Open Access Article
Open Access ArticleRational design of a ruthenium–cupin complex as an artificial ketone reductase
Koki
Matsumoto
,
Souto
Kitazawa
,
Ryusei
Matsumoto
,
Yoshitsugu
Morita
and
Nobutaka
Fujieda
 *
*
Department of Applied Biological Chemistry, Graduate School of Agriculture, Osaka Metropolitan University, 1-1 Gakuen-cho, Naka-ku, Sakai-shi, Osaka 599-8531, Japan. E-mail: fujieda@omu.ac.jp
First published on 21st February 2026
Abstract
A ruthenium(p-cymene)–cupin complex functioning as an artificial ketone reductase was developed through structure-guided engineering. Refinement of the primary and secondary coordination spheres based on a 1-His metal-binding motif enabled efficient asymmetric transfer hydrogenation of trifluoroacetophenone in water, affording up to 95% ee and 92% coupling efficiency.
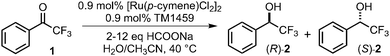
Optically active alcohols are important chiral building blocks that are widely used in the pharmaceutical, flavor, and fragrance industries.1–3 The development of efficient and stereoselective methods for their synthesis has long been a central challenge in the field of organic chemistry. Among various strategies, the asymmetric transfer hydrogenation (ATH) of ketones is one of the most effective approaches, with noble metal complexes of ruthenium (Ru), iridium (Ir), and rhodium (Rh) typically used as catalysts.4 Significant advances in ATH have been achieved since the pioneering work on asymmetric reduction using the Rh(I)-DIPAMP (DIPAMP = 1,2-bis[(2-methoxyphenyl)(phenylphosphino)]ethane) complex constructed by Knowles and co-workers5,6 and the Ru(II)-BINAP (BINAP = 2,2′-bis(diphenylphosphino)-1,1′-binaphthyl) complex reported by Noyori and co-workers.7 Subsequently, the Noyori–Ikariya catalyst, [RuCl(p-cymene)(TsDPEN)] (TsDPEN = N-(p-toluenesulfonyl)-1,2-diphenylethylenediamine), was developed as a more practical system, which is highly active toward prochiral ketones.8 This catalyst enables hydrogen transfer using isopropanol as both a hydrogen donor and a solvent, thus eliminating the need for high-pressure hydrogen gas.9,10 In this context, formic acid and sodium formate serve as attractive hydrogen sources in terms of sustainability, since they are inexpensive and easy-to-handle, producing only CO2 as a byproduct.11–13
For complementary catalytic systems, artificial metalloenzymes (ArMs) have attracted attention as an innovative class of hybrid catalysts that integrate the reactivity and tunability of transition-metal complexes with the biocompatibility and evolvability of protein scaffolds.14–18 Incorporating a metal complex into a protein framework enables catalysis in aqueous media, while the chiral environment of the active site affords excellent stereocontrol.19 Ward and co-workers developed a streptavidin-based system with a biotinylated Ru complex non-covalently incorporated, generating the first example of an artificial ketone reductase (AKRase).20 Since then, several other AKRase designs have been reported using sodium formate as a hydrogen donor,14,21 although competing H2 evolution can occur.11,22
Salmain and co-workers recently developed β-lactoglobulin- and lysozyme-based AKRases by direct coordination of Ru complexes to surface-exposed histidine (His) residues, achieving ATH activity in the presence of formate (Fig. 1A).23 Hartwig and co-workers and Ward and co-workers have more recently engineered human carbonic anhydrase24,25 and myoglobin,26 respectively, to create AKRases that use phenylsilane as an unconventional hydride source (Fig. 1A).
 | ||
| Fig. 1 (A) Recently reported AKRases. (B) Stepwise rational engineering of the Ru(p-cymene)-binding sites of TM1459 cupin proteins. | ||
We had previously engineered ArMs based on the homodimeric cupin protein TM1459 from Thermotoga maritima.27–30 This protein contains a 4-His metal-binding site that enables the straightforward incorporation of metal ions and is thus well suited for coordination chemistry.31 TM1459 exhibits high thermal stability, a compact molecular weight (13.1 kDa), and remarkable tolerance to amino-acid substitutions, which collectively make it an excellent platform for rational ArM development. In contrast to β-lactoglobulin and lysozyme, this protein possesses an appropriately sized hydrophobic cavity proximal to the metal-binding site that can encapsulate both the substrate and the Ru(p-cymene) complex within the protein interior rather than on the protein surface. We hypothesized that this could suppress nonspecific surface interactions and enable stereoselectivity to be controlled through spatial confinement. In this article, we present our rational engineering of a TM1459-based AKRase complexed with Ru(p-cymene) via site-directed mutation of the primary and secondary coordination spheres (Fig. 1B), resulting in a system capable of efficient and stereoselective ATH reaction.
First, we engineered the native 4-His tetrad of the TM1459 metal-binding site into a 2-His dyad that is well suited for anchoring the Ru(p-cymene) complex, mimicking the bidentate nitrogen ligand used in the Noyori–Ikariya catalyst. By mutating two of the four His residues (H52, H54, H58, and H92) to alanine, a panel of 2-His variants were prepared as reported previously (Table S1 and Fig. S1, S2, SI).28 However, none of these variants displayed clear ATH activity toward the aromatic ketone 2,2,2-trifluoroacetophenone 1 in the presence of Ru(p-cymene) (Table S2, SI). Therefore, we next expanded the protein cavity by producing 1-His “monodentate anchor” variants, in which three of the four His residues were replaced with alanine (Fig. 1B and Table S1 and Fig. S1, S2, SI).
In stark contrast to the 2-His variants, several 1-His mutants exhibited measurable catalytic activities and selectivities (Table 1, entries 2–5). In particular, the H54A/H58A/H92A mutant afforded the ATH product 2 with 74% (R) enantiomeric excess (ee) (Table 1, entry 2). Conversely, the H52A/H54A/H92A mutant (hereinafter 58H) produced the opposite S-enantiomer with 61% ee (Table 1, entry 4 and Fig. S3, SI). These results indicate that the position of the single coordinating His strongly dictates the chiral environment governing hydride delivery.
| Entry | TM1459 variant | Formate (equiv.) | Yield (%)c | eec (%) |
|---|---|---|---|---|
a Reaction conditions: TM1459 (0.15 mM), [Ru(p-cymene)Cl2]2 (0.15 mM), and substrate 1 (17 mM) in H2O/CH3CN (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) for 48 h at 40 °C and pH 5.
b Protein concentration was calculated based on the homodimeric form of TM1459.
c The yields and enantiomeric excesses (ee) were determined using chiral HPLC analysis.
d Potassium phosphate buffer (pH 6.0)/CH3CN (9 1) for 48 h at 40 °C and pH 5.
b Protein concentration was calculated based on the homodimeric form of TM1459.
c The yields and enantiomeric excesses (ee) were determined using chiral HPLC analysis.
d Potassium phosphate buffer (pH 6.0)/CH3CN (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) at 50 °C. 1) at 50 °C.
|
||||
| 1 | — | 12 | 47 | — |
| 2 | H54A/H58A/H92A | 12 | 63 | 74(R) |
| 3 | H52A/H58A/H92A | 12 | 41 | 54(S) |
| 4 | H52A/H54A/H92A (58H) | 12 | 31 | 61(S) |
| 5 | H52A/H54A/H58A | 12 | 21 | 7(R) |
| 6 | 58H/C106D | 12 | 33 | 4(R) |
| 7 | 58H/C106Q | 12 | 20 | 2(R) |
| 8 | 58H/C106H | 12 | 30 | 4(R) |
| 9 | 58H/C106M | 12 | 21 | 5(R) |
| 10 | 58H/C106E | 12 | 80 | 81(S) |
| 11 | 58H/C106E | 3 | 56 | 91(S) |
| 12 | R39M/58H/C106E | 3 | 43 | 88(R) |
| 13 | R39K/58H/C106E | 3 | 69 | 94(S) |
| 14d | 58H | 2 | 14 | 69(S) |
| 15d | 58H/C106E | 2 | 77 | 91(S) |
| 16d | R39M/58H/C106E | 2 | 54 | 85(R) |
| 17d | R39K/58H/C106E | 2 | 95 | 94(S) |
To further improve the performance of the 58H scaffold, we introduced an additional coordinating residue at Cys106, located adjacent to His58, and positioned it analogously to the bidentate ligand of the Noyori–Ikariya catalyst (Table S1 and Fig. 1B and Fig. S1, S2, SI). Substitutions with aspartate, glutamine, His, and methionine (Met) severely decreased the enantioselectivity (Table 1, entries 6–9), whereas 58H/C106E, bearing glutamate (Glu) at position 106, showed a substantial increase in both the yield and ee [80% yield, 81% ee (S); Table 1, entry 10]. Furthermore, when the sodium formate concentration was decreased from 12 to 3 equivalents, the activity of 58H dropped drastically (Table S3, entries 1–3), whereas 58H/C106E retained high stereoselectivity [91% ee (S)] and maintained a moderate yield (Table 1, entry 11 and Table S3, entries 4–6, SI). These observations suggest that Glu106 plays a critical role in activating the Ru center under low-formate conditions, possibly by tuning the local protonation environment and facilitating hydride transfer chemistry.
