Open Access Article
Open Access ArticleStructure-guided engineering of membrane-binding regions for surfactant-free solubilization of direct electron transfer-type alcohol dehydrogenase
Konatsu
Ichikawa
a,
Taiki
Adachi
*a,
Tomoko
Miyata
bc,
Fumiaki
Makino
bcd,
Keiichi
Namba
bc,
Yuki
Kitazumi
a,
Osamu
Shirai
 a and
Keisei
Sowa
a and
Keisei
Sowa
 *a
*a
aDivision of Applied Life Sciences, Graduate School of Agriculture, Kyoto University, Sakyo, Kyoto 606-8502, Japan. E-mail: sowa.keisei.2u@kyoto-u.ac.jp
bGraduate School of Frontier Biosciences, The University of Osaka, Suita, Osaka 565-0871, Japan
cJEOL YOKOGUSHI Research Alliance Laboratories, The University of Osaka, Suita, Osaka 565-0871, Japan
dJEOL Ltd, Akishima, Tokyo 196-8558, Japan
First published on 12th March 2026
Abstract
Membrane-bound alcohol dehydrogenase (ADH) from Gluconobacter oxydans is a direct electron transfer-type biocatalyst for ethanol oxidation. To improve its bioelectrocatalysis, membrane-binding regions of ADH were predicted, resulting in the construction of a soluble ADH variant by enzyme engineering. The variant was purified and characterized using structural and bioelectrochemical approaches.
Biocatalysts, such as microbes and enzymes, offer high selectivity in chemical reactions and operate under mild conditions (neutral pH, room temperature, and normal pressure). In bioelectrocatalysis, biocatalysts are artificially integrated into electrochemical systems, enabling efficient conversion between chemical and electrical energies.1–8 In particular, directly coupled reactions between oxidoreductases and electrodes are called direct electron transfer (DET)-type bioelectrocatalysis.9–17 Because artificial electron mediators are not required, DET-type reactions are ideal systems with high biocompatibility and energy efficiency. Therefore, the reaction is considered a promising core technology for applications in biosensors,18,19 bioreactors,20,21 and biofuel cells.5,6,22–24
Among enzymes involved in DET-type reactions (DET enzymes), membrane-bound dehydrogenases from acetic acid bacteria have particularly been investigated. To metabolize extracellular substrates, acetic acid bacteria possess various membrane-bound respiratory chain dehydrogenases on the cell membrane.25,26 In previous studies, the DET activities of alcohol dehydrogenase (ADH),27 aldehyde dehydrogenase (ALDH),28 fructose dehydrogenase (FDH),29 gluconate dehydrogenase,30 and lactate dehydrogenase31 have been reported. They commonly have a membrane-bound cytochrome c subunit containing hemes c (C subunit), which is responsible for electron transfer to a biological electron acceptor, ubiquinone (UQ), in vivo or to the electrode in the DET-type reaction.
However, DET-type membrane-bound enzymes have several limitations in industrial applications. First, heterologous overexpression of membrane-bound enzymes using alternative hosts such as Escherichia coli and yeast is challenging, because membrane-bound enzymes generally have high hydrophobicity and are frequently inactivated due to misfolding.32 Moreover, high-yield expression of membrane-bound proteins requires optimization of signal sequences and membrane-binding regions. Secondly, the solubilization of membrane-bound enzymes using surfactants often requires prolonged incubation, sometimes extending overnight, leading to increased purification costs and adverse effects on the enzymes. Additionally, surfactants coexisting with enzymes complicate DET-type reactions. A previous study reported that surfactants could affect the surface concentration of enzymes and enzyme orientation in DET-type reactions.33,34
Effective methods for addressing these problems are the deletion of membrane-bound subunits or regions, aiming to construct solubilized variants. For ADH, ALDH, FDH, and glucose dehydrogenase from Burkholderia cepacia, the variants truncating the C subunits were constructed.35–38 However, deletion of the C subunit decreased enzyme activity because the intermolecular electron transfer in the C subunit is important to enhance catalytic activity.38 Therefore, in the solubilization of the DET enzyme, deletion of the minimal membrane-binding region is more desirable. Membrane-bound DET-type hydrogenases have been solubilized using trypsin, which digests the membrane-anchoring site, allowing purification without any surfactants.39 An FDH variant with a double deletion of the two membrane-binding regions was also constructed.34 To rationally design soluble enzymes focusing on the membrane-binding regions, it is essential to understand the structural and sequence information related to membrane binding.
We focused on ADH from Gluconobacter oxydans, which is one of the most important membrane-bound respiratory chain dehydrogenases involved in acetic acid fermentation. The entire structure of ADH was elucidated using cryo-electron microscopy (cryo-EM) in 2023 (Fig. 1A).40 ADH consists of three subunits: a large subunit (L subunit) containing a pyrroloquinoline quinone (PQQ) and heme c (heme cL), a C subunit containing three hemes c (referred to as hemes 1c, 2c, and 3c from the N-terminus), and a small subunit (S subunit). Furthermore, the UQ-binding pocket in the C subunit was identified via structural analysis. However, the membrane-binding regions of ADH have not yet been identified.
 | ||
| Fig. 1 (A) 3D structure of ADH (PDB: 8GY2). (B) Enlarged view of predicted membrane-binding regions around UQ and heme 1c. | ||
In this study, we aimed to elucidate the membrane-binding regions in ADH and to construct a surfactant-free solubilized ADH variant (sADH) without decreasing the original DET activity. We focused on the structure of ADH (Fig. 1A) and predicted three membrane-binding regions in the C subunit. Single and multiple variants lacking these regions were constructed. The localization of enzyme activity was investigated using soluble and membrane fractions. The variant with the highest solubilization efficiency was defined as sADH and purified without surfactants. Furthermore, sADH was characterized by structural analysis using cryo-EM and electrochemical measurements.
To predict the membrane-binding regions in ADH, we focused on the following three points: (1) surfactant-derived noise in the cryo-EM map, (2) hydrophobic amino acid residues around the UQ-binding site, and (3) hydrophobic side chains facing outward. Fig. S1A shows the cryo-EM map of ADH; the dotted circle indicates the noise attributed to the surfactants used for solubilization. Moreover, UQ was located near the surfactant-derived noise (Fig. S1B). This result indicates that the corresponding part of the C subunit is hydrophobic and is involved in membrane binding. Near the UQ, there are many hydrophobic amino acid residues whose side chains face outward (Fig. 1B), whereas hydrophobic residues are generally located inside the enzyme. Based on the structural characteristics of ADH, three candidate membrane-binding regions (region 1: P77–I78, region 2: W159–F174, and region 3: G212–M215) were selected. The overall sequence of the C subunit and the predicted membrane-binding regions are shown in Fig. S2. Because the deletion of membrane-binding regions could result in unexpected structural changes and a decrease in enzymatic activity, glycine linkers were used to replace the deletion. Based on the predicted structure by AlphaFold3,41 the following three mutations were designed to minimize significant impact on the backbone structure of the original template: Δ1, P77–I78 replaced with a di-glycine linker; Δ2, W159–F174 replaced with a tetra-glycine linker; and Δ3, deletion of G212–M215 without a glycine linker (Fig. S3).
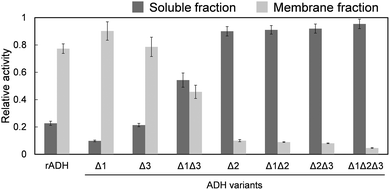
Wild-type recombinant ADH (rADH) and the single- and multiple-deleted ADH variants (Δ1, Δ2, Δ3, Δ1Δ2, Δ1Δ3, Δ2Δ3, and Δ1Δ2Δ3) were constructed. The total activities of ethanol oxidation in the soluble fractions and membrane fractions solubilized by Triton X-100 were investigated. Localization of the enzyme activity was evaluated using a crude enzyme solution (Fig. 2). Relative activity was calculated as the activity of each fraction divided by the total activity (soluble + membrane fractions). The ADH variants were arranged in order of increasing activity in the soluble fraction. While 77 ± 4% of the total activity was originally located in the membrane fraction of rADH, the Δ2 variant lacking the hydrophobic helical structure had 90 ± 3% of the total activity in the soluble fraction. In the Δ1 variant, the relative activity in the membrane fraction increased, indicating stronger binding between the enzymes and the membrane. This may be due to changes in the enzyme orientation on the membrane. Although the Δ1 and Δ3 variants exhibited no increase in relative activity in the soluble fraction, the solubilization efficiency of the Δ1Δ3 variant was higher than that of rADH. Therefore, regions 1 and 3 were only slightly involved in membrane binding. Among all the variants, the Δ1Δ2Δ3 variant showed the highest relative activity in the soluble fraction (95 ± 3%). Molecular dynamics (MD) simulations using the ADH–lipid complex also supported this discussion. After an adequate simulation period (300 ns), region 2 was located inside the lipid membrane, and regions 1 and 3 also faced the lipid membrane (Fig. S4). It should be noted that the present simulation does not include a quantum mechanical description of electrostatic interactions.
The variant exhibiting the highest relative activity in the soluble fraction (Δ1Δ2Δ3) was designated as solubilized ADH (sADH). We successfully purified it from the soluble fraction without any surfactants, whereas rADH was purified from the membrane fraction with 1% Triton X-100. The sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) results are shown in Fig. S5. Structural analysis of sADH was conducted using cryo-EM single particle image analysis. The analytical process is illustrated in Fig. S6. As shown in Fig. 3, two sADH structures were elucidated, and a conformational change in the structure around heme 1c was observed (Fig. S7). The region was suggested to be structurally flexible and unstable, given its lower local resolution compared with the rest of the structure (Fig. S6D and E). By contrast, most C subunits around hemes 2c and 3c were completely conserved. The specific activities of rADH and sADH were investigated using ferricyanide as an electron acceptor. The specific activity of sADH was 114 ± 5 U mg−1 and decreased to approximately one-third of that of rADH (323 ± 8 U mg−1). This can be attributed to the structural changes around heme 1c observed in the cryo-EM analysis which may have affected the redox reaction between the enzyme and ferricyanide.
Electrochemical measurements were performed on rADH and sADH. Multi-walled carbon nanotubes functionalized with carboxylic acid groups (CNT-COOH) were employed as porous electrode materials because carbon materials decorated with carboxylic acids have been reported to exhibit good compatibility with the DET-type reaction of ADH.40 A 10 µL aliquot of each enzyme solution (10 mg mL−1) was dropped onto CNT-COOH-modified glassy carbon electrodes (enzyme/CNT-COOH/GCE). As a control experiment to evaluate the effect of surfactants on sADH, sADH in the presence of 0.1% Triton X-100 was also adsorbed on electrodes using the same method. Fig. 4A shows the cyclic voltammograms (CVs) recorded at the enzyme/CNT-COOH/GCE. The statistical results are shown in Fig. S8. sADH/CNT-COOH/GCE showed clear DET activity, indicating that sADH was constructed and purified with maintaining its high DET activity. The current density of sADH/CNT-COOH/GCE was higher than that of rADH/CNT-COOH/GCE. This result indicates that the enzyme orientation of sADH on the electrode was improved compared with that of rADH owing to the mutation, which led to an increase in the surface concentration of the enzyme effective for DET. The trend in specific activity between rADH and sADH was reversed in the DET-type reaction. However, these two measurements involve different acceptors (ferricyanide and electrode) and electron transfer processes. Therefore, they cannot be directly compared.38 Focusing on the effect of surfactants, sADH/CNT-COOH/GCE prepared with an enzyme solution containing 0.1% Triton X-100 exhibited a slight increase in catalytic current density compared with sADH/CNT-COOH/GCE prepared without Triton X-100. An increase in specific activity of sADH in the presence of Triton X-100 was also observed in solution (Fig. S9), suggesting that the catalytic constant of sADH changes in the presence of Triton X-100. As shown in Fig. 4B, the stabilities of the electrodes were also investigated. The stability of sADH/CNT-COOH/GCE was slightly lower than that of rADH. This result suggests that the structural flexibility of sADH caused by mutations may lead to a decrease in the stability of catalytic current density. Moreover, sADH dissociation may be promoted by the co-adsorption of Triton X-100 on the electrode, resulting in a further decrease in stability. By contrast, the storage stability of the three enzymes was the same (Fig. S10). These results indicate that the mutation had no negative effects on the structural stability of sADH in solution.
To confirm the versatility of the solubilization approach used in this study, we investigated the sequence and structural homology between ADH and other DET enzymes from Gluconobacter sp. Here, we focused on ALDH and FDH, whose structures have already elucidated. The C subunits in ADH, ALDH, and FDH commonly have three hemes c and a UQ-binding site. The values of root-mean-square deviation (RMSD) with ADH were 1.142 and 1.534 Å for ALDH and FDH, respectively. As shown in Fig. 5, the structures of ADH, ALDH, and FDH exhibited high homology around their UQ-binding sites. Moreover, the structures corresponding to regions 1, 2, and 3 in ADH were highly conserved in ALDH and FDH, although they had additional clear membrane-bound helical structures. Structural conservation is generally considered to reflect functionally important regions.42 Therefore, the three regions investigated in this study may also play important roles in membrane binding of other DET enzymes.
In conclusion, three membrane-binding regions of ADH were predicted using the structural information. Based on the localization of enzyme activity and MD simulations, all regions were shown to be involved in membrane binding. sADH was successfully purified from the soluble fraction without surfactants, and structural and electrochemical characterizations were conducted. Although the C subunit has partial structural flexibility, sADH maintained the ethanol oxidation activity. Moreover, the sADH-modified electrode exhibited approximately twice the activity of the rADH-modified electrode. This study will lead to functional improvements in DET enzymes for industrial applications. This solubilization method will also help clarify the physiological functions of membrane-bound enzymes based on their structural homology. Moreover, incorporating additional computational approaches, such as density functional theory (DFT) calculations, would enable a more detailed discussion of the enzyme–lipid and enzyme–electrode interactions.
Conflicts of interest
There are no conflicts of interest.Data availability
The data supporting this article are included in the supplementary information (SI). Supplementary information: experimental section, cryo-EM analysis, molecular dynamics simulation, SDS-PAGE of the enzymes, structural alignment, and additional electrochemical data. See DOI: https://doi.org/10.1039/d6cc00143b.Acknowledgements
This research was supported by the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED under Grant Number JP25ama121003 to K. N., JSPS KAKENHI under grant numbers JP25K18412 to K. S. and JP24K17827 to T. A., the GteX Program of Japan under grant number JPMJGX23B4 to K. S., and FY 2022 Kusunoki 125 of Kyoto University 125th Anniversary Fund to K. S. We express our gratitude to Hirou Kaku, Koryu Ou, and Yasuyuki Hamano for their financial support. We also acknowledge the JEOL YOKOGUSHI Research Alliance Laboratories, The University of Osaka, for their support. We would also like to thank Editage (https://www.editage.com) for English language editing.References
- I. Willner, E. Katz and B. Willner, Electroanalysis, 1997, 9, 965–977 CrossRef CAS.
- K. Habermüller, M. Mosbach and W. Schuhmann, Fresenius’ J. Anal. Chem., 2000, 366, 560–568 CrossRef PubMed.
- S. Cosnier, A. J. Gross, A. Le Goff and M. Holzinger, J. Power Sources, 2016, 325, 252–263 CrossRef CAS.
- V. Fourmond and C. Léger, Curr. Opin. Electrochem., 2017, 1, 110–120 CrossRef CAS.
- N. Mano and A. de Poulpiquet, Chem. Rev., 2018, 118, 2392–2468 CrossRef CAS PubMed.
- X. Xiao, H. Q. Xia, R. Wu, L. Bai, L. Yan, E. Magner, S. Cosnier, E. Lojou, Z. Zhu and A. Liu, Chem. Rev., 2019, 119, 9509–9558 CrossRef CAS PubMed.
- H. Chen, O. Simoska, K. Lim, M. Grattieri, M. Yuan, F. Dong, Y. S. Lee, K. Beaver, S. Weliwatte, E. M. Gaffney and S. D. Minteer, Chem. Rev., 2020, 120, 12903–12993 CrossRef CAS PubMed.
- K. Kano, Biosci., Biotechnol., Biochem., 2022, 86, 141–156 CrossRef PubMed.
- C. Léger and P. Bertrand, Chem. Rev., 2008, 108, 2379–2438 CrossRef.
- M. Falk, Z. Blum and S. Shleev, Electrochim. Acta, 2012, 82, 191–202 CrossRef CAS.
- A. A. Karyakin, Bioelectrochemistry, 2012, 88, 70–75 CrossRef CAS PubMed.
- R. D. Milton and S. D. Minteer, J. R. Soc., Interface, 2017, 14, 20170253 CrossRef PubMed.
- P. Bollella, L. Gorton and R. Antiochia, Sensors, 2018, 18, 1319 CrossRef PubMed.
- I. Mazurenko, V. P. Hitaishi and E. Lojou, Curr. Opin. Electrochem., 2020, 19, 113–121 CrossRef CAS.
- F. A. Armstrong, Electrochim. Acta, 2021, 390, 138836 CrossRef CAS PubMed.
- O. Smutok, T. Kavetskyy and E. Katz, Curr. Opin. Electrochem., 2022, 31, 100856 CrossRef CAS.
- K. Sowa, J. Okuda-Shimazaki, E. Fukawa and K. Sode, Annu. Rev. Biomed. Eng., 2024, 26, 357–382 CrossRef CAS.
- D. R. Thevenot, K. Toth, R. A. Durst and G. S. Wilson, Biosens. Bioelectron., 2001, 16, 121–131 CrossRef CAS PubMed.
- P. Bollella and L. Gorton, Curr. Opin. Electrochem., 2018, 10, 157–173 CrossRef CAS.
- T. Krieg, A. Sydow, U. Schröder, J. Schrader and D. Holtmann, Trends Biotechnol., 2014, 32, 645–655 CrossRef CAS.
- R. D. Milton and S. D. Minteer, ChemPlusChem, 2017, 82, 513–521 CrossRef CAS PubMed.
- J. A. Cracknell, K. A. Vincent and F. A. Armstrong, Chem. Rev., 2008, 108, 2439–2461 CrossRef CAS PubMed.
- M. T. Meredith and S. D. Minteer, Annu. Rev. Anal. Chem., 2012, 5, 157–179 CrossRef CAS PubMed.
- I. Mazurenko, A. de Poulpiquet and E. Lojou, Curr. Opin. Electrochem., 2017, 5, 74–84 CrossRef CAS.
- C. Prust, M. Hoffmeister, H. Liesegang, A. Wiezer, W. F. Fricke, A. Ehrenreich, G. Gottschalk and U. Deppenmeie, Nat. Biotechnol., 2005, 23, 195–200 Search PubMed.
- J. Tkac, J. Svitel, I. Vostiar, M. Navratil and P. Gemeiner, Bioelectrochemistry, 2009, 76, 53–62 Search PubMed.
- T. Ikeda, D. Kobayashi, F. Matsushita, T. Sagara and K. Niki, J. Electroanal. Chem., 1993, 361, 221–228 Search PubMed.
- T. Adachi, Y. Kitazumi, O. Shirai and K. Kano, Electrochem. Commun., 2021, 123, 106911 Search PubMed.
- T. Ikeda, F. Matsushita and M. Senda, Biosens. Bioelectron., 1991, 6, 299–304 CrossRef CAS.
- S. Tsujimura, T. Abo, K. Matsushita, Y. Ano and K. Kano, Electrochemistry, 2008, 76, 549–551 CrossRef CAS.
- B. L. Treu and S. D. Minteer, Bioelectrochemistry, 2008, 74, 73–77 Search PubMed.
- F. Bernaudat, A. Frelet-Barrand, N. Pochon, S. Dementin, P. Hivin, S. Boutigny, J.-B. Rioux, D. Salvi, D. Seigneurin-Berny, P. Richaud, J. Joyard, D. Pignol, M. Sabaty, T. Desnos, E. Pebay-Peyroula, E. Darrouzet, T. Vernet, N. Rolland and H. W. van Veen, PLoS One, 2011, 6, e29191 Search PubMed.
- S. Kawai, T. Yakushi, K. Matsushita, Y. Kitazumi, O. Shirai and K. Kano, Electrochim. Acta, 2015, 152, 19–24 Search PubMed.
- T. Adachi, K. Ichikawa, T. Miyata, F. Makino, H. Tanaka, K. Namba and K. Sowa, ACS Electrochem., 2025, 1, 1649–1658 Search PubMed.
- K. Matsushita, T. Yakushi, H. Toyama, E. Shinagawa and O. Adachi, J. Biol. Chem., 1996, 271, 4850–4857 CrossRef CAS PubMed.
- S. Kawai, T. Yakushi, K. Matsushita, Y. Kitazumi, O. Shirai and K. Kano, Electrochem. Commun., 2014, 38, 28–31 CrossRef CAS.
- H. Yoshida, K. Kojima, M. Shiota, K. Yoshimatsu, T. Yamazaki, S. Ferri, W. Tsugawa, S. Kamitori and K. Sode, Struct. Biol., 2019, 75, 841–851 Search PubMed.
- K. Ichikawa, T. Adachi, Y. Kitazumi, O. Shirai and K. Sowa, ACS Catal., 2025, 15, 7283–7295 CrossRef CAS.
- Y. Higuchi, T. Yagi and N. Yasuoka, Structure, 1997, 5, 1671–1680 CrossRef CAS PubMed.
- T. Adachi, T. Miyata, F. Makino, H. Tanaka, K. Namba, K. Kano, K. Sowa, Y. Kitazumi and O. Shirai, ACS Catal., 2023, 13, 7955–7965 Search PubMed.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O’Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Nature, 2024, 630, 493–500 Search PubMed.
- C. Chothia and A. M. Lesk, EMBO J., 1986, 5, 823–826 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |