Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Structural domain plasticity drives Zn(II) distribution and buffering in metallothionein-3
Manuel David
Peris-Díaz
 ab,
Eduard H. T. M.
Ebberink
a,
Jan
Fiala
ab,
Eduard H. T. M.
Ebberink
a,
Jan
Fiala
 a,
Albert J. R.
Heck
a,
Albert J. R.
Heck
 a and
Artur
Krężel
a and
Artur
Krężel
 *b
*b
aBiomolecular Mass Spectrometry and Proteomics, Bijvoet Center for Biomolecular Research and Utrecht Institute for Pharmaceutical Sciences, Utrecht University, Utrecht 3584 CH, The Netherlands
bDepartment of Chemical Biology, Faculty of Biotechnology, University of Wroclaw, F. Joliot-Curie 14a, 50-383, Wrocław, Poland. E-mail: artur.krezel@uwr.edu.pl
First published on 3rd March 2026
Abstract
Complementary mass spectrometry-based approaches and molecular dynamics simulations were integrated to map Zn(II) distribution in partially metalated Zn4–6-metallothionein-3 species relevant to zinc metabolism. Sequential Zn(II) dissociation engages both the α- and β-domains through a structurally plastic, interdomain mechanism that underpins regulatory zinc buffering at physiological pZn.
In mammals, the metallothionein (MT) protein family comprises four major isoforms (MT1–MT4) and multiple MT1 sub-isoforms, which differ in metal-binding preferences, tissue distribution, and cellular localization.1 While MT1 and MT2 are ubiquitously expressed across all tissues, MT3 is primarily found in the brain. However, many reports suggest a critical role for MT3 in other tissues as well.2–4 In the central nervous system (CNS), MT3 regulates synaptic zinc signaling and protects against neurodegeneration, swapping Zn(II) ions for excess Cu(II)/Cu(I).5,6 Recent studies have expanded MT3's functional relevance beyond the CNS, for instance, Mo et al., showed that MT3 is the dominant MT isoform expressed in osteoclasts, and plays a vital role in osteoclastogenesis and bone metabolism by regulating intracellular Zn(II) concentrations.7 Salvagio et al. showed that MT3 was a source of zinc spikes in neurons, acting as a regulatory signal during neuronal development,8 and Tran et al. demonstrated a critical role for MT3 in Zn(II) homeostasis underlying protective immune functions.9 Yet the molecular basis of how MTs donate/accept Zn(II) remains poorly understood.1 This is largely because MTs are small (∼6–7 kDa), cysteine-rich proteins lacking stable secondary structure and aromatic residues, whereas Zn(II) is spectroscopically silent. Consequently, only one X-ray structure of mammalian MT (mixed complex of rat Cd5Zn2MT2) has been solved to date.10 In that structure, the protein folds into a dumbbell-shaped structure, with two metal/sulfur clusters located in two α and β domains.11 In the presence of soft or medium soft divalent metal ions (M(II)), the 20 cysteine residues (21 in MT1b) can assemble metal/sulfur clusters within the β- and α-domains resulting in M3Cys9 and M4Cys11 cores, respectively, when fully saturated.12–14 However, at physiological labile Zn(II) concentrations ([Zn(II)]labile) MT is present as partially metalated species (Zn4–6MT) due to differences in zinc sites’ affinity.1,15 The unsaturated nature of those species allows them to function as buffer molecules, as shown for MT2, keeping labile Zn(II) at the right pZn level (pZn = -log[Zn(II)]labile), and act as Zn(II) donors and acceptors when needed.15–19 How Zn(II) is distributed within MT3 across these Zn4–6MT3 species, and how this distribution affects properties, remains unknown and is a focus of this study.
MT3 also differs functionally from other MTs, as unlike MT1-2, MT3 is generally not strongly metal-inducible.1 Nevertheless, recent studies suggest MT3 is highly upregulated during osteoclast differentiation and is crucial for Zn(II) buffering during osteoclastogenesis.7,20 Structurally, MT3 is the only isoform with two β-domain prolines in the T5CPCP9 motif, and an acidic hexapeptide loop in the α-domain. Despite these differences, isothermal titration calorimetry (ITC) measurements of Zn(II) binding to MT3 revealed three thermodynamically distinct binding populations, similar to those observed for MT2.21
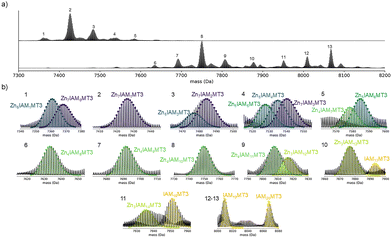
To investigate the structural and dynamical response of MT3 to Zn(II) binding and to define the microscopic organization of these partially Zn(II)-loaded MT3 species, we combined chemical labeling mass spectrometry (MS), top-down (TD)-MS, ion mobility-MS, and molecular dynamics (MD) simulations. The number of cysteine residues coordinating Zn(II) across the Zn1–7MT3 series was determined using a differential thiol-labeling strategy (Fig. S1). In the first step, iodoacetamide (IAM; +57.0215 Da per modified cysteine residue) was used to induce stepwise Zn(II) dissociation from Zn7MT3, and the resulting Zn7−xIAMyMT3 species were measured by native MS.22 IAM alkylates cysteine residues via an SN2 mechanism, forming S-carbamidomethyl cysteine adducts and thereby increasing the protein mass by +57.0215 Da per modified cysteine residue. As this reaction requires the availability of nucleophilic thiolates, the weakest Zn(II)-thiolate sites are expected to react first.23 To annotate the Zn(II):IAM stoichiometries, theoretical isotopic distributions for candidate compositions were simulated and compared with the experimental spectra, and the best-fitting model was selected (Fig. 1).
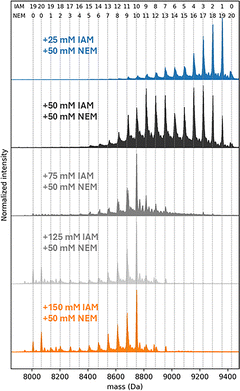
The deconvolved mass distribution of Zn7MT3 incubated with 25–50 mM IAM showed that up to two IAM molecules react without Zn(II) dissociation, suggesting that up to two cysteine residues are free or only weakly coordinated to Zn(II) (Fig. 1, peak 2–3). We therefore verified the number of Zn(II)-coordinating cysteine residues by chelating the remaining bound Zn(II) and labeling the newly accessible thiols with N-ethylmaleimide (NEM), which reacts with cysteine residues via Michael addition to form thioether adducts (+125.0477 Da per modified Cys). This provides a readout of the cysteine residues previously bound to Zn(II). These experiments confirm that 19 cysteine residues selectively coordinate all seven Zn(II) ions and that one cysteine is only weakly involved in Zn(II) coordination (Fig. 2).
Next, we aimed to identify the IAM-labeled cysteine residues in Zn7MT3 using native top-down MS. However, overlapping ions between Zn6IAM3MT35+ and Zn7IAM2MT35+ prevented quadrupole isolation and fragmentation of the latter species (Fig. 1b, peak 3). Instead, Zn7IAM1MT35+ ions were quadrupole-selected and subjected to collision-induced dissociation (CID). The resulting spectra showed limited backbone fragmentation, preventing localization of the IAM modification, most likely because Zn(II) shields the backbone from cleavage (Fig. S2). Only terminal b- and high-mass y-ions were observed, neither of which provided site-localization information.
We therefore performed TD CID-MS on the doubly labeled NEM19IAM2MT35+ ions, with IAM reporting on free cysteine residues and NEM identifying cysteines that had previously coordinated Zn(II). Specifically, quadrupole-isolated NEM19IAM2MT35+ ions were subjected to collisional activation (CA), generating b-fragment ions b6–b11 that show Cys6 and Cys8 are IAM-labeled. The y-fragment ions y60–y66 further corroborated this assignment (Fig. S3).
The third and fourth IAM modifications triggered Zn(II) dissociation, yielding Zn6IAM3MT3 and Zn6IAM4MT3 species (Fig. 1, peak no. 3–4). After dual labeling, we estimated that 16–17 cysteine residues predominantly coordinate the six Zn(II) ions (Fig. 2). When NEM16IAM4MT35+ ions were quadrupole-isolated and analyzed by TD CID-MS, the resulting spectra contained b-fragment ions b6–b10, consistent with IAM modification of Cys6 and Cys8, together with y-fragment ions y55–y66 that confirm this assignment. Although we could not unambiguously locate the remaining two Cys-IAM modifications, the overall fragmentation patterns hint that the seventh Zn(II) ion is bound in the β-domain (Fig. S4). Because the structural resolution was limited, we complemented these MS experiments with steered molecular dynamics (SMD) simulations. SMD was used to rank Zn(II) binding sites according to the work required for metal dissociation, thereby inferring plausible partially Zn(II)-loaded MT3 forms.
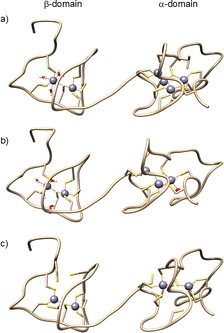
Consistent with the TD CID-MS results, SMD identified the weakest Zn(II) site as a β-domain site in which Zn(II) is coordinated by Cys6, Cys8, Cys22, and Cys25 (Fig. 3a). The resulting Zn6MT3 structure features a Zn2Cys6 cluster in the β-domain, with three cysteines completely free and solvent molecules entering the coordination sphere (Fig. S5).
Subsequent IAM modification led to dissociation of a second Zn(II) ion, forming Zn5IAM5–6MT3 species (Fig. 1b peak 4–5). Dual-labeling experiments determined that 14–15 cysteine residues coordinate the five Zn(II) ions (Fig. 2). TD CID–MS of NEM14IAM6MT35+ showed multiple C-terminal y-fragment ions that were IAM-labeled, indicating Zn(II) dissociation from the α-domain (Fig. S6). Moreover, y-fragment ions y52–y66 supported that Cys6 and Cys8 were also IAM-labeled. In line with this, SMD simulations identified Zn(II) dissociation from the α-domain, yielding a structure with a Zn3Cys9 cluster in the α-domain and a Zn2Cys6 cluster in the β-domain (Fig. 3b). Dissociation of the third Zn(II) ion generated Zn4IAM8–10MT35+ ions (Fig. 1, peak no. 6–9). Conventional top-down MS using CID did not provide any new additional fragmentation, either when comparing the singly labeled Zn4IAM8–10MT3 and Zn5IAM5–6MT3 species or when comparing the doubly labeled species NEM14IAM6MT3 and NEM10–12IAM10–8MT3. Electron-based fragmentation methods have previously been shown to enable metal-binding site localization in several metalloproteins.24 We therefore employed top-down electron transfer dissociation (ETD) and electron transfer/higher-energy collisional dissociation (EThcD), but only nondissociative electron transfer dissociation (ETnoD) was observed, leading to electron–capture charge reduction (Fig. S7). To improve fragmentation and site localization, we applied top-down ultraviolet photodissociation (UVPD) on quadrupole-isolated NEM10IAM10MT35+ ions, which yielded extensive protein backbone fragmentation and sequence coverage (Fig. S8).
The resulting fragmentation map shows that up to 3 cysteines are IAM-modified and 6 are NEM-modified in the β-domain, consistent with SMD simulations, which depict a Zn4MT3 structure in which the β-domain contains a Zn2Cys6 cluster and 3 free cysteine residues. TD-UVPD and SMD results point toward a configuration in which Zn(II) is redistributed between the α- and β-domains (Fig. 3c).
To assess whether this redistribution is accompanied by global conformational changes, we examined partially Zn(II)-loaded MT3 species by native travelling-wave ion mobility MS (TWIM-MS). Zn7MT3 shows a collision cross-section (TWCCSN2) distribution dominated by a single conformer centered at ∼1071 Å2 (Fig. S9). Zn(II) dissociation from Zn7MT3 to form Zn4MT3 does not induce a major shift to a more extended conformation, only a low-intensity conformer appears centred at ∼1180 Å2 (Fig. S9).
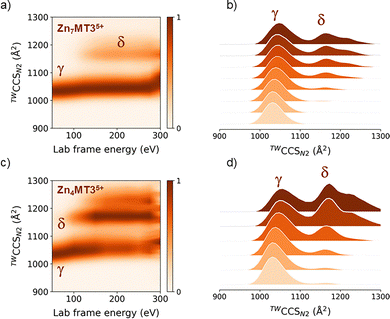
To probe conformational families and unfolding pathways, we applied collisional activation to quadrupole-selected ions and recorded TWIM-MS spectra (Fig. 4). CA of the ∼1071 Å2γ conformer leads to a minor δ conformer at ∼1180 Å2 (Fig. 4a).
The same conformer is also observed for quadrupole-selected Zn4MT35+ ions and becomes the predominant conformation upon activating conditions. Together, these data indicate that the ∼1180 Å2 conformer is present in all Zn(II)-loaded MT3 forms and is therefore not uniquely determined by metal content. While TWIM-MS reports on gas-phase conformational families and does not directly reflect in solution-phase structures, the broadly similar unfolding behavior observed for Zn7MT3 and Zn4MT3 suggests comparable global stability features. In combination with TD-UVPD and SMD results, this supports a model in which four Zn(II) ions are coordinated across both domains, since a configuration with one completely metal-free domain would be expected to induce a more pronounced conformational shift. Therefore, both the α- and β-domains appear to be required features of metallothioneins for regulatory Zn(II) buffering.
In our previous study, Zn7MT2 was observed to undergo complete unfolding to a highly extended conformation at ∼100 eV.23 By contrast, Zn7MT3 exhibits only partial unfolding, with the compact ∼1071 Å2γ conformer dominating the conformational landscape. This indicates that Zn7TM3 adopts a more stable conformation than Zn7MT2. This observation agrees with previous studies showing an enhanced resistance of MT3 to proteolysis, suggesting that MT3 is stabilized in a thermodynamically favorable conformation that may be relevant to its specific biological function.25–28 Despite this increased structural stability, MT3 binds Zn(II) more weakly than MT2.29–33 Experiments with EDTA demonstrate that Zn7MT3 has a higher metal-transfer potential than Zn7MT2, indicating that resistance to unfolding is not directly correlated with Zn(II)-binding affinity, but instead confers MT3 with unique functional properties, which might be connected to its intra- and extracellular functions.24,33 For example, studies using Zn(II)-reconstituted domain peptides have shown that S-nitrosothiols dissociate Zn(II) from both the α- and β-domains of MT3, but not from MT2.34 Consistently, cell-based studies demonstrate that MT3 provides significantly stronger protection against S-nitrosothiols, as it is the only metallothionein isoform containing consensus acid–base sequence motifs for S-nitrosylation in both domains. Functional studies further indicate that the N-terminal β-domain is necessary for growth-inhibition activity, whereas the C-terminal α-domain may be dispensable for GIF activity.35,36
Mutational studies identified two conserved prolines and a threonine within a distinctive TCPCP motif in the N-terminal β domain of MT3 as essential for GIF activity.29,35,37 This TCPCP motif, which is absent in other metallothionein isoforms, contributes to the unique structural and metal-binding properties of MT3. Through its β-domain, MT3 protects against copper toxicity by scavenging free Cu(II) ions or removing Cu(II) bound to β-amyloid and α-synuclein.38–40 The enhanced structural plasticity of the β-domain likely underpins these functions. Altogether, MT3 is a unique biologically active neuronal growth-inhibitory factor.
Overall, our study provides new insights into the architecture, stability, and dynamics of Zn(II)-MT3 and links these features to its metal buffering properties, underscoring its key role in regulating zinc and copper levels in the human body.
This research was supported by the National Science Centre of Poland (NCN) under the Opus grant no. 2021/43/B/NZ1/02961.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: the mass spectrometry data have been deposited in the Figshare repository (https://doi.org/10.6084/m9.figshare.31436293). See DOI: https://doi.org/10.1039/d6cc00567e.References
- A. Krężel and W. Maret, Chem. Rev., 2021, 121, 14594–14648 Search PubMed.
- Y. Uchida, K. Takio, K. Titani, Y. Ihara and M. Tomonaga, Neuron, 1991, 7, 337–347 CrossRef CAS PubMed.
- M. Vašák and G. Meloni, Int. J. Mol. Sci., 2017, 18, 1117 CrossRef PubMed.
- J. Y. Koh and S. J. Lee, Mol. Brain, 2020, 13, 116 CrossRef CAS PubMed.
- G. Meloni, V. Sonois, T. Delaine, L. Guilloreau, A. Gillet, J. Teissié, P. Faller and M. Vašák, Nat. Chem. Biol., 2008, 4, 366–372 CrossRef CAS PubMed.
- N. E. Wezynfeld, E. Stefaniak, K. Stachucy, A. Drozd, D. Płonka, S. C. Drew, A. Krężel and W. Bal, Angew. Chem., Int. Ed., 2016, 55, 8235–8238 Search PubMed.
- S. Mo, M. K. Kim, J. S. Jang, S. H. Lee, S. J. Hong, S. Jung and H. H. Kim, Exp. Mol. Med., 2024, 56, 1791–1806 Search PubMed.
- L. Salvagio, C. Zhang, B. E. Rue, N. Doris, C. Koehring, I. Tyler, R. S. Vargas, W. C. Oh and Y. Qin, J. Neurosci., 2025, 45, e0627252025 Search PubMed.
- N. B. Tran and S. J. Lee, Cell Death Discovery, 2025, 11, 45 Search PubMed.
- A. H. Robbins, D. E. McRee, M. Williamson, S. A. Collett, N. H. Xuong, W. F. Furey, B. C. Wang and C. D. Stout, J. Mol. Biol., 1991, 221, 1269–1293 CAS.
- W. Braun, M. Vašák, A. H. Robbins, C. D. Stout, G. Wagner, J. H. Kägi and K. Wüthrich, Proc. Natl. Acad. Sci. U. S. A., 1992, 89, 10124–10128 Search PubMed.
- D. W. Hasler, P. Faller and M. Vašák, Biochemistry, 1998, 37, 14966–14973 CrossRef CAS PubMed.
- P. Faller, D. W. Hasler, O. Zerbe, S. Klauser, D. R. Winge and M. Vašák, Biochemistry, 1999, 38, 10158–10167 CrossRef CAS PubMed.
- H. Wang, Q. Zhang, B. Cai, H. Li, K. H. Sze, Z. X. Huang, H. M. Wu and H. Sun, FEBS Lett., 2006, 580, 795–800 CrossRef CAS PubMed.
- A. Krężel and W. Maret, J. Am. Chem. Soc., 2007, 129, 10911–10921 CrossRef PubMed.
- A. Krężel and W. Maret, J. Biol. Inorg. Chem., 2006, 11, 1049–1062 CrossRef PubMed.
- K. Mosna, K. Jurczak and A. Krężel, Metallomics, 2023, 15, mfad061 CrossRef CAS PubMed.
- A. Krężel and W. Maret, J. Biol. Inorg. Chem., 2008, 13, 401–409 CrossRef PubMed.
- M. D. Peris-Díaz, R. Guran, C. Domene, V. de Los Rios, O. Zitka, V. Adam and A. Krężel, J. Am. Chem. Soc., 2021, 143, 16486–16501 CrossRef PubMed.
- S. Arisumi, T. Fujiwara, K. Yasumoto, T. Tsutsui, H. Saiwai, K. Kobayakawa, S. Okada, H. Zhao and Y. Nakashima, Cell Death Discovery, 2023, 9, 436 CrossRef CAS PubMed.
- M. C. Carpenter, A. Shami Shah, S. DeSilva, A. Gleaton, A. Su, B. Goundie, M. L. Croteau, M. J. Stevenson, D. E. Wilcox and R. N. Austin, Metallomics, 2016, 8, 605–617 CrossRef CAS PubMed.
- S. Tamara, M. A. den Boer and A. J. R. Heck, Chem. Rev., 2022, 122, 7269–7326 CrossRef CAS PubMed.
- M. D. Peris-Díaz, A. Barkhanskiy, E. Liggett, P. Barran and A. Krężel, Chem. Commun., 2023, 59, 4471–4474 RSC.
- P. Wongkongkathep, J. Y. Han, T. S. Choi, S. Yin, H. I. Kim and J. A. Loo, J. Am. Soc. Mass Spetrom., 2018, 29, 1870–1880 CrossRef CAS PubMed.
- R. Lakha, C. Hachicho, M. R. Mehlenbacher, D. E. Wilcox, R. N. Austin and C. L. Vizcarra, J. Inorg. Biochem., 2023, 242, 112157 CrossRef CAS PubMed.
- M. Knipp, G. Meloni, B. Roschitzki and M. Vašák, Biochemistry, 2005, 44, 3159–3165 CrossRef CAS PubMed.
- J. S. Calvo, R. L. E. Villones, N. J. York, E. Stefaniak, G. E. Hamilton, A. L. Stelling, W. Bal, B. S. Pierce and G. Meloni, J. Am. Chem. Soc., 2022, 144, 709–722 CrossRef CAS PubMed.
- A. T. Yuan and M. Stillman, FEBS J., 2025, 292, 619–634 CrossRef CAS PubMed.
- D. W. Hasler, L. T. Jensen, O. Zerbe, D. R. Winge and M. Vašák, Biochemistry, 2000, 39, 14567–14575 CrossRef CAS PubMed.
- M. C. Carpenter, A. Shami Shah, S. DeSilva, A. Gleaton, A. Su, B. Goundie, M. L. Croteau, M. J. Stevenson, D. E. Wilcox and R. N. Austin, Metallomics, 2016, 8, 605–617 CrossRef CAS PubMed.
- C. F. Quinn and D. E. Wilcox, Metallomics, 2024, 16, mfae041 CrossRef CAS PubMed.
- J. S. Calvo, V. M. Lopez and G. Meloni, Metallomics, 2018, 10, 1777–1791 CrossRef CAS PubMed.
- P. Palumaa, E. Eriste, O. Njunkova, L. Pokras, H. Jörnvall and R. Sillard, Biochemistry, 2002, 41, 6158–6163 Search PubMed.
- Y. Chen, Y. Irie, W. M. Keung and W. Maret, Biochemistry, 2002, 41, 8360–8367 Search PubMed.
- A. K. Sewell, L. J. Jensen, J. C. Erickson, R. D. Palmiter and D. R. Winge, Biochemistry, 1995, 34, 4740–4747 CrossRef CAS PubMed.
- S. Somji, S. H. Garrett, M. A. Sens and D. A. Sens, Toxicol. Sci, 2006, 90, 369–376 CrossRef CAS PubMed.
- B. Cai, Q. Zheng, X. C. Teng, D. Chen, Y. Wang, K. Q. Wang, G. M. Zhou, Y. Xie, M. J. Zhang, H. Z. Sun and Z. X. Huang, J. Biol. Inorg. Chem., 2006, 11, 476–482 CrossRef CAS PubMed.
- G. Meloni and M. Vašák, Free Radical Biol. Med., 2011, 50, 1471–1479 CrossRef CAS PubMed.
- G. Meloni, P. Faller and M. Vašák, J. Biol. Chem., 2007, 282, 16068–16078 CrossRef CAS PubMed.
- A. Melenbacher and M. J. Stillman, FEBS J., 2023, 290, 4316–4341 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |