 Open Access Article
Open Access ArticleA biocatalytic reticular framework via enzyme immobilization for environmental pollutant monitoring
Rui
Gao
a,
Xiaoxue
Kou
b,
Jing
Li
*c,
Siming
Huang
 d,
Jiehao
Yu
e,
Zhi-Wei
Li
d,
Jiehao
Yu
e,
Zhi-Wei
Li
 *e,
Guosheng
Chen
*e,
Guosheng
Chen
 b and
Gangfeng
Ouyang
b and
Gangfeng
Ouyang
 b
b
aInstitute for Advanced Study, Chengdu University, Chengdu 610106, People's Republic of China
bMOE Key Laboratory of Bioinorganic and Synthetic Chemistry, School of Chemistry, Sun Yat-sen University, Guangzhou 510275, People's Republic of China
cCollege of Food and Biological Engineering, Chengdu University, Chengdu 610106, People's Republic of China. E-mail: lijing@cdu.edu.cn
dGuangzhou Municipal and Guangzhou Province Key Laboratory of Molecular Target & Clinical Phamacology, the NMPA and State Key Laboratory of Respiratory Disease, School of Phamaceutical Sciences and the Fifth Affiliated Hospital, Guangzhou Medical University, Guangzhou 511436, People's Republic of China
eCollege of Chemistry and Materials, Jiangxi Normal University, Nanchang 330022, People's Republic of China. E-mail: lizhw69@mail.sysu.edu.cn
First published on 5th March 2026
Abstract
The growing prevalence of anthropogenic pollutants demands the advancement of environmental monitoring technologies. Biocatalytic sensing, leveraging enzymatic specificity and efficiency, offers a promising alternative, yet the structural fragility and instability of free enzymes limit its practical application. A transformative strategy involves immobilizing enzymes within engineered porous frameworks, such as metal–organic frameworks (MOFs) and covalent organic frameworks (COFs). These crystalline materials offer ultrahigh surface areas, tunable pore structures, and versatile surface chemistry, forming an ideal platform for constructing robust biohybrid sensing systems. They not only enable high enzyme loading but also establish a stabilized microenvironment that enhances enzymatic activity, stability, and reusability, while significantly improving catalytic selectivity and sensitivity. This review explores the strategic integration of enzymes with porous frameworks, detailing immobilization methodologies toward biosensing platforms. By examining their application in detecting pesticides, phenolic compounds, antibiotics, pathogens, and emerging contaminants, we highlight their potential to revolutionize environmental monitoring. Finally, we discuss current challenges and outline future directions to guide the development of sensitive, durable, and field-deployable analytical systems for sustainable environmental stewardship.
1. Introduction
The escalating release of environmental pollutants, fueled by rapid industrialization, population growth, and unsustainable production, has intensified ecological crises worldwide.1,2 The widespread use of pesticides,3 phenolics,4 and antibiotics,5 and emerging biological contaminants like pathogenic bacteria6 have contributed to pervasive environmental contamination. Regulatory mandates, such as those from the U.S. EPA, establish stringent maximum contaminant levels, necessitating the development of analytical methods capable of reliable, sensitive, and selective detection at trace concentrations (ppb to ppt) of pollutants in intricate matrices like wastewater, soil, and biological fluids.7 This scenario presents a quintessential analytical challenge, where achieving accurate quantification requires overcoming significant and variable background interference through exceptional selectivity and sensitivity.While conventional laboratory techniques such as chromatography coupled with mass spectrometry remain the gold standard for confirmatory multi-residue analysis due to their high accuracy, they are often limited by operational complexity, high cost, and poor suitability for real-time, on-site monitoring.8 Consequently, the field is increasingly shifting toward biosensing platforms that offer the potential for decentralized, rapid, and cost-effective environmental monitoring.
Enzyme-based biosensors represent a particularly promising alternative, leveraging the innate catalytic efficiency and molecular recognition specificity of enzymes, described as a “lock-and-key” mechanism, to achieve high selectivity even in complex sample environments.9–11 Their operation under mild, aqueous conditions further aligns with the principles of green analytical chemistry.12 However, the widespread deployment of native enzymes is significantly hampered by their intrinsic structural instability. Enzymes are conformationally labile biomacromolecules, and hence their catalytic activities are readily disrupted by non-physiological conditions commonly encountered in environmental samples, including extreme pH, temperature fluctuations, organic solvents, and chemical denaturants.13,14 This instability leads to signal drift, reduced shelf-life, and poor batch-to-batch reproducibility, compromising the accuracy, long-term reliability, and cost-effectiveness of the sensor, posing a major barrier to their real-world application.
Recent developments in enzyme immobilization have employed various nano- and microstructured carriers to improve enzymatic stability through spatial confinement, such as carbon-based materials, polymeric matrices, and porous nanoparticles.15–17 However, conventional supports often exhibit limited enzyme loading capacity and constrained pore accessibility, which consequently reduce catalytic efficiency. A transformative alternative has emerged with the use of porous reticular frameworks, including metal–organic frameworks (MOFs) and covalent organic frameworks (COFs).18–20 These materials offer an ideal platform for enzyme immobilization due to their high surface area, tunable porosity, and structural robustness.3,12,21–27 From an analytical standpoint, these frameworks deliver multiple critical advantages. First, the rigid porous architecture functions as a protective “exoskeleton” that confines enzymes and mitigates denaturation, thereby extending operational half-life and enhancing reusability. This directly translates into biosensors with improved durability, reproducibility, and cost-effectiveness.21,22 Second, well-defined mesopores facilitate efficient mass transfer of substrates and products, minimizing diffusion limitations that typically suppress apparent activity in conventional immobilized systems. As a result, catalytic turnover rates are maintained or even amplified, leading to stronger signal output and higher sensitivity. Moreover, the tailored pore dimensions can act as molecular sieves, selectively excluding interfering species from reaching the enzymatic active site.28,29 This “gatekeeping” function significantly enhances selectivity by reducing nonspecific interactions and background noise. Collectively, these attributes enable the design of robust, portable, and reusable biosensors suitable for decentralized, real-time environmental monitoring.
This review systematically summarizes recent advances in the rational design of biocatalytic reticular framework hybrids via enzyme immobilization for environmental sensing. We critically analyze key developments in functional integration strategies, such as surface adsorption, surface covalent anchoring, pore entrapment, and in situ encapsulation, and evaluate their impact on catalytic performance and operational stability. These engineered hybrids enable the highly sensitive and selective detection of a broad spectrum of environmental pollutants, including pesticides, antibiotics, phenolic compounds, and pathogenic microorganisms. Furthermore, the review identifies current limitations and outlines promising research directions aimed at enhancing real-world applicability. By integrating perspectives from materials science, analytical chemistry and environmental engineering, we provide a cross-disciplinary foundation for the next generation of smart, field-deployable biosensors.
2. Enzyme immobilization strategies
The enzyme immobilization strategies are broadly classified into two categories based on the enzyme's localization relative to the framework structure. This section will provide a concise overview of the distinct immobilization principles and their individual characteristics (Fig. 1).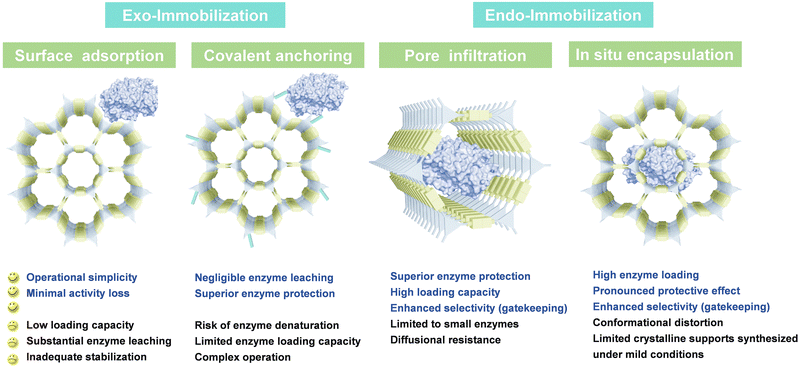 | ||
| Fig. 1 Schematic illustration of immobilization types of enzyme–porous reticular framework hybrids and their individual characteristics. | ||
2.1. Enzyme-on-porous reticular frameworks
Porous reticular frameworks, characterized by their tunable surface physicochemical properties and engineered functional groups, offer abundant binding sites for the surface bioconjugation of enzymes through supramolecular interactions. This approach demonstrates broad applicability and eliminates size constraints on the anchored biomolecules. In addition, it preserves the activity of the surface-immobilized enzymes due to minimal structural perturbation from the supporting matrix and optimal exposure to the surrounding microenvironment. This bioconjugation strategy primarily encompasses two mechanisms: surface adsorption via noncovalent interactions and covalent anchoring.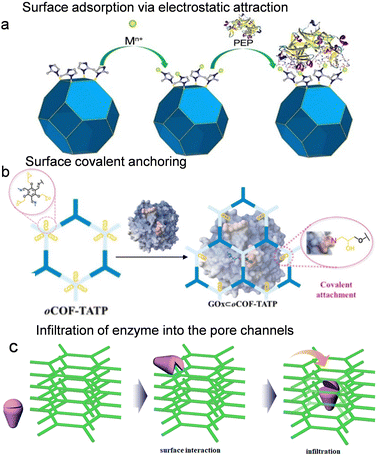 | ||
| Fig. 2 Enzyme immobilization on/into porous reticular frameworks via surface attachment and pore encapsulation methods. (a) Schematic illustration of surface adsorption via electrostatic attraction interactions of PEP to ZIF-8, reproduced from ref. 32 with permission from Wiley, Angew. Chem., Int. Ed., 2023, 62, e202216699, copyright 2023. (b) Covalent conjugation of glucose oxidase to COF carriers through surface-grafted epoxy groups, reproduced from ref. 34 with permission from American Chemical Society, Langmuir, 2025, 41, 11765–11775, copyright 2025. (c) Schematic illustration of the translocation of the enzyme into the pore channels, reproduced from ref. 35 with permission from American Chemical Society, J. Am. Chem. Soc., 2018, 140, 984–992, copyright 2018. | ||
From the standpoint of analytical performance, the retained activity, attributed to favorable enzyme orientation and reduced diffusion limitations for substrates, directly contributes to amplified signal generation, thereby lowering the limit of detection for target analytes and improving assay sensitivity. Nevertheless, the stability and reproducibility of such systems remain challenging under dynamic operational conditions. As mentioned earlier, environmental factors such as pH fluctuations and variations in ionic strength can dynamically perturb the adsorption equilibrium, potentially leading to substantial enzyme leaching over time. This desorption behavior poses a significant limitation for long-term or field applications.
2.2. Enzyme-in-porous reticular frameworks
This approach involves the encapsulation of enzymes within the nanopores or defective cavities of crystalline frameworks, fully leveraging the host architecture to provide physical protection and enhance biocatalyst stability under operational stressors.2.2.2.1. MOFs. ZIF-8 is widely used for enzyme encapsulation owing to its mild synthesis conditions (e.g., room temperature operation in aqueous media). It has been successfully applied to immobilize a variety of enzymes, including laccase (LAC),44 horseradish peroxidase (HRP),45 cytochrome c (Cyt c),46 acetylcholinesterase (AChE),47–49 and herbicide hydrolase.50 The resulting ZIF-8-enzyme composites have shown significant potential in the detection of pesticides and phenolic compounds, with the framework enhancing enzymatic stability under challenging operational conditions. However, ZIF-8 encapsulation faces several limitations: (1) restricted mass transport through its narrow pore apertures (∼3.4 Å), which hinders the diffusion of bulky substrates; and (2) structural instability under acidic conditions or in the presence of chelating agents. Recent studies suggest that such protective encapsulation behavior may extend beyond ZIF-8 to other MOFs, such as MOF-545-Fe,51 Tb-BTC,52 Zn-MOF-74,53 and UiO-66-F4,42 highlighting the broader role of MOFs in enhancing enzyme stability and facilitating substrate transport. For example, as illustrated in Fig. 3a, Zr4+ and tetrafluoroterephthalic acid can assemble into the ultrastable UiO-66-F4 under mild conditions, which was employed to co-encapsulate alkaline phosphatase (ALP) and HRP. The MOF shell constructed around the enzymes effectively preserved their activity under harsh environments, such as elevated temperature and extreme pH, while the proximity of the two enzymes facilitated efficient cascade catalysis. As a result, the ALP-HRP@MOF nanocomposites exhibited markedly improved catalytic efficiency, enhanced selectivity, and robust operational stability.42
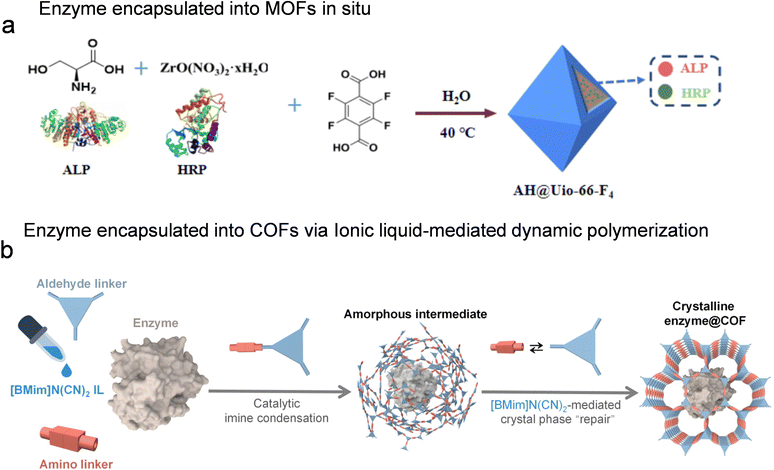 | ||
| Fig. 3 In situ encapsulation of enzyme in porous reticular frameworks. (a) Preparation of AH@UiO-66-F4in situ, reproduced from ref. 42 with permission from Elsevier, Food Chem., 2025, 493, 145602, copyright 2025. (b) Ionic liquid-mediated dynamic polymerization of COF, reproduced from ref. 29 with permission from Wiley, Angew. Chem., Int. Ed., 2024, 63, e202319876, copyright 2024. | ||
2.2.2.2. COFs. COFs exhibit superior chemical stability compared to MOFs due to their strong covalent linkages. However, conventional COF synthesis requires harsh conditions (elevated temperatures, organic solvents, and acid catalysts), which are not compatible for in situ enzyme encapsulation.28 To reduce the risk of enzyme denaturation during COF crystallization, Chen reported a MOF-based sacrificial templating strategy to construct enzyme–COF biocatalysts.54 This sacrificial templating concept can be extended to various COF hosts with similar properties. Although the mentioned template-protective assembly strategy could realize de novo encapsulation of enzymes into COFs, its complexity limits the extension to various COFs hosts. Their group reported a biofriendly and scalable one-pot synthesis strategy for enzyme–COF biocatalysts using acetic acid under aqueous solution, and this method could be extended to various COFs.55 However, due to the addition of enzyme-sensitive protonic acid, the practicality for different enzymes is still puzzled. Our group recently developed an aqueous-phase method that uses 2 µL of the 1-butyl-3-methylimidazolium dicyanamide ionic liquid catalyst to enable COF polymerization under enzyme-compatible conditions (Fig. 3b).29 The obtained highly crystalline enzyme–COF biocatalysts brought about the rigorous sieving effect and enhanced capability for degradation of pollutants compared to their poorly crystalline counterpart. In addition to liquid-phase methods, mechanochemistry provides new insights for fabricating enzyme–COF biocatalysts. This approach minimizes the consumption of organic solvents and strong acid/base catalysts, circumventing the harsh conditions that are typically necessary for COF crystallization and avoiding enzymes denaturation. In addition, we also report a mechanochemistry-guided assembly strategy for the construction of enzyme@COFs.56 The enzyme@COFs remain stable in a wider pH range (3–11), which expands the versatility and encourages further exploration in various environmental scenarios.
3. Environmental pollutant detection
The integration of enzymes with porous reticular frameworks has revolutionized the development of enzyme-based biosensors, greatly improving their performance in detecting environmental pollutants. These framework materials offer an ideal microenvironment for enzyme immobilization, which helps preserve catalytic activity and promotes efficient electron transfer and signal amplification. This section systematically examines the application of enzyme-immobilized porous reticular frameworks in the detection of diverse contaminants, including pesticides, phenolic compounds, antibiotics, microbial pathogens, and other emerging pollutants. Furthermore, the performance of representative hybrid biocatalysts is critically evaluated in terms of sensing efficacy (Table 1).| Pollutants | Targets | Porous reticular frameworks | Enzymes | Detection methods | Ranges | LOD | Ref. |
|---|---|---|---|---|---|---|---|
| Pesticides | Acephate | ZIF-8 | AChE/CHO/HRP | Colorimetric | 0.001–1 µM | 2.3 × 10−4 µM | 47 |
| Chlorpyrifos | ZIF-8 | AChE | Colorimetric | 1–50 nM | 0.61 nM | 64 | |
| Chemiluminescent | 1–40 nM | 0.13 nM | |||||
| Chlorpyrifos | Zn-MOF-74 | AChE | Colorimetric | 1.0–600 ng mL−1 | 1.0 ng mL−1 | 53 | |
| Carbaryl | COFThi-TFPB | AChE | Electrochemical | 2.2–60 µM | 0.22 µM | 66 | |
| Carbaryl | COFDhnda-BTH | AChE | Electrochemical | 0.48–35.0 µM | 0.16 µM | 67 | |
| DDVP | Ce/Zr-MOF | AChE/CHO | Colorimetric | 0.5–500 ng mL−1 | 0.32 ng mL−1 | 61 | |
| Ethyl paraoxon | ZIF-8 | AChE/HRP | Colorimetric | 6–800 ng mL−1 | 1.8 ng mL−1 | 49 | |
| Glyphosate | NH2-IMOF | AChE | Electrochemical | 1 × 10−15–1 × 10−9 M | 1.24 × 10−13 M | 68 | |
| Isocarbophos | ZIF-8 | AChE | Electrochemical | 0.5–100 ng mL−1 | 0.18 ng mL−1 | 65 | |
| Malathion | COF-DhaTab | AChE | Electrochemical | 1–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 nM 000 nM |
0.5 nM | 62 | |
| Malathion | COFTFPB-DBD | AChE | Electrochemical | 1 × 10−12–1 × 10−8 g L−1 | 3.0 × 10−13 g L−1 | 63 | |
| MP | ZIF-8 | AChE/CHO | Fluorescence | 1–5000 µg L−1 | 0.23 µg L−1 | 48 | |
| MP | La-MOF-NH2 | AChE | Electrochemical | 1.0 × 10−13–5.0 × 10−9 g mL−1 | 5.8 × 10−14 g mL−1 | 69 | |
| MP | MPNFs | AChE/CHO/HRP | Colorimetric | 0.1–1000 nM | 0.032 nM | 70 | |
| MP | MPNCs | AChE/CHO | Fluorescence | 0.05–1000 nM | 0.015 nM | 71 | |
| MP | p-COFs | AChE | Electrochemical | 1.9 × 10−9–3.8 × 10−5 M | 2.3 × 10−10 M | 72 | |
| MP | Tb-BTC | OPH | Fluorescence | 0.1–103 µM | 2.6 nM | 50 | |
| MP | UiO-66-NH2 | OPH | Fluorescence | 10–106 ng mL−1 | 10 ng mL−1 | 80 | |
| MP | Cu-MOF | Lipase | Electrochemical | 0.1–38 µM | 0.067 µM | 81 | |
| MP | ZIF-8 | Lipase BCL | Electrochemical | 0.1–38 mM | 0.28 mM | 82 | |
| MP | UiO-66 | Lipase BCL | Electrochemical | 5–40 µM | 0.84 µM. | 85 | |
| Trichlorfon | COF-LZU1 | AChE | Electrochemical | 0.2–19 ng mL−1 | 0.067 ng mL−1 | 73 | |
| Paraoxon | Aza-COFs | AChE | Electrochemical | 10–1000 ng mL−1 | 1.4 ng mL−1 | 74 | |
| Paraoxon | Cu3(THQ)2 | AChE | Electrochemical | 1–1000 ng mL−1 | 0.37 ng mL−1 | 75 | |
| Malathion Omethoate | ZIF-8 | AChE | Electrochemical | 3 pM-300 nM | 1.10 pM | 76 | |
| 4.69 pM-4.69 µM | 1.32 pM | ||||||
| Glyphosate | MOF-545-Fe | AChE | Electrochemical | 2.0–800 ng mL−1 | 0.65 ng mL−1 | 51 | |
| Omethoate | 0.5–800 ng mL−1 | 0.16 ng mL−1 | |||||
| Paraoxon | 1.0–800 ng mL−1 | 0.32 ng mL−1 | |||||
| MP | COF-Bta-NSs | AChE | Electrochemical | 5.0 × 10−13–1.0 × 10−8 g mL−1 | 2.04 × 10−13 g mL−1 | 77 | |
| Paraoxon | 1.0 × 10−12–1.0 × 10−8 g mL−1 | 7.94 × 10−13 g mL−1 | |||||
| Malathion | 5.0 × 10−11–1.0 × 10−7 g mL−1 | 5.37 × 10−12 g mL−1 | |||||
| Glyphosate | UiO-66-F4 | ALP/HRP | Colorimetric | 0.02–6.67 µM | 10.0 nM | 42 | |
| QpE | mZIF | QpeH/AOx | Colorimetric | 8.2–214.6 × 10−6 M | 2 × 10−6 M | 50 | |
| Nitrofen | UiO-66 | CRL | Electrochemical | 0–100 µM | 0.016 µM | 85 | |
| Nitrofen | ZIF-8 | BCL | Electrochemical | 0–114 µM | 0.46 µM | 86 | |
| Nitrofen | Cu-BTC | CRL | Electrochemical | 0–78 µM | 1.15 µM | 87 | |
| Nitrofen | COF-300 | CRL | Chemiluminescence | 0.02–50.0 µM | 11 nM | 90 | |
| Chlorpyrifos | HP-UiO-66-OH | Cyt c | Colorimetric | 10–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 pg mL−1 000 pg mL−1 |
4.63 pg mL−1 | 93 | |
| Imidacloprid | NSMOFs | HRP | Colorimetric | 0.02–1.1 ng mL−1 | 8.0 pg mL−1 | 91 | |
| Imidacloprid | MOFs-P | HRP | Smartphone | 0.001- 500 ng mL−1 | 0.0013 ng mL−1 | 92 | |
| Isocarbophos | COFs-PB | HRP | Colorimetric | 0.05–1000 ng mL−1 | 0.03 ng mL−1 | 90 | |
| Phenolic contaminants | Phenol | BHb | ZIF-8 | Colorimetric | 0–200 µM | 1 µM | 94 |
| Phenol | HRP/GOx | ZIF-8 | Colorimetric | 20–300 µM | 0.60 µM | 45 | |
| Phenol | HRP/GOx | H-ZIF-8- | Colorimetric | 0–100 µM | 0.86 µM | 96 | |
| Phenol | NiZn-MOF | Tyr | Electrochemical | 0.08–58.2 mM | 6.5 nM | 98 | |
| BPA | Cu-TCPP | Tyr | Electrochemical | 3.5 nM-8.9 µM | 1.2 nM | 97 | |
| BPA | ZIF-8 | LAC | Electrochemical | 0.01–0.4 mM | 1.95 × 10−3 mM | 44 | |
| BHA | ZIF-8 | HRP | Electrochemical | 0.1–1600 µM | 0.03 µM | 95 | |
| Catechol | ZIF-90 | LAC | Electrochemical | 20–400 µM | 1.86 µM | 99 | |
| Antibiotics | Tetracycline | UiO-66-NH2 | LAC | Electrochemical | 1.0 × 10−6–6.0 × 10−5 M | 8.94 × 10−7 M | 100 |
| Chloramphenicol | HRP | MAF-7 | Colorimetric | 0.005–6 ng mL−1 | 0.51 pg mL−1 | 101 | |
| Streptomycin | HRP | Ce/Zr-UiO66 | Colorimetric | 0.01–5000 ng mL−1 | 0.051 ng mL−1 | 102 | |
| Microbial pathogens and mycotoxins | Escherichia coli | Cyt c | ZIF-8 | Colorimetric | 1–106 CFU mL−1 | 1 CFU m−1 | 46 |
| Escherichia coli | GOx | mZIF-8 | Colorimetric | 10–108 CFU mL−1 | 10 CFU mL−1 | 103 | |
| S. aureus | 30–108 CFU mL−1 | 30 CFU mL−1 | |||||
| S. aureus | ZIF-90 | Urease | Colorimetric | 10–109 CFU mL−1 | 1.96 CFU mL−1 | 104 | |
| AFB1 | PCN-224 | HRP | Fluorescence and Colorimetric | 100 pg mL−1–10 ng mL−1 | 0.65 pg mL−1 | 109 | |
| AFB1 | HKUST-1 | GOx | Colorimetric | 0.01–0.1 ng mL−1 | 0.004 ng mL−1 | 105 | |
| Deoxynivalenol | GOx | PCN-224 | Fluorescence and Colorimetric | 0.01–1000 ng mL−1 | 0.0053 ng mL−1 | 108 | |
| 0–1000 ng mL−1 | 0.034 ng mL−1 | ||||||
| Other contaminants | As(V) | ACP/hemin | Zn-MOF | Fluorescence | 3.33–300 µg L−1 | 1.05 µg L−1 | 110 |
| H2S | PSSA-HRP | ZIF-8 | Chemiluminescence | 4.88–78.13 nM | 0.09 nM | 112 |
3.1. Pesticides
Pesticides have long been fundamental tools in global agriculture, playing a crucial role in safeguarding crop yield and quality by controlling pests, diseases, and weeds. However, the widespread and often excessive application of certain classes, such as organophosphorus pesticides (OPs) and carbamates, has led to their persistence in the environment and bioaccumulation across food chains, raising significant public health and ecological concerns.57,58 In the era of smart agriculture, which emphasizes precision and sustainability, the development of real-time, on-site monitoring methods for pesticide residues has become increasingly vital.3 To address this need, pesticide biosensors have evolved from conventional enzyme inhibition assays toward more sophisticated platforms, including those based on direct hydrolytic enzymes (e.g., OPH, QpeH, and lipase) and immunoassay-based detection (ELISA).AChE-based biosensors, whose detection mechanism for organophosphorus pesticides (OPs) is illustrated in Fig. 4a, represent one of the most extensively investigated platforms for pesticide detection.59,60 To overcome limitations in enzyme stability and signal amplification, recent studies have integrated AChE with MOFs and COFs. This integration substantially improves both the operational lifetime and shelf stability of the biosensors, which are crucial for field-deployable analytical devices.61–77 A representative example is the work by Cao et al., who co-encapsulated AChE, HRP, and choline oxidase (CHO) within a cruciate flower-like ZIF-8 via a one-step co-precipitation method (Fig. 4b). Using acetylcholine (ATCh) as the substrate, the pesticide acephate inhibits AChE activity, thereby disrupting the enzymatic cascade and decreasing hydrogen peroxide (H2O2) production, leading to a measurable color change for colorimetric detection (Fig. 4c). The well-defined porous structure of CF-ZIF-8 provided a favorable micro-environment that maintained enzyme activity and spatial organization. This multi-enzyme platform showed a good linear relationship between the acephate concentration and the inhibition rate in the range of 0.001–1 µM and achieved a detection limit of 0.23 nM for acephate (Fig. 4d), demonstrating high sensitivity.47
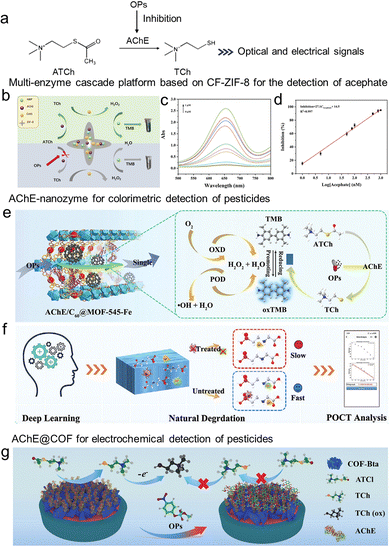 | ||
| Fig. 4 AChE inhibition-based sensors for pesticide detection through the immobilization of enzymes on porous reticular frameworks. (a) The mechanism of ACh enzyme inhibition reaction sensors for the detection of OPs. (b) Schematic diagram of a high-performance multi-enzyme cascade platform for detecting organophosphorus. (c) The ultraviolet spectra of organophosphorus at different concentrations detected by enzyme cascade platform. (d) The linear curve of inhibition rate and logarithm of acephate concentration, reproduced from ref. 47 with permission from Elsevier, Food Chem., 2022, 381, 132282, copyright 2022. (e) Principle of AChE/C60@MOF-545-Fe cascade flow reactor and (f) the application of a deep learning strategy, reproduced from ref. 51 with permission from Wiley, Adv. Funct. Mater., 2024, 35, 2419499, copyright 2024. (g) Detection mechanism of the electrochemical biosensing for OPs, reproduced from ref. 77 with permission from Wiley, Adv. Funct. Mater., 2023, 33, 2302917, copyright 2023. | ||
Beyond serving as immobilization supports, recent studies demonstrate a paradigm shift toward frameworks that actively participate in signal transduction, either through intrinsic enzyme-mimetic activity (e.g., Fe-based MOFs with peroxidase-like function) or by facilitating electron transfer in electrochemical systems. For example, Fe-based MOFs demonstrate peroxidase-like activity, enabling them to catalyze the oxidation of thiocholine (TCh), a hydrolysate of ATCh by AChE, into a chromogenic product. Inhibition of AChE by pesticides disrupts this catalytic process, producing a measurable signal change. Recent developments include multifunctional nanozymes such as C60@MOF-545-Fe, in which host–guest interactions enhance both oxidase- and peroxidase-like activities, enabling cascade catalysis without external energy input.51 The nanoconfined structure of this composite facilitates efficient molecular transport. When combined with AChE via supramolecular assembly, an enzyme–nanozyme continuous-flow reactor was constructed. This system not only allowed simultaneous detection of glyphosate, omethoate, and paraoxon but also discriminated among them via a three-channel sensor array (Fig. 4e). Furthermore, by integrating a YOLO v5-OPs deep learning model into a portable platform, direct terminal-based fitting equations were achieved, supporting rapid on-site analysis of organophosphorus pesticides (Fig. 4f). In parallel, confined catalytic systems such as V-TCPP(Fe)/AChE@ZIF have been designed for dual-mode colorimetric and chemiluminescence detection of pesticides like chlorpyrifos, achieving linear detection ranges of 1–50 nM (LOD 0.61 nM) and 1–40 nM (LOD 0.13 nM), respectively.64 Another colorimetric strategy leverages the reaction between TCh and 5,5′-dithiobis (2-nitrobenzoic acid) (DTNB), generating yellow 2-nitro-5-thiobenzoic acid (TNB) with a characteristic absorption at 412 nm. Based on this mechanism, AChE immobilized in Zn-MOF-74 was used to construct a biosensor that can detect chlorpyrifos with a limit of 1.0 ng mL−1, demonstrating high potential for trace-level quantification.53
In electrochemical biosensors, conductive MOFs or COFs serve as efficient electron-transfer mediators.74,75,77,78 When AChE is immobilized on such materials, the electroactive product thiocholine is directly oxidized on the conductive framework, promoting efficient electron transfer to the electrode surface. This “electron-wiring” effect enhances amperometric signal strength and overall detection sensitivity. For instance, ultrathin COF nanosheets (COF-Bta-NSs) with a thickness of approximately 1.95 nm and enriched nitrogen/sulfur-containing bithiazole units have been developed (Fig. 4g).77 The ordered porous structure with abundant unsaturated edge sites enables robust immobilization of AChE via supramolecular interactions and promotes local enrichment of analyte molecules. Moreover, the incorporation of N, S-rich bithiazole moieties enhances electrical conductivity, thereby improving biosensor detectability. Under optimized conditions, this biosensor exhibited exceptional performance in detecting OPs, featuring a broad detection range and ultralow detection limits (e.g., methyl parathion (MP): 2.04 × 10−13 g mL−1; paraoxon: 7.94 × 10−13 g mL−1; and malathion: 5.37 × 10−12 g mL−1), along with high selectivity and stability.
While AChE inhibition offers broad-spectrum detection, it inherently lacks selectivity among different pesticides. In contrast, direct hydrolytic enzymes provide superior specificity by recognizing distinct pesticide structures and catalyzing their hydrolysis. These enzymes enable the design of biosensing platforms that convert hydrolysis products into measurable signals, offering promising routes for selective and sensitive detection. Aryloxyphenoxypropionate (AOPP) herbicides are widely used for controlling grass weeds in broadleaf crops. Leveraging this specificity, Ma et al. developed a colorimetric biosensor by co-encapsulating a three-enzyme cascade that contains QpeH, alcohol oxidase (AOx), and peroxidase-mimicking magnetic ZIF-8 (mZIF-8) into a single framework.50 As outlined in Fig. 5a, the sensing mechanism involves the QpeH-catalyzed hydrolysis of the AOPP herbicide, followed by AOx oxidation of the released alcohol, with mZIF-8 finally catalyzing the formation of a colored product for detection. This biosensor demonstrated high sensitivity for quizalofop-P-ethyl (QpE), achieving a linear detection range of 8.2 to 214.6 µM and a detection limit of 2 µM.
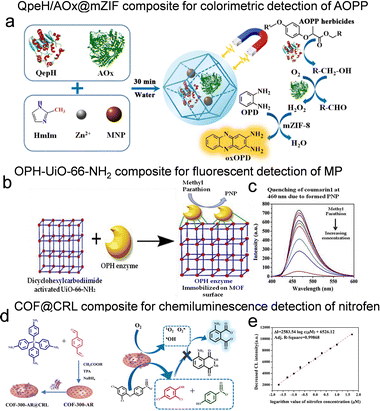 | ||
| Fig. 5 Direct catalytic enzyme-based sensors for pesticide detection through the immobilization of enzymes on porous reticular frameworks. (a) Schematic illustration of the one-pot synthesis of QpeH/AOx@mZIF composite with magnetic collection and its use as a biosensor for the rapid detection of AOPP herbicides, reproduced from ref. 50 with permission from American Chemical Society, ACS Appl. Mater. Interfaces, 2021, 13, 44329–44338, Copyright 2021. (b) Reaction mechanism of OPH6His/UiO-66-NH2 for fluorescence detection of MP. (c) The fluorescence intensity of MP at different concentrations, reproduced from ref. 80 with permission from Elsevier, Environ. Res., 2019, 174, 46–53, Copyright 2019. (d) Schematic diagram of COF-300-AR@CRL synthesis and chemiluminescence detection of nitrofen. (e) Linear calibration plot between the decreased chemiluminescence intensities and the logarithm values of nitrofen concentration in the 0.02–50.0 µM range, reproduced from ref. 88 with permission from Springer Nature. Microchim. Acta, 2023, 190, 492, Copyright 2023. | ||
MP, a highly toxic organophosphorus pesticide, poses significant risks to human health and the environment due to its neurotoxic effects and environmental persistence.79 Its detection has been effectively achieved using both OPH-based fluorescence sensing and lipase-based electrochemical approaches.52,80–83 For fluorescence-based detection, the immobilization of hexahistidine-tagged OPH (OPH6His) onto a zirconium-based MOF (UiO-66-NH2) has been demonstrated. In this system, the immobilized enzyme catalyzes the hydrolysis of MP to p-nitrophenol (PNP), with the PNP concentration being directly proportional to the original pesticide concentration (Fig. 5b). The fluorescent reporter coumarin 1 is subsequently introduced, whose emission is quantitatively quenched by PNP (Fig. 5c). This OPH6His/UiO-66-NH2 composite not only enhanced enzymatic activity by approximately 37% but also maintained stability over 60 days and allowed at least eight reuse cycles. The sensing platform achieved a wide detection range of 10–106 ng mL−1 with a quantification limit of 10 ng mL−1 and was successfully validated in spiked food samples.80 Electrochemical detection of MP has been realized using lipase immobilized on ZIF-8 and its amine-functionalized derivative. Lipase catalyzes the hydrolysis of MP to PNP, which is then electrochemically oxidized at the electrode interface. The lipase@ZIF-8/chitosan-modified glassy carbon electrode exhibited a low detection limit of 0.28 µM, a wide linear detection range, and good repeatability. The biosensor retained over 80% of its initial response after 20 days of storage, demonstrating excellent operational and storage stability. This system illustrates the effectiveness of MOF-immobilized lipase in facilitating both hydrolysis and subsequent electrochemical signal generation.82
As a representative diphenyl ether herbicide, nitrofen is widely used for weed control but poses considerable environmental and health hazards.84 Recent studies have reported the development of electrochemical enzyme sensors incorporating lipases embedded in materials such as UiO-66, ZIF-8 or Cu-BTC.85–88 These lipases catalyze the cleavage of the C–O–C bond in nitrofen, producing redox-active phenolic compounds. For example, Candida rugosa lipase (CRL) was immobilized on an amine-rich covalent organic framework (COF-300-AR) (Fig. 5d). The resulting COF-300-AR@CRL composite exhibited dual catalytic functions: the COF framework served as an oxidase mimic to catalyze the luminol-dissolved oxygen chemiluminescence reaction, while the immobilized CRL hydrolyzed nitrofen to release reducing phenolic compounds. These phenolic products effectively quenched the chemiluminescence signal, enabling the quantification of nitrofen in the range of 0.02–50.0 µM with a detection limit of 11 nM (Fig. 5e). This method displayed excellent precision (2.0% RSD for 50.0 nM nitrofen) and recyclability, and was successfully applied to vegetable and fruit samples.88
Despite the superior specificity of direct hydrolytic enzymes arising from their recognition of distinct pesticide structures, their practical application is hindered by a narrow spectrum of hydrolyzable substrates. ELISA presents a complementary and versatile alternative, offering robust analytical performance through the high specificity inherent in immunoreactions.89 This platform not only enables sensitive detection of environmental contaminants but also exhibits remarkable adaptability, as its target recognition can be readily tailored to diverse pesticides by simply exchanging the antibody employed. Yan et al. developed an enzyme-engineered assembly strategy for in situ enzyme encapsulation within COFs-PB via a covalent–polymerization–permeation mechanism.90 Capitalizing on this design, they fabricated an HRP@COFs-PB-based immunosensor for isocarbophos detection. In this competitive format, the coating antigen immobilizes on the plate, competing with the pesticide for primary antibody binding. The HRP-COFs-Ab2 conjugate recognizes the captured antibody and catalyzes the tetramethylbenzidine (TMB)/H2O2 chromogenic reaction, with signal intensity being inversely proportional to pesticide concentration (Fig. 6a). An obvious color attenuation of HRP-COFs composite-based ELISA is observed with an increase of isocarbophos concentration (Fig. 6b). Through G × B color processing, the authors achieved quantitative detection across 0.05–1000 ng mL−1 with a calculated LOD of 0.03 ng mL−1, outperforming other color-algorithm methods and highlighting its potential for on-site pesticide monitoring (Fig. 6c and d). Similarly, HRP immobilized in other MOFs has shown specific recognition capability in immunosensors for detecting imidacloprid.91,92 Notably, such immunosensors typically require secondary antibodies. However, Chen et al. developed an innovative immunoassay platform for colorimetric detection of chlorpyrifos (CPF) that circumvents this requirement.93 By modifying the ligand functional groups of MOFs, they precisely tuned the pore electronic properties and hydrophilic microenvironment, which enhanced both catalytic activity and stability of the immobilized enzyme system. The resulting HP-UiO-66-OH material not only serves as an enzyme immobilization platform but also functions as a recognition element for CPF through strong Zr–O–P interactions, thereby eliminating the need for secondary antibodies (Fig. 6e). As shown in Fig. 6f and g, the absorbance increased progressively with CPF concentration, exhibiting excellent linear correlation across the concentration range of 10–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 pg mL−1 when plotted against the logarithm of CPF concentration.
000 pg mL−1 when plotted against the logarithm of CPF concentration.
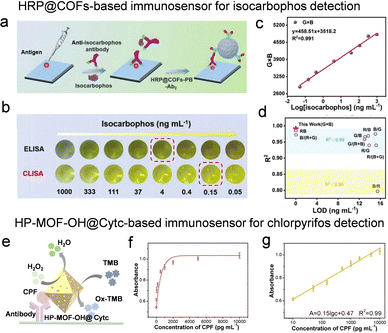 | ||
| Fig. 6 The immunosensor for pesticide detection through the immobilization of enzymes on porous reticular frameworks. (a) Schematic illustration of HRP@COFs-PB-based competitive immunosensor for imidacloprid detection. (b) The color change of HRP-COFs composite-based CLISA with the increase of isocarbophos concentration. (c) The relationship between the G × B value response and isocarbophos concentration. (d) Comparison of detection limits between the G × B algorithm and other algorithms, reproduced from ref. 90 with permission from Wiley, Adv. Funct. Mater., 2025, 35, 2420159, copyright 2025. (e) Schematic illustration of the HP-UiO-66-OH@Cytc-based immunoassay platform for CPF detection. (f) Calibration curve and (g) linear detection range of HP-UiO-66-OH@Cytc-based ELISA, reproduced from ref. 93 with permission from American Chemical Society, Anal. Chem., 2025, 97, 16019–16025, copyright 2025. | ||
A critical challenge in this field is the persistent disconnect between laboratory performance and real-world applicability. Despite exceptional detection limits reported under optimized conditions, most systems remain unvalidated in complex environmental matrices or under variable field conditions. Furthermore, the scalability and cost-effectiveness of these platforms have yet to be systematically explored.
3.2. Phenolic contaminants
Phenolic contaminants, extensively generated by industrial sectors such as textiles, plastics, and paper manufacturing, represent a major class of persistent environmental pollutants.4 Their significant ecological and human health risks, which arise from resistance to degradation, bioaccumulative potential, and multifaceted toxicity, underline the critical need for reliable analytical methods for their detection. Among various strategies, oxidoreductase-based biosensors have gained considerable attention as a sustainable and effective detection platform. For instance, a bovine hemeprotein–zeolitic imidazolate framework-8 hybrid was designed to exhibit enhanced peroxidase-like catalytic activity, improved operational stability, and facilitated enzyme reuse. Leveraging these properties, in the presence of phenol and 4-aminoantipyrine, a colorimetric biosensing platform was constructed for the fast and sensitive detection of phenol, achieving a detection limit of 1.0 µM and a wide linear detection range of 0–200 µM through naked-eye visualization.94 To further enhance performance, Au nanoparticles were introduced into hollow ZIF-8 improving its electrochemical behavior (Fig. 7a). The cavity of hollow ZIF-8 enhances the diffusion of substrates like butylated hydroxyanisole (BHA), while its superhydrophilic surface facilitates greater adsorption of reactant molecules, thereby accelerating electrochemical reactions. The electrochemical sensor demonstrates remarkable electrocatalytic activity toward BHA detection, with a broad linear response range from 0.1 to 1600 µmol L−1 and a detection limit as low as 0.03 µmol L−1.95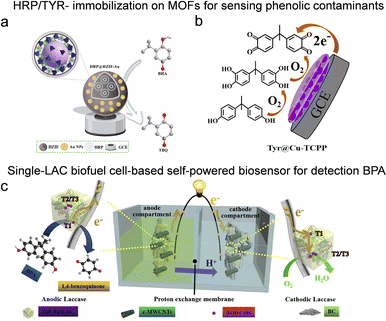 | ||
| Fig. 7 Enzyme-immobilized porous reticular framework-based sensors for phenolic contaminants detection. (a) Electrochemical oxidation mechanism of BHA, reproduced from ref. 94 with permission from American Chemical Society, ACS Appl. Mater. Interfaces, 2016, 8, 29052–29061. Copyright 2023. (b) Schematic mechanism of BPA biosensor with enhanced stability based on tyrosinase assembled between 2D Cu-TCPP nanofilms, reproduced from ref. 97 with permission from Elsevier, Chem. Eng. J., 2022, 446, 137001, copyright 2022. (c) Schematic illustration of a single-LAC biofuel cell with BPA as the fuel and O2 as the final electron acceptor, reproduced from ref. 44 with permission from Elsevier, Nano Energy, 2020, 68, 104308, copyright 2019. | ||
A significant challenge in peroxidase-based sensing is the reliance on H2O2, an essential but costly and unstable co-substrate. To circumvent the need for exogenous H2O2, innovative multi-enzyme systems have been developed. In a representative work,96 GOx and HRP were co-encapsulated within H-ZIF-8 to establish a cascaded biocatalytic system. This setup enables in situ H2O2 generation through GOx-catalyzed glucose oxidation, which subsequently activates spatially adjacent HRP for phenol detection. This system achieved a rapid colorimetric response with a linear detection range of 0–100 µM and a low LOD of 0.86 µM. Beyond HRP, the immobilization of laccase or tyrosinase (TYR) in porous crystalline frameworks has emerged as a powerful strategy for electrochemical sensing. Ma et al. developed an ultrasensitive electrochemical biosensor by assembling tyrosinase within two-dimensional Cu-TCPP nanosheets.97 As bisphenol A (BPA) is a natural substrate for tyrosinase, the enzyme specifically catalyzes its oxidation, generating a “signal-on” response (Fig. 7b), which allowed for BPA monitoring with an impressive LOD of 1.2 nM. Similarly, TYR immobilized on 2D bimetallic NiZn-MOF and laccase encapsulated in ZIF-90 (LAC@ZIF-90) have also demonstrated outstanding performance in the electrochemical detection of phenol and catechol, respectively.98,99
Conventional electrochemical sensing requires an external power source. To minimize energy consumption, a breaking approach integrates biosensing with energy harvesting, creating self-powered sensors. Li et al.44 developed highly flexible electrodes by encapsulating LAC in ZIF-8 and integrating them with a bacterial cellulose/carboxylated multi-walled carbon nanotube (BC/c-MWCNT) backbone (Fig. 7c). This BC/c-MWCNT/ZIF-8@LAC electrode was utilized as both the cathode and the anode in a single-enzyme enzymatic biofuel cell to construct a self-powered sensing platform for BPA. The oxidation of BPA by LAC generates an electric current, and the maximum power density of the EBFC was positively and linearly correlated with BPA concentration, achieving an LOD of 1.95 nM.
Despite notable innovations, most reported systems focus on model phenolic compounds (phenol, BHA, and BPA) with limited validation against complex mixtures of structurally similar contaminants commonly encountered in environmental samples. The selectivity of these platforms in real-world matrices remains inadequately characterized.
3.3. Antibiotics
The escalating overuse of antibiotics has resulted in the persistent presence of pharmaceutical contaminants in the environment and accelerated the spread of antimicrobial resistance (AMR). Residual antibiotics not only threaten the ecosystem but can also enter the human body through the food chain, further exacerbating AMR.5 Consequently, the development of accurate and reliable analytical methods for monitoring antibiotics is essential to mitigate these risks.LAC-based biosensors represent a promising approach for direct antibiotic detection. For instance, Zhong et al.100 successfully constructed a LAC-modified electrochemical biosensor for tetracycline (TC) detection in food samples. By immobilizing LAC within a mesoporous carbon sphere (MCS)@UiO-66-NH2 core–shell composite, they developed a sensing platform with enhanced enzymatic stability and electrocatalytic activity (Fig. 8a). The biosensor demonstrated satisfactory analytical performance, with a detection limit of 8.94 × 10−7 mol L−1, a linear detection range of 1.0 × 10−6 to 6.0 × 10−5 mol L−1, and good operational stability, making it suitable for screening antibiotic residues in complex food matrices.
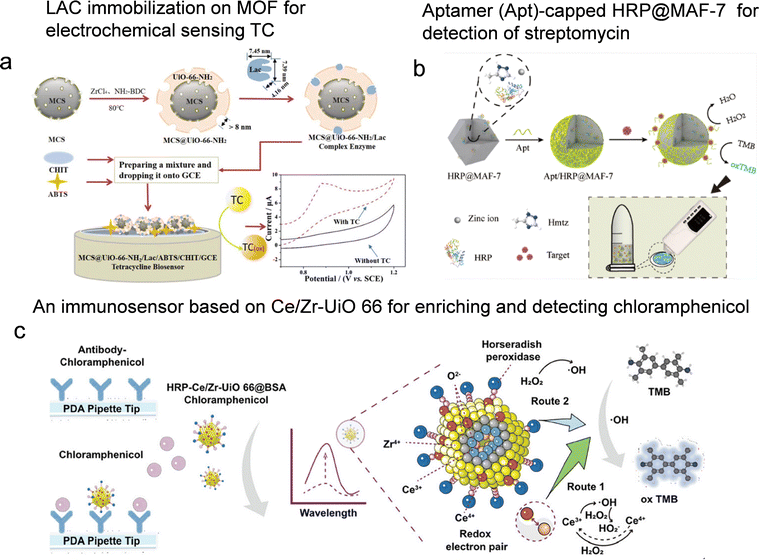 | ||
| Fig. 8 Enzyme immobilization on porous reticular framework-based sensors for antibiotics detection. (a) Illustration of the synthesis process of the MCS@UiO-66-NH2/LAC complex enzyme and construction of the MCS@UiO-66-NH2/LAC/ABTS/CHIT/GCE tetracycline biosensor, reproduced from ref. 100 with permission from Royal Society of Chemistry, Analyst, 2021, 146, 2825–2833, copyright 2021. (b) The principle of the Apt/HRP@MAF-7 sensor for on-site sensitive detection of targets, reproduced from ref. 101 with permission from American Chemical Society, Anal. Chem., 2020, 92, 14259–14266, copyright 2020, (c) schematic diagram of sensitive detection of chloramphenicol based on a dual-track multifunctional MOF-based biosensor with HRP-Ce/Zr-UiO-66@BSA-CAP and double catalysis of Ce3+/Ce4+ and HRP, reproduced from ref. 102 with permission from Wiley, Small, 2024, 20, e2309075, copyright 2024. | ||
LAC-based biosensors offer simplicity but are inherently limited to antibiotics that serve as enzyme substrates, restricting their applicability to a narrow subset of antibiotic classes. In contrast, aptamer- or antibody-based platforms provide broader applicability, as the recognition element can be tailored to any antibiotic by simply replacing the aptamer sequence. These systems typically utilize encapsulated enzymes for signal transduction and amplification, enabling highly sensitive detection across various sample types. A representative example is a portable colorimetric platform that integrates an aptamer-modified MOF for on-site antibiotic screening.101 Specifically, HRP was encapsulated in situ within the hydrophilic MAF-7, effectively suppressing enzyme aggregation and enhancing catalytic activity. Aptamers were modified on the material surface via π–π stacking, forming an intelligent Aptamer (Apt)/HRP@MAF-7 probe for targeted antibiotic recognition. During detection, the binding between the aptamer and the target triggers the release of HRP activity, catalyzing the oxidation of TMB (Fig. 8b). The resulting color signal was accurately quantified using a handheld colorimeter. Using streptomycin as a model analyte, the sensor exhibited a wide linear detection range from 0.005 to 6 ng mL−1 and a detection limit of 0.51 pg mL−1. This design can be adapted to different antibiotics by replacing the corresponding aptamer, making it highly valuable for field-deployable environmental and food safety monitoring.
To improve selectivity and reduce interference from complex sample matrices, pre-enrichment and separation of target analytes are effective strategies. Innovative platforms combining magnetic solid-phase extraction with enzymatic signal amplification have been developed. The researchers employed bimetallic MOF-based probes with dual functionality: efficient magnetic enrichment of target antibiotics and catalytic signal enhancement (Fig. 8c). In this system, Fe3O4@SiO2 serves as a magnetic core supporting Ce/Zr-UiO-66 with a high specific surface area, enabling efficient enrichment and rapid separation of chloramphenicol. Furthermore, the synergistic catalysis between HRP and Ce/Zr-UiO-66 allowed broad linear detection from 0.1 to 100 ng mL−1 on an immunosensing platform, achieving a sensitivity of 51.3 pg mL−1.102 It should be noted that this versatility comes at the cost of increased complexity and additional functionalization steps, a trade-off that must be considered for different application scenarios.
3.4. Microbial pathogens and mycotoxins
The contamination of water and food by microbial pathogens presents a substantial threat to global public health. According to the World Health Organization, waterborne diseases, such as cholera, dysentery, typhoid, and polio, account for approximately 3.4 million deaths annually.6 The rapid proliferation of pathogenic bacteria underscores the need for early and sensitive detection methods. Recently, biosensing platforms based on enzyme-immobilized porous reticular frameworks have emerged as promising tools for rapid, on-site pathogen monitoring.Gao et al.103 developed a portable pH-based biosensor utilizing GOx conjugated with mZIF-8. The mZIF-8/GOx conjugate was formed via electrostatic interactions. In this design, GOx activity remains suppressed until the target bacteria competitively bind to mZIF-8, triggering enzyme release and restoring catalytic function (Fig. 9a). The subsequent conversion of glucose to gluconic acid induces a measurable pH shift, enabling the detection of Escherichia coli and Staphylococcus aureus (S. aureus) with limits as low as 10 CFU mL−1 and 30 CFU mL−1, respectively. This low-cost and user-friendly biosensor is well-suited for monitoring the quality of contaminated drinking water. Similarly, Li et al.104 designed an enhanced immunoassay by encapsulating urease (U) within ZIF-90 through in situ synthesis (Fig. 9b). The resulting U@ZIF-90 composite was conjugated with antibodies for specific detection of S. aureus. The encapsulation not only increased enzyme loading and stability but also enhanced catalytic activity 3-fold due to the confinement effect. By quantifying urea hydrolysis, which induced pH changes detectable using a portable pH meter, the assay achieved a detection limit of 1.96 CFU mL−1 for S. aureus with high specificity. Its successful application in milk samples further demonstrated its potential for safeguarding infant food safety.
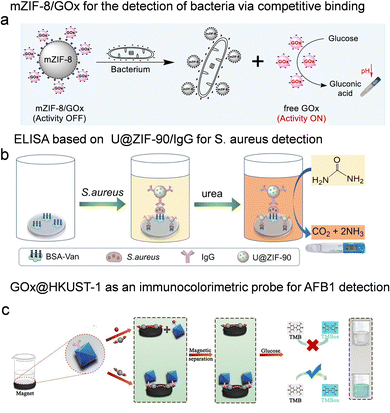 | ||
| Fig. 9 Enzyme immobilization on porous reticular framework-based sensors for detection of microbial pathogens and mycotoxins. (a) Schematic illustration of the working principle of mZIF-8/GOx conjugate for the detection of bacteria by using a pH meter as a readout, reproduced from ref. 103 with permission from American Chemical Society, ACS Appl. Mater. Interfaces, 2023, 15, 31224–31232, copyright 2023. (b) Illustration of the construction of ELISA based on U@ZIF-90/IgG for S. aureus detection with a pH meter as a readout, reproduced from ref. 104 with permission from Springer Nature, Microchim. Acta, 2022, 189, 358, copyright 2022. (c) Schematic illustration of the synthesis of GOx@HKUST-1 @antibody and measuring principle of AFB1 sensor, reproduced from ref. 105 with permission from Elsevier, Sens. Actuators B Chem., 2022, 369, 132362, copyright 2022. | ||
Beyond microbial pathogens, mycotoxins, which are secondary metabolites produced by certain fungi, also pose serious risks of contamination of water resources and agricultural products. These toxins can induce multiple adverse effects in humans, including causing damage to the neuroendocrine, gastrointestinal, reproductive, and immune systems.106 Thus, establishing highly sensitive and accurate detection methods is essential.
Aflatoxins (AFs), predominantly produced by Aspergillus flavus that contaminate crops and food stored under unsuitable conditions, rank among the most widely investigated mycotoxins.107 To enable sensitive and stable AF detection, Zhang et al.105 developed a multifunctional immunoprobe based on HKUST-1 (Fig. 9c). By employing an in situ encapsulation strategy, GOx and specific antibodies were incorporated into the framework. This composite material exhibits three key functions: enzyme immobilization, antibody support, and peroxidase-like activity. In the designed immunoassay, AFB1- and AFB1-coated magnetic beads compete for the binding sites on the immunoprobes. Unbound immunoprobes trigger a colorimetric cascade reaction: GOx catalyzes the oxidation of glucose to generate H2O2, which subsequently oxidizes TMB to form a blue-colored product. The intensity of this colorimetric signal is inversely proportional to AFB1 concentration. This method demonstrated a linear detection range of 0.01–0.1 ng mL−1 for AFB1, with a detection limit as low as 0.004 ng mL−1. Notably, the detection limit was 126-fold lower than that achieved by conventional ELISA.
Recognizing that single-mode detection is susceptible to environmental interference, recent work has embraced dual-signal platforms. These self-validating systems enhance reliability by providing orthogonal confirmation, a crucial feature for regulatory applications where false positives or negatives carry significant consequences. Hong et al. constructed a cascade catalytic system using PCN-224 loaded with Pt nanoparticles and GOx for the detection of deoxynivalenol.108 In this design, GOx catalyzes glucose to generate H2O2, which subsequently activates Pt nanoparticles to catalyze the TMB colorimetric reaction. Fluorescence signals are generated via Zr–O–P binding of fluorescent DNA. This dual-mode platform achieved limits of detection of 34 pg mL−1 (colorimetric) and 5.3 pg mL−1 (fluorescence), combining high sensitivity with reliability. Similarly, the same group also reported a dual-mode colorimetric and fluorescence method using PCN-224 loaded with HRP for sensing AFB1.109 Despite impressive detection limits, the stability of antibody-functionalized probes in complex food matrices remains problematic. Non-specific binding and batch-to-batch variability in antibody performance represent the underaddressed obstacles to commercialization.
3.5. Other contaminants
In addition to organic pollutants, well-designed porous framework-based enzyme biosensing systems have been effectively extended to the detection of other chemical contaminants, such as heavy metal ions and toxic gases. For instance, Xu et al. constructed a ratiometric fluorescent platform for detecting arsenate (As(V)) using a Zn-MOF co-encapsulating acid phosphatase (ACP) and hemin.110 This system operates on an enzyme-mediated signal switching mechanism: the inherent fluorescence of the MOF at 452 nm is quenched during the oxidation of o-phenylenediamine (OPD) to a fluorescent product (564 nm). The introduction of ascorbic acid 2-phosphate (AAP) suppresses this oxidation via enzymatic hydrolysis, restoring the original signal. As(V) inhibits ACP, preventing AAP hydrolysis and reactivating OPD oxidation, thereby enabling quantifiable As(V) detection with high selectivity, a linear detection range of 3.33–300 µg L−1, and a detection limit of 1.05 µg L−1 (Fig. 10a and b).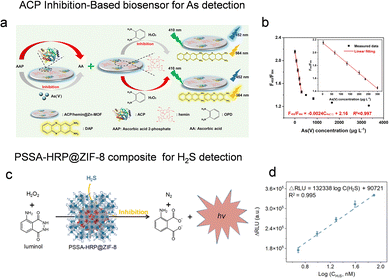 | ||
| Fig. 10 Enzyme immobilization on porous reticular framework-based sensors for heavy metal ions and toxic gases. (a) Schematic illustration of the fluorescence detection of inorganic As(V) by using ACP/hemin@Zn-MOF composite. (b) The relationship between the fluorescence ratio F452/F564 and the concentration of inorganic As(v), reproduced from ref. 110 with permission from Elsevier, J. Hazard. Mater., 2021, 412, 124407, copyright 2021. (c) Schematic illustration of H2S sensing based on the PSSA-HRP@ZIF-8 composite combined with the luminol-H2O2 system. (d) The linear detection range for H2S quantification, reproduced from ref. 112 with permission from the Royal Society of Chemistry, Chem. Sci., 2024, 15, 19609–19618, copyright 2024. | ||
Hydrogen sulfide (H2S), a significant bioactive substance and a typical environmental pollutant, poses serious health and environmental risks. High concentrations of H2S can cause respiratory irritation, and prolonged exposure may lead to unconsciousness or cardiac arrest; in addition, it contributes to air and water contamination as well as metal structure corrosion.111 To address this, we developed a chemiluminescence sensor based on polystyrolsulfon acid (PSSA)-HRP@ZIF-8. This biocomposite activates luminol to produce chemiluminescence, which is selectively suppressed in the presence of H2S due to inhibition of the biocatalyst (Fig. 10c). The resulting decrease in emission intensity allows for highly sensitive H2S detection, with a linear detection range from 4.88 nM to 5 µM and a remarkable detection limit of 0.09 nM (Fig. 10d), demonstrating suitability for environmental and biological monitoring.112
While proof-of-concept studies have demonstrated remarkable analytical performance, systematic investigation of cross-reactivity and interference from co-existing environmental constituents remains limited. This concern is particularly pronounced for enzyme inhibition-based sensors, where numerous environmental substances can interfere with color development, leading to potential false positives or inaccurate quantification. Consequently, selectivity claims require rigorous validation against panels of structurally similar compounds and common matrix components to establish reliable performance in complex environmental settings.
4. Conclusions and future perspectives
Environmental contaminants pose escalating threats to global ecosystem stability and public health, creating an urgent demand for advanced sensing platforms that enable accurate, on-site detection and quantification of pollutants. The integration of natural enzymes with porous reticular frameworks has established an innovative platform for developing robust hybrid biocatalysts. This nanobiotechnological approach synergistically combines exceptional catalytic functionality with structural programmability, providing a green route for constructing high-performance biocatalytic systems. These composites exhibit enhanced stability, superior catalytic performance, and remarkable detection sensitivity, offering promising avenues for real-time monitoring of diverse environmental contaminants. However, several scientific and technical challenges must be addressed before achieving widespread practical implementation (Fig. 11).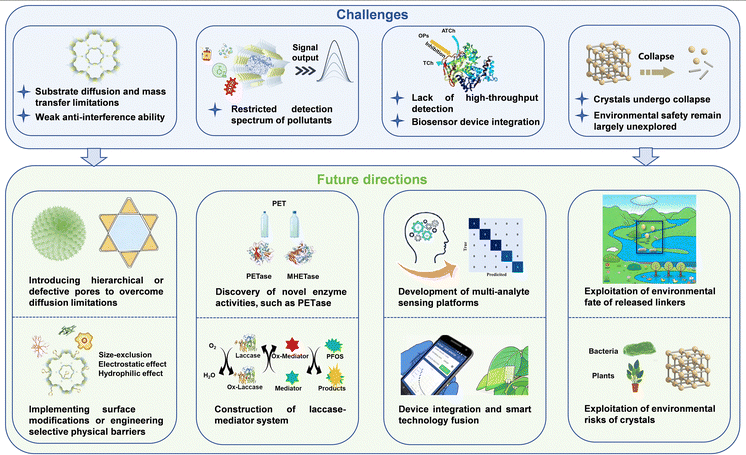 | ||
| Fig. 11 The challenges and future directions of enzyme–porous reticular framework hybrids for environmental pollutant monitoring. | ||
4.1. Substrate diffusion and mass transfer limitations
The confined pore architectures of porous reticular frameworks often hinder substrate accessibility to encapsulated enzymes, resulting in reduced catalytic efficiency compared to homogeneous reaction systems. Although reticular frameworks provide molecular-scale enzyme accessibility, mass transfer efficiency remains limited under diffusion-controlled conditions. Structural engineering strategies such as introducing hierarchical or defective pores via soft- or hard-templating methods can facilitate pollutant diffusion to enzymatic active sites without compromising framework stability.39,113,114 Additionally, optimizing surface functionalization enhances interfacial compatibility and molecular transport.93 The spatial preorganization of enzymes establishes a favorable channeling effect between proximal enzymes, significantly accelerating cascade reaction rates.115 Combining pore architecture modulation with biomolecular spatial design offers a viable route to overcome diffusion limitations and enhance biosensor detection sensitivity.4.2. Enhanced anti-interference capability for operation in complex matrices
The reliable deployment of sensors in real-world applications critically depends on their capacity to retain high performance when exposed to intricate sample environments. Sustaining catalytic activity and structural integrity under such conditions, particularly in the presence of salts, humic acids, and microorganisms, represents a formidable challenge. A major impediment is biofouling, i.e., the undesirable accumulation of proteins, biomolecules, and microbes on sensor surfaces. This non-specific adsorption can obstruct active sites, diminish selectivity, and ultimately degrade sensor reliability and operational lifespan.116 To mitigate these effects, systematic evaluation of anti-interference performance under laboratory-simulated real-matrix conditions is essential. Effective management of non-specific interactions necessitates deliberate interfacial engineering. Two principal strategies include: (1) implementing antifouling surface modifications: coating sensor interfaces with zwitterionic polymers or polyethylene glycol to form a bio-inert layer that minimizes non-specific adsorption of interfering species; and (2) engineering selective physical barriers:117 precisely tailoring pore apertures or fabricating hierarchical structures to sterically exclude larger matrix components while preserving permeability for target analytes.4.3. Expanding the detection spectrum
While enzyme-porous reticular framework hybrid sensors have shown effectiveness in detecting conventional targets, their broader application is limited by the inherent substrate specificity of the incorporated enzymes. To significantly expand the range of detectable analytes, particularly emerging contaminants such as perfluorinated compounds and microplastics, future work should focus on overcoming this enzymatic constraint. This can be accomplished through the discovery of novel enzyme activities or the rational engineering of existing enzymes to tailor their recognition and catalytic properties. For example, nano-polyethylene terephthalate is hydrolyzed synergistically by cutinase and lipase to terephthalic acid, which is then sensitively detected using bromocresol purple as an indicator.118 Additionally, LAC, especially when used with mediators, has demonstrated the ability to degrade perfluorinated compounds, suggesting its potential for sensing applications.1194.4. Development of multi-analyte sensing platforms
Environmental contamination typically involves complex pollutant mixtures, presenting particular challenges for AChE inhibition-based sensors where specific pollutant identification remains difficult. Future development should focus on multiplexed sensor arrays capable of simultaneous detection of multiple contaminants.49,120 These integrated platforms, comprising multiple responsive elements, offer advantages including rapid response, high scalability, and multi-functional compatibility. The incorporation of artificial intelligence (AI) technologies, particularly machine learning and deep learning algorithms, represents a crucial advancement. In systems employing colorimetric, electrochemical, or fluorescence-based detection, AI significantly enhances signal interpretation and noise reduction. More importantly, by leveraging pattern recognition and data fusion techniques, AI algorithms can deconvolute complex cross-reactive signals generated by sensor arrays, thereby enabling the precise identification and accurate quantification of each pollutant within high-throughput multiplexed assays. This capability is essential for resolving the composition of contaminant mixtures and avoiding false positives or negatives.121,122 Furthermore, AI-enabled platforms can analyze complex datasets to predict contamination risks, assess spatiotemporal pollution trends, and support real-time decision-making.4.5. Device integration and smart technology fusion
Translating laboratory biosensing platforms into robust, user-friendly, mass-producible devices presents a significant interdisciplinary challenge. The seamless integration of high-sensitivity biosensors into wearable systems or internet of things (IoT) networks enables continuous, real-time environmental monitoring. Recent advances in wearable sensor technology, initially developed for biomedical applications, offer promising avenues for environmental monitoring. The integration of flexible electronics with nanomaterials enables novel approaches for direct in situ detection of plant physiological signals.123 Plant nanobionic technology has successfully transformed wild spinach into living sensors for detecting nitroaromatic compounds in groundwater by embedding single-walled carbon nanotubes within leaf tissues.124 This innovative approach addresses quantitative calibration challenges and individual variability while demonstrating practical remote monitoring capabilities. The convergence of these platforms with smartphone interfaces, IoT networks, and AI algorithms enables real-time data transmission, automated analysis, and predictive modeling, transforming environmental monitoring into proactive, intelligent networks with significant implications for precision agriculture and environmental security.4.6. Exploitation of environmental safety
Given that the proposed biocatalytic frameworks are intended for environmental pollutant monitoring, where they may be directly deployed in natural water bodies or ecosystems, it is essential to evaluate not only their analytical performance but also their potential environmental risks. These porous crystals may undergo structural collapse after long-term use or upon exposure to complex environmental conditions, potentially releasing constituent metal ions and organic linkers into the surrounding water, which raises concern.125 Such degradation products could themselves act as contaminants, leading to secondary pollution that undermines the purpose of monitoring. Another critical consideration is the emerging evidence that the frameworks can migrate through the aquatic food chain and bioaccumulate in organisms.126 If scaffold fragments accumulate in biota, they may pose ecological hazards that outweigh the benefits of pollutant monitoring. At present, the environmental fate and long-term biosafety of these porous crystals remain largely unexplored. Therefore, future research efforts should prioritize investigating the ecotoxicity, degradation behavior, and bioaccumulation potential of these scaffolds to ensure the sustainable and safe application of this promising technology.Author contributions
All authors have discussed the organization and presentation of the content and participated in the writing and editing of this Review Article.Conflicts of interest
There are no conflicts to declare.Abbreviations
| MOFs | Metal–organic frameworks |
| COFs | Covalent organic frameworks |
| PEP | Pepsin |
| GOx | Glucose oxidase |
| MP-11 | Microperoxidase-11 |
| Tb-mesoMOF | Terbium-based mesoporous MOF |
| LAC | Laccase |
| HRP | Horseradish peroxidase |
| Cyt c | Cytochrome c |
| ALP | Alkaline phosphatase |
| AChE | Acetylcholinesterase |
| OPs | Organophosphorus pesticides |
| ATCh | Acetylcholine |
| CHO | Choline oxidase |
| H2O2 | Hydrogen peroxide |
| TCh | Thiocholine |
| LOD | Limit of detection |
| DTNB | 5,5′-dithiobis (2-nitrobenzoic acid) |
| TNB | 2-nitro-5-thiobenzoic acid |
| MP | Methyl parathion |
| QpeH | Aryloxyphenoxypropionate herbicide hydrolase |
| OPH | Organophosphate hydrolase |
| AOPP | Aryloxyphenoxypropionate |
| AOx | Alcohol oxidase |
| QpE | Quizalofop-P-ethyl |
| PNP | p-Nitrophenol |
| mZIF-8 | Magnetic ZIF-8 |
| CRL | Candida rugosa lipase |
| BCL | Burkholderia cepacia lipase |
| ELISA | Enzyme-linked immunosorbent assays |
| TMB | Tetramethylbenzidine |
| CPF | Chlorpyrifos |
| BHA | Butylated hydroxyanisole |
| TYR | Tyrosinase |
| BPA | Bisphenol A |
| AMR | Antimicrobial resistance |
| TC | Tetracycline |
| MCS | Mesoporous carbon sphere |
| Apt | Aptamer |
| S. aureus | Staphylococcus aureus |
| U | Urease |
| AFs | Aflatoxins |
| As(V) | Arsenate |
| ACP | Acid phosphatase |
| OPD | o-phenylenediamine |
| AAP | Ascorbic acid 2-phosphate |
| H2S | Hydrogen sulfide |
| PSSA | Polystyrolsulfon acid |
| AI | Artificial intelligence |
| IoT | Internet of Things |
Data availability
All data discussed in this review originate from the studies cited throughout the manuscript. Readers seeking detailed experimental procedures, analytical methods or supplementary data sets can refer to the corresponding publications listed in the References section.Acknowledgements
We acknowledge financial support from projects of the National Natural Science Foundation of China (22104159, 22174164, and 22506069), the Guangdong Basic and Applied Basic Research Foundation (2024B1515020070, 2023A1515011632, and 2022A1515010837), the National High-Level Talents Special Support Program-Young Talents (2024WRQB006), the Fundamental Research Funds for the Central Universities, Sun Yat-sen University (23lgbj005), the Science and Technology Planning Project of Sichuan Province (No. 2025NSFSC2058), the Guangdong Basic Research Center of Excellence for Functional Molecular Engineering (31000-42080002), and the Natural Science Foundation of Guangzhou City (202201011408).Notes and references
- V. K. Gaur, P. Sharma, R. Sirohi, M. K. Awasthi, C.-G. Dussap and A. Pandey, J. Hazard. Mater., 2020, 398, 123019 CrossRef CAS PubMed.
- M. M. McAlister, Q. Zhang, J. Annis, R. W. Schweitzer, S. Guidotti and J. R. Mihelcic, Environ. Sci. Technol., 2022, 56, 732–738 CrossRef CAS PubMed.
- Y. Wang, H. Li and X. Yan, TrAC, Trends Anal. Chem., 2025, 191, 118381 CrossRef CAS.
- R. Sridar, U. U. Ramanane and M. Rajasimman, Environ. Nanotechnol., Monit. Manage., 2018, 10, 388–393 Search PubMed.
- M. A. Cook and G. D. Wright, Sci. Transl. Med., 2022, 14, eabo7793 CrossRef CAS PubMed.
- F. Y. Ramírez-Castillo, A. Loera-Muro, M. Jacques, P. Garneau, F. J. Avelar-González, J. Harel and A. L. Guerrero-Barrera, Pathogens, 2015, 4, 307–334 CrossRef PubMed.
- C. K. Ho, A. Robinson, D. R. Miller and M. J. Davis, Sensors, 2005, 5, 4–37 CrossRef CAS.
- J. Devers, D. I. Pattison, A. B. Hansen and J. H. Christensen, Talanta, 2025, 282, 127032 CrossRef CAS PubMed.
- R. Pei, L. Ye and C. Jing, Biosens. Bioelectron., 2023, 229, 115244 CrossRef CAS PubMed.
- S. Liu, C. Nie, F. He, G. Wu, H. Wang, S. Li, C. Du, Z. Zheng, J. Cheng, Y. Shen and J. Cheng, Trends Food Sci. Technol., 2024, 150, 104597 CrossRef CAS.
- M. Bilal, A. K. Singh, H. M. N. Iqbal and G. Boczkaj, Chem. Eng. J., 2023, 474, 145020 CrossRef CAS.
- Z. Fu, L. Yang, Y. Ji, J. Xie and Z. Ding, Coordin. Chem. Rev., 2026, 548, 217157 CrossRef CAS.
- S. Feng, H. Hao Ngo, W. Guo, S. Woong Chang, D. Duc Nguyen, D. Cheng, S. Varjani, Z. Lei and Y. Liu, Bioresour. Technol., 2021, 335, 125278 CrossRef CAS PubMed.
- V. Karthik, P. Senthil Kumar, D.-V. N. Vo, P. Selvakumar, M. Gokulakrishnan, P. Keerthana, V. Audilakshmi and J. Jeyanthi, Environ. Chem. Lett., 2021, 19, 2331–2350 CrossRef CAS.
- R. A. Sheldon, A. Basso and D. Brady, Chem. Soc. Rev., 2021, 50, 5850–5862 RSC.
- L. Wang, X. Du, Y. Li, Y. Bai, T. Tang, J. Wu, H. Liang and D. Gao, Crit. Rev. Environ. Sci. Technol., 2023, 53, 1684–1708 CrossRef CAS.
- Z. Guo, J. J. Richardson, B. Kong and K. Liang, Sci. Adv., 2020, 6, eaaz0330 CrossRef CAS PubMed.
- A. P. Côté, A. I. Benin, N. W. Ockwig, M. O'Keeffe, A. J. Matzger and O. M. Yaghi, Science, 2005, 310, 1166–1170 CrossRef PubMed.
- H. Furukawa, K. E. Cordova, M. O’Keeffe and O. M. Yaghi, Science, 2013, 341, 1230444 CrossRef PubMed.
- B. Wang, R.-B. Lin, Z. Zhang, S. Xiang and B. Chen, J. Am. Chem. Soc., 2020, 142, 14399–14416 CrossRef CAS PubMed.
- Y. Feng, Y. Xu, S. Liu, D. Wu, Z. Su, G. Chen, J. Liu and G. Li, Coord. Chem. Rev., 2022, 459, 214414 CrossRef CAS.
- S. Huang, G. Chen and G. Ouyang, Chem. Soc. Rev., 2022, 51, 6824–6863 RSC.
- C. Doonan, R. Riccò, K. Liang, D. Bradshaw and P. Falcaro, Acc. Chem. Res., 2017, 50, 1423–1432 CrossRef CAS PubMed.
- X. Liu, M. Qiu, Y. Zhang, J. Pan, W. Zhang, L. Guo, X. Zhang and Y. Jiang, Adv. Colloid Interface Sci., 2025, 344, 103589 CrossRef CAS PubMed.
- M. Feng, C. Xing, Y. Jin, X. Feng, Y. Zhang and B. Wang, J. Am. Chem. Soc., 2024, 146, 32883–32905 CrossRef CAS PubMed.
- H. Wang, X. Kou, R. Gao, S. Huang, G. Chen and G. Ouyang, Environ. Sci. Technol., 2024, 58, 11869–11886 CrossRef CAS PubMed.
- L. Tong, S. Huang, G. Chen and G. Ouyang, Angew. Chem., Int. Ed., 2025, 64, e202421192 CrossRef CAS PubMed.
- W. Huang, H. Yuan, H. Yang, X. Ma, S. Huang, H. Zhang, S. Huang, G. Chen and G. Ouyang, Nat. Commun., 2023, 14, 3644 CrossRef CAS PubMed.
- R. Gao, X. Kou, L. Tong, Z. W. Li, Y. Shen, R. He, L. Guo, H. Wang, X. Ma, S. Huang, G. Chen and G. Ouyang, Angew. Chem., Int. Ed., 2024, 63, e202319876 CrossRef CAS PubMed.
- W. Tan, T. Wei, J. Huo, M. Loubidi, T. Liu, Y. Liang and L. Deng, ACS Appl. Mater. Interfaces, 2019, 11, 36782–36788 CrossRef CAS PubMed.
- R. J. Drout, L. Robison and O. K. Farha, Coord. Chem. Rev., 2019, 381, 151–160 Search PubMed.
- X. G. Yang, J. R. Zhang, X. K. Tian, J. H. Qin, X. Y. Zhang and L. F. Ma, Angew. Chem., Int. Ed., 2023, 62, e202216699 CrossRef CAS PubMed.
- F. L. Oliveira, S. P. de Souza, J. Bassut, H. M. Álvarez, Y. Garcia-Basabe, R. O. M. Alves de Souza, P. M. Esteves and R. S. B. Gonçalves, Chem. – Eur. J., 2019, 25, 15863–15870 CrossRef CAS PubMed.
- Y. Lu, X. Wu, H. Lou and Z. Li, Langmuir, 2025, 41, 11765–11775 CrossRef CAS PubMed.
- Q. Sun, C.-W. Fu, B. Aguila, J. Perman, S. Wang, H.-Y. Huang, F.-S. Xiao and S. Ma, J. Am. Chem. Soc., 2018, 140, 984–992 CrossRef CAS PubMed.
- C. Wang and K. Liao, ACS Appl. Mater. Interfaces, 2021, 13, 56752–56776 CrossRef CAS PubMed.
- R. Gao, X. Kou, S. Huang, G. Chen and G. Ouyang, ChemBioChem, 2024, 25, e202400339 CrossRef CAS PubMed.
- V. Lykourinou, Y. Chen, X.-S. Wang, L. Meng, T. Hoang, L.-J. Ming, R. L. Musselman and S. Ma, J. Am. Chem. Soc., 2011, 133, 10382–10385 Search PubMed.
- P. Li, Q. Chen, T. C. Wang, N. A. Vermeulen, B. L. Mehdi, A. Dohnalkova, N. D. Browning, D. Shen, R. Anderson, D. A. Gómez-Gualdrón, F. M. Cetin, J. Jagiello, A. M. Asiri, J. F. Stoddart and O. K. Farha, Chem, 2018, 4, 1022–1034 CAS.
- L. Guo, R. He, G. Chen, H. Yang, X. Kou, W. Huang, R. Gao, S. Huang, S. Huang, F. Zhu and G. Ouyang, J. Am. Chem. Soc., 2024, 146, 17189–17200 CrossRef CAS PubMed.
- S. Liang, X.-L. Wu, J. Xiong, M.-H. Zong and W.-Y. Lou, Coord. Chem. Rev., 2020, 406, 213149 CrossRef CAS.
- Y. Liu, J. Wang, X. Qin, G. Liu and H. Zhang, Food Chem., 2025, 493, 145602 CrossRef CAS PubMed.
- G. Chen, S. Huang, Y. Shen, X. Kou, X. Ma, S. Huang, Q. Tong, K. Ma, W. Chen, P. Wang, J. Shen, F. Zhu and G. Ouyang, Chem, 2021, 7, 2722–2742 CAS.
- X. Li, D. Li, Y. Zhang, P. Lv, Q. Feng and Q. Wei, Nano Energy, 2020, 68, 104308 CrossRef CAS.
- R. Xu, M. Liu, C. Yao and X. Xu, ACS Appl. Mater. Interfaces, 2025, 17, 5401–5409 CrossRef CAS PubMed.
- J. Liang, Z. Huang, K. Wang, L. Zhang, Y. Wan, T. Yang and H. Zeng, Talanta, 2023, 259, 124503 CrossRef CAS PubMed.
- X. Cao, Y. Guo, M. Zhao, J. Li, C. Wang, J. Xia, T. Zou and Z. Wang, Food Chem., 2022, 381, 132282 CrossRef CAS PubMed.
- D. Chen, L. Wang, J. Wei, T. Jiao, Q. Chen, M. Oyama, Q. Chen, X. Chen and X. Chen, Food Chem., 2024, 442, 138389 CrossRef CAS PubMed.
- J. Huang, L. Jiao, W. Xu, H. Wang, M. Sha, Z. Wu, W. Gu, L. Hu and C. Zhu, J. Hazard. Mater., 2022, 432, 128707 Search PubMed.
- H. Ma, M. Li, T. Yu, H. Zhang, M. Xiong and F. Li, ACS Appl. Mater. Interfaces, 2021, 13, 44329–44338 Search PubMed.
- Y. Bai, S. Nie, W. Gao, N. Li, P. Zhu, L. Zhang and J. Yu, Adv. Funct. Mater., 2024, 35, 2419499 CrossRef.
- J. Mehta, S. Dhaka, N. Bhardwaj, A. K. Paul, S. Dayananda, S.-E. Lee, K.-H. Kim and A. Deep, Sens. Actuators, B, 2019, 290, 267–274 Search PubMed.
- H. Li, Q. Lu, J. Shi, X. Zhang, P. Sun, X. Yan and G. Lu, Adv. Funct. Mater., 2023, 34, 2309383 CrossRef.
- M. Li, S. Qiao, Y. Zheng, Y. H. Andaloussi, X. Li, Z. Zhang, A. Li, P. Cheng, S. Ma and Y. Chen, J. Am. Chem. Soc., 2020, 142, 6675–6681 Search PubMed.
- Y. Zheng, S. Zhang, J. Guo, R. Shi, J. Yu, K. Li, N. Li, Z. Zhang and Y. Chen, Angew. Chem., Int. Ed., 2022, 61, e202208744 CrossRef CAS PubMed.
- R. Gao, N. Zhong, L. Tong, X. Kou, W. Huang, H. Yang, S. Huang, J. Wu, G. Chen and G. Ouyang, Cell Rep. Phys. Sci., 2022, 3, 101153 CrossRef CAS.
- J. Chen, Z. Guo, Y. Xin, Z. Gu, L. Zhang and X. Guo, Sci. Total Environ., 2023, 867, 161510 CrossRef CAS PubMed.
- Y. Chen, K. Yu, M. Hassan, C. Xu, B. Zhang, K. Y.-H. Gin and Y. He, Ecotoxicol. Environ. Saf., 2018, 166, 320–327 CrossRef CAS PubMed.
- H. Patel, D. Rawtani and Y. K. Agrawal, Trends Food Sci. Tech., 2019, 85, 78–91 CrossRef CAS.
- D.-M. Liu, B. Xu and C. Dong, TrAC, Trends Anal. Chem., 2021, 142, 116320 CrossRef CAS.
- C. Jin, S. Yang, J. Zheng, F. Chai and M. Tian, Food Chem., 2024, 459, 140369 CrossRef CAS PubMed.
- X. Wang, S. Yang, J. Shan and X. Bai, Int. J. Electrochem. Sci., 2022, 17, 220543 CrossRef CAS.
- H. Liang, L. Wang, Y. Yang, Y. Song and L. Wang, Biosens. Bioelectron., 2021, 193, 113553 CrossRef CAS PubMed.
- H. Chai, Y. Li, K. Yu, Z. Yuan, J. Guan, W. Tan, J. Ma, X. Zhang and G. Zhang, Anal. Chem., 2023, 95, 16383–16391 CrossRef CAS PubMed.
- L. Wen, N. Wang, Z. Liu, C.-A. Tao, X. Zou, F. Wang and J. Wang, Chemosensors, 2022, 10, 418 CrossRef CAS.
- Y. Luo, N. Wu, L. Wang, Y. Song, Y. Du and G. Ma, Biosensors, 2022, 12, 625 Search PubMed.
- Y. Xiao, N. Wu, L. Wang and L. Chen, Biosensors, 2022, 12, 899 Search PubMed.
- Q. Wu, Y. Wang, L. Wang, Y. Su, G. He, X. Chen, L. Hou, W. Zhang and Y. Y. Wang, ACS Appl. Mater. Interfaces, 2024, 16, 55802–55812 CAS.
- S. Dong, L. Peng, W. Wei and T. Huang, ACS Appl. Mater. Interfaces, 2018, 10, 14665–14672 Search PubMed.
- C. Gong, D. Wang and H. Zhao, Small, 2023, 19, e2304077 CrossRef PubMed.
- C. Gong, B. Chen, Y. Xing and H. Zhao, J. Hazard. Mater., 2024, 463, 132849 CrossRef CAS PubMed.
- K. Yang, H. Zhao, N. Li, Y. Wang, B. Sun, M. Cui and C. Zhang, Microchem. J., 2024, 203, 110945 CrossRef CAS.
- Y. Liu, M. Zhou, C. Jin, J. Zeng, C. Huang, Q. Song and Y. Song, Front. Chem., 2020, 8, 643 CrossRef CAS PubMed.
- K. Niu, Y. Zhang, J. Chen and X. Lu, ACS Sens., 2022, 7, 3551–3559 CrossRef CAS PubMed.
- K. Niu, P. Sun, J. Chen and X. Lu, Anal. Chem., 2022, 94, 17177–17185 CrossRef CAS PubMed.
- Y. Zhao, X. Lu and F. Gao, Anal. Biochem., 2022, 646, 114628 CrossRef CAS PubMed.
- W. Wei, S. Zhou, D. D. Ma, Q. Li, M. Ran, X. Li, X. T. Wu and Q. L. Zhu, Adv. Funct. Mater., 2023, 33, 2302917 CrossRef CAS.
- H. Zhao, X. Tan, H. Chai, L. Hu, H. Li, L. Qu, X. Zhang and G. Zhang, Chin. Chem. Lett., 2025, 36, 110571 CrossRef CAS.
- E. Xie, Z. Chen, X. Zhang, X. Zhang, L. Zheng, X. Wang and D. Zhang, J. Hazard. Mater., 2025, 492, 138159 CrossRef CAS PubMed.
- J. Mehta, S. Dhaka, A. K. Paul, S. Dayananda and A. Deep, Environ. Res., 2019, 174, 46–53 CrossRef CAS PubMed.
- Z. Wang, B. Ma, C. Shen and L.-Z. Cheong, Talanta, 2019, 197, 356–362 CrossRef CAS PubMed.
- B. Ma, L.-Z. Cheong, X. Weng, C.-P. Tan and C. Shen, Electrochim. Acta, 2018, 283, 509–516 CrossRef CAS.
- C. Liu, X. Wang, X. Jia, W. He, S. Zhang and H. Liu, Chem. Eng. J., 2025, 520, 166170 CrossRef CAS.
- M. Romero-Lopez, M. Oria, F. Ferrer-Marquez, M. F. Varela, K. Lampe, M. Watanabe-Chailland, L. Martinez and J. L. Peiro, Pediatr. Surg. Int., 2023, 39, 180 CrossRef PubMed.
- Y. Cheng, O. M. Lai, C. P. Tan, W. Panpipat, L. Z. Cheong and C. Shen, ACS Appl. Mater. Interfaces, 2021, 13, 4146–4155 CrossRef CAS PubMed.
- Y. Cheng, B. Ma, C.-P. Tan, O.-M. Lai, W. Panpipat, L.-Z. Cheong and C. Shen, Sens. Actuators, B, 2020, 321, 128477 CrossRef CAS.
- Y. Chen, G. Yuan, L. Tan, P. Wang, R. Lu and C. Wang, J. Inorg. Organomet. Polym. Mater., 2022, 32, 4401–4411 CrossRef CAS.
- Y. Ma and Y. Li, Microchim. Acta, 2023, 190, 492 CrossRef CAS PubMed.
- R.-I. Stefan-van Staden, R. G. Bokretsion, J. F. van Staden and H. Y. Aboul-Enein, Crit. Rev. Anal. Chem., 2015, 45, 2–31 CrossRef CAS.
- X. Yan, C. Wei, C. Su, X. Liu, P. Sun, Z. Xu, G. Lu, Y. Lin and H. Li, Adv. Funct. Mater., 2025, 35, 2420159 CrossRef CAS.
- D. Su, T. Wang, A. Li, Y. Ma, X. Liu, C. Wang, X. Jia, P. Sun, F. Liu, X. Yan and G. Lu, Adv. Funct. Mater., 2022, 32, 2204130 CrossRef CAS.
- X. Yan, D. Su, H. Li, X. Zhao, X. Liu, F. Liu, P. Sun and G. Lu, Adv. Funct. Mater., 2023, 33, 2215192 CrossRef CAS.
- G. Chen, W. Xu, W. Gu and C. Zhu, Anal. Chem., 2025, 97, 16019–16025 CrossRef CAS PubMed.
- Y. Yin, C. Gao, Q. Xiao, G. Lin, Z. Lin, Z. Cai and H. Yang, ACS Appl. Mater. Interfaces, 2016, 8, 29052–29061 CrossRef CAS PubMed.
- W. Sun, H. Chu, X. Zha, S. Lu and Y. Wang, ACS Appl. Nano Mater., 2023, 6, 15183–15192 CrossRef CAS.
- H. Liu, Y. Du, J. Gao, L. Zhou, Y. He, L. Ma, G. Liu, Z. Huang and Y. Jiang, Ind. Eng. Chem. Res., 2019, 59, 42–51 CrossRef.
- J. Ma, J. Yuan, Y. Xu, Y. Jiang, W. Bai and J. Zheng, Chem. Eng. J., 2022, 446, 137001 CrossRef.
- Y. Wen, R. Li, J. Liu, X. Zhang, P. Wang, X. Zhang, B. Zhou, H. Li, J. Wang, Z. Li and B. Sun, Anal. Chim. Acta, 2020, 1127, 131–139 CrossRef CAS PubMed.
- D. Li, Y. Cheng, H. Zuo, W. Zhang, G. Pan, Y. Fu and Q. Wei, J. Colloid Interface Sci., 2021, 603, 771–782 CrossRef CAS PubMed.
- X. Zhong, F. Wang, J. Piao and Y. Chen, Analyst, 2021, 146, 2825–2833 RSC.
- L. Wang, G. Liu, Y. Ren, Y. Feng, X. Zhao, Y. Zhu, M. Chen, F. Zhu, Q. Liu and X. Chen, Anal. Chem., 2020, 92, 14259–14266 CrossRef CAS PubMed.
- Y. Dong, N. Feng, P. Liu, Q. Wei, X. Peng, F. Jiang and Y. Chen, Small, 2024, 20, e2309075 CrossRef PubMed.
- L. Gao, J. Wen, Z. Huang, S. Sheng, F. Xu, G. Ma and H. Tan, ACS Appl. Mater. Interfaces, 2023, 15, 31224–31232 CrossRef CAS PubMed.
- S. Li, H. Xie, F. Xie, Q. Yi and H. Tan, Microchim. Acta, 2022, 189, 358 CrossRef CAS PubMed.
- C. Zhang, Y. Wang, Y. Sun, J. Duan, M. Wang, S. Ai and J. Hou, Sens. Actuators, B, 2022, 369, 132362 CrossRef CAS.
- D. K. Dey, J. I. Kang, V. K. Bajpai, K. Kim, H. Lee, S. Sonwal, J. Simal-Gandara, J. Xiao, S. Ali, Y. S. Huh, Y.-K. Han and S. Shukla, Crit. Rev. Food Sci. Nutr., 2023, 63, 8489–8510 CrossRef CAS PubMed.
- B. Wang, Y. Chen, Y. Wu, B. Weng, Y. Liu, Z. Lu, C. M. Li and C. Yu, Biosens. Bioelectron., 2016, 78, 23–30 CrossRef CAS PubMed.
- F. Hong, L. Ren and Y. Chen, Chem. Eng. J., 2023, 452, 139149 CrossRef CAS.
- L. Ren, F. Hong, L. Zeng and Y. Chen, ACS Appl. Mater. Interfaces, 2022, 14, 51234–51243 CrossRef CAS PubMed.
- X. Xu, Z. Luo, K. Ye, X. Zou, X. Niu and J. Pan, J. Hazard. Mater., 2021, 412, 124407 CrossRef CAS PubMed.
- S. Wang, V. Rohani, P. Leroux, C. Gracian, V. Nastasi and L. Fulcheri, Chemosphere, 2024, 364, 143174 CrossRef CAS PubMed.
- Y. Xu, H. Yang, A. Huang, L. Tong, W. Huang, G. Chen, W. Yi, S. Huang and G. Ouyang, Chem. Sci., 2024, 15, 19609–19618 RSC.
- Y. Tong, J. Yang, F. Xia and J. Gu, JACS Au, 2025, 5, 178–186 CrossRef CAS PubMed.
- J. Guo, X. Yue, Y. Hou, Y. Wang, Y. Liu, G. Liu, Y. He, L. Ma, L. Zhou and Y. Jiang, Int. J. Biol. Macromol., 2024, 278, 134641 CrossRef CAS PubMed.
- J. Song, W. He, H. Shen, Z. Zhou, M. Li, P. Su and Y. Yang, Chem. Eng. J., 2019, 363, 174–182 CrossRef CAS.
- J.-C. Lee, S. Y. Kim, J. Song, H. Jang, M. Kim, H. Kim, S. Q. Choi, S. Kim, P. Jolly, T. Kang, S. Park and D. E. Ingber, Nat. Commun., 2024, 15, 711 CrossRef CAS PubMed.
- L. Zhou, R. Yang, X. Li, N. Dong, B. Zhu, J. Wang, X. Lin and B. Su, J. Am. Chem. Soc., 2023, 145, 23727–23738 CrossRef CAS PubMed.
- W. Zhan, C. Wang, X. Yang, H. Li, S. Xiong and X. Li, Environ. Int., 2024, 186, 108598 CrossRef CAS PubMed.
- Q. Luo, X. Yan, J. Lu and Q. Huang, Environ. Sci. Technol., 2018, 52, 10617–10626 CrossRef CAS PubMed.
- S. Li, X. Chen, Q. Tian, J. Liu, Z. Tang and X. Niu, Anal. Chem., 2025, 97, 2537–2545 CrossRef CAS PubMed.
- T. Tian, D. Song, L. Zhen, Z. Bi, L. Zhang, H. Huang and Y. Li, Biosens. Bioelectron., 2025, 277, 117286 CrossRef CAS PubMed.
- Y. Li, F. Chen, Y. Liu, M. A. Khan, H. Zhao, H. Cao and D. Ye, Chem. Eng. J., 2025, 511, 162115 CrossRef CAS.
- H. Ibrahim, S. Moru, P. Schnable and L. Dong, ACS Sens., 2022, 7, 2293–2302 CrossRef CAS PubMed.
- M. H. Wong, J. P. Giraldo, S.-Y. Kwak, V. B. Koman, R. Sinclair, T. T. S. Lew, G. Bisker, P. Liu and M. S. Strano, Nat. Mater., 2017, 16, 264–272 CrossRef CAS PubMed.
- A. Tricoli, A. R. D. Lowe and T. Tsuzuki, Chem. Eng. J., 2021, 413, 127511 CrossRef.
- X. Luo, Z. Fang, K. Ren, H. Zou and Q. Liu, Environ. Sci. Technol., 2026, 60, 4658 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |








