Open Access Article
Open Access ArticleTransformation of tetranuclear [{MoO3(Lcat)Cu}2]2− into trinuclear [MoO2(Lcat)2Cu2] and possible implications for the formation of Bis(MPT)Mo by MobA
A. M. Buddhika
Chandima†
 a,
Umesh I.
Kaluarachchige Don
a,
Cassandra L.
Ward
a,
Umesh I.
Kaluarachchige Don
a,
Cassandra L.
Ward
 b,
Richard L.
Lord
b,
Richard L.
Lord
 *c and
Stanislav
Groysman
*c and
Stanislav
Groysman
 *a
*a
aDepartment of Chemistry, Wayne State University, 5101 Cass Ave., Detroit, MI 48202, USA. E-mail: groysman@wayne.edu
bLumigen Instrument Center, Wayne State University, 5101 Cass Avenue, Detroit, Michigan 48202, USA
cDepartment of Chemistry, Grand Valley State University, 1 Campus Drive, Allendale, Michigan 49401, USA. E-mail: lordri@gvsu.edu
First published on 2nd March 2026
Abstract
Many molybdoenzymes contain two molybdopterin (MPT) ligands, but the mechanism for the transformation of mono(MPT)Mo into bis(MPT)Mo is lacking. Using a catecholate-based system, we demonstrate that [MoVIO3(Lcat)]2− forms a dimeric structure, which spontaneously disproportionates to produce [MoVIO2(Lcat)2]2− and molybdate. This suggests that a similar transformation of [{MoO3(MPT)}2]4− into [MoO2(MPT)2]2− is feasible.
Molybdenum-dependent enzymes are ubiquitous in biology, being crucial for most forms of life.1 Nearly all molydoenzymes, and all known tungsten enzymes, utilize molybdopterin (MPT) cofactor at their active sites; MPT serves as a dithiolene ligand to the Mo (W) center.1,2 Certain molybdoenzymes (e.g. xanthine dehydrogenase and sulfite oxidase) utilize a single MPT cofactor (Fig. 1). However, many other molybdoenzymes (DMSO reductase family) and all tungstoenzymes incorporate two MPT cofactors per metal (for selected structures of molybdoenzymes, see Fig. 1). The enzymatic mechanism of molybdate insertion into MPT to form mono(MPT)Mo active sites has been interrogated in depth, including the detailed molecular pathway through which MPT reacts with the Mo(VI) precursor, molybdate.3,4 In contrast, while the biochemistry of the bis(MPT)Mo formation is well understood, the intricate molecular details of how bis(MPT)Mo (W) sites are formed are lacking.5–7 Leimkühler and coworkers demonstrated formation of bis(MPT)Mo by the enzyme MobA. The authors hypothesized that bis(MPT)Mo sites could form by disproportionation of two mono(MPT) sites [MoO3(MPT)] (Fig. 1); this reaction should also result in the formation of molybdate.8,9 As an alternative possibility, it was proposed that the formation of bis(MPT)Mo sites could take place in MobA by the combination of mono(MPT)Mo and free MPT.8 To our knowledge, these hypotheses have not been tested in a molecular model.
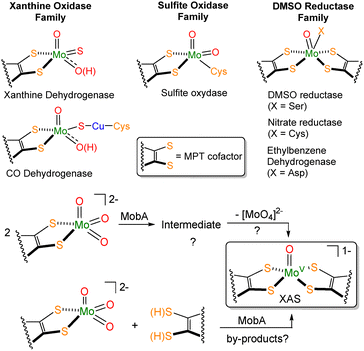 | ||
| Fig. 1 Three families of molybdoenzymes (top). Possible routes of MobA-mediated formation of bis(MPT)Mo sites from mono(MPT)Mo(VI) trioxo (bottom). | ||
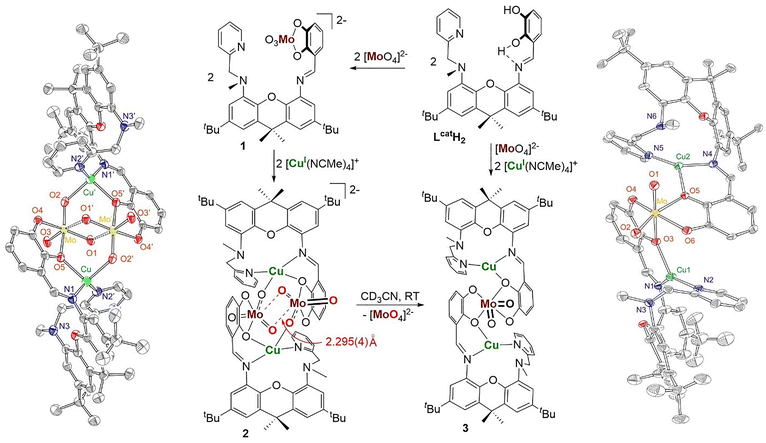
Our research focuses on structural and functional models of molybdoenzymes, including models of CO dehydrogenase (CODH) (Fig. 1).10–19 Mo-Cu CODH belongs to the xanthine dehydrogenase family of molybdoenzymes, as the Mo(VI) site is ligated by one MPT ligand, two oxo groups, and one sulfido that bridges to a nearby Cu(I) site.1,10,11 To develop these models, we employ heterodinucleating ligands that incorporate catecholate as a model for dithiolene.13 Catecholates exhibit many similar features to those of dithiolenes:20,21 they are redox non-innocent dichalcogenide donor ligands that generally form tightly bound metal chelates. While our previous studies have demonstrated bimetallic cooperativity, we were not able to isolate heterobimetallic complexes featuring nucleophilic [MoVIO3(cat)] (cat = catecholate) and Cu(I) in close proximity.22–24 Using a newly synthesized heterodinucleating catecholate/aminopyridine ligand LcatH2 (Fig. 2), we report the isolation and structural characterization of [{MoO3(Lcat)Cu}2]2−. More significantly, we demonstrate that this complex spontaneously and unimolecularly disproportionates in solution into the trinuclear mono-Mo bis(catecholate) complex [MoO2(Lcat)2Cu2] while releasing molybdate. This transformation could provide a molecular model for the related 2[MoO3(MPT)]2− → [MoO2(MPT)2]2− + [MoO4]2− reaction that likely occurs in MobA.
Treatment of LcatH2 with one equivalent of tetraethylammonium molybdate produced lemon-yellow (NEt4)2[MoO3(Lcat)] (1), which was obtained in 89% yield. Although we could not obtain the crystal structure of 1, the multinuclear NMR spectroscopic and high-resolution mass spectrometric (HRMS) characterization methods unequivocally supported its formation. The HRMS contains a molecular ion peak at m/z = 721.2164, whose isotopic distribution pattern correlates well with the expected isotopic distribution for [MoO2(OH)(Lcat)]− (see SI). 1 is stable in CD3CN demonstrating no visible decomposition for at least one week at room temperature. Closely related [MO3(bdt)]2− (M = Mo, W; bdt = benzene-1,2-dithiolate) were previously reported and were also shown to be stable under ambient conditions.25,26 Subsequent treatment of 1 with [Cu(NCMe)4](PF6) produced a dark orange solution. NMR investigation of the reaction carried out in CD3CN indicated the disappearance of 1 and the formation of a new product. Crystallization from CH3CN/ether at −33 °C produced dark orange crystals of 2, which were characterized by X-ray crystallography (Fig. 2), 1H and 13C NMR spectroscopy, and HRMS (see SI). The solid-state structure of 2 demonstrates the [Mo2O6(Lcat)2Cu2]2− composition, balanced by two tetraethylammonium ions. The structure can be viewed as a homodimer of two heterodinuclear “[MoO3(Lcat)Cu]−” components, [{MoO3(Lcat)Cu}2]2−. Within each of the heterodinuclear units, Mo(VI) and Cu(I) are held together by the same Lcat ligand with the metals interacting directly through a catecholate oxygen; in addition, there is a {MoVI![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O}–CuI bond (2.117(4) Å) between neighboring units. Furthermore, a weak interaction is observed between {MoVI
O}–CuI bond (2.117(4) Å) between neighboring units. Furthermore, a weak interaction is observed between {MoVI![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O} and MoVI of the neighboring complex (2.295(4) Å), resulting in a Mo2O2 “diamond” core. While this structure does not constitute a structural analogue of Mo–Cu CODH, it nevertheless captures the reactive Cu(I)-coordinated nucleophilic [MoVIO3(L)]2− fragment that is likely necessary (in the form of [MoVIO2S(MPT)]2−) for the nucleophilic attack at the Cu(I)-activated CO.
O} and MoVI of the neighboring complex (2.295(4) Å), resulting in a Mo2O2 “diamond” core. While this structure does not constitute a structural analogue of Mo–Cu CODH, it nevertheless captures the reactive Cu(I)-coordinated nucleophilic [MoVIO3(L)]2− fragment that is likely necessary (in the form of [MoVIO2S(MPT)]2−) for the nucleophilic attack at the Cu(I)-activated CO.
Complex 2 is stable in the solid state at −33 °C. In acetonitrile solution at room temperature, however, it undergoes facile decomposition. The decomposition was monitored by disappearance of the 1H NMR spectrum of 2 (Fig. S16) and formation of an orange precipitate. The acetonitrile-insoluble orange precipitate was recrystallized from dichloromethane/ether to yield X-ray quality crystals of complex 3. 3 was characterized by X-ray crystallography, 1H and 13C NMR spectroscopy, and HRMS. The solid-state structure of 3 reveals a trinuclear [MoO2(Lcat)2Cu2] composition (Fig. 2). The single Mo(VI) center is octahedral, bearing two oxo ligands and two catecholates. The Mo–oxo bonds in 3 are significantly shorter than in 2, consistent with less nucleophilic character. Each Lcat ligand incorporates trigonal planar Cu(I) coordinated by the pyridine and imine nitrogen donors, in addition to one catecholate oxygen donor that links it to Mo(VI). Complex 3 can also be synthesized directly by reaction of molybdate with two equivalents of LcatH2, followed by the addition of two equivalents of [Cu(NCMe)4](PF6) (Fig. 2).
In addition to the formation of 3, decomposition of 2 in acetonitrile produces molybdate. Molybdate remains soluble in acetonitrile and its presence was confirmed by HRMS (see SI). Thus, decomposition of 2 can be formulated as a disproportionation reaction likely driven by the relative instability/nucleophilicity of the Mo(VI) trioxo and precipitation of 3 from the reaction medium. We investigated the decomposition kinetics of 2 by 1H NMR spectroscopy. The reaction is first order as demonstrated by kinetic analysis (Fig. S16), with k = 8.0 × 10−6 s−1. Unimolecular decomposition of 2 suggests that 2 does not decompose into heterodinuclear dimers in solution. Instead, the transformation of 2 into 3 and [MoO4]2− is likely enabled by the tetranuclear core of 2, in which two Mo(VI) interact via the oxo groups.
Based on this evidence, we performed DFT calculations to probe intermediates along the reaction path from 2 to 3. Calculations were performed at the B3LYP-D4/def2-TZVP/SMD(MeCN)//BP86/def2-SVP/SMD(MeCN) level of theory in ORCA 6.1.0 (full details in the SI).27–41 A simplification of Lcat replaced the xanthene tBu and Me substituents with H, and NEt4+ counterions were omitted (i.e. calculated as anionic species). 2 was optimized as a dianion and is shown in Fig. 3. To be as unbiased as possible, linear scans along multiple Mo–O bonds in the diamond core of 2 that could lead to the release of molybdate were probed. These scans produced intermediates i–iii (Fig. 3); all three are consistent with experimental conditions, being endergonic by 14–23 kcal mol−1. Intermediate i features two five-coordinate molybdenum centers; while the structure is non-symmetric, neither Mo is yet to coordinate to both catecholates. Intermediate ii features a bridging molybdate between the left Cu and a Mo with two catecholates coordinated, one κ2 and one κ1. Intermediate iii similarly features molybdate seemingly poised to dissociate, but the right-most Cu is no longer bound to an oxo at the six-coordinate Mo. Of these three species, the one featuring a three- versus four-coordinate Cu on the right side of the structure is favored.
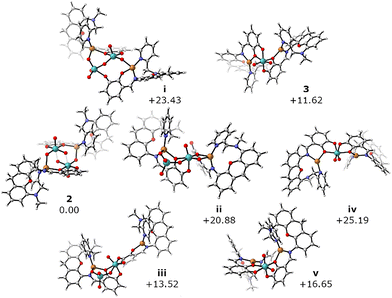 | ||
| Fig. 3 Geometry optimized structures with relative free energies (kcal mol−1). Molecular structures created with CylView.42 | ||
With asymmetric structures i–iii, one can imagine molybdate being lost from each of these intermediates. Deletion of molybdate and optimization of the resulting neutral structures resulted in trimetallic species 3, iv, and v from i, iii, and iv, respectively (Fig. 3). 3 has the non-Cu bound catecholate oxygens trans, iv has the Cu-bound catecholate oxygens trans, and v has all three types of oxygens cis to one another. Based on the trans influence, the weakest donor (Cu-bound catecholate) should be trans to the oxo functionalities. Indeed, 3 is the lowest in energy followed by v (+5 kcal mol−1), with iv much more endergonic (+14 kcal mol−1); this preference is observed in the crystal structure. While 2 → 3 + [MoO4]2− was computed to be endergonic by 11.62 kcal mol−1, our model cannot account for precipitation that helps drive this irreversible reaction. Analogous calculations for the benzenedithiolate analogue of our ligand suggest that this reaction would become exergonic by 21.64 kcal mol−1 in a ligand environment more similar to the enzyme (see Fig. S28). Furthermore, removal of Cu(I) from our catecholate system leads to the overall exergonic (−6.53 kcal mol−1) transformation ([Mo2O6(Lcat)2]4− → [MoO2(Lcat)2]2− + [MoO4]2−), suggesting that Cu(I) is not required for this reaction. The presence of Cu(I) allows us to observe the reactive tetrametallic intermediate, which would be undetectable in its absence.
Transformation of the tetranuclear complex [{MoO3(Lcat)Cu}2]2− into the trinuclear [MoO2(Lcat)2Cu2] and [MoO4]2− may explain the formation of bis(MPT) Mo in MobA. As stated above, the leading hypothesis proposed by Leimkühler and others was that the formation of the bis(MPT) Mo cofactor takes place via the disproportionation of [MoO3(MPT)]2−, along with formation of molybdate.8,9 Our results using a molecular model suggest that this transformation could be preceded by formation of the homodimer held together by Mo–O–Mo interactions (Fig. 4). While the enzyme likely does not utilize a secondary metal like Cu(I) to organize this dimer, this role could be played by the enzymatic environment in MobA. We also note that XAS indicated that the final product in MobA was [MoVO(MPT)], whereas we observe [MoVIO2(cat)2]2− as the product of disproportionation. It is feasible that the immediate product of MobA-catalyzed disproportionation, Mo(VI) dioxo, undergoes spontaneous reduction under physiological conditions. A related bis(MPT)Mo(VI) active site (of DMSO reductase) exhibited a Mo(VI)/Mo(V) redox couple at +161 mV vs. NHE at pH 8.43
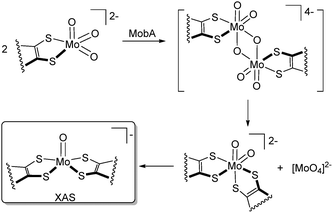 | ||
| Fig. 4 Possible mechanism of the transformation of the mono(MPT)Mo site into bis(MPT)Mo sites by MobA. | ||
In summary, we report the formation and spontaneous decomposition of a catecholate-ligated Mo(VI) trioxo complex. The complex was isolated in the dimeric “diamond-core” form stabilized by the presence of Cu(I). Kinetic measurements and DFT calculations suggest that its dimeric structure is likely responsible for the unimolecular disproportionation reaction that it undergoes, 2[{MoO3(Lcat)Cu}2]2− → [MoO2(Lcat)2Cu2] and [MoO4]2−. This reaction may serve as a functional model of the previously hypothesized disproportionation process in MobA, 2[MoO3(MPT)]2− → [MoO2(MPT)2]2− + [MoO4]2−, that is responsible for the formation of all bis(MPT) Mo active sites.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: SI includes NMR and mass spectra, X-ray collection and refinement details, computational details, and xyz structures for all optimized geometries. See DOI: https://doi.org/10.1039/d6cc00178e.CCDC 2521145 and 2521146 contain the supplementary crystallographic data for this paper.44a,b
Acknowledgements
S. G. and R. L. L. are grateful to the National Science Foundation (NSF) for current support through CHE-2348382. R. L. L. acknowledges the NSF for computational resources through MRI CHE-1919571 to the Midwest Undergraduate Computational Chemistry Consortium. Experimental characterization was carried out at the Lumigen Instrument Center of Wayne State University. This work made use of single-crystal XRD that was partially funded by the National Institutes of Health supplement grant #3R01EB027103- 02S1.References
- R. Hille, J. Hall and P. Basu, The Mononuclear Molybdenum Enzymes, Chem. Rev., 2014, 114, 3963–4038 CrossRef CAS PubMed.
- C. S. Seelmann, M. Willistein, J. Heider and M. Boll, Tungstoenzymes: Occurrence, Catalytic Diversity and Cofactor Synthesis, Inorganics, 2020, 8, 44 CrossRef CAS.
- R. R. Mendel and S. Leimkühler, The Biosynthesis of the Molybdenum Cofactors, J. Biol. Inorg. Chem., 2015, 20, 337–347 CrossRef CAS PubMed.
- C. Probst, J. Yang, J. Krausze, T. W. Hercher, C. P. Richers, T. Spatzal, K. Kc, L. J. Giles, D. C. Rees, R. R. Mendel, M. L. Kirk and T. Kruse, Mechanism of molybdate insertion into pterin-based molybdenum cofactors, Nat. Chem., 2021, 13, 758–765 CrossRef CAS PubMed.
- C. A. Temple and K. V. Rajagopalan, Mechanism of assembly of the bis(molybdopterin guanine dinucleotide)molybdenum cofactor in rhodobacter sphaeroides dimethyl sulfoxide reductase, J. Biol. Chem., 2000, 275, 40202–40210 CrossRef CAS PubMed.
- M. W. Lake, C. A. Temple, K. V. Rajagopalan and H. Schindelin, The crystal structure of the Escherichia coli MobA protein provides insight into molybdopterin guanine dinucleotide biosynthesis, J. Biol. Chem., 2000, 275, 40211–40217 CrossRef CAS PubMed.
- S. Reschke, B. R. Duffus, P. Schrapers, S. Mebs, C. Teutloff, H. Dau, M. Haumann and S. Leimkühler, Identification of YdhV as the First Molybdoenzyme Binding a Bis-Mo-MPT Cofactor in Escherichia coli, Biochemistry, 2019, 58, 2228–2242 CrossRef CAS PubMed.
- S. Reschke, K. G. V. Sigfridsson, P. Kaufmann, N. Leidel, S. Horn, K. Gast, C. Schulzke, M. Haumann and S. Leimkühler, Identification of a bis-molybdopterin intermediate in molybdenum cofactor biosynthesis in Escherichia coli, J. Biol. Chem., 2013, 288, 29736–29745 CrossRef CAS PubMed.
- K. Tiedemann, C. Iobbi-Nivol and S. Leimkühler, The Role of the Nucleotides in the Insertion of the bis-Molybdopterin Guanine Dinucleotide Cofactor into apo-Molybdoenzymes, Molecules, 2022, 27, 2993 CrossRef CAS PubMed.
- H. Dobbek, L. Gremer, R. Kiefersauer, R. Huber and O. Meyer, Catalysis at a Dinuclear [CuSMo(=O)OH] Cluster in a CO Dehydrogenase Resolved at 1.1-Å Resolution, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 15971–15976 CrossRef CAS PubMed.
- A. Majumdar, Structural and Functional Models in Molybdenum and Tungsten Bioinorganic Chemistry: Description of Selected Model Complexes, Present Scenario and Possible Future Scopes, Dalton Trans., 2014, 43, 8990–9003 RSC.
- N. P. Mankad and D. Ghosh, Biomimetic Studies of the Mo/Cu Active Site of CO Dehydrogenase, In Comprehensive Coordination Chemistry III, ed. E. C. Constable, G. Parkin, L. Que Jr, Elsevier, 2021, vol. 8, pp. 772–789 Search PubMed.
- U. I. Kaluarachchige Don, A. M. B. Chandima, D. Gelman, R. L. Lord and S. Groysman, Design and reactivity of early-late bimetallics as structural and functional models of Mo-Cu CODH, Comments Inorg. Chem., 2024, 44, 349–384 CrossRef CAS.
- C. Gourlay, D. J. Nielsen, J. M. White, S. Z. Knottenbelt, M. L. Kirk and C. G. Young, Paramagnetic Active Site Models for the Molybdenum−Copper Carbon Monoxide Dehydrogenase, J. Am. Chem. Soc., 2006, 128, 2164–2165 CrossRef CAS PubMed.
- C. Gourlay, D. J. Nielsen, D. J. Evans, J. M. White and C. G. Young, Models for Aerobic Carbon Monoxide Dehydrogenase: Synthesis, Characterization and Reactivity of Paramagnetic MoVO(μ-S)CuI Complexes, Chem. Sci., 2018, 9, 876–888 RSC.
- M. Takuma, Y. Ohki and K. Tatsumi, Sulfido-Bridged Dinuclear Molybdenum−Copper Complexes Related to the Active Site of CO Dehydrogenase: [(Dithiolate)Mo(O)S2Cu(SAr)]2− (Dithiolate = 1,2-S2C6H4, 1,2-S2C6H2-3,6-Cl2, 1,2-S2C2H4), Inorg. Chem., 2005, 44, 6034–6043 CrossRef CAS PubMed.
- D. Ghosh, D. Sinhababu, B. D. Santarsiero and N. P. Mankad, A W/Cu Synthetic Model for the Mo/Cu Cofactor of Aerobic CODH Indicates That Biochemical CO Oxidation Requires a Frustrated Lewis Acid/Base Pair, J. Am. Chem. Soc., 2020, 142, 12635–12642 CrossRef CAS PubMed.
- S. Groysman, A. Majumdar, S.-L. Zheng and R. H. Holm, Reactions of Monodithiolene Tungsten(VI) Sulfido Complexes with Copper(I) in Relation to the Structure of the Active Site of Carbon Monoxide Dehydrogenase, Inorg. Chem., 2010, 49, 1082–1089 CrossRef CAS PubMed.
- A. M. B. Chandima, G. Sgro, S. M. Hilditch, U. I. Kaluarachchige Don, C. L. Ward, D. P. Anderson, L. Herman, D. Gelman, R. L. Lord and S. Groysman, Heterodinuclear Cu(I)/Mo(VI) Chemistry with Bifunctional Dibenzobarrelene Ligands, Dalton Trans., 2025, 54, 14716–14727 RSC.
- W. Kaim and B. Schwederski, Non-innocent Ligands in Bioinorganic Chemistry—An Overview, Coord. Chem. Rev., 2010, 254, 1580–1588 CrossRef CAS.
- D. Coucouvanis, A. R. Paital, Q. Zhang, N. Lehnert, R. Ahlrichs, K. Fink, D. Fenske, A. K. Powell and Y. Lan, Synthesis, Electronic Structure, and Structural Characterization of the New, “Non-Innocent” 4,5-Dithio-Catecholate Ligand, Its Metal Complexes, and Their Oxidized 4,5-Dithio-o-quinone Derivatives, Inorg. Chem., 2009, 48, 8830–8844 CrossRef CAS PubMed.
- T. S. Hollingsworth, R. L. Hollingsworth, R. L. Lord and S. Groysman, Cooperative Bimetallic Reactivity of a Heterodinuclear Molybdenum-Copper Model of Mo-Cu CoDH, Dalton Trans., 2018, 47, 10017–10024 RSC.
- U. I. Kaluarachchige Don, S. S. Kurup, T. S. Hollingsworth, C. L. Ward, R. L. Lord and S. Groysman, S. Synthesis and Cu(I)/Mo(VI) reactivity of a bifunctional heterodinucleating ligand on a xanthene platform, Inorg. Chem., 2021, 60, 14655–14666 CrossRef CAS PubMed.
- U. I. Kaluarachchige Don, Z. Palmer, C. L. Ward, R. L. Lord and S. Groysman, Combining [MoVIO3] and [M0(CO)3] (M = Mo, Cr) fragments within the same complex: synthesis and reactivity of the single oxo- bridged heterobimetallics supported by xanthene-based heterodinucleating ligands, Inorg. Chem., 2023, 62, 15063–15075 CrossRef CAS PubMed.
- D. V. Partyka and R. H. Holm, Oxygen/Sulfur Substitution Reactions of Tetraoxometalates Effected by Electrophilic Carbon and Silicon Reagents, Inorg. Chem., 2004, 43, 8609–8616 CrossRef CAS PubMed.
- S. Groysman, J.-J. Wang, R. Tagore, S. C. Lee and R. H. Holm, A Biomimetic Approach to Oxidized Sites in the Xanthine Oxidoreductase Family: Synthesis and Stereochemistry of Tungsten(VI) Analogue Complexes, J. Am. Chem. Soc., 2008, 130, 12794–12807 CrossRef CAS PubMed.
- S. H. Vosko, L. Wilk and M. Nusair, Accurate Spin-Dependent Electron Liquid Correlation Energies for Local Spin Density Calculations: A Critical Analysis, Can. J. Phys., 1980, 58, 1200–1211 CrossRef CAS.
- C. Lee, W. Yang and R. G. Parr, Development of the Colle-Salvetti Correlation-Energy Formula into a Functional of the Electron Density, Phys. Rev. B, 1988, 37, 785–789 CrossRef CAS PubMed.
- A. D. Becke, Density-functional Thermochemistry. III. The Role of Exact Exchange, J. Chem. Phys., 1993, 98, 5648–5652 CrossRef CAS.
- P. J. Stephens, F. J. Devlin, C. F. Chabalowski and M. J. Frisch, Ab Initio Calculation of Vibrational Absorption and Circular Dichroism Spectra Using Density Functional Force Fields, J. Phys. Chem., 1994, 98, 11623–11627 CrossRef CAS.
- E. Caldeweyher, S. Ehlert, A. Hansen, H. Neugebauer, S. Spicher, C. Bannwarth and S. Grimme, A Generally Applicable Atomic-Charge Dependent London Dispersion Correction, J. Chem. Phys., 2019, 150, 154122 CrossRef PubMed.
- F. Weigend and R. Ahlrichs, Balanced Basis Sets of Split Valence, Triple Zeta Valence and Quadruple Zeta Valence Quality for H to Rn: Design and Assessment of Accuracy, Phys. Chem. Chem. Phys., 2005, 7, 3297–3305 RSC.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, Universal Solvation Model Based on Solute Electron Density and on a Continuum Model of the Solvent Defined by the Bulk Dielectric Constant and Atomic Surface Tensions, J. Phys. Chem. B, 2009, 113, 6378–6396 CrossRef CAS PubMed.
- A. D. Becke, Density-Functional Exchange-Energy Approximation with Correct Asymptotic Behavior, Phys. Rev. A, 1988, 38, 3098–3100 CrossRef CAS PubMed.
- J. P. Perdew, Density-Functional Approximation for the Correlation Energy of the Inhomogeneous Electron Gas, Phys. Rev. B, 1986, 33, 8822–8824 CrossRef PubMed.
- F. Neese, Software Update: The ORCA Program System—Version 6.0, WIREs Comput. Mol. Sci., 2025, 15, e70019 CrossRef.
- F. Neese, An Improvement of the Resolution of the Identity Approximation for the Formation of the Coulomb Matrix, J. Comput. Chem., 2003, 24, 1740–1747 CrossRef CAS PubMed.
- F. Neese, F. Wennmohs, A. Hansen and U. Becker, Efficient, Approximate and Parallel Hartree-Fock and Hybrid DFT Calculations. A “Chain-of-Spheres” Algorithm for the Hartree-Fock Exchange, Chem. Phys., 2009, 356, 98–109 CrossRef CAS.
- M. Garcia-Ratés and F. Neese, Effect of the Solute Cavity on the Solvation Energy and Its Derivatives within the Framework of the Gaussian Charge Scheme, J. Comput. Chem., 2020, 41, 922–939 CrossRef PubMed.
- B. Helmich-Paris, B. de Souza, F. Neese and R. Izsák, An Improved Chain of Spheres for Exchange Algorithm, J. Chem. Phys., 2021, 155, 104109 CrossRef CAS PubMed.
- F. Neese, The SHARK Integral Generation and Digestion System, J. Comput. Chem., 2023, 44, 381–396 CrossRef CAS PubMed.
- C. Y. Legault, CYLview20, Université de Sherbrooke, 2020 (https://www.cylview.org) Search PubMed.
- K.-F. Aguey-Zinsou, P. V. Bernhardt, A. G. McEwan and J. P. Ridge, The first non-turnover voltammetric response from a molybdenum enzyme: direct electrochemistry of dimethylsulfoxide reductase from Rhodobacter capsulatus, J. Biol. Inorg. Chem., 2002, 7, 879–883 CrossRef CAS PubMed.
- (a) CCDC 2521145: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2qmg83; (b) CCDC 2521146: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2qmg94.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |