Open Access Article
Open Access ArticleA meso-acyl-BODIPY-based fluorogenic probe for the selective monitoring of aldo–keto reductase 1C1 in living cells
Yeri Kim†
a,
Le Bich Hang Pham†b,
Jae Yeol Leea,
Jeeyeon Lee *b and
Youngmi Kim
*b and
Youngmi Kim *a
*a
aDepartment of Chemistry and Research Institute of Basic Sciences, Kyung Hee University, 26 Kyungheedae-ro, Dongdaemun-gu, Seoul, 02447, Korea. E-mail: youngmi.kim@khu.ac.kr
bResearch Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, 1 Gwanak-ro, Gwanak-gu, Seoul 08826, Korea. E-mail: jyleeut@snu.ac.kr
First published on 31st March 2026
Abstract
The BODIPY-based fluorogenic probe 1-CO enabled selective detection of AKR1C1 activity over related isoforms via ketone reduction, affording 50-fold fluorescence enhancement, nanomolar sensitivity, and real-time imaging in living cells.
Aldo–keto reductases (AKRs) constitute a large superfamily of NAD(P)(H)-dependent oxidoreductases that reduce a broad range of endogenous and xenobiotic carbonyl compounds, including steroids, prostaglandins, sugars, lipid peroxidation products, and numerous environmental or pharmaceutical agents.1 Within this superfamily, the human AKR1C subfamily consists of four highly homologous isoforms (AKR1C1–AKR1C4) that share over 86% sequence identity yet exhibit distinct substrate preferences and tissue distribution.2 Acting as 3-, 17-, and 20-ketosteroid reductases to varying degrees, these enzymes serve as key “pre-receptor” regulators of steroid hormone action.3 Among these isoforms, AKR1C1 (20α(3α)-hydroxysteroid dehydrogenase, 20α-HSD) holds distinct physiological and clinical significance.4 As the predominant 20-ketosteroid reductase in humans, it converts progesterone into its inactive form, 20α-hydroxyprogesterone, a critical step in pregnancy maintenance and the timing of parturition.5 In the central nervous system, AKR1C1 regulates neuroactive steroid levels that modulate γ-aminobutyric acid A (GABAA) receptor activity, thereby influencing mood, stress responses, and neurological function.6 Beyond these roles, AKR1C1 is frequently overexpressed in various malignancies, including breast, ovarian, lung, and prostate cancers,7 where it promotes tumor progression8 and contributes to chemotherapeutic resistance.9
Despite its broad clinical relevance, the selective study of AKR1C1 remains challenging. AKR1C1 shares high sequence identity with its isoforms—most notably AKR1C2, differing by only seven amino acids with a single non-conserved residue (Leu54 vs. Val54) in the active site.2,3c,4b,10 This structural near-identity makes it exceptionally difficult to selectively measure AKR1C1 activity using traditional methods, such as spectrophotometric monitoring of NADPH consumption2,11 or immunoassays,12 which lack the isoform selectivity, sensitivity, and spatiotemporal resolution required for complex biological systems. Alternative radiochemical methods using radiolabeled physiological substrates (e.g., [14C]5α-DHT, [14C]progesterone, or [3H]progesterone) offer higher sensitivity but are discontinuous, labor-intensive, and incompatible with high-throughput screening applications.2,13 To address these limitations, fluorogenic probes have emerged as powerful tools for real-time imaging of enzyme activity.14 Sames and colleagues pioneered push–pull chromophores wherein the enzymatic reduction of a nonfluorescent carbonyl (“Coumberone”) to a fluorescent alcohol (“Coumberol”) perturbs a π-conjugated system, dramatically altering emission properties.15 While these coumarin-based probes enabled visualization of cellular AKR1C2/AKR1C3 activity, their fluorescence is intrinsically sensitive to solvent polarity. In heterogeneous biological systems, polar environments stabilize their charge-separated excited states, leading to variable background signals that confound quantitative measurements.16 Furthermore, to our knowledge, no fluorogenic probe has been validated for the selective detection of AKR1C1 over its homologs. In this work, we report a BODIPY-based fluorogenic probe (1-CO, Scheme 1b) that enables selective detection of AKR1C1 activity. The probe exhibits excellent chemical stability, and a strong “turn-on” response upon AKR1C1-mediated reduction, with high sensitivity and isoform selectivity. These properties allow high-contrast imaging of endogenous AKR1C1 activity in living cells with minimal off-target interference, providing a versatile tool for probing AKR1C1 function in hormone-dependent diseases and drug resistance.
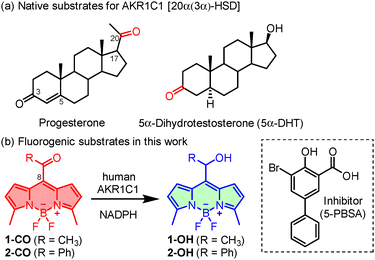 | ||
| Scheme 1 (a) Representative natural AKR1C1 substrates. (b) Activation of fluorogenic probes 1-CO and 2-CO via AKR1C1/NADPH-dependent reduction to 1-OH and 2-OH. Structure of the AKR1C1 inhibitor. | ||
The design of 1-CO harnesses the substrate specificity of AKR1C1 (20α-HSD), which catalyzes the NADPH-dependent reduction of progesterone to its 20α-hydroxy metabolite and converts other ketosteroids, such as 5α-dihydrotestosterone (5α-DHT), to the corresponding 3α-hydroxy metabolites (Scheme 1a). Rather than mimicking the full steroid framework, the probes incorporate a compact reducible ketone inspired by the C20 carbonyl of progesterone. Accordingly, the BODIPY-based probes 1-CO (R = CH3) and 2-CO (R = Ph) were constructed with a meso-acyl group as the enzyme-reactive motif (Scheme 1b), with 2-CO included as a sterically expanded analogue to examine how increased meso-acyl size influences enzyme recognition and thus isoform selectivity. This design exploits the distinct photophysical differences between meso-acyl BODIPYs and their reduced 2° alcohol derivatives (1-OH and 2-OH). As we recently reported,17 the sp2-hybridized meso-acyl form exhibits weak red emission with a large Stokes shift, resulting from a facile distortion in the excited state that accelerates non-radiative decay. Enzymatic reduction to the corresponding sp3-hybridized form suppresses this distortion pathway and restores the characteristic bright green BODIPY fluorescence, providing a robust red-to-green optical readout for AKR1C1-mediated reduction.
Probes 1-CO and 2-CO were synthesized in moderate yields (Schemes S1 and S2). To confirm the proposed fluorescence “turn-on” behavior required for enzymatic sensing, the corresponding reduction products (1-OH and 2-OH) were independently prepared as controls. In phosphate buffer (10 mM, pH 7.4, 1% DMSO), 1-CO and 2-CO showed absorption maxima at 516 nm (ε = 4.0 × 104 M−1 cm−1) and 518 nm (ε = 3.3 × 104 M−1 cm−1), respectively (Fig. 1a). Importantly, the absorption and emission maxima of 1-CO remained essentially unchanged across solvents with widely different polarities (Fig. S8)17 and in dioxane/water mixtures of varying compositions (Fig. S4). This is an advantage over push–pull dye systems for quantitative measurements in heterogeneous biological environments. Both probes displayed broad emission bands in the red region, with maxima centered at 624 nm (1-CO) and 603 nm (2-CO), and low quantum yields (ΦF = 0.011 and 0.033, respectively). In contrast, the reduced alcohol products 1-OH and 2-OH had slightly blue-shifted absorption (λabs = 510 nm and 515 nm, respectively) and markedly blue-shifted emission maxima (λem = 525 nm and 529 nm, respectively). Critically for sensing applications, the brightness (ε × ΦF) increased 132-fold and 36-fold for 1-OH and 2-OH, respectively, providing a robust photophysical basis for a turn-on fluorescence response upon enzymatic reduction.
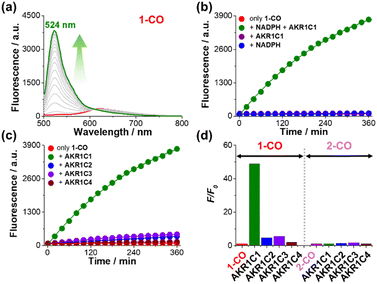
Given the marked emission difference between meso-acylated probes and their 2° alcohol products, the AKR1C1-mediated reduction of 1-CO and 2-CO was evaluated. Under optimized conditions (10 mM phosphate buffer, pH 7.4, 1% DMSO, 50 µM NADPH, and 37 °C; see the SI for details), incubation of 1-CO (4 µM) with AKR1C1 (100 nM) produced a time-dependent increase in green fluorescence at 524 nm, yielding a 50-fold enhancement after 6 h and reaching a plateau by 10 h (Fig. 2a, b and Fig. S21). ESI-MS analysis confirmed 1-OH as the primary enzymatic reduction product (Fig. S22), validating the proposed fluorescence activation mechanism. Control experiments showed negligible fluorescence increase in the absence of either AKR1C1 or NADPH (Fig. 2b), demonstrating that reduction of the probe is strictly enzyme- and cofactor-dependent. Additionally, 1-CO remained stable in the assay buffer over the same period (red, Fig. 2b), excluding non-specific reduction or degradation. Importantly, 1-CO showed excellent selectivity for AKR1C1 over the closely related AKR1C2–C4 isoforms (Fig. 2c and d). While AKR1C1 induced a robust 50-fold fluorescence increase, the other isoforms generated substantially weaker responses: AKR1C2 (F/F0 = 5), AKR1C3 (F/F0 = 6), and AKR1C4 (F/F0 = 2). The ∼10-fold preference for AKR1C1 over AKR1C2—the most homologous isoform—underscores the probe's suitability for isoform-specific detection in biological applications. In stark contrast, 2-CO showed minimal enzymatic activation across all isoforms (F/F0 = 1.1–1.8); however, slightly higher responses were observed for AKR1C2 and AKR1C3 (Fig. 2d and Fig. S24–S27). This pronounced performance difference likely reflects the impact of the meso-substituent on probe–enzyme interactions: molecular docking simulations indicate that the bulkier benzoyl group is poorly accommodated within the enzyme's active site (Fig. S35). This is not a surprise considering that AKR1C1's native substrate is the exocyclic acetyl group at the 17-position of progesterone.
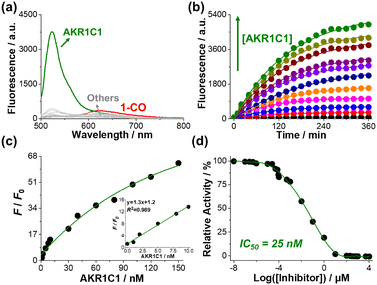
The response of 1-CO to potential interferences was assessed to ensure its practical utility in biological environments (Fig. 3a). Under the assay conditions (6 h, 37 °C), 1-CO (4 µM) showed strong fluorescence enhancement only with AKR1C1, whereas hydrolytic enzymes (trypsin, esterase, lipase), amino acids and thiols (GSH, Cys, HCy, Gly, Lys, Trp, His), reactive oxygen species (H2O2, HOCl, ˙OH), and common biomolecules (glucose, BSA) generated negligible signals. These results demonstrate that 1-CO retains exceptional selectivity for AKR1C1 even in the presence of common cellular interferents. Quantitative analysis was performed by monitoring fluorescence changes across AKR1C1 concentrations of 0–150 nM (Fig. 3b and c). Fluorescence at 524 nm increased with both incubation time and enzyme concentration, with a linear response over 0–10 nM (R2 = 0.989; Fig. 3c, inset). The detection limit was 0.14 nM (3σ/slope, 6 h), confirming the high sensitivity of 1-CO for AKR1C1 quantification. Enzyme kinetic analysis (Fig. S31–S33) gave a Michaelis constant (KM) of 2.43 µM for 1-CO, comparable to the native substrate progesterone (2.65 µM) and substantially lower than 5α-DHT (80.6 µM),2 indicating favorable binding affinity. The catalytic rate constant (kcat) was 2.7 × 10−3 min−1, yielding a catalytic efficiency (kcat/KM) of 1.1 mM−1 min−1. Although catalytic efficiency was lower for 1-CO than for progesterone (109 mM−1 min−1) or 5α-DHT (8 mM−1 min−1), the large quantum yield increase upon reduction enabled a strong fluorogenic response despite moderate turnover.
The utility of 1-CO for the screening of enzyme inhibitors was tested using the well-characterized substrate-competitive AKR1C1 inhibitor 3-bromo-5-phenylsalicylic acid (5-PBSA; Ki = 4 nM).11,18 Preincubation of AKR1C1 with 5-PBSA (0–10 mM) produced dose-dependent suppression of 1-CO's fluorescence response, yielding an IC50 of 24.6 ± 0.5 nM (Fig. 3d). The corresponding Ki (∼9 nM, calculated using the Cheng–Prusoff equation),19 was in reasonable agreement with the reported value, establishing 1-CO as a reliable fluorescence-based platform for high-throughput AKR1C1 inhibitor screening. Building on the performance of 1-CO in cell-free assays and inhibitor screening, whether the probe could detect endogenous AKR1C1 activity in living cells was next examined. HeLa (cervical cancer) and A549 (lung cancer) cells were incubated with 1-CO (2 µM) and imaged by confocal microscopy using red (Ex/Em = 561/566–635 nm) and green (Ex/Em = 488/493–560 nm) channels. Time-lapse imaging over 6 h revealed clear changes in both cell lines, with decreasing red and increasing green fluorescence (Fig. 4a and Fig. S36a), indicating efficient cell permeability and intracellular reduction of 1-CO to 1-OH. Line-scan analysis at 6 h confirmed dominant green emission (Fig. 4b and Fig. S36b), consistent with enzymatic probe activation. Quantitative analysis revealed pronounced cell-type-dependent kinetics (Fig. 4c and Fig. S36c). HeLa cells showed a modest green-to-red fluorescence (G/R) ratio increase (0.3 → 2.4) over 6 h, whereas A549 cells exhibited much rapid and extensive conversion, reaching a G/R ratio of ∼5.5. These differences aligned with endogenous AKR1C1 expression levels: western blots confirmed substantially higher AKR1C1 expression in A549 cells relative to HeLa cells (Fig. 4d). As elevated AKR1C1 is implicated in proliferation and metastasis of non-small cell lung cancer,8 these results highlight the potential of 1-CO for metabolic profiling of AKR1C1-driven cancer phenotypes.
To confirm that probe activation occurs at sites of endogenous enzyme activity, co-localization studies were performed using ER-Tracker Blue-White. Following incubation with 1-CO (2 µM, 6 h), the resulting green fluorescence from enzymatically generated 1-OH exhibited high spatial overlap with the ER marker in both cell lines (Pearson's r = 0.92 for HeLa and 0.94 for A549) (Fig. S37). This ER-specific localization is consistent with the known subcellular distribution of AKR1C1,20 confirming that probe conversion occurs predominantly at the site of enzyme expression. Cytotoxicity assays further demonstrated excellent biocompatibility, with >90% cell viability maintained after 24 h exposure to up to 8 µM 1-CO (Fig. S38). Target engagement was validated using the AKR1C1-selective inhibitor 5-PBSA. Pretreatment with 5-PBSA (10 µM, 3 h) markedly reduced both green and red fluorescence in HeLa and A549 cells (Fig. S39a and b), without altering AKR1C1 protein levels (Fig. S39c), confirming that intracellular probe activation is specifically driven by endogenous AKR1C1. Collectively, these results establish 1-CO as a powerful tool for quantitative, real-time visualization of AKR1C1 activity in living cells, offering broad utility for probing enzyme-driven metabolic processes in complex biological environments.
In summary, the polarity-independent BODIPY-based fluorogenic probe 1-CO enables the highly sensitive and selective detection of AKR1C1 activity. It combines excellent photophysical properties, strong isoform selectivity, low cytotoxicity, and robust cell permeability to enable real-time monitoring of endogenous AKR1C1 dynamics in living cells. These features establish 1-CO as a powerful molecular tool for studying AKR1C biology and for high-throughput discovery of AKR1C1-targeted inhibitors.
This research was supported by National Research Foundation of Korea (NRF) grants funded by the Korean government (MSIT, No. RS-2025-16063709 to J. L. and RS-2024-00353907 to Y. K.).
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: experimental details, additional spectroscopic data and photographs. See DOI: https://doi.org/10.1039/d6cc00473c.References
- (a) T. M. Penning and J. E. Drury, Arch. Biochem. Biophys., 2007, 464, 241 CrossRef CAS PubMed; (b) O. A. Barski, S. M. Tipparaju and A. Bhatnagar, Drug Metab. Rev., 2008, 40, 553 CrossRef CAS; (c) R. D. Mindnich and T. M. Penning, Hum. Genomics, 2009, 3, 362 CrossRef CAS PubMed; (d) T. M. Penning, Chem.-Biol. Interact., 2015, 234, 236 CrossRef CAS PubMed.
- T. M. Penning, M. E. Burczynski, J. M. Jez, C.-F. Hung, H.-K. Lin, H. Ma, M. Moore, N. Palackal and K. Ratnam, Biochem. J., 2000, 351, 67 CrossRef CAS.
- (a) M. J. Bennett, R. H. Albert, J. M. Jez, H. Ma, T. M. Penning and M. Lewis, Structure, 1997, 5, 799 CrossRef CAS PubMed; (b) T. M. Penning, D. R. Bauman, Y. Jin and T. L. Rizner, Mol. Cell. Endocrinol., 2007, 265–266, 77 CrossRef CAS PubMed; (c) T. L. Rižner and T. M. Penning, Steroids, 2014, 79, 49 CrossRef PubMed; (d) T. M. Penning, P. Wangtrakuldee and R. J. Auchus, Endocr. Rev., 2019, 40, 447 CrossRef.
- (a) Y. Zhang, I. Dufort, P. Rheault and V. Luu-The, J. Mol. Endocrinol., 2000, 25, 221 CAS; (b) P. Brožič, S. Turk, A. O. Adeniji, J. Konc, D. Janežič, T. M. Penning, T. L. Rižner and S. Gobec, J. Med. Chem., 2012, 55, 7417 CrossRef PubMed; (c) X. Chu, S. He, Y. Liu, Y. Liu, F. Feng, Q. Guo, L. Zhao and H. Sun, Chem.-Biol. Interact., 2022, 351, 109746 CrossRef CAS PubMed.
- (a) R. P. Piekorz, S. Gingras, A. Hoffmeyer, J. N. Ihle and Y. Weinstein, Mol. Endocrinol., 2005, 19, 431 CrossRef CAS PubMed; (b) M. Paul, T. Zakar, J. Phung, A. Gregson, A. P. Barreda, T. A. Butler, F. R. Walker, C. Pennell, R. Smith and J. W. Paul, Reprod. Sci., 2023, 30, 2512 CrossRef CAS PubMed.
- (a) R. H. Purdy, A. L. Morrow, P. H. Moore Jr. and S. M. Paul, Proc. Natl. Acad. Sci. U. S. A., 1991, 88, 4553 CrossRef CAS PubMed; (b) S. M. Paul and R. H. Purdy, FASEB J., 1992, 6, 2311 CrossRef CAS PubMed; (c) Y. Higaki, N. Usami, S. Shintani, S. Ishikura, O. El-Kabbani and A. Hara, Chem.-Biol. Interact., 2003, 143–144, 503 CrossRef CAS PubMed.
- (a) C.-M. Zeng, L.-L. Chang, M.-D. Ying, J. Cao, Q.-J. He, H. Zhu and B. Yang, Front. Pharmacol., 2017, 8, 119 Search PubMed; (b) R. Matsumoto, M. Tsuda, K. Yoshida, M. Tanino, T. Kimura, H. Nishihara, T. Abe, N. Shinohara, K. Nonomura and S. Tanaka, Sci. Rep., 2016, 6, 34625 CrossRef CAS PubMed.
- (a) H. Zhu, L.-L. Chang, F.-J. Yan, Y. Hu, C.-M. Zeng, T.-Y. Zhou, T. Yuan, M.-D. Ying, J. Cao, Q.-J. He and B. Yang, Theranostics, 2018, 8, 676 CrossRef CAS; (b) L.-L. Chang, P.-H. Lu, W. Yang, Y. Hu, L. Zheng, Q. Zhao, N.-M. Lin and W.-Z. Zhang, Transl. Oncol., 2022, 20, 101421 CrossRef PubMed.
- (a) T. Matsunaga, A. Hojo, Y. Yamane, S. Endo, O. El-Kabbani and A. Hara, Chem.-Biol. Interact., 2013, 202, 234 CrossRef CAS PubMed; (b) K.-A. Jung, B.-H. Choi, C.-W. Nam, M. Song, S.-T. Kim, J. Y. Lee and M.-K. Kwak, Toxicol. Lett., 2013, 218, 39 CrossRef CAS PubMed; (c) T. M. Penning, Chem. Res. Toxicol., 2017, 30, 162 Search PubMed; (d) W.-M. Chang, Y.-C. Chang, Y.-C. Yang, S.-K. Lin, P. M.-H. Chang and M. Hsiao, J. Exp. Clin. Cancer Res., 2019, 38, 245 Search PubMed; (e) T. M. Penning, S. Jonnalagadda, P. C. Trippier and T. L. Rižner, Pharmacol. Rev., 2021, 73, 1150 CrossRef CAS PubMed.
- T. M. Penning, J. Steroid Biochem. Mol. Biol., 2011, 125, 46 CrossRef CAS PubMed.
- O. El-Kabbani, P. J. Scammells, J. Gosling, U. Dhagat, S. Endo, T. Matsunaga, M. Soda and A. Hara, J. Med. Chem., 2009, 52, 3259 CrossRef CAS PubMed.
- Q. Ji, C. Aoyama, Y.-D. Nien, P. I. Liu, P. K. Chen, L. Chang, F. Z. Stanczyk and A. Stolz, Cancer Res., 2004, 64, 7610 CrossRef CAS PubMed.
- (a) T. L. Rižner, H. K. Lin and T. M. Penning, Chem.-Biol. Interact., 2003, 143–144, 401 CrossRef PubMed; (b) T. L. Rižner, H. K. Lin, D. M. Peehl, S. Steckelbroeck, D. R. Bauman and T. M. Penning, Endocrinology, 2003, 144, 2922 CrossRef PubMed; (c) S. Blanchette, K. Blouin, C. Richard, P. Dupont, V. Luu-The and A. Tchernof, J. Clin. Endocrinol. Metab., 2005, 90, 264 CrossRef CAS.
- (a) T. Komatsu and Y. Urano, Anal. Sci., 2015, 31, 257 CrossRef CAS PubMed; (b) W. Chyan and R. T. Raines, ACS Chem. Biol., 2018, 13, 1810 CrossRef CAS PubMed; (c) X. Wu, W. Shi, X. Li and H. Ma, Acc. Chem. Res., 2019, 52, 1892 CrossRef CAS PubMed; (d) J. Zhang, X. Chai, X.-P. He, H.-J. Kim, J. Yoon and H. Tian, Chem. Soc. Rev., 2019, 48, 683 RSC; (e) K. Fujita and Y. Urano, Chem. Rev., 2024, 124, 4021 CrossRef CAS PubMed.
- (a) D. J. Yee, V. Balsanek and D. Sames, J. Am. Chem. Soc., 2004, 126, 2282 CrossRef CAS PubMed; (b) D. J. Yee, V. Balsanek, D. R. Bauman, T. M. Penning and D. Sames, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 13304 CrossRef CAS PubMed; (c) M. Halim, D. J. Yee and D. Sames, J. Am. Chem. Soc., 2008, 130, 14123 CrossRef CAS; (d) S. Jockusch, Q. Zheng, G. S. He, H. E. Pudavar, D. J. Yee, V. Balsanek, M. Halim, D. Sames, P. N. Prasad and N. J. Turro, J. Phys. Chem. C, 2007, 111, 8872 CrossRef CAS.
- C. Reichardt, Chem. Rev., 1994, 94, 2319 CrossRef CAS.
- Y. Kim and Y. Kim, Org. Lett., 2025, 27, 6710 CrossRef CAS PubMed.
- (a) O. El-Kabbani, P. J. Scammells, T. Day, U. Dhagat, S. Endo, T. Matsunaga, M. Soda and A. Hara, Eur. J. Med. Chem., 2010, 45, 5309 CrossRef CAS PubMed; (b) O. El-Kabbani, U. Dhagat and A. Hara, J. Steroid Biochem. Mol. Biol., 2011, 125, 105 CrossRef CAS.
- Y. Cheng and W. H. Prusoff, Biochem. Pharmacol., 1973, 22, 3099 CrossRef CAS PubMed.
- https://www.proteinatlas.org/ENSG00000187134-AKR1C1.
Footnote |
| † Y. K. and L. B. H. P. contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |