Open Access Article
Open Access ArticleFabrication of a self-assembling peptide for the inhibition of cancer cell proliferation by targeting LAT1
Yuki
Koba
a,
Kanta
Ogura
a,
Masahiko
Nakamoto
 *ab and
Michiya
Matsusaki
*ab and
Michiya
Matsusaki
 *a
*a
aDivision of Applied Chemistry, Graduate School of Engineering, Osaka University Suita, 2-1 Yamada-oka, Osaka 565-0871, Japan. E-mail: m-nakamoto@chem.eng.osaka-u.ac.jp; m-matsus@chem.eng.osaka-u.ac.jp
bCenter for Future Innovation (CFi), Faculty of Engineering, The University of Osaka, 2-1 Yamada-oka, Suita, Osaka 565-0871, Japan
First published on 12th March 2026
Abstract
Self-assembling peptides targeting overexpressed proteins and/or enzymes on the cancer cell membrane have emerged as promising strategies for effective and selective cancer therapy. We developed a self-assembling peptide targeting L-type amino acid transporter (LAT1), LffVLKK-4Phe, that selectively inhibits cancer cell proliferation by inhibiting LAT1. This work demonstrates a self-assembling peptide-based strategy to inhibit cancer cell proliferation via suppressing amino acid transport.
Overexpressed membrane-associated proteins/enzymes promote cancer cell survival and proliferation.1–3 Therefore, nanomedicines including polymer ligands4–8 and self-assembling peptides9–11 that lead to cancer cell death through direct multivalent interactions with these proteins/enzymes and/or enzyme-responsive membrane disruption/intracellular stress induction have emerged as promising strategies for effective and selective cancer therapy. In this context, enzyme-responsive12,13 or ligand-directed14–17 self-assembling peptides that display adaptive multivalent binding interfaces with target membrane-associated proteins/enzymes can be promising for overcoming the spatial heterogeneity of cancer cells. For example, Li et al. reported that self-assembling peptides conjugated with a carbonic anhydrase IX (CAIX)-targeting motif inhibited cancer growth through multivalent interactions arising from nanofiber formation driven by CAIX-mediated peptide accumulation on the cancer cell membrane.14 Recently, transporters have attracted increasing attention as promising targets for selective cancer interventions,18,19 since they play fundamental roles in metabolic transformation and energy homeostasis through nutrient uptake.20,21 However, to the best of our knowledge, little has been reported on self-assembling peptides that inhibit cancer cell proliferation by directly suppressing transporter-mediated substance uptake through multivalent interactions.
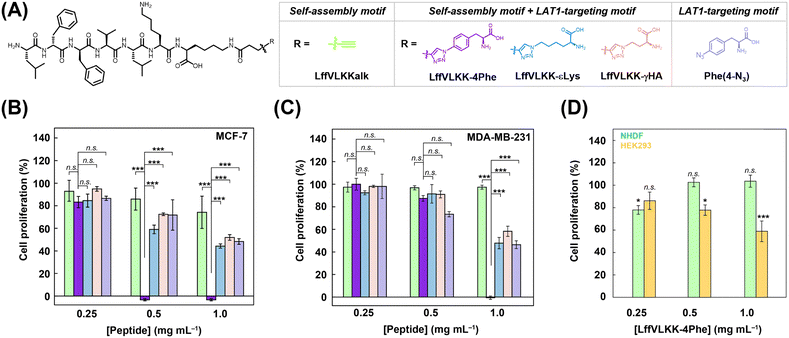
In this study, the L-type amino acid transporter (LAT1) was selected as the target protein because it plays an important role in the uptake of amino acids essential for proliferation.22,23 We designed a self-assembling peptide conjugated to the L-phenylalanine (Phe) motif as an LAT1 ligand (LffVLKK-4Phe) (Fig. 1). Here, we demonstrated that LffVLKK-4Phe inhibits cancer cell proliferation by suppressing LAT1-mediated amino acid transport. The findings of this study provide a strategy for designing self-assembling peptides that inhibit cancer cell proliferation by suppressing amino acid transport.
 | ||
| Fig. 1 (A) A Chemical structure and an illustration of a self-assembling peptide targeting LAT1, LffVLKK-4Phe. (B) Conceptual illustration of this study. | ||
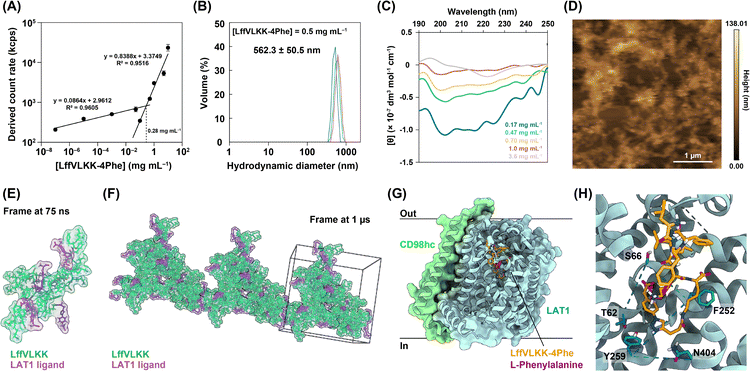
First, a self-assembling sequence24–26 (LffVLKKalk) was synthesized by solid-phase peptide synthesis (SI 2 and Fig. S1–S3). Subsequently, 4-azido-L-phenylalanine (Phe(4-N3)) was introduced as an LAT1-targeting motif by copper-catalyzed azide–alkyne cycloaddition, yielding LffVLKK-4Phe (SI 3 and Fig. S4–S11). The critical self-assembly concentration (CSC)8 of LffVLKK-4Phe in Dulbecco's phosphate-buffered saline (D-PBS, pH 6.8) was determined to be 0.28 mg mL−1 (238 µM) based on the derived count rate from dynamic light scattering (DLS) measurements27 (Fig. 2A). Furthermore, the dynamics arising from cooperative diffusion28 were observed with increasing concentrations (Fig. S12A–G). These results indicated the concentration-dependent self-assembly behaviour of LffVLKK-4Phe. The hydrodynamic diameter of LffVLKK-4Phe at 0.5 mg mL−1 was 562.3 ± 50.5 nm (Fig. 2B). Circular dichroism (CD) spectra suggested that LffVLKK-4Phe shifted toward a β-sheet-rich secondary structure upon increasing the concentration (Fig. 2C). A decrease in [θ] values at high concentrations could be because of the light scattering and the formation of large peptide assemblies29 (Fig. S14). The morphology of drop-cast, air-dried LffVLKK-4Phe (1.0 mg mL−1) was examined by atomic force microscopy (AFM), revealing that nano-fibrous and network-like structures (Fig. 2D), whereas nanoparticle-like structures were observed at lower concentrations (0.25 and 0.5 mg mL−1) (Fig. S15). The concentration-dependent self-assembly of LffVLKK-4Phe was also confirmed by fluorescence microscopy (Fig. S16). In addition, Nile red staining of the assemblies indicated that the self-assembly of LffVLKK-4Phe provided a hydrophobic environment (Fig. S17).
For further understanding, molecular dynamics (MD) simulations of peptide self-assembly were performed using GROMACS software30 (SI 5 and Fig. S18). The LAT1 ligands of the assemblies derived from the six molecules of LffVLKK-4Phe were located outside the assembly (Fig. 2E), indicating that LffVLKK-4Phe in the assembly state could multivalently interact with LAT1. After 1 µs MD simulations, the fibrous assembly of LffVLKK-4Phe was observed (Fig. 2F). Next, the binding capacity of LffVLKK-4Phe to LAT1 was investigated through docking simulations using AutoDock Vina31 (SI 5). The binding of LffVLKK-4Phe to the substrate pocket of LAT1 (PDB ID: 8kdn) was observed (Fig. 2G and H). The free energy change of LffVLKK-4Phe through binding was lower than that of Phe (LffVLKK-4Phe: −9.3 kcal mol−1; Phe: −6.3 kcal mol−1, respectively), suggesting the LAT1-targeting capability of LffVLKK-4Phe. Taken together, experimental results and simulations suggest that the self-assembly of LffVLKK-4Phe could be promoted by increasing its local concentration on the cancer cell membrane in response to the overexpression of LAT1. This assembly is expected to provide a large binding surface on the cancer cell membrane.
Next, we investigated the proliferative inhibitory effect of LffVLKK-4Phe on human breast cancer cell line, MCF-7, overexpressing LAT1,32 under hypoxic conditions at pH 6.833 (SI 6). To explore the effect of the LAT1-targeting self-assembly, LffVLKKalk as a self-assembly motif alone; LffVLKK-4Phe, LffVLKK-εLys, and LffVLKK-γHA as LAT1-targeted peptides; and Phe(4-N3) as the LAT1 ligand alone were tested (Fig. 3A and Fig. S19). Firstly, to understand the importance of LAT1 targeting, the proliferative inhibitory effect of LffVLKK-4Phe and LffVLKKalk was compared. LffVLKK-4Phe showed a considerably high proliferative inhibitory effect at 0.5 and 1.0 mg mL−1 after treatment for 72 h (Fig. 3B). In contrast, LffVLKKalk showed marginal efficacy at all tested concentrations (Fig. 3B) despite its comparable CSC (0.43 mg mL−1) with that of LffVLKK-4Phe (Fig. S13). These results suggest that LAT1 targeting was required to inhibit MCF-7 cell proliferation. Next, to reveal the suitable structure as an LAT1 ligand, the proliferative inhibitory effect of LffVLKK-4Phe, LffVLKK-εLys, and LffVLKK-γHA were assessed. LffVLKK-4Phe exhibited significantly higher proliferative inhibition than LffVLKK-εLys and LffVLKK-γHA (Fig. 3B). In addition, the CSC of LffVLKK-4Phe in PBS was lower than those of LffVLKK-γHA and LffVLKK-εLys (0.34 mg mL−1 and not determined, respectively) (Fig. S13). These results suggest that the Phe motif, which is favorable for promoting self-assembly and interacting with LAT1 would be important for the effective inhibition. To understand the importance of the assembly-induced ligand multivalency, the proliferative inhibitory effect of LffVLKK-4Phe and Phe(4-N3) were evaluated. The proliferative inhibitory effect of LffVLKK-4Phe was significantly greater than that of Phe(4-N3) (Fig. 3B), indicating that the inhibitory effect of the LAT1 ligand could be enhanced by an increase in multivalency via self-assembly. Moreover, the proliferative inhibitory effect of LffVLKK-4Phe against LAT1-positive human breast cancer cell line, MDA-MB-231, was also explored, and a similar trend to that observed in MCF-7 cells was confirmed (Fig. 3C). LffVLKK-4Phe exhibited higher efficacy in MCF-7 cells than in MDA-MB-231 cells (Fig. 3C). This may be because of differences in LAT1 expression, amino acid uptake, and/or phenotype. Indeed, it has been reported that mRNA levels of LAT1 are higher in MCF-7 cells than MDA-MB-231 cells.32 In addition, we confirmed that MCF-7 cells exhibited greater amino acid uptake than MDA-MB-231 cells (Fig. S20). Cytotoxicity in healthy cells was further evaluated using normal human dermal fibroblasts, NHDF cells, and human embryonic kidney 293, HEK293 cells, under normoxic conditions at pH 7.4. LffVLKK-4Phe did not affect cell proliferation of NHDF at any concentrations tested (Fig. 3D). Although HEK293 cell proliferation was reduced to approximately 60% at 1.0 mg mL−1 (Fig. 3D), cancer cell proliferation was almost completely inhibited (Fig. 3B and C). These results suggest a high selectivity LffVLKK-4Phe for cancer cells.
To elucidate the inhibitory mechanism of LffVLKK-4Phe on MCF-7 cell proliferation, amino acid uptake assays using boronophenylalanine (BPA) and lactate dehydrogenase (LDH) cytotoxicity assays were performed. At the terminal state, disrupted cell morphology was observed at 0.5 and 1.0 mg mL−1 (Fig. S19), implying that LffVLKK-4Phe induced cell membrane disruption in MCF-7 cells at the later stage. To gain mechanistic insights, early-stage cellular events (30 min) in MCF-7 cells were examined because the terminal stage (72 h), which is associated with complete cell death, suppresses amino acid uptake and leads to extensive LDH release. The uptake of BPA via LAT1 in MCF-7 cells was significantly inhibited after 30 min of treatment with LffVLKK-4Phe in a concentration-dependent manner (Fig. 4A), suggesting that LffVLKK-4Phe suppressed LAT1-mediated amino acid transport activity at an early stage of MCF-7 proliferative inhibition. At the same time point, slight increased LDH release was observed from MCF-7 cells treated with LffVLKK-4Phe (Fig. 4B). These results suggest that at an early stage, the proliferative inhibition of MCF-7 cells is predominantly caused by LAT1 inhibition, whereas membrane disruption occurs as an accompanying effect. This could be due to the self-assembly of LffVLKK-4Phe in response to the overexpression of LAT1 in the MCF-7 cell membrane.
In conclusion, we developed a self-assembling peptide targeting LAT1, LffVLKK-4Phe, which selectively inhibited the proliferation of MCF-7 and MDA-MB-231 cells. These results indicated that LffVLKK-4Phe inhibited cancer cell proliferation by suppressing LAT1-mediated amino acid transport, accompanied by cell membrane disruption. The findings of this study provide a strategy for designing self-assembling peptides that can inhibit cancer cell proliferation by adapting to the spatial heterogeneity of cancer cell membranes.
Author contributions
Y. K. and M. N. designed the study, the main conceptual ideas, and the proof outline. Y. K. and M. N. designed the experiments. Y. K. and K. O. collected data. Y. K. and M. N. interpreted the data. Y. K. and M. N. wrote the manuscript. M. M. and M. N. supervised this study. All the authors discussed the results and commented on the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article are included in the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cc00782a.Acknowledgements
This work was supported by Grant-in-Aid for JSPS Fellows (25KJ1746), JST FOREST (JPMJFR232E), and JSPS Scientific Research for Transformative Research Areas (A) “Bottom-up Biotech” (JP 24H01137). We appreciate Dr Zhang Zhuying in The University of Osaka for AFM observations.References
- R. A. Cairns, I. S. Harris and T. W. Mak, Nat. Rev. Cancer, 2011, 11, 85 CrossRef CAS PubMed.
- G. D. Grass, L. B. Tolliver, M. Bratoeva and B. P. Toole, J. Biol. Chem., 2013, 288, 26089 CrossRef CAS PubMed.
- H. M. Becker, Br. J. Cancer, 2020, 122, 157 CrossRef CAS PubMed.
- M. Morales-Cruz, Y. Delgado, B. Castillo, C. M. Figueroa, A. M. Molina, A. Torres, M. Milián and K. Griebenow, Drug Des., Dev. Ther., 2019, 13, 3753 CrossRef CAS PubMed.
- P. Šácha, T. Knedlík, J. Schimer, J. Tykvart, J. Parolek, V. Navrátil, P. Dvořáková, F. Sedlák, K. Ulbrich, J. Strohalm, P. Majer, V. Šubr and J. Konvalinka, Angew. Chem., Int. Ed., 2016, 55, 2356 CrossRef PubMed.
- Y. Koba, M. Nakamoto and M. Matsusaki, ACS Appl. Mater. Interfaces, 2022, 14, 51790 CrossRef CAS PubMed.
- Y. Koba, M. Nakamoto, M. Nagao, Y. Miura and M. Matsusaki, Nano Lett., 2024, 24, 14206 CrossRef CAS PubMed.
- R. Sakamoto, Y. Koba, M. Nakamoto, T. Fukuta, K. Kadota and M. Matsusaki, ACS Appl. Mater. Interfaces, 2025, 17, 59726 CrossRef CAS PubMed.
- Y. Kuang, J. Shi, J. Li, D. Yuan, K. A. Alberti, Q. Xu and B. Xu, Angew. Chem., Int. Ed., 2014, 53, 8104 CrossRef CAS PubMed.
- J. Shi, X. Du, D. Yuan, J. Zhou, N. Zhou, Y. Huang and B. Xu, Biomacromolecules, 2014, 15, 3559 CrossRef CAS PubMed.
- K. Morita, K. Nishimura, S. Yamamoto, N. Shimizu, T. Yashiro, R. Kawabata, T. Aoi, A. Tamura and T. Maruyama, JACS Au, 2022, 2, 2023 CrossRef CAS PubMed.
- J. Zhan, Y. Cai, S. He, L. Wang and Z. Yang, Angew. Chem., Int. Ed., 2018, 57, 1813 CrossRef CAS PubMed.
- Y. Ding, D. Zheng, L. Xie, X. Zhang, Z. Zhang, L. Wang, Z.-W. Hu and Z. Yang, J. Am. Chem. Soc., 2023, 145, 4366 CrossRef CAS PubMed.
- J. Li, K. Shi, Z. F. Sabet, W. Fu, H. Zhou, S. Xu, T. Liu, M. You, M. Cao, M. Xu, X. Cui, B. Hu, Y. Liu and C. Chen, Sci. Adv., 2019, 5, eaax0937 CrossRef CAS PubMed.
- J. Li, Y. Fang, Y. Zhang, H. Wang, Z. Yang and D. Ding, Adv. Mater., 2021, 33, 2008518 CrossRef CAS PubMed.
- M.-D. Wang, G.-T. Lv, H.-W. An, N.-Y. Zhang and H. Wang, Angew. Chem., Int. Ed., 2022, 61, e202113649 CrossRef CAS PubMed.
- H. Yin, Y. Hua, S. Feng, Y. Xu, Y. Ding, S. Liu, D. Chen, F. Du, G. Liang, W. Zhan and Y. Shen, Adv. Mater., 2024, 36, 2308504 CrossRef CAS PubMed . (ligand for ALX, conc.).
- X. Wu, J. Shen, X. Jiang, H. Han, Z. Li, Y. Xiang, D. Yuan and J. Shi, Chem. Eng. J., 2024, 493, 152479 CrossRef CAS.
- Y. Lee, C. Jin, R. Ohgaki, M. Xu, S. Ogasawara, R. Warshamanage, K. Yamashita, G. Murshudov, O. Nureki, T. Murata and Y. Kanai, Nat. Commun., 2025, 16, 1635 CrossRef CAS PubMed.
- Z. C. Nwosu, M. G. Song, M. P. di Magliano, C. A. Lyssiotis and S. E. Kim, Oncogene, 2023, 42, 711 CrossRef CAS PubMed.
- R. Xia, H.-F. Peng, X. Zhang and H.-S. Zhang, Int. J. Biol. Macromol., 2024, 260, 129646 CrossRef CAS PubMed.
- Y. Kanai, H. Segawa, K. Miyamoto, H. Uchino, E. Takeda and H. Endou, J. Biol. Chem., 1998, 273, 23629 CrossRef CAS PubMed.
- P. Häfliger and R.-P. Charles, Int. J. Mol. Sci., 2019, 20, 2428 CrossRef PubMed.
- D. Iglesias, M. Melle-Franco, M. Kurbasic, M. Melchionna, M. Abrami, M. Grassi, M. Prato and S. Marchesan, ACS Nano, 2018, 12, 5530 CrossRef CAS PubMed.
- G. Chen, T. Xu, Y. Yan, Y. Zhou, Y. Jiang, K. Melcher and H. E. Xu, Acta Pharmacol. Sin., 2017, 38, 1205 CrossRef CAS PubMed.
- S. Marchesan, L. Waddington, C. D. Easton, D. A. Winkler, L. Goodall, J. Forsythe and P. G. Hartley, Nanoscale, 2012, 4, 6752 RSC.
- K. Moroishi, M. Nakamoto, S. Fujita, M. Kawahara, R. Katayama and M. Matsusaki, Mater. Today Bio, 2026, 37, 102752 CrossRef CAS PubMed.
- T. Kureha, H. Minato, D. Suzuki, K. Urayama and M. Shibayama, Soft Matter, 2019, 15, 5390 RSC.
- J. Kardos, M. P. Nyiri, É. Moussong, F. Wien, T. Molnár, N. Murvai, V. Tóth, H. Vadászi, J. Kun, F. Jamme and A. Micsonai, Protein Sci., 2025, 34, e70066 CrossRef CAS PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19 CrossRef.
- O. Trott and A. J. Olson, J. Comput. Chem., 2010, 31, 455 CrossRef CAS PubMed.
- D. B. Shennan, J. Thomson, I. F. Gow, M. T. Travers and M. C. Barber, Biochim. Biophys. Acta, Biomembr., 2004, 1664, 206 CrossRef CAS PubMed.
- O. Warburg, F. Wind and E. Negelein, J. Gen. Physiol., 1927, 8, 519 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |