Mechanistic principles of antimicrobial peptides uncovered by charge density-based machine learning
Hrushikesh
Malshikare
 ab,
U. Deva
Priyakumar
ab,
U. Deva
Priyakumar
 c,
Prathit
Chatterjee
c,
Prathit
Chatterjee
 *c and
Durba
Sengupta
*c and
Durba
Sengupta
 *ab
*ab
aPhysical and Materials Chemistry Division, CSIR-National Chemical Laboratory, Dr Homi Bhabha Road, Pune 411008, India. E-mail: d.sengupta.ncl@csir.res.in
bAcademy of Scientific and Innovative Research (AcSIR), Ghaziabad 201002, India
cCenter for Computational Natural Sciences and Bioinformatics, International Institute of Information Technology, Hyderabad 500032, India. E-mail: prathit.chatterjee@ihub-data.iiit.ac.in
First published on 27th January 2026
Abstract
Antimicrobial peptides (AMPs) are emerging as potent alternatives to conventional antibiotics, yet their diverse nature due to divergent mechanisms of action hinders rational design. Here, we present an electrostatics-stratified computational framework that uncovers key physicochemical principles governing AMP activity. Experimentally validated peptides were grouped by average charge per residue (i.e., the charge/length of the peptide) and analyzed through integrated sequence-, structure-, and chemistry-based descriptors. Distinct molecular signatures emerged across electrostatic regimes: low-charge/length peptides rely on amphipathic organization via structural compactness, whereas the intermediate-charge/length peptides exhibit balanced hydrophobicity and electrostatics. The high-charge peptides couple strong cationic attraction with lipophilicity and tryptophan anchoring to mainly disrupt membranes. Interestingly, hydrophobic moment, which is a measure of the amphipathicity, is found to be important in all three classes of AMPs. This study identifies distinguishing features of AMP sub-groups and suggests design guidelines for developing selective and potent next-generation AMPs.
Antimicrobial resistance (AMR) has become a growing concern and a major global health crisis. Bacterial AMR was reported to contribute to an estimated 1.27 million deaths globally in 2019, with projections indicating that the deaths attributable to AMR could escalate to 10 million by 2050.1,2 This alarming reality has driven urgent exploration of alternative therapeutic approaches, with antimicrobial peptides (AMPs) emerging as one of the most promising candidates due to their broad-spectrum activity, rapid bactericidal actions, and fundamentally distinct mechanisms of action compared to conventional antibiotics.3,4 Despite their immense therapeutic potential, clinical translation of AMPs has been hampered by significant knowledge gaps in understanding their complex and diverse mechanisms of action.5 A well-studied class of AMPs are the cationic AMPs that primarily target bacterial membranes, but they often exhibit multiple modes of membrane association that are less understood.6 Furthermore, many AMPs are multifunctional and bind intracellular targets or modulate immune responses.7 Previous experimental studies have highlighted a few relationships between the physicochemical properties of the peptides and their antimicrobial activity. For instance, it has been shown that the net charge and the number of positively charged residues significantly affect the antimicrobial and hemolytic activities of α-helical AMPs.8 Increasing the charge of magainin 2 from +3 to +5 improved the antibacterial activity, but an increase to +6 or +7 led to increased hemolytic activity and loss of antimicrobial activity.9 Furthermore, chain length has been shown to modulate the structure and stability of antimicrobial peptides.10 Understanding these molecular principles is critical for rational AMP design, as well as for identifying the functional features that distinguish active peptides from inactive analogs.11
The advent of artificial intelligence (AI) and machine learning (ML) has revolutionized AMP research, enabling researchers to decode complex relationships between peptide sequence, structure, and biological activity.12–14 The incorporation of deep learning architectures and generative models has further advanced the design of novel peptide sequences with desired functional profiles, moving beyond the constraints of conventional library screening.15 Recent developments include species-specific predictive models and AI-driven feature selection that highlights key physicochemical and sequence-based descriptors linked to antimicrobial activity.16
While prior computational efforts have predominantly focused on maximizing predictive accuracy, fewer studies have systematically dissected how specific physicochemical parameters contribute to functional differentiation. Addressing this gap, we use a stratified classification framework to identify the molecular signatures underlying AMP function, with a focus on physicochemical, structural, and sequence-derived properties. AMPs as well as non-active peptides (non-AMPs) were stratified into three datasets, based on their charge per residue (i.e. the charge/length ratio). Subsequently, three independent ML binary classifiers were trained on these datasets to distinguish AMPs from non-AMPs. By ranking feature importance within each subgroup, we identified key physicochemical determinants that appear to align with known mechanisms of action. This integrative approach not only enables accurate AMP classification but also helps predict AMP signatures based on experimentally established biophysical modes of action, providing design principles for developing novel peptides with optimized potency and selectivity.
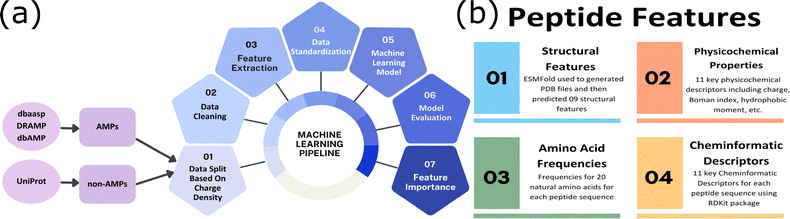
Experimentally validated AMPs were compiled from the dbAMP,17 DRAMP,18 and DBAASP19 databases and curated. Non-active peptide sequences were curated from the UniProt database. The overall ML workflow is illustrated in Fig. 1. The peptides were grouped into low (−0.3 to 0.1), moderate (0.1–0.25), and high (0.25–0.75) charge/length sub-groups based on the ratio of net charge to peptide length (Fig. 2). To ensure a fair comparison, the non-active peptides were explicitly matched to the AMPs based on the charge/length ratio. Since the charge-length distribution in AMPs is uneven (Fig. 2C), these boundaries create regions that are more balanced and sufficiently populated with both AMPs and non-AMPs. Each peptide was represented using 51 features derived from four descriptor sub-groups (Fig. 1b). The structural descriptors such as the secondary structure fraction were calculated from structures predicted using ESMFold.20 Short AMPs often sample dynamic ensembles; the ESMFold-derived structures should be viewed as sequence-encoded structural propensities. Similarly, physicochemical properties, such as hydrophobic moment and Boman index, cheminformatic features and sequence-based pseudo composition were calculated (see the SI). The dataset was first partitioned into three charge-density subgroups, and then an 80–20 train–test split was applied within each subgroup. Multiple ML models such as Random Forest (RF)21 and eXtreme Gradient Boosting XGB22 were implemented using Scikit-learn. All cross-validation was performed exclusively within the training set using stratified 10-fold CV, and the model had no access to the test data during fitting or hyperparameter selection. The features that contribute most to the prediction of AMPs in the three sub-groups were then analyzed by calculating the average SHAP (SHapley Additive exPlanations)23 values for the best-performing (XGBoost) model. Based on an ablation study described in the SI, we report the top five features that retain substantial predictive power of the model. Detailed descriptions of all methods are provided in the SI.
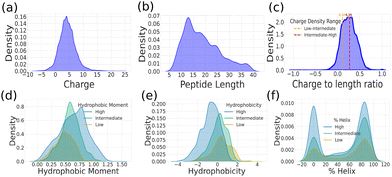
The comprehensive dataset of experimentally validated AMPs compiled in this study revealed a broad distribution of charge and peptide length (Fig. 2a and b) (charge ranges between −5 and +15 and peptide lengths from 5 to 35 residues), in line with their considerable electrostatic and structural diversity. To concomitantly account for these two parameters, we considered charge/length (net charge per residue) as a composite descriptor. From the resulting distribution (Fig. 2c), AMPs were broadly grouped into low (−0.30 to 0.1), intermediate (0.1 to 0.25), and high (0.25 to 0.75) charge/length sub-groups. Across these three sub-groups, several of the descriptors, such as the hydrophobic moment, hydrophobicity, and helical content, are broadly distributed and exhibit overlapping values (Fig. 2d–f). High-charge/length peptides exhibited larger hydrophobic moments but lower overall hydrophobicity, reflecting their increased polarity and electrostatic character. When comparing AMPs with non-AMPs, AMPs displayed a bimodal helicity distribution (Fig. S1), indicating that they often adopt either highly helical or completely unstructured conformations.
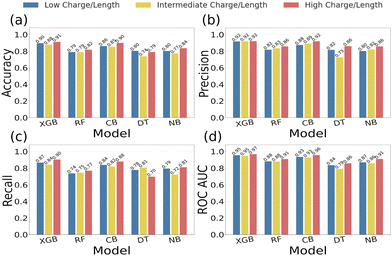
Multiple ML binary classification models were subsequently trained on the three charge/length-based sub-groups (low, intermediate, and high) to evaluate classification performance (see Fig. 3). Among all the tested models, XGBoost demonstrated the best overall performance across the three charge/length sub-groups. For the low and intermediate-charge density subgroups, the model had a training accuracy of 0.91 and a test accuracy of 0.89 in both cases. The high-charge density subgroup showed a training accuracy of 0.92 and a test accuracy of 0.91. A full set of performance metrics for all the evaluated models is provided in the SI and Table S1. To further assess the robustness and generalizability of the model, we performed additional validation, including feature-ablation, evaluation on an independent external dataset, and redundancy reduction using CD-HIT at defined sequence-identity thresholds (see SI). The strong performance of XGBoost reflects its ability to capture non-linear patterns and complex feature interactions within heterogeneous peptide datasets.
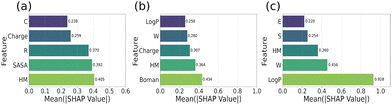
The distinguishing features of AMP sub-groups were then analyzed from the average SHAP values. The top 5 features for AMP prediction within each charge/length subgroup are shown in Fig. 4. The distributions of these features for AMPs and non-AMPs are shown in Fig. S2–S4 in the SI, and the corresponding absolute SHAP value distributions are given in Fig. S5, SI. Within the low-charge/length peptide sub-group, the SHAP index identified the hydrophobic moment, solvent-accessible surface area (SASA), arginine content, charge and cysteine content as key descriptors (Fig. 4a). These findings indicate that low-charge/length peptides that possess a minimal net positive charge adopt compact and balanced amphipathic conformations. To validate the feature-importance indices, we calculated the distribution of these features compared to those of non-active peptides (Fig. S2). Indeed, AMPs exhibit a right-shifted hydrophobic moment distribution and a left-shifted SASA distribution, indicating overall reduced solvent exposure. Interestingly, the arginine (R) frequency was lower in AMPs than in non-AMPs (Fig. S2c), suggesting that they may not target membranes as their main mode of action. Indeed, low-charge/length peptides such as Microcin J25 (MccJ25)24 are often internalized via different transport systems. Together, these features highlight that structural compactness and balanced amphipathic organization distinguish the active low-charge/length AMPs from inactive peptides.
In the intermediate charge/length sub-group, Boman index was identified as an important AMP signature, followed by hydrophobic moment, net charge, tryptophan content and LogP values (Fig. 4b). Comparison of the distribution of these metrics with non-active peptides (Fig. S3) shows that in general AMPs exhibit lower Boman index values, indicating reduced nonspecific binding potential. In addition, comparatively higher hydrophobic moment values of AMPs (Fig. S3c) reflect stronger amphipathic character that favors membrane interactions. Surprisingly, although the distribution of the charge/length was the same between the active and non-active peptides in this class, the overall net charge is highlighted as an important feature. In addition, the tryptophan content (W) was higher in a sub-group of AMPs, consistent with the partitioning of Trp side chains at the membrane–water interface. In conjunction, the lower Boman index together with these properties suggests that AMPs in this subgroup are less likely to interact non-specifically with protein partners. For example, indolicidin (charge/length 0.23) exhibits both enhanced membrane translocation and intracellular DNA binding.25 In contrast, magainin 2 (charge/length 0.17) mainly forms membrane pores and acts at the membrane interfaces.26 Together, these findings suggest that intermediate-charge/length AMPs may function through a balanced interplay of amphipathic alignment, moderate electrostatics, and structural adaptability, enabling both membrane-disruptive and intracellular modes of action.
Within the high-charge/length subgroup, SHAP values identified LogP as a critical factor (Fig. 4c). Furthermore, tryptophan content (W), hydrophobic moment, and the frequencies of S and E were associated with antimicrobial activity (Fig. 4c). AMPs in this subgroup, despite their high charge, combine global lipophilicity with residue-specific anchoring, enabling membrane interactions. A comparison of these values with those of inactive peptides (Fig. S4) reveals that AMPs display a right-shifted LogP distribution compared to non-AMPs, reflecting enhanced lipophilicity and a greater tendency to partition into the membrane. Similar to the intermediate sub-group, tryptophan (W) frequency distributions show a marked enrichment in AMPs. The hydrophobic moment further contributes by reinforcing amphipathic organization, promoting stable interfacial binding and orientation. In contrast, the presence of polar residues such as serine and glutamic acid is a feature of inactive peptides. These physicochemical trends align with previous findings linking lipophilicity (LogP) to overall antimicrobial potency.27 For instance, protegrin28 (charge/length: 0.33) and tritrpticin (0.38), a Trp-rich peptide,29 induce membrane leakage through the formation of pores. Collectively, these observations highlight that LogP, Trp content, and hydrophobic moment constitute a complementary set of descriptors integrating global lipophilicity, residue-specific anchoring, and amphipathic organization.
In conclusion, this study establishes a charge/length-stratified ML framework that suggests distinct mechanistic strategies underlying AMP activity. As the charge/length increases, the dominant mechanism of action appears to transition from intracellular targeting to membrane disruption, with intermediate-charge/length peptides exhibiting features of both. Interestingly, hydrophobic moment, a measure of amphipathicity, is found to be important in all three subgroups of AMPs. Low-charge/length peptides may favor compact, amphipathically balanced structures that facilitate intracellular interactions, whereas high-charge/length peptides seem to achieve potent membrane permeabilization through electrostatic attraction and residue-specific anchoring. These findings indicate that the AMP function is likely governed not by a single descriptor but by combinations of features. In the low-charge/length subgroup, the balance of properties appears to be important, whereas in the high-charge subgroup, membrane partitioning seems particularly critical. Taken together, the results fit well with established biophysical principles, and the data-supported patterns we identify provide mechanistic insights that can guide rational AMP design and help tune potency and selectivity across different charge/length subgroups.
D. S. and U. D. P.: conceptualization; P. C.: supervision; H. M.: data curation and implementation.
Conflicts of interest
There are no conflicts to declare.Data availability
All scripts and analysis notebooks used in this study are publicly available at https://github.com/hm619/AMP-ML-Signatures and https://github.com/Sengupta-NCL/AMP-signature.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5cc06374d.Acknowledgements
H. M. acknowledges support from a CSIR-Senior Research Fellowship for his doctoral studies. D. S. gratefully acknowledges the support of the Bioinformatics Center from DBT (BT/PR40123/BTIS/137/47/2022), India, hosted at CSIR-NCL, India. P. C. and U. D. P. acknowledge IHub-Data, International Institute of Information Technology, Hyderabad, India. We thank members of our research groups for discussions.References
- C. J. Murray, K. S. Ikuta, F. Sharara, L. Swetschinski, G. R. Aguilar, A. Gray, C. Han, C. Bisignano, P. Rao and E. Wool, et al. , Lancet, 2022, 399, 629–655 Search PubMed.
- W. H. Organization, Global antimicrobial resistance and use surveillance system (GLASS) report 2022, World Health Organization, 2022.
- M. Mahlapuu, C. Björn and J. Ekblom, Crit. Rev. Biotechnol., 2020, 40, 978–992 CrossRef CAS PubMed.
- L.-j Zhang and R. L. Gallo, Curr. Biol., 2016, 26, R14–R19 CrossRef CAS PubMed.
- K. A. Brogden, Nat. Rev. Microbiol., 2005, 3, 238–250 Search PubMed.
- L. T. Nguyen, E. F. Haney and H. J. Vogel, Trends Biotechnol., 2011, 29, 464–472 CrossRef CAS PubMed.
- N. Mookherjee, M. A. Anderson, H. P. Haagsman and D. J. Davidson, Nat. Rev. Drug Discovery, 2020, 19, 311–332 CrossRef CAS PubMed.
- L. M. Yin, M. A. Edwards, J. Li, C. M. Yip and C. M. Deber, J. Biol. Chem., 2012, 287, 7738–7745 CrossRef CAS PubMed.
- M. Dathe, H. Nikolenko, J. Meyer, M. Beyermann and M. Bienert, FEBS Lett., 2001, 501, 146–150 CrossRef CAS PubMed.
- N. Phambu, B. Almarwani, A. M. Garcia, N. S. Hamza, A. Muhsen, J. E. Baidoo and A. Sunda-Meya, Biophys. Chem., 2017, 227, 8–13 Search PubMed.
- C. D. Fjell, J. A. Hiss, R. E. Hancock and G. Schneider, Nat. Rev. Drug Discovery, 2012, 11, 37–51 Search PubMed.
- H. Lv, Y. Zhang, J.-S. Wang, S.-S. Yuan, Z.-J. Sun, F.-Y. Dao, Z.-X. Guan, H. Lin and K.-J. Deng, Briefings Bioinf., 2022, 23, bbab486 Search PubMed.
- G. Wang, X. Li and Z. Wang, Nucleic Acids Res., 2016, 44, D1087–D1093 Search PubMed.
- A. Cesaro, M. Bagheri, M. Torres, F. Wan and C. De La Fuente-Nunez, Expert Opin. Drug Discovery, 2023, 18, 1245–1257 Search PubMed.
- M. Salem, A. Keshavarzi Arshadi and J. S. Yuan, BMC Bioinf., 2022, 23, 389 Search PubMed.
- G. He, Q. He, J. Cheng, R. Yu, J. Shuai and Y. Cao, Int. J. Mol. Sci., 2024, 25, 7237 CrossRef CAS PubMed.
- J.-H. Jhong, L. Yao, Y. Pang, Z. Li, C.-R. Chung, R. Wang, S. Li, W. Li, M. Luo and R. Ma, et al. , Nucleic Acids Res., 2022, 50, D460–D470 Search PubMed.
- G. Shi, X. Kang, F. Dong, Y. Liu, N. Zhu, Y. Hu, H. Xu, X. Lao and H. Zheng, Nucleic Acids Res., 2022, 50, D488–D496 CrossRef CAS PubMed.
- M. Pirtskhalava, A. A. Amstrong, M. Grigolava, M. Chubinidze, E. Alimbarashvili, B. Vishnepolsky, A. Gabrielian, A. Rosenthal, D. E. Hurt and M. Tartakovsky, Nucleic Acids Res., 2021, 49, D288–D297 CrossRef CAS PubMed.
- Z. Lin, H. Akin, R. Rao, B. Hie, Z. Zhu, W. Lu, N. Smetanin, R. Verkuil, O. Kabeli and Y. Shmueli, et al. , Science, 2023, 379, 1123–1130 Search PubMed.
- L. Breiman, Mach. Learn., 2001, 45, 5–32 Search PubMed.
- T. Chen and C. Guestrin, Proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining, 2016, pp. 785-794.
- E. Winter, Handbook of game theory with economic applications, 2002, 3, pp. 2025–2054 Search PubMed.
- A. Bellomio, P. A. Vincent, B. F. de Arcuri, R. N. Farias and R. D. Morero, J. Bacteriol., 2007, 189, 4180–4186 Search PubMed.
- H. Ngo Van, H. Luong Xuan, H. Le Viet, H. B. T. Phuong, Y. Do Hai, N. Q. Thang, T. Truong Thanh, T. V. Yen, T. N. Minh and L. N. Van, et al. , PLoS One, 2025, 20, e0331796 CrossRef CAS PubMed.
- M. V. Volovik and O. V. Batishchev, Biomolecules, 2024, 14, 1118 CrossRef CAS PubMed.
- M. Van der Walt, D. S. Moller, R. J. Van Wyk, P. M. Ferguson, C. K. Hind, M. Clifford, P. Do Carmo Silva, J. M. Sutton, A. J. Mason and M. J. Bester, et al. , ACS Omega, 2024, 9, 26030–26049 Search PubMed.
- D. Bolintineanu, E. Hazrati, H. T. Davis, R. I. Lehrer and Y. N. Kaznessis, Peptides, 2010, 31, 1–8 CrossRef CAS PubMed.
- D. I. Chan, E. J. Prenner and H. J. Vogel, Biochim. Biophys. Acta, Biomembr., 2006, 1758, 1184–1202 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |