Ancestral sequence reconstruction reveals new functional fluorinases and mechanistic insights into enzymatic fluorination
Shreyas
Supekar†
 *a,
Wan Lin
Yeo†
*a,
Wan Lin
Yeo†
 b,
Elaine
Tiong†
b,
Elaine
Tiong†
 c,
Juliana
Rizal
c,
Juliana
Rizal
 d,
Ee Lui
Ang
d,
Ee Lui
Ang
 *be,
Fong Tian
Wong
*be,
Fong Tian
Wong
 *cd,
Yee Hwee
Lim
*cd,
Yee Hwee
Lim
 *ce and
Hao
Fan
*ce and
Hao
Fan
 *aef
*aef
aBioinformatics Institute (BII), Agency for Science, Technology and Research (A*STAR), 30 Biopolis Street, #07-01 Matrix Building, Singapore 138671, Republic of Singapore. E-mail: supekars@a-star.edu.sg
bSingapore Institute of Food and Biotechnology Innovation (SIFBI), Agency for Science, Technology and Research (A*STAR), 31 Biopolis Way, #04-01 Nanos, Singapore 138669, Republic of Singapore. E-mail: ang_ee_lui@a-star.edu.sg
cInstitute of Sustainability for Chemicals, Energy and Environment (ISCE2), Agency for Science, Technology and Research (A*STAR), 8 Biomedical Grove, #07-01 Neuros Building, Singapore 138665, Republic of Singapore. E-mail: wong_fong_tian@a-star.edu.sg; lim_yee_hwee@a-star.edu.sg
dInstitute of Molecular and Cell Biology (IMCB), Agency for Science, Technology and Research (A*STAR), 61 Biopolis Drive, Proteos #07-06, Singapore, 138673, Republic of Singapore
eSynthetic Biology Translational Research Program, Yong Loo Lin School of Medicine, National University of Singapore, 10 Medical Drive, Singapore 117597, Republic of Singapore
fInstitute for Molecular and Cellular Therapeutics, Chinese Institutes for Medical Research, Beijing 100069, China. E-mail: fanhao@cimrbj.ac.cn
First published on 4th February 2026
Abstract
Fluorinases catalyze carbon–fluorine bond formation and represent a rare but valuable enzyme class. We employed ancestral sequence reconstruction to expand fluorinase diversity, identifying seven functional enzymes (28–79% conversion). Comparative analysis revealed previously uncharacterized residues crucial for fluorination, enabling activity rescue in a low-activity ancestral variant.
The carbon–fluorine bond is widely used in pharmaceuticals, agrochemicals, and materials due to its unique properties, including enhanced metabolic stability and membrane permeability.1–3 Although fluorine is plentiful in nature, enzymes that incorporate fluorine into organic molecules are extremely rare, with only a few fluorinases identified so far.4–8 Fluorinase catalyzes nucleophilic SN2 substitution of S-adenosyl-L-methionine (SAM) with inorganic fluoride (F−) to produce 5′-fluoro-5′-deoxyadenosine (5′-FDA) and L-methionine.9 Fluorinase functions as a trimer with the active site at the interface of monomers (Fig. 1).10 Although fluorinases and chlorinases catalyze halide incorporation via similar mechanisms, their substrate specificities are notably asymmetric. Fluorinase from Streptomyces sp. MA37 (FLA_MA37),5 the most efficient fluorinase characterized to date, can also catalyze chlorination; however, naturally occurring chlorinases cannot perform fluorination.
Given the scarcity of natural fluorinases and their industrial relevance, there is significant interest in expanding their repertoire through protein engineering approaches.1,11 Traditional methods like directed evolution and rational design have yielded incremental improvements but are constrained by limited sequence space.12,13
Ancestral sequence reconstruction (ASR) enables exploration of deep evolutionary sequence space to access functional proteins beyond the reach of traditional engineering.14–18 In this study, we apply ASR to fluorinases to expand the biocatalytic toolkit for fluorination. Here, we identified seven catalytically active ancestral fluorinases, with the best ancestral variant matching the activity of top-performing natural fluorinase FLA_MA37, while exhibiting improved F−/Cl− selectivity. Comparative sequence analysis revealed key residues for fluorination, validated through targeted mutations that rescued activity in a low-performing ancestor, achieving ca. 2.2-fold improvement. MD simulations were also employed to identify structural determinants for fluorinase function. These findings demonstrate that ASR effectively expands accessible sequence space beyond traditional enzyme engineering approaches, providing novel biocatalysts and mechanistic insights into these rare enzymes.
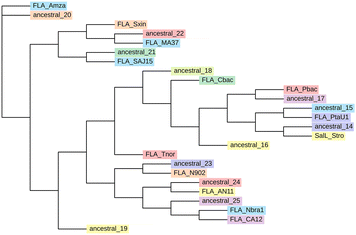
To identify candidate sequences for ancestral reconstruction, we started with the fluorinase from Streptomyces sp. MA37 (FLA_MA37, Uniprot: Q70GK9, PDB: 5B6I).5 The query FLA_MA37 sequence was searched against NCBI NR database using PSI-BLAST to identify fluorinase homologues. Homologues were filtered for 30–90% sequence identity, appropriate length, and presence of catalytic residues (D16, Y77, S158, D210, N215; Fig. 1), then clustered at 90% identity yielding thirteen representative sequences for ASR. The candidate set comprised thirteen selected homologs including more recently discovered fluorinases from S. xinghaiensis, S. sp. MA37, Actinoplanes sp. N902-109 and Methanosaeta sp. PtaU1 (Table S1).7,19,20 The candidate set also contained a chlorinase from S. tropica (SalL)13 which exhibits distant homology to fluorinases but lacks any fluorination function. The sequences shared 58–94% identity at the amino acid level. Multiple sequence alignment (MSA) of the candidate set was performed with MAFFT.21 FastTREE22 was used to construct an initial a maximum likelihood phylogenetic tree for the candidate set MSA (Fig. S1).
The twelve ancestral nodes (ancestral_14 through ancestral_25) from the phylogenetic tree representing the divergence points in fluorinase evolution were inferred using iqtree23 suite (Fig. 2). High reconstruction confidence was confirmed with >90% of positions showing posterior probabilities >0.8 (Table S2; SI data). Ancestors shared 38–94% identity with extant homologs (Fig. S2 and Table S3). Reconstructed ancestral_20 was excluded from our ancestral set as it shares 99% sequence similarity with ancestral_19, yielding eleven putative fluorinases for characterization.
Based our previous work on naturally occurring fluorinases,5 we optimized solubility conditions to ensure successful expression and characterization. We expressed and tested the activity of two distinct ancestral sequences, specifically ancestral_25 and ancestral_15, using various solubility tags (Table S4). NT1124 and SUMO25 solubility tags were chosen were for the expression of the remaining nine ancestral sequences (Table S5).
Fluorination activity (monitoring 5′-FDA formation) of the ancestral enzymes were evaluated using high-throughput (HTP) lysates with SAM and F− as substrates. Nine reconstructed enzymes (ancestral_16-25) demonstrated fluorination activity, while ancestral_14 and ancestral_15 showed no detectable product formation (Table S5). Chloride (Cl−) was also tested to check for substrate promiscuity. All active variants retained F− selectivity over Cl− (Table S5).
To quantify the catalytic performance, we purified the top active variants and determined conversion yields by HPLC (Table 1; see SI methods). We find that all the seven tested fluorinases retained their preference for F− over Cl−. The best performing ancestral variant, ancestral_22, exhibited a fluorination capacity comparable to the best known fluorinase FLA_MA37 (Table 1), while showing higher F−/Cl− selectivity. To our knowledge, this is the first demonstration that non-natural fluorinases, including ancestral_16, ancestral_22, and ancestral_25, can achieve performance comparable to natural enzymes, with improvement in selectivity. Protein thermal shift assays revealed that most ancestral variants exhibit comparable or better thermostability compared to FLA_MA37 (Tm = 48.5 °C), with ancestral_16 showing the highest stability (Tm = 64.8 °C). Notably, the best-performing variant ancestral_22 exhibits slightly reduced thermostability (Tm = 46.2 °C), suggesting a potential trade-off between catalytic efficiency and protein stability. Equally important, the addition of these seven new fluorinases to the previous narrow set of 20 known fluorinases expands the available fluorinase repertoire, enabling the identification of new sites for improving enzymatic fluorination.
| Variant | % conversion (24h) | F/Cl 24h selectivity | Protein melting temperature (Tm) | |
|---|---|---|---|---|
| Fluorination | Chlorination | |||
| FLA_MA37 | 82.20 ± 0.30 | 15.21 ± 0.16 | 5.40 | 48.5 ± 0.1 |
| A_16 | 28.57 ± 0.03 | 3.67 ± 0.01 | 7.79 | 64.8 ± 0.1 |
| A_19 | 41.34 ± 0.63 | 11.37 ± 0.05 | 3.64 | 55.8 ± 0.0 |
| A_21 | 51.76 ± 0.74 | 19.01 ± 0.10 | 2.72 | 50.5 ± 0.4 |
| A_22 | 79.17 ± 0.87 | 12.49 ± 0.25 | 6.34 | 46.2 ± 0.2 |
| A_23 | 66.07 ± 0.15 | 14.68 ± 0.18 | 4.50 | 53.2 ± 0.0 |
| A_24 | 66.88 ± 0.19 | 15.16 ± 0.19 | 4.41 | 54.5 ± 0.1 |
| A_25 | 52.49 ± 0.17 | 8.44 ± 0.08 | 6.22 | 59.2 ± 0.2 |
| A_16_M13 | 51.14 ± 0.61 | 9.25 ± 0.09 | 5.53 | 62.0 ± 0.1 |
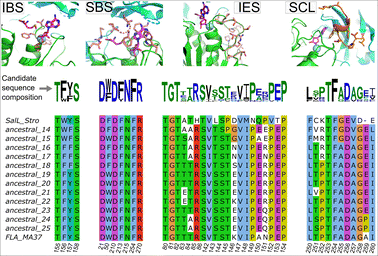
To identify sequence–function relationships, we examined across four key components of fluorinase: (1) the ion binding site (IBS), (2) the SAM-binding site (SBS), (3) the ion egress site (IES), and (4) the SAM capping loop (SCL) (Fig. 3).
In the IBS, we find that residues T155 and S158 (residue numbering is based on 1RQP) are conserved in all reconstructed sequences (Fig. 3). Ancestral_14 and ancestral_15, which exhibit no fluorination activity contain the F156W substitution. Interestingly, ancestral_14 and ancestral_15 are the predecessors to SalL_Stro and FLA_PtaU1, respectively which also exhibit no activity in our fluorination assays. Substitution Y157F found in ancestral_14–18 also leads to weak or no activity. These sequence-activity correlations suggest that F156W substitution is detrimental to fluorination, while Y157F reduces fluorination efficiency.
For the SBS residues, all positions except W50 are conserved. W50F (Fig. 3, 1RQP numbering) substitution was similarly present in the dead ancestral_14 variant, or the low activity ancestral_16–18 variants, mirroring a similar effect as seen in Y157F in the IBS, indicating W50 might be of functional importance for fluorination. In the IES, a recently identified chain of residues proposed to function as a channel for guiding F− to the active site, most residues are conserved. As described previously,7 T80 and T82, that are crucial to fluorination function are conserved in all predicted ancestors. Similarly, another crucial residue R85, proposed to be the “hook” that guides F− ion to the IBS is also conserved in all ancestral sequences. Some variation is seen at T83, N151 and K146 (Fig. 3, 1RQP numbering), especially for moderate to good variants ancestral_19–25. A lysine residue at position 146, present in FLA_MA37 and ancestral_22 seems to improve fluorinase function, while K146G substitution, present in ancestral_14 and ancestral_15 leads to abolishment of fluorination function.
The SCL is a 11-residue loop in proximity of the cofactor SAM, that could serve as a gating element that opens to allow SAM access to the fluorinase active site. The role of SCL has not been probed in fluorinase studies yet. In the SCL, while most residues are conserved, we find the substitution A259P (1RQP numbering) in ancestral_24 and ancestral_25 that is not present in other predicted ancestors. Notably, other known fluorinases possess E259 at this position, while the best-performing natural fluorinase, FLA_MA37, uniquely contains A259. At the position 250, SalL_Stro, ancestral_14 and ancestral_15 (no fluorination activity) contain F, while L250 is conserved in all other ancestral sequences and all known fluorinases with fluorination activity. Another interesting variation is seen at the position 252 (Fig. 3, 1RQP numbering), where the chlorinase SalL_Stro exhibits a lysine residue and the low activity ancestral_14–17 contain an arginine residue in this position. Conversely, the better performing ancestral sequences and FLA_MA37 contain a proline residue in this position. We thus hypothesize that L250 and P252 are important for fluorination, and mutations at position 250 and 252 might lead to abolishment or severe reduction of fluorination activity, respectively.
To investigate whether the identified key residues from sequence–function analysis play a critical role in fluorination activity, we probed ancestral_16, that exhibits low fluorination activity and contains substitutions at positions 50 (SBS), 157 (IBS), and 252 (SCL). With the goal of enhancing its fluorination capability, we introduced three single mutations into ancestral_16, namely: F50W (M1), F157Y (M2), and R252P (M3), along with corresponding double mutants M12, M13, M23, and the triple mutant M123. Our high-throughput assays showed that these mutations did not enhance chlorination, which remains undetectable in these enzymes (Table S7). The M1 mutant showed only a slight improvement, while M2 and M3 exhibited no fluorination activity. In contrast, while the mutants M12, M23, and M123 maintained their activity, the mutant M13 (F50W, R252P) mutant exhibited a vast increase in fluorination activity compared to ancestral_16 (∼4.3-fold). Upon validation using purified proteins, the M13 mutant showed at least 1.8-fold improvement in fluorination. (Table 1).
To elucidate the mechanistic principles underlying the fluorination capabilities of the predicted ancestral fluorinases, we performed comparative molecular dynamics (MD) simulations of FLA_MA37, ancestral_22, ancestral_16, and the single, double, and triple mutants of ancestral_16 (M1, M2, M3, M12, M13, M23, M123). Homology models were built with Schrodinger26 using Streptomyces cattleya fluorinase (PDB ID: 1RQP) as template. Following equilibration, triplicate all-atom MD simulations (3 × 300 ns) were performed for each system using Desmond.27 Analysis was performed with MDAnalysis.28
The high solvation energy of the fluoride ion renders it a weak nucleophile in the presence of water, potentially hindering its fluorination function. To examine this effect, we calculated water permeation in the vicinity of S158 (SBS) from MD simulations. FLA_MA37 exhibits the least permeation, followed by ancestral_22 and M13, while ancestral_16, M23 and M123 show the highest water occupancy (Table 2). Remarkably, this with the experimental observations of fluorination (Table 1 and S5, S7), suggesting that active site hydrophobicity contributes to catalytic efficiency.
| Variant | Water permeation | |
|---|---|---|
| Trimer (%) | Monomer (%) | |
| FLA_MA37 | 23.1 | 7.7 |
| Ancestral_22 | 42.51 | 16.14 |
| Ancestral_16 | 78.93 | 32.82 |
| Ancestral_16_m1 | 58.66 | 28.51 |
| Ancestral_16_m2 | 64.11 | 25.93 |
| Ancestral_16_m3 | 62.78 | 27.32 |
| Ancestral_16_m12 | 58.39 | 27.25 |
| Ancestral_16_m13 | 45.7 | 21.88 |
| Ancestral_16_m23 | 78.07 | 43.6 |
| Ancestral_16_m123 | 86.74 | 44.09 |
The variants ancestral_16, M2, M3, M12, and M23 show elevated SAM and SBS dynamics, consistent with reduced activity (Fig. S3–S8 and Table S8, see SI for details). To probe mutations at positions 50 (SBS), 157 (IBS), and 252 (SCL), we analyzed key intermolecular distances to SAM or fluoride (Fig. S9). FLA_MA37 exhibits the most native-like geometry, followed by ancestral_22 and M13. While position 157 shows little conformational variability, position 50 is highly divergent. M1 and M12 fail to form native-like SAM contacts, whereas M13 and M123 restore proper SAM positioning. Position 252 reveals an unexpected mechanism for mutants, where the sidechain of R252 reorients to form a cation–π interaction with Y232, destabilizing the β-sheet spanning residues 232–252 and increasing flexibility in the SAM capping loop (Fig. S9–S12).
MD simulations show that single mutants M1, M2, and M3 fail to rescue activity, highlighting the composite nature of the defects. Among double mutants, M13 uniquely restores activity by improving SAM π-stacking while eliminating the disruptive R252–Y232 interaction, yielding native-like SAM positioning and reduced water permeation. Although M123 shows native-like protein and SAM dynamics, it exhibits high water permeation, indicating that active-site dehydration is the dominant determinant of fluorination activity (Fig. S3–S8 and Table 2 and S8).
Despite sequence divergence, fluorination activity is maintained across diverse Streptomyces and Actinoplanes species. Ancestral_14 and ancestral_15 represent evolutionary branches leading to inactivity, suggesting fluorination can be lost through few substitutions (F156W, Y157F, W50F, L250F, R252). The asymmetry between fluorinases (that can chlorinate) and chlorinases (that cannot fluorinate) likely reflects the greater catalytic challenge of fluorination due to fluoride's high solvation energy. Our water permeation analysis confirms that active fluorinases have highly hydrophobic active sites, a requirement for fluorination but not for chlorination. Ancestral_22 matches FLA_MA37 activity with marginal improvement in selectivity, suggesting fluorination emerged early and has been evolutionarily conserved.
In summary, fluorination activity persists across diverse ancestral fluorinases but is highly sensitive to substitutions that increase active-site hydration or destabilize SAM binding. Our integrated evolutionary, mutational, and molecular dynamics analyses reveal that strict control of active-site dehydration and SAM positioning are key constraints governing biological C–F bond formation. This work expands the accessible fluorinase sequence space and provides a mechanistic framework for future engineering efforts toward biofluorination.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as supporting information (SI) and supporting data. Supplementary information is available. See DOI: https://doi.org/10.1039/d5cc06378g.Acknowledgements
The authors thank Ravi Kumar Verma for insightful discussions and inputs. The authors acknowledge funding support from Agency for Science, Technology and Research (A*STAR), Singapore (C211917003, C233017006, and C211917010), and Institute for Molecular and Cellular Therapeutics, Chinese Institutes for Medical Research, Beijing, China.References
- M. Inoue, Y. Sumii and N. Shibata, ACS Omega, 2020, 5, 10633–10640 CrossRef CAS PubMed.
- T. Okazoe, Proc. Jpn. Acad., Ser. B, 2009, 85, 276–289 CrossRef CAS PubMed.
- J. Wang, M. Sánchez-Roselló, J. L. Aceña, C. del Pozo, A. E. Sorochinsky, S. Fustero, V. A. Soloshonok and H. Liu, Chem. Rev., 2014, 114, 2432–2506 CrossRef CAS PubMed.
- C. Dong, F. Huang, H. Deng, C. Schaffrath, J. B. Spencer, D. O’Hagan and J. H. Naismith, Nature, 2004, 427, 561–565 Search PubMed.
- H. Sun, W. L. Yeo, Y. H. Lim, X. Chew, D. J. Smith, B. Xue, K. P. Chan, R. C. Robinson, E. G. Robins, H. Zhao and E. L. Ang, Angew. Chem., Int. Ed., 2016, 55, 14277–14280 CrossRef CAS PubMed.
- X. Feng, Y. Cao, W. Liu and M. Xian, Front. Bioeng. Biotechnol., 2022, 10 DOI:10.3389/fbioe.2022.881326.
- I. Pardo, D. Bednar, P. Calero, D. C. Volke, J. Damborský and P. I. Nikel, ACS Catal., 2022, 12, 6570–6577 CrossRef CAS PubMed.
- R. K. Verma, W. L. Yeo, E. Tiong, E. L. Ang, Y. H. Lim, F. T. Wong and H. Fan, Chem. Sci., 2025, 16, 10610–10619 RSC.
- C. D. Cadicamo, J. Courtieu, H. Deng, A. Meddour and D. O’Hagan, ChemBioChem, 2004, 5, 685–690 CrossRef CAS PubMed.
- T. Kittilä, P. Calero, F. Fredslund, P. T. Lowe, D. Tezé, M. Nieto-Domínguez, D. O’Hagan, P. I. Nikel and D. H. Welner, Microb. Biotechnol., 2022, 15, 1622–1632 Search PubMed.
- S. Purser, P. R. Moore, S. Swallow and V. Gouverneur, Chem. Soc. Rev., 2008, 37, 320–330 RSC.
- H. M. Senn, D. O’Hagan and W. Thiel, J. Am. Chem. Soc., 2005, 127, 13643–13655 CrossRef CAS PubMed.
- A. S. Eustáquio, F. Pojer, J. P. Noel and B. S. Moore, Nat. Chem. Biol., 2008, 4, 69–74 CrossRef PubMed.
- E. A. Gaucher, S. Govindarajan and O. K. Ganesh, Nature, 2008, 451, 704–707 Search PubMed.
- M. J. Harms and J. W. Thornton, Nat. Rev. Genet., 2013, 14, 559–571 Search PubMed.
- D. L. Trudeau and D. S. Tawfik, Curr. Opin. Biotechnol., 2019, 60, 46–52 CrossRef CAS PubMed.
- Y. Gumulya, J.-M. Baek, S.-J. Wun, R. E. S. Thomson, K. L. Harris, D. J. B. Hunter, J. B. Y. H. Behrendorff, J. Kulig, S. Zheng, X. Wu, B. Wu, J. E. Stok, J. J. De Voss, G. Schenk, U. Jurva, S. Andersson, E. M. Isin, M. Bodén, L. Guddat and E. M. J. Gillam, Nat. Catal., 2018, 1, 878–888 CrossRef CAS.
- B. S. Jones, C. M. Ross, G. Foley, N. Pozhydaieva, J. W. Sharratt, N. Kress, L. S. Seibt, R. E. S. Thomson, Y. Gumulya, M. A. Hayes, E. M. J. Gillam and S. L. Flitsch, Angew. Chem., Int. Ed., 2024, 63, e202314869 CrossRef CAS PubMed.
- L. Ma, Y. Li, L. Meng, H. Deng, Y. Li, Q. Zhang and A. Diao, RSC Adv., 2016, 6, 27047–27051 RSC.
- H. Deng, L. Ma, N. Bandaranayaka, Z. Qin, G. Mann, K. Kyeremeh, Y. Yu, T. Shepherd, J. H. Naismith and D. O’Hagan, ChemBioChem, 2014, 15, 364–368 Search PubMed.
- K. Katoh and D. M. Standley, Mol. Biol. Evol., 2013, 30, 772–780 CrossRef CAS PubMed.
- M. N. Price, P. S. Dehal and A. P. Arkin, PLoS One, 2010, 5, e9490 Search PubMed.
- L.-T. Nguyen, H. A. Schmidt, A. von Haeseler and B. Q. Minh, Mol. Biol. Evol., 2015, 32, 268–274 Search PubMed.
- T. K. M. Nguyen, M. R. Ki, R. G. Son and S. P. Pack, Appl. Microbiol. Biotechnol., 2019, 103, 2205–2216 Search PubMed.
- J. G. Marblestone, S. C. Edavettal, Y. Lim, P. Lim, X. Zuo and T. R. Butt, Protein Sci., 2006, 15, 182–189 Search PubMed.
- M. P. Jacobson, D. L. Pincus, C. S. Rapp, T. J. F. Day, B. Honig, D. E. Shaw and R. A. Friesner, Proteins, 2004, 55, 351–367 CrossRef CAS PubMed.
- K. J. Bowers, D. E. Chow, H. Xu, R. O. Dror, M. P. Eastwood, B. A. Gregersen, J. L. Klepeis, I. Kolossvary, M. A. Moraes, F. D. Sacerdoti, J. K. Salmon, Y. Shan and D. E. Shaw, in SC ’06: Proceedings of the 2006 ACM/IEEE Conference on Supercomputing, 2006, pp. 43–43.
- N. Michaud-Agrawal, E. J. Denning, T. B. Woolf and O. Beckstein, J. Comput. Chem., 2011, 32, 2319–2327 Search PubMed.
Footnote |
| † Equal contributions. |
| This journal is © The Royal Society of Chemistry 2026 |