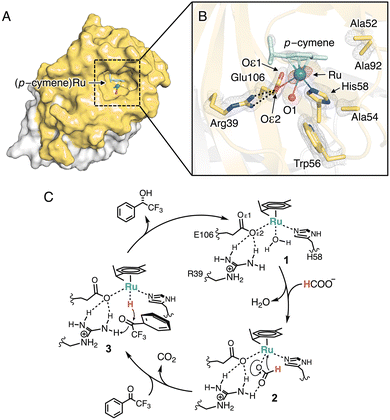
To elucidate the molecular basis of the enhanced reactivity of 58H/C106E, we determined its crystal structure at 1.26 Å resolution (PDB code: 9XS8, Fig. 2A, B and Tables S4–S6, SI). The overall fold was nearly identical to that of the Fe-bound wild type (RMSD = 0.135 over 197 Cα atoms; PDB code: 9JEU; Fig. S4A, SI). Moreover, a clear electron density corresponding to the Ru(p-cymene) complex, including an anomalous peak, was observed proximal to the Glu106 residue introduced in chain A (Fig. 2B). This structure revealed a well-defined Ru coordination environment: the Nε atom of His58 binds Ru (Ru–N, 2.14 ± 0.04 Å), whereas Glu106 engages in a monodentate κ1-Oε2(syn) interaction (Ru–Oε2, 3.13 ± 0.05 Å; Table S6, SI), and a single water molecule (O1) occupies another coordination site (Ru–O1, 2.81 ± 0.04 Å; Fig. 2B). The carboxylate group of Glu106 likely modulates the local charge distribution, thereby increasing the pKa of the coordinating water molecules and preventing their deprotonation. This facilitates ligand exchange from the water molecules to the formate ions, because hydroxide is less likely to dissociate from the Ru center. (Fig. 2C, 1→2).32,33
Unexpectedly, the Oε2 atom of Glu106 formed a stable hydrogen bond with the guanidino group of arginine at position 39 (Arg39), positioning the positively charged residue near the Ru center. This arrangement may assist in effective capture of a formate anion through hydrogen bonding during hydride abstraction from formate (Fig. 2C, 2). Furthermore, it may play a role in the hydride transfer and protonation for the substrate ketone 1 as in the case of the bifunctional Noyori–Ikariya catalyst (Fig. 2C, 3→1).34
To investigate the contributions of Arg39 and Glu106, we substituted Arg39 with either Met (which lacks hydrogen-bonding ability) or lysine (Lys, which maintains a basic side chain) (Table S1 and Fig. S1, S2, SI). At 3 equivalents of sodium formate, the R39M/58H/C106E variant showed a lower yield than that of 58H/C106E (Table 1, entry 12 and Fig. S3, SI), but higher activity than that of the parent 58H (Table S3, entry 3, SI), supporting the hypothesis that both Arg39 and Glu106 contribute to catalytic enhancement. Notably, R39M/58H/C106E produced the inverted R-enantiomer with 88% ee. In its crystal structure, the substrate-binding cavity was more spatially accessible around position 39 than that of 58H/C106E (Tables S4–S6 and Fig. S4BC, SI), possibly favoring the Si-face attack of substrate 1.
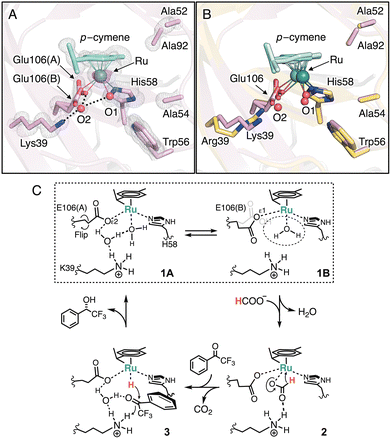
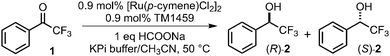
By contrast, the R39K/58H/C106E exhibited a higher yield (69%) and higher enantioselectivity [94% ee (S), Fig. S3, SI] than those of Arg39-containing 58H/C106E [56% yield, 91% ee (S); Table 1, entries 13 vs. 11]. The crystal structure of R39K/58H/C106E at 1.09 Å resolution revealed the presence of an additional water molecule (O2, Fig. 3A, B) that bridged the Nζ atom of Lys39, the Oε2 atom of Glu106(A), and Ru-coordinating water O1 through a hydrogen-bond network (Fig. 3C, 1A). Glu(A) engages in a monodentate κ1-Oε2(syn) interaction with Ru as in the case of 58H/C106E. Surprisingly, we also observed an alternative rotamer Glu106(B), coordinating with Ru in a monodentate κ1-Oε1(syn) manner and displacing the water molecule O2 (Fig. 3C, 1B). In this configuration, the O1-binding site became slightly more accessible, as the distance of Ru–Oε1 [Glu106(B), 2.47 ± 0.05 Å] was longer than that of Ru–Oε2[Glu106(A), 2.11 ± 0.02 Å], and the Oε(Glu106)–Ru–Nε(His58) bite angle increased (83.6 ± 0.7° → 87.7 ± 0.9°, Table S6 and Fig. 3C, 1B, dotted circle). This subtle opening may allow a closer approach of formate to the Ru center, thereby accelerating hydride abstraction (Fig. 3C, 2). Additionally, at neutral pH, the ammonium group of Lys is a slightly stronger proton donor than the guanidinium group of Arg, which may further facilitate protonation of substrate 1 (Fig. 3C, 3). Consistent with these structural features, the catalytic performance of R39K/58H/C106E exhibited pronounced pH dependence (Table S7, SI). At pH 6.0 and 50 °C, R39K/58H/C106E delivered 95% yield and 94% ee (S) (Table 1, entry 17), even with only 2 equivalents of sodium formate. Under these conditions, the order of yields (R39K/58H/C106E > 58H/C106E > R39M/58H/C106E > 58H; Table 1, entries 14–17) correlates well with the structural and mechanistic features described above.
To evaluate the chemical selectivity of the Ru-hydride species, we quantified the amount of formate remaining after the reaction at stoichiometric formate concentration and compared it with the alcohol yield (Table 2). A strong correlation was observed, indicating that hydride abstraction from formate was the rate-limiting step in the catalytic cycle (Fig. 3C, 2→3). Remarkably, R39K/58H/C106E achieved a 71% yield and 95% ee (S) even with 1 equivalent of sodium formate (Table 2, entry 4). Variants containing basic residues at position 39 (Arg or Lys) showed 92% and 84% coupling efficiency, respectively (Table 2, entries 2 and 4), indicating that the Ru-coordination geometry together with the hydrogen-bond network promoted hydride transfer to substrate 1 while suppressing H2 evolution (Fig. 3C, 3→1). Consistent with the stronger proton-donating ability of the ammonium group of Lys than the guanidium group of Arg, R39K/58H/C106E exhibited a slightly lower coupling efficiency than that of 58H/C106E despite its higher alcohol yield.
| Entry | TM1459 variant | Alcohol yield (%)c | ee (%)c | FC (%)d | CE (%)e |
|---|---|---|---|---|---|
a Reaction conditions: TM1459 (0.15 mM), [Ru(p-cymene)Cl2]2 (0.15 mM), substrate 1 (17 mM), and sodium formate (1 equivalent) in potassium phosphate buffer (pH 6.0)/CH3CN (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) for 48 h at 50 °C.
b Protein concentration was calculated based on the homodimeric form of TM1459.
c The yields and enantiomeric excesses (ee) were determined using chiral HPLC analysis.
d Formate consumption (FC) was determined by anion-exchange chromatography.
e Coupling efficiency [CE (%)] = [Product (mol)/formate ion consumed (mol)] × 100. 1) for 48 h at 50 °C.
b Protein concentration was calculated based on the homodimeric form of TM1459.
c The yields and enantiomeric excesses (ee) were determined using chiral HPLC analysis.
d Formate consumption (FC) was determined by anion-exchange chromatography.
e Coupling efficiency [CE (%)] = [Product (mol)/formate ion consumed (mol)] × 100.
|
|||||
| 1 | 58H | 12 | 73 | 26 | 46 |
| 2 | 58H/C106E | 58 | 88 | 63 | 92 |
| 3 | R39M/58H/C106E | 33 | 84 | 48 | 69 |
| 4 | R39K/58H/C106E | 71 | 95 | 84 | 84 |
In conclusion, we have demonstrated a rational and structure-guided strategy for developing Ru-based AKRases using the cupin protein TM1459 as a robust protein scaffold. Remodeling of the primary coordination sphere into a 1-His monodentate anchor site, followed by precise tuning of the secondary coordination sphere, enabled productive Ru(p-cymene) binding and efficient ATH activity in aqueous media. Glu106 incorporation and residue 39 modulation substantially enhanced the stereoselectivity, coupling efficiency, and catalytic turnover. Crystallographic analyses revealed a dynamic Ru coordination mode and hydrogen-bond network that governed Ru–hydride formation and stereocontrolled hydride transfer. Our study showcases a method for unlocking the full potential of ArMs-based synthetic systems and provides a general blueprint for developing biocompatible and environmentally benign ArMs capable of high-performance asymmetric reduction catalysis.
Koki Matsumoto: conceptualization, methodology, investigation, and writing – original draft. Souto Kitazawa: methodology and investigation. Ryusei Matsumoto: investigation and formal analysis (X-ray crystallography). Yoshitsugu Morita: investigation and formal analysis (X-ray crystallography). Nobutaka Fujieda: conceptualization, supervision, funding acquisition, and writing – review and editing.
Conflicts of interest
There are no conflicts to declare.Data availability
The protein structures were deposited in the Protein Data Bank: 58H/C106E structure (PDB code: 9XS8), R39K/58H/C106E structure (PDB code: 9XS9), and R39M/58H/C106E structure (PDB code: 9XSA).All data supporting the findings of this study are provided in the article and the supplementary information (SI). Supplementary information: general procedures, site-directed mutagenesis, protein expression and purification, protein X-ray crystallography, and chemical synthesis. See DOI: https://doi.org/10.1039/d5cc07188g.
Acknowledgements
N. F. thanks the JSPS and MEXT, Japan (JSPS KAKENHI Grant Numbers JP21H01954, JP24K01503, and JP25H02285) and The Asahi Glass Foundation (Research Encouragement Grant). R. M. thanks the JSPS and MEXT, Japan (JSPS KAKENHI Grant Number JP24KJ1907). We are deeply grateful to Prof. Genji Kurisu of the Institute for Protein Research, Osaka University, for his assistance with the crystallographic analysis and for kindly providing computational resources. This study was performed using the synchrotron beamline BL44XU at SPring-8 (Harima, Japan) under the Cooperative Research Program of the Institute for Protein Research, Osaka University (proposal numbers 2023B6832, 2024A6929, and 2025B6525). We thank Dr E. Yamashita, Dr K. Sakurai, and Dr A. Nakagawa of SPring-8 BL44XU for their support with the crystallographic data collection. We sincerely appreciate the generous guidance on DNA manipulation provided by Ms Noriko Kure of Osaka Metropolitan University.References
- E. Brenna, C. Fuganti, F. G. Gatti and S. Serra, Chem. Rev., 2011, 111, 4036–4072 CrossRef CAS PubMed.
- S. Wu, R. Snajdrova, J. C. Moore, K. Baldenius and U. T. Bornscheuer, Angew. Chem., Int. Ed., 2021, 60, 88–119 Search PubMed.
- I. Agranat, H. Caner and J. Caldwell, Nat. Rev. Drug Discovery, 2002, 1, 753–768 CrossRef CAS PubMed.
- D. Wang and D. Astruc, Chem. Rev., 2015, 115, 6621–6686 CrossRef CAS PubMed.
- B. D. Vineyard, W. S. Knowles, M. J. Sabacky, G. L. Bachman and D. J. Weinkauff, J. Am. Chem. Soc., 1977, 99, 5946–5952 CrossRef CAS.
- W. S. Knowles, Acc. Chem. Res., 1983, 16, 106–112 CrossRef CAS.
- R. Noyori, M. Ohta, Y. Hsiao, M. Kitamura, T. Ohta and H. Takaya, J. Am. Chem. Soc., 1986, 108, 7117–7119 CrossRef CAS.
- S. Hashiguchi, A. Fujii, J. Takehara, T. Ikariya and R. Noyori, J. Am. Chem. Soc., 1995, 117, 7562–7563 CrossRef CAS.
- R. Noyori and S. Hashiguchi, Acc. Chem. Res., 1997, 30, 97–102 CrossRef CAS.
- C. Wang, X. Wu and J. Xiao, Chem. – Asian J., 2008, 3, 1750–1770 CrossRef CAS PubMed.
- R. Nie, Y. Tao, Y. Nie, T. Lu, J. Wang, Y. Zhang, X. Lu and C. C. Xu, ACS Catal., 2021, 11, 1071–1095 Search PubMed.
- Y. Wei, Y. Liang, R. Luo and L. Ouyang, Org. Biomol. Chem., 2023, 21, 7484–7497 RSC.
- X. Wu and J. Xiao, Chem Comm, 2007, 2449–2466 RSC.
- F. Schwizer, Y. Okamoto, T. Heinisch, Y. Gu, M. M. Pellizzoni, V. Lebrun, R. Reuter, V. Köhler, J. C. Lewis and T. R. Ward, Chem. Rev., 2018, 118, 142–231 Search PubMed.
- Y. Yu, C. Hu, L. Xia and J. Wang, ACS Catal., 2018, 8, 1851–1863 CrossRef CAS.
- R. B. Leveson-Gower, C. Mayer and G. Roelfes, Nat. Rev. Chem., 2019, 3, 687–705 Search PubMed.
- C. Van Stappen, Y. Deng, Y. Liu, H. Heidari, J.-X. Wang, Y. Zhou, A. P. Ledray and Y. Lu, Chem. Rev., 2022, 122, 11974–12045 CrossRef CAS PubMed.
- I. Morita and T. R. Ward, Curr. Opin. Chem. Biol., 2024, 81, 102508 CrossRef CAS PubMed.
- T. Vornholt, F. Leiss-Maier, W. J. Jeong, C. Zeymer, W. J. Song, G. Roelfes and T. R. Ward, Nat. Rev. Methods Primers, 2024, 4, 78 Search PubMed.
- C. Letondor, N. Humbert and T. R. Ward, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 4683–4687 Search PubMed.
- S. T. Goralski and M. J. Rose, Curr. Opin. Chem. Biol., 2022, 66, 102096 CrossRef CAS PubMed.
- C. Luo, L. Li, X. Yue, P. Li, L. Zhang, Z. Yang, M. Pu, Z. Cao and M. Lei, RSC Adv., 2020, 10, 10411–10419 RSC.
- J. D. J. Cázares-Marinero, C. Przybylski and M. Salmain, Eur. J. Inorg. Chem., 2018, 1383–1393 CrossRef.
- P. Ji, J. Park, Y. Gu, D. S. Clark and J. F. Hartwig, Nat. Chem., 2021, 13, 312–318 CrossRef CAS PubMed.
- R. Chen, C. S. Kayrouz, E. McAmis, D. S. Clark and J. F. Hartwig, Angew. Chem., Int. Ed., 2024, 63, e202407111 Search PubMed.
- X. Zhang, D. Chen, J. Stropp, R. Tachibana, Z. Zou, D. Klose and T. R. Ward, Chemistry, 2024, 10, 2577–2589 CrossRef CAS.
- N. Fujieda, T. Nakano, Y. Taniguchi, H. Ichihashi, H. Sugimoto, Y. Morimoto, Y. Nishikawa, G. Kurisu and S. Itoh, J. Am. Chem. Soc., 2017, 139, 5149–5155 CrossRef CAS PubMed.
- N. Fujieda, H. Ichihashi, M. Yuasa, Y. Nishikawa, G. Kurisu and S. Itoh, Angew. Chem., Int. Ed., 2020, 59, 7717–7720 Search PubMed.
- R. Matsumoto, S. Yoshioka, M. Yuasa, Y. Morita, G. Kurisu and N. Fujieda, Chem. Sci., 2023, 14, 3932–3937 RSC.
- N. Fujieda, A. Matsuo and S. Itoh, Chem. – Eur. J., 2024, 30, e202402803 Search PubMed.
- N. Fujieda, K. Ishihama, H. Ichihashi, S. Yanagisawa, G. Kurisu and S. Itoh, Chem. – Asian J., 2025, 20, e202401191 CrossRef CAS PubMed.
- L. Dadci, H. Elias, U. Frey, A. Hoernig, U. Koelle, A. E. Merbach, H. Paulus and J. S. Schneider, Inorg. Chem., 1995, 34, 306–315 CrossRef CAS.
- S. Ogo, T. Abura and Y. Watanabe, Organometallics, 2002, 21, 2964–2969 CrossRef CAS.
- T. Ikariya and A. J. Blacker, Acc. Chem. Res., 2007, 40, 1300–1308 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |