Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Protein nanoarchitectonics at liquid-related interfaces: physicochemical communications with biological processes
Katsuhiko
Ariga
 *ab,
Jingwen
Song
*ab,
Jingwen
Song
 *cd and
Kohsaku
Kawakami
*cd and
Kohsaku
Kawakami
 de
de
aResearch Center for Materials Nanoarchitectonics, National Institute for Materials Science (NIMS), 1-1 Namiki, Tsukuba 305-0044, Japan. E-mail: ARIGA.Katsuhiko@nims.go.jp
bGraduate School of Frontier Sciences, The University of Tokyo, 5-1-5 Kashiwa-no-ha, Kashiwa 277-8561, Japan
cKey Laboratory of Multifunctional Nanomaterials and Smart Systems, Suzhou Institute of Nano-tech and Nano-bionics (SINANO), Chinese Academy of Sciences (CAS), Suzhou 215123, P. R. China
dResearch Center for Macromolecules and Biomaterials, National Institute for Materials Science (NIMS), 1-1 Namiki, Tsukuba 305-0044, Ibaraki, Japan
eGraduate School of Science and Technology, University of Tsukuba, 1-1-1 Tennodai, Tsukuba 305-8577, Ibaraki, Japan
First published on 19th January 2026
Abstract
Building functional material structures that resemble living systems from atoms and molecules is a highly challenging process. However, a sophisticated yet expeditious approach can be formulated by utilizing biological materials characterized by meticulously regulated nanostructures that function as fundamental units. Proteins are regarded as the most promising candidates. The nanoarchitectonics of proteins in interfacial environments involving liquids such as water is a highly attractive and feasible approach. In this feature review article, we introduce systems that can be regarded as protein nanoarchitectonics at each of the following interface types: gas–liquid, solid–liquid and liquid–liquid (oil–water and water–water). The air–water interface is an ideal medium for organising water-soluble proteins into ultrathin films and also serves as a simple model system. Research into protein nanoarchitectonics at solid–liquid interfaces appears to be evolving to a higher level, with a focus on solving problems and revealing new functions of proteins. The oil–water interface also presents opportunities for the organization of biological materials, and various fundamental characterization studies of its interfaces have been conducted. The water–water interface is considered the optimal environment in which to apply knowledge gained from model systems such as the air–liquid and solid–liquid interfaces to more realistic biological systems. Furthermore, physicochemical communication between interfaces has been shown to regulate complex processes, including cell differentiation and coupling biochemical reactions with device functions. Pursuing such methodologies may facilitate the creation of functional systems that exhibit characteristics analogous to those of living organisms.
1. Introduction
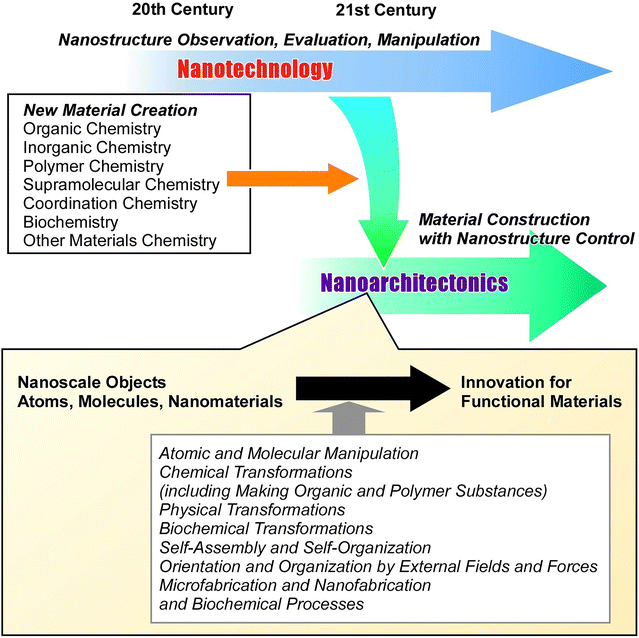
In recent years, humanity has made significant progress in developing cyberspace and the technology that supports it, including information technology, device technology and artificial intelligence. However, real-world materials are truly essential for supporting our lives. Functional materials are being developed to address various issues, including those related to energy,1 the environment,2 and healthcare.3 As new problems emerge, it seems that the development of necessary material functions is limitless. Much research effort appears to be dispersed and expanding. An ultimate goal is not always apparent. This is because material development has tended to be carried out individually, rather than towards creating a unique, ultimate methodology. In this introduction, we would like to present the background and propose “protein nanoarchitectonics at liquid-related interfaces” as a potential solution.Making useful materials needs a few things. We can roughly divide these into three steps. The first step is to create materials with the desired properties. The second step is to control their microstructures and nanostructures. The third step is to build these materials into functional materials. Materials science has developed a lot along this line. In the past, people collected materials from nature, but in the 20th century, we learned how to make new materials using chemistry. While there are different types of chemistry, such as organic chemistry,4 inorganic chemistry,5 polymer chemistry,6 supramolecular chemistry,7 coordination chemistry,8 biochemistry,9 and other materials chemistry,10 these fields have developed together as academic subjects. Along with the development of functional materials, it has become clear that controlling nanostructures is crucial for creating materials with better functions and performances.11 Many attempts to investigate nanostructures and their physical properties have made under the umbrella concept of nanotechnology. We have made great strides in observing12 and controlling13 structures at the atomic and molecular levels, as well as in measuring physical properties and characteristics at the nano and micro levels.14 We have clear strategies for making unit materials and controlling nanostructures.
The next step is the construction of highly functional materials through putting together nanostructured materials. This project has attracted a lot of research interest. For example, it involves the formation of various self-assembled structures,15 the creation of ordered porous materials such as metal–organic frameworks (MOFs),16 covalent organic frameworks (COFs),17 and nanoporous materials,18 and the control of crystalline structures.19 Interface chemistry has made a big impact on this field, with self-assembly monolayers (SAMs),20 the Langmuir–Blodgett (LB) method,21 and layer-by-layer (LbL) assembly22 being key techniques for thin film assembly. However, these approaches have been developed separately and on their own. The idea of nanotechnology has not yet resulted in advances in science and technology at the nano level. To build working structures, we need to use a combined idea, a post-nanotechnology approach.23
Nanoarchitectonics is the answer (Fig. 1). It is a comprehensive combined concept for building useful structures from very small parts, such as atoms, molecules and nanomaterials.24 It suggests making materials by combining atomic and molecular manipulation, chemical transformations (including making organic and polymer substances), physical transformations, biochemical transformations, self-assembly and self-organization, orientation and organization caused by external fields and forces, microfabrication and nanofabrication, and biochemical processes.25 Richard Feynman came up with the idea of nanotechnology in the middle of the 20th century26 and Masakazu Aono came up with nanoarchitectonics at the start of the 21st century.27 Matter is basically made up of atoms and molecules, so this method could be used for all materials. If the super unification theory that unifies all forces is the theory of everything that physics aims for,28 then nanoarchitectonics could become a method for everything in materials science.29 Of course, this may be considered an exaggeration. However, just as physics strives toward achieving its ultimate goal, perhaps material chemistry could also have some dreamlike objective. Material chemistry tends to advance through specialized research. It is meaningful to consider a research trajectory that establishes concepts and moves toward a unified goal. It is with this hope that we have made this proposal here. In fact, the term nanoarchitectonics is spreading across numerous fields. The concept of assembling functional materials from atoms and molecules to create functional materials systems could become a functional concept in many disciplines. Recent examples clearly show that numerous studies and papers spanning fields from basic sciences dealing with molecules and atoms, to energy, environment, and medicine are being published under the name of nanoarchitectonics.25
Which material systems demonstrate higher functionality? Looking broadly at existing functional systems, the answer lies not in those created by humanity, but in naturally occurring functional systems. There we can observe rational, efficient functional systems that appear to far surpass materials we have artificially constructed. In a living system, the way it works is organized so that it is efficient and selective, and the flow of energy, electrons and information is controlled. All of this is achieved at room temperature and pressure. Also, several functions work together. This performance is achieved by organizing the best possible units into the best possible organization.30 This is the direction that the creation of functional materials in nanoarchitectonics should be aimed at.31 Life has developed in the way it has done because of the selection and survival of the best structures over very long periods of time, spanning billions of years. Nanoarchitectonics must achieve this within a realistic time span of several decades. Of course, this approach is not confined to the narrow goal of mimicking biological functions. Rather, it seeks to harness the essence of forming sophisticated functional units—the kind found in biological systems—and apply it to the architecture of functional material structures.
The objective is indeed one of considerable difficulty. The construction of functional material structures comparable to living systems from atoms and molecules will be a highly challenging process. It is imperative that a sophisticated yet expeditious approach be formulated, namely the utilization of biological materials characterized by meticulously regulated nanostructures that function as the fundamental units. Proteins are regarded as the most promising biological materials. Proteins are composed of units with intricately controlled internal structures that perform highly efficient and selective functions.32 This strategy involves the utilization of biological evolution to produce the desired materials, followed by the application of nanoarchitectonics to arrange them into specific structures. However, it is important to note that the mere organization of proteins is not a sufficient strategy in itself. It is imperative to establish an environment that restricts their movement and dispersion. This is the interfacial environment. It is notable that a significant proportion of proteins are water-soluble and operate within aqueous environments. Consequently, the nanoarchitectonics of proteins in interfacial environments involving liquids, such as water, is a highly attractive and feasible approach. The nanoarchitectonics of proteins at liquid interfaces may hold the key to unlocking the methodology for developing ultimate functional materials.
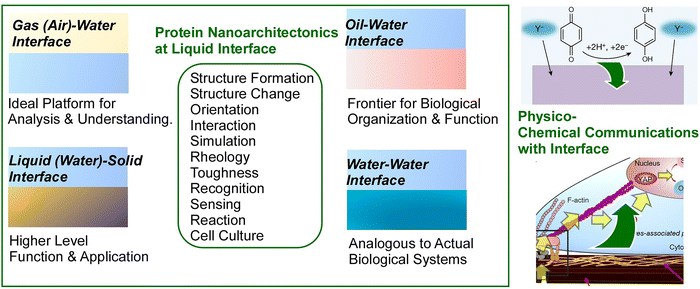
From this standpoint, the present feature article discusses the subject of “protein nanoarchitectonics at liquid-related interfaces”. In the ensuing sections, we introduce systems that can be regarded as protein nanoarchitectonics at each interface type: gas–liquid interfaces, solid–liquid interfaces, and liquid–liquid interfaces (oil–water interfaces and water–water interfaces). Moreover, in the section entitled “Physico-chemical communications with biological processes at liquid interfaces”, the subject of physicochemical communications is discussed, with the development of interfacial phenomena into more sophisticated functions being a particular focus. The overarching objective of this structure is twofold: firstly, to identify the characteristics and commonalities of protein assembly at various liquid interfaces and, secondly, to demonstrate the potential for such organisations to lead to the emergence of more alternating functions. Consequently, the examples provided herein are not intended to be exhaustive; they have been selected from recent research examples to elucidate the aforementioned discussion. In the final section, we offer a concise discussion of the necessary steps to advance these efforts towards the creation of ultimate functional materials. This feature article does not aim to simply list recent examples; rather, it seeks to identify trends in protein organization and function at liquid-related interfaces. The methods employed in the creation of interfaces are varied, with many undergoing constant development and refinement. The comprehension of interface-specific trends in organization and function provides a foundation for the design of interfaces that can achieve desired outcomes. In the final section, a summary of the characteristics of protein nanoarchitectonics at different interfaces is provided, derived from the numerous examples discussed. In addition, the principal objective of this feature article is not to provide an exhaustive overview of all instances of “protein nanoarchitectonics at liquid-related interfaces”, but rather to illustrate that these subjects constitute a prospective approach to establishing methodologies for the development of ultimate materials.
2. Gas–liquid interfaces
The air–water interface is a significant medium for the formation of ultrathin films of numerous materials. This is actually a powerful approach to organizing protein molecules into two-dimensional systems as an nanoarchitectonics approach from protein molecules into useful thin film nanoarchitectures. A typical example is a Langmuir monolayer composed of lipid molecules, which serves as a model for biological membranes and provides an ideal platform for studying interactions with various biomolecules dissolved in the aqueous phase.33 It has been demonstrated that model substances, including peptides, amino acids, nucleic acids and glycans, are capable of assembling into thin films at the air–water interface.34 Furthermore, proteins can be used as a target for the organization of functional materials at the air–water interface.35 However, the elevated surface tension at the air–water interface has the potential to induce protein denaturation. A number of pioneering examples for the prevention of this phenomenon have long been devised. As one pioneering example, illustrated in Fig. 2, the Langmuir–Blodgett (LB) method for enzymes utilizes a complex comprising glucose oxidase and lipid molecules.36 The complex of glucose oxidase and lipid molecules can be dissolved in organic solvents and spread on the water surface, similar to common substances such as lipids, to form a Langmuir monolayer. The presence of lipid molecules has been demonstrated to circumvent the effects of the high surface tension of water. It has been demonstrated that this method does not result in interfacial denaturation of glucose oxidase. This was well confirmed by the high activity of glucose oxidase in sensor organization. The development of a glucose sensor is facilitated by the layering of a monolayer on an electrode. This example is not the only documented instance of this phenomenon; it has been reported that the air–water interface is a promising site for the organization of biomolecules, such as proteins, into ultrathin films.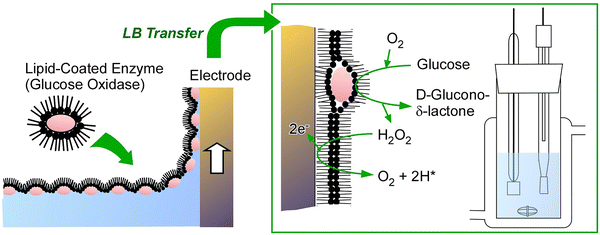 | ||
| Fig. 2 Langmuir–Blodgett (LB) method for a complex comprising glucose oxidase and lipid molecules (left) and application of the prepared LB film as a glucose sensor (right). | ||
The process of forming proteins into ultrathin films has been shown to increase the potential for their application as functional materials. In addition to fundamental research, there has been a recent emergence of application-oriented research focusing on the organization of proteins at the air–water interface. The following part shows some recent examples. For instance, Wang, Shao, and their coworkers have developed a method for efficiently fabricating large-area, robust, and free-standing nanofilms at the air–water interface using silk fibroin and lysozyme as building blocks (Fig. 3).37 This is the interfacial cooperative assembly of silk fibroin and lysozyme. The strong intermolecular interactions between these two proteins, in conjunction with the abundance of intra- and intermolecular β-sheet motifs, facilitate the formation of robust, ultrathin nanofilms. When a solution of lysozyme and tris(2-carboxyethyl) phosphine (TCEP) was added to an aqueous silk fibroin solution at room temperature, a floating silk fibroin/lysozyme nanofilm spontaneously formed at the air–water interface after approximately one hour of incubation. This nanofilm, with a thickness of merely 50 nm, demonstrated noteworthy mechanical robustness and self-supporting properties in ambient air. Moreover, this methodology can be applied to construct ultrasensitive photonic structures and organic–inorganic hybrid multilayer films with excellent optical and mechanical properties. Furthermore, the integration of functionalization into the protein interfacial assembly process offers the potential to add new functions to these two-dimensional (2D) protein systems by incorporating functional dopants, such as bioactive molecules, conductive nanomaterials, and photothermal and photochemical moieties, within versatile protein formats. Such versatile all-protein-based nanofilms can be fabricated, offering a combination of ultrathinness, robustness, large-area uniformity, self-supporting properties, high flexibility, versatile transferability, self-adhesion, rapid and tuneable responses, and multifunctionality. These properties have the potential to create new possibilities for wearable devices, medical implants, and brain–machine interfaces. The combination of bottom-up protein interfacial construction, top-down fabrication techniques, and programmable stimuli-responsive behaviours may pave the way for the development of a variety of adaptive and multifunctional optical, electronic, and mechanical devices.
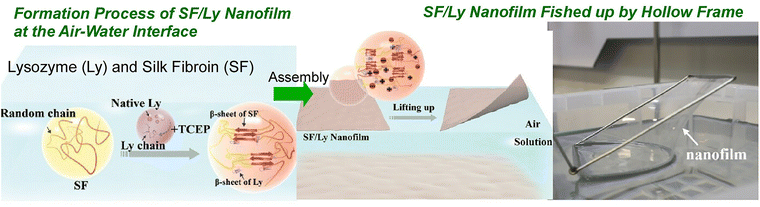 | ||
| Fig. 3 Fabrication of large-area, robust, and free-standing protein nanofilms at the air–water interface using silk fibroin and lysozyme as building blocks. TCEP: tris(2-carboxyethyl) phosphine (TCEP). Reprinted with permission from ref. 37 Copyright 2024 Wiley-VCH. | ||
Foams containing gas bubbles are also media that constitute air–water interfaces. It has been demonstrated that plant proteins, when incorporated as components, function as stabilizers of multiphase food materials, including edible foams and emulsions. It is imperative to investigate the mechanisms that govern the lifetime of foams stabilized by complex surface-active species, such as plant-derived proteins. Experiments conducted at the length scale of the thin liquid film separating two adjacent bubbles have the potential to facilitate a more profound comprehension of the mechanisms by which plant proteins stabilize foams, which is a significant material for food applications. Chatzigiannakis et al. employed the dynamic thin film balance method to investigate the equilibrium properties and dynamic behaviour of thin liquid films of foams stabilized with proteins from two major plant protein sources, namely pea and rapeseed (Fig. 4).38 The objective of this study is to examine protein-stabilized interfaces and to enhance comprehension of the foam stabilization mechanisms prevalent among diverse surface-active species. The half-life of such foams is proportional to the film thickness, which is closely related to the steric and electrostatic interactions occurring between the respective films in equilibrium. Local effects have been demonstrated to play a pivotal role in the process of film rupture. It has been demonstrated that both average film properties and local interfacial structures play a role in the process of aggregation. This finding provides a method by which to engineer plant protein-based membranes with complex and non-uniform thicknesses to be rupture-resistant, an essential element for stable foam formation. It is possible to precisely engineer foam stability by controlling interface and membrane structure, by accessing all relevant length scales (i.e. molecular, interfacial, membrane, cellular and foam).
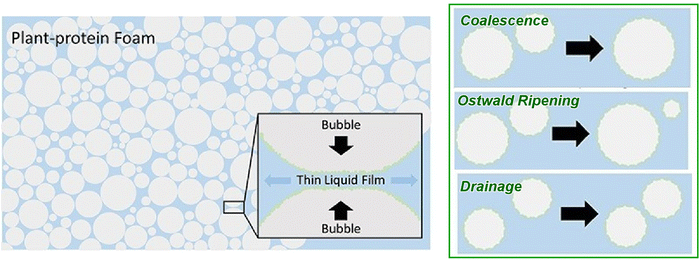 | ||
| Fig. 4 Dynamic behaviour of thin liquid films of foams stabilized with proteins from two major plant protein sources, namely pea and rapeseed: coalescence, Ostwald Ripening, drainage. Reproduced under terms of the CC-BY license from ref. 38 Copyright 2025 Elsevier. | ||
In other studies, Hinderink et al. blended pea protein isolate, which was employed as a model legume protein, with dairy proteins (whey protein isolate or sodium caseinate).39 Utilizing an automated drop tensiometer, dilational rheometry was conducted at the air–water and oil–water interfaces to characterize the behaviour and structure of the interfacial films. Guo and coworkers utilized a force tensiometer to monitor the decline in surface tension and the interplay between surfactants and proteins at the interface, with the objective of correlating the protein surface behaviour with interfacial stability.40 These investigations provide valuable insights into how proteins organize and stabilize interfacial films, which is critical for understanding phenomena like foam or emulsion stability. Similarly, the interfacial properties of therapeutic proteins play a crucial role in determining their stability in formulation.
Due to their amphiphilic nature, monoclonal antibodies form viscoelastic, gel-like layers at the air–water interface. It has been demonstrated that this property can result in the undesirable aggregation of proteins in therapeutic formulations. Qian, Wagner, and coworkers have demonstrated that the interfacial viscoelasticity and surface pressure of monoclonal antibodies at the air–water interface are correlated with their long-term stability.41 A potential molecular mechanism was proposed to explain the relationship between interfacial elastic modulus and long-term stability. In addition, Wagner and coworkers utilized a multifaceted approach encompassing surface tension measurements, interfacial rheology, capillary viscosities, and neutron reflectivity measurements to investigate the surface activity of a model surfactant, Polysorbate 20 (PS20), and monoclonal antibody systems (Fig. 5).42 In the domain of pharmaceuticals, surfactants such as PS20 are incorporated into protein formulations with the objective of enhancing protein stability and reducing monoclonal antibody adsorption at interfaces. The need to quantify this process was paramount. The adsorption of monoclonal antibodies into surfactant-free solutions resulted in the formation of a monolayer exhibiting significant viscoelastic properties. Conversely, the adsorption of PS20 leads to a reduction in interfacial viscosity to an undetectable level, thereby safeguarding the air–water interface from monoclonal antibody adsorption. This finding is further substantiated by the quantitative analysis, which demonstrates that PS20, significantly exceeding its critical micelle concentration (CMC), can impede the adsorption and subsequent formation of an elastic film by monoclonal antibodies at the air–water interface. Furthermore, it was observed that the addition of PS20 after the formation of a monoclonal antibody film at the interface reduced this viscoelasticity. The findings of this study may offer valuable insights for the development of formulations that exhibit enhanced stability and are suitable for extended storage periods.
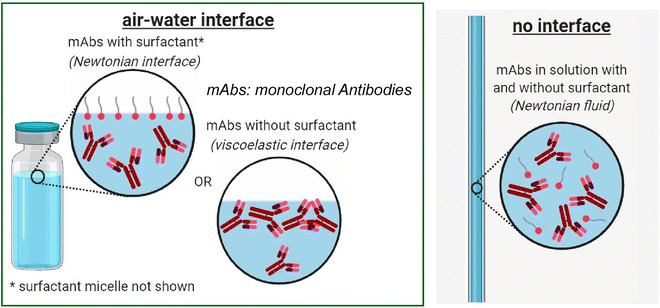 | ||
| Fig. 5 Behaviours of monoclonal antibodies with and without a model surfactant, Polysorbate 20 (PS20) at the air–water interface, where the adsorption of monoclonal antibodies into surfactant-free solutions resulted in the formation of a monolayer exhibiting significant viscoelastic properties, but the adsorption of PS20 leads to a reduction in interfacial viscosity. Reprinted with permission from ref. 42 Copyright 2020 American Chemical Society. | ||
In a similar vein, Furst and coworkers evaluated the interfacial shear rheology of a model antibody, anti-streptavidin immunoglobulin-1 (AS-IgG1), at the air–water interface.43 Maintaining formulation stability during manufacturing, storage, and clinical administration is pivotal to the development of protein therapeutics, such as monoclonal antibodies. Utilizing passive microrheology techniques, Dhar and coworkers investigated the correlation between alterations in surface adsorption of a monoclonal antibody standard and changes in surface rheology, as well as in interface-induced protein particle formation occurring beyond the visible range.44
The organization of proteins at the air–water interface also has applications in analytical techniques. Cryo-electron microscopy is a powerful tool for elucidating the high-resolution structure of biomacromolecules in solution. However, the preferential orientation, dissociation, or denaturation of biomacromolecules induced by the air–water interface during cryo-vitrification remains a limiting factor for many specimens. Qiao, Sun and coworkers developed a cryo-electron microscopy support membrane based on 2D crystals of the hydrophobin HFBI in order to address this issue (Fig. 6).45 A novel type of cryo-electron microscopy grid was developed by coating the amphiphilic hydrophobin HFBI onto a holey ANTA foil grid. The hydrophilic surface of the HFBI membrane has been shown to adsorb protein particles through electrostatic interactions, thus allowing the formation of ice layers of sufficient thinness for the collection of reliable data. In addition, it has been demonstrated that the particle orientation distribution can be modulated through the adjustment of the pH level. The optimal pH is closely approximated to the isoelectric point of the target complex. Utilizing this support membrane, the cryo-EM structures of catalase (2.29 Å) and influenza hemagglutinin trimer (2.56 Å) were determined, which exhibited strong preferable orientations when conventional cryo-vitrification protocols were employed. It is anticipated that this approach will have extensive applications in future cryo-EM studies. From a general perspective, the following idea is also interesting for interpretation of this research. This research engages in nanoarchitectonics across two size domains. That is, research into material-level nanoarchitectonics—organizing proteins into two-dimensional films—serves as a means to analyse the molecular-level nanoarchitecture of proteins.
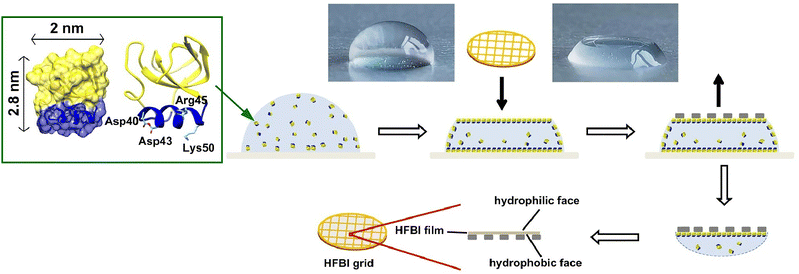 | ||
| Fig. 6 Use of a novel type of cryo-electron microscopy grid developed by coating the amphiphilic hydrophobin HFBI onto a holey foil grid for adsorption of protein particles through electrostatic interactions, thus allowing the formation of ice layers of sufficient thinness for the collection of reliable data. Reproduced under terms of the CC-BY license from ref. 45, 2021 Springer-Nature. | ||
The air–water interface has been demonstrated to offer an ideal platform for the assembly of water-soluble proteins into ultrathin films. Basic research conducted in the 20th century, the aim of which was to devise methods to prevent protein denaturation, has now been developed into a variety of practical applications. Whilst it is not feasible to provide coverage of all of the aforementioned examples, this section presents a wide range of potential applications. These include the creation of robust nanofilms for the development of various adaptive and multifunctional optical, electronic, and mechanical devices; the stabilization of multiphase food materials such as edible vegetable protein foams and emulsions; the interfacial properties for aqueous formulations of dairy proteins and protein drugs; the evaluation of various properties to prevent undesired aggregation of proteins in therapeutic formulations such as monoclonal antibodies; and the development of interfacial processes to support the elucidation of high-resolution structures of biopolymers using cryo-electron microscopy. The applications of this technology are diverse, encompassing devices, food, pharmaceuticals, and analytical technologies. This finding underscores the notion that the simple model field of the air–water interface serves as an exceptional platform for the evaluation of protein organization, while also meeting a multitude of requirements. Demand is known to be diverse, and further developments are anticipated.
3. Solid–liquid interfaces
The immobilization of proteins from the solution phase onto solid surfaces is of significant importance in a variety of applications. The process of assembling nanofilms on solid surfaces is a subject of particular intrigue. Layer-by-layer (LbL) assembly is a simple and versatile method for fabricating thin films on solid substrates (Fig. 7A).46 The most prevalent LbL technique is grounded in electrostatic interactions. In summary, when a solid substrate is immersed in a solution of a substance, such as a polyelectrolyte, which possesses an opposite charge to that of the substrate, the substance in the solution adsorbs as a nanofilm through electrostatic interactions. Overadsorption, a process which has been observed to reverse the charge on the adsorbed surface, is another phenomenon of relevance to this method. When the substrate with this thin film is immersed in a solution of a substance with the opposite charge, the substance adsorbs as a nanofilm, and the surface charge is reversed. Consequently, by repeating this process, it is possible to construct a multitude of thin films. The layering sequence can also be modified freely, as long as the established charge rules are followed. The process is uncomplicated, and the stacking process can be performed expeditiously with just a beaker and tweezers. It is also possible to utilize the spin coating and spraying methods.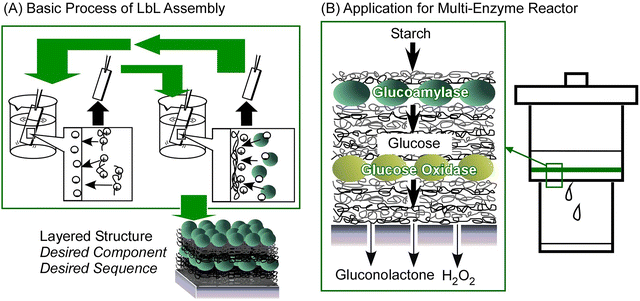 | ||
| Fig. 7 (A) The basic process of layer-by-layer (LbL) assembly and (B) application of the prepared LbL films in a multi-enzyme reactor. | ||
The aforementioned mechanism is contingent on the surface charge, a property that is intrinsic to a multitude of materials, and its applicability is extensive, encompassing polymers, dyes and a variety of nanomaterials. In particular, the majority of biomolecules, such as proteins, possess surface charges, rendering them suitable candidates for the LbL method. Proteins have been shown to be capable of being layered with a high degree of universality,47 a property that is also exhibited by DNA,48 charged polysaccharides,49 and biomembrane models.50 Interactions employed for layering are not confined to electrostatic interactions; hydrogen bonding,51 coordination interactions,52 covalent bonding,53 molecular inclusion,54 charge-transfer complex formation,55 and stereocomplex formation56 are also utilized. In particular, the specific recognition between sugars and lectins,57 and the specific interaction between biotin and avidin,58 can be utilized for LbL layering. It is evident that the LbL method is a highly efficacious technique for the assembly of biomolecules as thin films on solid substrates. It has also been demonstrated that nanoreactors can be fabricated by means of a rational layering process involving multiple enzymes (Fig. 7B).59 The LbL method was proposed and proven useful in the late 20th century, and it continues to be actively researched today as a method for assembling proteins on solid substrates. In this context, it is essential to acknowledge the LbL method as a means for the assembly of proteins as thin films on solid substrates. The progress of this method has been comprehensively documented in numerous reviews.60 In this section, the focus will be on the presentation of recent topics, with the assistance of additional research examples.
In the field of protein organization at solid–liquid interfaces, attention is focused not only on the simple immobilization of proteins but also on the accompanying changes in protein secondary structure. In particular, alterations in secondary structure have been associated with functions such as the regulation of surface adhesion properties. For instance, proteins are utilized in the synthesis of robust bioadhesive materials. In many of these cases, β-sheet stacking has been recognized as a key feature closely related to protein interfacial adhesion. Ren, Yang, and coworkers have challenged this commonly held belief and proposed a model of protein interfacial adhesion mediated by α-helices (Fig. 8).61 This study employed bovine serum albumin (BSA), a prevalent α-helical protein in nature, as a model protein, which contains approximately 60% helical structure. The reduction of disulfide bonds in BSA has been shown to result in the formation of random coil structures from unfolded BSA, with the concomitant dragging of the α-helices and their subsequent assembly at the solid–liquid interface. Following the unlocking of the high-energy α-helices, the interfacial adhesion of unfolded protein chains is driven by interactions between the abundant functional groups exposed from the α-helices and the interface, as well as by intermolecular disulfide bridges in the adsorption layer. Subsequently, hydrophobic residues in the α-helices become exposed, penetrating the hydration layer at the solid–liquid interface. Subsequently, hydrophilic and hydrophobic residues deposit at random to achieve interfacial adhesion. The initially organized layer is enriched in α-helical secondary structures, reinforced by intermolecular disulfide bonds, resulting in the organization of a graded layered protein layer. The process of β-sheet stacking is subject to a gradual transformation from α-helices, occurring in an evolving manner. The ultrafast surface priming process has been shown to induce the stepwise stacking of unfolded proteins at the interface, forming protein nanofilms with controlled monolayers, few layers, or multiple molecular layers. Protein nanofilms with tuneable molecular layers and stable interfacial adhesion on various solid surfaces may exhibit great application value in the fields of biointerfaces, advanced materials technology, and so forth. Furthermore, these findings may provide unprecedented insights into the development of creative biomedicine, biotechnology, and advanced material systems, which could lead to the effective management of amyloid-related bioadhesion problems.
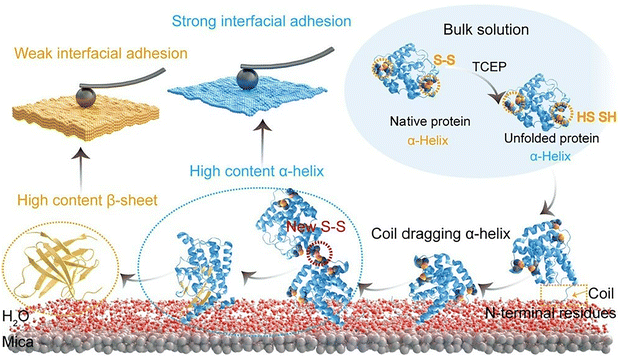 | ||
| Fig. 8 Protein adhesion pathway from the bulk solution to the solid–liquid interface by the α-helix-mediated interfacial adhesion model accompanied with changes in secondary structures and disulfide bonding, which was investigated using AFM probe force measurements. Reprinted with permission from ref. 61 Copyright 2023 American Chemical Society. | ||
The significant disparity in elastic modulus between a metal coating and a polymer substrate gives rise to interfacial mismatch. This issue has the potential to have a significant impact on the stability of flexible devices. Consequently, enhancing the adhesion stability of metal layers on inert polymer substrates, while concurrently preventing delamination, has emerged as a pivotal challenge. In a recent study, Liu, Yang, and coworkers have proposed the use of an ultrathin protein nanofilm, synthesized by disulfide bond-reducing protein aggregation, as a potent adhesive layer to enhance adhesion between polymer substrates and metal coatings (Fig. 9).62 Such an ultrathin protein nanofilm directly connects the two interfaces with a thickness of approximately a single molecule, thereby enhancing interfacial adhesion strength and reducing the contribution of cohesive forces in the overall adhesive system. In this approach, the structural control of protein aggregation induced by the cleavage of disulfide bonds in BSA allows for precise control of the nanofilm thickness. Consequently, an ultrathin adhesive layer measuring approximately 5 to 15 nm was obtained. This nearly monolayer adhesive layer significantly improves interfacial adhesion strength through an α-helix-mediated multisite protein adhesion mechanism and efficient contact between the flexible protein chains and the substrate. Moreover, this approach virtually eliminates the contribution of aggregation by directly interacting two adjacent surfaces with a single molecular chain. The adhesion of multifunctional metal coatings to flexible polymer substrates was achieved through the utilization of an ultrathin protein nanofilm as an adhesive layer, by means of ion sputtering. This method resulted in unparalleled adhesion stability, even under repeated mechanical deformation. Applications of this method include reversible transparency control, strain-responsive encryption, reusable optical sensing, and wearable capacitive touch sensors, facilitating precise control of robotic arms. Moreover, by supplying a biocompatible solution with elevated surface activity and diminished cohesion, it may also promote the development of hybrid devices with stable metal nanocoatings.
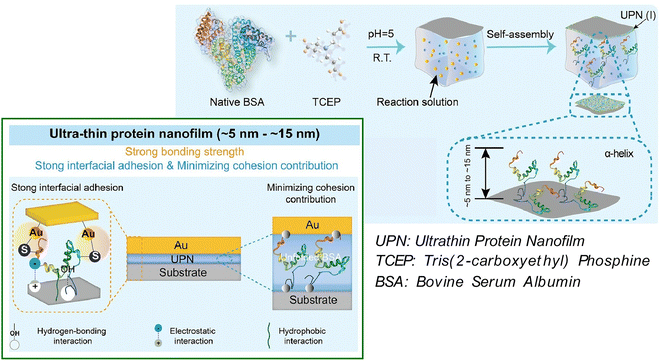 | ||
| Fig. 9 Use of an ultrathin protein nanofilm, synthesized by disulfide bond-reducing protein aggregation, as a potent adhesive layer to enhance adhesion between polymer substrates and metal coatings. Reprinted with permission from ref. 62 Copyright 2024 Wiley-VCH. | ||
While not specifically focused on the organisation of proteins on solid surfaces, there has also been research on the solid–liquid interface, i.e. on the surface of proteins in water. One of the subjects of this research is ice-nucleation active bacteria. It has been demonstrated that bacteria capable of ice-nucleation can promote the growth of ice more effectively than any other known substance. When suspended in the atmosphere, they have been observed to promote ice nucleation in clouds and potentially affect global precipitation patterns. In consideration of their environmental significance, it is imperative to elucidate the molecular mechanism of freezing by ice-nucleating proteins. Weidner and coworkers conducted research in order to elucidate the structural basis of the interaction between water and the ice nucleating protein Z (InaZ) from the ice-nucleation active bacterium Pseudomonas syringae (P. syringae) (Fig. 10).63 Utilizing the techniques of vibrational sum-frequency generation and two-dimensional infrared spectroscopy, the researchers demonstrated that the ice-active repeats of InaZ adopt a β-helical structure in solution and on the water surface. Through the integration of vibrational sum-frequency generation experimental data with molecular dynamics simulations and spectral calculations, it was demonstrated that InaZ undergoes reorientation at low temperatures. The following schematic diagram illustrates the orientation of proteins. It has been demonstrated that InaZ exhibits reorientation at low temperatures, becoming more tilted at 5 °C. In the context of densely packed protein layers, this cold orientation has been demonstrated to enhance the interaction of the ice-nucleation active site with water. In addition, direct experimental evidence has been provided that InaZ adopts a β-helical structure in solution as well as in protein layers interfacing with water. This reorientation and conformational change may enhance water interactions and thus increase the efficiency of ice nucleation. The data presented herein support the hypothesis that InaZ is the driving force behind the water ordering observed at low temperatures in P. syringae. When InaZ is cooled from room temperature to the “operating temperature” of the ice-nucleating protein, it reorients at the interface, thereby increasing the exposure of the ice-nucleation active site. The activation of this process under the influence of temperature has been shown to increase the ordering of water molecules, leading to the formation of ice crystals on the surface.
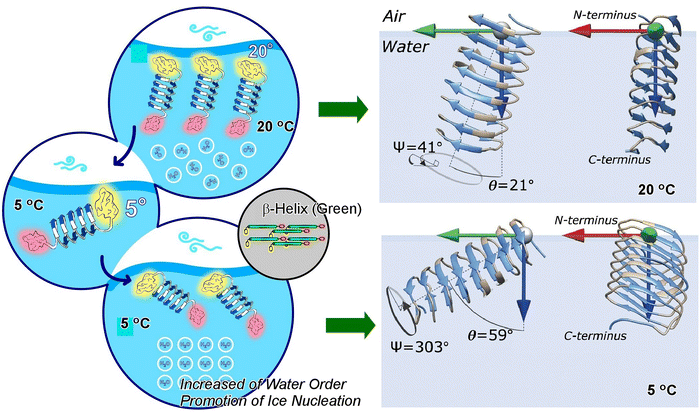 | ||
| Fig. 10 (Top) At room temperature, the ice-nucleating sites are buried in the protein film resulting in relatively low water order, but (bottom) at the low-temperature protein with the β-helix axis parallel to the surface, the ice-nucleating sites are exposed to the interfacial water layer with increased water order. Reproduced under terms of the CC-BY license from ref. 63, 2021 Springer-Nature. | ||
The subject of protein assembly on solid surfaces has been extensively researched through the utilization of well-established techniques, including the LbL method,64 the Langmuir–Blodgett method,65 and the supported lipid bilayer (SLB) method.66 In this section, we propose a series of examples that do not utilize the aforementioned popular methods. The prevailing focus has been on the stabilization of proteins and the successful assembly of these molecules. However, recent examples have demonstrated a growing interest in more detailed secondary structural changes. In other words, researchers are exploring the potential functions of proteins, rather than simply immobilizing their well-known functions on solid substrates. Protein nanoarchitectonics at solid–liquid interfaces appears to have evolved to a higher level, with research focused on elucidating unsolved questions and revealing new functions inherent in proteins.
4. Liquid–liquid interface
Liquid–liquid interfaces, which exhibit more flexible and ambiguous boundaries than gas–liquid or mixed-liquid interfaces, are not necessarily straightforward to analyse or control, and many aspects remain unexplored, rendering them a frontier in the field. However, as liquid–liquid phase separation structures have been shown to play an important role in biological activities, the importance of liquid–liquid interfaces for biomolecules is increasingly recognized.67 In the ensuing sections, we will introduce examples of protein organization at two liquid–liquid interfaces: an oil–water interface and a water–water interface.4.1. Oil–water interface
One interface that can readily form a liquid–liquid interface is that formed by a mixture of immiscible solvents, such as organic solvents and water. These interfaces are also referred to as oil–water interfaces. It is evident that biological materials, such as proteins, typically do not require the utilization of organic solvents. These materials may undergo a process of denaturation, thereby precluding the use of organic solvents as a predominant medium for protein assembly. Nevertheless, in recent years, the significance of oil–water interfaces in biology has been acknowledged, and pioneering research endeavours have been undertaken.68 Research on protein adsorption behaviour at oil–water interfaces has ranged from basic research to application-oriented research.The control and manipulation of protein behaviour at interfaces is intimately connected to a wide range of physicochemical and biological phenomena and technological processes. Protein adsorption at oil–water interfaces is of fundamental importance in many biological and biotechnological processes. For instance, the prediction and control of the properties of functional foods, particularly those based on emulsions stabilized with dairy proteins, requires a comprehensive understanding of the adsorption-induced changes in proteins and the interface itself. Consequently, a comprehensive molecular-level understanding of the mechanisms of protein adsorption at interfaces is imperative. McGrath et al. utilized atomistic molecular dynamics simulations to evaluate the adsorption characteristics of β-lactoglobulin at different oil–water interfaces (Fig. 11), and then compared the outcomes with those of a preceding study employing the hydrophobic oil decane.69 In this study, the oil phases employed included octanol, which exhibited slight hydrophilic properties, and triolein, which exhibited significantly higher hydrophilicity. Utilizing molecular dynamics simulations, the approach and adsorption of β-lactoglobulin to the oil–water interface was tracked with atomic precision. The behaviour of both the surface approach and adsorption mechanisms was contingent upon the hydrophilicity of the oil and the interfacial tension of the oil–water interface. The nature of adsorption, the resulting structural changes, and the energy driving force varied significantly depending on the type of oil. The simulation results were consistent with the experimental findings, including the observation that proteins undergo significant structural changes upon adsorption to hydrophobic surfaces, resulting in the flattening of the protein and exposure of hydrophobic interior residues. In contrast, hydrophilic surfaces formed thicker, native-protein-like adsorption layers. The structural and mechanistic details behind each adsorption mechanism were revealed. The interfacial tension and the relative hydrophilicity of various oils were found to be factors determining the adsorption properties of β-lactoglobulin. A more profound comprehension of the forces that govern protein adsorption behaviour at oil–water interfaces is anticipated to facilitate the implementation and engineering of protein/oil systems, including functional foods and drug delivery systems.
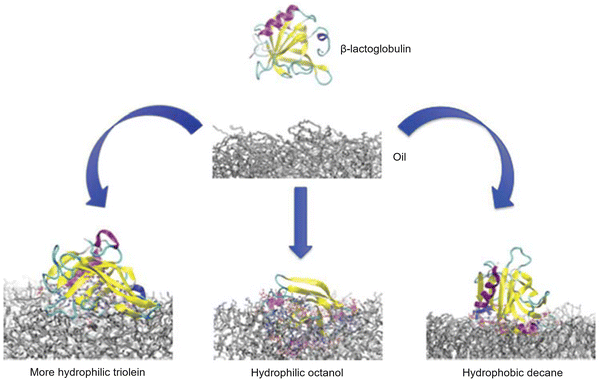 | ||
| Fig. 11 Atomistic molecular dynamics simulations to evaluate the adsorption characteristics of β-lactoglobulin at three distinct oil–water interfaces. At the more hydrophilic triolein interface (moderate tension), β-LG remains mainly in the aqueous phase, forming only limited contacts with hydrophilic residues. At the hydrophilic octanol interface (lowest interfacial tension), β-LG penetrates deeply into the oil phase. In contrast, at the hydrophobic decane interface (highest tension), β-LG undergoes structural rearrangement, exposing hydrophobic residues toward the oil phase. Reprinted with permission from ref. 69 Copyright 2016 American Chemical Society. | ||
The adsorption of protein layers at oil–water interfaces is of crucial importance for the formation and stability of various emulsions, for example, in technological applications and the storage of biological lipids. Bergfreund, Fischer, and coworkers conducted a study to evaluate the effect of high-purity oil–water interfaces on the formation and properties of β-lactoglobulin adsorbed layers (Fig. 12).70 Utilizing the principles of drop profile tensiometry and interfacial rheometry, the researchers measured the adsorption rate, swelling ratio, and interfacial shear modulus. It was found that the adsorption properties of globular proteins differed depending on the molecular size, flexibility, hydrophobicity, polarity, and polarizability of oils. The greater the interaction between polar oils and the hydrophilic outer surface of native β-lactoglobulin, the slower the protein unfolding. This resulted in a decrease in surface pressure and a slower formation of a viscoelastic network. The viscoelastic response of equilibrium β-lactoglobulin films to swelling ratio and shear modulus was consistent for all aliphatic oils due to their mutual non-polar nature. Conversely, polar oils infiltrated the protein network, thereby functioning as emollients, culminating in a comparatively less robust interfacial layer. It was hypothesized that higher polarity would result in lower swelling and shear modulus values. To elaborate, the precise customization of the protein network at the interface of immiscible fluids is paramount for the creation of bespoke interfacial systems. It is imperative to comprehend the factors that influence interfacial protein films, as this knowledge is essential for the formulation of pharmaceuticals, cosmetics, food emulsions, and biological lipid storage containers.
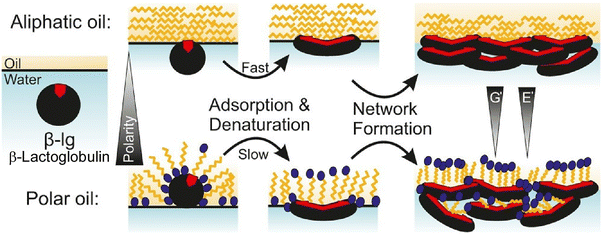 | ||
| Fig. 12 Formation of β-lactoglobulin adsorbed layers at the oil–water interface with aliphatic oil (top) and polar oil (bottom). Reprinted with permission from ref. 70 Copyright 2018 American Chemical Society. | ||
Wooster, Aguilar, and their coworkers employed a combination of synchrotron radiation circular dichroism spectroscopy, front-face fluorescence spectroscopy, and dual polarization interferometry to characterize the conformation and geometry of β-lactoglobulin upon adsorption at two oil–water interfaces: hexadecane–water and tricaprylin–water.71 Subsequent to adsorption at both oil–water interfaces, β-lactoglobulin underwent a β-sheet-to-α-helix transition, concomitantly losing its globular tertiary structure. This study provides insight into the structural properties of proteins adsorbed at oil–water interfaces and has important implications for the formulation and manufacturing of emulsions stabilized by globular proteins. Bergfreund, Fischer, and coworkers investigated the effect of oil polarity on globular protein networks at the oil–water interface.72 The formation dynamics and mechanical properties of interfacial networks of different globular proteins, lysozyme, β-lactoglobulin, and BSA were studied using interfacial shear and dilational rheometry. It was established that oil polarity exerts a substantial influence on network formation, the degree of interfacial protein unfolding, the location of interfacial proteins, and the resulting network strength. This may enable the prediction of protein emulsion stabilization, the tailoring of interfacial layers with desirable mechanical properties, and the preservation of protein structure and function upon adsorption.
Gautrot and colleagues utilised interfacial shear rheology to quantify the change in interfacial shear modulus as a function of the composition in BSA/β-casein mixed assemblies at the interface between the fluorinated oil Novec 7500 and phosphate-buffered saline (Fig. 13).73 Furthermore, neutron reflectivity and fluorescence recovery after photobleaching were utilised to characterise the structure and surface diffusion of the corresponding protein assemblies. The dynamics exhibited by these two proteins at the oil–water interface were found to be markedly divergent. The prevailing role of β-casein in regulating interfacial shear dynamics was observed, exhibiting a strong correlation with the pronounced asymmetry in the adsorption of these two proteins. Despite exhibiting comparable surface densities, BSA and β-casein assemblies at the oil–water interface exhibited interfacial shear storage moduli differing by almost two orders of magnitude and markedly distinct viscoelastic profiles. Furthermore, it was found that coadsorption and sequential adsorption processes could be utilized to further tune the interfacial shear dynamics. The comprehension of complex mixed protein assemblies and their dynamics may offer benefits that extend beyond the domain of formulation science. For instance, it could facilitate the elucidation of alterations in the mechanical strength of corresponding interfaces in tissue culture and physiological conditions.
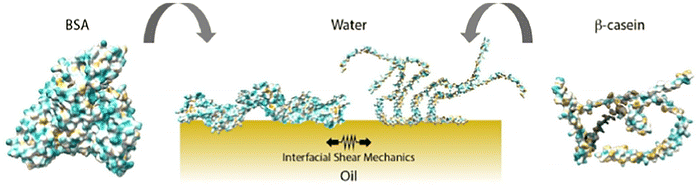 | ||
| Fig. 13 BSA/β-casein mixed assemblies at the interface between the fluorinated oil Novec 7500 and phosphate-buffered saline. Reproduced under terms of the CC-BY license from ref. 73 Copyright 2024 Elsevier. | ||
The cultivation of cells at the interface between two immiscible liquids, such as oil droplets in water, is a promising approach to increase the yield of adherent cells while eliminating the use of microplastics.74 Such a process necessitates cell adhesion to an interface that is stabilized and reinforced by protein nanosheets. These nanosheets exhibit high elasticity and also display cell adhesive ligands capable of binding to integrin receptors. Gautrot and colleagues have demonstrated that the coassembly of supercharged albumin with the cosurfactant pentafluorobenzoyl chloride results in the formation of strong, elastic protein nanosheets (Fig. 14).75 These nanosheets have been shown to mediate extracellular matrix protein adsorption and cell adhesion. The adsorption of extracellular matrix protein to the resulting supercharged nanosheets was quantified by surface plasmon resonance and fluorescence microscopy. The present study has demonstrated that supercharged albumin plays a dual role as both a scaffold protein-structured liquid–liquid interface and a substrate for capturing extracellular matrix molecules. Furthermore, the adhesion and proliferation of primary human epidermal stem cells on bioemulsions stabilized with supercharged nanosheets on droplets was investigated. The process of focal adhesion formation is ordinarily subject to regulation by the substrate stiffness to which the cells are adherent. However, it is the nanoscale mechanics that govern these processes, rather than the bulk mechanical properties. This concept has been applied to cell adhesion to liquid substrates, such as fluorophilic oils and silicone oils, where cell culture is enabled by assembling rigid, elastic polymer or protein nanosheets at the corresponding interfaces. The surfactant-reinforced supercharged protein nanosheets offer a combination of high interfacial modulus, elasticity, and the ability to promote adhesion to the extracellular matrix. This process has been shown to promote cell adhesion and focal adhesion formation, despite the absence of an underlying rigid or elastic substrate. The demonstration of long-term proliferation of mesenchymal stem cells at such interfaces may pave the way for the application of these systems in bioreactors.
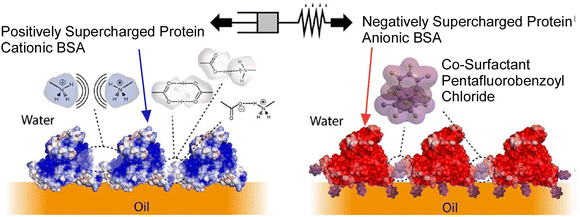 | ||
| Fig. 14 Coassembly of supercharged albumin with the cosurfactant pentafluorobenzoyl chloride for the formation of strong, elastic protein nanosheets. Reproduced under terms of the CC-BY license from ref. 75, 2023 American Chemical Society. | ||
Furthermore, investigations are underway to ascertain the processes involved in protein modification at oil–water interfaces. In systems where the interface is predominant, lipid peroxidation and interfacial protein alignment frequently coexist. The role of interfaces in the promotion of lipid-derived electrophiles, particularly α,β-unsaturated aldehydes, in the extensive modification of proteins, and their impact on protein behaviour, remains to be elucidated. Zhu and coworkers synthesized an yne-ACR probe in order to simulate lipid-derived electrophiles.76 They then proceeded to study the effect of this modification on whey protein in an interface-dominated system model consisting of n-hexadecane and water (Fig. 15). Their research demonstrated that interfaces play a pivotal role in the mediation of protein modification. In contrast to homogeneous systems, interfaces offer a distinct environment for protein modification, resulting in varied aggregation behaviours between interfacial and non-adsorbed proteins. All-atom molecular dynamics simulations have revealed that protein modification disrupts the stability of protein aggregates at interfaces, promotes redistribution between the interface and bulk phase, and alters interfacial activity. This approach enhanced our comprehension of the manner in which interfaces influence target protein modification by lipid-derived electrophiles under particular conditions. Such interfacial studies may highlight the importance of recognizing the role of interfaces in various natural processes and leveraging them more effectively for technological advances.
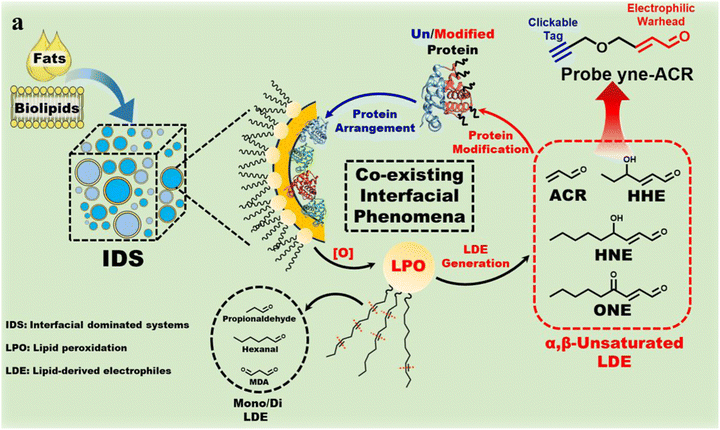 | ||
| Fig. 15 Schematic illustration of lipid peroxidation (LPO) and protein modification co-occurring at oil–water interfaces in interfacial-dominated systems (IDSs). A clickable probe (yne-ACR) mimics α,β-unsaturated lipid-derived electrophiles (LDEs) to visualize protein modification at the interface. Reprinted with permission from ref. 76 Copyright 2025 American Chemical Society. | ||
The oil–water interface represents a frontier for the organization of biological materials, and various fundamental characterizations of the interface are currently being conducted. Protein adsorption to the interface and the accompanying changes in secondary structure, as well as the evaluation of various mechanical properties and protein modification behaviour, have been observed. Active basic research is being conducted on these interfaces, with a focus on emerging applications in food and healthcare, cell culture, and biotechnology. A range of analytical techniques, including interfacial rheometry, synchrotron radiation circular dichroism spectroscopy, fluorescence spectroscopy, dual-polarization interferometry, neutron reflectometry, and fluorescence recovery, are being applied to these analyses. Furthermore, simulations such as all-atom molecular dynamics have also made valuable contributions. The field of protein nanoarchitectonics at oil–water interfaces offers considerable potential for further development and will likely continue to be an attractive area of research.
4.2. Water–water interface
It is evident that, due to the absence of organic solvents within actual biological systems, the phenomenon of oil–water interfaces is non-existent in realistic biological environments. However, it is becoming increasingly evident that liquid–liquid phase separation plays a significant role in biological activities. Consequently, interfacial structures consisting exclusively of aqueous solutions, i.e., water–water interfaces, are becoming a subject of significant interest. The scientific community has recently dedicated significant attention to the study of liquid–liquid phase separation and coacervate formation involving biological materials, such as proteins. Numerous examples of this phenomenon have already been documented.77 Further elaboration on these can be found in other sources. The present study does not intend to explore these issues in any further depth. In this section, we will introduce a limited number of examples of protein organization at water–water interfaces for the purpose of further discussion.The construction of functional synthetic microcompartments at the water–water interface is useful for deepening our understanding of cellular processes and is important for developing applications in various fields. Naz, Arslan, and coworkers have developed lipoproteasomes, which are microscale compartments composed of BSA and stearoyl chloride nanoconjugates. These were synthesized using thiourea coupling chemistry (Fig. 16).78 Protein–lipid nanocomplexes were first assembled into lipoproteasomes. Subsequently, hydrophobic chains were grafted onto BSA, followed by Pickering emulsification, crosslinking, oil removal via dialysis, and final stabilization in the aqueous phase. These microcapsules are capable of encapsulating and retaining water-soluble macromolecules and exhibit enzyme-driven communication reminiscent of protocells. In addition, these lipoproteasomes display molecular-weight-dependent permeability and remarkable stability, making them highly promising for targeted drug delivery, biosensing, and synthetic biology. Moreover, the integration of stimuli-responsive components could further enhance the functionality of lipoproteasomes, enabling dynamic and controllable behaviors.
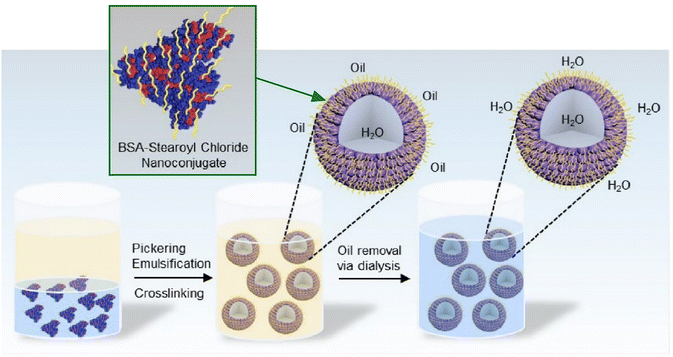 | ||
| Fig. 16 Assembly of protein–lipid nanoconjugates into lipoproteinosomes involves hydrophobic chain introduction in BSA, followed by lipoproteinosome formation via Pickering emulsification, cross-linking, and their ultimate transfer to the aqueous phase. Reprinted with permission from ref. 78 Copyright 2025 American Chemical Society. | ||
Collectively, such lipoproteasomes represent a potential platform for constructing next-generation protocells capable of performing complex, regulated biological functions.
The process of coacervation via liquid–liquid phase separation presents a distinctive opportunity to address the challenge of designing multifunctional nanostructured biomaterials. Protein-polysaccharide coacervates have been identified as a promising approach for targeting biomaterial scaffolds. It is imperative to enhance the mechanical and chemical stability of the condensates. Mezzenga and coworkers investigated the structuring of cationic protein amyloids and anionic linear polysaccharides through coacervation (Fig. 17).79 By converting natural proteins into amyloid fibrils, they overcame the limitations on stability and demonstrated the achievement of interfacial self-assembly of biomaterials with precisely controlled structure and properties. Coacervation phase diagrams were constructed under different conditions, and the hierarchical structures of the resulting materials were systematically characterized. Utilizing semiflexible amyloid fibrils derived from β-lactoglobulin, a predominant protein constituent of whey, as the polycation, and hyaluronic acid, a long-chain linear natural polysaccharide, as the polyanion, coacervates were formed at the water–water interface. These coacervates exhibited highly ordered asymmetric structures, with amyloid fibrils on one side and polysaccharides on the other. This material exhibits significantly superior properties in comparison to homologous coacervates derived from the same components but containing proteins in their native form. This particular amyloid polysaccharide coacervate is a unique and effective biomaterial with versatile applications in internal medicine. Specifically, the efficacy of these coacervates in the prevention of gastric ulcers was demonstrated through the validation of their therapeutic efficacy as artificial microparticles in in vivo assays. Moreover, these materials have the potential to extend far beyond therapy to include tissue engineering scaffolds, biomaterial templates, and membranes for molecular separations.
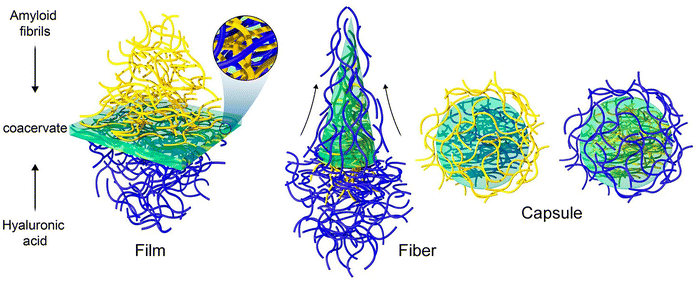 | ||
| Fig. 17 Formation of various co-assembly structures through structuring of cationic protein amyloids and anionic linear polysaccharides (hyaluronic acid) through interfacial coacervation. Reproduced under terms of the CC-BY license from ref. 79, 2023 Springer-Nature. | ||
This section presents two examples of protein-organized structures at water–water interfaces. The first method involves the assembly of proteins at a well-defined oil–liquid interface, with the oil phase being replaced by an aqueous phase. The second example pertains to the formation of protein-organized structures through coacervate formation, a process that arises from liquid–liquid phase separation within an aqueous system. The interfacial environment has been utilized in the creation of various structures. Furthermore, there have been endeavours to induce alterations in the secondary structure of proteins at the interface, with the objective of achieving enhanced functionality. These findings bear a resemblance to the nanoarchitectonics of proteins at various interfaces, as discussed in the preceding sections. The water–water interface, where both interfaces are aqueous, offers greater flexibility in approaching the interface of water-soluble biological materials, potentially enabling a greater variety in the tailoring of their components and structures. The potential applications of this technology are manifold. The practical applications of these systems encompass targeted drug delivery, biosensing, microreactors, tissue engineering scaffolds, molecular separation, and biomedical uses such as the prevention of gastric ulcers. Additionally, it finds application in synthetic biology, enzyme-driven communication, protocell signalling, and cell origin research. The organization of protein at the water–water interface has been shown to share similar characteristics with that observed at other interfaces. While the principles and characteristics are largely similar, the water–water interface exhibits characteristics that are most akin to those of actual biological systems. The water–water interface may be the most suitable environment for applying knowledge gained from model systems, such as air–liquid and solid–liquid interfaces, to more realistic biological systems.
5. Physico-chemical communications with biological processes at liquid interfaces
In the preceding sections, an introduction has been made to the organization, analysis and function of proteins in a variety of interfacial environments. It is anticipated that these organizations will be incorporated into more sophisticated functions. The implementation of these advanced functions will necessitate the collaboration of functional units, as observed in biological systems. It can be concluded that the transmission of physicochemical signals from biological units at interfaces will be a significant factor. In this study, we will examine bioprocesses and physicochemical communication at liquid interfaces using several examples.As demonstrated by the aforementioned examples, the oil–water interface has the capacity to function as a site for protein organization. The organized protein layer has the capacity to communicate signals to attached cells and control cell differentiation. As demonstrated by Jia et al., a protein monolayer is formed at the interface between a perfluorocarbon and an aqueous liquid.80 This monolayer is strong enough for cells to adhere and spread. This environment offers new possibilities for the optimization of cell culture materials. In this study, two different perfluorocarbons, namely perfluorodecalin and perfluorotributylamine, were utilized. The strength of the serum protein monolayer formed at the oil–water interface varies depending on the fluorocarbon. The insertion of fibronectin into the protein monolayer resulted in the observation of human mesenchymal stem cells exhibiting an increased spreading area and larger focal adhesions at the perfluorotributylamine–water interface (Fig. 18A). This finding indicates that cells do not directly sense the bulk stiffness of the perfluorocarbon liquid, but rather sense the nanometer-level stiffness of the protein nanosheet at the liquid interface. Stem cells interact with their substrates to regulate their behaviour and fate. The forces exerted by cells have been shown to alter the morphology and stiffness of the extracellular matrix microenvironment. It is evident that extracellular matrix remodelling exerts a feedback effect that regulates the behaviour and fate of stem cells. The concept of adaptive biomaterials based on protein monolayers assembled at liquid–liquid interfaces is considered a novel one. Liquids possess the capacity to flow and reconfigure their shape, thereby exhibiting a unique response mechanism that is not possible with solid counterparts. Protein assemblies at liquid–liquid interfaces have been observed to dynamically adapt to cell-generated forces through interfacial interference and nanoscale spatial rearrangement. Stretched fibronectin assemblies have been demonstrated to promote the development of extended focal adhesions, enhance focal adhesion kinase activation, and promote neural differentiation. This finding thus demonstrates the existence of a feedback mechanism linking dynamic extracellular matrix mechanics, biological signalling, and the long-term fate of stem cells. The communication of physicochemical signals from protein layers formed at oil–interfaces has been demonstrated to induce biological signals and regulate cell fate. From a pragmatic standpoint, this approach has the potential to stimulate the neural differentiation of human mesenchymal stem cells, thereby obviating the necessity for costly growth factors or intricate fabrication processes. The implementation of such a mechanism could represent a significant advancement in the domain of neural tissue engineering.
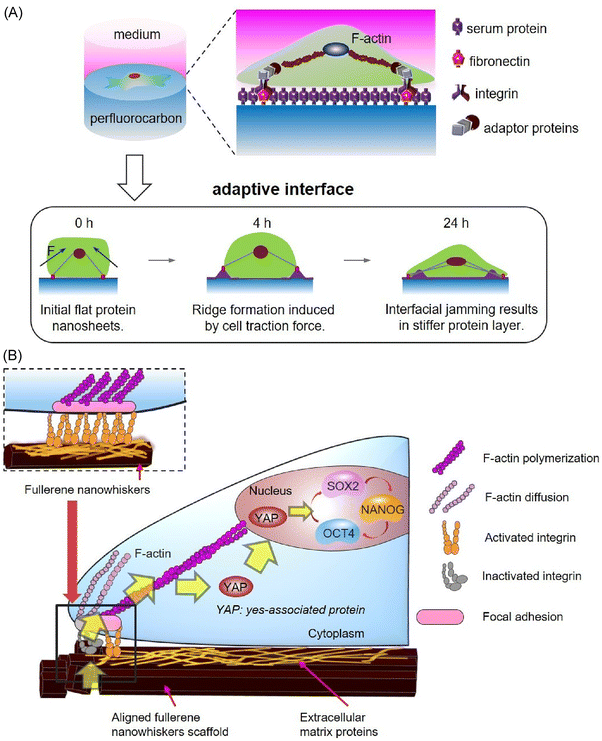 | ||
| Fig. 18 Physico-chemical communication from interfaces to control the differentiation fate of stem cells: (A) at the perfluorotributylamine–water interface; (B) on a solid–liquid interface covered with aligned fullerene nanowhiskers. Reprinted with permission from ref. 80c Copyright 2020 Elsevier and reprinted with permission from ref. 81b Copyright 2020 Wiley-VCH. | ||
Furthermore, the regulation of cells through the communication of physicochemical signals from interfaces has also been observed in solid–liquid interface systems. Song et al. demonstrated enhanced self-renewal and multipotency of mesenchymal stem cells (MSCs) through cell culture on a solid–liquid interface covered with aligned fullerene nanowhiskers (Fig. 18B).81 The aligned 1D hard nanocarbon structure of fullerene nanowhiskers exerts a significant influence on cell proliferation behaviours, including spreading, orientation, and focal adhesions. The appropriate affinity between mesenchymal stem cells and partially hydrophobic surfaces covered with fullerene nanowhiskers results in focal adhesions suitable for cell contraction and nuclear localization of yes-related protein, leading to the maintenance of multipotency of mesenchymal stem cells. In this particular instance, the physicochemical characteristics of the interface are responsible for the communication of cellular biosignals, thereby exerting a substantial control over cell behaviour. Conversely, cells cultivated on conventional flat surfaces, such as glass, manifest pronounced focal adhesion regions accompanied by elevated cytoskeletal tension. This phenomenon impedes the proliferation of mesenchymal stem cells and induces an activated state, accompanied by undesirable differentiation and loss of stemness. The surface with patterned hydrophobic fullerene nanowhiskers demonstrated here changes this conventional wisdom. From a pragmatic standpoint, the sustained maintenance of multipotency and a stem-cell-like phenotype constitutes a pivotal prerequisite for stem cell-based therapeutic interventions. The fabrication of patterned surfaces of aligned fullerene nanowhiskers over large centimetre-sized areas represents a significant advancement in the field, with potential applications in tissue engineering and in vitro proliferation of stem cells in clinical settings.
Finally, we present an example of coupling physicochemical communication and biochemical reactions at a water–solid interface, albeit unrelated to biomolecular systems. This example couples the doping of an organic semiconductor with proton-coupled electron transfer, a mechanism utilised in numerous pathways within cellular respiration, including water oxidation in photosynthesis and nitrogen fixation in oxygen reduction. Ishii, Yamashita, and their coworkers integrated a process referred to as proton-coupled electron transfer with doping at a polymer-organic semiconductor interface (Fig. 19).82 This represents a novel approach that integrates the proton activity of the medium with the chemical doping process of the organic semiconductor. A thin film of an organic semiconducting polymer (poly[2,5-bis(3-tetradecylthiophen-2-yl)thieno[3,2-b]thiophene], PBTTT-C14) is immersed in an aqueous solution containing a benzoquinone/hydroquinone redox couple capable of proton-coupled electron transfer. This results in p-type doping via an interfacial communication process. The efficient chemical doping of polymer organic semiconductors can be achieved through the synergistic reaction of proton-coupled electron transfer and hydrophobic ion intercalation. The doping level can be precisely controlled in pH-controlled aqueous solutions, resulting in several orders of magnitude improvement in conductivity. This method is also innovative in that it can be performed in aqueous solutions on a standard laboratory bench. The utilization of redox reagents in the context of chemical doping has been employed for the purpose of enhancing the properties of organic semiconductors. However, it should be noted that redox reagents are prone to degradation in the presence of water and air, which limits their use to inert gas and vacuum environments. Conversely, the implementation of this process in an aqueous environment, as observed in the present system, successfully circumvents the conventional challenges associated with the doping of organic semiconductors. The underlying mechanisms of these processes are predicated on the coupling of interfacial chemical equilibrium with membrane electrical conductance. Proton-coupled electron transfer is a prevalent process in biological systems with the potential to contribute to bioelectronic applications. Consequently, it may be utilized as a foundation for biomolecular electronics. In addition to the chemical equilibrium demonstrated in this particular instance, a significant number of enzymes and coenzymes frequently undergo redox reactions. The aforementioned concept may be expanded upon by considering the coupling of protein nanoarchitectonics at liquid-related interfaces with physicochemical device functions.
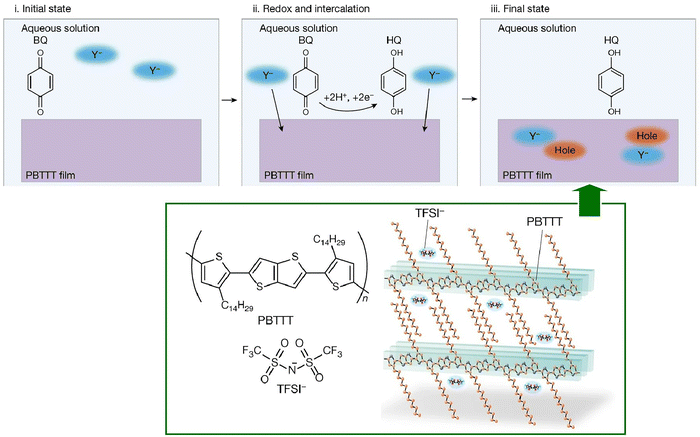 | ||
| Fig. 19 Coupling of proton-coupled electron transfer with doping at a polymer–organic semiconductor interface. Reprinted with permission from ref. 82 Copyright 2023 Springer-Nature. | ||
The examples presented here demonstrate the potential for physico-chemical communication from interfaces to control advanced functions, such as cell differentiation, and to facilitate the development of new methodologies for integrating biochemical reactions with device functions. Whilst these illustrations are less complex than the multifarious actual biological functional systems, they demonstrate that nanoarchitectonics at liquid interfaces has the capacity to facilitate the coordination of functional signals observed in biological systems. The pursuit of such methodologies may offer the prospect of the creation of functional systems that exhibit characteristics similar to those of living organisms.
6. Summary and future perspectives
In order to construct highly functional material systems, it is necessary to synthesize fundamental functional units, control their nanostructure, and assemble them into rational organizations. This constitutes the fundamental principle underpinning the development of functional materials through nanoarchitectonics. The ideal model for this is the functional systems found in living organisms. Natural systems have evolved over billions of years. The objective of nanoarchitectonics is to achieve a comparable outcome within a few decades. This objective is extremely challenging. A fruitful approach to this issue would be the utilization of functional units that have already been established by nature. In this case, proteins have been identified as particularly promising candidates. Proteins are powerful functional units, possessing highly advanced nanostructures that exhibit a wide range of functions. The organization of these elements through nanoarchitectonics has emerged as a pivotal strategy for the creation of more ideal functional material systems.In order to organize functional units in a more strategic and rational manner, it is necessary to establish an appropriate medium for this purpose. Interfaces are a particularly effective media for protein nanoarchitectonics. Proteins ordinarily function in aqueous environments; as such, interfaces involving liquids, especially aqueous media, are of crucial significance. The present article expounds on several recent examples of protein nanoarchitectonics at various liquid interfaces (Fig. 20). This feature article does not aim to simply list recent examples, but rather seeks to identify trends in protein organization and function at interfaces involving liquids. Techniques for creating interfaces are diverse, and many have been developed. Understanding interface-specific trends in organization and function provides guidance for designing interfaces to obtain the desired silk. Below, we summarize the characteristics of interface-specific protein nanoarchitectonics derived from this feature article.
The air–water interface is an ideal medium for the organization of water-soluble proteins into ultrathin films, and also serves as a simple model system. Consequently, it is an optimal environment for the evaluation and analysis of protein organization. The organization of proteins from the liquid phase onto solid surfaces has been the subject of extensive research. The present research is not confined to the immobilization of well-known protein functions on solid substrates; it also explores the potential functions of proteins. Protein nanoarchitectonics at solid–liquid interfaces appears to be evolving to a higher level, with research focused on elucidating unsolved problems and revealing new functions inherent in proteins. The oil–water interface also poses a frontier for the organization of biological materials, and various fundamental characterizations of these interfaces have been conducted. The adsorption of proteins to interfaces, and the accompanying changes in secondary structure, have been observed, as well as the evaluation of various mechanical properties and protein modification behaviour. The water–water interface exhibits characteristics that are most analogous to those of actual biological systems. The water–water interface is considered the optimal environment for the application of knowledge gained from model systems, such as air–liquid and solid–liquid interfaces, to more realistic biological systems. As summarized here, this feature article classifies interfaces from a physicochemical perspective. It is also noteworthy that these characteristics apply to material systems related to actual liquid interfaces. The insights gained here can be extended to interfaces formed by polymer–inorganic composites83 and emulsion systems.84
The present feature article focuses on examples that clearly reflect the characteristics of each interface. Nevertheless, these illustrations underscore the numerous commonalities between interfaces, thereby overshadowing their divergent characteristics. In certain interfaces, endeavours are underway to immobilize proteins themselves, in addition to controlling alterations in their secondary structure, thereby engendering properties that are not inherently functional. The latent capabilities of proteins are being utilized. Interfacial environments provide an ideal platform for protein analysis, where simulations such as molecular dynamics are being utilized. Furthermore, it has been demonstrated that interactions and phenomena at interfaces can be developed into more advanced functions. For instance, physicochemical communication at interfaces has been demonstrated to regulate advanced biological functions, including cell differentiation. Moreover, new methodologies can be developed that couple biochemical reactions with device functions. The significance of the coordination of functional signals is demonstrated by these systems. The pursuit of such methodologies may facilitate the creation of functional systems that exhibit characteristics analogous to those of living organisms.
A plethora of captivating research endeavours have been undertaken in the realm of nanoarchitectonics of proteins at liquid interfaces. In comparison with the multi-component and multi-functional systems present in living organisms, artificially assembled systems remain primitive. In order to construct highly functional material systems that approach those of living organisms, it is necessary to create systems in which a multitude of functional components and physicochemical interactions are rationally integrated with biomaterials. It is imperative that this process is undertaken on the basis of accumulated knowledge. The construction of more complex systems necessitates the incorporation of additional elements. The design of such systems cannot be accomplished simply through the application of existing experience, knowledge, and the prevailing theories that govern them. The process necessitates the extraction of conclusions from a substantial corpus of data. Furthermore, proteins and their assemblies are characterized by irregular and complex structures and behaviours, which renders them extremely challenging. Fortunately, humanity has acquired a new tool in the form of artificial intelligence (AI). Indeed, there is active research in progress on chemical control and material synthesis methods using machine learning.85 The combination of nanoarchitectonics and artificial intelligence has also been mooted.86 The implementation of this information technology is expected to be a key factor in the advancement of the nanoarchitectonics of proteins and tissues, which are characterized by increasingly sophisticated functions. In particular, interfaces involving liquids are ideal locations for the flexible organization of protein functional units. If AI can provide design guidelines and methodologies, then it will be possible to create complex yet highly functional material systems. The evolutionary process of life, which required prolonged periods of selection and elimination to create optimal structures, can now be replicated by AI in an exceedingly brief timeframe. At that juncture, nanoarchitectonics will evolve into a methodology that facilitates the instantaneous creation and assembly of advanced functions.
Author contributions
J. S.: conceptualization, writing, review & editing. K. K.: project administration, review & editing. K. A.: conceptualization, writing, review & editing, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
Data availability is not applicable to this article as no new data were created or analysed in this study.Acknowledgements
This study was partially supported by the Japan Society for the Promotion of Science KAKENHI (Grant Numbers JP23H05459 and JP25H00898).Notes and references
- (a) M. N. Lakhan, A. Hanan, A. Hussain, I. A. Soomro, Y. Wang, M. Ahmed, U. Aftab, H. Sun and H. Arandiyan, Chem. Commun., 2024, 60, 5104–5135 Search PubMed; (b) T. Nakamura, Y. Kondo, N. Ohashi, C. Sakamoto, A. Hasegawa, S. Hu, M. A. Truong, R. Murdey, Y. Kanemitsu and A. Wakamiya, Bull. Chem. Soc. Jpn., 2024, 97, uoad025 CrossRef CAS; (c) P. A. Shinde, Q. Abbas, N. R. Chodankar, K. Ariga, M. A. Abdelkareem and A. G. Olabi,, J. Energy Chem., 2023, 79, 611–638 CrossRef CAS; (d) G. Chen, M. Isegawa, T. Koide, Y. Yoshida, K. Harano, K. Hayashida, S. Fujita, K. Takeyasu, K. Ariga and J. Nakamura, Angew. Chem., Int. Ed., 2024, 63, e202410747 CrossRef CAS PubMed; (e) W. Yoshimune, Bull. Chem. Soc. Jpn., 2024, 97, uoae046 CrossRef CAS.
- (a) N. Chen, M. Hu, L. Gou, L. Tan, D. Zhao and H. Feng, Bull. Chem. Soc. Jpn., 2024, 97, uoae030 CrossRef CAS; (b) S. Liang, H. Feng, N. Chen, B. Wang, M. Hu, X. Huang, K. Yang and Y. Gu, Bull. Chem. Soc. Jpn., 2024, 97, uoae054 Search PubMed; (c) S. Miyazaki, N. Ogiwara, C. A. Nagasaka, K. Takiishi, M. Inada and S. Uchida, Bull. Chem. Soc. Jpn., 2024, 97, uoae105 CrossRef CAS; (d) A. Jancik-Prochazkova and K. Ariga, Research, 2025, 8, 0624 CrossRef CAS PubMed; (e) A. Mariania and G. Malucelli, Chem. Commun., 2025, 61, 2173–2189 RSC.
- (a) X.-Y. Guo, L. Yi, J. Yang, H.-W. An, Z.-X. Yang and H. Wang, Chem. Commun., 2024, 60, 2009–2021 RSC; (b) P. H. Syaifie, M. A. F. Nasution, I. Rahmawati, E. Saepudin and T. A. Ivandini, Bull. Chem. Soc. Jpn., 2024, 97, uoad007 Search PubMed; (c) T. Miura, T. R. Malla, L. Brewitz, A. Tumber, E. Salah, K. J. Lee, N. Terasaka, C. D. Owen, C. Strain-Damerell, P. Lukacik, M. A. Walsh, A. Kawamura, C. J. Schofield, T. Katoh and H. Suga, Bull. Chem. Soc. Jpn., 2024, 97, uoae018 CrossRef CAS PubMed; (d) K. Matsunaga, M. Takahashi, T. Kagaya, D. Takahashi and K. Toshima, Bull. Chem. Soc. Jpn., 2024, 97, uoae057 CrossRef CAS; (e) C. Zhao, W. Song, J. Wang, X. Tang and Z. Jiang, Chem. Commun., 2025, 61, 1962–1977 RSC.
- (a) Y. Nakao, Bull. Chem. Soc. Jpn., 2024, 97, uoae027 CrossRef CAS; (b) Y. Hayashi, Bull. Chem. Soc. Jpn., 2024, 97, uoae039 CrossRef CAS; (c) B. Maedaa and K. Murakami, Chem. Commun., 2024, 60, 2839–2864 Search PubMed; (d) Y. Inokuma,, Bull. Chem. Soc. Jpn., 2024, 97, uoae072 Search PubMed; (e) T. Yasukawa, Bull. Chem. Soc. Jpn., 2024, 97, uoae076 Search PubMed.
- (a) D. A. Buschmann, H. Hirai and T. Tsukuda, Inorg. Chem. Front., 2024, 11, 6694–6710 Search PubMed; (b) J. Yang, Y. Sekizawa, X. Shi, K. Ijiro and H. Mitomo, Bull. Chem. Soc. Jpn., 2024, 97, uoae073 CrossRef CAS; (c) K. Sasaki, Appl. Phys. Express, 2024, 17, 090101 CrossRef CAS; (d) R. Tahawy, M. Esmat, H. El-Hosainy, F. E. Farghaly, E.-S. A. Abdel-Aal, F. I. El-Hosiny and Y. Ide, Bull. Chem. Soc. Jpn., 2024, 97, uoae079 CrossRef CAS.
- (a) J. Kim, J. H. Kim and K. Ariga, Joule, 2017, 1, 739–768 CrossRef CAS; (b) C. Liu, N. Morimoto, L. Jiang, S. Kawahara, T. Noritomi, H. Yokoyama, K. Mayumi and K. Ito, Science, 2021, 372, 1078–1081 CrossRef CAS PubMed; (c) T. Yamamoto, A. Takahashi and H. Otsuka, Bull. Chem. Soc. Jpn., 2024, 97, uoad004 CrossRef CAS; (d) Y. Kim, K. Iimura and N. Tamaoki, Bull. Chem. Soc. Jpn., 2024, 97, uoae034 Search PubMed; (e) M. Kamigaito, Bull. Chem. Soc. Jpn., 2024, 97, uoae069 CrossRef CAS.
- (a) S. Datta, Y. Kato, S. Higashiharaguchi, K. Aratsu, A. Isobe, T. Saito, D. D. Prabhu, Y. Kitamoto, M. J. Hollamby, A. J. Smith, R. Dalgliesh, N. Mahmoudi, L. Pesce, C. Perego, G. M. Pavan and S. Yagai, Nature, 2020, 583, 400–405 CrossRef CAS PubMed; (b) M. Yamaguchi, M. de la, H. Tomás, A. Fujiwara, R. Oketani, K. Okubo, K. Oka, N. Tohnai, A. Douhal and I. Hisaki, Bull. Chem. Soc. Jpn., 2024, 97, uoae004 CrossRef CAS; (c) H. Fujimoto, T. Hirao and T. Haino, Bull. Chem. Soc. Jpn., 2024, 97, uoad016 CrossRef CAS; (d) M. Kataria and S. Seki, Chem. – Eur. J., 2025, 31, e202403460 CrossRef CAS PubMed; (e) R. Kudo, R. Kawai, S. Takamiya, S. Datta, H. Hanayama, N. Hara, H. Tamiaki and S. Yagai, Chem. Commun., 2025, 61, 8427–8430 Search PubMed.
- (a) T. D. Bennett and S. Horike, Nat. Rev. Mater., 2018, 3, 431–440 Search PubMed; (b) M. Nakai, K. Asano, K. Shimada, K. Kanno, Y. Nakabayashi, L. Alba, Y. Funahashi, S. Yano and H. Ishida, Bull. Chem. Soc. Jpn., 2024, 97, uoad017 Search PubMed; (c) M. Yuasa, S. Nakamura and H. Tamiaki, Bull. Chem. Soc. Jpn., 2024, 97, uoae029 Search PubMed; (d) D. W. Burke, M. Yamashita, Z. Wang, M. Kuzumoto, K. Urayama, K. Saito and S. Furukawa, Chem. Sci., 2025, 16, 8509–8522 Search PubMed; (e) H. Narita, A. Virovets, H.-W. Lerner, M. Wagner and S. Yamaguchi, Chem. Sci., 2025, 16, 8764–8771 Search PubMed.
- (a) M. Komiyama, K. Yoshimoto, M. Sisido and K. Ariga, Bull. Chem. Soc. Jpn., 2017, 90, 967–1004 CrossRef; (b) S. d Kalyana Sundaram, M. M. Hossain, M. Rezki, K. Ariga and S. Tsujimura, Biosensors, 2023, 13, 1018 CrossRef CAS PubMed; (c) M. Imai, J. Uchiyama, I. Takemura-Uchiyama, S. Matsuzaki, Y. Niko, S. Hadano and S. Watanabe, Bull. Chem. Soc. Jpn., 2024, 97, uoad010 CrossRef CAS; (d) M. Panchabashini, H. Pedige, A. Sugawara and H. Uyama, Bull. Chem. Soc. Jpn., 2024, 97, uoae065 CrossRef; (e) L. Sutrisno, G. J. Richards, J. D. Evans, M. Matsumoto, X. Li, K. Uto, J. P. Hill, M. Taki, S. Yamaguchi and K. Ariga, Sci. Adv., 2025, 11, eadz6633 CrossRef CAS PubMed.
- (a) K. Ariga, S. Akakabe, R. Sekiguchi, M. L. Thomas, Y. Takeoka, M. Rikukawa and M. Yoshizawa-Fujita, ACS Omega, 2024, 9, 22203–22212 Search PubMed; (b) A. Kuzume and K. Yamamoto, Bull. Chem. Soc. Jpn., 2024, 97, uoae022 Search PubMed; (c) M. Minakshi, A. Samayamanthry, J. Whale, R. Aughterson, P. A. Shinde, K. Ariga and L. K. Shrestha, Chem. – Asian J., 2024, 19, e202400622 Search PubMed; (d) K. Sato, M. Gon, K. Tanaka and Y. Chujo, Bull. Chem. Soc. Jpn., 2024, 97, uoae040 Search PubMed; (e) Z. Guo, C. Pan, A. Shinohara and T. Nakanishi, Sci. Technol. Adv. Mater., 2025, 26, 2515007 CrossRef PubMed.
- (a) K. Saitow, Bull. Chem. Soc. Jpn., 2024, 97, uoad002 Search PubMed; (b) S. Hasegawa, Appl. Phys. Express, 2024, 17, 050101 Search PubMed; (c) K. Hashiguchi, A. Maruo, T. Soeda, S. Iwane and H. Jippo, Bull. Chem. Soc. Jpn., 2024, 97, uoae131 CrossRef CAS; (d) K. Hayashida, J. Nakamura and K. Takeyasu, Angew. Chem., Int. Ed., 2025, 64, e202502702 CrossRef CAS PubMed; (e) H. Imahori and M. Akiyama, Mater. Horiz., 2025, 12, 92–102 Search PubMed.
- (a) Y. Sugimoto, P. Pou, M. Abe, P. Jelinek, R. Pérez, S. Morita and Ó. Custance, Nature, 2007, 446, 64–67 Search PubMed; (b) H. Mogi, R. Wakabayashi, S. Yoshida, Y. Arashida, A. Taninaka, K. Iwaya, T. Miura, O. Takeuchi and H. Shigekawa, Appl. Phys. Express, 2024, 17, 015003 Search PubMed; (c) T. Nakamuro, Bull. Chem. Soc. Jpn., 2024, 97, uoae082 Search PubMed; (d) M. Takeguchi, K. Mitsuishi and A. Hashimoto, Appl. Phys. Express, 2024, 17, 085001 Search PubMed.
- (a) Y. Okawa and M. Aono, Nature, 2001, 409, 683–684 Search PubMed; (b) C. Manzano, W.-H. Soe, H. S. Wong, F. Ample, A. Gourdon, N. Chandrasekhar and C. Joachim, Nat. Mater., 2009, 8, 576–579 CrossRef CAS PubMed; (c) S. Kawai, O. Krejčí, T. Nishiuchi, K. Sahara, T. Kodama, R. Pawlak, E. Meyer, T. Kubo and A. S. Foster, Sci. Adv., 2020, 6, eaay8913 CrossRef CAS PubMed; (d) K. H. Au-Yeung, S. Sarkar, T. Kühne, O. Aiboudi, D. A. Ryndyk, R. Robles, N. Lorente, F. Lissel, C. Joachim and F. Moresco, ACS Nano, 2023, 17, 3128–3134 CrossRef CAS PubMed.
- (a) S. Tsubokura, S. Kawano, Y. Imai, T. Ueda, K. Nakamoto, H. Noma, H. Hioki and T. Minato, Appl. Phys. Express, 2024, 17, 035003 CrossRef CAS; (b) H. Fujiwara, C. Bareille, M. Okawa, S. Shin and T. Taniuchi, Appl. Phys. Express, 2024, 17, 086505 Search PubMed; (c) T. Tahara, Bull. Chem. Soc. Jpn., 2024, 97, uoae012 CrossRef CAS; (d) R. Otsuka, K. Shimizu, H. Wakabayashi and S. Watanabe, Appl. Phys. Express, 2024, 17, 115501 Search PubMed.
- (a) K. Ariga, M. Nishikawa, T. Mori, J. Takeya, L. K. Shrestha and J. P. Hill, Sci. Technol. Adv. Mater., 2019, 20, 51–95 Search PubMed; (b) T. Ito, M. Nakagawa and T. Kawai, Bull. Chem. Soc. Jpn., 2024, 97, uoae075 CrossRef CAS; (c) T. Kurita and K. Numata, Phys. Chem. Chem. Phys., 2024, 26, 28776–28792 RSC; (d) H. Nabika, Bull. Chem. Soc. Jpn., 2025, 98, uoaf048 Search PubMed; (e) T. Sugawara, M. Matsuo and T. Toyota, Bull. Chem. Soc. Jpn., 2025, 98, uoae134 CrossRef CAS.
- (a) J. Zhao, T. Wu, Q. He, H. Fang, M. Liu, Y. Liu, Z. Zhou, J. Zhang and W. Yang, Bull. Chem. Soc. Jpn., 2024, 97, uoae015 CrossRef CAS; (b) B. Gao, F. Bi, Z. Zhou, Y. Zhang, J. Wei, X. Lv, B. Liu, Y. Huanga and X. Zhang, Chem. Commun., 2024, 60, 7455–7458 Search PubMed; (c) K. Adpakpang, P. Ponchai, L. Pukdeejorhor, K. Faungnawakij and S. Bureekaew, Bull. Chem. Soc. Jpn., 2024, 97, uoae016 Search PubMed; (d) T. Kitao, Bull. Chem. Soc. Jpn., 2024, 97, uoae103 CrossRef CAS; (e) K. Ariga, M. Rezki, K. Suzuki-Nagata, T. Mikawa and S. Tsujimura, Chem. Commun., 2025, 61, 12309–12312 Search PubMed.
- (a) D. Jiang, X. Xu, Y. Bando, S. M. Alshehri, M. Eguchi, T. Asahi and Y. Yamauchi, Bull. Chem. Soc. Jpn., 2024, 97, uoae074 CrossRef CAS; (b) R. Nakatani, T. Irie, S. Das, Q. Fang and Y. Negishi, ACS Appl. Mater. Interfaces, 2025, 17, 24701–24729 Search PubMed; (c) J. Fu, J.-Y. Kang, W. Gao, Z.-W. Huang, L.-Q. Kong, K. Xie, Q.-H. Zhu, G.-H. Zhang, G.-H. Tao and L. He, Chem. Commun., 2025, 61, 2235–2256 Search PubMed; (d) T. Irie, S. Das, Q. Fang and Y. Negishi, J. Am. Chem. Soc., 2025, 147, 1367–1380 Search PubMed; (e) R. Xu, X. Xu, A. Alowasheeir, Y. Yamauchi and D. Jiang, Bull. Chem. Soc. Jpn., 2025, 98, uoaf053 CrossRef CAS.
- (a) A. Alowasheeir, M. Eguchi, Y. Fujita, K. Tsuchiya, R. Wakabayashi, T. Kimura, K. Ariga, K. Hatano, N. Fukumitsu and Y. Yamauchi, Bull. Chem. Soc. Jpn., 2024, 97, uoae099 Search PubMed; (b) Y. Ishizaki-Betchaku, S. Yamamoto, S. Nagano and M. Mitsuishi, Jpn. J. Appl. Phys., 2025, 64, 030801 CrossRef; (c) M. Han, T. Nagaura, J. Kim, S. M. Alshehri, T. Ahamad, Y. Bando, A. Alowasheeir, Y. Asakura and Y. Yamauchi, Bull. Chem. Soc. Jpn., 2025, 98, uoae136 Search PubMed; (d) T. Hikino, H. Mochizuki, T. Matsuno, K. Kuroda and A. Shimojima, Chem. Commun., 2025, 61, 7337–7340 RSC; (e) P. Kowalczyk, Alexander V. Neimark, Sylwester Furmaniak, Artur P. Terzyk and K. Kaneko,, Carbon, 2025, 245, 120737 Search PubMed.
- (a) O. Oki, H. Yamagishi, Y. Morisaki, R. Inoue, K. Ogawa, N. Miki, Y. Norikane, H. Sato and Y. Yamamoto, Science, 2022, 377, 673–678 Search PubMed; (b) Y. Hirotsu, R. Sekiguchi, Y. Takeoka, M. Rikukawa and M. Yoshizawa-Fujita, Bull. Chem. Soc. Jpn., 2024, 97, uoae101 Search PubMed; (c) J. Koga, Y. Chiashi, A. Nakamura, T. Akiba, H. Takahashi, T. Shimojima, S. Ishiwata and K. Ishizaka, Appl. Phys. Express, 2024, 17, 042007 Search PubMed; (d) K. Ikeda, T. Matsuo, K. Yano and S. Hayashi, Bull. Chem. Soc. Jpn., 2024, 97, uoae115 CrossRef CAS; (e) K. Bulgarevich and K. Takimiya, CrystEngComm, 2025, 27, 4776–4786 RSC.
- (a) T. Seki, Bull. Chem. Soc. Jpn., 2024, 97, bcsj.20230219 Search PubMed; (b) T. Hayashi and G. V. Latag, ACS Appl. Nano Mater., 2025, 8, 8570–8587 CrossRef CAS; (c) M. Zharnikov, Y. Shoji and T. Fukushima, Acc. Chem. Res., 2025, 58, 312–324 Search PubMed; (d) T. Tsuji, Y. Machida, S. De Feyter, K. Tahara and Y. Tobe, J. Phys. Chem. C, 2025, 129, 7894–7902 CrossRef CAS; (e) O. Vu, J. Song, H. Sugimoto and M. Fujii, ACS Appl. Nano Mater., 2025, 8, 14802–14810 CrossRef CAS.
- (a) R. Makiura,, Coord. Chem. Rev., 2022, 469, 214650 CrossRef; (b) K. Ariga, Chem. Mater., 2023, 35, 5233–5254 Search PubMed; (c) R. Terui, Y. Otsuki, Y. Shibasaki and A. Fujimori, Bull. Chem. Soc. Jpn., 2024, 97, uoae050 CrossRef CAS; (d) J. Song, A. Jancik-Prochazkova, K. Kawakami and K. Ariga, Chem. Sci., 2024, 15, 18715–18750 RSC; (e) Y. Shioda, I. Fujii and A. Fujimori, Langmuir, 2025, 41, 9959–9966 CrossRef CAS PubMed.
- (a) G. Decher, Science, 1997, 277, 1232–1237 CrossRef CAS; (b) K. Ariga, J. P. Hill and Q. Ji, Phys. Chem. Chem. Phys., 2007, 9, 2319–2340 Search PubMed; (c) J. J. Richardson, M. Björnmalm and F. Caruso,, Science, 2015, 348, aaa2491 CrossRef PubMed; (d) H. Tsunematsu, K. Nishibashi, E. Yamamoto, M. Kobayashi, T. Yoshida and M. Osada, Appl. Phys. Express, 2024, 17, 015002 Search PubMed; (e) K. Ariga, J. Song and K. Kawakami, Chem. Commun., 2024, 60, 2152–2167 RSC.
- K. Ariga, Nanoscale Horiz., 2021, 6, 364–378 Search PubMed.
- (a) K. Ariga, Q. Ji, W. Nakanishi, J. P. Hill and M. Aono, Mater. Horiz., 2015, 2, 406–413 RSC; (b) L. Cao, Y. Huang, B. Parakhonskiy and A. G. Skirtach, Nanoscale, 2022, 14, 15964–16002 RSC; (c) G. Chen, T. Koide, J. Nakamura and K. Ariga, Small Methods, 2025, 9, 2500069 Search PubMed; (d) H.-X. Wang, X. Zhu and M. Liu, Chem. Soc. Rev., 2025, 54, 5516–5550 RSC.
- (a) K. Ariga, J. Li, J. Fei, Q. Ji and J. P. Hill, Adv. Mater., 2016, 28, 1251–1286 CrossRef CAS PubMed; (b) A. Jancik-Prochazkova and K. Ariga, ACS Sustainable Chem. Eng., 2025, 13, 6900–6917 CrossRef CAS; (c) M. Sharma, A. Kumar, D. Sajwan, K. Kumari, B. P. Mishra and V. Krishnan, Adv. Sustainable Syst., 2025, 9, 2400903 CrossRef CAS; (d) Y. Song, W. Zhu, C. Wang, Z. Li, R. Xin, D. Wu, X. Ma, H. Wang, X. Wang, S. Li, J. Kim, Q. Sun, M. Kim and Y. Yamauch, Adv. Mater., 2025, 37, 2504364 CrossRef CAS PubMed; (e) S. Chintalapati and E. Miyako, Carbon, 2025, 238, 120252 CrossRef CAS.
- (a) R. P. Feynman, Eng. Sci., 1960, 23, 32–36 Search PubMed; (b) M. Roukes, Sci. Am., 2001, 285, 48–51 CrossRef CAS PubMed.
- K. Ariga, J. Song and K. Kawakami, Chem. – Asian J., 2025, e00836 CrossRef CAS PubMed.
- R. B. Laughlin and D. Pines, Proc. Natl. Acad. Sci. U. S. A., 2000, 97, 28–31 CrossRef CAS PubMed.
- (a) K. Ariga and R. Fakhrullin, Bull. Chem. Soc. Jpn., 2022, 95, 774–795 CrossRef CAS; (b) K. Ariga, Bull. Chem. Soc. Jpn., 2024, 97, uoad001 CrossRef CAS.
- (a) P. Jordan, P. Fromme, H. T. Witt, O. Klukas, W. Saenger and N. Krauß, Nature, 2001, 411, 909–917 CrossRef CAS PubMed; (b) K. N. Ferreira, T. M. Iverson, K. Maghlaoui, J. Barber and S. Iwata, Science, 2004, 303, 1831–1838 CrossRef CAS PubMed.
- (a) K. Ariga, Mater. Chem. Front., 2017, 1, 208–211 RSC; (b) K. Ariga, X. Jia, J. Song, J. P. Hill, D. T. Leong, Y. Jia and J. Li, Angew. Chem., Int. Ed., 2020, 59, 15424 CrossRef CAS PubMed; (c) K. Ariga and Y. Yamauchi, Chem. – Asian J., 2020, 15, 718–728 CrossRef CAS PubMed.
- (a) L. Li, Y. Zhang, Y. Wu, Z. Wang, W. Cui, C. Zhang, J. Wang, Y. Liu and P. Yang, Adv. Funct. Mater., 2024, 34, 2315509 CrossRef CAS; (b) H. Hagiwara, E. Fujii, T. Iwabuchi and T. Takahashi, Bull. Chem. Soc. Jpn., 2024, 97, uoae019 CrossRef CAS; (c) C. Fu, Z. Wang, X. Zhou, B. Hu, C. Li and P. Yang, Chem. Soc. Rev., 2024, 53, 1514–1551 RSC; (d) M. Minamide, M. Tsurushima, R. Koga, K. Hasegawa, Y. Kurosawa, T. Tsuchida and S. Goto, Bull. Chem. Soc. Jpn., 2024, 97, uoae058 CrossRef CAS.
- (a) K. Ariga and T. Kunitake, Acc. Chem. Res., 1998, 31, 371–378 CrossRef CAS; (b) K. Ariga, Phys. Chem. Chem. Phys., 2020, 22, 24856–24869 RSC; (c) O. N. Oliveira Jr., L. Caseli and K. Ariga, Chem. Rev., 2022, 122, 6459–6513 CrossRef CAS PubMed.
- (a) F. J. Pavinatto, L. Caseli and O. N. Oliveira Jr., Biomacromolecules, 2010, 11, 1897–1908 CrossRef CAS PubMed; (b) Y. Yamada, K. Xu and A. Fujimori, ChemistrySelect, 2023, 8, e202302340 CrossRef CAS.
- (a) A. P. Girard-Egrot, S. Godoy and L. J. Blum, Adv. Colloid Interface Sci., 2005, 116, 205–225 CrossRef CAS PubMed; (b) G. Nerath, D. A. Oliveira, J. R. Siqueira Jr and L. Caseli, ACS Appl. Mater. Interfaces, 2025, 17, 13018–13028 CrossRef CAS PubMed.
- (a) Y. Okahata, T. Tsuruta, K. Ijiro and K. Ariga, Langmuir, 1988, 4, 1373–1375 CrossRef CAS; (b) Y. Okahata, T. Tsuruta, K. Ijiro and K. Ariga, Thin Solid Films, 1989, 180, 65–72 CrossRef CAS.
- B. Miao, Y. Wang, Y. Liu, G. Da, Y. Wang, X. Chen and Z. Shao, Adv. Funct. Mater., 2024, 34, 2312324 CrossRef CAS.
- E. Chatzigiannakis, J. Yang, L. M. C. Sagis and C. V. Nikiforidis, J. Colloid Interface Sci., 2025, 683, 408–419 CrossRef CAS PubMed.
- E. B. A. Hinderink, L. Sagis, K. Schroën and C. C. Berton-Carabin, Colloids Surf., B, 2020, 192, 111015 CrossRef CAS PubMed.
- Y. Wang, T. Wang, Q. Chen, W. Zhou and J. Guo, Mol. Pharm., 2023, 20, 2536–2544 CrossRef CAS PubMed.
- K. G. Pham, B. R. Thompson, T. Wang, S. Samaddar, K. K. Qian, Y. Liu and N. J. Wagner, J. Phys. Chem. B, 2023, 127, 9724–9733 CrossRef CAS PubMed.
- Y. S. Tein, Z. Zhang and N. J. Wagner, Langmuir, 2020, 36, 7814–7823 Search PubMed.
- C. V. Wood, V. I. Razinkov, W. Qi, C. J. Roberts, J. Vermant and E. M. Furst, Langmuir, 2023, 39, 7775–7782 CrossRef CAS PubMed.
- E. L. N. Escobar, M. C. Vaclaw, J. T. Lozenski and P. Dhar, Langmuir, 2024, 40, 4789–4800 CrossRef CAS PubMed.
- H. Fan, B. Wang, Y. Zhang, Y. Zhu, B. Song, H. Xu, Y. Zhai, M. Qiao and F. Sun, Nat. Commun., 2021, 12, 7257 CrossRef CAS PubMed.
- (a) Z. Tang, Y. Wang, P. Podsiadlo and N. Kotov, Adv. Mater., 2006, 18, 3203–3224 CrossRef CAS; (b) G. Rydzek, Q. Ji, M. Li, P. Schaaf, J. P. Hill, F. Boulmedais and K. Ariga,, Nano Today, 2015, 10, 138–167 CrossRef CAS; (c) K. Ariga, Y. Lvov and G. Decher, Phys. Chem. Chem. Phys., 2022, 24, 4097–4115 RSC; (d) K. Ariga, Materials, 2025, 18, 654 CrossRef CAS PubMed.
- (a) Y. Lvov, K. Ariga and T. Kunitake, Chem. Lett., 1994, 2323–2326 CrossRef CAS; (b) Y. Lvov, K. Ariga, I. Ichinose and T. Kunitake, J. Am. Chem. Soc., 1995, 117, 6117–6123 CrossRef CAS.
- (a) Y. M. Lvov, Z. Lu, J. B. Schenkman, X. Zu and J. F. Rusling, J. Am. Chem. Soc., 1998, 120, 4073–4080 CrossRef CAS; (b) C. M. Jewell and D. M. Lynn, Adv. Drug Delivery Rev., 2008, 60, 979–999 CrossRef CAS PubMed.
- (a) L. Richert, P. Lavalle, E. Payan, X. Z. Shu, G. D. Prestwich, J.-F. Stoltz, P. Schaaf, J.-C. Voegel and C. Picart, Langmuir, 2004, 20, 448–458 Search PubMed; (b) C. F. V. Sousa, L. P. G. Monteiro, J. M. M. Rodrigues, J. Borges and J. F. Mano, J. Mater. Chem. B, 2023, 11, 6671–6684 Search PubMed.
- (a) K. Katagiri, R. Hamasaki, K. Ariga and J. Kikuchi, J. Am. Chem. Soc., 2002, 124, 7892–7893 CrossRef CAS PubMed; (b) K. Katagiri, R. Hamasaki, K. Ariga and J. Kikuchi, Langmuir, 2002, 18, 6709–6711 CrossRef CAS.
- (a) W. B. Stockton and M. F. Rubner, Macromolecules, 1997, 30, 2717–2725 CrossRef CAS; (b) B.-S. Kim, S. W. Park and P. T. Hammond, ACS Nano, 2008, 2, 386–392 CrossRef CAS PubMed.
- (a) H. Lee, L. J. Kepley, H. G. Hong and T. E. Mallouk, J. Am. Chem. Soc., 1988, 110, 618–620 CrossRef CAS; (b) M. Haga, K. Kobayashi and K. Terada,, Coord. Chem. Rev., 2007, 251, 2688–2701 CrossRef CAS.
- (a) M. Li, S. Ishihara, M. Akada, M. Liao, L. Sang, J. P. Hill, V. Krishnan, Y. Ma and K. Ariga, J. Am. Chem. Soc., 2011, 133, 7348–7351 Search PubMed; (b) G. Rydzek, T. G. Terentyeva, A. Pakdel, D. Golberg, J. P. Hill and K. Ariga, ACS Nano, 2014, 8, 5240–5248 CrossRef CAS PubMed.
- A. Ikeda, T. Hatano, S. Shinkai, T. Akiyama and S. Yamada, J. Am. Chem. Soc., 2001, 123, 4855–4856 Search PubMed.
- (a) Y. Shimazaki, M. Mitsuishi, S. Ito and M. Yamamoto, Langmuir, 1997, 13, 1385–1387 CrossRef CAS; (b) Y. Shimazaki, M. Mitsuishi, S. Ito and M. Yamamoto, Langmuir, 1998, 14, 2768–2773 CrossRef CAS.
- (a) T. Serizawa, K. Hamada, T. Kitayama, N. Fujimoto, K. Hatada and M. Akashi, J. Am. Chem. Soc., 2000, 122, 1891–1899 CrossRef CAS; (b) T. Serizawa, K. Hamada and M. Akashi, Nature, 2004, 429, 52–55 CrossRef CAS PubMed.
- (a) Y. Lvov, K. Ariga, I. Ichinose and T. Kunitake, J. Chem. Soc., Chem. Commun., 1995, 2313–2314 RSC; (b) Y. Lvov, K. Ariga, I. Ichinose and T. Kunitake, Thin Solid Films, 1996, 284, 797–801 CrossRef.
- (a) J. Anzai, H. Takeshita, Y. Kobayashi, T. Osa and T. Hoshi, Anal. Chem., 1998, 70, 811–817 CrossRef CAS PubMed; (b) J. Anzai, Y. Kobayashi, N. Nakamura, M. Nishimura and T. Hoshi, Langmuir, 1999, 15, 221–226 CrossRef CAS.
- M. Onda, Y. Lvov, K. Ariga and T. Kunitake,, J. Ferment. Bioeng., 1996, 82, 502–506 CrossRef CAS.
- (a) H. Ai, S. A. Jones and Y. M. Lvov, Cell Biochem. Biophys., 2003, 39, 23–43 CrossRef CAS PubMed; (b) M. H. Iqbal, H. Kerdjoudj and F. Boulmedais, Chem. Sci., 2024, 15, 9408–9437 RSC; (c) B. H. Vilsinski, A. C. de Oliveira, P. R. Souza and A. F. Martins, Prog. Org. Coat., 2024, 196, 108720 Search PubMed.
- Y. Zhang, Y. Liu, Y. Liu, P. Zuo, S. Miao, B. Hu, Y. Kang, W. Liu, Q. Yang, H. Ren and P. Yang, J. Am. Chem. Soc., 2023, 145, 17125–17135 CrossRef CAS PubMed.
- Y. Zhang, H. Ren, C. Linghu, J. Zhang, A. Gao, H. Su, S. Miao, R. Qin, B. Hu, X. Chen, M. Deng, Y. Liu and P. Yang, Adv. Mater., 2024, 36, 2412378 CrossRef CAS PubMed.
- S. J. Roeters, T. W. Golbek, M. Bregnhøj, T. Drace, S. Alamdari, W. Roseboom, G. Kramer, T. Šantl-Temkiv, K. Finster, J. Pfaendtner, S. Woutersen, T. Boesen and T. Weidner, Nat. Commun., 2021, 12, 1183 CrossRef CAS PubMed.
- (a) G. Decher, B. Lehr, K. Lowack, Y. Lvov and J. Schmitt,, Biosens. Bioelectron., 1994, 9, 677–684 CrossRef CAS; (b) G. Delechiave, M. O. Silva, M. F. de Castro-Amarante, D. E. M. Camarena, A. A. Venceslau-Carvalho, L. C. S. Ferreira and L. H. Catalani, ACS Appl. Nano Mater., 2024, 7, 4068–4077 CrossRef CAS.
- (a) K. Ariga, T. Nakanishi and T. Michinobu, J. Nanosci. Nanotechnol., 2006, 6, 2278–2301 CrossRef CAS PubMed; (b) M. Zaborowska-Mazurkiewicz, N. Kraśkiewicz, P. Sekuła and R. Bilewicz, J. Phys. Chem. B, 2025, 129, 8742–8753 CrossRef CAS PubMed.
- (a) J. Salafsky, J. T. Groves and S. G. Boxer, Biochemistry, 1996, 35, 14773–14781 CrossRef CAS PubMed; (b) K. Glasmästar, C. Larsson, F. Höök and B. Kasemo,, J. Colloid Interface Sci., 2002, 246, 40–47 CrossRef PubMed.
- (a) Y. Shin and C. P. Brangwynne, Science, 2017, 357, eaaf4382 CrossRef PubMed; (b) S. Alberti, A. Gladfelter and T. Mittag,, Cell, 2019, 176, 419–434 CrossRef CAS PubMed; (c) J. Song, K. Kawakami and K. Ariga, Adv. Colloid Interface Sci., 2025, 339, 103420 Search PubMed.
- (a) K. Minami, T. Mori, W. Nakanishi, N. Shigi, J. Nakanishi, J. P. Hill, M. Komiyama and K. Ariga, ACS Appl. Mater. Interfaces, 2017, 9, 30553–30560 CrossRef CAS PubMed; (b) X. Jia, J. Song, W. Lv, J. P. Hill, J. Nakanishi and K. Ariga, Nat. Commun., 2022, 13, 3110 CrossRef CAS PubMed.
- D. Zare, J. R. Allison and K. M. McGrath, Biomacromolecules, 2016, 17, 1572–1581 CrossRef CAS PubMed.
- J. Bergfreund, P. Bertsch, S. Kuster and P. Fischer, Langmuir, 2018, 34, 4929–4936 CrossRef CAS PubMed.
- J. Zhai, T. J. Wooster, S. V. Hoffmann, T.-H. Lee, M. A. Augustin and M.-I. Aguilar, Langmuir, 2011, 27, 9227–9236 CrossRef CAS PubMed.
- J. Bergfreund, M. Diener, T. Geue, N. Nussbaum, N. Kummer, P. Bertsch, G. Nyström and P. Fischer, Soft Matter, 2021, 17, 1692–1700 RSC.
- A. Chrysanthou, M. Bosch-Fortea, C. Nadal, A. Zarbakhsh and J. E. Gautrot, J. Colloid Interface Sci., 2024, 674, 379–391 Search PubMed.
- (a) L. Peng and J. E. Gautrot, Mater. Today Bio, 2021, 12, 100159 CrossRef CAS PubMed; (b) T. Ueki, K. Uto, S. Yamamoto, R. Tamate, Y. Kamiyama, X. Jia, H. Noguchi, K. Minami, K. Ariga, H. Wang and J. Nakanishi, Adv. Mater., 2024, 36, 2310105 CrossRef CAS PubMed.
- A. Chrysanthou, H. Kanso, W. Zhong, L. Shang and J. E. Gautrot, ACS Appl. Mater. Interfaces, 2023, 15, 2760–2770 CrossRef CAS PubMed.
- W. Gong, K. Jiang, T. Yang, Y. Liang, J. Tu, J. Li, F. Liu, S. Ou and B. Zhu, ACS Nano, 2025, 19, 15596–15616 Search PubMed.
- (a) S. Rekhi, C. G. Garcia, M. Barai, A. Rizuan, B. S. Schuster, K. L. Kiick and J. Mittal, Nat. Chem., 2024, 16, 1113–1124 CrossRef CAS PubMed; (b) I. B. A. Smokers, B. S. Visser, A. D. Slootbeek, W. T. S. Huck and E. Spruijt, Acc. Chem. Res., 2024, 57, 1885–1895 CrossRef CAS PubMed; (c) P. Wen, H. Huang, R. Zhang, H. Zheng, T. Liang, C. Zhuang, Q. Wu, J. Wang, F. Liu, K. Zhang, W. Wu, K. He, F. Liu, H. Li and Z. Gu, Nat. Chem., 2025, 17, 279–288 Search PubMed.
- U. A. A. Abbas, M. Naz, I. Sargin and G. Arslan, Langmuir, 2025, 41, 6843–6849 Search PubMed.
- M. Peydayesh, S. Kistler, J. Zhou, V. Lutz-Bueno, F. D. Victorelli, A. B. Meneguin, L. Spósito, T. M. Bauab, M. Chorilli and R. Mezzenga, Nat. Commun., 2023, 14, 1848 CrossRef CAS PubMed.
- (a) X. Jia, K. Minami, K. Uto, A. C. Chang, J. P. Hill, T. Ueki, J. Nakanishi and K. Ariga, Small, 2019, 15, 1804640 CrossRef PubMed; (b) X. Jia, K. Minami, K. Uto, A. C. Chang, J. P. Hill, J. Nakanishi and K. Ariga, Adv. Mater., 2020, 32, 1905942 CrossRef CAS PubMed; (c) J. Song, X. Jia and K. Ariga,, Mater. Today Bio, 2020, 8, 100075 CrossRef PubMed; (d) X. Jia, J. Song, W. Lv, J. P. Hill, J. Nakanishi and K. Ariga, Nat. Commun., 2022, 13, 3110 Search PubMed.
- (a) J. Song, X. Jia, K. Minami, J. P. Hill, J. Nakanishi, L. K. Shrestha and K. Ariga, ACS Appl. Nano Mater., 2020, 3, 6497–6506 Search PubMed; (b) J. Song, X. Jia and K. Ariga, Small Methods, 2020, 4, 2000500 Search PubMed.
- (a) M. Ishii, Y. Yamashita, S. Watanabe, K. Ariga and J. Takeya, Nature, 2023, 622, 285–291 Search PubMed; (b) T. Ohashi, M. Ishii, J. Takeya, K. Ariga and Y. Yamashita, Small, 2025, 21, e09278 CrossRef CAS PubMed.
- I. Ishmukhametov, S. Batasheva, S. Konnova, Y. Lvov and R. Fakhrullin,, Appl. Clay Sci., 2025, 270, 107775 CrossRef CAS.
- (a) D. J. McClements,, Curr. Opin. Colloid Interface Sci., 2004, 9, 305–313 Search PubMed; (b) Y. Liu, Q. Wu, J. Zhang, W. Yan and X. Mao,, Int. J. Biol. Macromol., 2024, 261, 129795 CrossRef CAS PubMed; (c) W. Shu, W. Shi, H. Xie, S. Wang, Q. Zhang, K. Ouyang, F. Xiao and Q. Zhao,, Food Chem., 2025, 479, 143732 Search PubMed.
- (a) W. Yospanya, A. Matsumura, Y. Imasato, T. Itou, Y. Aoki, H. Nakazawa, T. Matsui, T. Yokoyama, M. Ui, M. Umetsu, S. Nagatoishi, K. Tsumoto, Y. Tanaka and K. Kinbara, Bull. Chem. Soc. Jpn., 2024, 97, uoae090 Search PubMed; (b) Y. Dai, Z. Zhang, D. Wang, T. Li, Y. Ren, J. Chen and L. Feng, Adv. Mater., 2024, 36, 2310455 CrossRef CAS PubMed; (c) K. Nishio, K. Shibata and T. Mizoguchi, Appl. Phys. Express, 2024, 17, 037002 Search PubMed; (d) T. Sadakane, K. Nakata, K. Suda and D. Yokogawa, Bull. Chem. Soc. Jpn., 2024, 97, uoae121 Search PubMed; (e) Y. Zhan, X. Ren, S. Zhao and Z. Guo,, J. Power Sources, 2025, 629, 235992 CrossRef CAS.
- (a) W. Chaikittisilp, Y. Yamauchi and K. Ariga, Adv. Mater., 2022, 34, 2107212 CrossRef CAS PubMed; (b) L. R. Oviedo, V. R. Oviedo, M. O. Martins, S. B. Fagan and W. L. da Silva, J. Nanopart. Res., 2022, 24, 157 CrossRef; (c) B. V. Parakhonskiy, J. Song and A. G. Skirtach, Adv. Colloid Interface Sci., 2025, 343, 103546 CrossRef CAS PubMed; (d) H. Bae, H. Ji, K. Konstantinov, R. Sluyter, K. Ariga, Y. H. Kim and J. H. Kim, Adv. Mater., 2025, e10239 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |