Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Squaratides: a tunable platform for anion binding in peptide-inspired macrocycles
Farhad Ali.
Mohammed
 ab,
Hua
Tong
a,
Chris S.
Hawes
ab,
Hua
Tong
a,
Chris S.
Hawes
 c,
Shayon
Bhattacharya
c,
Shayon
Bhattacharya
 bd,
Damien
Thompson
bd,
Damien
Thompson
 be,
Katrina A.
Jolliffe
be,
Katrina A.
Jolliffe
 f and
Robert B. P.
Elmes
f and
Robert B. P.
Elmes
 *abg
*abg
aDepartment of Chemistry, Maynooth University, National University of Ireland, Maynooth, Co, Kildare, Ireland. E-mail: robert.elmes@mu.ie
bSSPC – the Research Ireland Centre for Pharmaceuticals, University of Limerick, V94 T9PX, Limerick, Ireland
cSchool of Chemical and Physical Sciences, Keele University, Keele, ST5 5BG, UK
dDepartment of Biological Sciences, Bernal Institute, University of Limerick, V94 T9PX, Limerick, Ireland
eDepartment of Physics, Bernal Institute, University of Limerick, V94 T9PX, Limerick, Ireland
fARC Centre of Excellence for Innovations in Peptide and Protein Science, School of Chemistry, University of Sydney, 2006, NSW, Australia
gKathleen Lonsdale Institute for Human Health Research, Maynooth University, National University of Ireland, Co. Kildare, W23 F2H6, Maynooth, Ireland
First published on 26th March 2026
Abstract
We report a new class of sequence-defined peptidomimetic macrocycles, termed squaratides, that incorporate squaramide units into a tunable peptide backbone. These modular receptors can bind acetate and chloride ions where sidechain functionality can control affinity and stoichiometry.
Designed artificial receptors that mimic natural systems in their ability to bind selectively to a target ion1–4 provide molecular connectivity for supramolecular assemblies for diverse applications ranging from sustainable electronics to molecular medicine.5,6 Nature's modular protein construction enables exquisite anion recognition through complementary hydrogen-bonding interactions (e.g., the hydroxymethyl and guanidino sidechains of Ser and Arg, respectively) reinforced by backbone amides, creating preorganised cavities that bind anions with remarkable affinity and selectivity.7–9
Several groups have reported sequence-defined scaffolds that incorporate abiotic elements to enhance molecular recognition. For example, Jolliffe and co-workers designed cyclic and linear peptide based receptors for sulphate recognition7,8,10,11 while Kubik et al. incorporated 3-aminobenzoic acids and 6-aminopicolinic acids into peptidomimetic macrocycles for cation and anion recognition.12–14 Knipe et al. synthesised α-helical and β-strand peptidomimetics based on pyrimidines, while Meisel and Hamilton reported cavitand-like molecular containers derived from 2,4-dialkoxy-meta-aminomethylbenzoic acid (MAMBA) monomers.15 Alfonso et al. synthesised a family of tris-pyridine-decorated cyclic peptides that self-assemble to carry out chloride recognition and transport.16,17 The common feature among these examples is the judicious incorporation of abiotic building blocks into peptide scaffolds to expand the chemical design space.7,12
However, few sequence-defined scaffolds integrate additional hydrogen-bonding capacity directly into the peptide backbone, even though doing so could improve anion affinity and selectivity.18–20 Here, we report a first in class family of sequence-defined macrocyclic peptidomimetics (Fig. 1(i)), termed “squaratides”. These modular receptors embed squaramide units directly within the peptidomimetic backbone while allowing programmable sidechain functionality. Our design rationale was to couple the strong, directional hydrogen-bond donating properties of squaramides with the structural preorganisation imposed by macrocyclisation, while retaining amino acid sidechains to tune the platforms chemical properties systematically. We present a general synthetic route to create squaratides and comprehensively characterise their structures and anion-binding behaviour using NMR, IR and CD spectroscopy, X-ray crystallography, and DFT calculations. We demonstrate that macrocyclisation enhances preorganisation and thus binding selectivity toward biologically relevant anions such as acetate and chloride, and that amino acid sidechain identity profoundly influences both binding affinity and stoichiometry. Together, these findings establish squaratides as a versatile new platform for programmable anion recognition.
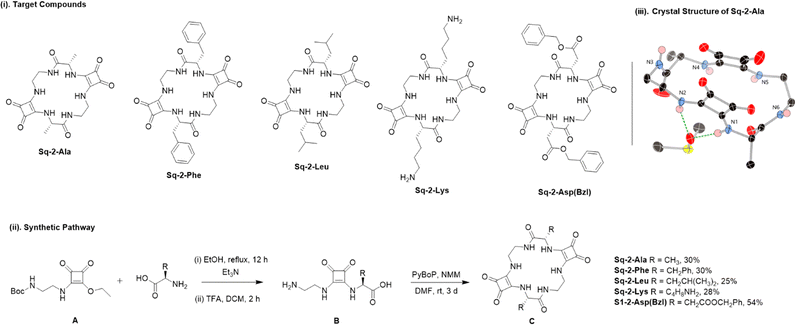 | ||
| Fig. 1 (i). Chemical structures of target cyclic squaratides. (ii) General synthetic strategy towards cyclic squaratides. (iii) Crystal structure of Sq-2-Ala. | ||
The synthesis of cyclic squaratides is outlined in Fig. 1(ii) where a modular, convergent strategy was employed in which diethyl squarate was sequentially functionalised with ethylenediamine and amino acid building blocks, followed by intramolecular macrocyclisation to afford a series of the sequence-defined peptidomimetic receptors. Briefly, reaction of diethyl squarate with N-Boc-ethylenediamine furnished the monosubstituted intermediate A, which was subsequently reacted with the desired amino acid before TFA initiated Boc deprotection yielded monomer B. Dimerisation in the presence of benzotriazol-1-yloxytripyrrolidinophosphonium hexafluorophosphate (PyBOP), afforded a series of cyclic squaratides C in yields from 25–54%.
Single crystals of Sq-2-Ala grown from dilute DMSO-d6 confirmed the cyclic structure by X-ray crystallography. The structure was solved in the chiral monoclinic space group P21 as a bis-DMSO solvate. As shown in Fig. 1(iii), the squaramide units adopt a syn conformation with both N–H donor pairs (N1, N2 and N4, N5) oriented in the same direction. Macrocycle puckering offsets the cyclobutene rings such that each functions independently as a bis-bidentate, rather than tetradentate, hydrogen-bond donor toward a single oxygen acceptor. The alanine methyl groups project towards the macrocycle periphery which suggests that side chain functionalisation may not sterically hinder anion binding.
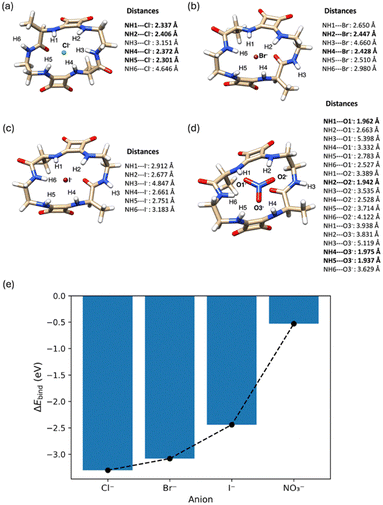
To provide a molecular-level framework for the anion-binding behaviour of the cyclic squaratide platform, DFT calculations were performed on host–guest complexes of Sq-2-Ala with Cl−, Br−, I−, and NO3−. Geometries were optimised in the gas phase from the experimental XRD structure, followed by counterpoise (CP) correction for basis set superposition error (BSSE) (see SI). The resulting BSSE-corrected binding energies (ΔEbind) enable comparison across the anion series. The calculated energies follow the order Cl− > Br− > I− ≫ NO3−, with ΔEbind values of −3.30, −3.08, −2.44, and −0.53 eV, respectively (Fig. 2e). It should be noted that these binding energies are calculated for gas-phase complexes and therefore represent an upper theoretical limit to host–guest interactions in the absence of solvent. In solution, competition with solvent molecules, particularly strongly coordinating solvents such as DMSO, is expected to weaken the effective binding interactions observed experimentally. This affinity trend reflects progressively weaker binding with increasing halide size and decreasing basicity, while NO3− binds much more weakly due to poor geometric complementarity with the preorganised hydrogen-bond donor array of Sq-2-Ala. Analysis of the optimised geometries provides further insight into this selectivity (Fig. 2a–d). In the halide complexes, Cl− and Br− form multiple short N–H⋯X− contacts, several <2.5 Å (bold in Fig. 2), consistent with strong directional hydrogen bonding. In contrast, I− shows fewer short contacts and more elongated N–H⋯I− interactions, consistent with its weaker binding energy. For NO3−, the anion remains offset from the macrocyclic cavity centre, forming only one or two short N–H⋯O contacts while the remaining donors remain uncoordinated, resulting in much weaker stabilisation. Side-view representations (Fig. S100, SI) show that halides sit close to the squaramide core of Sq-2-Ala, whereas NO3− resides significantly above the hydrogen-bond donor plane. This separation limits cooperative binding and accounts for the predicted energetic penalty for NO3− relative to the halides. Although absolute binding energies are likely overestimated in the gas phase, the BSSE-corrected calculations reproduce the experimentally relevant relative anion preference of Sq-2-Ala for monovalent anions reported below. These calculations provide a structural and energetic framework for interpreting the NMR titration data, supporting a model in which strong directional squaramide hydrogen bonding underpins chloride recognition, whereas larger or geometrically mismatched anions interact only weakly and are not selectively bound. Additional computational studies of competing solvent binding and alternative guest species, including DMSO, fluoride (with possible deprotonation pathways), and hydroxide, are underway and will be reported in future work. Guided by the structural analysis and modelling, the binding behaviour of Sq-2-Ala was investigated in DMSO-d6 by 1H NMR spectroscopy. Initial screening against eight tetrabutylammonium (TBA) salts of halides and oxoanions (F−, Cl−, Br−, I−, SO42−, AcO−, H2PO4−, NO3−) revealed varying responses as a function of anion concentration. Deprotonation occurred upon addition of F−, evidenced by a downfield triplet at ca. 16 ppm characteristic of hydrogen difluoride (HF2−) formation and consistent with fluoride-induced deprotonation of the squaramide NH groups.21 In contrast, minimal shifts were observed for I−, Br− and NO3−, indicating negligible binding consistent with previous reports of weak iodide and nitrate binding.22,23 Although gas-phase DFT calculations predict moderate Br− binding relative to Cl−, the negligible experimental binding likely reflects solvent competition and reduced hydrogen-bond directionality in solution, highlighting the tendency of gas-phase calculations to overestimate absolute binding energies.
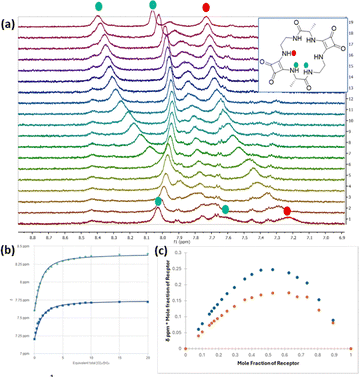
The weak responses observed for the larger halides and NO3− are consistent with the reduced directional hydrogen bonding and geometric mismatch identified in the DFT calculations. Minor perturbations were observed in the presence of SO42−, contrasting with the high sulfate affinity reported for other cyclic squaramides24 and suggesting steric or geometric constraints within the squaratide cavity that hinder optimal sulfate coordination. Upon incremental addition of AcO−, the amide NH signal shifted upfield then downfield, while the squaramide NH signals shifted downfield (Δppm ≈ 1.5 for both). This behaviour suggests a concentration-dependent process in which weak, transient interactions at low AcO− concentrations—likely involving conformational rearrangement and steric fit of the small methyl group (vdW radius ∼0.2 nm, similar to the halide series, Fig. 2e)—are followed by direct hydrogen bonding at higher anion concentrations (Fig. 3a). Upon Cl− addition, the NH signals exhibited similar but smaller shifts than for AcO− (Δppm ≈ 0.6 and 0.9). Association constants for Cl− and AcO− were determined by fitting the squaramide NH chemical shift changes as a function of anion concentration using BindFit.25
Sq-2-Ala exhibited 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry toward both anions, with higher affinity for Cl− (Ka = 692 M−1) than AcO− (Ka = 227 M−1). This chloride affinity is comparable to highly electron-deficient diaryl squaramides and suggests that macrocyclic preorganisation, rather than increased intrinsic hydrogen-bond donor strength, underpins the observed binding enhancement.26 The 1
1 binding stoichiometry toward both anions, with higher affinity for Cl− (Ka = 692 M−1) than AcO− (Ka = 227 M−1). This chloride affinity is comparable to highly electron-deficient diaryl squaramides and suggests that macrocyclic preorganisation, rather than increased intrinsic hydrogen-bond donor strength, underpins the observed binding enhancement.26 The 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding model was supported by Job-plot analyses showing maxima at 0.5 mole fraction (Fig. 3b and c) and by independent fitting using Musketeer,27 which yielded comparable association constants. Variation of side-chain functionality significantly modulates binding affinity and stoichiometry across the receptor series. For example, Sq-2-Phe displayed a 2
1 binding model was supported by Job-plot analyses showing maxima at 0.5 mole fraction (Fig. 3b and c) and by independent fitting using Musketeer,27 which yielded comparable association constants. Variation of side-chain functionality significantly modulates binding affinity and stoichiometry across the receptor series. For example, Sq-2-Phe displayed a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (host:guest) binding mode (K11 = 8.3 × 103 M−1, K21 = 3 × 103 M−1), supported by Job-plot analysis. However, deviations in the fitting residuals suggest coexisting equilibria or multiple binding geometries, and the extracted constants should be interpreted with caution. This behaviour likely arises from hydrophobic effects and π–π interactions that complicate anion binding. Nevertheless, all receptors show significant Cl− and AcO− binding under these conditions, underscoring the role of side-chain polarity and steric profile in tuning anion recognition (Table 1).
1 (host:guest) binding mode (K11 = 8.3 × 103 M−1, K21 = 3 × 103 M−1), supported by Job-plot analysis. However, deviations in the fitting residuals suggest coexisting equilibria or multiple binding geometries, and the extracted constants should be interpreted with caution. This behaviour likely arises from hydrophobic effects and π–π interactions that complicate anion binding. Nevertheless, all receptors show significant Cl− and AcO− binding under these conditions, underscoring the role of side-chain polarity and steric profile in tuning anion recognition (Table 1).
| Receptor | Cl− (M−1) | Jobs plot | AcO− (M−1) | Jobs plot |
|---|---|---|---|---|
| Sq-2-Ala | K a = 692 (±6) | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
K a = 227 (±3) M−1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| Sq-2-Phe | K a = 366 (±4) | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
K 11 = 8.3 × 103 (±69) K21 = 3 × 103 (±68) | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| Sq-2-Leu | K a = 558 (±2) | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
K 21 > 104 | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| Sq-2-Lys | K 11 = 1.8 × 103 (±3) | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
K a = 333 (±12) | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| K 21 = 703 (±9) | ||||
| Sq-2-Asp(Bzl) | K 21 > 104 | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
K 11 > 104 | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
To further investigate the complex binding behaviour observed during anion titrations, a concentration-dependent 1H NMR study of Sq-2-Phe was conducted over 0.25–20 mM (Fig. S95, SI). The 1H NMR spectra of these squaratides were previously noted to be poorly resolved in DMSO-d6 despite LCMS evidence of compound purity, suggesting that conformational heterogeneity and/or molecular self-assembly may contribute to the observed behaviour. For Sq-2-Phe, broad concentration-dependent changes were observed, particularly in the aromatic and squaramide NH regions. Increasing concentration led to (i) chemical shift changes, (ii) significant signal broadening, particularly for NH resonances, and (iii) the appearance of new signals. These features are characteristic of supramolecular self-association, consistent with intermolecular squaramide hydrogen bonding and π–π stacking of phenylalanine side chains. In contrast, while Sq-2-Ala also displayed poorly resolved 1H NMR spectra, chemical shifts were largely concentration-invariant across the same range (Fig. S96, SI), suggesting that this receptor does not engage in aggregation to the same extent but conformational isomerism (including cis–trans amide isomerism as well as anti/anti- and anti/syn- conformations of bis-secondary squaramides) is likely to be a contributor. This observation further highlights the role of amino-acid identity in governing supramolecular behaviour.
To assess the influence of macrocyclisation, the acyclic analogue ASq-2-Phe was synthesised (Scheme S2, SI). Although its 1H NMR spectra indicated conformational heterogeneity typical of acyclic peptidomimetics and foldamers,28,29 concentration-dependent studies showed minimal aggregation, with resonances remaining relatively sharp across 0.25–20 mM (Fig. S97, SI). This contrast indicates that cyclisation in Sq-2-Phe enforces preorganisation and spatial proximity between squaramide and aromatic units, promoting ordered aggregation, whereas the greater flexibility of ASq-2-Phe suppresses this behaviour. 1H NMR titration of ASq-2-Phe with TBA AcO− revealed complex, multi-site binding behaviour. Larger downfield shifts at the methyl ester terminus relative to the Boc-protected end indicate uneven binding along the flexible scaffold. Up to one equivalent of AcO−, binding occurs mainly at the squaramide units, with backbone amide NHs engaging only at higher anion loadings. No satisfactory single binding model could be fitted, underscoring the impact of reduced preorganisation. Nevertheless, although less rigid and therefore less discriminating, the acyclic squaratide dimers retain binding ability and provide a platform that, with appropriate modification, may yield useful conformationally adaptable receptor systems.
In summary, we report squaratides, a new peptidomimetic platform that incorporates squaramide hydrogen-bond donors into a peptide-inspired backbone. Side-chain functionality governs self-assembly and anion binding behaviour, tuning affinity, stoichiometry and selectivity, with preference observed for AcO− and strong Cl− binding across the series. The modularity of this scaffold makes it a promising platform for potential bioinspired therapeutic peptidomimetics and synthetic anion transport systems.
R. B. P. E., D. T., and K. A. J. conceived and designed the study. F. A. M., C. S. H, S. B., D. T., K.A. J. and R. B. P. E. wrote the manuscript. R. B. P. E. and D. T. supervised the study. F. A. M. and H. T. synthesised the compounds under study and carried out all spectroscopic titrations. C. S. H. carried out X-ray crystallographic analysis. S. B. and D. T. carried out computational analysis. All authors discussed the results and commented on the manuscript.
Conflicts of interest
There are no conflicts to declare.Data availability
All relevant experimental details, characterisation data, computational data and binding titration data are presented in the main text and supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cc00836d.CCDC 2287336 contains the supplementary crystallographic data for this paper.30
Acknowledgements
R. E. and D. T. acknowledge support from Research Ireland (RI) under award number 12/RC/2275_P2 (SSPC) which is co-funded under the European Regional Development Fund. F. M. acknowledges the award of a Hume Scholarship from Maynooth University. R. E. also acknowledges funding from RI for the funding of the NMR facility (12/RI/2346/SOF) through the Research Infrastructure Programme and the Advion Compact Mass Spec through the Opportunistic Infrastructure Fund (16/RI/3399). D. T. and S. B. acknowledge supercomputing resources at the RI/Higher Education Authority Irish Centre for High-End Computing (ICHEC).References
- N. Busschaert, C. Caltagirone, W. Van Rossom and P. A. Gale, Chem. Rev., 2015, 115, 8038–8155 CrossRef CAS PubMed.
- L. K. Macreadie, A. M. Gilchrist, D. A. McNaughton, W. G. Ryder, M. Fares and P. A. Gale, Chemistry, 2022, 8, 46–118 CrossRef CAS.
- P. A. Gale, E. N. W. Howe and X. Wu, Chemistry, 2016, 1, 351–422 CrossRef CAS.
- F. A. Mohammed, T. Xiao, L. Wang and R. B. P. Elmes, Chem. Commun., 2024, 60, 11812–11836 RSC.
- Y. Chen, S. Guerin, H. Yuan, J. O’Donnell, B. Xue, P.-A. Cazade, E. U. Haq, L. J. W. Shimon, S. Rencus-Lazar, S. A. M. Tofail, Y. Cao, D. Thompson, R. Yang and E. Gazit, J. Am. Chem. Soc., 2022, 144, 3468–3476 CrossRef CAS PubMed.
- L. Chen, S. N. Berry, X. Wu, E. N. W. Howe and P. A. Gale, Chemistry, 2020, 6, 61–141 CrossRef CAS.
- R. B. P. Elmes and K. A. Jolliffe, Chem. Commun., 2015, 51, 4951–4968 RSC.
- G. E. Sergeant and K. A. Jolliffe, Supramol. Chem., 2020, 32, 233–237 CrossRef CAS.
- J. Bartl and S. Kubik, ChemPlusChem, 2020, 85, 963–969 CrossRef CAS PubMed.
- R. B. P. Elmes, K. K. Y. Yuen and K. A. Jolliffe, Chem. – Eur. J., 2014, 20, 7373–7380 CrossRef CAS PubMed.
- P. G. Young and K. A. Jolliffe, Org. Biomol. Chem., 2012, 10, 2664–2672 RSC.
- S. Kubik, Molecules, 2022, 27, 2821 CrossRef CAS PubMed.
- S. Kubik, C. Reyheller and S. Stüwe, J. Inclusion Phenom. Macrocyclic Chem., 2005, 52, 137–187 CrossRef CAS.
- S. Kubik, Chem. Soc. Rev., 2009, 38, 585–605 RSC.
- J. W. Meisel, C. T. Hu and A. D. Hamilton, Org. Lett., 2019, 21, 7763–7767 CrossRef CAS PubMed.
- N. Rodríguez-Vázquez, M. Amorín, I. Alfonso and J. R. Granja, Angew. Chem., Int. Ed., 2016, 55, 4504–4508 CrossRef PubMed.
- A. Fuertes, M. Amorín and J. R. Granja, Chem. Commun., 2020, 56, 46–49 RSC.
- S. S. Burade, S. V. Pawar, T. Saha, N. Kumbhar, A. S. Kotmale, M. Ahmad, P. Talukdar and D. D. Dhavale, Beilstein J. Org. Chem., 2019, 15, 2419–2427 CrossRef CAS PubMed.
- V. Diemer, L. Fischer, B. Kauffmann and G. Guichard, Chem. – Eur. J., 2016, 22, 15684–15692 CrossRef CAS PubMed.
- L. Martínez-Crespo, E. C. Escudero-Adán, A. Costa and C. Rotger, Chem. – Eur. J., 2018, 24, 17802–17813 CrossRef PubMed.
- H. Tong, F. Ali Mohammed and R. B. P. Elmes, Results Chem., 2024, 11, 101777 CrossRef CAS.
- L. K. Kumawat, C. Wynne, E. Cappello, P. Fisher, L. E. Brennan, A. Strofaldi, J. J. McManus, C. S. Hawes, K. A. Jolliffe, T. Gunnlaugsson and R. B. P. Elmes, ChemPlusChem, 2021, 86, 1058–1068 CrossRef CAS PubMed.
- R. B. Elmes, P. Turner and K. A. Jolliffe, Org. Lett., 2013, 15, 5638–5641 CrossRef CAS PubMed.
- L. Qin, A. Hartley, P. Turner, R. B. P. Elmes and K. A. Jolliffe, Chem. Sci., 2016, 7, 4563–4572 RSC.
- D. Brynn Hibbert and P. Thordarson, Chem. Commun., 2016, 52, 12792–12805 RSC.
- N. Busschaert, I. L. Kirby, S. Young, S. J. Coles, P. N. Horton, M. E. Light and P. A. Gale, Angew. Chem., Int. Ed., 2012, 51, 4426–4430 CrossRef CAS PubMed.
- D. O. Soloviev and C. A. Hunter, Chem. Sci., 2024, 15, 15299–15310 RSC.
- C. Acconcia, A. Paladino, M. D. Valle, B. Farina, A. Del Gatto, S. D. Gaetano, D. Capasso, M. T. Gentile, G. Malgieri, C. Isernia, M. Saviano, R. Fattorusso, L. Zaccaro and L. Russo, Int. J. Mol. Sci., 2022, 23, 11039 CrossRef CAS PubMed.
- J. Damjanovic, J. Miao, H. Huang and Y. S. Lin, Chem. Rev., 2021, 121, 2292–2324 CrossRef CAS PubMed.
- CCDC 2287336: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2gs51h.
| This journal is © The Royal Society of Chemistry 2026 |