Open Access Article
Open Access ArticleAsymmetric synthesis of propargylic alcohols using bifunctional glucose dehydrogenases
Rohit Kumar,
Simran Bhatia,
Samantha J. Tambrini and
Guangde Jiang *
*
Department of Pharmaceutical Sciences, Eugene Applebaum College of Pharmacy and Health Sciences, Wayne State University, Detroit, MI 48201, USA. E-mail: guangde@wayne.edu
First published on 8th April 2026
Abstract
Herein, two bifunctional glucose dehydrogenases (GDHs) are reported that can efficiently catalyze the ketone reduction of α-ketoalkynes while regenerating NADPH for the ketone reduction process. The GDH-catalysed reaction can accept multiple types of α-ketoalkynes, including one particularly bulky substrate that typical alcohol dehydrogenases (ADHs) cannot efficiently work on. Notably, the developed biocatalytic approach allows gram-scale synthesis and the access to both (R) and (S) enantiomers of the products.
The last decade has witnessed a tremendous advancement in the field of biocatalysis.1 Unlike traditional chemical transformations, the biocatalytic approaches provide mild and green alternatives for the synthesis of complex molecules with excellent chemo- and regioselectivities. A new trend in recent years is the exploration of full biocatalytic potential of those well-studied types of enzymes through using nonnative substrates or special reaction conditions.2 It demonstrates that even those frequently studied enzymes can still surprise us with unexpected chemical transformations. Glucose dehydrogenases (GDHs) represent one such type of enzyme. It is primarily used for NAD(P)H regeneration for various biocatalytic reactions. However, in recent years, GDHs have been reported as a multifunctional biocatalyst in several chemical transformations3 such as carbonyl (aldehyde and ketone) reduction,4 imine reduction,4c,5 cyclohexanol dehydrogenation,6 and ketosteroid isomerization.6 Although GDH has been utilized as a multifunctional biocatalyst in a few chemical reactions, the full potential of GDH as a multifunctional biocatalyst still remains unexplored.
Propargylic alcohols represent a pivotal class of functionalized alkynes that have broad application in cyclization reactions, click chemistry, and as versatile building blocks for the synthesis of complex molecules.7 Notably, many bioactive compounds, including natural products8 and marketed pharmaceuticals,9 feature this functional motif. For example, the bioactive natural products falcarinol,10 panaxydol,11 and capillinol are all propargylic alcohols. In addition, therapeutic agents such as ethisterone, mestranol, levonorgestrel9 also contain a propargyl alcohol moiety (Fig. S1).
The chemical synthesis of chiral propargylic alcohols from readily available propargylic ketones has been reported in only a limited number of studies,12 with relatively few catalytic systems available to achieve high levels of enantioselectivity, highlighting the need for further development in this area. On the other hand, enzymatic synthesis of chiral propargylic alcohols offers an attractive alternative for achieving high regio- and enantioselectivity.
Despite prior efforts13 to utilize enzymes for the synthesis of chiral propargylic alcohols using propargylic ketones as substrates, these approaches have been constrained either by a limited substrate scope or by the necessity of multiple enzymes. To our knowledge, the enzymatic synthesis of chiral propargylic alcohols facilitated by a single enzyme has not been reported to date. Herein, we report that two bifunctional GDHs can efficiently synthesize various chiral propargylic alcohols using propargylic ketones as substrates.
Although previous studies have demonstrated that alcohol dehydrogenases (ADHs) can reduce alkynones to the corresponding alcohols,13a,b most of the reported substrates are relatively small, and the bulkier substrate containing two aromatic rings cannot be efficiently utilized by them. One such bulky propargylic ketone, 1a, poses a particular challenge for transformation by ADHs.12 Thus, we tested the substrate with RasADH from a Ralstonia species12 that have been reported to work on the reduction of propargylic ketones to alcohols and found only 18% conversion of the substate could be achieved while the enantioselectivity is relatively low (82![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 18 e.r.) (Table 1, entry 2). Interestingly, when we employed the GDH from Bacillus subtilis (BsGDH) for the reaction, excellent enantioselectivity (>99
18 e.r.) (Table 1, entry 2). Interestingly, when we employed the GDH from Bacillus subtilis (BsGDH) for the reaction, excellent enantioselectivity (>99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 e.r.) could be achieved despite the low conversion yield (Table 1, entry 3).
1 e.r.) could be achieved despite the low conversion yield (Table 1, entry 3).
| Entry | Variation from standard conditions | 2a yieldb (%) | e.r. |
|---|---|---|---|
| a (Standard conditions): 1a (5 mM), BsGDH_L95V (1 mol%), NADP+ (1 mM), D-glucose (100 mM), 50 mM NaPi buffer pH 7.0, 30 °C, 24 h, n.d. = not detected.b NMR yields using 1,1,2,2-tetrachloroethane as internal standard (Fig. S3). | |||
| 1 | None | 56 | >99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| 2 | RasADH | 18 | 82![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 18 18 |
| 3 | BsGDH | 23 | >99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| 4 | BsGDH_Q252K | 33 | >99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| 5 | OsGDH | 67 | >99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| 6 | Without BsGDH_L95V | n.d. | — |
| 7 | Without NADP+ | n.d. | — |
| 8 | Without D-glucose | n.d. | — |
| 9 | 50 mM NaPi buffer pH 6.0 | 19 | >99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| 10 | 50 mM NaPi buffer pH 8.0 | 33 | >99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
We further tested two mutants of BsGDH, BsGDH_L95V and BsGDH_Q252K (Fig. S2), which were reported to be able to selectively reduce cyclohexanone derivatives to corresponding alcohols with excellent efficiency.4d The results indicate that both mutants have excellent enantioselectivities (>99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 e.r.) while BsGDH_L95V could provide higher yield (Table 1, entries 1 and 4). Additionally, we found a GDH from Oscillatoria sp. (OsGDH) can also catalyze this reaction. The tested OsGDH was more efficient than BsGDH_L95V, affording the product in 67% yield (determined by NMR) with excellent stereocontrol (Table 1, entry 5). Various control experiments were performed to verify GDHs can indeed contribute to the transformation. When no enzyme was added to the reaction, no product was detected (Table 1, entry 6). Similarly, no product was observed when the reaction was carried out in the absence of NADP+ and D-glucose (Table 1, entries 7 and 8). We also found the pH value of the reaction buffer plays an essential role in influencing conversion yield, as both lowering and increasing the pH resulted in decreased product yields, while no impact on stereoselectivity was observed in either case (Table 1, entries 9 and 10).
1 e.r.) while BsGDH_L95V could provide higher yield (Table 1, entries 1 and 4). Additionally, we found a GDH from Oscillatoria sp. (OsGDH) can also catalyze this reaction. The tested OsGDH was more efficient than BsGDH_L95V, affording the product in 67% yield (determined by NMR) with excellent stereocontrol (Table 1, entry 5). Various control experiments were performed to verify GDHs can indeed contribute to the transformation. When no enzyme was added to the reaction, no product was detected (Table 1, entry 6). Similarly, no product was observed when the reaction was carried out in the absence of NADP+ and D-glucose (Table 1, entries 7 and 8). We also found the pH value of the reaction buffer plays an essential role in influencing conversion yield, as both lowering and increasing the pH resulted in decreased product yields, while no impact on stereoselectivity was observed in either case (Table 1, entries 9 and 10).
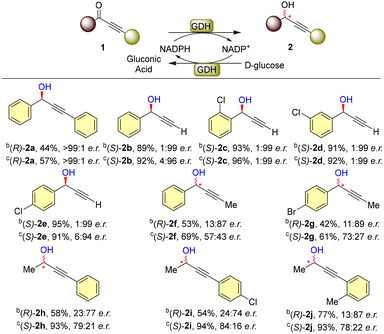
With the optimized reaction conditions, we explored the substrate scope of the developed reaction (Scheme 1) at 0.1 mmol scale and both enzymes (BsGDH_L95V and OsGDH) were tested for all the substrates. The substrate 1a worked well under the developed conditions providing the corresponding alcohol (R)-2a) in moderate isolated yields (44% with BsGDH_L95V and 57% with OsGDH) with excellent stereocontrol (>99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 e.r.). Terminal keto-alkynes (1b–e) worked very well under the developed conditions both in terms of yields and enantiomeric ratios with the two enzymes. The unsubstituted terminal alkynone (1b) provided the product (2b) in 89% and 92% yields with 99
1 e.r.). Terminal keto-alkynes (1b–e) worked very well under the developed conditions both in terms of yields and enantiomeric ratios with the two enzymes. The unsubstituted terminal alkynone (1b) provided the product (2b) in 89% and 92% yields with 99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 96
1 and 96![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 e.r., using BsGDH_L95V and OsGDH, respectively. Notably, substitutions on terminal keto-alkynes had minimal impact on the reaction outcome, as both GDHs provided excellent yields (91–96%) and outstanding enantiomeric ratios (up to >99
4 e.r., using BsGDH_L95V and OsGDH, respectively. Notably, substitutions on terminal keto-alkynes had minimal impact on the reaction outcome, as both GDHs provided excellent yields (91–96%) and outstanding enantiomeric ratios (up to >99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) of the products (2c–e). In addition to terminal alkynes, internal alkynes (1f–j) were also tested and were found tolerable for the developed methodology albeit in lower yields and lower e.r. compared to terminal keto-alkynes. Overall, OsGDH efficiently reduced the propargylic ketones and provided the corresponding products in better yields compared to BsGDH_L95V. Surprisingly, for internal ketoalkynes, BsGDH_L95V and OsGDH provided the alcohol products with different stereoselectivity. Internal ketoalkynes 1f–g led to the formation of products 2f–g in moderate yields with both GDHs. However, the stereochemical outcomes of the products differed, as BsGDH_L95V provided products with better enantiomeric ratios compared to OsGDH (2f and 2g).
1) of the products (2c–e). In addition to terminal alkynes, internal alkynes (1f–j) were also tested and were found tolerable for the developed methodology albeit in lower yields and lower e.r. compared to terminal keto-alkynes. Overall, OsGDH efficiently reduced the propargylic ketones and provided the corresponding products in better yields compared to BsGDH_L95V. Surprisingly, for internal ketoalkynes, BsGDH_L95V and OsGDH provided the alcohol products with different stereoselectivity. Internal ketoalkynes 1f–g led to the formation of products 2f–g in moderate yields with both GDHs. However, the stereochemical outcomes of the products differed, as BsGDH_L95V provided products with better enantiomeric ratios compared to OsGDH (2f and 2g).
The OsGDH-catalyzed reduction was significantly influenced by substituent effects, as the reduction of 1f and 1g provided the products with distinct enantiomeric ratios. Furthermore, para substitution on the aryl ring of the substrate contributed to increased enantioselectivity, as a higher enantiomeric ratio was obtained with substrate 1f compared to the unsubstituted substrate 1g. Other types of internal alkynones (1h–j) were also tested with both the BsGDH mutant and OsGDH. BsGDH_L95V provided the products (2h–j) in moderate yields whereas excellent yields were obtained when OsGDH was utilized. For the internal ketoalkynes (1h–j), moderate stereoselectivity was observed with both BsGDH_L95V and OsGDH.
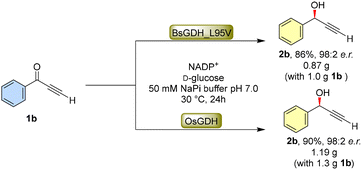
To demonstrate the efficiency and scalability of the current biocatalytic methodology, gram-scale reactions were performed (Scheme 2) using substrate 1b with slightly modified reaction conditions. Specifically, following our laboratory's typical procedures for expressing BsGDH_L95V or OsGDH in Escherichia coli (E. coli) BL21 cells, the corresponding wet cells containing either BsGDH_L95V or OsGDH were collected and utilized for the biocatalytic reduction of substrate 1b. The corresponding product (2b) was obtained in 86% isolated yield for BsGDH_L95V and 90% isolated yield for OsGDH. Both GDHs can lead to the product with enantiomeric ratio of 98![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, highlighting the scalability and effectiveness of the enzymatic reaction.
2, highlighting the scalability and effectiveness of the enzymatic reaction.
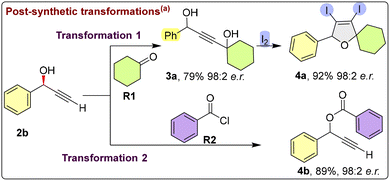
To demonstrate the synthetic utility of the enantioenriched propargylic alcohols, representative post-synthetic transformations were investigated (Scheme 3). The chiral alkynyl alcohol 2b was converted into an iodine-containing five-membered cyclic alcohol14 (4a) through a two-step sequence, proceeding via an isolable intermediate (3a) obtained in good yield and high enantiomeric excess, followed by iodine-promoted tandem cyclization without erosion of stereochemical integrity. In addition, benzoyl protection of the hydroxyl group of 2b furnished ester 4b in high yield while fully preserving enantioselectivity.15
Collectively, these transformations underscore the robustness of the stereocenter and highlight the potential of these products as versatile chiral building blocks for further synthetic elaboration.
To have a better understanding of the reaction mechanism, we performed deuterium labelling experiment (Scheme 3). When the reaction was carried out in the presence of deuterated D-glucose, 91% deuterium incorporation was observed at α-position of the enzymatic product (2b-d1) (Scheme 4 and Fig. S4).
 | ||
| Scheme 4 Deuterium labelling experiment. Reaction conditions: 1b (0.1 mmol), BsGDH_L95V (1 mol%), NADP+ (1 mM), D-glucose 1,2,3,4,5,6,6-d7 (100 mM), 50 mM NaPi buffer pH 7.0, 30 °C, 24 h. | ||
Based on these results and previous reports8,12 about multifunctional GDHs, we propose a plausible reaction mechanism (Fig. S5). First, the cofactor NADP+ is reduced to NADPH by GDH. The substrate is then likely stabilized by the amino acid residues Ser and Tyr within the catalytic pocket. The reaction is initiated by hydride transfer from NADPH to the carbonyl carbon of the substrate, followed by the proton transfer from Tyr to the carbonyl oxygen to generate the corresponding alcohol. The oxidized cofactor NADP+ is subsequently regenerated to NADPH by GDH, thereby completing the catalytic cycle.
In summary, we have utilized the dual functionality of GDHs for the synthesis of various chiral propargylic alcohols. This dual role of GDHs streamlines the reaction process and enhances reaction efficiency. Moreover, this methodology enabled the synthesis of a wide range of chiral propargylic alcohols, especially terminal substrates, in high yields and excellent enantiomeric ratios. Remarkably, the bifunctional GDHs can not only work on those substrates of ADHs but also go beyond them and work on bulkier substrates. To demonstrate the practicality and scalability of the developed methodology, a gram-scale synthesis was also carried out using E. coli cells containing GDHs, affording the product in good yield and good enantioselectivity, thereby underscoring the method's potential for large-scale applications in synthetic chemistry. Furthermore, the synthetic utility of the enantioenriched propargylic alcohols was demonstrated through post-synthetic modifications. A representative substrate underwent a two-step iodine-promoted cyclization to afford an iodine-containing five-membered cyclic alcohol with retention of enantiomeric purity, while benzoyl protection of the hydroxyl group proceeded smoothly without loss of stereochemical integrity. In addition, mechanistic experiments were performed to elucidate the reaction mechanism. Overall, the current findings further shed light on the biocatalytic versatility of GDHs. It underlines that GDHs may have a broader capability for biocatalysis, and they could be further engineered to improve catalytic efficiency and substrate tolerance to enhance their utility for chemical transformations. With the advancement of machine learning algorithms and the increasing accumulation of knowledge in enzyme engineering, this will likely open up exciting opportunities to further explore and harness GDHs for various synthetic applications.
Conceptualization: R. K. and G. J. data curation: R. K., S. J. T. and G. J. formal analysis: R. K. and G. J. software: S. B. funding acquisition: G. J. investigation: all authors. methodology: R. K. and G. J. supervision: G. J. writing: R. K. and G. J.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: details on the synthesis of racemic alcohols, general reaction procedures, NMR spectra and HPLC spectrums. See DOI: https://doi.org/10.1039/d5cc07322g.Acknowledgements
We gratefully acknowledge financial support from Wayne State University for this study.References
- (a) U. Hanefeld, F. Hollmann and C. E. Paul, Chem. Soc. Rev., 2022, 51, 594–627 RSC; (b) S. P. France, R. D. Lewis and C. A. Martinez, JACS Au, 2023, 3, 715–735 CrossRef CAS PubMed; (c) E. L. Bell, W. Finnigan, S. P. France, A. P. Green, M. A. Hayes, L. J. Hepworth, S. L. Lovelock, H. Niikura, S. Osuna and E. Romero, Nat. Rev. Methods Primers, 2021, 1, 1–21 Search PubMed.
- (a) X. Huang, B. Wang, Y. Wang, G. Jiang, J. Feng and H. Zhao, Nature, 2020, 584, 69–74 CrossRef CAS PubMed; (b) X. Huang, J. Feng, J. Cui, G. Jiang, W. Harrison, X. Zang, J. Zhou, B. Wang and H. Zhao, Nat. Catal., 2022, 5, 586–593 CrossRef CAS; (c) G. Jiang, C. Huang, W. Harrison, H. Li, M. Zhou and H. Zhao, Angew. Chem., Int. Ed., 2023, 62, e202302125 CrossRef CAS PubMed; (d) L. Wang, Y. Wu, J. Hu, D. Yin, W. Wei, J. Wen, X. Chen, C. Gao, Y. Zhou and J. Liu, Nat. Commun., 2024, 15, 5737 CrossRef CAS PubMed.
- R. Kumar, S. J. Tambrini and G. Jiang, ChemBioChem, 2025, 26, e202400716 CrossRef CAS PubMed.
- (a) S. M. Husain, M. A. Schätzle, S. Lüdeke and M. Müller, Angew. Chem., Int. Ed., 2014, 53, 9806–9811 CrossRef CAS PubMed; (b) D. Conradt, M. A. Schätzle, S. M. Husain and M. Müller, ChemCatChem, 2015, 7, 3116–3120 Search PubMed; (c) S. Roth, P. Stockinger, J. Steff, S. Steimle, V. Sautner, K. Tittmann, J. Pleiss and M. Müller, ChemBioChem, 2020, 21, 2615–2619 Search PubMed; (d) G. Jiang, J. Lu, M. Zhou, W. Harrison and H. Zhao, ACS Catal., 2024, 14, 7074–7079 CrossRef CAS; (e) Q. Yan, X. Zhang, Y. Chen, B. Guo, P. Zhou, B. Chen, Q. Huang and J.-B. Wang, ACS Catal., 2022, 12, 6746–6755 CrossRef CAS; (f) J. González-Rodríguez, S. González-Granda, H. Kumar, O. Alvizo, L. Escot, H. C. Hailes, V. Gotor-Fernández and I. Lavandera, Angew. Chem., Int. Ed., 2024, 63, e202410283 CrossRef PubMed; (g) A. E. Wolder, C. M. Heckmann, P.-L. Hagedoorn, D. J. Opperman and C. E. Paul, ACS Catal., 2024, 14, 15713–15720 CrossRef CAS PubMed.
- (a) X. Yi, F. Kleinbeck, C. Ching, L. Boghospor, S. Gomes, O. Alvizo, T. Allmendinger, J. Fell, N. Subramanian and M. Li, ACS Catal., 2024, 14, 7087–7096 CrossRef CAS; (b) S. Roth, M. B. Kilgore, T. M. Kutchan and M. Müller, ChemBioChem, 2018, 19, 1849–1852 CrossRef CAS PubMed.
- J. Dai, Z. Wu, Z. Liu, C. Li, L. Zhu, H. Chen and X. Chen, Appl. Microbiol. Biotechnol., 2024, 108, 399 CrossRef CAS PubMed.
- (a) R. Abonia, D. Insuasty and K. K. Laali, Molecules, 2023, 28, 3379 CrossRef CAS PubMed; (b) H. C. Kolb and K. B. Sharpless, Drug Discovery Today, 2003, 8, 1128–1137 CrossRef CAS PubMed.
- (a) M. N. Islam, R. J. Choi, H. A. Jung, S. H. Oh and J. S. Choi, Arch. Pharmacal. Res., 2016, 39, 340–349 Search PubMed; (b) F. Liu, J. Zhong, S. Li, M. Li, L. Wu, Q. Wang, J. Mao, S. Liu, B. Zheng and M. Wang, J. Nat. Prod., 2016, 79, 244–247 CrossRef CAS PubMed.
- (a) F. Z. Stanczyk, D. F. Archer and B. R. Bhavnani, Contraception, 2013, 87, 706–727 CrossRef CAS PubMed; (b) L. McRobb, D. J. Handelsman, R. Kazlauskas, S. Wilkinson, M. D. McLeod and A. K. Heather, J. Steroid Biochem. Mol. Biol., 2008, 110, 39–47 Search PubMed.
- S. Purup, E. Larsen and L. P. Christensen, J. Agric. Food Chem., 2009, 57, 8290–8296 CrossRef CAS PubMed.
- M.-Y. Kim, B. Jeong, G.-S. Lee, H. Jeon, Y. M. Yang, H. Yang and Y.-H. J. I. I. Han, Int. Immunopharmacol., 2024, 128, 111565 Search PubMed.
- (a) N. Arai, H. Satoh, N. Utsumi, K. Murata, K. Tsutsumi and T. Ohkuma, Org. Lett., 2013, 15, 3030–3033 CrossRef CAS PubMed; (b) Y. Sun, C. Lu, B. Zhao and M. Xue, J. Org. Chem., 2020, 85, 10504–10513 Search PubMed; (c) K. Matsumura, S. Hashiguchi, T. Ikariya and R. Noyori, J. Am. Chem. Soc., 1997, 119, 8738–8739 Search PubMed.
- (a) S. González Granda, D. Méndez Sánchez, I. Lavandera and V. Gotor Fernández, ChemCatChem, 2020, 12, 520–527 Search PubMed; (b) X. Sang, F. Tong, Z. Zeng, M. Wu, B. Yuan, Z. Sun, X. Sheng, G. Qu, M. Alcalde and F. Hollmann, Org. Lett., 2022, 24, 4252–4257 CrossRef CAS PubMed; (c) T. Schubert, W. Hummel and M. Müller, Angew. Chem., Int. Ed., 2002, 41, 634–637 CrossRef CAS; (d) T. Schubert, W. Hummel, M. R. Kula and M. Müller, Euro. J. Org. Chem., 2001, 2001, 4181–4187 Search PubMed.
- (a) F. Yang, T. Jin, M. Bao and Y. Yamamoto, Tetrahedron, 2011, 67, 10147–10155 CrossRef CAS; (b) K.-G. Ji, H.-T. Zhu, F. Yang, A. Shaukat, X.-F. Xia, Y.-F. Yang, X.-Y. Liu and Y.-M. Liang, J. Org. Chem., 2010, 75, 5670–5678 CrossRef CAS PubMed; (c) K. G. Ji, H. T. Zhu, F. Yang, X. Z. Shu, S. C. Zhao, X. Y. Liu, A. Shaukat and Y. M. Liang, Chem. Eur. J., 2010, 16, 6151–6154 CrossRef CAS PubMed.
- X. Huang, T. de Haro and C. Nevado, Chem. Eur. J., 2009, 15, 5904–5908 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |