Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Differential self-assembly of sequence-isomeric phosphoestamers
James Williamson a,
Tomasz Piskorz
a,
Tomasz Piskorz b,
Bini Claringboldc,
Alexandra R. Paul
b,
Bini Claringboldc,
Alexandra R. Paul c,
Nikita Harvey
c,
Nikita Harvey a,
Fernanda Duarte
a,
Fernanda Duarte b and
Christopher J. Serpell
b and
Christopher J. Serpell *a
*a
aDepartment of Biological and Pharmaceutical Chemistry, School of Pharmacy, University College London, 29-39 Brunswick Square, London, WC1N 1AX, UK. E-mail: chris.serpell@ucl.ac.uk
bDepartment of Chemistry, University of Oxford, Chemistry Research Laboratory, Mansfield Road, Oxford, OX1 1TA, UK
cSchool of Natural Sciences, University of Kent, Ingram Building, Canterbury, Kent, CT2 7NH, UK
First published on 11th February 2026
Abstract
Phosphoestamers self-assemble into an array of superstructures according to their sequence and thus provide a useful model for understanding sequence/structure relationships. To explore this, we synthesised all tetrameric phosphoestamers composed of equal ratios of C12 and HEG monomers and examined their self-assemblies in a combined experimental/computational workflow.
Sequence-defined macromolecules exhibit differences in superstructure,1 material properties,2 and function3 according to changes in monomer order. This allows macromolecule behaviour and self-assembly to be programmed into the primary sequence. Nature is inherently sequence-defined and uses modular biopolymers to achieve function from sequence via 3D structure.4 As a result, sequence definition is the key to biological recognition, catalysis, self-replication, complex self-assembly, and the heredity processes of all organisms.5,6
Poly/oligophosphoesters are ideal candidates for the exploration of sequence definition in the synthetic milieu.7 Polyphosphoesters can incorporate many monomers beyond nucleosides,8 and the anionic backbone facilitates water solubility and provides opportunities for responsive behaviour with cations. Most importantly, oligo/polyphosphoesters that are sequence-defined (phosphoestamers) can be readily produced on the solid-phase using the phosphoramidite method, the principal technique in the synthesis of DNA and the gold standard for sequence control, due to its outstanding efficiency.9 Recently, nucleic acids of up to 1728 monomers in length have been made using this technology.10
We are currently a long way from programming many-kDa phosphoestamers which mimic the structures and functions of proteins. While studies on the self-assembly and folding behaviour of phosphoestamers are reported,11–14 the specific relationship of sequence to behaviour is as yet poorly understood. Nonetheless, sequence/function relationships have been established, such as the selective inhibition of protein–protein interactions.15 Previously, we explored the self-assembly of sequence-isomeric phosphoestamers composed of hydrophobic dodecane diol (C12) and hydrophilic hexa(ethylene glycol) (HEG) monomers, which are commercially available.13 Using two 20mer sequence isomers of these monomers (C1210-HEG10 and (C12-HEG)10), we observed self-assemblies that were dynamic, responsive, and sequence-programmable.13 However, there was considerable sequence-space left unexplored.
Herein, we have sought to further understand sequence/self-assembly relationships in phosphoestamers by synthesising all possible sequence isomers of tetramers containing an equal ratio of C12 and HEG monomers, and studying their self-assembly through diffusion ordered NMR spectroscopy (DOSY) and molecular dynamics (MD) simulations. Tetramers were chosen because 4mers are a lower limit of sequence-defined self-assembly in systems with a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio of two monomers,16 while also having the capacity to form hierarchical self-assemblies,17 being readily computationally tractable,18 and providing substantial sequence variety using just four sequences. Study of fragments, be it peptides as protein substructures, or oligonucleotides for genomic DNA structure, has long proved a fruitful approach (without ignoring the possibility for emergent higher order structures). It was anticipated that the diblock (C122-HEG2), alternating ((C12-HEG)2), and two XYYX symmetric tetramers (C12-HEG-HEG-C12 and HEG-C12-C12-HEG) would produce a more diverse range of sequence-defined self-assemblies.
1 ratio of two monomers,16 while also having the capacity to form hierarchical self-assemblies,17 being readily computationally tractable,18 and providing substantial sequence variety using just four sequences. Study of fragments, be it peptides as protein substructures, or oligonucleotides for genomic DNA structure, has long proved a fruitful approach (without ignoring the possibility for emergent higher order structures). It was anticipated that the diblock (C122-HEG2), alternating ((C12-HEG)2), and two XYYX symmetric tetramers (C12-HEG-HEG-C12 and HEG-C12-C12-HEG) would produce a more diverse range of sequence-defined self-assemblies.
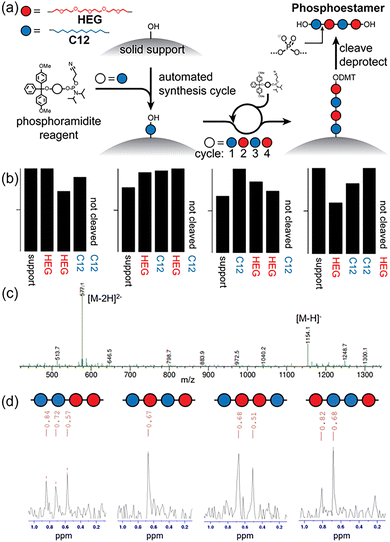
The tetramers were synthesised on a 1 µmol scale by the phosphoramidite method, using controlled pore glass (CPG) supports via the UnyLinker (Fig. 1a).19 Coupling success was measured semi-quantitatively using in-line monitoring of the outgoing DMT cation concentration by optical absorption at 500 nm (Fig. 1b) and phosphoestamers were cleaved from the solid support using 28% ammonium hydroxide at 60 °C. The concentration of the samples was determined by cleaving the final DMT protecting group in 80% acetic acid and accurately measuring the optical absorption at 500 nm compared to a standard.13,14 The phosphoestamers were then purified by extraction of the DMT side products into dichloromethane, before successful synthesis was confirmed by mass spectrometric identification of the molecular ion [M − H]− and [M − 2H]2− peaks at m/z 1154 and 577, respectively (Fig. 1c and Fig. S3–S6).
Phosphorus-31 NMR was employed to characterise the tetramer sequences (Fig. 1d and Fig. S2). The C12-PO4-C12 orthophosphate environment appeared at 0.82–0.84 ppm, C12-PO4-HEG at 0.67–0.72 ppm, and HEG-PO4-HEG at 0.51–0.57 ppm. To our knowledge, this is the first time that 31P NMR has been applied to identify sequence signatures in phosphoestamers.
We then proceeded to study aggregate size via diffusion coefficients using DOSY NMR.20 The diffusion coefficient varies according to molecular weight21 and the morphologies of isomeric macromolecules.22 DOSY NMR is highly sensitive and can differentiate between aggregates in a mixture based on their translational diffusion.23 If two chemical environments share the same diffusion coefficient, they are likely to be part of the same compound/aggregate.23
For the phosphoestamers, the diffusion coefficients were extracted from two signals in the 1H NMR to increase the accuracy of our measurements. These were the main resonances of the alkoxy protons (3.60–3.70 ppm), and the alkyl protons (1.20–1.30 ppm) (Fig. S7), which correspond to the HEG and C12 monomers, respectively. 1H DOSY NMR was trialled for the tetramers at 10 µM, 100 µM, and 1 mM in pure D2O. The tetramers at 1 mM gave the strongest signals (Fig. S8–S19) and their diffusion coefficients were calculated (Table 1).
| Tetramer sequence | Pure D2O diffusion coefficient (µm2 s−1) | 1 M NH4OAc (aq.) diffusion coefficient (µm2 s−1) |
|---|---|---|
| C122-HEG2 | 230 ± 9 | 163 ± 3 |
| (C12-HEG)2 | 247 ± 5 | 213 ± 9 |
| C12-HEG-HEG-C12 | 232 ± 3 | 173 ± 3 |
| HEG-C12-C12-HEG | 241 ± 4 | 192 ± 8 |
At 1 mM in D2O, the sequence-isomeric phosphoestamers displayed poorly separated translational diffusion across sequences. Diffusion coefficients of approximately 240 µm2 s−1 can be indicative of free molecule diffusion (e.g. the fluorescent dye Cy5 diffuses at 280 µm2 s−1 in aqueous solutions24). This suggests that we observed isolated chains, which would be consistent with the trianionic nature of the molecules resulting in mutual repulsion, regardless of sequence.
We then added cations to encourage self-assembly through screening of repulsive electrostatic interactions between phosphates, similar to the use of cations in nucleic acid hybridisation.25 This previously resulted in responsive behaviour for the 20mers.13 Ammonium cations (NH4+) in the form of ammonium acetate (NH4OAc) were chosen for this because NH4OAc, unlike Na+ or Mg2+ buffers, can be evaporated from the samples and because NH4OAc has been used before in self-assembly studies by DOSY NMR.26 NH4OAc solution (1 M) was prepared using D2O and the tetramers (1 mM) were dissolved in NH4OAc solution before acquiring 1H DOSY NMR spectra (Fig. S20–S31). The results showed significant differences in the translational diffusion of the tetramers (Table 1). Following the addition of NH4OAc, all the phosphoestamer diffusion coefficients decreased relative to those in pure D2O. This was expected because salts, like NH4OAc, are known to affect diffusion coefficients.27
More significant was the increasing distinction between diffusion coefficients for each phosphoestamer, particularly C122-HEG2 and (C12-HEG)2. With NH4OAc, the diffusion coefficient of the diblock tetramer was 50 µm2 s−1 lower than that of the alternating tetramer and gave considerable evidence that the diblock sequence formed bigger self-assemblies, which diffuse at a slower rate. This is consistent with work on the 20mers in which the diblock sequence produced larger superstructures than the alternating sequence.13 The symmetric phosphoestamers C12-HEG-HEG-C12 and HEG-C12-C12-HEG formed aggregates that diffused faster than C122-HEG2, but more slowly than (C12-HEG)2. Additionally, the diffusion coefficients correlated well with the average block length within each tetramer (Fig. S32).
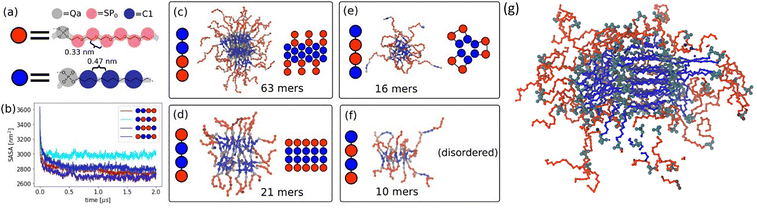
Although DOSY NMR has shown that sequences can be used to fine-tune the assembly of amphiphilic phosphoestamers, it does not provide information on the structural details of the assemblies. We found that because of the relatively small size of these molecules, we were unable to observe them reliably by TEM or AFM, but MD simulations produced models consistent with the experimental DOSY data. A coarse-grained system consisting of the tetramers in water was simulated using standard parameters for the C12 alkane chain and phosphate group (SI) and Rossi et al. parameterisation of HEG (Fig. 2a).28 All simulations reached equilibrium after 1.0 µs, as indicated by the flattening of the solvent-accessible surface area (SASA, Fig. 2b). Analysis of the results (Fig. 2c–f) shows that C122-HEG2 forms the heaviest structures (63mers) with compact and ordered cores, as well as antiparallel oriented C12 chains. A similar, though smaller structure was observed for HEG-C12-C12-HEG, which forms 21mer clusters with compact and ordered cores. The C12 chains in the centre are parallel to each other, while hydrophilic (HEG) groups are solvated in water. The most compact but discrete structure is formed by C12-HEG-HEG-C12. These tetramers form small micelle-like structures (16mers) with the chains in a U-shape such that the hydrophobic C12 groups could be buried in the core, and hydrophilic HEG chains form loops solvated in water. Sporadic cross-links between micelles were also seen with this sequence, in cases where one of the chains was linear. (C12-HEG)2 seems not to form an ordered structure, but a network that spans over the whole system and crosses periodic boundaries with small clusters of no more than 10 phosphoestamers. Snapshots of the entire frames can be found in the SI (Fig. S34).
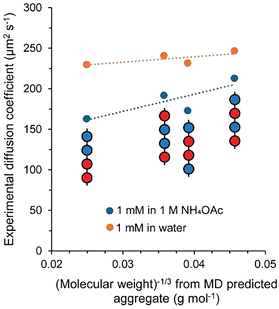
To compare the computational data against the DOSY NMR, the experimental diffusion coefficients (D) were plotted versus the average molecular weight (Mw) of computationally predicted aggregates using the linear relationship between diffusion coefficient and the reciprocal cube root of Mw, D = f(Mw−1/3).29 This was plotted for the tetramers both with and without NH4OAc (Fig. 3) and the results again highlighted that cation driven self-assembly gave greater differences in the translational diffusion of the tetramers, reflective of more extensive self-assembly. This was also observed by plotting the calculated hydrodynamic radii against the radius of gyration of the largest cluster simulated for each tetramer (Fig. S37).
When NH4OAc was used, the translational diffusion of the diblock and alternating tetramers was shown to correlate well with the aggregates predicted by MD simulations. The low diffusion coefficient of C122-HEG2 is consistent with the predicted interdigitated C12 chains leading to compact stacks of layers, which would diffuse slowly. At the other end of the scale, (C12-HEG)2 displays rapid translational diffusion even with the addition of NH4+ cations, validating the small, disordered systems predicted by the MD simulations. These tetramers only have small patches of hydrophobic surface, and as a result, the aggregates are weak and cannot reach critical mass. This causes them to diffuse more similarly to free particle systems than the other sequences. The intermediate sequences, C12-HEG-HEG-C12 and HEG-C12-C12-HEG, form clusters neither as large as C122-HEG2, nor as small as (C12-HEG)2, which is broadly consistent with the computational data, although their order is not the same. Since a linear line can be plotted from the ends through HEG-C12-C12-HEG in both cases, it is likely that C12-HEG-HEG-C12 is the sequence behaving more anomalously, being unexpectedly slow in diffusion (see also Fig. S38 and S39). Gratifyingly, a potential mechanism for this is already presented in the computational model – while C12 termini are folded back into the micelles, some are not, and form cross-links with other micelles, resulting in larger overall aggregates.
Since the coarse-grained models can be reconciled with the experimental results, we were interested in whether we could observe finer structural details. An all-atom simulation (50 ns), back-mapped from the coarse-grained model of the most well-defined system (C122-HEG2) was performed (Fig. 2g) to give insights into potential secondary structure motifs that phosphoestamers could form and thus make a small step closer to development of protein-like structures. The simulation shows that the interior of the clusters is composed of a stack of layered C12-PO4-C12 chains, with the phosphates in register with each other. This structure is reminiscent of lipid bilayers, and is in contrast to the general structure of polymer star micelles, in which there is a radial arrangement of chains, as well as related modelled systems where C12 chains double-back after every phosphoester.30
In summary, we have used phosphoramidite chemistry to produce sequence-defined tetrameric phosphoestamers, making every sequence containing two C12 and two HEG units. DOSY NMR and MD simulations have shown that in pure water, there is little difference due to mutual repulsion arising from the anionic phosphates, but countercations screen the repulsion, and substantial differences in self-assembly occur according to sequence. Hydrophobic aggregation of C12 chains is the primary mechanism for self-assembly. C122-HEG2 forms the largest structures, appearing somewhat like star micelles, but with a parallel, in register, arrangement of interdigitated chains in the core. HEG-C12-C12-HEG also forms micelle-like structures, with parallel C12 chains, but smaller in this case since each chain has a hydrophilic section at both termini and thus spans the whole assembly. C12-HEG-HEG-C12, on the other hand, mainly folds back on itself to satisfy hydrophobic aggregation, giving smaller structures, although on occasion this folding does not occur, resulting in cross-linking with a second aggregate. (C12-HEG)2 appears to be sufficiently able to shield its hydrophobic sections without extensive self-assembly.
This is the first time that self-assembly of non-nucleosidic polyphosphoesters has been studied at this level of detail and the results highlight new motifs which could support the future, bottom-up design of protein-like phosphoestamers that take advantage of the possibilities of the phosphoramidite method.
The authors are grateful to the University of Kent, Centauri Therapeutics, and UCL as well as Rosetrees Trust PhD Project Grant M743 and PhDPlus Project PhD2022\100050 for funding. The authors also thank the UCL School of Pharmacy Nuclear Magnetic Resonance Core Facility (RRID:SCR_027123) for use of the equipment to carry out DOSY NMR experiments.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: synthetic protocols, MS, DOSY spectra, and supplementary MD. See DOI: https://doi.org/10.1039/d5cc06986f.References
- L. Escobar, D. Sun, M. Dhiman and C. A. Hunter, Chem. Commun., 2025, 61, 504–507 Search PubMed.
- J. V. Hoorde, N. Badi and F. E. D. Prez, Polym. Chem., 2024, 15, 4319–4326 RSC.
- Y. Song, C. Sun, C. Tian, H. Ming, Y. Wang, W. Liu, N. He, X. He, M. Ding, J. Li, F. Luo, H. Tan and Q. Fu, Chem. Sci., 2022, 13, 5353–5362 RSC.
- A. J. DeStefano, R. A. Segalman and E. C. Davidson, JACS Au, 2021, 1, 1556–1571 Search PubMed.
- J.-F. Lutz, M. Ouchi, D. R. Liu and M. Sawamoto, Science, 2013, 341, 1238149 Search PubMed.
- J.-F. Lutz, J.-M. Lehn, E. W. Meijer and K. Matyjaszewski, Nat. Rev. Mater., 2016, 1, 16024 Search PubMed.
- N. Appukutti and C. J. Serpell, Polym. Chem., 2018, 9, 2210–2226 RSC.
- A. Al Ouahabi, L. Charles and J.-F. Lutz, J. Am. Chem. Soc., 2015, 137, 5629–5635 CrossRef CAS PubMed.
- M. H. Caruthers, Acc. Chem. Res., 1991, 24, 278–284 CrossRef CAS.
- Y. Yin, R. Arneson, Y. Yuan and S. Fang, Chem. Sci., 2025, 16, 1966–1973 RSC.
- M. Vybornyi, Y. Vyborna and R. Häner, Chem. Soc. Rev., 2019, 48, 4347–4360 RSC.
- K. Pérez De Carvasal, N. Aissaoui, G. Vergoten, G. Bellot, J.-J. Vasseur, M. Smietana and F. Morvan, Chem. Commun., 2021, 57, 4130–4133 RSC.
- N. Appukutti, J. R. Jones and C. J. Serpell, Chem. Commun., 2020, 56, 5307–5310 RSC.
- N. Appukutti, A. H. de Vries, P. G. Gudeangadi, B. R. Claringbold, M. D. Garrett, M. R. Reithofer and C. J. Serpell, Chem. Commun., 2022, 58, 12200–12203 RSC.
- B. Claringbold, S. Vance, A. R. Paul, J. Williamson, M. D. Garrett and C. J. Serpell, Chem. Sci., 2025, 16, 113–123 RSC.
- S. Hanessian, X. Luo, R. Schaum and S. Michnick, J. Am. Chem. Soc., 1998, 120, 8569–8570 CrossRef CAS.
- Y. Fan, J. Li, M. Jiang, J. Zhao, L. He, Y. Wang and F. Shao, Nanoscale, 2024, 16, 17964–17973 RSC.
- X.-J. Zhai, M.-Y. Luo, X.-M. Luo, X.-Y. Dong, Y. Si, C. Zhang, Z. Han, R. Han, S.-Q. Zang and T. C. W. Mak, Nat. Commun., 2024, 15, 9155 CrossRef CAS PubMed.
- V. T. Ravikumar, R. K. Kumar, P. Olsen, M. N. Moore, R. L. Carty, M. Andrade, D. Gorman, X. Zhu, I. Cedillo, Z. Wang, L. Mendez, A. N. Scozzari, G. Aguirre, R. Somanathan and S. Berneès, Org. Process Res. Dev., 2008, 12, 399–410 CrossRef CAS.
- D. H. Wu, A. D. Chen and C. S. Johnson, J. Magn. Reson., Ser. A, 1995, 115, 260–264 CrossRef CAS.
- E. Ruzicka, P. Pellechia and B. C. Benicewicz, Anal. Chem., 2023, 95, 7849–7854 CrossRef CAS PubMed.
- S. Habuchi, S. Fujiwara, T. Yamamoto and Y. Tezuka, Polym. Chem., 2015, 6, 4109–4115 RSC.
- P. Groves, Polym. Chem., 2017, 8, 6700–6708 RSC.
- M. Iwai, C.-G. Pack, Y. Takenaka, Y. Sako and A. Nakano, Sci. Rep., 2013, 3, 2833 CrossRef PubMed.
- Z.-J. Tan and S.-J. Chen, Biophys. J., 2006, 90, 1175–1190 CrossRef CAS PubMed.
- K. Salorinne, T. Lahtinen, S. Malola, J. Koivisto and H. Häkkinen, Nanoscale, 2014, 6, 7823–7826 RSC.
- B. Filova, L. Musilova, A. Mracek, M. L. Ramos, L. M. P. Veríssimo, A. J. M. Valente and A. C. F. Ribeiro, J. Mol. Liq., 2020, 304, 112728 CrossRef CAS.
- G. Rossi, J. Barnoud and L. Monticelli, J. Phys. Chem. Lett., 2014, 5, 241–246 CrossRef CAS PubMed.
- D. P. Valencia and F. J. González, Electrochem. Commun., 2011, 13, 129–132 CrossRef CAS.
- T. Trinh, C. Liao, V. Toader, M. Barłóg, H. S. Bazzi, J. Li and H. F. Sleiman, Nat. Chem., 2018, 10, 184–192 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |