Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A solid-phase click2 strategy for chromophore–DNA conjugates and their application as a light harvesting system
Andreas J. Schmidt ,
Annabel M. L. Knispel
,
Annabel M. L. Knispel and
Hans-Achim Wagenknecht
and
Hans-Achim Wagenknecht *
*
Karlsruhe Institute of Technology (KIT), Institute of Organic Chemistry, Fritz-Haber-Weg 6, Karlsruhe 76131, Germany. E-mail: wagenknecht@kit.edu
First published on 13th March 2026
Abstract
A stepwise and orthogonal approach of two independent click reactions directly on the solid phase gives easy access to doubly modified DNA. Firstly, 5′-ethynyl-2′-deoxyuridine reacts directly with an azide-modified porphyrin; secondly, 5′-azido-2′-deoxyuridine is generated in situ from 5′-iodo-2′-deoxyuridine, which enables a click reaction with an alkyne-functionalized pyrene. The doubly modified DNA was characterized as a light-harvesting system.
Copper-catalyzed azide–alkyne cycloaddition (CuAAC)1,2 has emerged as a robust and efficient tool for postsynthetic functionalization of biomolecules, in particular DNA, bypassing the need for complex synthesis of modified phosphoramidites as building blocks for each DNA modification and simplifying the purification of modified DNA.3–7 Conventionally, this is achieved using alkyne-bearing building blocks, such as 5′-ethynyl-2′-deoxyuridine (EdU) reacting with organic azides for labelling.8 The approach vice versa requires the synthesis of a DNA modified with an azide group. However, the inherent instability of many azides during automated DNA synthesis based on phosphorous(III) often restricts this approach and limits it to rare examples in the literature.5,9 We solved this issue by synthesizing DNA containing 5-iodo-2′-deoxyuridine (IdU) within the sequence, which can selectively and postsynthetically be modified by ethynyl-modified Nile red as a fluorescent label after in situ formation of the intermediate 5-azido-2′-deoxyuridine (AdU).3 Building on this approach, we report herein the combination of EdU and IdU for two independent orthogonal click reactions resulting in a dual click2 postsynthetic DNA labelling strategy with full sequence control (Fig. 1).
The desired oligonucleotides DNA1, DNA2 and DNA3 were designed with a random sequence, the chromophore modifications in the middle and the chromophores in DNA3 separated by two intervening base pairs, and were synthesized using the readily accessible phosphoramidites of commercially available EdU and IdU for the corresponding precursors DNA1′, DNA2′ and DNA3′. Standard coupling protocols ensured efficient incorporation of both artificial nucleotides into the strand.10
Afterwards, the presynthesized oligonucleotides (1 µmol) were not cleaved from the solid support (controlled pore glass, CPG). The first postsynthetic modification was achieved by reacting the ethynyl group of EdU as part of the sequence of DNA1′ and DNA3′ with the porphyrin azide 1 (10 equiv.) by means of CuAAC (50 equiv. sodium ascorbate, 11 equiv. TBTA ligand,11 30 equiv. Cu(CH3CN)4PF6, 15 h at room temperature). As anticipated and determined by HPLC, the first click modification proceeded with near-quantitative efficiency. To introduce 1-ethynylpyrene 2 as the first or second label, the azide group of AdU was generated in situ from the IdU precursor as part of the presynthesized DNA2′ or DNA3′ by treatment with sodium azide (200 µmol in DMSO, 1 h at 50 °C). The resulting oligonucleotide was immediately reacted with 1-ethynylpyrene 2 (50 equiv.) in a click reaction using the same conditions as for the formation of DNA1. The selection of pyrene for the second chromophore modification is strategically significant. (i) Its UV/Vis absorption occurs well separated from the porphyrin. (ii) 1-Ethnylpyrene is commercially available. (iii) Pyrenes are able to induce an electron transfer in DNA.12,13 In conclusion, the pyrene/porphyrin pair is well suited for light-harvesting systems by spectroscopic means.14,15 Successful modification to DNA1–DNA3 was evidenced by ESI mass spectrometry. Interestingly, MS analysis of DNA1 and DNA3 showed the coordination of Cu(II), presumably by the presence of the porphyrin as a ligand.
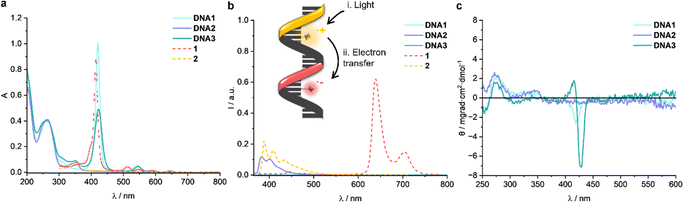
Following successful synthesis of the chromophore–DNA conjugates DNA1, DNA2 and DNA3, their photophysical properties were characterized using methods of optical spectroscopy, including UV/vis absorption spectroscopy, steady-state and time-resolved fluorescence spectroscopy, and circular dichroism (CD) spectroscopy (Fig. 2). In the case of pyrene, we could determine a melting temperature of 40 °C for double-stranded DNA2. However, the bulky porphyrin modification strongly interferes with annealing and no stable hybridization was observed with complementary unmodified counterstrands to DNA1 and DNA3, and consequently the melting temperatures could not be determined. Consequently, we used only the single strands DNA1, DNA2 and DNA3 for characterization. Their UV/Vis absorption spectra (Fig. 2a) display the characteristic nucleic acid maximum at λ = 260 nm together with the pyrene band at λ = 350 nm and an intense porphyrin Soret band at λ = 419 nm, accompanied by Q-bands in the range of λ = 550–600 nm.16,17 The single strands show also the DNA-characteristic helicity as the circular dichroism (Fig. 2c) reveals the characteristic Cotton effect of B-DNA with a positive band at λ = 275 nm and a negative band at λ = 250 nm. A negative Cotton effect within the Soret band characterizes the spatial orientation of the porphyrin macrocycle when intercalated between nucleobase pairs. Significant exciton splitting in the range of λ = 400–450 nm was detected, due to strong ground-state interactions between the two different chromophores in DNA3.18–20 Minor intensity changes in the base regions merely indicate local structural adjustments in the immediate vicinity of the modification sites.17 Obviously, two intervening base pairs between the chromophores are sufficient. Compared to chromophores 1 and 2, the fluorescence spectra (Fig. 2b) show pronounced emission quenching due to covalent binding of these chromophores to DNA1 and DNA2. A significant attenuation of porphyrin fluorescence occurs in the singly labelled DNA2. In the doubly labelled DNA3, both the porphyrin and pyrene emission are almost completely quenched. This indicates highly efficient non-radiative decay pathways of the excited state of the chromophores. Time-resolved fluorescence measurements by TCSPC corroborate this interpretation, revealing a drastically shortened average lifetime of τ = 0.13 ns for the emission of DNA3 at λem = 400 nm when excited at λexc = 370 nm (ns-pulsed LED) compared to typical lifetimes of the unbound pyrene derivative 2 with τ0 = 6.4 ns, respectively, measured at identical concentration in DMSO (due to solubility limitations of the chromophore in aqueous media).21–24 This corresponds to a quenching efficiency E exceeding 95%, calculated according to E = 1 − (τ/τ0), where τ and τ0 denote the fluorescence lifetimes of the chromophore covalently bound to DNA (τ) and the chromophore without DNA (τ0), respectively.
The estimated quenching rate constant of k = 0.16 × 109 s−1 lies within the typical range reported for ultrafast photoinduced electron processes.23,25,26 A qualitative Rehm–Weller estimation23 based on the redox potentials, ΔGET = Eox − Ered − E00 (neglecting the Coulomb interaction EC) of the chromophores supports the thermodynamic feasibility of a photoinduced charge separation in this bichromophore DNA: this charge separation reduces the porphyrin by one electron and oxidizes pyrene, based on the reduction potential of tetraphenylporphyrin Ered = −1.05 V (vs. SCE),27 E00 = 3.25 V28,29 and the oxidation potential Eox = 1.28 V (vs. SCE) of pyrene28,29 yielding a strongly exergonic driving force ΔGET = −0.92 V.29
In conclusion, we have developed a novel, stepwise orthogonal click2 strategy performed directly on a solid support after automated DNA synthesis. This approach leverages readily accessible phosphoramidite precursors (EdU and IdU) to incorporate potentially any chromophore modified with an azide or ethynyl function. We representatively tested this synthetic method with a porphyrin and a pyrene that labelled the DNA efficiently. The spectroscopic data indicate that the DNA helix acts as a pre-organized scaffold that positions donor and acceptor chromophores at a distance of approximately 1–1.5 nm, thereby promoting ultrafast electron transfer processes. The almost complete fluorescence quenching in the doubly modified DNA cannot be attributed to extensive structural denaturation or exciton delocalization, but rather to a highly efficient, DNA-mediated non-radiative relaxation mechanism. Our method expands the toolbox for synthesizing complex, multi-chromophoric DNA assemblies suitable for applications in light-harvesting systems and molecular electronics.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: all experimental data, procedures for syntheses and spectroscopic experiments. See DOI: https://doi.org/10.1039/d6cc00319b.Acknowledgements
Financial support from the Deutsche Forschungsgemeinschaft (DFG) (grant Wa 1386/27-1), the Fonds der chemischen Industrie (FCI) and the Karlsruhe Institute of Technology (KIT) is gratefully acknowledged.References
- C. W. Tornoe, C. Christensen and M. Meldal, J. Org. Chem., 2002, 67, 3057–3064 CrossRef CAS PubMed.
- V. V. Rostovstev, L. G. Green, V. V. Fokin and K. B. Sharpless, Angew. Chem., Int. Ed., 2002, 41, 2596–2599 CrossRef.
- C. Beyer and H.-A. Wagenknecht, Chem. Commun., 2010, 46, 2230–2231 RSC.
- K. Krell, D. Harijan, D. Ganz, L. Doll and H.-A. Wagenknecht, Bioconjugate Chem., 2020, 31, 990–1011 CrossRef CAS PubMed.
- A. El-Sagheer and T. Brown, Chem. Rev., 2010, 39, 1388–1405 CAS.
- M. Shelbourne, X. Chen, T. Brown and A. H. El-Sagheer, Chem. Commun., 2011, 47, 6257–6259 RSC.
- O. S. Wolfbeis, Angew. Chem., Int. Ed., 2007, 46, 2980–2982 CrossRef PubMed.
- K. Gutsmiedl, D. Fazio and T. Carell, Chem. – Eur. J., 2010, 16, 6877–6883 CrossRef CAS PubMed.
- M. Aigner, M. Hartl, K. Fauster, J. Steger, K. Bister and R. Micura, ChemBioChem, 2011, 12, 47–51 CrossRef CAS PubMed.
- M. B. Walunj and S. G. Srivatsan, Bioconjugate Chem., 2020, 31, 2513–2521 CrossRef CAS PubMed.
- T. R. Chan, R. Hilgraf, K. B. Sharpless and V. V. Fokin, Org. Lett., 2004, 6, 2853–2855 CrossRef CAS PubMed.
- H.-A. Wagenknecht, Angew. Chem., Int. Ed., 2003, 42, 2454–2460 CrossRef CAS PubMed.
- N. Amann, T. Fiebig and H.-A. Wagenknecht, Chem. – Eur. J., 2002, 8, 4877–4883 CrossRef CAS PubMed.
- E. Stulz, Acc. Chem. Res., 2017, 50, 823–831 CrossRef CAS PubMed.
- H. S. Lee, J. H. Han and D. W. Cho, Phys. Chem. Chem. Phys., 2017, 19, 27123–27131 RSC.
- J. S. Lindsey, Acc. Chem. Res., 2010, 43, 300–311 CrossRef CAS PubMed.
- W. Saenger, Principles of nucleic acid structure, Springer Science & Business Media, 2013 Search PubMed.
- M. Şener, J. Strümpfer, J. Hsin, D. Chandler, S. Scheuring, C. N. Hunter and K. Schulten, ChemPhysChem, 2011, 12, 518–531 CrossRef PubMed.
- A. Olaya-Castro and G. D. Scholes, Int. Rev. Phys. Chem., 2011, 30, 49–77 Search PubMed.
- S. G. Telfer, T. M. McLean and M. R. Waterland, Dalton Trans., 2011, 40, 3097–3108 RSC.
- D. Inan, R. K. Dubey, W. F. Jager and F. C. Grozema, J. Phys. Chem. C, 2019, 123, 36–47 CrossRef CAS PubMed.
- M. L. Perrin, F. Prins and D. Dulić, Angew. Chem., Int. Ed., 2011, 50, 11223–11226 CrossRef CAS PubMed.
- D. Rehm and A. Weller, Isr. J. Chem., 1970, 8, 259–271 CrossRef CAS.
- M. R. Wasielewski, Acc. Chem. Res., 2009, 42, 1910–1921 CrossRef CAS PubMed.
- M. R. Wasielewski, Chem. Rev., 1992, 92, 435–461 CrossRef CAS.
- R. A. Marcus, J. Electroanal. Chem., 1997, 438, 251–259 CrossRef CAS.
- J. R. Diers, C. Kirmaier, M. Taniguchi, J. S. Lindsey, D. F. Bocian and D. Holten, Phys. Chem. Chem. Phys., 2021, 23, 19130–19140 RSC.
- T. L. Netzel, M. Zhao, K. Nafisi, J. Headrick, M. S. Sigman and B. E. Eaton, J. Am. Chem. Soc., 1995, 117, 9119–9128 CrossRef CAS.
- N. Amann, E. Pandurski, T. Fiebig and H.-A. Wagenknecht, Angew. Chem., Int. Ed., 2002, 41, 2978–2980 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |