial survival under antibiotic stress. Collectively, these adaptive features account for nearly 75% of bacterial infections in clinical settings.104,105 Such growing challenges underscore the urgent need for resistance-proof drug discovery and the development of effective novel antibacterials and alternative therapeutic strategies.
From host-defense peptides to antimicrobial polymer: bridging nature and design
Host-defense peptides
HDPs, also known as antimicrobial peptides (AMPs), are key components of innate immunity in multicellular organisms. Boman and colleagues106 were the first to discover HDPs, identifying cecropins in the humoral immune system of the silk moth (Hyalophora cecropia). These peptides exhibited broad-spectrum activity against pathogens, including Gram-positive and Gram-negative bacteria as well as fungi.106 Since this landmark discovery in 1981, the field of HDPs has expanded rapidly, with thousands of peptides now catalogued in the AMP database (APD6: https://aps.unmc.edu/).Structurally, HDPs are short amphiphilic peptides composed of 10–50 amino acids and characterized by a net positive charge.26 Their sequences usually include cationic residues (e.g., lysine, arginine), hydrophobic residues (e.g., leucine, valine), and neutral hydrophilic residues (e.g., serine, asparagine). This composition enables them to adopt amphipathic conformations that selectively target bacterial membranes.26 The cationic residues promote electrostatic interactions with negatively charged bacterial surfaces, while hydrophobic residues insert into lipid domains to destabilize the bilayer, and neutral hydrophilic residues enhance solubility and stabilize secondary structures, thereby supporting efficient membrane interaction.31,107–110 Notably, their bactericidal activity relies on membrane lysis, a physical mechanism that bacteria rarely evade through mutation, making HDPs promising candidates for combating MDR pathogens.26,30–32,52,111,112
Despite their considerable potential, the clinical translation of HDPs remains challenging. Their therapeutic performance is limited by the high costs and technical complexity associated with peptide discovery, optimization, and large-scale production.113,114In vivo efficacy is further hindered by susceptibility to enzymatic degradation and poor pharmacokinetic stability.37,115,116
Synthetic antimicrobial polymers
In parallel with the discovery of HDPs, such as cecropins and magainins, the field of polymer-based disinfectants began to emerge.108,117 These polymers were first applied in surface disinfection, water purification, and textile industries. In 1984, Ikeda synthesized a series of cationic polymers based on poly(vinyl benzyl ammonium chloride) (BAC), which is widely recognized as a breakthrough in the development of cationic APs.117 Like HDPs, these early copolymers combined cationic groups, most commonly quaternary ammonium salts (QAS), with hydrophobic moieties such as long alkyl chains to kill pathogens via membrane disruption (Fig. 3).117 | ||
| Fig. 3 The convergence of concepts from HDPs and polymer disinfectants has inspired the development of peptide-mimetic APs. Representative examples highlight key milestones, from naturally derived peptides (e.g., magainin-2) to early polymer disinfectants (e.g., poly(BAC)) and subsequent advances in β-peptides, biocidal polymers, and HDP-mimic APs. Chemical structures are adapted and modified from ref. 118 under the terms of the CC BY-NC 3.0 license, Royal Society of Chemistry, copyright 2018. | ||
Since then, research on APs has expanded rapidly, and many studies have been reported. The pioneering contributions of Kenawy,118 DeGrado,119 Tew,120 Kuroda,120 Gellman,121 and others demonstrated that polymers bearing both cationic and hydrophobic substituents can mimic HDPs (Fig. 3).79 These polymers electrostatically interact with bacterial membranes, targeting the cytoplasmic membrane in Gram-positive bacteria and the outer membrane in Gram-negative species—leading to pore formation or membrane deformation, causing bacterial lysis with no resistance development.24,79,122
A wide range of polymerization strategies have been employed to develop such HDPs mimic biomaterials. These include conventional radical polymerization as well as advanced methods such as ring-opening polymerization (ROP), and ring-opening metathesis polymerization (ROMP). In addition, RDRP techniques such as ATRP, RAFT, and photoinduced energy/electron transfer-RAFT (PET-RAFT) have been widely applied.24,40,79 These synthetic routes enable the production of biomacromolecules with precise structural control on a large scale and at relatively low cost, while conferring resistance to proteolytic degradation and improved stability and bioavailability under physiological conditions.24,50,107,123
Mechanism of actions: host-defense peptides/antimicrobial polymers
The antimicrobial mechanisms exhibited by HDPs and APs provide valuable insight for the rational design of novel membrane-active agents to combat MDR pathogens. Understanding the structural differences between microbial and mammalian cell membranes is also essential to explaining the preferential activity, or selectivity, of HDPs and APs toward microbial cells. Because different microbial families have distinct cell wall compositions, these variations account for the species-dependent activity of membrane-active macromolecules. The main structural differences between mammalian membranes, Gram-positive and Gram-negative bacteria, and yeast (fungi) are summarized in Fig. 4. | ||
| Fig. 4 Schematic comparison of mammalian and microbial cell membranes. Reproduced from ref. 110 with permission from Elsevier, copyright 2012. | ||
Mammalian cell membranes are largely composed of zwitterionic phospholipids, cholesterol, and glycosaminoglycans, which contribute to an overall neutral surface charge. In contrast, bacterial membranes are enriched in acidic phospholipids and anionic components, giving them a net negative charge that is typically stabilized by divalent cations such as Mg2+ and Ca2+.40 In Gram-positive bacteria, the cell wall is dominated by a thick peptidoglycan layer (∼20–80 nm), accounting for up to 90% of the wall structure. Negatively charged teichoic acids (a polymer stitching peptidoglycan mesh from different layers together), attached to the cytoplasmic membrane and extending outward across the peptidoglycan network, contribute additional anionic character to the cell surface. In Gram-negative bacteria, the peptidoglycan layer is much thinner (∼10 nm, ∼15–20% of the wall) and is shielded by an asymmetric outer membrane enriched in lipopolysaccharides, which further reinforce the overall negative surface charge (Fig. 4).124,125
Yeast, unlike bacteria, lack peptidoglycan and instead possesses a rigid, multilayered cell wall composed of a chitin–glucan network (approximately 140 nm thick) overlaid with mannoproteins (glycoprotein). These glycoproteins contain phosphodiester linkages that impart a uniformly anionic character at physiological pH (Fig. 4).125,126 Despite differences in cell wall architecture, yeast membranes share organizational similarities with bacterial membranes, including a pronounced negative surface charge.77 These structural characteristics render bacterial77,127,128 and fungal77,129,130 cell surfaces more negatively charged than mammalian cell membranes. This strong anionic character acts as a primary target for cationic HDPs and APs, driving passive selectivity toward microbial cells through electrostatic interactions, which facilitates preferential binding and subsequent membrane disruption.24
Building on this understanding, reviews by Spaar et al.,131 Vogel et al.,132 Wang et al.,127 Kuroda et al.,107 and Tang et al.128 have outlined several mechanistic models for HDPs and APs, while a comprehensive 2022 report by Li et al.77 classified these into three major categories encompassing 15 distinct subtypes (Fig. 5). These are: (I) membrane destructive models including barrel-stave model (formation of transmembrane channels/pores),133 carpet model (adsorption of HDPs/APs in parallel with membrane surface like a carpet),134,135 toroidal pore model (formation of pore-lining),136 disordered toroidal pore model,137 electroporation model triggered by asymmetric charge distribution distribution;138 (II) non-destructive membrane disturbance including membrane thinning model,139 nonlytic membrane depolarization model (reduction of membrane potential),140 membrane thickening model,141 anionic phospholipid clustering domains;142 and (III) intracellular targeting mechanisms such as binding nucleic acids,143 inhibiting nucleic acid and protein synthesis,144 inhibiting enzyme activity,145 blocking cell division and proliferation and inhibiting cell wall synthesis (Fig. 5).146,147
 | ||
| Fig. 5 Structured overview of HDP/AP modes of action. Reproduced from ref. 77 with permission from American Chemical Society, copyright 2022. | ||
Among the widely reported mechanisms, membrane-active models—encompassing both destructive disruption and non-destructive disturbance are particularly appealing for the design of APs, as bacterial mutations rarely result in fundamental alterations to membrane architecture. As noted above, these mechanisms primarily rely on electrostatic interactions between the cationic residues of HDPs or synthetic mimetic polymers and the negatively charged microbial membrane. Consistent with their efficacy, 5 of the 7 US FDA-approved HDPs are membrane-active peptides (Table 1).24,148 Such charge-driven interactions explain the selectivity of these agents by enabling preferential targeting of microbial over mammalian membranes; however, excessive cationic density can occasionally cause mammalian cytotoxicity (e.g., hemolysis), thereby reducing selectivity.36,138,149,150
| Peptide name | Type | Activity | Mechanism | Molecular weight (g mol−1) | T 1/2 | FDA approval year |
|---|---|---|---|---|---|---|
| Notes: The antimicrobial peptide gramicidin is limited to topical use due to mammalian cell toxicity,151 while others, such as colistin (polymyxin E) and vancomycin, despite potent antibacterial activity, are associated with nephrotoxicity.152,153 This underscores the challenge of achieving antibacterial selectivity and biocompatibility —the limitation this review seeks to overcome. Methicillin-resistant Staphylococcus aureus (MRSA). Vancomycin-resistant enterococci (VRE). | ||||||
| Colistin | Cyclic lipopeptide | G− bacteria, including MDR strains | Membrane lysis | 1155 | ∼5 h | 1962 |
| Dalbavancin | Lipoglycopeptide | G+ bacteria, including MDR (e.g., MRSA) | Inhibitor of cell wall synthesis | 1817 | ∼14 days | 2014 |
| Daptomycin | Cyclic lipopeptide | G+ bacteria, including MRSA and VRE | Membrane lysis | 1621 | 8–9 h | 2003 |
| Gramicidin | Linear peptide | Gram+ bacteria, some Gram− bacteria; toxic to mammalian cells | Membrane poration | 1882 | — | 1955 |
| Oritavancin | Lipoglycopeptide | G+ bacteria, including MRSA and VRE | (a) Membrane lysis and (b) inhibitor of cell wall synthesis | 1793 | ∼195.4 h | 2014 |
| Telavancin | Lipoglycopeptide | G+ bacteria, including MRSA | (a) Membrane lysis and (b) inhibitor of cell wall synthesis | 1756 | ∼8 h | 2009 |
| Vancomycin | Lipoglycopeptide | G+ bacteria, including MRSA | Inhibitor of cell wall synthesis | 1449 | 7.5 days | 1983 |
Selectivity challenges in host-defense peptides/antimicrobial polymers
The selectivity of HDPs and APs for bacterial over mammalian membranes derives from differences in membrane composition and the spatial organization of surface components (Fig. 4).128 Although the net negative surface charge of bacterial envelopes can be exploited for selective targeting, achieving high specificity within a mammalian host remains challenging. Local bacterial infection sites can strongly influence the binding and antimicrobial activity of HDPs and their synthetic mimics. In addition, the mammalian extracellular matrix and body fluids contain abundant charged species that attenuate electrostatic attractions between cationic peptides and bacterial membranes.31,154–157 Contributors to this screening effect include small inorganic ions and proteins, as well as proteoglycan components such as glycosaminoglycans. Collectively, these factors divert both peptides and polymers into non-productive binding sinks instead of bacterial membranes, thereby reducing apparent selectivity, and increasing toxicity to mammalian cells.The selectivity trade-off is evident from both clinical and model examples. Colistin (polymyxin E), an intravenous peptide antibiotic, displays potent antibacterial activity but also marked nephrotoxicity, which restricts its therapeutic use.152 This toxicity reflects insufficient selectivity, including interactions with membranes of human kidney proximal tubular cells.83,158 Another example is melittin, a 26-amino acid peptide composed of approximately 46% hydrophobic residues and carrying a net positive charge at physiological pH. Consequently, it lyses bacterial cells and human red blood cells (RBCs) at comparable concentrations.159 Similar behavior has been reported for polymethacrylate,35,160 where an increase in the hydrophobic mass fraction enhances hydrophobic interactions at the expense of electrostatic selectivity, leading to nonspecific membrane disruption and pronounced hemolysis.35 Beyond hydrophobicity, excessive positive charge in HDP-mimetic polymers can also induce cytotoxicity and compromise selectivity, thereby limiting their clinical potential.36,37,161
Moreover, microbial cells generally exhibit greater mechanical robustness than mammalian cells, meaning that poorly balanced macromolecular designs may inflict more damage on host cells.110 To address this, AP research prioritizes strong antibacterial activity with minimal mammalian toxicity. Consequently, these considerations have driven systematic optimization of key polymer design parameters to deliver potent and safe APs for biomedical applications.
Structural determinants of antibacterial activity and biocompatibility
Achieving the rational design of synthetic polymers that exhibit high selectivity—effectively targeting and killing microbes while sparing mammalian cells—remains a formidable and major objective. When developing these macromolecules, several key elements must be incorporated. First, the polymer should interact with microbial surfaces. Second, it requires an appropriate density of cationic groups to promote adhesion to the negatively charged bacterial envelope. Third, hydrophobic moieties are needed to enable insertion into or disruption of the cellular membrane. In addition, these macromolecules must discriminate between microbial and mammalian cells to avoid host toxicity.110A range of molecular factors collectively determine bioactivity of synthetic polymers. These include amphiphilic balance, cationic charge density, backbone chemistry, charge distribution, hydrophobicity, hydrophilicity, end-group functionality, molecular weight, sequence definition, and polymer architecture. In the following sections, APs are discussed as representative systems, with insights drawn from the literature to identify critical structural and compositional parameters that can guide the rational design of next-generation membrane-active APs with enhanced efficacy and biocompatibility.
Amphiphilic balance
Optimizing the balance between cationic charge and hydrophobic content, often referred to as the amphiphilic balance, is a key design principle for binary APs (Fig. 6).When a polymer is highly cationic but lacks sufficient hydrophobicity, it can bind electrostatically to the negatively charged bacterial surface but fails to insert effectively into the lipid bilayer.108,118,160,162 An excessive positive charge can also cause hemagglutination of RBCs, posing significant risks for biomedical use. In contrast, APs with predominant hydrophobic character and limited cationic functionality often display strong membrane-disruptive activity but poor selectivity, damaging both bacterial and mammalian cells. Highly hydrophobic polymers also exhibit low aqueous solubility, which further limits their biological performance. | ||
| Fig. 6 Optimizing amphiphilic balance is central to the design of antibacterial methacrylate copolymers. Adjusting the ratio of cationic to hydrophobic comonomers modulates antibacterial potency and compatibility with RBCs. Adapted from ref. 118 under the terms of the CC BY-NC 3.0 license, Royal Society of Chemistry, copyright 2018. | ||
Between these extremes, specific compositions achieve a favorable balance between antibacterial potency and cytocompatibility. For example, poly(methacrylate) statistical copolymers reported by Kuroda and coworkers,35 demonstrated that an approximately 10-mer containing 40% methyl side chains and 60% aminoethyl side chains exhibited potent antibacterial activity with minimal hemolysis. In contrast, the fully cationic homopolymer (0% methyl side chains) was inactive against E. coli, whereas more hydrophobic variants (>70% methyl side chains) showed hemolytic behavior and poor solubility (Fig. 6).
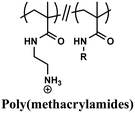
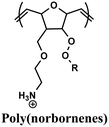
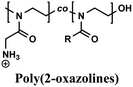
Comparable composition–activity trends have been observed for various polymer backbones (Table 2), including additional poly(methacrylates),163 poly(methacrylamides),164 nylon-3 copolymers derived from β-lactam monomers,121,165,166 poly(norbornenes),120,167–169 poly(carbonates),170 and poly(2-oxazolines).171 However, each polymer system requires independent optimization to identify its optimal balance between charge and hydrophobicity. Continued studies in physical polymer chemistry are essential to transform these case-by-case findings into general design principles for creating polymers with precisely tuned structures and functions.
| General structure | R-group | Polymerization technique | Bacteria tested | Toxicity tested (in vitro/in vivo) | Ref. |
|---|---|---|---|---|---|
| Notes: Methicillin-sensitive Staphylococcus aureus (MSSA), clinical isolates from chronic wound infections from human patients (CW2, and CW4). | |||||

|
Methyl, ethyl, butyl, benzyl | Free-radical | E. coli | RBCs (in vitro) | 35,119,172 |
|
Butyl, hexyl | Free-radical | E. coli and S. aureus | RBCs/HEp-2 cells, (in vitro) | 164 |

|
— | ROP | E. coli, B. subtilis, S. aureus, E. facium | RBCs (in vitro) | 121 |
|
Methyl, ethyl, propyl, butyl, Isopentyl, hexyl | ROMP | E. coli, S. aureus | RBCs (in vitro) | 168 |

|
— | ROP | E. coli, S. aureus, P. aeruginosa | RBCs (in vitro) | 170 |

|
— | RAFT | MRSA, MSSA, E. coli, S. epidermidis | RBCs (in vitro) | 163 |
|
Ethyl, propyl | ROP | MRSA, MSSA, CW2, CW4 | RBCs, keratinocyte (HaCaT), fibroblast (3T3), G. mellonella (in vitro/in vivo) | 171 |
Design of cationic groups
A defining feature of HDPs is the source of their positive charge: basic amino acid residues such as lysine and arginine are protonated at physiological pH, imparting a net cationic character.108 In contrast, many polymer disinfectants use permanently charged QAS units as the cationic functionality. In APs, cationic groups promote adsorption to anionic components of bacterial membranes and can induce polymers to adopt globally amphiphilic conformations that enhance bioactivity.24,165 Owing to these essential roles, cationic groups have attracted sustained attention in AP research.49,53,65,173To design ideal synthetic APs, polymer chemists have systematically explored the type of cationic centre, its spatial arrangement along the chain, and the effects of chain length and charge density on antibacterial activity and biocompatibility.53,107,163,174–176 Focusing specifically on the cationic centre, the functional moieties used in amphiphilic APs are commonly classified into two groups: nitrogen-based classes (ammonium and iminium) and non-nitrogen-based classes (sulfonium and phosphonium) (Fig. 7).24 However, inspired by the amino acid chemistry of natural HDPs, nitrogen-based cationic groups are often preferred over non-nitrogen alternatives.24
Ammonium group
In nitrogen-based cationic centres, primary, secondary, tertiary, and quaternary ammonium groups are among the most widely used cationic functionalities in antibacterial polymers (Fig. 7). Polymers bearing quaternary ammonium groups possess intrinsic, pH-independent positive charges, whereas the cationic form of primary, secondary, and tertiary ammonium groups is generated only through protonation of the corresponding amines.177 To investigate how the nature of amine moieties influences antibacterial activity and selectivity toward bacterial over mammalian cells, different research groups, including Kuroda et al.,53 Boyer et al.,49,65 Liu et al.,178 Wong et al.,23 and Hammond et al.,179 among others, have conducted systematic studies.For example, Kuroda and coworkers53 prepared a library of amphiphilic statistical copolymers containing both cationic and hydrophobic side chains. These were synthesized by copolymerizing amine-functionalized methacrylate monomers with varying proportions of alkyl methacrylates via free radical polymerization. The study demonstrated that copolymers incorporating primary and tertiary amine groups displayed strong antibacterial activity and selectivity against E. coli over RBCs. In contrast, copolymers with quaternary ammonium groups exhibited antibacterial activity against E. coli only when the hydrophobic content of the polymer was sufficiently high.53 Consistent with Kuroda's observation that primary amine groups enhance antibacterial performance, Boyer and coworkers extended this concept to ternary polymer systems.49 They prepared three distinct polymer libraries containing primary amine, tertiary amine, and quaternary ammonium moieties, comprising 120 polymers synthesized by high-throughput PET-RAFT polymerization. Their study confirmed that polymers bearing primary amines exhibited the strongest antibacterial activity (MIC90 = 32–128 µg mL−1), particularly against Gram-negative P. aeruginosa.
In a recent study, Liu et al.178 also examined the effect of amine group variation (primary, secondary, or tertiary) on the antibacterial and biocompatibility performance of HDP-mimicking β-peptide polymers synthesized via ROP (Fig. 8A; P1–P6).178 Their results demonstrated that secondary ammonium groups were superior to both primary and tertiary ammonium groups as cationic moieties in antibacterial β-peptide polymers (Fig. 8B–D). The lead polymer, a homopolymer bearing secondary amino groups (P4), exhibited potent antibacterial activity, with MIC values ranging from 6.25–12.5 µg mL−1 against E. coli, VRE, and MRSA, along with the highest selectivity (SI = 64) reflected by excellent hemocompatibility (HC10 = 400 µg mL−1) (Fig. 8B–D). Beyond in vitro assays, P4 also displayed strong activity against antibiotic-resistant pathogens and achieved high therapeutic efficacy in vivo, including treatment of MRSA-induced wound infections and keratitis, while showing low acute dermal toxicity and minimal corneal epithelial cytotoxicity (Fig. 8E–H). Consistent with these observations, a 2024 study by Wong et al.23 further reported that secondary amine macromolecules exhibited stronger antibacterial and membrane-disruptive properties than their primary amine counterparts. In addition, secondary amine polymers were less toxic and displayed higher TI values.23 Overall, these studies indicate that secondary amines confer advantages in both antimicrobial potency and selectivity. In contrast, Hammond et al.179 observed that amine type had little influence on activity in α-peptide systems, suggesting that variations in amphiphilic architecture may explain these discrepancies and emphasising the need for further evaluation of structural parameters to optimize antimicrobial performance.75
 | ||
| Fig. 8 In vitro and in vivo bioactivity of secondary amine pendant β-peptide polymers. (A) Chemical structures of synthesized β-peptide polymers with variable amine groups (primary, secondary, or tertiary). (B)–(D) Antibacterial activity, hemolytic activity, and SI of polymers. (E) Schematic of acute dermal toxicity, full-thickness wound infection, and contact lens-induced microbial keratitis models in mice. (F) Representative skin appearance in acute dermal toxicity analysis after treatment with P4. (G) Histological analysis of acute dermal toxicity after 3 days of P4 treatment (H&E staining). (H) Histological analysis of corneal toxicity following P4 treatment (H&E staining). This figure is reproduced and modified from ref. 178 with permission from American Chemical Society, copyright 2022. | ||
Apart from ammonium, other cationic groups have also gained considerable attention in determining the bioactivity of APs. To systematically evaluate the effect of different cationic functionalities, Boyer and coworkers65 synthesized a library of 31 statistical amphiphilic ternary polymers using the PET-RAFT polymerization technique. These copolymers were designed to incorporate three distinct components: (I) a cationic group, drawn either from nitrogen-based classes such as quaternary ammonium salts (N), primary amines, (K) or guanidinium groups (R), or from non-nitrogen-based classes such as sulfonium (S); (II) a hydrophilic monomer, chosen as either N-hydroxyethyl acrylamide (Hm) or oligo(ethylene glycol) methyl ether acrylamide (Pm); and (III) a hydrophobic monomer, N-isopentyl acrylamide (Im). The molar ratios of the components were carefully adjusted to probe their structure–activity relationships (SAR) (Fig. 9). Based on the identity of the cationic component, the polymer library was organized into five families: lysine-mimicking (K-family), arginine-mimicking (R-family), quaternary ammonium (N-family), sulfonium (S-family), and a hybrid K–R-family that combined lysine- and arginine-type cationic groups in varying ratios (Fig. 9).
 | ||
| Fig. 9 In vitro evaluation of ternary amphiphilic polymers. (A) Antibacterial activity, expressed as minimum inhibitory concentration (MIC), against Gram-negative strains Pseudomonas aeruginosa PAO1 (wild type) and PA37 (MDR), Escherichia coli (EC) K12, and Gram-positive Staphylococcus aureus (SA) ATCC 29213. (B) Biocompatibility profiles assessed through hemolytic activity toward sheep red blood cells (HC50) and cytotoxicity toward mouse embryonic fibroblasts (MEFs) using the alamarBlue metabolic assay (IC50). Adapted from ref. 65 under the terms of the CC BY 4.0 license, Wiley, copyright 2022. | ||
Analysis of the library showed that the balance of cationic functionality is a decisive factor governing antibacterial activity and selectivity (Fig. 9A and B). Among the cationic groups tested, the lysine-mimicking monomer (primary ammonium) paired with the hydrophilic P-monomer (mPEG-acrylamide) produced the most selective APs. In particular, K55-P1530 and K50-P2030 exhibited strong antibacterial activity (MIC = 16–32 µg mL−1 against P. aeruginosa PAO1, 32–64 µg mL−1 against multidrug-resistant PA37, and 32 µg mL−1 against E. coli K12) while maintaining excellent biocompatibility (HC50 = >2000 µg mL−1; IC50 = 500 µg mL−1), yielding SI (HC50) >83 and SI (IC50) 21 (Fig. 9A and B). Collectively, these results indicate that polymers containing 50–60% cationic groups, with a cationic-to-hydrophobic ratio of 1.4–2 and an appropriate hydrophilic-to-hydrophobic balance, deliver the most favorable selectivity toward mammalian cells while retaining strong antibacterial potency against Gram-negative bacteria (Fig. 9A and B).65
Beyond the choice of cationic groups, other molecular features also play critical roles in determining the biological activity of synthetic APs.107 For example, antibacterial activity and biocompatibility are strongly influenced by the cationic spacer arm, which is defined as the distance between the cationic centre and the polymer backbone. Kuroda and colleagues180 demonstrated this by investigating amphiphilic statistical copolymers containing primary amines with side chain spacers of 2-aminoethylene, 4-aminobutylene, and 6-aminohexylene, prepared via radical polymerization (Fig. 10A).180 Their results showed that copolymers with 4-aminobutylene side chains achieved higher antibacterial activity than those with 2-aminoethylene spacers, while avoiding undesirable toxicity (Fig. 10B–D). In contrast, extending the spacer to 6-aminohexylene yielded copolymers with strong antibacterial potency but also significant hemolytic activity (Fig. 10B–D). The 4-aminobutylene spacer was therefore identified as the optimal length, particularly when combined with hydrophobic ethyl methacrylate (EMA) in a roughly 70/30 ratio, resulting in maximal antimicrobial efficacy and minimal hemolysis, thereby achieving high selectivity (Fig. 10E). Copolymers of this composition also demonstrated rapid bactericidal kinetics and broad-spectrum activity against both Gram-positive and Gram-negative bacteria.180
 | ||
| Fig. 10 (A) Synthetic route for amphiphilic methacrylate statistical copolymers carrying cationic side chain spacer arms of varying lengths. (B) and (C) Antibacterial activity against E. coli and S. aureus, respectively. (D) Hemolytic activity (HC50). (E) Selectivity toward E. coli and S. aureus over human red blood cells. Reproduced from ref. 180 with permission from American Chemical Society, copyright 2012. | ||
Expanding on the influence of molecular architecture, Gellman and colleagues181 explored stereochemistry as another structural determinant by synthesizing novel nylon-3 polymers incorporating stereoisomeric monoethyl (ME) subunits in cis and trans configurations. Although both stereoisomers maintained an equivalent overall amphiphilic balance, their biological activity profiles differed markedly (Fig. 11A). Despite showing similar antibacterial potency against both Gram-negative and Gram-positive bacteria (3.1 to >200 µg mL−1; Fig. 11B), their abilities to disrupt eukaryotic membranes varied substantially. At the highest tested concentration (500 µg mL−1), ME-cis:TM induced nearly complete hemolysis (∼95–100%) and significant HeLa cell lysis (∼60–65%), whereas ME-trans:TM caused only moderate membrane disruption (∼20–30%; Fig. 11C and D). Complementary experiments using planar lipid bilayers and synthetic liposomes revealed that eukaryotic toxicity stemmed from polymer-mediated pore formation. These observations indicate that stereochemical variation influences polymer chain conformation and membrane interactions, even in heterochiral systems. Collectively, these findings highlight stereochemistry and spacer length as critical design parameters to consider when optimizing the selectivity of antibacterial polymers.181
 | ||
| Fig. 11 (A) Changing subunit stereochemistry alters the biological activity profile of nylon-3 polymers. (B) MIC values of ME-cis and ME-trans copolymers. (C) Hemolysis of human red blood cells after 1 and 6 h. (D) HeLa cell lysis after 1 and 6 h. Adapted from ref. 181 under the terms of the CC BY 4.0 license, American Chemical Society, copyright 2021. Notes: Tetramethyl (TM), Luria–Bertani medium (LB), EZ rich defined medium (EZRDM), and brain heart infusion medium (BHI). | ||
Staphylococcus aureus is a frequent cause of skin infections, and efforts to prevent community-associated, drug-resistant infections remain less effective than those implemented in hospital settings.182 A key factor influencing infection is the difference in skin pH: healthy human skin maintains an acidic environment due to the acid mantle, typically ranging from pH 5.4–5.9, although reported values vary depending on measurement methods.183 This acidic milieu suppresses bacterial proliferation and protease activity that can damage tissue.183 In contrast, infected sites approach neutrality because of exposure of subcutaneous tissue.184
To investigate the biological impact of this change, Hong, Kuroda, and coworkers154 studied a cationic amphiphilic statistical copolymer composed of aminobutyl methacrylate and EMA.154 The polymer displayed pH-dependent antibacterial activity, being highly effective against clinical isolates of drug-resistant S. aureus at neutral pH (MIC = 15–20 µg mL−1), but inactive under acidic conditions (MIC = >200 µg mL−1 at pH 5.5). Importantly, it retained low hemolytic activity toward human RBCs and showed no significant cytotoxicity against human dermal fibroblasts across the tested pH range (5.5–7.4). These findings demonstrate that the copolymer can selectively target bacteria over mammalian cells in a pH-responsive manner.154
In parallel with environmental sensitivity, the density of cationic charge within the polymer backbone also plays a critical role in determining biological performance. While most of the APs discussed above typically incorporate a single cation per monomer unit, the Tew group185 demonstrated that varying the number of positively charged groups per repeat unit has a pronounced effect on hemolytic activity. They developed norbornene monomers carrying one, two, or three Boc-protected amine functionalities, which were polymerized via ROMP and subsequently deprotected to yield polynorbornenes containing one, two, or three charges per monomer (polyA1–3).185 A clear trend was observed: polyA1, with a single amine, was highly hemolytic (<1 µg mL−1), reflecting its nonselective activity and increased hydrophobicity due to the isobutenyl backbone. In contrast, incorporation of additional amine groups markedly reduced hemolytic activity, with HC50 values of 700 µg mL−1 for polyA2 and 500 µg mL−1 for polyA3, while antibacterial efficacy was maintained. MIC90 values against E. coli and S. aureus showed minimal variation across the series, confirming that increased charge density decreased mammalian cell toxicity without compromising antibacterial potency.185
Iminium group
Another important nitrogen-based cationic centre is the iminium group, which includes functional classes such as guanidinium, imidazolium, pyridinium, triazolium, and thiazolium (Fig. 7). Unlike ammonium groups, where the positive charge is localized, iminium groups possess delocalized charges distributed across π bonds or aromatic conjugated systems.177 This electronic feature enhances their interaction with negatively charged bacterial membranes, thereby improving antibacterial activity while maintaining selectivity over mammalian cell membranes. Among the different functional classes of iminium groups listed here, guanidinium salts are the most widely employed as cationic sources in amphiphilic antibacterial copolymers because of their close structural resemblance to the natural amino acid arginine.24,40Arginine-rich peptides generally show superior antimicrobial activity compared to lysine-rich analogs,186 partly due to their ability to act as cell-penetrating agents that translocate across membranes and reach intracellular targets.187,188 The guanidinium group is resonance-stabilized, with delocalization of the positive charge across three nitrogen atoms.189 Because the pKa of arginine (∼12.5) is higher than that of lysine (∼10.5), polymers containing multiple guanidinium groups remain highly protonated under physiological conditions.162 In addition, the bidentate hydrogen bonding between guanidinium and membrane phosphates further reinforces these interactions, contributing to selectivity. Similar effects have been observed in synthetic guanidinylated polymers.190
Several studies have demonstrated the superior biological performance of guanidinium-functionalized polymers. Tew and coworkers191 synthesized polyguanidinium oxanorbornene (PGON) from norbornene monomers via ROMP. PGON exhibited strong antibacterial activity (MIC90 = 6–50 µg mL−1) compared with primary amine-based norbornene polymers (MIC90 = 25–400 µg mL−1) against both Gram-negative and Gram-positive bacteria, while showing minimal hemolysis (HC50 = 1500 µg mL−1) relative to the primary amine-based analogs (1–2150 µg mL−1). Interestingly, PGON did not disrupt membranes in vesicle dye-leakage or microscopy assays, suggesting a distinct mechanism of action. Its behavior, reminiscent of cell-penetrating peptides, highlights the potential of guanidinium-containing macromolecules as highly selective antibacterial agents.191
Consistent with these findings, Locock and colleagues163 converted primary amine-based polymethacrylates into guanidinium-functionalized derivatives, allowing direct comparison between the two classes.163 Polymers with pendant guanidinium groups displayed stronger antimicrobial activity against S. epidermidis, S. aureus, E. coli, and C. albicans, along with reduced hemolysis relative to their amine analogs (Fig. 12).163 Ikeda and coworkers reported acrylate homopolymers and copolymers containing biguanide side chains, which demonstrated broad-spectrum antimicrobial activity against both Gram-positive and Gram-negative bacteria.192 More broadly, guanidinium-functionalized polymers have been found to be effective against all ESKAPE pathogens as well as critical fungal species.193–195 Their strong affinity for phospholipid membranes contributes to enhanced antimicrobial activity while simultaneously reducing host toxicity.75
 | ||
| Fig. 12 Comparison of antimicrobial activity and hemolysis between guanidine-functionalized polymers (PG series) and primary amine-functionalized polymers (PA series). Reproduced from ref. 163 with permission from American Chemical Society, copyright 2013. | ||
The unique properties of guanidinium allow APs to operate through multiple mechanisms of action. Yang et al. demonstrated that guanidinium-functionalized polycarbonates exert antibacterial effects by disrupting membranes and translocating into cells, leading to the precipitation of cytosolic components (Table 3).43,193 Bai and coworkers196 developed an oligoguanidine of peptide-like size that combined membrane permeabilization with DNA binding, showing enhanced selectivity, synergy with conventional antibiotics, and a low propensity for resistance development, along with effective in vivo performance in a mouse wound infection model of MDR P. aeruginosa (Table 3). In addition, Yang and Hedrick showed that combining quaternary ammonium and guanidinium homopolymers produced additive or synergistic antibacterial activity (Table 3).197
| Guanidine-based polymer | Spacer/R-group | Test subject | MIC (µg mL−1) | HC50/IC50 (µg mL−1) | Ref. |
|---|---|---|---|---|---|
| a Multidrug-resistant strains. | |||||

|
Ethyl, propyl, butyl, pentyl, cyclohexyl, phenyl, benzyl, propyl | S. aureus, E. coli, P. aeruginosa | 3.9–62.5 (SA), 3.9–125 (EC), 15.6–125 (PA) | 62.5–>8000 (HC50) | 193 |

|
Benzyl, ethyl, butyl, isobutyl, hexyl | S. aureus, A. baumannii 1789, E. coli, K. pneumoniae, P. aeruginosa | 7.8–15.6 (SA, AB, EC), 15.6–31.3 (KP, PA) | 170–>2000 (HC50) | 43 |

|
— | S. aureus, K. pneumoniae, A. baumannii, P. aeruginosa | 2 (SA), 1 (KP), 0.5 (AB), 1 (PA)a | 638 (HC50), 14–16 (IC50) | 196 |

|
— | S. aureus, A. baumannii 1789, K. pneumoniae, P. aeruginosa, E. coli | 3.9 (SA), 7.8 (AB, EC, KP), 15.6 (PA) | — | 197 |
Imidazole side chains, inspired by histidine-rich HDPs, represent another important source of cationic charge in APs. Recently, imidazole groups have been explored as cationic sources for antibacterial polymers. Using an ultrafast ROP combined with click chemistry, Tang, and coworkers198 synthesized imidazolium-based block copolypeptides (PPGn-PILm). These α-helical copolypeptides readily self-assembled into positively charged nanoscale micelles, which displayed strong antimicrobial activity and excellent hemocompatibility.198 The lead polymer, PPG34-PIL70, exhibited low MIC (25 µg mL−1) against both Gram-positive S. aureus and Gram-negative E. coli. Importantly, it showed negligible hemolysis and low cytotoxicity toward NIH 3T3 fibroblasts and 293T kidney cells, even at concentrations twice the MIC. Live/dead fluorescence assays further confirmed high cell viability, highlighting its favorable safety profile. Overall, they claimed PPG34-PIL70 represents a promising candidate for anti-infective applications combining broad-spectrum potency with excellent mammalian cell compatibility (Fig. 13).198
 | ||
| Fig. 13 Antibacterial activity and biocompatibility of imidazolium-based block copolymers. (A) MICs against S. aureus and E. coli. (B) Hemolysis profile at different polymer concentrations. (C) Viability of NIH 3T3 fibroblasts and (D) 293T kidney cells exposed to polymers at 50–400 µg mL−1. (E) Live/dead fluorescence images of untreated NIH 3T3 cells and (F) cells treated with PPG34-PIL70 (50 µg mL−1), showing predominantly viable cells (green). Reproduced from ref. 198 with permission from American Chemical Society, copyright 2021. | ||
Yan and colleagues199 synthesized imidazolium-type ionic liquid monomers and their corresponding poly(ionic liquid) (PIL) membranes. By studying the impact of chemical structure on antibacterial activity, they found that activity increased with higher charge density (double cation) but decreased with longer carbon chain length.199 In addition, Yang and coworkers200 prepared a series of polycarbonates bearing propyl or hexyl side chains quaternized with different nitrogen-containing heterocycles (pyridinium and imidazolium; Fig. 14). These polymers exhibited substantially enhanced anti-infective properties against bacterial and fungal strains compared to their analogs quaternized with trimethylamine (TMA). The improvement in efficacy, as reflected in their MIC values (Table 4), was attributed to a more favorable balance between cationic charge and hydrophobicity. Notably, hemolysis assays further confirmed their excellent hemocompatibility, with many polymers displaying high HC50 values (>1000 µg mL−1) and pronounced selectivity, with some achieving >250-fold preference for S. aureus over mammalian RBCs (Table 4). Acting predominantly through membrane-lytic mechanisms, these polymers present a reduced risk of resistance development and hold strong promise as potent antimicrobial agents.200
 | ||
| Fig. 14 Chemical structures of polycarbonates prepared by organocatalyzed ROP, functionalized with different nitrogen-containing heterocycles (Q = MeImdz, EtImdz, BuImdz, Pyr, or DMAP). Adapted and redrawn from ref. 200 with permission from American Chemical Society, copyright 2014. | ||
| Polymer | MIC [(mg L−1), (Selectivity)] | ||||
|---|---|---|---|---|---|
| S. aureus (Gram +) | E. coli (Gram −) | P. aeruginosa (Gram −) | C. albicans (fungus) | HC50 | |
| a Trimethylamine (TMA) was used for quaternization to prepare cationic polymer analogs serving as reference compounds for comparison. | |||||
| PrBr(MeImdz) | 500 (>2) | 1000 (>1) | >1000 | 250 (>4) | >1000 |
| PrBr(EtImdz) | 31 (>32) | 250 (>4) | >1000 | 250 (>4) | >1000 |
| PrBr(BuImdz) | 8 (∼125) | 31 (∼32) | 250 (∼4) | 250 (∼4) | ∼1000 |
| PrBr(Pyr) | 63 (>16) | 125 (>8) | 1000 (>1) | 125 (>8) | >1000 |
| PrBr(DMAP) | 8 (>125) | 63 (>16) | 125 (>8) | 250 (>4) | >1000 |
| PrBr(TMA)a | 500 (>2) | 1000 (>1) | >1000 | 500 (>2) | >1000 |
| HexBr(MeImdz) | <4 (>250) | 31 (>32) | 250 (>4) | 125 (>8) | >1000 |
| HexBr(EtImdz) | <4 (>250) | 16 (>64) | 250 (>4) | 125 (>8) | >1000 |
| HexBr(BuImdz) | <4 (>250) | 8 (∼62) | 31 (∼16) | 125 (∼8) | ∼1000 |
| HexBr(Pyr) | <4 (>250) | 31 (>32) | 125 (>8) | 125 (>8) | >1000 |
| HexBr(DMAP) | <4 (>250) | 31 (>16) | 31 (>16) | 250 (>2) | >500 |
| HexCl(TMA)a | 125 (>8) | 125 (>8) | 1000 (>1) | 250 (>4) | >1000 |
In a separate study, the Sen group201 was the first to report the pronounced influence of spatial positioning of the cationic charge and hydrophobic tail on the bioactivity of APs. They designed a series of amphiphilic pyridinium–methacrylate copolymers with variations in the relative placement of the positive charge and the alkyl tail (Fig. 15).201 Evaluation of their antibacterial and hemolytic properties showed that the separate-centered configuration enhanced membrane disruption, thereby increasing both antimicrobial and hemolytic activity, while the same-centered configuration reduced hemotoxicity and resulted in improved selectivity (Fig. 15).201
 | ||
| Fig. 15 Effect of “same-center” and “separate-center” positioning on the antibacterial and hemolytic activity of homologous polymers. Reproduced from ref. 201 with permission from Wiley-VCH GmbH, copyright 2008. | ||
The Fernández-García, López and coworkers202 synthesized two series of antimicrobial polymethacrylates incorporating mono- and bis-cationic quaternary ammonium groups via controlled N-alkylation of pendent thiazole- and triazole moieties. In this design, the azole side chains were alkylated with butyl iodide, allowing precise regulation of the degree of quaternization (DQ) from 10% to 100%. When the DQ exceeded 50%, the polymers displayed striking antibacterial potency against both P. aeruginosa and S. aureus (MIC <10 µg mL−1), while simultaneously maintaining exceptional hemocompatibility (HC50 >5000 µg mL−1), resulting in an outstanding SI. Time–kill studies further revealed rapid bactericidal action, achieving a 3-log reduction in CFU mL−1 within just 15 minutes. Scanning electron microscopy confirmed the membrane-disruptive nature of these polymers, showing surface roughening and pronounced bacterial aggregation after treatment.202
Sulfonium group
Sulfur compounds play essential roles in cellular signalling and defense, motivating interest in sulfur-based functionalities for antimicrobial design. Drawing on this inspiration, synthetic chemists have investigated sulfonium groups as cationic centers in APs. Kanazawa et al.203 pioneered this approach by synthesizing polymers in which sulfur atoms acted as sources of positive charge. Since then, sulfonium-containing compounds have gained significant attention in the development of amphiphilic APs and other therapeutic systems.204–206 Their structural and electronic resemblance to quaternary ammonium groups provides a useful framework for charge incorporation, while offering unique opportunities for chemical diversification. Early studies reported that sulfonium-based polymers exhibited stronger activity against Gram-positive than Gram-negative bacteria.204–206 More recently, advances in sequence-defined polysulfonium architectures have demonstrated broad-spectrum activity,207 including potent efficacy against antibiotic-resistant pathogens.206,208Expanding on these findings, the Rao group209 recently reported main-chain sulfonium-containing polymers with AB-alternating sequences, introducing a new class of sulfonium-based macromolecules. These polymers exhibited broad-spectrum bactericidal activity against clinically relevant pathogens while demonstrating excellent biocompatibility. Notably, they were highly effective against MRSA, with MBCs ranging from 1.25 to 10 µg mL−1 and caused no detectable hemolysis even at concentrations up to 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 µg mL−1. This remarkable profile corresponded to a SI exceeding 2000, highlighting their strong therapeutic potential.209 Building on this success, the same group210 later developed sulfonium-based homopolymers incorporating cationic sulfonium units separated by alkane spacers in the main-chain.210 These polysulfoniums displayed potent antimicrobial activity against bacteria (E. coli, S. aureus) and fungi (C. albicans), with MICs between 0.5 and 32 µg mL−1. Furthermore, optimized compositions effectively disrupted biofilms, achieving 80–90% biomass reduction and >99% eradication of C. albicans and E. coli in 3-day mature biofilms at 2 × MIC, while maintaining excellent hemocompatibility (Fig. 16).210
000 µg mL−1. This remarkable profile corresponded to a SI exceeding 2000, highlighting their strong therapeutic potential.209 Building on this success, the same group210 later developed sulfonium-based homopolymers incorporating cationic sulfonium units separated by alkane spacers in the main-chain.210 These polysulfoniums displayed potent antimicrobial activity against bacteria (E. coli, S. aureus) and fungi (C. albicans), with MICs between 0.5 and 32 µg mL−1. Furthermore, optimized compositions effectively disrupted biofilms, achieving 80–90% biomass reduction and >99% eradication of C. albicans and E. coli in 3-day mature biofilms at 2 × MIC, while maintaining excellent hemocompatibility (Fig. 16).210
 | ||
| Fig. 16 Schematic illustration of main-chain sulfonium-containing homopolymers (PS+) showing broad-spectrum antimicrobial activity, excellent hemocompatibility, and effective biofilm eradication at low MIC values. Reproduced from ref. 210 with permission from American Chemical Society, copyright 2021. | ||
Encouraged by the strong bioactivity of main-chain sulfonium polymers, the Klinger group recently extended this concept by shifting focus toward side-chain functionalization.41 While main-chain systems had already shown excellent antibacterial activity and biocompatibilities, the SAR of side-chain sulfonium polymers remained largely unexplored. To address this gap, they designed a library of sulfonium-based terpolymers that incorporated different hydrophobic (aliphatic or aromatic) groups alongside hydrophilic polyethylene glycol (PEG) segments, while keeping overall chain lengths comparable (Fig. 17). Crucially, the study examined how the spatial arrangement of cationic and hydrophobic groups on the backbone, specifically same-center versus different-center configurations, influenced biological activity (Fig. 17A and B).41 Bactericidal assays against both Gram-positive and Gram-negative bacteria revealed that same-center polymers displayed superior activity compared to different-center analogs with similar cLogP values. However, higher cLogP values in both polymer types were associated with increased hemolysis (Fig. 17A and B), thereby compromising overall polymer selectivity. Notably, sulfonium-based APs demonstrated enhanced bactericidal activity and selectivity relative to their quaternary ammonium counterparts.41 Complementary results have also been reported for other recently developed sulfonium-derived APs.175,211
 | ||
| Fig. 17 Chemical structures and bioactivity of sulfonium-based terpolymers prepared with two architectures: (A) same-center and (B) different-center designs showing antibacterial and hemolytic activities. Adapted from ref. 41 under the terms of the CC BY 3.0 license, Royal Society of Chemistry, copyright 2025. | ||
Fu, Whittaker, and coworkers212 found that sulfonium polymers displayed greater antibacterial potency against both E. coli (MIC90 = 125–250 µg mL−1) and S. aureus (MIC90 = 2000 µg mL−1), outperforming their ammonium-based analogs (MIC90 = 2000–>8000 µg mL−1 against E. coli and >8000 µg mL−1 against S. aureus) and phosphonium-based analogs (MIC90 = 250–>8000 µg mL−1 against E. coli and >8000 µg mL−1 against S. aureus). However, all classes of cationic polymers exhibited some degree of hemolytic activity, with sulfonium polymers inducing the highest levels, followed by phosphonium and then ammonium systems. A similar trend was observed for cytotoxicity toward mammalian cells. Despite this drawback, sulfonium polymers maintained a comparatively higher therapeutic index (TI = 72), reflecting a balance between their potent antibacterial efficacy and associated toxicity.212
These findings highlight both the promise and the challenges of sulfonium chemistry for antibacterial applications, emphasizing the need for further molecular design strategies to enhance antibacterial performance while reducing hemolysis and mammalian cytotoxicity.
Phosphonium group
Phosphonium groups, positioned below nitrogen in the periodic table, carry a stronger cationic character than ammonium, which enhances their electrostatic interaction with negatively charged bacterial membranes and thereby improves antibacterial potency.79,213,214 This principle was first demonstrated by the Kanazawa group,215 who reported the use of phosphonium moieties as cationic charge centers in amphiphilic antibacterial polymers. Comparative studies revealed that polymers with identical backbones, but different cationic substituents displayed markedly different activity, with phosphonium-based salts exhibiting antibacterial activity up to two orders of magnitude greater than their quaternary ammonium analog.215Further advances were made by Gillies et al.,173 who investigated phosphonium-functionalized block copolymer micelles as intrinsically antibacterial polymer assemblies. In their design, phosphonium cations of varying alkyl chain lengths were conjugated to the termini of poly(ethylene oxide)–polycaprolactone block copolymers, which subsequently self-assembled into micelles in aqueous solution. The antibacterial performance, measured as MBC, was strongly influenced by the length of the phosphonium alkyl chain, with distinct trends observed between S. aureus and E. coli. Notably, the most active micellar assemblies achieved bactericidal activity without causing hemolysis at concentrations above their MBC values, demonstrating selective disruption of bacterial membranes while sparing RBCs.173
Design of hydrophobic group
It is commonly recognized that the hydrophobic domains of cationic APs play a decisive role in disrupting microbial membranes. Once adsorbed to the cell surface, these groups penetrate the lipid bilayer, destabilize membrane integrity, and cause cytoplasmic leakage, ultimately leading to microbial death. As a result, the length, branching, and chemical nature of hydrophobic substituents are critical factors in monomer design. Both the local hydrophobicity of individual monomers and the overall hydrophobicity of the polymer are key determinants of antibacterial potency and cytotoxicity.35,109,174,216 Achieving the right hydrophilic to hydrophobic balance is therefore essential for the design of amphiphilic antimicrobial copolymers, as highlighted in the preceding section (Fig. 6). This balance can be tuned at either the monomer or polymer level through three main approaches: (I) tailoring monomer structures, such as spacer length and hydrophobic group type; (II) adjusting the ratio of hydrophobic to other functional groups in copolymers; and (III) incorporating neutral hydrophilic units, such as PEG.31,108,118The first strategy focuses on modulating hydrophobicity at the local level by careful selection of hydrophobic monomer structures, thereby fine-tuning the overall hydrophilic–hydrophobic balance. Most APs employ alkyl chains as hydrophobic substituents, and the length and type of these pendant groups play a crucial role in determining bioactivity.24,40 For example, Yang and Hedrick's group217 developed a library of polycarbonates in which the length of the alkyl spacer between the quaternary ammonium group and the polymer backbone was systematically varied.217 Their findings (Fig. 18) showed that the MICs against multiple pathogens decreased as the spacer length increased from three to eight carbons. Within this range, elongation of the alkyl chain enhanced the hydrophobic character of the polymers, thereby promoting stronger interactions with bacterial membranes and improving antimicrobial performance.217 Similar observations have been reported by other groups across a variety of polymer backbones,218–220 reinforcing the critical role of hydrophobic monomer design in antimicrobial efficacy.
 | ||
| Fig. 18 MIC values of antimicrobial cationic polycarbonates bearing different chain lengths of alkyl groups against various pathogens. Adapted and redrawn from ref. 217 with permission from American Chemical Society, copyright 2013. | ||
In a related study, the Halder group demonstrated that amide- and ester-containing cationic amphiphilic polymers with adjustable side-chain hydrophobicity can regulate both antibacterial efficacy and cytotoxicity (Fig. 19A).221 Their findings indicated that amide polymers could act as potent antibacterial agents even at lower levels of hydrophobicity, whereas ester-based polymers required relatively higher hydrophobicity to achieve comparable effectiveness (Fig. 19B). At elevated hydrophobic content, both amide and ester polymer showed similar biological profiles in terms of membrane-disruptive antibacterial activity and host cell toxicity. In contrast, at reduced hydrophobicity, both types were less cytotoxic, but amide polymers retained significant antibacterial and membrane activity compared to ester polymers (Fig. 19B).221
 | ||
| Fig. 19 Chemical structure and bioactivity of maleimide-based polymer. (A) Amide- and ester-containing amphiphilic polymers with tunable hydrophobicity. (B) Antibacterial activity (MIC), hemolytic activity (HC50), and selectivity index of the corresponding cationic polymers. Reproduced and redrawn from ref. 221 with permission from American Chemical Society, copyright 2016. | ||
In addition to the effect of chain length, the structural nature of the hydrophobic group also has a significant impact on bioactivity. Although linear alkyl chains are the most common hydrophobic units in cationic APs, cyclic and fused-ring structures can also be incorporated. To explore the role of cyclic groups, Gellman group synthesized nylon-3 statistical copolymers containing either cyclohexane substituents or their acyclic analogs.222 Polymers bearing cyclohexane rings displayed enhanced antibacterial activity (MIC = 1.6–50 µg mL−1) along with low hemolytic activity compared to those with structurally similar acyclic groups (MIC = 3.13–>200 µg mL−1).222 Later, Yang and coworkers developed a broader library of polymers featuring diverse spacer groups, including both aliphatic and aromatic structures.193 Their findings demonstrated that polymers with aliphatic spacers generally exhibited superior antibacterial activity and hemocompatibility relative to their aromatic counterparts. However, increasing the alkyl chain length from ethyl to pentyl or cyclohexyl was associated with greater hemolysis, reflecting the increased hydrophobicity of these polymers.193
Following this approach, the same group recently reported guanidinium-functionalized polycarbonate statistical copolymers prepared by organocatalytic ROP to investigate the influence of hydrophobic side chains (ethyl, propyl, isopropyl, benzyl, and hexyl) on antimicrobial performance and selectivity.43 Across this series, the polymers showed broadly comparable MICs and MBCs against a wide range of microbial strains, although polymers bearing more hydrophobic substituents demonstrated faster bactericidal action. At a hydrophobic comonomer content of 20 mol%, the polymers exhibited diminished selectivity, which was attributed to an increase in hemolytic activity.43
The second and third design approaches emphasize control at the global polymer level, where the hydrophobic component modulates bioactivity through its contribution to the overall amphiphilic balance in antimicrobial copolymers. In binary APs, an excess of cationic content reduces the proportion of hydrophobic groups, which can compromise antimicrobial potency. Overly cationic polymers may also induce RBC agglutination223 and disrupt host cellular processes, leading to toxicity.224–227 Conversely, polymers with excessive hydrophobicity can damage both bacterial and mammalian membranes indiscriminately, while also suffering from poor aqueous solubility, ultimately reducing their bioavailability and antimicrobial effectiveness.
To systematically examine the impact of hydrophobicity, the Boyer group constructed a library of ternary antibacterial polymers via PET-RAFT polymerization, varying both the hydrophobic monomer type (linear, branched, cyclic, or aromatic) and the chain length (C5, C7, or C9) (Fig. 20A and B).109 The polymers were tested against four bacterial strains and RBCs to evaluate their antibacterial potency and hemocompatibility (Table 5). Clear structure–bioactivity trends were identified. For polymers with comparable degree of polymerization (DP = 40) and monomer ratios, increasing side-chain length reduced antibacterial activity, with C9 derivatives being less effective than C5 and C7 analogs.109
 | ||
| Fig. 20 (A) Chemical structures of the monomers used: eight hydrophobic monomers [N-isopentylacrylamide (Im), N-pentylacrylamide (Pm), N-heptylacrylamide (Hm), N-butyl-N-propylacrylamide (Pbm), N-cycloheptylacrylamide (Cpm), N-(cyclohexylmethyl)acrylamide (Cxm), N-benzylacrylamide (Bm), and N-nonylacrylamide (Nm)], the cationic monomer tert-butyl (2-acrylamidoethyl)carbamate, the hydrophilic monomer HEAm, and the RAFT agent BTPA. (B) Reaction scheme for the synthesis of antibacterial polymers by PET-RAFT. (C) Correlation between cLogP of polymers and their MIC values against PAO1. MIC values >128 µg mL−1 were plotted as 256 µg mL−1 for ease of viewing. Brown ●, polymers from reference;49 green ▼, polymers from references;64,228 blue ⧫, polymers from the reported study.109 cLogP values were calculated using a theoretical DP of 10. Reproduced from ref. 109 with permission from American Chemical Society, copyright 2020. | ||
| Family of polymers | Hydrophobic monomer (cLogP) | Polymer | Monomer ratio (x![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) y y![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) z) z) |
cLogP of representative oligomer | MIC of polymer (µg mL−1) | HC50 (µg mL−1) | |||
|---|---|---|---|---|---|---|---|---|---|
| PAO1 | PA 27853 | EC K12 | SA 29213 | ||||||
| I-family | Im (1.40) | I40-1040 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40 40 |
2.7 | 16 | 16–32 | 16 | >256 | 141.5 ± 6 |
| I40-1535 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 35 35 |
1.55 | 16 | 16–32 | 16 | >256 | >2000 | ||
| I40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
0.39 | 16–32 | 32–64 | 32 | >256 | >2000 | ||
| I40-2525 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25 |
−0.77 | 256 | >256 | 128–256 | >256 | No hemolysis | ||
| I40-3020 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
−1.93 | >256 | >256 | >256 | >256 | No hemolysis | ||
| P-family | Pm (1.53) | P40-1040 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40 40 |
3.22 | 32 | 32 | 16 | >256 | 87 ± 9 |
| P40-1535 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 35 35 |
2 | 32 | 32 | 16 | >256 | ∼2000 | ||
| P40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
0.78 | 32 | 32–64 | 32–64 | >256 | >2000 | ||
| H-family | Hm (2.59) | H40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
3.95 | 64 | 128 | 64 | >256 | 1150 ± 392 |
| H40-2525 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25 |
2.2 | 64 | 64 | 64 | >256 | >2000 | ||
| H40-3020 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
0.45 | 64–128 | 128–256 | 128 | >256 | >2000 | ||
| Pb-family | Pbm (2.37) | Pb40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
3.7 | 32 | 32–64 | 16–32 | >256 | 523 ± 209 |
| Pb40-2525 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25 |
2 | 32–64 | 32–64 | 32–64 | >256 | >2000 | ||
| Pb40-3020 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
0.29 | 64 | 128 | 128–256 | >256 | >2000 | ||
| Cp-family | Cpm(2.01) | Cp40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
2.2 | 32–64 | 64 | 32 | >256 | 123 ± 13 |
| Cp40-2525 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25 |
0.74 | 32-64 | 32–64 | 16-32 | >256 | ∼2000 | ||
| Cx-family | Cxm (2.07) | Cx40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
2.38 | 32 | 32–64 | 32 | >256 | 131 ± 20 |
| Cx40-2525 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 25 |
0.89 | 32–64 | 64 | 16–32 | >256 | ∼2000 | ||
| B-family | Bm (1.19) | B40-1535 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 35 35 |
1.11 | 32–64 | 64 | 64–128 | >256 | >2000 |
| B40-1040 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40 40 |
2.23 | 16–32 | 32 | 32 | >256 | 961 ± 425 | ||
| B40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
0.05 | 64 | 64–128 | 256 | 256 | >2000 | ||
| N-family | Nm (3.65) | N40-2030 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
7.13 | >256 | >256 | >256 | >256 | 257 ± 37 |
| N40-3020 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
2.57 | 128–256 | 256 | 128–256 | 256 | >2000 | ||
| N40-3515 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 35 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 15 |
0.29 | 128–256 | 256 | 128–256 | >256 | No hemolysis | ||
Hydrophobic content also had a strong influence on toxicity. Increasing hydrophobic fraction from 10 mol% to 40 mol% led to greater hemolysis, particularly as chain length increased from C5 to C9 (Table 5). For instance, Pm-based polymers (P40-2030) showed HC50 >2000 µg mL−1, compared with Hm-based polymers (H40-2030, HC50 = 1150 µg mL−1) and Nm-based polymers (N40-2030, HC50 = 257 µg mL−1) (Table 5). These results demonstrate that both hydrophobicity and hydrophobic content are key parameters of blood compatibility. Within same chain length series, branched hydrophobic polymer group showed enhanced antibacterial efficacy relative to their linear counterparts. Among all the variants, isopentyl-based monomers (Im) offered the most favorable balance between antibacterial performance and hemocompatibility. Overall, linear and branched alkyl side chains conferred greater antibacterial activity than cyclic or aromatic groups. Importantly, the group analyzed the cLogP values of statistical APs from their earlier studies49,64,228 and compared them with data from the reported work by plotting them against the MIC values for P. aeruginosa (PAO1) (Fig. 20C).109 Consistent with their previous observations, the antibacterial activity of these polymers showed a strong correlation with the cLogP values. Although some variation in activity was observed, polymers with poor antibacterial performance (MIC >128 µg mL−1) consistently exhibited cLogP values outside the range of 0–6 (Fig. 20C). Notably, polymers with calculated cLogP values between 0 and 2 achieved the best compromise, combining strong anti-infective properties with minimal hemotoxicity.109
Design of neutral hydrophilic group
A complementary strategy to mitigate the toxicity of antibacterial polymers is the incorporation of electrically neutral, hydrophilic moieties into the polymer backbone or side chains, thereby refining the amphiphilic balance. HDPs achieve selective antimicrobial activity through a combination of cationic, hydrophobic, and neutral hydrophilic amino acids such as serine. While cationic residues enable electrostatic binding and hydrophobic residues promote membrane insertion, neutral hydrophilic residues help temper activity and enhance overall biocompatibility.Drawing inspiration from this design, synthetic polymers have increasingly adopted the third neutral hydrophilic groups—such as PEG or sugar units, to reduce nonspecific interactions and hemolysis while maintaining antimicrobial potency. A pioneering example was reported in 2007 by Youngblood and colleagues,229 who copolymerized quaternized poly(vinylpyridine) with hydroxyethyl methacrylate and PEG methyl ether methacrylate using free radical polymerization technique. Polymers bearing hydrophilic units showed diminished hemolytic activity with maintained or even enhanced antibacterial performance.229 Later, the Yang group converted hydrophobic groups of primary amine-functionalized polymethacrylates into hydrophilic moieties to improve biocompatibility.174 Encouraged by these findings, the Boyer group developed a series of statistical amphiphilic ternary copolymers to systematically probe the role of neutral hydrophilic groups in antibacterial activity and toxicity.27 Unlike hydrophobic groups, which directly disrupt bacterial membranes, hydrophilic substituents exert an indirect but crucial influence by modulating the overall hydrophobic/hydrophilic balance and global amphiphilicity.27 This effect is consistent with atomistic simulations by Vemparala230 and Rani,231 which highlighted the importance of water–polymer interactions in tuning activity profiles.230,231 This indicated that the structural characteristics of hydrophilic groups—including chain length, flexibility, and intrinsic hydrophilicity, are key parameters that determine selectivity in APs. In particular, the Boyer group found that polymers incorporating PEG as the neutral hydrophilic component exhibited markedly higher biocompatibility and selectivity than those containing shorter, less flexible groups such as 4-acryloylmorpholine or N-hydroxyethyl acrylamide.27
To further leverage the advantages of PEG in antibacterial polymer design, Chattopadhyay and coworkers232 synthesized a series of graft copolymers by chemoselectively attaching acrylate-terminated hydrophobic PEG chains to a chitosan backbone at the C2–NH2 position (Fig. 21).232 These PEG–chitosan graft copolymers exhibited potent and selective antibacterial potency against both Gram-negative and Gram-positive bacteria, while maintaining excellent hemocompatibility (Fig. 21). Notably, bacterial selectivity was strongly influenced by grafting density. For instance, low-density grafts (CHT-g-OPEG10) preferentially targeted E. coli over S. aureus, as indicated by a MIC (S. aureus)/MIC (E. coli) ratio of 1.68. Conversely, higher-density grafts (CHT-g-OPEG30) shifted activity toward S. aureus, with a MIC (E. coli)/MIC (S. aureus) ratio of 2.14. Together, these results demonstrate that modulation of PEG grafting density offers a safe and tunable approach for controlling antibacterial specificity in chitosan-based copolymers.232
 | ||
| Fig. 21 Structure and biological performance of PEG–chitosan graft copolymers. Reproduced from ref. 232 with permission from American Chemical Society, copyright 2023. | ||
In addition to graft copolymer strategies, PEG has also been incorporated directly as a side chain to further optimize the amphiphilic balance of APs. In a recent 2024 study, Jańczewski et al. introduced PEG side chains into the repeating units of APs to simultaneously enhance activity and minimize toxicity. These polymers exhibited strong and broad-spectrum efficacy against E. coli, S. aureus, and C. albicans, with the PEG substitution not only improving biocompatibility but also significantly enhancing antimicrobial performance.38
An alternative to PEG is the use of sugar-based macromolecules as neutral hydrophilic components in APs. Their natural abundance, hydroxyl-rich architecture, and excellent biocompatibility make them attractive candidates for enhancing selectivity. Notably, E. coli express mannose-binding proteins on their pili, which facilitate adhesion to glycoconjugates on mammalian cell surfaces.233 Leveraging this feature, the Ragogna group developed amphiphilic polyphosphonium polymers functionalized with pendant mannose moieties, hypothesizing that this modification would enhance bacterial binding and killing specificity toward E. coli.214 In their study, they reported the first synthesis of carbohydrate-containing phosphonium monomers and polymers, featuring either mannoside or glucoside substituents combined with hydrophobic hexyl chains, alongside a tris(hydroxypropyl)phosphonium-based control polymer. Unexpectedly, the control polymer, despite lacking hydrophobic alkyl chains, displayed potent antibacterial activity against both Gram-negative and Gram-positive bacteria, while also showing minimal hemolysis, thereby increasing overall selectivity and safety.214
Aligned with this sugar-based strategy, the Li group developed a different class of cationic peptidopolysaccharides by grafting antimicrobial polypeptides onto a polysaccharide backbone.234 Using thiol–ene click chemistry, they coupled methacrylate-terminated poly(lysine-random-phenylalanine) (Me-KnFm) chains to thiolated dextran, generating Dex-g-KnFm copolymers.234 These polymers exhibited potent broad-spectrum antimicrobial activity against Gram-negative, Gram-positive, and fungal strains, with MIC values ranging from 31 to 500 µg mL−1, and did not induce resistance in MRSA. In addition, Dex-g-KnFm copolymers showed negligible hemolysis and good in vitro biocompatibility with murine myoblast (C2C12) cells. Among the series, one polymer emerged as the most effective candidate, displaying over 200-fold greater selectivity than the corresponding polypeptide molecules. Importantly, this optimized copolymer also achieved strong in vivo antibacterial performance, with more than a 3-log reduction in bacterial load in a mouse sepsis model.234
Extending this work, the same group in 2025 addressed bacterial liver abscesses,235 an increasingly critical clinical challenge exacerbated by poor antibiotic penetration and multidrug resistance, by developing a hepatotropic pullulan-based peptidopolysaccharide designed for targeted liver delivery and broad-spectrum antibacterial action (Fig. 22).235 Through precise molecular engineering to balance antibacterial potency with hemocompatibility, the lead copolymer PP11 demonstrated outstanding efficacy against all ESKAPE pathogens, including MDR strains, while maintaining exceptionally high HC50 values and a SI exceeding 647. PP11's membrane-disruptive mechanism minimized the risk of resistance development. Importantly, ex vivo and in vivo studies confirmed pronounced hepatotropic targeting and potent antibacterial performance in a murine liver abscess model induced by ESBL-producing E. coli, achieving >99.9% bacterial reduction and significant suppression of pro-inflammatory cytokines IL-6 and IL-1β. A four-week biosafety assessment further verified the absence of toxicity across major organs and preservation of normal hepatic and renal function.235
 | ||
| Fig. 22 (A) Chemical structure and therapeutic concept of the hepatotropic pullulan-based copolymer (PP11), exhibiting broad-spectrum antibacterial activity against ESKAPE pathogens with minimal risk of drug resistance. (B) Targeted treatment of ESBL-producing E. coli-induced bacterial liver abscess using PP11, demonstrating efficient hepatotropic targeting, potent antibacterial efficacy, and low systemic toxicity. Reproduced from ref. 235 with permission from Elsevier, copyright 2025. | ||
Parallel to these developments, insights from HDPs have highlighted the role of neutral polar residues in governing selectivity. Serine (polar but uncharged) is one of the most frequently occurring amino acids in HDPs, while glycine is more abundant in HDPs compared with typical proteins. Inspired by these observations, Gellman and coworkers designed ternary nylon-3 copolymers incorporating serine-like or glycine-like neutral subunits together with hydrophobic and cationic components (Fig. 23).166 Compared with earlier binary cationic–hydrophobic nylon-3 copolymers, these ternary systems exhibited enhanced anti-infective effects with reduced hemolysis, underscoring the importance of neutral polar moieties in achieving a favorable balance between efficacy and biocompatibility.166
 | ||
| Fig. 23 Chemical structures of ternary nylon-3 copolymers incorporating neutral polar subunits. Adapted and redrawn from ref. 166 under the terms of the CC-BY-NC-ND license, American Chemical Society, copyright 2014. | ||
Overall, the strategic incorporation of neutral hydrophilic groups has proven to be an effective route for enhancing the safety profile of antimicrobial copolymers while preserving or even strengthening their antibacterial potency. Such design principles hold considerable promise for advancing next-generation APs that combine high efficacy with excellent biocompatibility.
Design of end group
For polymers of high molecular weight, the contribution of chain-end groups to the overall amphiphilic balance is generally considered negligible. However, in the case of HDP-mimetic antibacterial polymers, which are often designed as relatively short oligomers, the identity of the terminal groups can have a pronounced influence on biological properties. To address this, controlled polymerization techniques such as ROP, RAFT, and ATRP have been widely applied, providing precise control over end-group fidelity and enabling systematic evaluation of their role in antimicrobial potency and biocompatibility.118One of the earliest examples was reported by Maynard group,236 who examined the toxicity of PEG–acrylate-based polymers synthesized via RAFT polymerization using various chain-transfer agents. Their results demonstrated that polymer toxicity was strongly linked to the presence of thiol chain ends, and that this toxicity could be alleviated by masking the free thiol functionality.236 Complementing this work, Gellman and colleagues investigated nylon-3 copolymers and revealed that the nature of end groups substantially influenced hemolytic behavior, with more hydrophobic termini leading to increased RBC lysis.165
Expanding on these findings, Michl and coworkers conducted a broader study by synthesizing a series of eight RAFT-derived cationic methacrylate polymers containing either amine (PA1–4) or guanidine (PG1–4) pendant groups (Fig. 24).237 In their work, the chemical nature of both the R- and Z-end groups of the RAFT agent was systematically varied to investigate their influence on polymer bioactivity (Table 6).237 The resulting polymers were evaluated for hemocompatibility and antimicrobial activity against several clinically significant microorganisms, including a strongly biofilm-forming strain of Staphylococcus epidermidis, a vancomycin- and methicillin-resistant Staphylococcus aureus strain (VISA), and the opportunistic fungal pathogen Candida albicans (Table 6).237 The findings revealed that the R-end group played the predominant role in dictating polymer-associated hemotoxicity in polymers containing either amine or guanidine pendant groups. Notably, replacing the anionic cyanovaleric acid R-group in PA1 with a neutral isobutyronitrile R-group (PA3) resulted in more than a twentyfold increase in RBC lysis (Table 6). In contrast, the Z-end group exerted a stronger influence on antimicrobial potency. Polymers incorporating a long, hydrophobic dodecylsulfanyl Z-group demonstrated markedly enhanced activity against VISA, S. epidermidis, and C. albicans (PA1 and PG2) compared to analogs with shorter ethylsulfanyl substituents or those lacking a ZCS2-group (Fig. 24 and Table 6).237
 | ||
| Fig. 24 Synthesis process of statistical APs containing amine (PA1–PA4) or guanidine (PG1–PG4) pendant groups that vary in both RAFT Z- and R-groups. Adapted from ref. 237 with permission from Royal Society of Chemistry, copyright 2014. | ||
| Polymer | MIC (µg mL−1) | Hemolysis (%) | ||
|---|---|---|---|---|
| VISA | S. epidermidis | C. albicans | ||
| Notes: PA1–4 and PG1–4 correspond to amine and guanidine pendent groups, respectively. Vancomycin and methicillin-resistant strain of Staphylococcus aureus (VISA). Hemolysis was determined as the percentage of lysed cells at the MIC concentration of S. epidermidis. | ||||
| PA1 | 32 | 32 | 32 | 1.2 |
| PA2 | 64 | 32 | 256 | 1.2 |
| PA3 | 32 | 32 | 32 | 26.2 |
| PA4 | 128 | 32 | 128 | 3.3 |
| PG1 | 16 | 16 | 32 | 13.4 |
| PG2 | 32 | 16 | 64 | 10.3 |
| PG3 | 32 | 32 | 128 | 22.5 |
| PG4 | 32 | 32 | 64 | 13.4 |
In a related work, Velkov and Whittaker investigated antibacterial polyacrylates.238,239 Unlike polymethacrylate systems, acrylate-based polymers possess more flexible backbones, which influence their biological activity profiles. Using ATRP, they systematically varied both the type of cationic functionality (primary amines, QAS, or guanidinium groups) and the terminal group derived from the initiator. For very short acrylate oligomers, the presence of a hydrophobic dodecyl end group resulted in a marked increase in antibacterial potency, consistent with the observations of Michl and colleagues.237–239
Taken together, while end groups are often considered minor structural elements, they can have a noticeable influence in low MW oligomers. Adjusting these terminal groups may help balance antibacterial activity and biocompatibility and thus represents a factor worth considering in the design APs.
Molecular weight
The relationship between molecular weight (MW) and antibacterial efficacy tends to be more pronounced against Gram-positive bacteria than Gram-negative strains. This difference is largely attributed to the thick, highly cross-linked peptidoglycan network present in Gram-positive cell walls, which functions as a barrier restricting the penetration of large membrane-active macromolecules. This phenomenon, often referred to as the “sieving effect,” was first described by Lienkamp.240 In addition, polymer MW plays a critical role in determining biocompatibility, highlighting its dual importance in shaping both antibacterial activity and cytocompatibility.For example, Yang and coworkers170 synthesized a series of statistical polycarbonates with different MWs via ROP and found that Gram-positive bacteria displayed the “sieving effect,” whereas this behavior was not observed in Gram-negative strains. The antibacterial activity against S. aureus was strongly MW-dependent: lower-MW polymers were more effective, with MIC values of 63, 125, and 500 mg L−1 for 6 Kda, 10 Kda, and 16 Kda polymers (each with ∼50% charge density), respectively. In contrast, in Gram-negative bacteria, higher-MW polymers (16 Kda and 10 Kda) exhibited greater activity than the lower-MW analog (6 Kda), an effect linked to the enhanced hydrophobicity of the higher-MW polymers.170
In a separate study, Mishra et al.241 investigated amphiphilic methacrylamide statistical copolymers synthesized via free-radical copolymerization, employing aminopropyl methacrylamide (A) as the cationic component and benzylmethacrylamide (B) as the hydrophobic segment.241 The hydrophobic content was maintained at approximately 20 mol%, while the MW (average number DP) was systematically varied (DP = 10, 17, 27, 29, and 66) to assess antibacterial and cytotoxic properties. The results revealed a strong MW influence on antibacterial efficacy for both E. coli and S. aureus strains studied. MIC values decreased with increasing DP, indicating enhanced activity of the higher-DP polymers (Fig. 25A).241 Unlike previous study,170 no evidence of a sieving effect was observed against Gram-positive bacteria, as both Gram-positive and Gram-negative strains exhibited improved activity with increasing MW.241 However, cytotoxicity also increased with DP: polymers (AB-20)10, (AB-20)17, and (AB-20)27 caused only a 5–10% reduction in metabolic activity at 1 × and 8 × MICs, whereas higher-DP polymers (AB-20)44 and (AB-20)66 reduced cell metabolic activity by ∼70% at 1 × MIC (Fig. 25B).241
 | ||
| Fig. 25 (A) MICs of AB polymers against E. coli and S. aureus. (B) Cell viability of human juvenile fibroblasts after 24 h incubation with AB polymers at concentrations corresponding to 1 × and 8 × MICs. Adapted from ref. 241 under the terms of the CC BY-NC-ND 4.0 license, American Chemical Society, copyright 2021. | ||
Comparable MW-dependent trends were later confirmed in a more recent study by Jańczewski and coworkers, who investigated linear poly(trimethylenimine),242 further reinforcing the critical role of MW in balancing antibacterial efficacy with biocompatibility.
Amphiphilic antimicrobial polymer topologies
The topology of a polymer defines the spatial arrangement of its monomer units along the chain, which determines how hydrophobic and hydrophilic domains are distributed within the macromolecule. Variations in this arrangement influence the polymer's physicochemical characteristics and its mode of interaction with bacterial membranes. By adjusting these spatial features, the selectivity of APs can also be modulated. Advances in controlled and living polymerization methods have made it possible to precisely regulate chain topology, allowing for the preparation of well-defined cationic APs with tunable structures and functions.24,40,177,243,244Depending on the synthetic strategy, APs can adopt a wide range of architectures.24 These include simple linear or cyclic structures, as well as more complex sequence-defined architectures (such as block and alternating copolymers) and nonlinear topologies (such as hyperbranched, star, and bottle brush polymers) Each configuration provides a distinct organization of functional groups and chain mobility, resulting in a diverse range of bioactivities.
Linear polymers
Linear polymers represent the simplest and most extensively investigated topological form among APs. Their straightforward architecture provides a versatile platform for designing amphiphilic copolymers through a variety of modern polymerization techniques. Depending on the placement of functional groups, linear APs can be categorized into main-chain and side-chain types. In main-chain systems, cationic functionalities are incorporated directly along the polymer backbone, while in side-chain systems they are attached as pendant groups to the main-chain. This structural distinction strongly influences polymer conformation, charge distribution, and ultimately biological performance.Main-chain or side-chain cationic polymers
In main-chain cationic polymers, multiple cationic centers are directly incorporated along the polymer backbone, resulting in a high local charge density that enhances electrostatic attraction to negatively charged bacterial membranes.177 In a study, Zhang and coworkers reported an imidazolium-based main-chain oligomer (IBN-1) (Fig. 26A) and investigated its antimicrobial performance and biocompatibility.245 IBN-1 effectively inhibited the growth of various pathogenic bacteria and fungi. As shown in Fig. 26B–D, treatment with IBN-1 significantly suppressed Bacillus subtilis (BS) growth at concentrations of 3.9 and 15.6 µg mL−1 and exhibited rapid bactericidal kinetics against MRSA and Klebsiella pneumoniae (KP). Despite its strong antimicrobial activity, the polymer showed negligible hemolytic effects toward RBCs, even at higher concentrations (Fig. 26E and F).245 In in vivo evaluations using a S. aureus infection model, IBN-1 displayed good safety with no observable toxicity at doses up to 0.6 mg kg−1, and an LD50 of 1.72 mg mL−1, corresponding to a high TI (Fig. 26G).245 Similar findings were reported in recent studies by the Rao209 and Khan groups,207 where main-chain polysulfonium polymers exhibited potent antibacterial and antifungal activities against both planktonic and biofilm-associated pathogens, along with negligible hemolytic effects.207,209 | ||
| Fig. 26 (A) Chemical structure of the imidazolium-based main-chain oligomer IBN-1. (B) Growth curves of BS treated with IBN-1 at concentrations of 3.9 and 15.6 µg mL−1. (C) and (D) Time-dependent bactericidal activity of IBN-1 against MRSA and KP, respectively. (E) Optical micrograph of RBCs after incubation with IBN-1 in PBS at 37 °C for 1 h. (F) Hemolysis rate of RBCs determined from hemoglobin release measured at 576 nm. (G) Survival curves of mice infected with S. aureus (8 × 109 CFU) and treated with different doses of IBN-1. Reproduced from ref. 245 with permission from Elsevier, copyright 2012. | ||
In an elaborated study, Yan et al. performed a direct comparison between small-molecule cationic compounds and their corresponding polymeric analogs to evaluate the influence of polymer architecture on bioactivity.246 They synthesized imidazolium, quaternary ammonium, and 1,4-diazabicyclo[2.2.2]octane-1,4-diium (DABCO-diium) cation-based small molecules, together with their side-chain and main-chain cationic polymer derivatives.246 The results demonstrated that main-chain polymers exhibited markedly higher antibacterial activity and superior hemocompatibility than the side-chain counterparts, underscoring the influence of charge localization along the polymer backbone.246
In addition to the previously established classical pathway for intracellular uptake of APs, which is primarily driven by electrostatic interactions with negatively charged bacterial membranes and often results in membrane disruption, an alternative mechanism has recently been proposed by the Park team involving N-heterocyclic carbene (NHC)-mediated translocation.247 This pathway enables main-chain oligoimidazolium (OIM) polymers to efficiently access the bacterial cytosol, thereby reaching intracellular DNA targets. OIM polymers containing an acidic C(2)–H group can undergo transient deprotonation under physiological conditions, generating short-lived, neutral, and hydrophobic NHC species (Fig. 27(i)). As only a fraction of the imidazolium units are converted, the polymer adopts a dynamic amphiphilic configuration composed of both positively charged imidazolium groups and neutral carbene moieties. This amphiphilic transition is critical, as it facilitates insertion into the hydrophobic interior of bacterial membranes (Fig. 27(ii)), where the carbene species are stabilized in the low-water environment. Translocation across the membrane is further enhanced by the bacterial membrane potential, which drives OIM polymers toward the inner leaflet (Fig. 27(iii)). At the cytosolic interface, the NHC species are quenched by water and revert to their cationic imidazolium state, enabling the polymer to pass into the cytosol (Fig. 27(iv)). Once internalized, OIM polymers exert their antibacterial activity through condensation with bacterial DNA. Importantly, cytosolic uptake strongly correlates with antibacterial efficacy: derivatives capable of efficient NHC formation exhibit low MIC values and high intracellular accumulation, whereas those unable to form carbene species fail to enter the cytosol and show negligible antibacterial activity.247
 | ||
| Fig. 27 Proposed mechanism of NHC-mediated uptake of antibacterial polymers. Adapted from ref. 247 under the terms of the CC-BY-NC-ND license, Springer Nature, copyright 2025. | ||
In contrast to main-chain APs, the synthesis of side-chain polymers benefits from a broad range of living and controlled polymerization techniques, allowing the creation of structurally diverse polymer libraries with tunable molecular characteristics.46,248 These advances in controlled polymerization methodologies enable precise adjustment of molecular parameters—such as molecular weight, monomer composition, and chain architecture, thereby facilitating the production of well-defined copolymers with predictable topology and functionality.50,249
Statistical polymers
First, it is important to clarify that in this section, we refer to statistical copolymers, i.e., copolymers with unspecified monomer sequences, in contrast to more defined macromolecular architectures. This designation does not imply a truly random monomer distribution in the strict IUPAC definition, as copolymerization parameters (r1, r2) are generally not determined. Statistical copolymers represent one of the most extensively explored polymeric architectures in the development of APs. Their straightforward synthesis, which does not require precise sequence control, together with their proven antibacterial performance, has made them a fundamental design strategy for developing amphiphilic APs.24,40 Most reported antimicrobial copolymers are derived from functionalized poly(methacrylates), poly(methacrylamides), poly(acrylates), poly(acrylamides), poly(oxazolines), poly(β-lactams), poly(norbornenes), and poly(carbonates). Moreover, recent research efforts have focused on elucidating how specific structural features within these copolymers influence antibacterial activity and compatibility with host cells.250Cyclic polymers
A relatively unexplored topology in APs is the cyclic architecture, owing to the complex and labor-intensive procedures required to obtain highly purified cyclic polymers (CPs). Characterized by ring-like structures without chain ends, these polymers display distinctive physicochemical properties, including smaller hydrodynamic size, enhanced stability, and slower degradation than their linear analogs. Their constrained topology also affects chain mobility and self-assembly behavior, contributing to improved colloidal stability, extended circulation time, and higher drug-loading efficiency. Owing to these advantages, CPs are emerging as promising candidates for biomedical and antibacterial applications.For example, the Grayson group251 demonstrated that polymer architecture plays a crucial role in modulating cytotoxicity. They showed that cyclic poly(ethylene imine) exhibits markedly lower toxicity toward normal human cells compared with its branched analogs.251 Building on the concept of topology-driven bioactivity, Alabi and coworkers later synthesized a library of oligothioetheramide (oligoTEA) macrocycles through a one-pot acid-catalyzed cascade reaction.61 This versatile method enabled the rapid generation of more than 20 macrocyclic structures with diverse compositions. Biological evaluations revealed that oligoTEA macrocycles containing only 2–3 cationic charge centers displayed potent antibacterial activity against both Gram-positive and Gram-negative bacteria.61
Extending this understanding, Duan et al. reported the synthesis of cyclic polymethacrylates with various compositions via intrachain click cyclization of α-alkyne–ω-azido hetero-difunctional linear precursors prepared by ATRP (Fig. 28).60 The resulting cationic CPs demonstrated enhanced affinity toward negatively charged bacterial membranes, promoting more efficient membrane disruption compared to their linear analogs. Consequently, these non-linear binary polymers exhibited superior antibacterial potency against both Gram-negative and Gram-positive bacteria (Table 7), while displaying slightly reduced cytotoxicity relative to the corresponding linear counterparts.60
 | ||
| Fig. 28 Synthesis scheme of linear and CPs. Adapted from ref. 60 with permission from Royal Society of Chemistry, copyright 2020. | ||
| Polymers | MIC (µg mL−1) | |
|---|---|---|
| E. coli | S. aureus | |
| Linear P(DMAEMA + 70-co-MMA30) | 417 | 156 |
| Cyclic P(DMAEMA + 70-co-MMA30) | 130 | 78 |
| Linear P(DMAEMA + 55-co-MMA45) | 156 | 78 |
| Cyclic P(DMAEMA + 55-co-MMA45) | 78 | 40 |
| Linear P(DMAEMA + 40-co-MMA60) | 39 | 78 |
| Cyclic P(DMAEMA + 40-co-MMA60) | 13 | 36 |
| Linear P(DMAEMA + 50-co-BMA50) | 69 | 156 |
| Cyclic P(DMAEMA + 50-co-BMA50) | 33 | 78 |
| Linear P(DMAEMA + 40-co-BMA60) | 78 | 261 |
| Cyclic P(DMAEMA + 40-co-BMA60) | 26 | 126 |
| Linear P(DMAEMA + 30-co-BMA70) | 209 | 625 |
| Cyclic P(DMAEMA + 30-co-BMA70) | 52 | 304 |
| Linear P(DMAEMA + 70-co-tBMA30) | 78 | 78 |
| Cyclic P(DMAEMA + 70-co-tBMA30) | 33 | 40 |
| Linear P(DMAEMA + 60-co-tBMA40) | 78 | 78 |
| Cyclic P(DMAEMA + 60-co-tBMA40) | 39 | 50 |
| Linear P(DMAEMA + 50-co-tBMA50) | 113 | 313 |
| Cyclic P(DMAEMA + 50-co-tBMA50) | 20 | 156 |
Inspired by the earlier binary cyclic APs, the Boyer group for the first time synthesized cyclic terpolymers to evaluate their antibacterial performance and biocompatibility.252 Initially, the team designed a benzaldehyde-functionalized RAFT agent (Fig. 29), which was employed to synthesize amphiphilic cationic linear terpolymers via thermal RAFT polymerization. These statistical linear precursors were subsequently cyclized through a hetero-Diels–Alder click reaction to yield cyclic terpolymers (Fig. 29). The resulting terpolymers displayed potent anti-infective activity (MIC90 = 8–256 µg mL−1), effectively eradicating MDR Gram-negative Pseudomonas aeruginosa PA37 more efficiently than conventional antibiotics such as gentamicin and ciprofloxacin. Moreover, the cyclic terpolymers rapidly disrupted bacterial membranes, achieving 99.99% bacterial killing within 15 minutes, and exhibited superior biocompatibility than their linear counterparts, highlighting their excellent safety profile (CPs > LPs).252 Similar biocompatibility trends were also reported for polynorbornene-based cyclic polymers synthesized via a combination of ring-expansion metathesis polymerization and click chemistry, which showed higher selectivity than their linear analogs.253
 | ||
| Fig. 29 Synthesis scheme of linear and cyclic terpolymer. Adapted from ref. 252 under the terms of the CC-BY license, Royal Society of Chemistry, copyright 2024. | ||
Recently, the Jańczewski group254 redirected the conventional step-growth polymerization route used for preparing highly antimicrobial ionenes toward the synthesis of macrocyclic QAS (MQAs) by applying a high-dilution strategy. Biological evaluations demonstrated that the resulting MQAs exhibited strong antimicrobial activity and markedly higher selectivity (SI = 11–91) compared with their linear polymeric counterparts (SI = 3–7).254
Overall, continued advancements in synthetic methodologies and polymer science are anticipated to drive significant innovation in the synthesis of CPs.255,256 These developments will enable deeper exploration of cyclic topologies and enhance their contribution to APs research, paralleling the success of naturally derived cyclic lipopeptide antibiotics.
Advanced architecture design
Driven by the structural complexity and functional efficiency of natural proteins, researchers are advancing the design of amphiphilic APs with higher-order architectures. Progress in polymer chemistry has enabled the controlled synthesis of macromolecules with sophisticated sequence-defined architectures (such as block and alternating copolymers) as well as nonlinear topologies, including hyperbranched, dendritic, star, and bottle brush structures, surpassing the limitations of traditional linear or statistical systems. These complex architectures provide versatile platforms for optimizing amphiphilic balance, enhancing antimicrobial potency, and improving biocompatibility through precise spatial organization of functional groups.Block copolymers
The synthesis of block copolymers often relies on controlled/living polymerization strategies, including ROP, ATRP, RAFT, and PET-RAFT, allowing the preparation of well-defined architectures with tailored amphiphilic segments.24,40,128,257,258 These polymers exhibit distinct segregation between hydrophobic and hydrophilic domains, providing an effective means to regulate amphiphilic balance and promote self-assembly into ordered nanostructures. Such structural organization differentiates them from homopolymers and random copolymers, where chain composition is less spatially controlled.128Georges and colleagues investigated the antibacterial properties of all-siloxane block and statistical copolymers containing quaternary ammonium functionalities.259 Both copolymer types showed strong bactericidal activity in aqueous media against E. coli and S. aureus, with no notable difference in antibacterial potency between the block and statistical arrangements.259 Later, this observation was further supported by Liu260 and Kuroda groups.261 For instance, Kuroda studied amphiphilic block and random copolymers of poly(vinyl ether) derivatives synthesized via cationic polymerization (Fig. 30A).261 In this study, block and random copolymers with comparable monomer compositions exhibited similar antibacterial effects against E. coli (Fig. 30B). However, the block copolymers showed no detectable hemolytic activity even at the concentrations up to 1000 µg mL−1 (Fig. 30C). This high hemocompatibility is attributed to the ability of the block copolymers to form single-molecule cationic particles that shield their hydrophobic domains, thereby reducing direct interactions with RBC membranes (Fig. 30A).261
 | ||
| Fig. 30 (A) Structures of block and random copolymers and their interactions with bacterial and RBC membranes. (B) Bactericidal activity (BC99.9) against E. coli for block and random copolymers plotted as a function of the molar percentage of hydrophobic isobutyl vinyl ether (IBVE) units. (C) Hemolysis profiles of block and random copolymers at increasing polymer concentrations. Reproduced from ref. 261 with permission from American Chemical Society, copyright 2011. | ||
Extending this understanding, Boyer and colleagues55 further emphasized the importance of monomer sequence and local organization within block copolymers relative to the global composition of random copolymers. By examining a series of 32 well-defined multiblock ternary copolymers synthesized in a one-pot process via PET-RAFT polymerization, they observed that both antibacterial activity and hemocompatibility were closely governed by the monomer arrangement within blocks. In particular, the localized hydrophobic-to-hydrophilic ratio within amphiphilic domains was found to play a decisive role in determining bioactivity (Table 8).55
| Chemical structure | Polymerization technique | Test subject | MIC (µg mL−1) | HC50/IC50 (µg mL−1) | Ref. |
|---|---|---|---|---|---|
| a Biocidal concentration for 99.9% killing. | |||||

|
ATRP | E. coli | 54–97 (EC) | 250–1290 (HC50) | 260 |
| S. aureus | 34–67 (SA) | ||||

|
Cationic Polymerization | E. coli | 1.6–2.4a | >1000 (HC50) | 261 |

|
PET-RAFT | P. aeruginosa | 32–64 | 1000 (HC50) | 55 |
| E. coli | (PA, EC) | ||||
| A. baumannii | 128 (AB) | ||||

|
ATRP | S. aureus | 16–64 (SA) | >2500 (HC50) | 262 |
| S. epidermidis | 16–32 (SE) | ||||
| P. aeruginosa | 250 (PA) | ||||
| C. parapsilosis | 8–16 (CP) | ||||

|
RAFT | E. coli | 32–512 (EC) | >1024 (HC10) | 263 |
| P. aeruginosa | 32–64 (PA) | 330–>1024 (IC50, 3T3), 76–179 (IC50, Caco2) | |||
| S. aureus | 8–128 (SA) | ||||
| S. epidermidis | 4–32 (SE) | ||||
Fernández-García and coworkers262 employed ATRP to synthesize well-defined block and statistical copolymers composed of methacrylate units and a glycomonomer. The resulting copolymers exhibited broad-spectrum antimicrobial activity against Gram-negative and Gram-positive bacteria as well as yeast. Incorporation of carbohydrate-based pendant groups significantly improved hemocompatibility while preserving strong antimicrobial efficacy (Table 8).262 Hartlieb and Perrier further explored how the spatial distribution of hydrophobic and cationic functionalities affects the antimicrobial performance and selectivity of polymer systems.46 Using RAFT polymerization, they synthesized statistical (S), diblock (D), and highly segmented multiblock copolymers (M) with controlled architectures (Fig. 31A).46 Their results revealed that diblock and multiblock copolymers exhibited superior antibacterial activity compared to statistical analogs (Table 9). Notably, in polymers containing a lower proportion of cationic monomer, the multiblock design (M30) achieved remarkably enhanced selectivity toward P. aeruginosa (SI = >128) and Staphylococcus epidermidis (SI = >256) (Table 9), demonstrating how segmental organization can be leveraged to optimize antibacterial potency and biocompatibility (Fig. 31B).
 | ||
| Fig. 31 (A) Schematic illustration of the molecular structures, compositions, and segmental arrangements of the synthesized polymers. (B) Illustration of the interactions of different copolymer architectures with bacterial and mammalian cell membranes. Reproduced from ref. 46 with permission from American Chemical Society, copyright 2017. | ||
| Sample | HC10 (µg mL−1) | cH (µg mL−1) | MIC (µg mL−1) | Selectivity | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| E. coli (µg mL−1) | P. aeruginosa (µg mL−1) | S. aureus (µg mL−1) | S. epidermidis (µg mL−1) | E. coli | P. aeruginosa | S. aureus | S. epidermidis | |||
| Notes: cH is the lowest concentration polymer concentration at which visible RBC aggregation is observed. Selectivity is calculated as the ratio of the hemocompatibility concentration (the lower value between HC10 and cH) to the MIC for each bacterial strain. | ||||||||||
| H0 | >1024 | >1024 | >1024 | >1024 | >1024 | >1024 | >1 | >1 | >1 | >1 |
| H100 | 512 | 16 | 4 | 4 | 4 | 32 | 4 | 4 | 4 | 0.5 |
| S30 | >1024 | 32 | >1024 | >1024 | >1024 | 32 | 0.03 | 0.03 | 0.03 | 1 |
| S50 | >1024 | 32 | 64 | 128 | 8 | 2 | 0.5 | 0.25 | 4 | 16 |
| S70 | >1024 | 32 | 64 | 64 | 4 | 2 | 0.5 | 0.5 | 8 | 16 |
| M30 | >1024 | >1024 | 128 | 8 | 64 | 4 | >8 | >128 | >16 | >256 |
| M50 | >1024 | 32 | 1024 | 64 | 32 | 8 | 0.03 | 0.5 | 1 | 4 |
| M70 | >1024 | 32 | 1024 | 32 | 4 | 4 | 0.03 | 1 | 8 | 8 |
| D30 | >1024 | >1024 | 512 | 32 | 128 | 32 | >2 | >32 | >8 | >32 |
| D50 | >1024 | >1024 | 64 | 64 | 8 | 4 | >16 | >16 | >128 | >256 |
| D70 | >1024 | >1024 | 32 | 32 | 8 | 4 | >32 | >32 | >128 | >256 |
More recently, in a 2025 study, Klinger and coworkers211 developed sulfonium-based amphiphilic block copolymers (BCPs) composed of a cationic sulfonium-containing block with a neutral hydrophilic PEG methacrylate block to enhance cytocompatibility. These BCPs exhibited potent antimicrobial activity and retained membrane-disruptive capability. When used in combination with conventional antibiotics such as penicillin G and ciprofloxacin, the polymers produced pronounced synergistic effects, significantly lowering the effective doses of both agents while maintaining strong antibacterial efficacy and minimal cytotoxicity.211
Alternating copolymers
Polymers with alternating monomer sequences have attracted attention as antibacterial agents because they offer exceptional control over sequence order and molecular uniformity. However, the limited availability of monomer pairs that naturally form alternating structures through chain-growth polymerization has restricted the number of studies on such systems. Although step-growth approaches can generate alternating architectures, they typically yield polymers with broad MWDs.118Parker and Sampson reported that ROMP of 1-substituted cyclobutenes and cyclohexenes affords alternating copolymers with well-defined periodic spacing between functional groups.264 These materials, incorporating regularly positioned cationic and hydrophobic moieties, displayed significantly enhanced antibacterial performance and very low hemolytic activity compared with random analogs, achieving over 100-fold selectivity in the most effective examples. Notably, increasing the hydrophobicity between adjacent cationic units did not substantially increase cytotoxicity, which contrasts with trends observed for other AP systems.264 Complementary study by the Haldar group on alternating maleimide–isobutylene265 copolymers further reinforced these findings, demonstrating that precise alternation of functional groups represents a promising yet underexplored strategy for developing highly active and biocompatible antibacterial polymers.265
Hyperbranched polymers
Hyperbranched polymers (HPs) are highly branched macromolecules with a three-dimensional globular architecture and abundant terminal functional groups.59,266–268 Their unique structure offers high surface functionality and tunable amphiphilic balance, making them excellent candidates for APs design.59To investigate this, Kanai and colleagues269 reported the first synthesis of hyperbranched poly(cyanurateamine) and poly(triacrylatetrimine) using a Michael addition reaction. The resulting water-soluble polymers exhibited broad-spectrum antibacterial activity against both Gram-positive and Gram-negative bacteria. Notably, poly(triacrylatetrimine) demonstrated superior activity against antibiotic- and antiseptic-resistant strains such as P. aeruginosa and Serratia marcescens compared with poly(cyanurateamine).269 Similarly, the Yang group270 developed long-subchain hyperbranched poly(aminoethyl acrylate) (lhb-PAEA), which achieved >99.99% killing of E. coli and >98% killing of S. aureus at concentrations of 4 µg mL−1. The lhb-PAEA polymers also displayed excellent biocompatibility, with hemolysis below 35% even at concentrations up to 1024 µg mL−1.270
Inspired by these advances, the Boyer group systematically investigated the influence of polymer topology and chain length on the antibacterial and hemolytic performance of ternary APs (Fig. 32).59 The HPs demonstrated superior bioactivity compared with their linear random and block counterparts. In particular, the hyperbranched structure containing 2-ethylhexyl (EH) hydrophobic side chains exhibited comparable antimicrobial activity (MIC = 64 µg mL−1) and more than a fourfold increase in hemocompatibility (HC50 = >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 µg mL−1) compared to the linear analogs, with no observable hemagglutination. Moreover, these HPs displayed potent bactericidal effects, eliminating ≥99% of planktonic and 90% of biofilm P. aeruginosa cells.59 Consistent with their findings, other reported HP systems have also displayed strong antibacterial activity, low hemolysis, and superior selectivity compared with their linear counterparts.271–273
000 µg mL−1) compared to the linear analogs, with no observable hemagglutination. Moreover, these HPs displayed potent bactericidal effects, eliminating ≥99% of planktonic and 90% of biofilm P. aeruginosa cells.59 Consistent with their findings, other reported HP systems have also displayed strong antibacterial activity, low hemolysis, and superior selectivity compared with their linear counterparts.271–273
 | ||
| Fig. 32 Effect of topology (random, block, and HP) on bioactivity of polymers. Adapted from ref. 59 with permission from Royal Society of Chemistry, copyright 2018. | ||
Star polymers
Star polymers (SPs) comprise multiple polymeric arms radiating from a central core, forming a well-defined and compact three-dimensional architecture.56,274–276 This structural precision enables controlled multivalent presentation of functional groups, which enhances membrane interactions and antimicrobial efficacy. By carefully tuning parameters such as arm length, composition, and arm number, SPs can be engineered to achieve an optimal balance between antibacterial potency and cytocompatibility.277Several pioneering studies by Qiao,15,57,278 Park,279,280 Yang,281,282 Simpson,57,283 and Perrier56,284 have demonstrated the potential of star-shaped APs. In addressing the urgent challenge posed by MDR Gram-negative “superbugs,” Simpson and Qiao's group developed a series of star-shaped polymers termed structurally nanoengineered antimicrobial peptide polymers (SNAPPs) (Fig. 33A).57 Their study systematically investigated how two key structural parameters—the number of arms and the arm length affect antibacterial activity and biocompatibility (Fig. 33B).57 The findings revealed that increasing both arm number and arm length enhanced antibacterial potency, attributed to the higher local density of polypeptide arms and the increased α-helical content within the structure. The SNAPP architecture was found to directly influence bacterial membrane disruption and killing mechanisms. Although cytotoxicity also rose with greater arm number and length, TI analysis identified a 16-arm SNAPP and a more easily synthesized 4-arm variant as the most effective and biocompatible candidates. In vivo assessments further confirmed that the optimal SNAPP exhibited excellent safety, showing no evidence of systemic toxicity in treated mice.57
 | ||
| Fig. 33 (A) Synthesis SNAPPs via NCA-ROP. (B) Arm number and arm length of SNAPPs. Reproduced from ref. 57 with permission from Wiley, copyright 2018. | ||
Park et al. developed a novel class of antibacterial SPs measuring 14–26 nm in diameter, composed of mixed polylysine and glycopolymer arms. These polymers exhibited potent antibacterial activity against Gram-positive bacteria, including MRSA and VRE, achieving low MIC values of 16 µg mL−1. Importantly, they remained non-hemolytic (HC50) more than 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 µg mL−1 and demonstrated excellent compatibility with host cells.
000 µg mL−1 and demonstrated excellent compatibility with host cells.
Inspired by these highly biocompatible antibacterial SPs, Boyer group subsequently conducted a comprehensive investigation into the effect of polymer topology on biological performance by synthesizing 27 amphiphilic polymers, including linear polymers (LPs), hyperbranched polymers (HPs), and 3- and 4-arm star polymers (SPs), using PET-RAFT polymerization (Fig. 34).285 Comparative evaluation revealed that HPs with identical hydrophobic-to-hydrophilic ratios exhibited 2–4-fold higher blood compatibility than LPs, while SPs with lower hydrophobic content were non-hemolytic and maintained strong anti-infective properties against Gram-negative bacteria comparable to both LPs and HPs (MIC90 = 32–64 µg mL−1). Overall, SPs demonstrated the most favorable balance between antibacterial efficacy and hemocompatibility, following the selectivity trend SPs > HPs > LPs.285
 | ||
| Fig. 34 Synthesis scheme of LPs, HPs, and 4-arm SPs prepared via PET-RAFT polymerization. Adapted from ref. 285 under the terms of the CC-BY license, Elsevier, copyright 2024. | ||
Recently, Perrier group. developed a library of amphiphilic SPs and their linear counterparts using acrylamide-based monomers synthesized via RAFT polymerization (Fig. 35).56 The copolymers were systematically varied in monomer composition and molecular weight to examine how architectural differences influence biological performance. The statistical SP (S-SP 25) exhibited significantly enhanced antibacterial activity against P. aeruginosa (MIC = 32 µg mL−1) compared with its linear analog (S-LP 100; MIC = 512 µg mL−1) (Fig. 35). Electron microscopy revealed that the star topology promoted bacterial cell aggregation, accounting for its increased antibacterial efficacy, but also led to higher RBCs aggregation than the linear polymers. To address this, repositioning the cationic block toward the core of the star structure effectively minimized cell aggregation while preserving strong antibacterial activity, particularly in the smallest SP.56
 | ||
| Fig. 35 Structure of star and linear copolymer. Adapted from ref. 56 under the terms of the CC-BY 4.0 license, American Chemical Society, copyright 2023. | ||
In a subsequent study, the same group284 employed an arm-first synthesis strategy to construct a series of amphiphilic cationic core-cross-linked star (CCS) copolymers through RAFT polymerization.284 These copolymers incorporated both diblock and homopolymer arms. The results revealed that transforming from diblock to miktoarm CCS topologies led to improved antibacterial efficacy. Although none of the synthesized polymers exhibited hemolytic activity toward sheep RBCs, an increased tendency for RBC aggregation was observed with higher surface exposure of cationic functionalities. Interestingly, increasing the number of arms did not further enhance antibacterial potency compared to 4-arm systems, likely due to the partial shielding of cationic groups within the polymer core, which reduced their membrane-interaction capacity. Nonetheless, miktoarm structures showed marginally improved performance in overcoming this limitation.284
Collectively, extensive research has shown that amphiphilic antibacterial copolymers featuring star or dendritic star architectures exhibit greater antimicrobial efficacy and improved biocompatibility compared with their linear analogs, reflecting higher selectivity.157,286–288 This enhanced performance is often attributed to the intrinsic multivalency and compact nature of the star architecture, where multiple terminal groups create locally concentrated active sites. Such spatial organization can synergistically promote the initial adsorption of polymers onto bacterial membranes, enhancing disruption efficiency. Conversely, the restricted flexibility of the polymer arms may reduce nonspecific interactions with host cell membranes, resulting in minimal hemolytic activity without compromising anti-infective potency.
Bottle brush polymers
This section focuses exclusively on one-dimensional (1D) polymeric antibacterial brushes, including molecular brush or bottle brush topologies that mimic the structural organization of HDPs, while excluding surface-grafted coatings and antifouling systems that fall outside the scope of this literature review. Bottle brush polymers (BPs) are composed of a linear backbone densely grafted with polymeric side chains, typically approaching one graft per repeating unit. This dense grafting restricts side-chain mobility and stretches the backbone, giving rise to unique characteristics such as worm-like conformations, compact molecular size, and pronounced chain-end effects.289,290 These structural attributes confer distinctive physicochemical and biological properties, making BPs attractive platforms for designing next-generation antimicrobial materials.Despite their potential, only a limited number of studies have explored antibacterial BPs, primarily due to the complexity and time-intensive nature of their synthesis. Notably, Hartlieb and coworkers reported a few examples of bottle brush-type antibacterial polymers.62,63,291 In one study, they developed HDP mimics BPs synthesized via a combination of RAFT and ROMP technique (Fig. 36).62 This design enabled the multivalent presentation of antibacterial subunits, which significantly enhanced anti-infective potency and hemocompatibility, thereby improving selectivity toward bacterial over mammalian cells. Membrane integrity assays confirmed that the BPs caused rapid and potent bacterial membrane disruption. Overall, these multivalent BPs exhibited superior antibacterial activity and selectivity compared to their linear counterparts.62
 | ||
| Fig. 36 Synthesis scheme of bottle brush polymer. (A) Synthesis of macromonomers via RAFT polymerization of acrylamides using a norbornene-functionalized chain transfer agent (CTA), followed by polymerization through ROMP to form bottle brush copolymers. (B) Preparation of homograft bottle brush copolymers (B-S-XX) from macromonomers incorporating both monomers in a statistical manner (XX denotes the BocAEAm content in percentage). (C) Formation of heterograft bottle brush copolymers (B-H-XX) by ROMP of macromonomers composed of homopolymers in varying ratios. (D) Schematic illustration showing the interaction of deprotected cationic bottle brush copolymers with bacterial membranes, leading to membrane disruption. Reproduced from ref. 62 with permission from American Chemical Society, copyright 2020. | ||
In a subsequent study, the same group explored how anisotropy within bottle brush structures influences their bioactivity.63 Using a one-pot RAFT polymerization approach, they synthesized a series of BPs with constant chemical composition but systematically varied aspect ratios by adjusting backbone and side-chain lengths (backbone length vs. graft length). It was found that BPs with longer, more flexible side chains were more hydrophilic and exhibited enhanced intramolecular hydrogen bonding, leading to self-segregation into unimolecular micelles with hydrophobic cores and cationic outer shells. This structural organization enhanced bacterial membrane interactions and antibacterial properties against Gram-positive and Gram-negative bacteria, while maintaining excellent biocompatibility, achieving SI of 640.63
Overall, bottle brush polymers represent a promising frontier in AP design, as their tunable topology and multivalent functionality enable strong antibacterial efficacy with minimal cytotoxicity. Their structural precision, controllable anisotropy, and high selectivity position them as compelling therapeutic candidates and an exciting direction for future research in AP development.
Stimuli-responsive and self-immolative antimicrobial polymers
Recent reviews by Shabat,292 Park,293 Gillies,294 Hartlieb,244 and Tang295 have highlighted the growing significance of stimuli-responsive and self-immolative polymers (SIPs) in biomedical science and as innovative antimicrobial platforms. Stimuli-responsive polymers undergo controlled structural changes in response to pH, redox potential, light, or enzymatic triggers, enabling spatiotemporal regulation of antibacterial activity and on-demand degradation to minimize long-term cytotoxicity.Within this class, SIPs are distinguished by their ability to depolymerize in a cascade manner upon activation by a specific trigger.268 They consist of labile repeating units capped with a stimuli-sensitive end group that acts as a molecular “lock.”294 Once this end-cap is cleaved by enzymatic, acidic, or reductive stimuli—the polymer undergoes head-to-tail self-immolation, breaking down completely into benign small molecules. This controlled degradation mechanism enables the creation of transient antimicrobial materials that activate selectively in infection environments and degrade harmlessly after use.
To address the broader environmental challenge, where large quantities of antibiotics are consumed each year and a significant portion is released into the environment, researchers have focused on designing APs that are both effective and degradable. Recently, Zhang and colleagues reported a series of imidazolium-based polymers incorporating pH-degradable linkers (Fig. 37).69 These polymers exhibited broad-spectrum antimicrobial activity against both Gram-negative (MIC = 8 and 31 µg mL−1) and Gram-positive bacteria (MIC = 4 and 8 µg mL−1), as well as C. albicans (31–125 µg mL−1). Moreover, the incorporation of degradable linkers facilitated environmental breakdown into inactive small molecules. Both the polymers and their degradation products did not induce bacterial resistance and showed moderate biodegradation in surface water.69
 | ||
| Fig. 37 Chemical structure of pH-degradable imidazolium-based APs. Reproduced from ref. 69 with permission from American Chemical Society, copyright 2021. | ||
To mitigate the persistent toxicity and selectivity challenges associated with cationic APs, Wong et al. developed a β-galactosidase-responsive antibacterial dendron,296 in which cationic groups were enzymatically masked and subsequently unveiled through β-galactosidase activation to regenerate primary ammonium.296 This approach led to an approximately fourfold enhancement in anti-infective efficacy, accompanied by low hemolysis.296
Polymers capable of degrading in a specifically triggered manner offer advantages over those that undergo uncontrolled or passive degradation. A notable class of such materials is SIPs, which depolymerize completely upon exposure to a specific stimulus. Leveraging this concept, the Palermo group reported the first example of antibacterial SIPs.297 They synthesized poly(benzyl ether)s containing pendant allyl side chains and silyl ether end-caps (responsive to fluoride ions). The side chains were subsequently modified with cysteamine hydrochloride through photo-initiated thiol–ene chemistry. The resulted cationic poly(benzyl ether)s demonstrated broad-spectrum antibacterial activity, although they were initially highly toxic. Upon activation, the polymers underwent stimulus-induced depolymerization into low MW fragments, which retained bactericidal efficacy while significantly reducing hemolytic activity.297
Aligning with the emerging direction of designing smart and degradable APs,70,73 growing research efforts have focused on lipoic acid (LA)-based degradable systems. As an abundant, naturally occurring, disulfide-containing compound, LA provides a versatile platform for constructing redox-responsive and biodegradable polymers. Utilizing these dynamic disulfide linkages, the Becker and Tang groups advanced the field of degradable antibacterial polymers in 2025.73 Recognizing that HDP-mimetic linear polymers, although widely explored in preclinical studies, often suffer from limited stability and undesired toxicity, they proposed that fully degradable cyclic architectures could offer a promising alternative. To address the synthetic challenges historically associated with cyclic systems, they developed an efficient one-pot cascade approach for generating cationic cyclic oligo(disulfides) (CCOs) from lipoic acid derivatives (Fig. 38).73 This method involves an initial ROP step followed by in situ cyclo-depolymerization, yielding well-defined cyclic oligomers with intrinsic redox-responsiveness. The resulting CCOs demonstrated broad-spectrum antibacterial activity, achieving a 5.43-log reduction in E. coli within 5 minutes, and notably did not induce bacterial resistance in vitro. Furthermore, by exploiting glutathione-triggered degradation, the cytotoxicity of CCOs was significantly reduced. Fine-tuning of the cationic-to-hydrophilic balance further enhanced serum stability, resulting in an impressive SI (HC50/MIC = >1280) against MRSA. In an infected animal model, CCOs also exhibited strong in vivo antibacterial performance.73
 | ||
| Fig. 38 Structure and bioactivity of cyclic antibacterial polymer. Reproduced from ref. 73 with permission from American Chemical Society, copyright 2025. | ||
Following this recent progress in LA-based degradable antimicrobial systems, the Boyer group also developed a series of degradable, disulfide-containing ternary antibacterial polymers incorporating benzyl lipoate—a derivative of lipoic acid synthesized via RAFT polymerization (Fig. 39).67 In this design, benzyl lipoate serves as the hydrophobic moiety and is copolymerized with cationic and hydrophilic monomers. Most of the resulting polymers displayed potent antimicrobial activity against MDR P. aeruginosa (MIC90 = 64–256 µg mL−1), along with enhanced hemocompatibility (HC50 = 1000–2000 µg mL−1) and clear redox-responsive degradability.67
 | ||
| Fig. 39 Scheme of terpolymer degradation under redox condition. Adapted from ref. 67 under the terms of the CC-BY license, Wiley, copyright 2025. | ||
Extending this degradable design strategy to mitigate the toxicity commonly associated with HDPs and cationic APs, the same group further diversified polymer topology by developing ‘arm-first’ star polymers (SPs) incorporating disulfide-based core cross-linkers (Fig. 40A).298 This architectural evolution enabled the integration of structural complexity with stimuli-responsive degradability.
 | ||
| Fig. 40 (A) Structure and bioactivity of degradable “arm-first” SP. (B) Bacterial membrane activity of LP-30, SP-30-C2, and colistin. (B) Outer membrane sensitization of E. coli (EC K12) at 1 × MIC to the lytic action of sodium deoxycholate, expressed as the percentage optical density (485 nm) of treated samples relative to the negative control (PBS containing sodium deoxycholate). (C)–(E) Inner membrane disruption study on E.coli where the membrane permeability variation, as quantified by a dimensionless constant (the fluorescence fold change between the treatment group and the negative control group (PBS), measured at excitation and emission wavelengths of 544 and 622 nm, respectively). Reproduced from ref. 298 with permission from American Chemical Society, copyright 2025. | ||
Comparative antibacterial evaluations demonstrated that LPs exhibited overall superior antibacterial activity against all three Gram-negative bacterial strains when compared with both homo- and miktoarm SP analogs. Notably, the lead copolymers LP-30 and degradable SP-30-C2 displayed potent activity against the multidrug-resistant P. aeruginosa strain PA37, with MIC90 values of 32 and 64 µg mL−1, respectively. Both polymers induced rapid and efficient disruption of the outer and inner bacterial membranes, outperforming the reference antimicrobial peptide colistin, as confirmed by membrane permeabilization assay (Fig. 40B–E). Importantly, despite their slightly reduced antibacterial potency, SPs exhibited markedly enhanced hemocompatibility, with HC50 values in the range of 1000–2000 µg mL−1, even at elevated hydrophobic contents (30%). This reflected into significantly improved SI relative to their linear counterparts, underscoring the critical role of polymer topology in balancing antimicrobial efficacy and biocompatibility. Furthermore, incorporation of disulfide bonds within the SP cores imparted redox-triggered degradability, enabling efficient structural disassembly under reducing conditions (Fig. 40A).298 Such stimuli-responsive behavior highlights the potential of these SPs to act as degradable antibacterial platforms or delivery vehicles, capable of releasing active polymer arms in infection-relevant environments while promoting controlled biodegradation and minimizing long-term systemic toxicity.298
Taken together, stimuli-responsive and self-immolative APs offer precise control over bactericidal activity, enhancing both efficacy and biocompatibility while reducing long-term cytotoxicity associated with polymer accumulation. These smart materials represent a promising strategy to combat resistant pathogens through spatiotemporally regulated antimicrobial action and improved safety profiles.
Non-amphiphilic antimicrobial polymers
It is well established from the studies discussed above that classical APs typically exhibit amphiphilic architectures comprising cationic and hydrophobic components, which play a central role in governing antibacterial activity, biocompatibility, and selectivity. Additional hydrophilic segments can be incorporated to modulate solubility and biological performance. However, recent work by Hartlieb et al. challenges this long-standing paradigm by demonstrating that amphiphilicity is not an absolute requirement for antimicrobial activity in poly(meth)acrylate and poly(meth)acrylamide systems.299 In their study, non-amphiphilic poly(arylamides), composed of cationic monomers and hydrophilic comonomers (Fig. 41) and entirely lacking hydrophobic residues, were shown to achieve potent and selective antibacterial activity. Notably, the least amphiphilic polymer, MAM70 (Fig. 41), emerged as one of the most promising APs, exhibiting a MIC50 of 22.9 ± 2.8 µg mL−1 against E. coli. Instead of relying on the conventional mechanism of hydrophobic insertion into bacterial lipid bilayers, these polymers utilize hydrogen-bond-mediated interactions with bacterial membrane headgroups. This interaction induces membrane disorder and promotes a distinct polymer clustering effect at the membrane interface. Such clustering enables multivalent interactions that effectively destabilise bacterial membranes while minimising nonspecific toxicity toward mammalian cells, as evidenced by HC10 = 10240 µg mL−1 and CC50 = 1006 ± 100 µg mL−1, resulting in high selectivity (448 ± 54) and a TI of 44 ± 6.9 (Fig. 41). This emerging design strategy expands the accessible structural space of (meth)acrylate/(meth)acrylamide-based antimicrobial polymers and highlights that, beyond amphiphilicity, alternative non-hydrophobic interactions can be exploited to fine-tune antibacterial efficacy and selectivity, enabling the development of potent yet biocompatible APs.299 | ||
| Fig. 41 Chemical structure and bioactivity of the non-amphiphilic MAM70 antimicrobial polymer. Adapted and redrawn from ref. 299 under the terms of the CC-BY license, Wiley, copyright 2025. | ||
Summary
Over the past decades, the rational design of synthetic APs has undergone a remarkable transformation—from empirical formulations to precisely engineered molecular platforms that mimic HDPs as alternatives to drug-resistant microorganisms. Despite their potent antibacterial efficacy, the excessive cationic charge density of both HDPs and APs often promotes nonspecific electrostatic interactions with mammalian cell membranes, leading to toxicity such as hemolysis. Thanks to recent advances in polymer science and controlled polymerization techniques, it has become possible to precisely tailor morphology, structure, and the incorporation of stimuli-responsive properties, providing deeper insight into how these parameters collectively influence anti-infective efficacy and biocompatibility. With these advancements, the molecular landscape of biocompatible APs has become increasingly well defined, reflecting a more comprehensive understanding of SAR.Overall, achieving optimal selectivity relies on fine-tuning several molecular design principles. These include the careful choice and arrangement of monomeric units, appropriate hydrophobic–hydrophilic balance, control over chain length and copolymer composition, adjustment of molecular weight, and modulation of stereochemical configuration. Incorporation of neutral hydrophilic moieties is equally important to minimize nonspecific interactions with mammalian cells while preserving antibacterial activity. Furthermore, moving beyond conventional linear random copolymers to more defined architectures, such as block or alternating sequences, has shown to markedly enhance bioactivity. Recent studies also highlight that manipulating polymer topology toward advanced structures such as star, hyperbranched, or cyclic architectures can further improve antibacterial efficacy and biocompatibility. In addition, the development of stimuli-responsive or degradable APs that break down into low MW fragments represents a promising strategy to prevent long-term accumulation in biological systems and mitigate systemic toxicity. Importantly, emerging non-amphiphilic polymer systems offer an alternative design paradigm, expanding beyond traditional amphiphilicity-driven mechanisms and providing new opportunities to achieve selective antibacterial activity.
Challenges, and future perspectives
Challenges
Despite major advances in developing synthetic HDP-mimicking APs, their clinical translation remains limited. While many APs show excellent in vitro performance, the “real deal” lies in demonstrating activity in vivo, where biological complexity presents greater challenges. Bridging the gap between laboratory studies and clinical application requires detailed investigations of pharmacokinetics, long-term exposure, and toxicity. In particular, systematic in vivo toxicity studies, which are largely absent from most recent AP research, are essential for ensuring safety and advancing APs toward clinical use.Beyond biological evaluation, several structural, mechanistic, and regulatory issues continue to constrain the clinical development of APs. Their polydisperse and nonpeptidic nature, combined with broad antibacterial mechanisms, makes the development of bacterial resistance relatively unlikely, which is a major advantage over conventional antibiotics. However, this same structural heterogeneity complicates quality control and regulatory approval. The absence of molecular uniformity challenges the establishment of reproducible structure–activity relationships and makes it difficult to predict biological outcomes with precision.
To prepare for reaching these goals, several key future perspective directions can guide the next phase of AP research. These include biodegradable AP platforms, responsive AP systems, synergistic and combination therapies, and data-driven and artificial intelligence (AI)-assisted design.
Future perspective
Alongside these synthetic approaches, biodegradable APs can also be synthesized from biopolymer-derived backbones such as polysaccharides304 and polypeptides,305 or from nature-inspired frameworks such as poly(lipoic acid), all of which offer inherent biodegradability and can be chemically modified to introduce antimicrobial functionalities. For example, lipoic-acid–based polymers possess dynamic disulfide-rich backbones that can degrade or depolymerize under biologically relevant reducing environments, providing versatile platforms for designing anti-infective materials.67,73
This direction is particularly relevant for the future design of APs intended to help address AMR, as well as those used for repeated dosing, topical application, or implantation, where prolonged polymer retention may pose safety risks. Developing materials with predictable degradation profiles, non-toxic degradation products, and sustainable raw-material origins will support biosafety, facilitate regulatory acceptance, and promote environmentally responsible antimicrobial materials.
In these smart systems, antimicrobial agents are encapsulated within a protective matrix and are released only in the presence of pathogenic bacteria. Infection sites exhibit biochemical features that differ markedly from healthy tissue, including acidification caused by bacterial metabolites (e.g., lactic and acetic acids)307 and the secretion of extracellular enzymes. For example, S. aureus produces several proteolytic enzymes, including two cysteine proteinases, metalloproteinase, and serine glutamyl endopeptidase,308 along with lower levels of hyaluronidase.309 These infection-specific factors can be harnessed to trigger specific and localized antimicrobial release while minimizing exposure to healthy cells, thereby improving selectivity and providing promising directions for future smart AP development.
In conclusion, this review has summarized a comprehensive overview of the development and underlying mechanisms of antibiotic resistance, as well as the evolution of antibacterial polymers from HDP-inspired structural design toward precise biofunction control. Key advances in optimizing structural determinants for antibacterial activity and biocompatibility, encompassing both compositional and topological features, have been discussed. Emerging strategies based on stimuli-responsive and self-immolative polymers that enable on-demand degradation to minimize long-term cytotoxicity are highlighted, along with recent progress in non-amphiphilic polymer systems exhibiting distinct antibacterial mechanisms. Furthermore, future design directions involving synergistic and combination therapies, as well as data-driven and AI-assisted approaches for pathogen-specific polymer design, hold strong potential to address the remaining challenges of toxicity while maintaining or even enhancing antimicrobial potency, thereby improving overall selectivity.
Author contributions
Md Aquib: conceptualization, investigation, writing – original draft, review & editing. Vinod Kumar Kannaujiya: writing–review & editing. Cyrille Boyer: conceptualization, resources, project administration, writing–review and editing, and supervision.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
Md Aquib would like to acknowledge the University International Postgraduate Award (UIPA) from the University of New South Wales (UNSW) for its support. Cyrille Boyer is the recipient of an Australian Research Council-Australian Laureate Fellowship (project number FL220100016) funded by the Australian Government.References
- M. Cully, Public health: The politics of antibiotics, Nature, 2014, 509(7498), S16–S17 CrossRef CAS PubMed.
- C. Nathan, Resisting antimicrobial resistance, Nat. Rev. Microbiol., 2020, 18(5), 259–260 CrossRef CAS PubMed.
- N. van Rhijn, S. Arikan-Akdagli, J. Beardsley, F. Bongomin, A. Chakrabarti, S. C. Chen, T. Chiller, A. L. Colombo, N. P. Govender and A. Alastruey-Izquierdo, Beyond bacteria: the growing threat of antifungal resistance, The Lancet, 2024, 404(10457), 1017–1018 CrossRef CAS PubMed.
- R. Laxminarayan, A. Duse, C. Wattal, A. K. Zaidi, H. F. Wertheim, N. Sumpradit, E. Vlieghe, G. L. Hara, I. M. Gould and H. Goossens, Antibiotic resistance—the need for global solutions, Lancet Infect. Dis., 2013, 13(12), 1057–1098 CrossRef PubMed.
- G. Muteeb, M. T. Rehman, M. Shahwan and M. Aatif, Origin of antibiotics and antibiotic resistance, and their impacts on drug development: A narrative review, Pharmaceuticals, 2023, 16(11), 1615 CrossRef CAS PubMed.
- R. Wise, Antimicrobial resistance: priorities for action, J. Antimicrob. Chemother., 2002, 49(4), 585–586 CrossRef CAS PubMed.
- Y. Hsia, M. Sharland, C. Jackson, I. C. Wong, N. Magrini and J. A. Bielicki, Consumption of oral antibiotic formulations for young children according to the WHO Access, Watch, Reserve (AWaRe) antibiotic groups: an analysis of sales data from 70 middle-income and high-income countries, Lancet Infect. Dis., 2019, 19(1), 67–75 CrossRef PubMed.
- F. O. Gudda, M. G. Waigi, E. S. Odinga, B. Yang, L. Carter and Y. Gao, Antibiotic-contaminated wastewater irrigated vegetables pose resistance selection risks to the gut microbiome, Environ. Pollut., 2020, 264, 114752 CrossRef CAS PubMed.
- M.-C. Danner, A. Robertson, V. Behrends and J. Reiss, Antibiotic pollution in surface fresh waters: Occurrence and effects, Sci. Total Environ., 2019, 664, 793–804 CrossRef CAS.
- M. Naghavi, S. E. Vollset, K. S. Ikuta, L. R. Swetschinski, A. P. Gray, E. E. Wool, G. R. Aguilar, T. Mestrovic, G. Smith and C. Han, Global burden of bacterial antimicrobial resistance 1990–2021: a systematic analysis with forecasts to 2050, The Lancet, 2024, 404(10459), 1199–1226 CrossRef.
- Antimicrobial resistance. 2024. https://www.who.int/europe/news-room/fact-sheets/item/antimicrobial-resistance (accessed September 19, 2024).
- Organization, W. H. 2021 antibacterial agents in clinical and preclinical development: an overview and analysis. 2022.
- M. S. Butler, V. Gigante, H. Sati, S. Paulin, L. Al-Sulaiman, J. H. Rex, P. Fernandes, C. A. Arias, M. Paul and G. E. Thwaites, Analysis of the clinical pipeline of treatments for drug-resistant bacterial infections: despite progress, more action is needed, Antimicrob. Agents Chemother., 2022, 66(3), e01991–01921 CrossRef CAS.
- S. Di Bella, D. R. Giacobbe, A. E. Maraolo, V. Viaggi, R. Luzzati, M. Bassetti, F. Luzzaro and L. Principe, Resistance to ceftazidime/avibactam in infections and colonisations by KPC-producing Enterobacterales: a systematic review of observational clinical studies, J. Global Antimicrob. Resist., 2021, 25, 268–281 CrossRef CAS PubMed.
- S. J. Lam, N. M. O'Brien-Simpson, N. Pantarat, A. Sulistio, E. H. Wong, Y.-Y. Chen, J. C. Lenzo, J. A. Holden, A. Blencowe and E. C. Reynolds, Combating multidrug-resistant Gram-negative bacteria with structurally nanoengineered antimicrobial peptide polymers, Nat. Microbiol., 2016, 1(11), 1–11 Search PubMed.
- A. Pop-Vicas, J. Strom, K. Stanley and E. M. D’Agata, Multidrug-resistant Gram-negative bacteria among patients who require chronic hemodialysis, Clin. J. Am. Soc. Nephrol., 2008, 3(3), 752–758 CrossRef CAS PubMed.
- O. L. Davies and S. Bennett WHO publishes list of bacteria for which new antibiotics are urgently needed. WHO Newsletters 2017.
- A. H. Delcour, Outer membrane permeability and antibiotic resistance, Biochim. Biophys. Acta, Proteins Proteomics, 2009, 1794(5), 808–816 CrossRef CAS PubMed.
- J. M. Blair, M. A. Webber, A. J. Baylay, D. O. Ogbolu and L. J. Piddock, Molecular mechanisms of antibiotic resistance, Nat. Rev. Microbiol., 2015, 13(1), 42–51 CrossRef CAS PubMed.
- S. I. Miller, Antibiotic resistance and regulation of the Gram-negative bacterial outer membrane barrier by host innate immune molecules, mBio, 2016, 7(5), 01541 CrossRef PubMed.
- C. De La Fuente-Nunez, A. Cesaro and R. E. Hancock, Antibiotic failure: Beyond antimicrobial resistance, Drug Resist. Updates, 2023, 71, 101012 CrossRef CAS PubMed.
- V. Kundi, Y. Jin, A. Chandrasekaran, A. Muralidhar, H. Masood, C. Boyer and P. Kumar, Machine Learning Prediction of Antibacterial Activity of Block Copolymers, ACS Appl. Nano Mater., 2024, 7(8), 8939–8948 CrossRef CAS.
- Z. Shao, H. Luo, T. H. Q. Nguyen and E. H. Wong, Effects of Secondary Amine and Molecular Weight on the Biological Activities of Cationic Amphipathic Antimicrobial Macromolecules, Biomacromolecules, 2024, 25(10), 6899–6912 CrossRef CAS.
- P. Pham, S. Oliver and C. Boyer, Design of antimicrobial polymers, Macromol. Chem. Phys., 2023, 224(3), 2200226 CrossRef CAS.
- N. Mookherjee, M. A. Anderson, H. P. Haagsman and D. J. Davidson, Antimicrobial host defence peptides: functions and clinical potential, Nat. Rev. Drug Discovery, 2020, 19(5), 311–332 CrossRef CAS PubMed.
- R. E. Hancock and H.-G. Sahl, Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies, Nat. Biotechnol., 2006, 24(12), 1551–1557 CrossRef CAS PubMed.
- P. Pham, S. Oliver, E. H. Wong and C. Boyer, Effect of hydrophilic groups on the bioactivity of antimicrobial polymers, Polym. Chem., 2021, 12(39), 5689–5703 RSC.
- E. H.-L. Chen, C.-H. Wang, Y.-T. Liao, F.-Y. Chan, Y. Kanaoka, T. Uchihashi, K. Kato, L. Lai, Y.-W. Chang and M.-C. Ho, Visualizing the membrane disruption action of antimicrobial peptides by cryo-electron tomography, Nat. Commun., 2023, 14(1), 5464 CrossRef CAS.
- J. Tan, J. Tay, J. Hedrick and Y. Y. Yang, Synthetic macromolecules as therapeutics that overcome resistance in cancer and microbial infection, Biomaterials, 2020, 252, 120078 CrossRef CAS PubMed.
- M. Zasloff, Antimicrobial peptides of multicellular organisms, Nature, 2002, 415(6870), 389–395 CrossRef CAS PubMed.
- H. Takahashi, G. A. Caputo and K. Kuroda, Amphiphilic polymer therapeutics: An alternative platform in the fight against antibiotic resistant bacteria, Biomater. Sci., 2021, 9(8), 2758–2767 RSC.
- A. G. Elliott, J. X. Huang, S. Neve, J. Zuegg, I. A. Edwards, A. K. Cain, C. J. Boinett, L. Barquist, C. V. Lundberg and J. Steen, An amphipathic peptide with antibiotic activity against multidrug-resistant Gram-negative bacteria, Nat. Commun., 2020, 11(1), 3184 CrossRef CAS PubMed.
- B. L. Bray, Large-scale manufacture of peptide therapeutics by chemical synthesis, Nat. Rev. Drug Discovery, 2003, 2(7), 587–593 CrossRef CAS PubMed.
- M. Magana, M. Pushpanathan, A. L. Santos, L. Leanse, M. Fernandez, A. Ioannidis, M. A. Giulianotti, Y. Apidianakis, S. Bradfute and A. L. Ferguson, The value of antimicrobial peptides in the age of resistance, Lancet Infect. Dis., 2020, 20(9), e216–e230 CrossRef CAS PubMed.
- K. Kuroda, G. A. Caputo and W. F. DeGrado, The role of hydrophobicity in the antimicrobial and hemolytic activities of polymethacrylate derivatives, Chem. – Eur. J., 2009, 15(5), 1123–1133 CrossRef CAS.
- Z. Shao, K. Hakobyan, J. Xu, R. Chen, N. Kumar, M. Willcox and E. H. Wong, Photoinduced Unveiling of Cationic Amine: Toward Smart Photoresponsive Antimicrobial Polymers, ACS Appl. Polym. Mater., 2023, 5(10), 8735–8743 CrossRef CAS.
- P. Askari, M. H. Namaei, K. Ghazvini and M. Hosseini, In vitro and in vivo toxicity and antibacterial efficacy of melittin against clinical extensively drug-resistant bacteria, BMC Pharmacol. Toxicol., 2021, 22(1), 42 CrossRef CAS PubMed.
- J. M. Kurzyna, R. J. Kopiasz, M. Paul, M. Flont, P. Baranowska, J. Mierzejewska, K. Drężek, W. Tomaszewski, E. Jastrzębska and D. Jańczewski, Unlocking the Potential: PEGylation and Molecular Weight Reduction of Ionenes for Enhanced Antifungal Activity and Biocompatibility, Macromol. Biosci., 2024, 24(9), 2400032 CrossRef CAS PubMed.
- Z. Deng, R. Zhang, J. Gong, Z. Zhang, L. Zhang, Z. Qiu, P. Alam, J. Zhang, Y. Liu and Y. Li, Unveiling the role of alkyl chain in boosting antibacterial selectivity and cell biocompatibility, JACS Au, 2025, 5(2), 675–683 CrossRef CAS PubMed.
- Z. Cheng and P. Raffa, From design to application: amphiphilic copolymers as antimicrobial materials, Mater. Adv., 2025, 6(15), 4939–4968 RSC.
- S. Kanwal, U. B. A. Aziz, E. Quaas, K. Achazi and D. Klinger, Sulfonium-based polymethacrylamides for antimicrobial use: influence of the structure and composition, Biomater. Sci., 2025, 13(4), 993–1009 RSC.
- Y. Geng, Y. Yuan, Y. Bao, S. Huang, X. Wang, L. Huang, C. She, X. Gong and M. Xiong, pH window for high selectivity of ionizable antimicrobial polymers toward bacteria, ACS Appl. Mater. Interfaces, 2023, 15(18), 21781–21791 CrossRef CAS.
- J. Tan, Y. Zhao, J. L. Hedrick and Y. Y. Yang, Effects of hydrophobicity on antimicrobial activity, selectivity, and functional mechanism of guanidinium-functionalized polymers, Adv. Healthcare Mater., 2022, 11(7), 2100482 CrossRef CAS PubMed.
- S. J. Richards, A. Jones, R. M. Tomás and M. I. Gibson, Photochemical “In-Air” Combinatorial Discovery of Antimicrobial Co-polymers, Chem. – Eur. J., 2018, 24(52), 13758–13761 CrossRef CAS PubMed.
- C. Stubbs, T. Congdon, J. Davis, D. Lester, S.-J. Richards and M. I. Gibson, High-throughput tertiary amine deoxygenated photopolymerizations for synthesizing polymer libraries, Macromolecules, 2019, 52(20), 7603–7612 CrossRef CAS PubMed.
- A. Kuroki, P. Sangwan, Y. Qu, R. Peltier, C. Sanchez-Cano, J. Moat, C. G. Dowson, E. G. Williams, K. E. Locock and M. Hartlieb, Sequence control as a powerful tool for improving the selectivity of antimicrobial polymers, ACS Appl. Mater. Interfaces, 2017, 9(46), 40117–40126 CrossRef CAS PubMed.
- P. Gurnani, T. Floyd, J. Tanaka, C. Stubbs, D. Lester, C. Sanchez-Cano and S. Perrier, PCR-RAFT: rapid high throughput oxygen tolerant RAFT polymer synthesis in a biology laboratory, Polym. Chem., 2020, 11(6), 1230–1236 RSC.
- C. Yang, K. B. Wu, Y. Deng, J. Yuan and J. Niu, Geared toward applications: A perspective on functional sequence-controlled polymers, ACS Macro Lett., 2021, 10(2), 243–257 CrossRef CAS PubMed.
- P. R. Judzewitsch, L. Zhao, E. H. Wong and C. Boyer, High-throughput synthesis of antimicrobial copolymers and rapid evaluation of their bioactivity, Macromolecules, 2019, 52(11), 3975–3986 CrossRef CAS.
- C. Boyer, V. Bulmus, T. P. Davis, V. Ladmiral, J. Liu and S. Perrier, Bioapplications of RAFT polymerization, Chem. Rev., 2009, 109(11), 5402–5436 CrossRef CAS PubMed.
- V. K. Kannaujiya, T. Zhang, M. Aquib and C. Boyer, Oxygen-tolerant photo-RAFT enables in-situ synthesis of protein-based nanoparticles, Eur. Polym. J., 2024, 221, 113518 CrossRef.
- A. Jain, L. S. Duvvuri, S. Farah, N. Beyth, A. J. Domb and W. Khan, Antimicrobial polymers, Adv. Healthcare Mater., 2014, 3(12), 1969–1985 CrossRef CAS PubMed.
- E. F. Palermo and K. Kuroda, Chemical structure of cationic groups in amphiphilic polymethacrylates modulates the antimicrobial and hemolytic activities, Biomacromolecules, 2009, 10(6), 1416–1428 CrossRef CAS PubMed.
- A. Kuroki, A. K. Tchoupa, M. Hartlieb, R. Peltier, K. E. Locock, M. Unnikrishnan and S. Perrier, Targeting intracellular, multi-drug resistant Staphylococcus aureus with guanidinium polymers by elucidating the structure-activity relationship, Biomaterials, 2019, 217, 119249 CrossRef CAS PubMed.
- P. R. Judzewitsch, T. K. Nguyen, S. Shanmugam, E. H. Wong and C. Boyer, Towards sequence-controlled antimicrobial polymers: effect of polymer block order on antimicrobial activity, Angew. Chem., Int. Ed., 2018, 57(17), 4559–4564 CrossRef CAS PubMed.
- S. Laroque, R. n Garcia Maset, A. Hapeshi, F. Burgevin, K. E. Locock and S. Perrier, Synthetic star nanoengineered antimicrobial polymers as antibiofilm agents: Bacterial membrane disruption and cell aggregation, Biomacromolecules, 2023, 24(7), 3073–3085 CrossRef CAS PubMed.
- S. J. Shirbin, I. Insua, J. A. Holden, J. C. Lenzo, E. C. Reynolds, N. M. O'Brien-Simpson and G. G. Qiao, Architectural effects of star-shaped “structurally nanoengineered antimicrobial peptide polymers”(SNAPPs) on their biological activity, Adv. Healthcare Mater., 2018, 7(21), 1800627 CrossRef PubMed.
- H. Gmedhin, M. Aquib, N. Corrigan, M. D. Lenardon and C. Boyer, Photoactivated antifungal polymers prepared by PET-RAFT polymerization, Chem. Sci., 2026, 17, 3178–3188 RSC.
- R. Namivandi-Zangeneh, R. J. Kwan, T.-K. Nguyen, J. Yeow, F. L. Byrne, S. H. Oehlers, E. H. Wong and C. Boyer, The effects of polymer topology and chain length on the antimicrobial activity and hemocompatibility of amphiphilic ternary copolymers, Polym. Chem., 2018, 9(13), 1735–1744 RSC.
- J. Xu, L. Pu, J. Ma, S. K. Kumar and H. Duan, Antibacterial properties of synthesized cyclic and linear cationic copolymers, Polym. Chem., 2020, 11(41), 6632–6639 RSC.
- M. Porel, D. N. Thornlow, N. N. Phan and C. A. Alabi, Sequence-defined bioactive macrocycles via an acid-catalysed cascade reaction, Nat. Chem., 2016, 8(6), 590–596 CrossRef CAS PubMed.
- S. Laroque, M. Reifarth, M. Sperling, S. Kersting, S. Klöpzig, P. Budach, J. Storsberg and M. Hartlieb, Impact of multivalence and self-assembly in the design of polymeric antimicrobial peptide mimics, ACS Appl. Mater. Interfaces, 2020, 12(27), 30052–30065 CrossRef CAS PubMed.
- A. C. Lehnen, S. Kogikoski Jr, T. Stensitzki, A. AlSawaf, A. M. Bapolisi, M. Wolff, J. De Breuck, H. M. Müller-Werkmeister, S. Chiantia and I. Bald, Anisotropy in Antimicrobial Bottle Brush Copolymers and Its Influence on Biological Activity, Adv. Funct. Mater., 2024, 34(10), 2312651 CrossRef CAS.
- T.-K. Nguyen, S. J. Lam, K. K. Ho, N. Kumar, G. G. Qiao, S. Egan, C. Boyer and E. H. Wong, Rational design of single-chain polymeric nanoparticles that kill planktonic and biofilm bacteria, ACS Infect. Dis., 2017, 3(3), 237–248 CrossRef CAS PubMed.
- P. Pham, S. Oliver, D. T. Nguyen and C. Boyer, Effect of cationic groups on the selectivity of ternary antimicrobial polymers, Macromol. Rapid Commun., 2022, 43(21), 2200377 CrossRef CAS PubMed.
- C. Y. Ang, S. Y. Tan, C. Teh, J. M. Lee, M. F. E. Wong, Q. Qu, L. Q. Poh, M. Li, Y. Zhang and V. Korzh, Redox and pH dual responsive polymer based nanoparticles for in vivo drug delivery, Small, 2017, 13(7), 1602379 CrossRef PubMed.
- A. Giri, M. Aquib, A. Choudhury, V. K. Kannaujiya, J. L. Lim, Z. Gu, M. D. Lenardon and C. Boyer, Lipoic Acid Based Redox-Responsive Degradable Antimicrobial Polymers, Macromol. Rapid Commun., 2025, e00224 CrossRef CAS PubMed.
- R. Gadaga, Z. Varanaraja, J. Lefley, T. Rheinberger, F. R. Wurm and C. R. Becer, Degradable Polymer Architectures Based on Polyphosphonate, Poly (2-oxazoline), and Poly (2-oxazine), Macromolecules, 2024, 57(20), 9853–9866 CrossRef CAS.
- D. S. Lim, Y. Yuan and Y. Zhang, pH-degradable polymers as impermanent antimicrobial agents for environmental sustainability, ACS Appl. Bio Mater., 2021, 4(2), 1544–1551 CrossRef CAS PubMed.
- X. Li, S. Ilk, J. A. Linares-Pastén, Y. Liu, D. B. Raina, D. Demircan and B. Zhang, Synthesis, enzymatic degradation, and polymer-miscibility evaluation of nonionic antimicrobial hyperbranched polyesters with indole or isatin functionalities, Biomacromolecules, 2021, 22(5), 2256–2271 CrossRef CAS PubMed.
- J. Liu, H. Duong, M. R. Whittaker, T. P. Davis and C. Boyer, Synthesis of functional core, star polymers via RAFT polymerization for drug delivery applications, Macromol. Rapid Commun., 2012, 33(9), 760–766 CrossRef CAS PubMed.
- J. A. Syrett, D. M. Haddleton, M. R. Whittaker, T. P. Davis and C. Boyer, Functional, star polymeric molecular carriers, built from biodegradable microgel/nanogel cores, Chem. Commun., 2011, 47(5), 1449–1451 RSC.
- J. Guo, S. Zhang, Y. Tao, W. Zheng, H. Cheng, H. Li, Z. Wang, Y. Gou, J. Zhu and L. Li, Synthesis of Cationic Cyclic Oligo (disulfide) s via Cyclo-Depolymerization: A Redox-Responsive and Potent Antibacterial Reagent, J. Am. Chem. Soc., 2025, 147(8), 6772–6785 CrossRef CAS PubMed.
- Z. Cheng and P. Raffa, From design to application: amphiphilic copolymers as antimicrobial materials, Mater. Adv., 2025, 6(15), 4939–4968 RSC.
- Z. Si and M. B. Chan-Park, Chemical Innovations of Antimicrobial Polymers for Combating Antimicrobial Resistance, ACS Biomater. Sci. Eng., 2025, 11(5), 2470–2480 CrossRef CAS PubMed.
- Z. Si and M. B. Chan-Park, Recent Advances in Designing Antimicrobial Polymers with Diverse Uptake and Killing Mechanisms, Prog. Polym. Sci., 2026, 102080 CrossRef CAS.
- L. Yu, K. Li, J. Zhang, H. Jin, A. Saleem, Q. Song, Q. Jia and P. Li, Antimicrobial peptides and macromolecules for combating microbial infections: from agents to interfaces, ACS Appl. Bio Mater., 2022, 5(2), 366–393 CrossRef CAS PubMed.
- G. A. Kropp, C. N. McMillian, J. D. Mase, S. Kandagiri, E. S. Nowak and M. D. Schulz, Recent developments in antimicrobial polymers for biofilm inhibition, Chem. Commun., 2026 10.1039/D5CC06617D.
- Z. Si, W. Zheng, D. Prananty, J. Li, C. H. Koh, E.-T. Kang, K. Pethe and M. B. Chan-Park, Polymers as advanced antibacterial and antibiofilm agents for direct and combination therapies, Chem. Sci., 2022, 13(2), 345–364 RSC.
- J. Chen, F. Wang, Q. Liu and J. Du, Antibacterial polymeric nanostructures for biomedical applications, Chem. Commun., 2014, 50(93), 14482–14493 RSC.
- H. Zhao, J. Sun, Y. Cheng, S. Nie and W. Li, Advances in peptide/polymer antimicrobial assemblies, J. Mater. Chem. B, 2025, 13(5), 1518–1530 RSC.
- Y. Jiang, Y. Chen, Z. Song, Z. Tan and J. Cheng, Recent advances in design of antimicrobial peptides and polypeptides toward clinical translation, Adv. Drug Delivery Rev., 2021, 170, 261–280 CrossRef CAS PubMed.
- P. Salas-Ambrosio, A. Tronnet, P. Verhaeghe and C. Bonduelle, Synthetic polypeptide polymers as simplified analogues of antimicrobial peptides, Biomacromolecules, 2020, 22(1), 57–75 CrossRef PubMed.
- S. Paul, S. Verma and Y.-C. Chen, Peptide dendrimer-based antibacterial agents: synthesis and applications, ACS Infect. Dis., 2024, 10(4), 1034–1055 CrossRef CAS PubMed.
- A. Talebi Bezmin Abadi, A. A. Rizvanov, T. Haertlé and N. L. Blatt, World Health Organization report: current crisis of antibiotic resistance, Bionanoscience, 2019, 9(4), 778–788 CrossRef.
- T. U. Berendonk, C. M. Manaia, C. Merlin, D. Fatta-Kassinos, E. Cytryn, F. Walsh, H. Bürgmann, H. Sørum, M. Norström and M.-N. Pons, Tackling antibiotic resistance: the environmental framework, Nat. Rev. Microbiol., 2015, 13(5), 310–317 CrossRef CAS PubMed.
- S. I. Hay, P. C. Rao, C. Dolecek, N. P. Day, A. Stergachis, A. D. Lopez and C. J. Murray, Measuring and mapping the global burden of antimicrobial resistance, BMC Med., 2018, 16(1), 78 CrossRef PubMed.
- R. Namivandi-Zangeneh, E. H. Wong and C. Boyer, Synthetic antimicrobial polymers in combination therapy: tackling antibiotic resistance, ACS Infect. Dis., 2021, 7(2), 215–253 CrossRef CAS PubMed.
- (AMR), A. R. D. WHO Strategic Priorities on Antimicrobial Resistance: Preserving antimicrobials for today and tomorrow. 18 May 2022. https://www.who.int/publications/i/item/9789240041387 (accessed 12 October 2025).
- J. A. Ayukekbong, M. Ntemgwa and A. N. Atabe, The threat of antimicrobial resistance in developing countries: causes and control strategies, Antimicrob. Resist. Infect. Control, 2017, 6(1), 47 CrossRef PubMed.
- P. I. Rafailidis and D. Kofteridis, Proposed amendments regarding the definitions of multidrug-resistant and extensively drug-resistant bacteria, Expert Rev. Anti-Infect. Ther., 2022, 20(2), 139–146 CrossRef CAS PubMed.
- A.-P. Magiorakos, A. Srinivasan, R. B. Carey, Y. Carmeli, M. Falagas, C. Giske, S. Harbarth, J. Hindler, G. Kahlmeter and B. Olsson-Liljequist, Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance, Clin. Microbiol. Infect., 2012, 18(3), 268–281 CrossRef CAS PubMed.
- T. Li, Z. Wang, J. Guo, C. de la Fuente-Nunez, J. Wang, B. Han, H. Tao, J. Liu and X. Wang, Bacterial resistance to antibacterial agents: Mechanisms, control strategies, and implications for global health, Sci. Total Environ., 2023, 860, 160461 CrossRef CAS PubMed.
- A. Coates, Y. Hu, R. Bax and C. Page, The future challenges facing the development of new antimicrobial drugs, Nat. Rev. Drug Discovery, 2002, 1(11), 895–910 CrossRef CAS PubMed.
- H. Lee, K. W. Yun, H. J. Lee and E. H. Choi, Antimicrobial therapy of macrolide-resistant Mycoplasma pneumoniae pneumonia in children, Expert Rev. Anti-Infect. Ther., 2018, 16(1), 23–34 CrossRef CAS PubMed.
- A. M. Sefton, Mechanisms of antimicrobial resistance: their clinical relevance in the new millennium, Drugs, 2002, 62(4), 557–566 CrossRef CAS PubMed.
- S. Sandoval-Motta and M. Aldana, Adaptive resistance to antibiotics in bacteria: a systems biology perspective, Wiley Interdiscip. Rev.:Syst. Biol. Med., 2016, 8(3), 253–267 Search PubMed.
- J. Xuan, W. Feng, J. Wang, R. Wang, B. Zhang, L. Bo, Z.-S. Chen, H. Yang and L. Sun, Antimicrobial peptides for combating drug-resistant bacterial infections, Drug Resist. Updates, 2023, 68, 100954 CrossRef CAS PubMed.
- M. O. Sommer, C. Munck, R. V. Toft-Kehler and D. I. Andersson, Prediction of antibiotic resistance: time for a new preclinical paradigm?, Nat. Rev. Microbiol., 2017, 15(11), 689–696 CrossRef CAS PubMed.
- D. Mehta, V. Saini, B. Aggarwal, A. Khan and A. Bajaj, Unlocking the bacterial membrane as a therapeutic target for next-generation antimicrobial amphiphiles, Mol. Aspects Med., 2021, 81, 100999 CrossRef CAS PubMed.
- C. Walsh, Molecular mechanisms that confer antibacterial drug resistance, Nature, 2000, 406(6797), 775–781 CrossRef CAS PubMed.
- C. González-Bello, Antibiotic adjuvants–A strategy to unlock bacterial resistance to antibiotics, Bioorg. Med. Chem. Lett., 2017, 27(18), 4221–4228 CrossRef PubMed.
- A. Prinzi and R. Rohde The role of bacterial biofilms in antimicrobial resistance. American Society for Microbiology, 2023.
- S. Miquel, R. Lagrafeuille, B. Souweine and C. Forestier, Anti-biofilm activity as a health issue, Front. Microbiol., 2016, 7, 592 Search PubMed.
- M. Jamal, W. Ahmad, S. Andleeb, F. Jalil, M. Imran, M. A. Nawaz, T. Hussain, M. Ali, M. Rafiq and M. A. Kamil, Bacterial biofilm and associated infections, J. Chin. Med. Assoc., 2018, 81(1), 7–11 CrossRef PubMed.
- H. Steiner, D. Hultmark, Å. Engström, H. Bennich and H. G. Boman, Sequence and specificity of two antibacterial proteins involved in insect immunity, Nature, 1981, 292(5820), 246–248 CrossRef CAS PubMed.
- H. Takahashi, G. A. Caputo, S. Vemparala and K. Kuroda, Synthetic random copolymers as a molecular platform to mimic host-defense antimicrobial peptides, Bioconjugate Chem., 2017, 28(5), 1340–1350 CrossRef CAS PubMed.
- E. F. Palermo and K. Kuroda, Structural determinants of antimicrobial activity in polymers which mimic host defense peptides, Appl. Microbiol. Biotechnol., 2010, 87(5), 1605–1615 CrossRef CAS PubMed.
- P. T. Phuong, S. Oliver, J. He, E. H. Wong, R. T. Mathers and C. Boyer, Effect of hydrophobic groups on antimicrobial and hemolytic activity: Developing a predictive tool for ternary antimicrobial polymers, Biomacromolecules, 2020, 21(12), 5241–5255 CrossRef CAS PubMed.
- A. C. Engler, N. Wiradharma, Z. Y. Ong, D. J. Coady, J. L. Hedrick and Y.-Y. Yang, Emerging trends in macromolecular antimicrobials to fight multi-drug-resistant infections, Nano Today, 2012, 7(3), 201–222 CrossRef CAS.
- A. Som, S. Vemparala, I. Ivanov and G. N. Tew, Synthetic mimics of antimicrobial peptides, Pept. Sci., 2008, 90(2), 83–93 CrossRef CAS PubMed.
- N. Wiradharma, U. Khoe, C. A. Hauser, S. V. Seow, S. Zhang and Y.-Y. Yang, Synthetic cationic amphiphilic α-helical peptides as antimicrobial agents, Biomaterials, 2011, 32(8), 2204–2212 CrossRef CAS PubMed.
- A. K. Marr, W. J. Gooderham and R. E. Hancock, Antibacterial peptides for therapeutic use: obstacles and realistic outlook, Curr. Opin. Pharmacol., 2006, 6(5), 468–472 CrossRef CAS PubMed.
- Y. Wu, K. Chen, J. Wang, M. Chen, Y. Chen, Y. She, Z. Yan and R. Liu, Host defense peptide mimicking antimicrobial amino acid polymers and beyond: Design, synthesis and biomedical applications, Prog. Polym. Sci., 2023, 141, 101679 CrossRef CAS.
- L. Cresti, G. Cappello and A. Pini, Antimicrobial peptides towards clinical application—A long history to be concluded, Int. J. Mol. Sci., 2024, 25(9), 4870 CrossRef CAS PubMed.
- D. Wei and X. Zhang, Biosynthesis, bioactivity, biotoxicity and applications of antimicrobial peptides for human health, Biosaf. Health, 2022, 4(02), 118–134 CrossRef.
- T. Ikeda, S. Tazuke and Y. Suzuki, Biologically active polycations, 4. Synthesis and antimicrobial activity of poly (trialkylvinylbenzylammonium chloride) s, Macromol. Chem. Phys., 1984, 185(5), 869–876 CrossRef CAS.
- C. Ergene, K. Yasuhara and E. F. Palermo, Biomimetic antimicrobial polymers: Recent advances in molecular design, Polym. Chem., 2018, 9(18), 2407–2427 RSC.
- K. Kuroda and W. F. DeGrado, Amphiphilic polymethacrylate derivatives as antimicrobial agents, J. Am. Chem. Soc., 2005, 127(12), 4128–4129 CrossRef CAS PubMed.
- M. F. Ilker, K. Nüsslein, G. N. Tew and E. B. Coughlin, Tuning the hemolytic and antibacterial activities of amphiphilic polynorbornene derivatives, J. Am. Chem. Soc., 2004, 126(48), 15870–15875 CrossRef CAS PubMed.
- B. P. Mowery, S. E. Lee, D. A. Kissounko, R. F. Epand, R. M. Epand, B. Weisblum, S. S. Stahl and S. H. Gellman, Mimicry of antimicrobial host-defense peptides by random copolymers, J. Am. Chem. Soc., 2007, 129(50), 15474–15476 CrossRef CAS PubMed.
- R. n Garcia Maset, L. Pasquina-Lemonche, A. Hapeshi, L. A. Clifton, J. K. Hobbs, F. Harrison, S. Perrier and S. C. Hall, Assessing the mechanism of action of synthetic nanoengineered antimicrobial polymers against the bacterial membrane of Pseudomonas aeruginosa, Biomacromolecules, 2025, 26(10), 6854–6868 CrossRef CAS PubMed.
- J. Phommalysack-Lovan, Y. Chu, C. Boyer and J. Xu, PET-RAFT polymerisation: towards green and precision polymer manufacturing, Chem. Commun., 2018, 54(50), 6591–6606 RSC.
- B. Alberts; D. Bray; K. Hopkin; A. D. Johnson; J. Lewis; M. Raff; K. Roberts and P. Walter, Essential cell biology, Garland Science, 2015 Search PubMed.
- A. Cassone, D. Kerridge and E. F. Gale, Ultrastructural changes in the cell wall of Candida albicans following cessation of growth and their possible relationship to the development of polyene resistance, Microbiology, 1979, 110(2), 339–349 CrossRef CAS PubMed.
- J. Ruiz-Herrera, M. Victoria Elorza, E. Valentín and R. Sentandreu, Molecular organization of the cell wall of Candida albicans and its relation to pathogenicity, FEMS Yeast Res., 2006, 6(1), 14–29 CrossRef CAS PubMed.
- L.-L. Li, H.-W. An, B. Peng, R. Zheng and H. Wang, Self-assembled nanomaterials: design principles, the nanostructural effect, and their functional mechanisms as antimicrobial or detection agents, Mater. Horiz., 2019, 6(9), 1794–1811 RSC.
- M. S. Ganewatta and C. Tang, Controlling macromolecular structures towards effective antimicrobial polymers, Polymer, 2015, 63, A1–A29 CrossRef CAS.
- M. Rautenbach, A. M. Troskie and J. A. Vosloo, Antifungal peptides: To be or not to be membrane active, Biochimie, 2016, 130, 132–145 CrossRef CAS PubMed.
- S. Schaefer, T. T. P. Pham, S. Brunke, B. Hube, K. Jung, M. D. Lenardon and C. Boyer, Rational design of an antifungal polyacrylamide library with reduced host-cell toxicity, ACS Appl. Mater. Interfaces, 2021, 13(23), 27430–27444 CrossRef CAS PubMed.
- T. Salditt, C. Li and A. Spaar, Structure of antimicrobial peptides and lipid membranes probed by interface-sensitive X-ray scattering, Biochim. Biophys. Acta, Biomembr., 2006, 1758(9), 1483–1498 CrossRef CAS PubMed.
- L. T. Nguyen, E. F. Haney and H. J. Vogel, The expanding scope of antimicrobial peptide structures and their modes of action, Trends Biotechnol., 2011, 29(9), 464–472 CrossRef CAS PubMed.
- G. Ehrenstein and H. Lecar, Electrically gated ionic channels in lipid bilayers, Q. Rev. Biophys., 1977, 10(1), 1–34 CrossRef CAS PubMed.
- Y. Pouny, D. Rapaport, A. Mor, P. Nicolas and Y. Shai, Interaction of antimicrobial dermaseptin and its fluorescently labeled analogs with phospholipid membranes, Biochemistry, 1992, 31(49), 12416–12423 CrossRef CAS PubMed.
- E. Gazit, A. Boman, H. G. Boman and Y. Shai, Interaction of the mammalian antibacterial peptide cecropin P1 with phospholipid vesicles, Biochemistry, 1995, 34(36), 11479–11488 CrossRef CAS PubMed.
- S. J. Ludtke, K. He, W. T. Heller, T. A. Harroun, L. Yang and H. W. Huang, Membrane pores induced by magainin, Biochemistry, 1996, 35(43), 13723–13728 CrossRef CAS PubMed.
- H. Leontiadou, A. E. Mark and S. J. Marrink, Antimicrobial peptides in action, J. Am. Chem. Soc., 2006, 128(37), 12156–12161 CrossRef CAS PubMed.
- D. I. Chan, E. J. Prenner and H. J. Vogel, Tryptophan-and arginine-rich antimicrobial peptides: structures and mechanisms of action, Biochim. Biophys. Acta, Biomembr., 2006, 1758(9), 1184–1202 CrossRef CAS PubMed.
- S. Ludtke, K. He and H. Huang, Membrane thinning caused by magainin 2, Biochemistry, 1995, 34(51), 16764–16769 CrossRef CAS PubMed.
- J. A. Silverman, N. G. Perlmutter and H. M. Shapiro, Correlation of daptomycin bactericidal activity and membrane depolarization in Staphylococcus aureus, Antimicrob. Agents Chemother., 2003, 47(8), 2538–2544 CrossRef CAS PubMed.
- G. Pabst, S. L. Grage, S. Danner-Pongratz, W. Jing, A. S. Ulrich, A. Watts, K. Lohner and A. Hickel, Membrane thickening by the antimicrobial peptide PGLa, Biophys. J., 2008, 95(12), 5779–5788 CrossRef CAS PubMed.
- J. Oreopoulos, R. F. Epand, R. M. Epand and C. M. Yip, Peptide-induced domain formation in supported lipid bilayers: direct evidence by combined atomic force and polarized total internal reflection fluorescence microscopy, Biophys. J., 2010, 98(5), 815–823 CrossRef CAS PubMed.
- C. B. Park, H. S. Kim and S. C. Kim, Mechanism of action of the antimicrobial peptide buforin II: buforin II kills microorganisms by penetrating the cell membrane and inhibiting cellular functions, Biochem. Biophys. Res. Commun., 1998, 244(1), 253–257 CrossRef CAS PubMed.
- A. Patrzykat, C. L. Friedrich, L. Zhang, V. Mendoza and R. E. Hancock, Sublethal concentrations of pleurocidin-derived antimicrobial peptides inhibit macromolecular synthesis in Escherichia coli, Antimicrob. Agents Chemother., 2002, 46(3), 605–614 CrossRef CAS PubMed.
- M. Dangkulwanich, C. Raetz and A. Williams, Structure guided design of an antibacterial peptide that targets UDP-N-acetylglucosamine acyltransferase, Sci. Rep., 2019, 9(1), 3947 CrossRef PubMed.
- S. Somma, L. Gastaldo and A. Corti, Teicoplanin, a new antibiotic from Actinoplanes teichomyceticus nov. sp, Antimicrob. Agents Chemother., 1984, 26(6), 917–923 CrossRef CAS PubMed.
- L. D. Saravolatz, G. E. Stein and L. B. Johnson, Telavancin: a novel lipoglycopeptide, Clin. Infect. Dis., 2009, 49(12), 1908–1914 CrossRef CAS PubMed.
- C. H. Chen and T. K. Lu, Development and challenges of antimicrobial peptides for therapeutic applications, Antibiotics, 2020, 9(1), 24 CrossRef CAS PubMed.
- K. Beck, J. Nandy and M. Hoernke, Strong Membrane Permeabilization Activity Can Reduce Selectivity of Cyclic Antimicrobial Peptides, J. Phys. Chem. B, 2025, 129(9), 2446–2460 CrossRef CAS PubMed.
- S. Khemaissa, A. Walrant and S. Sagan, Tryptophan, more than just an interfacial amino acid in the membrane activity of cationic cell-penetrating and antimicrobial peptides, Q. Rev. Biophys., 2022, 55, e10 CrossRef CAS PubMed.
- J. Swierstra, V. Kapoerchan, A. Knijnenburg, A. van Belkum and M. Overhand, Structure, toxicity and antibiotic activity of gramicidin S and derivatives, Eur. J. Clin. Microbiol. Infect. Dis., 2016, 35(5), 763–769 CrossRef CAS PubMed.
- A. Ordooei Javan, S. Shokouhi and Z. Sahraei, A review on colistin nephrotoxicity, Eur. J. Clin. Pharmacol., 2015, 71(7), 801–810 CrossRef CAS PubMed.
- E. J. Filippone, W. K. Kraft and J. L. Farber, The nephrotoxicity of vancomycin, Clin. Pharm. Ther., 2017, 102(3), 459–469 CrossRef CAS PubMed.
- S. Hong, H. Takahashi, E. T. Nadres, H. Mortazavian, G. A. Caputo, J. G. Younger and K. Kuroda, A cationic amphiphilic random copolymer with ph-responsive activity against methicillin-resistant Staphylococcus aureus, PLoS One, 2017, 12(1), e0169262 CrossRef PubMed.
- W. F. Walkenhorst, Using adjuvants and environmental factors to modulate the activity of antimicrobial peptides, Biochim. Biophys. Acta, Biomembr., 2016, 1858(5), 926–935 CrossRef CAS PubMed.
- J. J. Lopez Cascales, S. Zenak, J. García de La Torre, O. G. Lezama, A. Garro and R. D. Enriz, Small cationic peptides: influence of charge on their antimicrobial activity, ACS Omega, 2018, 3(5), 5390–5398 CrossRef CAS PubMed.
- D. Pranantyo, L. Q. Xu, Z. Hou, E.-T. Kang and M. B. Chan-Park, Increasing bacterial affinity and cytocompatibility with four-arm star glycopolymers and antimicrobial α-polylysine, Polym. Chem., 2017, 8(21), 3364–3373 RSC.
- X. Jiang, S. Zhang, M. A. Azad, K. D. Roberts, L. Wan, B. Gong, K. Yang, B. Yuan, H. Uddin and J. Li, Structure–interaction relationship of polymyxins with the membrane of human kidney proximal tubular cells, ACS Infect. Dis., 2020, 6(8), 2110–2119 CrossRef CAS PubMed.
- K. Matsuzaki, S. Yoneyama and K. Miyajima, Pore formation and translocation of melittin, Biophys. J., 1997, 73(2), 831–838 CrossRef CAS PubMed.
- K. Kuroda and G. A. Caputo, Antimicrobial polymers as synthetic mimics of host-defense peptides, Wiley Interdiscip. Rev.:Nanomed. Nanobiotechnol., 2013, 5(1), 49–66 CAS.
- Y. Liang, X. Zhang, Y. Yuan, Y. Bao and M. Xiong, Role and modulation of the secondary structure of antimicrobial peptides to improve selectivity, Biomater. Sci., 2020, 8(24), 6858–6866 RSC.
- C. Ergene and E. F. Palermo, Antimicrobial synthetic polymers: An update on structure-activity relationships, Curr. Pharm. Des., 2018, 24(8), 855–865 CrossRef CAS PubMed.
- K. E. Locock, T. D. Michl, J. D. Valentin, K. Vasilev, J. D. Hayball, Y. Qu, A. Traven, H. J. Griesser, L. Meagher and M. Haeussler, Guanylated polymethacrylates: a class of potent antimicrobial polymers with low hemolytic activity, Biomacromolecules, 2013, 14(11), 4021–4031 CrossRef CAS PubMed.
- E. F. Palermo, I. Sovadinova and K. Kuroda, Structural determinants of antimicrobial activity and biocompatibility in membrane-disrupting methacrylamide random copolymers, Biomacromolecules, 2009, 10(11), 3098–3107 CrossRef CAS PubMed.
- B. P. Mowery, A. H. Lindner, B. Weisblum, S. S. Stahl and S. H. Gellman, Structure− activity relationships among random nylon-3 copolymers that mimic antibacterial host-defense peptides, J. Am. Chem. Soc., 2009, 131(28), 9735–9745 CrossRef CAS PubMed.
- S. Chakraborty, R. Liu, Z. Hayouka, X. Chen, J. Ehrhardt, Q. Lu, E. Burke, Y. Yang, B. Weisblum and G. C. Wong, Ternary nylon-3 copolymers as host-defense peptide mimics: beyond hydrophobic and cationic subunits, J. Am. Chem. Soc., 2014, 136(41), 14530–14535 CrossRef CAS PubMed.
- G. J. Gabriel, J. A. Maegerlein, C. F. Nelson, J. M. Dabkowski, T. Eren, K. Nüsslein and G. N. Tew, Comparison of facially amphiphilic versus segregated monomers in the design of antibacterial copolymers, Chem. – Eur. J., 2009, 15(2), 433–439 CrossRef CAS PubMed.
- K. Lienkamp, A. E. Madkour, A. Musante, C. F. Nelson, K. Nusslein and G. N. Tew, Antimicrobial polymers prepared by ROMP with unprecedented selectivity: a molecular construction kit approach, J. Am. Chem. Soc., 2008, 130(30), 9836–9843 CrossRef CAS PubMed.
- K. Lienkamp, A. E. Madkour, K. N. Kumar, K. Nüsslein and G. N. Tew, Antimicrobial polymers prepared by ring-opening metathesis polymerization: manipulating antimicrobial properties by organic counterion and charge density variation, Chem. – Eur. J., 2009, 15(43), 11715–11722 CrossRef CAS PubMed.
- Y. Qiao, C. Yang, D. J. Coady, Z. Y. Ong, J. L. Hedrick and Y.-Y. Yang, Highly dynamic biodegradable micelles capable of lysing Gram-positive and Gram-negative bacterial membrane, Biomaterials, 2012, 33(4), 1146–1153 CrossRef CAS PubMed.
- M. Concilio, R. Garcia Maset, L. P. Lemonche, V. Kontrimas, J. I. Song, S. K. Rajendrakumar, F. Harrison, C. R. Becer and S. Perrier, Mechanism of action of oxazoline-based antimicrobial polymers against staphylococcus aureus: in vivo antimicrobial activity evaluation, Adv. Healthcare Mater., 2023, 12(29), 2301961 CrossRef CAS PubMed.
- I. Sovadinova, E. F. Palermo, R. Huang, L. M. Thoma and K. Kuroda, Mechanism of polymer-induced hemolysis: nanosized pore formation and osmotic lysis, Biomacromolecules, 2011, 12(1), 260–268 CrossRef CAS PubMed.
- B. Hisey, P. J. Ragogna and E. R. Gillies, Phosphonium-functionalized polymer micelles with intrinsic antibacterial activity, Biomacromolecules, 2017, 18(3), 914–923 CrossRef CAS PubMed.
- X. Yang, K. Hu, G. Hu, D. Shi, Y. Jiang, L. Hui, R. Zhu, Y. Xie and L. Yang, Long hydrophilic-and-cationic polymers: A different pathway toward preferential activity against bacterial over mammalian membranes, Biomacromolecules, 2014, 15(9), 3267–3277 CrossRef CAS PubMed.
- Z. Zhang, X. Wang, J. Liu, H. Yang, H. Tang, J. Li, S. Luan, J. Yin, L. Wang and H. Shi, Structural Element of Vitamin U-Mimicking Antibacterial Polypeptide with Ultrahigh Selectivity for Effectively Treating MRSA Infections, Angew. Chem., 2024, 136(7), e202318011 CrossRef.
- L. Liu, K. Xu, H. Wang, P. K. J. Tan, W. Fan, S. S. Venkatraman, L. Li and Y.-Y. Yang, Self-assembled cationic peptide nanoparticles as an efficient antimicrobial agent, Nat. NANO, 2009, 4, 457–463 CrossRef CAS PubMed.
- Y. Yang, Z. Cai, Z. Huang, X. Tang and X. Zhang, Antimicrobial cationic polymers: From structural design to functional control, Polym. J., 2018, 50(1), 33–44 Search PubMed.
- Y. Qian, S. Deng, Z. Cong, H. Zhang, Z. Lu, N. Shao, S. A. Bhatti, C. Zhou, J. Cheng and S. H. Gellman, Secondary amine pendant β-peptide polymers displaying potent antibacterial activity and promising therapeutic potential in treating MRSA-induced wound infections and keratitis, J. Am. Chem. Soc., 2022, 144(4), 1690–1699 CrossRef CAS PubMed.
- A. C. Engler, A. Shukla, S. Puranam, H. G. Buss, N. Jreige and P. T. Hammond, Effects of side group functionality and molecular weight on the activity of synthetic antimicrobial polypeptides, Biomacromolecules, 2011, 12(5), 1666–1674 CrossRef CAS.
- E. F. Palermo, S. Vemparala and K. Kuroda, Cationic spacer arm design strategy for control of antimicrobial activity and conformation of amphiphilic methacrylate random copolymers, Biomacromolecules, 2012, 13(5), 1632–1641 CrossRef CAS PubMed.
- L. Liu, K. C. Courtney, S. W. Huth, L. A. Rank, B. Weisblum, E. R. Chapman and S. H. Gellman, Beyond amphiphilic balance: Changing subunit stereochemistry alters the pore-forming activity of nylon-3 polymers, J. Am. Chem. Soc., 2021, 143(8), 3219–3230 CrossRef CAS PubMed.
- R. Dantes, Y. Mu, R. Belflower, D. Aragon, G. Dumyati, L. H. Harrison, F. C. Lessa, R. Lynfield, J. Nadle and S. Petit, National burden of invasive methicillin-resistant Staphylococcus aureus infections, United States, 2011, JAMA Intern. Med., 2013, 173(21), 1970–1978 Search PubMed.
- M.-H. Schmid-Wendtner and H. C. Korting, The pH of the skin surface and its impact on the barrier function, Skin Pharmacol. Physiol., 2006, 19(6), 296–302 CrossRef PubMed.
- J. L. Matousek and K. L. Campbell, A comparative review of cutaneous pH, Vet. Dermatol., 2002, 13(6), 293–300 CrossRef PubMed.
- Z. M. Al-Badri, A. Som, S. Lyon, C. F. Nelson, K. Nüsslein and G. N. Tew, Investigating the effect of increasing charge density on the hemolytic activity of synthetic antimicrobial polymers, Biomacromolecules, 2008, 9(10), 2805–2810 CrossRef CAS PubMed.
- G. Zou, E. de Leeuw, C. Li, M. Pazgier, C. Li, P. Zeng, W.-Y. Lu, J. Lubkowski and W. Lu, Toward understanding the cationicity of defensins: Arg and Lys versus their noncoded analogs, J. Biol. Chem., 2007, 282(27), 19653–19665 CrossRef CAS PubMed.
- R. Abes, A. Arzumanov, H. Moulton, S. Abes, G. Ivanova, M. Gait, P. Iversen and B. Lebleu, Arginine-rich cell penetrating peptides: Design, structure–activity, and applications to alter pre-mRNA splicing by steric-block oligonucleotides, J. Pept. Sci., 2008, 14(4), 455–460 CrossRef CAS.
- R. Brock, The uptake of arginine-rich cell-penetrating peptides: putting the puzzle together, Bioconjugate Chem., 2014, 25(5), 863–868 CrossRef CAS PubMed.
- F. Sgolastra, B. M. Deronde, J. M. Sarapas, A. Som and G. N. Tew, Designing mimics of membrane active proteins, Acc. Chem. Res., 2013, 46(12), 2977–2987 CrossRef CAS PubMed.
- K. E. Locock, Bioinspired polymers: antimicrobial polymethacrylates, Aust. J. Chem., 2016, 69(7), 717–724 CrossRef CAS.
- G. J. Gabriel, A. E. Madkour, J. M. Dabkowski, C. F. Nelson, K. Nüsslein and G. N. Tew, Synthetic mimic of antimicrobial peptide with nonmembrane-disrupting antibacterial properties, Biomacromolecules, 2008, 9(11), 2980–2983 CrossRef CAS PubMed.
- T. Ikeda, H. Yamaguchi and S. Tazuke, New polymeric biocides: synthesis and antibacterial activities of polycations with pendant biguanide groups, Antimicrob. Agents Chemother., 1984, 26(2), 139–144 CrossRef CAS.
- W. Chin, G. Zhong, Q. Pu, C. Yang, W. Lou, P. F. De Sessions, B. Periaswamy, A. Lee, Z. C. Liang and X. Ding, A macromolecular approach to eradicate multidrug resistant bacterial infections while mitigating drug resistance onset, Nat. Commun., 2018, 9(1), 917 CrossRef PubMed.
- X. Ding, C. Yang, W. Moreira, P. Yuan, B. Periaswamy, P. F. de Sessions, H. Zhao, J. Tan, A. Lee and K. X. Ong, A macromolecule reversing antibiotic resistance phenotype and repurposing drugs as potent antibiotics, Adv. Sci., 2020, 7(17), 2001374 CrossRef CAS.
- W. Jiang, M. Zhou, Z. Cong, J. Xie, W. Zhang, S. Chen, J. Zou, Z. Ji, N. Shao and X. Chen, Short guanidinium-functionalized poly (2-oxazoline) s displaying potent therapeutic efficacy on drug-resistant fungal infections, Angew. Chem., 2022, 134(17), e202200778 CrossRef.
- Z. Chen, C. Zhou, Y. Xu, K. Wen, J. Song, S. Bai, C. Wu, W. Huang, Q. Cai and K. Zhou, An alternatingly amphiphilic, resistance-resistant antimicrobial oligoguanidine with dual mechanisms of action, Biomaterials, 2021, 275, 120858 CrossRef CAS PubMed.
- J. Leong, C. Yang, J. Tan, B. Q. Tan, S. Hor, J. L. Hedrick and Y. Y. Yang, Combination of guanidinium and quaternary ammonium polymers with distinctive antimicrobial mechanisms achieving a synergistic antimicrobial effect, Biomater. Sci., 2020, 8(24), 6920–6929 RSC.
- Z. Shi, X. Zhang, Z. Yu, F. Yang, H. Liu, R. Xue, S. Luan and H. Tang, Facile synthesis of imidazolium-based block copolypeptides with excellent antimicrobial activity, Biomacromolecules, 2021, 22(6), 2373–2381 CrossRef CAS PubMed.
- Z. Zheng, Q. Xu, J. Guo, J. Qin, H. Mao, B. Wang and F. Yan, Structure–antibacterial activity relationships of imidazolium-type ionic liquid monomers, poly (ionic liquids) and poly (ionic liquid) membranes: effect of alkyl chain length and cations, ACS Appl. Mater. Interfaces, 2016, 8(20), 12684–12692 CrossRef CAS PubMed.
- V. W. L. Ng, J. P. K. Tan, J. Leong, Z. X. Voo, J. L. Hedrick and Y. Y. Yang, Antimicrobial polycarbonates: Investigating the impact of nitrogen-containing heterocycles as quaternizing agents, Macromolecules, 2014, 47(4), 1285–1291 CrossRef CAS.
- V. Sambhy, B. R. Peterson and A. Sen, Antibacterial and hemolytic activities of pyridinium polymers as a function of the spatial relationship between the positive charge and the pendant alkyl tail, Angew. Chem., 2008, 120(7), 1270–1274 CrossRef.
- R. Tejero, D. López, F. López-Fabal, J. L. Gómez-Garcés and M. Fernández-García, High efficiency antimicrobial thiazolium and triazolium side-chain polymethacrylates obtained by controlled alkylation of the corresponding azole derivatives, Biomacromolecules, 2015, 16(6), 1844–1854 CrossRef CAS PubMed.
- A. Kanazawa, T. Ikeda and T. Endo, Antibacterial activity of polymeric sulfonium salts, J. Polym. Sci., Part A: Polym. Chem., 1993, 31(11), 2873–2876 CrossRef CAS.
- M. Hirayama, The antimicrobial activity, hydrophobicity and toxicity of sulfonium compounds, and their relationship, Biocontrol Sci., 2011, 16(1), 23–31 CrossRef CAS PubMed.
- J. A. Feliciano, A. J. Leitgeb, C. L. Schrank, R. A. Allen, K. P. Minbiole, W. M. Wuest and R. G. Carden, Trivalent sulfonium compounds (TSCs): Tetrahydrothiophene-based amphiphiles exhibit similar antimicrobial activity to analogous ammonium-based amphiphiles, Bioorg. Med. Chem. Lett., 2021, 37, 127809 CrossRef CAS PubMed.
- B. Zhang, M. Li, M. Lin, X. Yang and J. Sun, A convenient approach for antibacterial polypeptoids featuring sulfonium and oligo (ethylene glycol) subunits, Biomater. Sci., 2020, 8(24), 6969–6977 RSC.
- J. Oh and A. Khan, Main-chain polysulfonium salts: Development of non-ammonium antibacterial polymers similar in their activity to antibiotic drugs vancomycin and kanamycin, Biomacromolecules, 2021, 22(8), 3534–3542 CrossRef CAS PubMed.
- D. Guan, F. Chen, Y. Qiu, B. Jiang, L. Gong, L. Lan and W. Huang, Sulfonium, an underestimated moiety for structural modification, alters the antibacterial profile of vancomycin against multidrug-resistant bacteria, Angew. Chem., 2019, 131(20), 6750–6754 CrossRef.
- Y. Hu, J. Zhao, J. Zhang, Z. Zhu and J. Rao, Broad-spectrum bactericidal activity and remarkable selectivity of main-chain sulfonium-containing polymers with alternating sequences, ACS Macro Lett., 2021, 10(8), 990–995 CrossRef CAS PubMed.
- X. Wang, G. Wang, J. Zhao, Z. Zhu and J. Rao, Main-chain sulfonium-containing homopolymers with negligible hemolytic toxicity for eradication of bacterial and fungal biofilms, ACS Macro Lett., 2021, 10(12), 1643–1649 CrossRef CAS PubMed.
- S. Kanwal, O. Staudhammer, U. B. A. Aziz, E. Quaas, J. Rademann and D. Klinger, Sulfonium-Based Antimicrobial Block Copolymers: Influence of Hydrophobicity on Biological Activity and Antibiotic Synergy, Macromol. Rapid Commun., 2025, e00421 CrossRef PubMed.
- Y. Gong, X. Xu, M. Aquib, Y. Zhang, W. Yang, Y. Chang, H. Peng, C. Boyer, A. K. Whittaker and C. Fu, Ammonium, phosphonium, and sulfonium polymers for antimicrobial applications: a comparative study, ACS Appl. Polym. Mater., 2024, 6(12), 6966–6975 CrossRef CAS.
- J. Sun, M. Li, M. Lin, B. Zhang and X. Chen, High antibacterial activity and selectivity of the versatile polysulfoniums that combat drug resistance, Adv. Mater., 2021, 33(41), 2104402 CrossRef CAS PubMed.
- T. J. Cuthbert, B. Hisey, T. D. Harrison, J. F. Trant, E. R. Gillies and P. J. Ragogna, Surprising antibacterial activity and selectivity of hydrophilic polyphosphoniums featuring sugar and hydroxy substituents, Angew. Chem., Int. Ed., 2018, 57(39), 12707–12710 CrossRef CAS PubMed.
- A. Kanazawa, T. Ikeda and T. Endo, Novel polycationic biocides: synthesis and antibacterial activity of polymeric phosphonium salts, J. Polym. Sci., Part A: Polym. Chem., 1993, 31(2), 335–343 CrossRef CAS.
- H. Mortazavian, L. L. Foster, R. Bhat, S. Patel and K. Kuroda, Decoupling the functional roles of cationic and hydrophobic groups in the antimicrobial and hemolytic activities of methacrylate random copolymers, Biomacromolecules, 2018, 19(11), 4370–4378 CrossRef CAS PubMed.
- A. C. Engler, J. P. Tan, Z. Y. Ong, D. J. Coady, V. W. Ng, Y. Y. Yang and J. L. Hedrick, Antimicrobial polycarbonates: investigating the impact of balancing charge and hydrophobicity using a same-centered polymer approach, Biomacromolecules, 2013, 14(12), 4331–4339 CrossRef CAS PubMed.
- G. Lu, D. Wu and R. Fu, Studies on the synthesis and antibacterial activities of polymeric quaternary ammonium salts from dimethylaminoethyl methacrylate, React. Funct. Polym., 2007, 67(4), 355–366 CrossRef CAS.
- B. Dizman, M. O. Elasri and L. J. Mathias, Synthesis and antimicrobial activities of new water-soluble bis-quaternary ammonium methacrylate polymers, J. Appl. Polym. Sci., 2004, 94(2), 635–642 CrossRef CAS.
- Y. He, E. Heine, N. Keusgen, H. Keul and M. Möller, Synthesis and characterization of amphiphilic monodisperse compounds and poly (ethylene imine) s: Influence of their microstructures on the antimicrobial properties, Biomacromolecules, 2012, 13(3), 612–623 CrossRef CAS PubMed.
- D. S. Uppu, S. Samaddar, J. Hoque, M. M. Konai, P. Krishnamoorthy, B. R. Shome and J. Haldar, Side chain degradable cationic–amphiphilic polymers with tunable hydrophobicity show in vivo activity, Biomacromolecules, 2016, 17(9), 3094–3102 CrossRef CAS.
- S. Chakraborty, R. Liu, J. J. Lemke, Z. Hayouka, R. A. Welch, B. Weisblum, K. S. Masters and S. H. Gellman, Effects of cyclic vs acyclic hydrophobic subunits on the chemical structure and biological properties of nylon-3 copolymers, ACS Macro Lett., 2013, 2(8), 753–756 CrossRef CAS PubMed.
- E. Moreau, M. Domurado, P. Chapon, M. Vert and D. Domurado, Biocompatibility of polycations: in vitro agglutination and lysis of red blood cells and in vivo toxicity, J. Drug Targeting, 2002, 10(2), 161–173 CrossRef CAS PubMed.
- R. Bottega and R. M. Epand, Inhibition of protein kinase C by cationic amphiphiles, Biochemistry, 1992, 31(37), 9025–9030 CrossRef CAS PubMed.
- Y. Kim, S. Binauld and M. H. Stenzel, Zwitterionic guanidine-based oligomers mimicking cell-penetrating peptides as a nontoxic alternative to cationic polymers to enhance the cellular uptake of micelles, Biomacromolecules, 2012, 13(10), 3418–3426 CrossRef CAS PubMed.
- H. Lv, S. Zhang, B. Wang, S. Cui and J. Yan, Toxicity of cationic lipids and cationic polymers in gene delivery, J. Controlled Release, 2006, 114(1), 100–109 CrossRef CAS PubMed.
- D. Fischer, Y. Li, B. Ahlemeyer, J. Krieglstein and T. Kissel, In vitro cytotoxicity testing of polycations: influence of polymer structure on cell viability and hemolysis, Biomaterials, 2003, 24(7), 1121–1131 CrossRef CAS PubMed.
- R. Namivandi-Zangeneh, Z. Sadrearhami, D. Dutta, M. Willcox, E. H. Wong and C. Boyer, Synergy between synthetic antimicrobial polymer and antibiotics: a promising platform to combat multidrug-resistant bacteria, ACS Infect. Dis., 2019, 5(8), 1357–1365 CrossRef CAS PubMed.
- P. H. Sellenet, B. Allison, B. M. Applegate and J. P. Youngblood, Synergistic activity of hydrophilic modification in antibiotic polymers, Biomacromolecules, 2007, 8(1), 19–23 CrossRef CAS PubMed.
- G. Rani, K. Kuroda and S. Vemparala, Aggregation of methacrylate-based ternary biomimetic antimicrobial polymers in solution, J. Phys.: Condens. Matter, 2020, 33(6), 064003 CrossRef PubMed.
- G. Rani, K. Kuroda and S. Vemparala, Towards designing globular antimicrobial peptide mimics: role of polar functional groups in biomimetic ternary antimicrobial polymers, Soft Matter, 2021, 17(8), 2090–2103 RSC.
- J. D. Katiyar, A. Halder, M. Avais, H. Aidasani, O. Mukherjee and S. Chattopadhyay, Hydrophobic PEGylation of Chitosan: A Graft Copolymer Approach toward Developing Nontoxic Antimicrobial Chitosan, ACS Appl. Polym. Mater., 2023, 5(12), 9742–9750 CrossRef CAS.
- J. Bouckaert, J. Berglund, M. Schembri, E. De Genst, L. Cools, M. Wuhrer, C. S. Hung, J. Pinkner, R. Slättegård and A. Zavialov, Receptor binding studies disclose a novel class of high-affinity inhibitors of the Escherichia coli FimH adhesin, Mol. Microbiol., 2005, 55(2), 441–455 CrossRef CAS PubMed.
- Y. Chen, L. Yu, B. Zhang, W. Feng, M. Xu, L. Gao, N. Liu, Q. Wang, X. Huang and P. Li, Design and synthesis of biocompatible, hemocompatible, and highly selective antimicrobial cationic peptidopolysaccharides via click chemistry, Biomacromolecules, 2019, 20(6), 2230–2240 CrossRef CAS PubMed.
- L. Yu, J. Chen, P. Peng, R. Ding, K. Wang, C. Yan, M. Sun, Z. Du, C. Boyer and P. Li, Broad-spectrum Antibacterial Peptidopolysaccharides for Targeted Therapy of a Drug-resistant Bacterial Liver Abscess, Acta Biomater., 2025, 206, 371–384 CrossRef CAS PubMed.
- C.-W. Chang, E. Bays, L. Tao, S. N. Alconcel and H. D. Maynard, Differences in cytotoxicity of poly (PEGA) s synthesized by reversible addition–fragmentation chain transfer polymerization, Chem. Commun., 2009, 3580–3582 RSC.
- T. D. Michl, K. E. Locock, N. E. Stevens, J. D. Hayball, K. Vasilev, A. Postma, Y. Qu, A. Traven, M. Haeussler and L. Meagher, RAFT-derived antimicrobial polymethacrylates: elucidating the impact of end-groups on activity and cytotoxicity, Polym. Chem., 2014, 5(19), 5813–5822 RSC.
- J. L. Grace, J. X. Huang, S.-E. Cheah, N. P. Truong, M. A. Cooper, J. Li, T. P. Davis, J. F. Quinn, T. Velkov and M. R. Whittaker, Antibacterial low molecular weight cationic polymers: Dissecting the contribution of hydrophobicity, chain length and charge to activity, RSC Adv., 2016, 6(19), 15469–15477 RSC.
- J. L. Grace, A. G. Elliott, J. X. Huang, E. K. Schneider, N. P. Truong, M. A. Cooper, J. Li, T. P. Davis, J. F. Quinn and T. Velkov, Cationic acrylate oligomers comprising amino acid mimic moieties demonstrate improved antibacterial killing efficiency, J. Mater. Chem. B, 2017, 5(3), 531–536 RSC.
- K. Lienkamp, K. N. Kumar, A. Som, K. Nüsslein and G. N. Tew, “Doubly selective” antimicrobial polymers: how do they differentiate between bacteria?, Chem. – Eur. J., 2009, 15(43), 11710–11714 CrossRef CAS PubMed.
- A. Tyagi and A. Mishra, Optimal balance of hydrophobic content and degree of polymerization results in a potent membrane-targeting antibacterial polymer, ACS Omega, 2021, 6(50), 34724–34735 CrossRef CAS.
- J. Pachla, R. J. Kopiasz, G. Marek, W. Tomaszewski, A. Głogowska, K. Drezek, S. Kowalczyk, R. Podgórski, B. Butruk-Raszeja and T. Ciach, Polytrimethylenimines: Highly potent antibacterial agents with activity and toxicity modulated by the polymer molecular weight, Biomacromolecules, 2023, 24(5), 2237–2249 CrossRef CAS PubMed.
- S. Oliver, L. Zhao, A. J. Gormley, R. Chapman and C. Boyer, Living in the fast lane—High throughput controlled/living radical polymerization, Macromolecules, 2018, 52(1), 3–23 CrossRef.
- H. Javadi, A. C. Lehnen and M. Hartlieb, Bioinspired Cationic Antimicrobial Polymers, Angew. Chem., Int. Ed., 2025, 64(24), e202503738 CrossRef CAS PubMed.
- L. Liu, Y. Huang, S. N. Riduan, S. Gao, Y. Yang, W. Fan and Y. Zhang, Main-chain imidazolium oligomer material as a selective biomimetic antimicrobial agent, Biomaterials, 2012, 33(33), 8625–8631 CrossRef CAS PubMed.
- J. Guo, J. Qin, Y. Ren, B. Wang, H. Cui, Y. Ding, H. Mao and F. Yan, Antibacterial activity of cationic polymers: side-chain or main-chain type?, Polym. Chem., 2018, 9(37), 4611–4616 RSC.
- C. H. Koh, M. R. Lambu, C. Tan, G. Wei, Z. Y. Kok, K. Zhang, Q. H. N. Vu, M. Panneerselvam, Y. J. Ooi and S. H. Tan, Carbene formation as a mechanism for efficient intracellular uptake of cationic antimicrobial carbon acid polymers, Nat. Commun., 2025, 16(1), 6460 CrossRef CAS PubMed.
- M.-N. Antonopoulou, R. Whitfield, N. P. Truong, D. Wyers, S. Harrisson, T. Junkers and A. Anastasaki, Concurrent control over sequence and dispersity in multiblock copolymers, Nat. Chem., 2022, 14(3), 304–312 CrossRef CAS.
- F. Perin, A. Motta and D. Maniglio, Amphiphilic copolymers in biomedical applications: Synthesis routes and property control, Mater. Sci. Eng., C, 2021, 123, 111952 CrossRef CAS PubMed.
- H. Takahashi, I. Sovadinova, K. Yasuhara, S. Vemparala, G. A. Caputo and K. Kuroda, Biomimetic antimicrobial polymers—Design, characterization, antimicrobial, and novel applications, Wiley Interdiscip. Rev.:Nanomed. Nanobiotechnol., 2023, 15(3), e1866 CAS.
- M. A. Cortez, W. T. Godbey, Y. Fang, M. E. Payne, B. J. Cafferty, K. A. Kosakowska and S. M. Grayson, The synthesis of cyclic poly (ethylene imine) and exact linear analogues: an evaluation of gene delivery comparing polymer architectures, J. Am. Chem. Soc., 2015, 137(20), 6541–6549 CrossRef CAS PubMed.
- M. Aquib, W. Yang, L. Yu, V. K. Kannaujiya, Y. Zhang, P. Li, A. Whittaker, C. Fu and C. Boyer, Effect of cyclic topology versus linear terpolymers on antibacterial activity and biocompatibility: antimicrobial peptide avatars, Chem. Sci., 2024, 15(45), 19057–19069 RSC.
- C. Gonnot, M. B. H. Mahboob, M. Aumond, J. R. Tait, K. Nay, K. Nazemi, H. Floyd, J. Zuegg, F. Boeda and C. B. Landersdorfer, Cyclic and linear cationic polymers based on metathesis polymerization for antibacterial and antifungal Applications, Eur. Polym. J., 2025, 230, 113875 CrossRef CAS.
- R. J. Kopiasz, M. Dranka, W. Tomaszewski, P. Kowalska, B. Butruk-Raszeja, K. Drezek, J. Mierzejewska, T. Ciach and D. Janczewski, Antimicrobial Macrocycles–Synthesis, Characterization, and Activity Comparison with Their Linear Polycationic Analogues, Biomacromolecules, 2024, 25(12), 7814–7827 CrossRef CAS PubMed.
- H. Luo and E. H. Wong, Antimicrobial Cyclic Peptidomimetics, Macromol. Rapid Commun., 2025, e00569 CrossRef PubMed.
- P. Salas-Ambrosio, A. Tronnet, M. Since, S. Bourgeade-Delmas, J.-L. Stigliani, A. Vax, S. Lecommandoux, B. Dupuy, P. Verhaeghe and C. Bonduelle, Cyclic poly (α-peptoid) s by lithium bis (trimethylsilyl) amide (LiHMDS)-mediated ring-expansion polymerization: simple access to bioactive backbones, J. Am. Chem. Soc., 2021, 143(10), 3697–3702 CrossRef CAS PubMed.
- H. Gmedhin, S. Schaefer, N. Corrigan, P. Wu, Z. Gu, M. D. Lenardon and C. Boyer, Effect of Defined Block Sequence Terpolymers on Antifungal Activity and Biocompatibility, Macromol. Biosci., 2025, 25(4), 2400429 CrossRef CAS PubMed.
- M. Godzina, R. Terracciano, Z. Varanaraja, D. Mackinnon and C. R. Becer, Self-assembled nanoparticles of PEG and poly (2-oxazoline) based lactide block copolymers, Eur. Polym. J., 2024, 221, 113517 CrossRef CAS.
- G. Sauvet, W. Fortuniak, K. Kazmierski and J. Chojnowski, Amphiphilic block and statistical siloxane copolymers with antimicrobial activity, J. Polym. Sci., Part A: Polym. Chem., 2003, 41(19), 2939–2948 CrossRef CAS.
- Y. Wang, J. Xu, Y. Zhang, H. Yan and K. Liu, Antimicrobial and hemolytic activities of copolymers with cationic and hydrophobic groups: A comparison of block and random copolymers, Macromol. Biosci., 2011, 11(11), 1499–1504 CrossRef CAS PubMed.
- Y. Oda, S. Kanaoka, T. Sato, S. Aoshima and K. Kuroda, Block versus random amphiphilic copolymers as antibacterial agents, Biomacromolecules, 2011, 12(10), 3581–3591 CrossRef CAS PubMed.
- M. Álvarez-Paino, A. Muñoz-Bonilla, F. López-Fabal, J. L. Gómez-Garcés, J. P. Heuts and M. Fernández-García, Effect of glycounits on the antimicrobial properties and toxicity behavior of polymers based on quaternized DMAEMA, Biomacromolecules, 2015, 16(1), 295–303 CrossRef PubMed.
- K. Agnes, S. Parveen, Q. Yue, P. Raoul, S.-C. Carlos, M. John, W. E. GL, L. K. ES, H. Matthias and P. Sebastien, Sequence Control as a Powerful Tool for Improving the Selectivity of Antimicrobial Polymers, 2017.
- A. Song, S. G. Walker, K. A. Parker and N. S. Sampson, Antibacterial studies of cationic polymers with alternating, random, and uniform backbones, ACS Chem. Biol., 2011, 6(6), 590–599 CrossRef CAS PubMed.
- D. Uppu, M. Bhowmik, S. Samaddar and J. Haldar, Cyclization and unsaturation rather than isomerisation of side chains govern the selective antibacterial activity of cationic-amphiphilic polymers, Chem. Commun., 2016, 52(25), 4644–4647 RSC.
- G. Hayes and C. R. Becer, Hyperbranched poly (2-oxazoline) s via bisfunctional crosslinker, Eur. Polym. J., 2022, 181, 111678 CrossRef CAS.
- S. Vandewalle, S. Wallyn, S. Chattopadhyay, C. R. Becer and F. Du Prez, Thermoresponsive hyperbranched glycopolymers: Synthesis, characterization and lectin interaction studies, Eur. Polym. J., 2015, 69, 490–498 CrossRef CAS.
- G. Liu, G. Zhang, J. Hu, X. Wang, M. Zhu and S. Liu, Hyperbranched self-immolative polymers (h SIPs) for programmed payload delivery and ultrasensitive detection, J. Am. Chem. Soc., 2015, 137(36), 11645–11655 CrossRef CAS PubMed.
- T. Kanai, D. Thirumoolan, R. Mohanram, K. Vetrivel and K. A. Basha, Antimicrobial activity of hyperbranched polymers: Synthesis, characterization, and activity assay study, J. Bioact. Compat. Polym., 2015, 30(2), 145–156 CrossRef CAS.
- S. Q. Chen, L. Xu, C. He, P. Y. Li, X. X. Lu, J. M. Li, H. J. Li, W. D. He and L. Yang, Long-subchain hyperbranched poly (aminoethyl acrylate): A potent antimicrobial polymer with low hemolytic toxicity, J. Polym. Sci., Part A: Polym. Chem., 2016, 54(21), 3462–3469 CrossRef CAS.
- K. A. Gibney, I. Sovadinova, A. I. Lopez, M. Urban, Z. Ridgway, G. A. Caputo and K. Kuroda, Poly (ethylene imine) s as antimicrobial agents with selective activity, Macromol. Biosci., 2012, 12(9), 1279–1289 CrossRef CAS PubMed.
- N. Pasquier, H. Keul, E. Heine, M. Moeller, B. Angelov, S. Linser and R. Willumeit, Amphiphilic branched polymers as antimicrobial agents, Macromol. Biosci., 2008, 8(10), 903–915 CrossRef CAS PubMed.
- C. R. Arza, X. Li, S. Ilk, Y. Liu, D. Demircan and B. Zhang, Biocompatible non-leachable antimicrobial polymers with a nonionic hyperbranched backbone and phenolic terminal units, J. Mater. Chem. B, 2022, 10(39), 8064–8074 RSC.
- G. Hayes, B. Drain and C. R. Becer, Multiarm core Cross-Linked Star-Shaped poly (2-oxazoline) s using a bisfunctional 2-Oxazoline monomer, Macromolecules, 2021, 55(1), 146–155 CrossRef.
- A. Monaco, B. Drain and C. R. Becer, Detailed GPC analysis of poly (N-isopropylacrylamide) with core cross-linked star architecture, Polym. Chem., 2021, 12(36), 5229–5238 RSC.
- J. Hu, R. Qiao, M. R. Whittaker, J. F. Quinn and T. P. Davis, Synthesis of star polymers by RAFT polymerization as versatile nanoparticles for biomedical applications, Aust. J. Chem., 2017, 70(11), 1161–1170 CrossRef CAS.
- J. M. Ren, T. G. McKenzie, Q. Fu, E. H. Wong, J. Xu, Z. An, S. Shanmugam, T. P. Davis, C. Boyer and G. G. Qiao, Star polymers, Chem. Rev., 2016, 116(12), 6743–6836 CrossRef CAS PubMed.
- S. J. Lam, E. H. Wong, N. M. O’Brien-Simpson, N. Pantarat, A. Blencowe, E. C. Reynolds and G. G. Qiao, Bionano interaction study on antimicrobial star-shaped peptide polymer nanoparticles, ACS Appl. Mater. Interfaces, 2016, 8(49), 33446–33456 CrossRef CAS.
- E. H. Wong, M. M. Khin, V. Ravikumar, Z. Si, S. A. Rice and M. B. Chan-Park, Modulating antimicrobial activity and mammalian cell biocompatibility with glucosamine-functionalized star polymers, Biomacromolecules, 2016, 17(3), 1170–1178 CrossRef CAS PubMed.
- Y. Pu, Z. Hou, M. M. Khin, R. Zamudio-Vázquez, K. L. Poon, H. Duan and M. B. Chan-Park, Synthesis and antibacterial study of sulfobetaine/quaternary ammonium-modified star-shaped poly [2-(dimethylamino) ethyl methacrylate]-based copolymers with an inorganic core, Biomacromolecules, 2016, 18(1), 44–55 CrossRef PubMed.
- C. Yang, S. Krishnamurthy, J. Liu, S. Liu, X. Lu, D. J. Coady, W. Cheng, G. De Libero, A. Singhal and J. L. Hedrick, Broad-Spectrum Antimicrobial Star Polycarbonates Functionalized with Mannose for Targeting Bacteria Residing inside Immune Cells, Adv. Healthcare Mater., 2016, 5(11), 1272–1281 CrossRef CAS PubMed.
- N. Wiradharma, S. Q. Liu and Y. Y. Yang, Branched and 4-Arm Starlike α-Helical Peptide Structures with Enhanced Antimicrobial Potency and Selectivity, Small, 2012, 8(3), 362–366 CrossRef CAS PubMed.
- W. Li, S. Hadjigol, A. R. Mazo, J. Holden, J. Lenzo, S. J. Shirbin, A. Barlow, S. Shabani, T. Huang and E. C. Reynolds, Star-peptide polymers are multi-drug-resistant Gram-positive bacteria killers, ACS Appl. Mater. Interfaces, 2022, 14(22), 25025–25041 CrossRef CAS PubMed.
- S. Laroque, K. E. Locock and S. Perrier, Cationic Star Polymers Obtained by the Arm-First Approach-Influence of Arm Number and Positioning of Cationic Units on Antimicrobial Activity, Biomacromolecules, 2024, 26(1), 190–200 CrossRef PubMed.
- M. Aquib, S. Schaefer, H. Gmedhin, N. Corrigan, V. A. Bobrin and C. Boyer, Shape matters: Effect of amphiphilic polymer topology on antibacterial activity and hemocompatibility, Eur. Polym. J., 2024, 205, 112698 CrossRef CAS.
- Y.-F. Chen, Y.-D. Lai, C.-H. Chang, Y.-C. Tsai, C.-C. Tang and J.-S. Jan, Star-shaped polypeptides exhibit potent antibacterial activities, Nanoscale, 2019, 11(24), 11696–11708 RSC.
- M. Pan, C. Lu, M. Zheng, W. Zhou, F. Song, W. Chen, F. Yao, D. Liu and J. Cai, Unnatural amino-acid-based star-shaped poly (L-Ornithine) s as emerging long-term and biofilm-disrupting antimicrobial peptides to treat Pseudomonas aeruginosa-infected burn wounds, Adv. Healthcare Mater., 2020, 9(19), 2000647 CrossRef CAS PubMed.
- I. Mukherjee, A. Ghosh, P. Bhadury and P. De, Leucine-based polymer architecture-induced antimicrobial properties and bacterial cell morphology switching, ACS Omega, 2018, 3(1), 769–780 CrossRef CAS PubMed.
- L. Yin, L. Liu and N. Zhang, Brush-like polymers: design, synthesis and applications, Chem. Commun., 2021, 57(81), 10484–10499 RSC.
- W.-L. Chen, R. Cordero, H. Tran and C. K. Ober, 50th anniversary perspective: Polymer brushes: Novel surfaces for future materials, Macromolecules, 2017, 50(11), 4089–4113 CrossRef CAS.
- A.-C. Lehnen, A. M. Bapolisi, M. Krass, A. AlSawaf, J. Kurki, S. Kersting, H. Fuchs and M. Hartlieb, Shape matters: Highly selective Antimicrobial Bottle Brush copolymers via a one-pot RAFT polymerization approach, Biomacromolecules, 2022, 23(12), 5350–5360 CrossRef CAS PubMed.
- O. Shelef, S. Gnaim and D. Shabat, Self-immolative polymers: an emerging class of degradable materials with distinct disassembly profiles, J. Am. Chem. Soc., 2021, 143(50), 21177–21188 CrossRef CAS PubMed.
- Y. Li, J. Liu, J. He, A. Dey, V. D. Bui and J. H. Park, Recent advances in stimuli-responsive self-immolative polymers for drug delivery and molecular imaging, Chem. Mater., 2024, 36(9), 4054–4077 CrossRef CAS.
- J. Gong, B. Tavsanli and E. R. Gillies, Self-immolative polymers: From synthesis to applications, Annu. Rev. Mater. Res., 2024, 54(1), 47–73 CrossRef CAS.
- J. Guo, Y. Tao, Z. Du, S. Zhang, W. Zheng, Z. Wang, Z. Yi, Y. Gou and W. Tang, Stimuli-Responsive Antimicrobial Polymer Systems: From Structural Design to Biomedical Applications, Giant, 2025, 100366 CrossRef CAS.
- Z. Shao, Y. D. Xu, H. Luo, K. Hakobyan, M. Zhang, J. Xu, M. H. Stenzel and E. H. Wong, Smart Galactosidase-Responsive Antimicrobial Dendron: Towards More Biocompatible Membrane-Disruptive Agents, Macromol. Rapid Commun., 2024, 45(19), 2400350 CrossRef CAS PubMed.
- C. Ergene and E. F. Palermo, Cationic poly (benzyl ether) s as self-Immolative antimicrobial polymers, Biomacromolecules, 2017, 18(10), 3400–3409 CrossRef CAS.
- M. Aquib, L. Zong, V. K. Kannaujiya, T. Quan, Z. Gu and C. Boyer, Stimuli-Responsive “Arm-First” Star Copolymers for Antibacterial Activity and Biocompatibility, ACS Appl. Polym. Mater., 2025, 7(24), 16847–16858 CrossRef CAS.
- A. M. Bapolisi, A. C. Lehnen, M. Wolff, J. Kramer, S. Kogikoski Jr, R. Steinbrecher, N. Michler, A. Kiesow, I. Bald and M. Obry, Non-Amphiphilic Antimicrobial Polymers, Angew. Chem., Int. Ed., 2025, 64(33), e202507564 CrossRef CAS PubMed.
- W. Chin, C. Yang, V. W. L. Ng, Y. Huang, J. Cheng, Y. W. Tong, D. J. Coady, W. Fan, J. L. Hedrick and Y. Y. Yang, Biodegradable broad-spectrum antimicrobial polycarbonates: Investigating the role of chemical structure on activity and selectivity, Macromolecules, 2013, 46(22), 8797–8807 CrossRef CAS.
- S. Mankoci, R. L. Kaiser, N. Sahai, H. A. Barton and A. Joy, Bactericidal peptidomimetic polyurethanes with remarkable selectivity against Escherichia coli, ACS Biomater. Sci. Eng., 2017, 3(10), 2588–2597 CrossRef CAS PubMed.
- H. Wang, M. Langner and S. Agarwal, Biodegradable aliphatic–aromatic polyester with antibacterial property, Polym. Eng. Sci., 2016, 56(10), 1146–1152 CrossRef CAS.
- M. Mizutani, E. F. Palermo, L. M. Thoma, K. Satoh, M. Kamigaito and K. Kuroda, Design and synthesis of self-degradable antibacterial polymers by simultaneous chain-and step-growth radical copolymerization, Biomacromolecules, 2012, 13(5), 1554–1563 CrossRef CAS PubMed.
- W. Janik, M. Nowotarski, D. Y. Shyntum, A. Banaś, K. Krukiewicz, S. Kudła and G. Dudek, Antibacterial and biodegradable polysaccharide-based films for food packaging applications: Comparative study, Materials, 2022, 15(9), 3236 CrossRef CAS PubMed.
- M. A. DeWit and E. R. Gillies, A cascade biodegradable polymer based on alternating cyclization and elimination reactions, J. Am. Chem. Soc., 2009, 131(51), 18327–18334 CrossRef CAS PubMed.
- L. Tallet, V. Gribova, L. Ploux, N. E. Vrana and P. Lavalle, New smart antimicrobial hydrogels, nanomaterials, and coatings: Earlier action, more specific, better dosing?, Adv. Healthcare Mater., 2021, 10(1), 2001199 CrossRef CAS PubMed.
- X. Wang, L. Song, J. Zhao, R. Zhou, S. Luan, Y. Huang, J. Yin and A. Khan, Bacterial adaptability of enzyme and pH dual-responsive surface for infection resistance, J. Mater. Chem. B, 2018, 6(46), 7710–7718 RSC.
- L. Shaw, E. Golonka, J. Potempa and S. J. Foster, The role and regulation of the extracellular proteases of Staphylococcus aureus, Microbiology, 2004, 150(1), 217–228 CrossRef CAS PubMed.
- B. Skalka, Hyaluronidase test in the diagnosis of staphylococci, Vet. Med., 1985, 30(6), 373–378 CAS.
- H. Jiang, Y. Li, X. Cao, Y. Liu, Z. Zhang, Y. Hu, B. Yu, H. Hu and Y. Shen, The Hyperbranched Polymer as a Medical Rising Star: From Synthesis to Biomedical Applications, Available at SSRN 5091940.
- S. Omidi, Z. Rafiee and A. Kakanejadifard, Design and synthesis of curcumin nanostructures: Evaluation of solubility, stability, antibacterial and antioxidant activities, Bioorg. Chem., 2021, 116, 105308 CrossRef CAS PubMed.
- J. Luo, Y. Jiang, Y. Lu, J. Liu, Y. Tian, M. Zhao, P. Yang and W. Wu, Conjugated Hyperbranched Polymer Photosensitizer Based Covalently Linked Micelles for Broad-Spectrum Bacterial Treatment, Adv. Healthcare Mater., 2025, e04005 Search PubMed.
- X. Wang, Y.-Q. Wang and D.-C. Wu, Facile fabrication of hyperbranched polyacetal quaternary ammonium with pH-responsive curcumin release for synergistic antibacterial activity, Chin. J. Polym. Sci., 2023, 41(4), 564–573 CrossRef CAS.
- W. K. W. A. Khodir, M. W. Ismail, S. A. Hamid, R. Daik, D. Susanti, M. Taher and V. Guarino, Synthesis and Characterization of Ciprofloxacin Loaded Star-Shaped Polycaprolactone–Polyethylene Glycol Hydrogels for Oral Delivery, Micromachines, 2023, 14(7), 1382 CrossRef PubMed.
- Y. Gong, Q. Peng, Y. Qiao, D. Tian, Y. Zhang, X. Xiong, M. He, X. Xu and B. Shi, Hyperbranched polylysine exhibits a collaborative enhancement of the antibiotic capacity to kill Gram-negative pathogens, Antibiotics, 2024, 13(3), 217 CrossRef CAS PubMed.
- T. Wu, M. Zhou, J. Zou, Q. Chen, F. Qian, J. Kurths, R. Liu and Y. Tang, AI-guided few-shot inverse design of HDP-mimicking polymers against drug-resistant bacteria, Nat. Commun., 2024, 15(1), 6288 CrossRef CAS PubMed.
- S. Han and J. Wu, Artificial intelligence (AI) meets biomaterials and biomedicine, Smart Mater. Med., 2024, 5(2), 251–255 CAS.
- J. Roh, K.-T. Huang, A. Choudhury, T. Quan, C.-J. Huang, P. Kumar and C. Boyer, Machine Learning-Guided Design of High-Performance Antifungal Polymers against Candida albicans, ACS Polymers Au, 2025, 5(5), 557–569 CrossRef CAS PubMed.
- A. Tiihonen, S. J. Cox-Vazquez, Q. Liang, M. Ragab, Z. Ren, N. T. P. Hartono, Z. Liu, S. Sun, C. Zhou and N. C. Incandela, Predicting antimicrobial activity of conjugated oligoelectrolyte molecules via machine learning, J. Am. Chem. Soc., 2021, 143(45), 18917–18931 CrossRef CAS PubMed.
- J. Roh, C. Boyer and P. V. Kumar, Uncovering Key Characteristics of Antibacterial Peptides through Machine Learning, Macromol. Rapid Commun., 2025, e00583 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |





