Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Internalization of antibiotics by the multi-resistant bacteria Acinetobacter baumannii through the CarO outer membrane porin
Florent
Barbault
* and
Antonio
Monari
 *
*
Université Paris Cité and CNRS, ITODYS, F-75006, Paris, France. E-mail: antonio.monari@u-paris.fr; florent.barbault@u-paris.fr
First published on 30th January 2026
Abstract
Antibiotic resistance represents an emerging global health threat. In this contribution, we report all-atom molecular dynamics simulations of a bacterial protein in Acinetobacter baumanii. By using enhanced sampling we show the favorable internalization of the antibiotic via the CarO porin. This result correlates with the knockout or the mutation of the porin as a resistance mechanism.
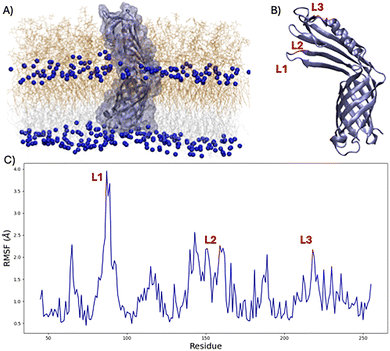
The discovery of antibiotics has revolutionized medicine, allowing the treatment and management of life-threatening diseases and the efficient control of bacterial-based epidemics.1 This includes diseases like the plague or cholera, which have been mostly eradicated at least in developed countries, although sporadic surges still occur periodically.2 Yet almost no novel antibiotics have been introduced to the market or entered clinical use in recent years. This situation, combined with their widespread use, has driven the emergence of resistance mechanisms, which significantly diminish their effectiveness.3,4 These mechanisms may encompass diverse strategies including collective organization, such as biofilm formation.5 Specific mutations leading to the production of antibiotic degrading enzymes or efflux pumps to expel antibiotics have also been reported. Furthermore, the knock-out or inactivation by specific mutations of membrane porins is also employed as a surviving strategy. The latter is particularly efficient in the case of Gram-negative bacteria, which are characterized by an outer lipid membrane, surrounding the periplasmic region and the inner membrane.6,7 Indeed, the outer membrane, which is also composed of lipopolysaccharides (LPS), offers an efficient barrier to the spontaneous internalization of external molecules, in addition to being involved in adhesion and migration via its specific membrane proteins. The emergence of multi-resistant bacterial strains, which are insensitive to different classes of antibiotics, is particularly problematic and requires particular attention.3 This is even more stringent in the case of bacteria responsible for nosocomial infections, which often affect fragile or immunocompromised patients.8 Indeed, it is expected that in the next few years the number of deaths due to antibiotic resistance will rise considerably, reaching the value of 40 million in 2050.9 The development of novel therapeutic strategies, which may also include photodynamic inactivation, is therefore urgently needed. The World Health Organization (WHO) has defined a list of bacteria, known under the acronym ESKAPE, which comprises priority pathogens exhibiting multi-resistance and for which the development of new antibiotics is necessary. Among the members of the ESKAPE family, one particularly critical component is the Gram-negative Acinetobacter baumannii.10 While A. baumannii is usually benign in healthy and immunologically active patients it may develop severe infections in compromised individuals.11 Its spread in nosocomial environments, combined with the pronounced adaptability and resistance mechanisms, makes this bacterium one of the most important targets to be tackled. A. baumannii nosocomial infection in nosocomial environments may also correlate with bad prognosis in ventilator induced pneumonia.12 The defence and resistance mechanisms of A. baumannii are diverse and include shielding from the host immune response via the interaction with endogenous proteins like fibronectin13 or the formation of biofilms.14 The outer membrane of A. baumannii is rich in different proteins, including porins, signalling agents, and adhesion mediators.15,16 Porins allow the internalization of fundamental nutrients in the periplasmic space and are characterized by differential selectivity towards ionic or small organic molecules, including antibiotics.17 The selectivity of porins is usually related to the size of their central channel and its electrostatic environment, allowing to differentiate between anions and cations.6,18 Interestingly, some porins are also directly involved in reinforcing membrane structural stability and in providing adhesion capacity.13,19 Among the different porins in A. baumannii a crucial role is played by CarO,20 which is related to the induction of carbapenem resistance.21 Indeed, the bacteria may either suppress the expression of CarO or produce non-functional protein variants via point mutations or truncation. This leads to a highly reduced internalization of carbapenems, which correlates with antibiotic resistance and increased virulence of the bacterial strains. Therefore, understanding the mechanism of the antibiotic internalization through this channel is of high importance to understand the resistance mechanisms. The structure of CarO has been experimentally resolved22 showing a rather particular arrangement especially compared to other porins (Fig. 1). In addition to a common transmembrane channel composed of a rigid β-barrel arrangement, a glove-like structure is present in the extracellular region composed of both rigid β-sheets, one α-helix, and more flexible loops. Interestingly, the glove resides in the sugar rich region of the external membrane leaflet and may be involved in assisting internalization by preferentially selecting transferable compounds and funnelling them towards the channel entrance. A disordered and unresolved periplasmatic tail is also present and comprises 44 amino acids at the N-terminus of the protein. The periplasmatic tail may also interact with the peptidoglycans leading to enhanced structural stability of the outer membrane. Because of its importance in antibiotic resistance, different studies have considered CarO selectivity and transport of antibiotics,21,23 conducted both at an experimental and computational level, including via all atom molecular dynamics (MD).24 However, contrasting results over the involvement of CarO in the transport of the imipenem (IMP) carbapenem antibiotic have been reported.25 While in vivo results20,23 have correlated the knock-down of CarO with IMP resistance, its penetration through CarO sensitized liposomes in vitro has not been fully confirmed yet.25 Thus, in this contribution, we leverage the potentiality of all atom MD simulations complementing equilibrium dynamics with free-energy methods and enhanced sampling approaches to get the potential of mean force (PMF) describing the internalization of IMP via Caro. Importantly, our model system involves a realistic Gram-negative outer membrane, showing asymmetry between the extracellular and the periplasmatic leaflet. The inclusion of specific A. baumannii LPS in the external leaflet is also fundamental to fully describe the interactions in the glove region and rationalize the internalization driving force. The initial system has been built using the Charmm-gui web tools,26 and protein and lipids have been described using the charm force field. Water buffers including 0.15 M physiological concentration of NaCl have been added. All the MD simulations have been performed using the NAMD code.27,28 The PMF has been obtained with the Umbrella Sampling (US) methods as implemented in the Colvar module29 for a total sampling of 6.5 µs. Images have been rendered using VMD.30 The full computational procedure can be found in the SI.
From the 700 ns equilibrium simulation (Fig. 1), we observe that the β-barrel region of CarO is favourably inserted in the membrane core, leading to a stable arrangement. This is also confirmed by the analysis of the time evolution of the root mean square deviation (RMSD), which is provided in the SI. The glove region, which is a peculiar characteristic of CarO, extends farther away from the membrane, encompassing the sugar region, and is also globally stable. Interestingly, this global stability masks a more complicated behaviour with slight, yet significant variations in the flexibility profile of the protein, as can be observed in the per residue Root Mean Square Fluctuation (RMSF) reported in Fig. 1C.
Indeed, while the RMSF for the amino acids constituting the β-barrel is particularly low and remains below the 2.0 Å limit, some flexibility peaks appear. This involves most notably the triad S87, V88, and N89, which are positioned at the unstructured end of loop 1 (L1, see Fig. 1B for the correspondence in the protein structure). A secondary region of higher flexibility involves the loop 2 (L2) edge. Differently from L1, this region encompasses a larger number of amino acids going from K139 to E162, yet without achieving the flexibility maximum observed for L1. Finally, a third flexibility peak is present in correspondence with G219 partially extending up to G125, which corresponds to the terminal part of the loop 3 (L3) region. Importantly, the additional higher flexibility domains correspond to either the truncated C- and N-terminal regions or the short loops connecting the barrel β-sheets at the internal leaflet polar heads. As previously mentioned, the glove region extends towards the outer leaflet polar heads and directly interacts with the external sugars.
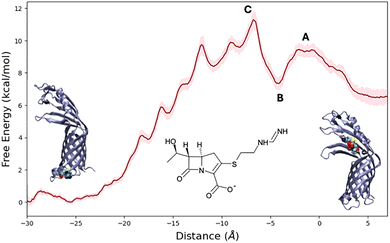
Furthermore, from the equilibrium MD one can appreciate the establishment of a water channel inside CarO's β-barrel domain, which can be seen in the SI. While the channel appears globally continuous allowing communication between the periplasmatic and the extracellular spaces, it is also narrow. Indeed, only one molecule of water can occupy the channel space. The diameter of the pore has been estimated to be comprised between 3 and 4 Å. Therefore, questions may arise as to the capacity of CarO to efficiently favour the internalization of organic molecules such as antibiotics, without requiring a considerable reorganization energy penalty. Indeed, contrasting experimental evidence has been provided regarding the capacity of CarO to efficiently internalize carbapenem and specifically IMP.25 To rationalize the internalization of the IMP antibiotic through CarO, we have performed US enhanced sampling along the distance of the center of mass of IMP and the outer membrane polar heads projected on the normal to the membrane plane. Note that the deprotonated form of the carboxylic acid of IMP has been considered solely, due to its predominance at neutral pH. This strategy allows obtaining the free energy profile for the antibiotic insertion into the periplasmic space of the bacterium. The obtained free energy profile is reported in Fig. 2.
One can immediately see that the specific composition of the bacterial membrane induces a strong asymmetry, which leads to a consistent driving force for the internalization of IMP. As a matter of fact, the antibiotics are more stable by about 6.5 kcal mol−1 when residing in the periplasmic space (negative values of the collective variable) than when being positioned in the extracellular sugar region and more specifically in the glove space (positive values of the collective variable). This result is not surprising due to the specific composition of the bacterial external membrane as compared to more usual lipid bilayers. Upon interacting with CarO, IMP should first bypass a very slight energy barrier (∼3 kcal mol−1, point A in Fig. 2), which corresponds to the slightly hydrophobic entrance of the channel. By going deeper into the protein, we observe the emergence of a well-defined free energy minimum characterized by a stabilization of about 3 kcal mol−1 (point B in Fig. 2). The presence of such an intermediate state is also coherent with previous equilibrium MD simulations24 and may be regarded as a factor favoring the accumulation of the compounds to be internalized in the vicinity of the channel. To leave the intermediate state, IMP should overcome a free energy barrier of about 5 kcal/mol and reach the transition state (point C in Fig. 2) which represents the veritable rate-limiting step governing the whole internalization process. Indeed, and rather surprisingly, no other free energy barrier can be found on the pathway diving deeper into the porin channel and reaching the periplasmatic space. In contrast, an important free energy gradient can be observed imposing a driving force towards the internalization amounting at about 12 kcal mol−1. Interestingly some local maxima in the PMF can be observed; however, without exceeding 1.0-1.5 kcal mol−1. These points correspond to local deformations of the β-barrel, which are necessary to overcome the steric hindrance due to the passage of IMP. However, they do not impose energy penalties which may significantly slow down the internalization of the antibiotic. The peculiar free energy landscape identified by our PMF also confirms that the bacterial porin is clearly designed to favor the unidirectional internalization instead than the expulsion of exogenous compounds. Indeed, while the internalization is energetically favorable and only requires overcoming a small to moderate 5 kcal mol−1 barrier, the expulsion of an internalized IMP would necessitate passing a barrier of about 11 kcal mol−1. On top of these considerations, we may also add that the concentration of IMP or nutrients is expected to be higher in the extracellular regions than in the periplasmatic space, thus further justifying their internalization.
In Fig. 3, we report three significant snapshots for the main transition and intermediate states, as previously described. It appears evident that the small free energy maximum at A may be connected from one side to the reduced polarity of the entrance of the channel and to the reduction of the entropic factors due to the confinement of IMP at the entrance of the channel. In contrast, the stabilization at the intermediate state B is mainly due to favorable electrostatic and hydrogen bond interactions, mainly involving the carboxylate moiety of IMP. As also shown in the zoomed-in inlay at this position, the COO− is stabilized by forming a strong and persistent salt bridge with the NH3+ group of K278. An additional stabilization is also provided by the formation of a rather specific hydrogen bond with N76. From the distribution of the atom-atom distances describing these interactions and reported in Fig. 3D we may see that the salt bridge with the lysine forms a sharp and extremely persistent distribution having an almost ideal Gaussian shape. In contrast, N76 is less persistent and the interaction is temporarily loosened or lost as indicated by the tail in the distribution spanning larger distances and by the more pronounced width of the distribution. The subsequent free energy maximum at position C is mainly due to the breaking of this favorable conformation and by the less important polar character of the channel. On the other hand, the almost spontaneous sliding of IMP towards the periplasmatic region is essentially due to the polar nature of the amino acids surrounding the β-barrel and to the small or rather moderate reorganization free energy. It has been reported that knock-out of CarO or mutations affecting the channel entrance area induce IMP and carbapenem resistance in A. baumannii.22,25 This evidence is well substantiated by our MD simulations since CarO favors the passage of IMP through the outer membrane of the bacteria. In some of the resistant strains, the CarO sequence is truncated, missing the glove region and presenting a suboptimal channel gate. This may lead to an inefficient funnelling of the antibiotic to the porin, notably by perturbing its interaction at the level of the intermediate state.
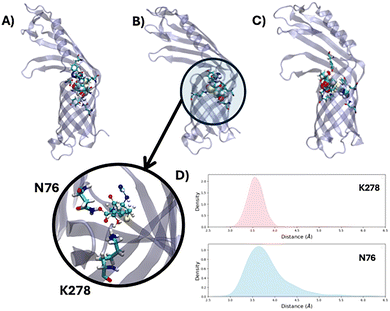 | ||
| Fig. 3 Representative snapshot of IMP (in van der Waals representation) interacting with CarO (transparent cartoon) at the first transition state (A), at the intermediate state (B), and the second transition state (C). Note that these positions of the free energy landscape are also evidenced with the same letters in Fig. 2. The zoomed-in view of IMP interacting with two key amino acids (N76 and K28) is also reported as an inlay, together with the distribution of the distances between the COO–Carbon atom and the lateral chain nitrogen of the two amino acids as a Kernel Density Estimation (D) analyzed in the US window constraining the collective variable to the value of −6.0 Å. The corresponding time evolution is provided in the SI. | ||
The effects of the membrane asymmetry may also reconcile the contrasting experimental results and notably explain why the internalization of IMP has not been observed in artificial liposomes, which may not be able to provide the necessary driving force.25 We have provided evidence of the capacity of CarO to favor internalization of carbapenems and specifically IMP. The internalization process is spontaneous, also due to the bacterial membrane asymmetry, and requires bypassing only a very moderate activation free energy. An intermediate state is found in the vicinity of the channel gate and may participate in accumulating the antibiotic at the entrance of the porin. Our results underly the crucial role of CarO in assuring antibiotic transport and are coherent with the fact that its suppression or mutation correlates with the emergence of antibiotic resistance. Our study sets the base to a better understanding of key processes related to multi-drug resistance in a critical ESKAPE bacterium, particularly involved in nosocomial infections. Thus, they may participate in the quest to develop viable therapeutic or prophylactic strategies to counteract the emergence of resistant strains.
The authors thank GENCI, Explor, and the Platform P3MB for the computational resources.
Conflicts of interest
There are no conflicts to declare.Data availability
Data available at Zenodo server, DOI https://doi.org/10.5281/zenodo.17769189.Supplementary information (SI): extended computational methodology, RMSD and density profile, description of the collective variable, and analysis of the free energy convergence. See DOI: https://doi.org/10.1039/d5cc06876b.
References
- M. I. Hutchings, A. W. Truman and B. Wilkinson, Curr. Opin. Microbiol., 2019, 51, 72–80 Search PubMed.
- T. Butler, Clin. Infect. Dis., 2009, 49, 736–742 CrossRef PubMed.
- N. Macesic, A.-C. Uhlemann and A. Y. Peleg, Lancet, 2025, 405, 257–272 Search PubMed.
- A. Gauba and K. M. Rahman, Antibiotics, 2023, 12, 1590 Search PubMed.
- W. C. Reygaert, AIMS Microbiol., 2018, 4, 482–501 CAS.
- J. D. Prajapati, U. Kleinekathöfer and M. Winterhalter, Chem. Rev., 2021, 121, 5158–5192 CrossRef CAS PubMed.
- N. C. Rosas and T. Lithgow, Trends Microbiol., 2022, 30, 544–552 Search PubMed.
- H. A. Khan, F. K. Baig and R. Mehboob, Asian Pac. J. Trop. Biomed., 2017, 7, 478–482 CrossRef.
- M. Naghavi, S. E. Vollset and K. S. Ikuta, et al. , Lancet, 2024, 404, 1199–1226 CrossRef PubMed.
- A. Howard, M. O’Donoghue, A. Feeney and R. D. Sleator, Virulence, 2012, 3, 243–250 CrossRef PubMed.
- H. Mukhopadhyay, A. Bairagi, A. Mukherjee, A. K. Prasad, A. D. Roy and A. Nayak, Curr. Res. Microb. Sci., 2025, 8, 100331 Search PubMed.
- J. Inchai, C. Pothirat, C. Bumroongkit, A. Limsukon, W. Khositsakulchai and C. Liwsrisakun, J. Intensive Care, 2015, 3, 9 CrossRef PubMed.
- L. Vasseur, F. Barbault and A. Monari, Chem. – Eur. J., 2025, 31, e00874 Search PubMed.
- J. Choudhary and M. Shariff, Sci. Rep., 2025, 15, 33892 Search PubMed.
- S. R. Uppalapati, A. Sett and R. Pathania, Front. Microbiol., 2020, 11, 589234 Search PubMed.
- S. Skariyachan, N. Taskeen, M. Ganta and B. Venkata Krishna, Crit. Rev. Microbiol., 2019, 45, 315–333 CrossRef PubMed.
- D. Nie, Y. Hu, Z. Chen, M. Li, Z. Hou, X. Luo, X. Mao and X. Xue, J. Biomed. Sci., 2020, 27, 26 CrossRef CAS PubMed.
- R. Benz, Current Topics in Membranes and Transport, 1984, pp. 199–219 Search PubMed.
- D. Scribano, E. Cheri, A. Pompilio, G. Di Bonaventura, M. Belli, M. Cristina, L. Sansone, C. Zagaglia, M. Sarshar, A. T. Palamara and C. Ambrosi, Commun. Biol., 2024, 7, 948 Search PubMed.
- G. Labrador-Herrera, A. J. Pérez-Pulido, R. Álvarez-Marín, C. S. Casimiro-Soriguer, T. Cebrero-Cangueiro, J. Morán-Barrio, J. Pachón, A. M. Viale and M. E. Pachón-Ibáñez, Virulence, 2020, 11, 1727–1737 CrossRef CAS PubMed.
- M. A. Mussi, A. S. Limansky and A. M. Viale, Antimicrob. Agents Chemother., 2005, 49, 1432–1440 CrossRef CAS PubMed.
- M. Zahn, T. D’Agostino, E. Eren, A. Baslé, M. Ceccarelli and B. van den Berg, J. Mol. Biol., 2015, 427, 2329–2339 CrossRef CAS PubMed.
- S. Royer, P. A. De Campos, B. F. Araújo, M. L. Ferreira, I. R. Gonçalves, D. W. D. F. Batistão, R. T. E. S. Brígido, L. T. Cerdeira, L. G. Machado, C. S. De Brito, P. P. Gontijo-Filho and R. M. Ribas, PLoS One, 2018, 13, e0198643 Search PubMed.
- E. Sariyer, Res. Microbiol., 2022, 173, 103966 CrossRef CAS PubMed.
- M. Zahn, S. P. Bhamidimarri, A. Baslé, M. Winterhalter and B. van den Berg, Structure, 2016, 24, 221–231 CrossRef CAS PubMed.
- S. Jo, T. Kim, V. G. Iyer and W. Im, J. Comput. Chem., 2008, 29, 1859–1865 CrossRef CAS PubMed.
- J. C. Phillips, R. Braun, W. Wang, J. Gumbart, E. Tajkhorshid, E. Villa, C. Chipot, R. D. Skeel, L. Kalé and K. Schulten, J. Comput. Chem., 2005, 26, 1781–1802 CrossRef CAS PubMed.
- J. C. Phillips, D. J. Hardy, J. D. C. Maia, J. E. Stone, J. V. Ribeiro, R. C. Bernardi, R. Buch, G. Fiorin, J. Hénin, W. Jiang, R. McGreevy, M. C. R. Melo, B. K. Radak, R. D. Skeel, A. Singharoy, Y. Wang, B. Roux, A. Aksimentiev, Z. Luthey-Schulten, L. V. Kalé, K. Schulten, C. Chipot and E. Tajkhorshid, J. Chem. Phys., 2020, 153, 044130 CrossRef CAS PubMed.
- G. Fiorin, M. L. Klein and J. Hénin, Mol. Phys., 2013, 111, 3345–3362 CrossRef CAS.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graphics, 1996, 14, 33–38 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |