Decoding fold robustness and thermostability in Thermococcus AMP phosphorylase and its DPBB domains
Khushboo
Bhagat
 and
Aditya K.
Padhi
and
Aditya K.
Padhi
 *
*
Laboratory for Computational Biology & Biomolecular Design, School of Biochemical Engineering, Indian Institute of Technology (BHU) Varanasi, Varanasi-221005, Uttar Pradesh, India. E-mail: aditya.bce@iitbhu.ac.in
First published on 10th December 2025
Abstract
We dissect thermostability and fold resilience in Thermococcus AMP phosphorylase and its DPBB domains using multiscale simulations, energetic profiling, and rational redesign. Comparative structural-dynamics analyses across homologs uncover adaptation-linked paradigms integrating sequence divergence, rugged thermodynamics, and fold plasticity, highlighting evolutionary strategies driving thermostability and functional robustness under extreme physicochemical conditions.
The emergence of complex modern proteins traces back to ancient prototype folds that served as evolutionary building blocks.1 Among these, fold architectures such as the Rossmann, ferredoxin, and the double-psi β-barrel (DPBB) represent a structurally intricate and evolutionarily resilient domain present in key enzymes like RNA polymerases, β-secretases, and AMP phosphorylases.2–7
AMP phosphorylase (EC 2.4.2.57), prominently characterized in the hyperthermophilic archaeon Thermococcus kodakarensis (T. kodakarensis), catalyzes the reversible phosphorolysis of AMP and other nucleotides, operating efficiently above 80 °C (353 K). The enzyme comprises a tripartite domain organization, with its N-terminal DPBB domain contributing significantly to thermostability through a six-stranded β-barrel structure flanked by α-helices and ψ loops.8 Despite its prominence, the molecular underpinnings of thermostability within the DPBB domain and full-length enzyme environment across hyperthermophilic genera remain largely unresolved. To address this gap, we present a multiscale framework linking evolutionary signals to energetic landscapes, enabling analysis and rational redesign of Thermococcus AMP phosphorylase and its DPBB domain to elucidate stability and structure-dynamics coupling.
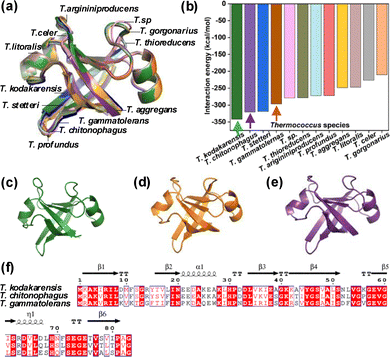
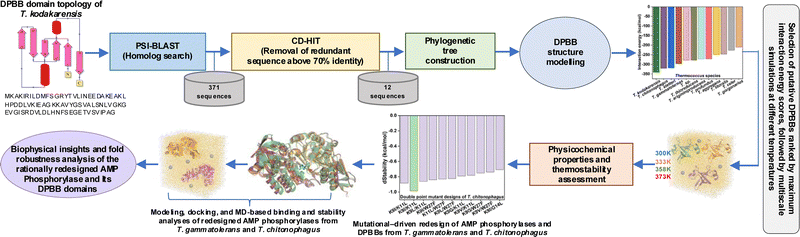
The DPBB domain sequence from T. kodakarensis AMP phosphorylase (UniProt: Q5JCX3) was used to identify homologs via PSI-BLAST.9 This resulted in 371 homologous DPBB sequences across archaeal genera and 93 sequences from the Thermococcus genus. CD-HIT clustering at 70% identity yielded 12 non-redundant homolog sequences.10 Next, Maximum Likelihood guided phylogenetic analysis was carried out, followed by AlphaFold3-based structural modeling and validation via Ramachandran analysis since structures of the DPBB domains were not experimentally resolved (Fig. S1). Structural superposition revealed a conserved DPBB fold, highlighting evolutionary preservation across Thermococcus species (Fig. 1a).
Residue–residue interaction energies of the DPBB domains, calculated using INTAA identified T. kodakarensis, T. gammatolerans, and T. chitonophagus as having the highest intramolecular interaction energies (Fig. 1b).11 Representative structures of which are illustrated in Fig. 1c–e. Here, T. chitonophagus and T. gammatolerans represent Thermococcus chitonophagus and Thermococcus gammatolerans, respectively.
Multiple sequence alignment revealed that T. chitonophagus and T. gammatolerans shared 72% and 57% sequence identity with T. kodakarensis, respectively. Conserved residues were primarily located in the core β-strands and α-helices known to support DPBB structural stability (Fig. 1f). Phylogenetic analysis underscored evolutionary divergence within the domain, with T. chitonophagus exhibiting the closest relationship to T. kodakarensis (divergence value = 0.33), followed by T. gammatolerans (0.541) (Fig. S2). Considering the sequences with the highest IEs from Thermococcus species and sequence diversity, (T. kodakarensis, T. chitonophagus, and T. gammatolerans) were further aligned using Clustal Omega, followed by visual refinement with ESPript v3.0 to highlight conserved motifs and secondary structure elements.12,13
Next, to assess conformational flexibility and stability of the DPBB domains, we performed multiscale molecular dynamics (MD) simulations—both coarse-grained MD (CGMD) and all-atom MD (AAMD)—across four different temperatures (300 K, 333K, 358 K, 373 K), totalling 18 µs of sampling. These temperatures were chosen based on prior experimental studies that evaluated thermostability of AMP phosphorylase in T. kodakarensis, where the enzymes were incubated at 25–85 °C to assess heat-induced aggregation and solubility retention.8 The CGMD simulations demonstrated that T. kodakarensis DPBB exhibited the lowest RMSD across all temperatures compared to T. gammatolerans and T. chitonophagus (Fig. S3). AAMD simulations reinforced these observations with finer resolution, showing T. kodakarensis maintained superior stability at 300 K, 333 K, 358 K, and 373 K (Fig. S4a, d, g and j). Next, per-residue root mean square fluctuation (RMSF) profiles revealed reduced atomic flexibility for T. kodakarensis. At 300 K, its average RMSF was 0.25 nm (CGMD) and 0.11 nm (AAMD), compared to higher values for the other species (Fig. S3b and S4b). This trend persisted at elevated temperatures (Fig. S3e, h, k and S4e, h, k). RMSF analysis indicated higher flexibility in T. gammatolerans and T. chitonophagus due to sequence-specific substitutions in loops and secondary structures. Finally, Radius of gyration (Rg) analysis representing compactness of the systems confirmed T. kodakarensis remained more compact across 300–373 K (Fig. S3c, f, i, l and S4c, f, i, l). Next, Principal component analysis (PCA) and free energy landscapes (FELs) were used to extract collective motions and energy minima from the simulations. The PCA and FEL analyses confirmed the trends in global motions and stability. Covariance matrix trace values from CGMD at 300 K showed T. kodakarensis displayed lower conformational flexibility (15.39 nm2) than T. gammatolerans (18.37 nm2) and T. chitonophagus (17.11 nm2) (Fig. S5a–c). With increasing temperature, trace values rose but remained lowest for T. kodakarensis at 333 K (20.82 nm2), 358 K (22.96 nm2), and 373 K (26.33 nm2) (Fig. S5d–i and Fig. 2a–c). FELs revealed temperature-dependent conformational states, with T. kodakarensis occupying deeper energy minima (Fig. S6). AAMD simulations showed parallel trends (Fig. S7a–l).
Solvent accessible surface area (SASA) assessment revealed T. kodakarensis consistently exhibited lower SASA profiles under thermal stress (Fig. 2d). Hydrogen bond analysis showed T. kodakarensis maintained the highest average count at all temperatures (Fig. 2e), while salt bridge analysis revealed consistently higher electrostatic interactions (Fig. 2f). Secondary structure analysis demonstrated remarkable thermal resilience in T. kodakarensis, maintaining stable β-sheet content (∼43%), consistent α-helix content (∼8%), and modest 3-helix representation (4–5%). In contrast, T. gammatolerans showed declining β-sheet content, and T. chitonophagus exhibited the highest disruption (Table S1).
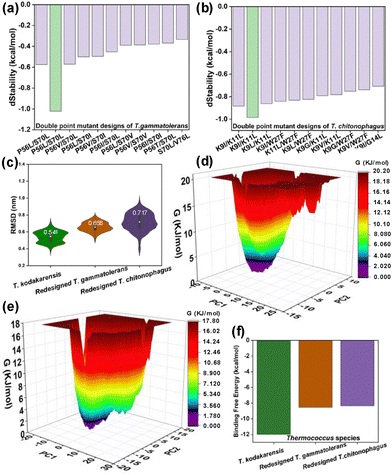
Comparative mutational profiling using MAESTROweb identified destabilizing residues in T. gammatolerans and T. chitonophagus, which were rationally redesigned for both isolated DPBB domains and domains embedded in the full-length AMP phosphorylase (Fig. S8).14 Next, the Residue Scan approach was used to identify potential stabilizing mutations, and mutants were evaluated in terms of improved relative dStability.15 Systematic in silico mutagenesis generated 76 single-point and 2400 double-point mutants for T. gammatolerans, and 152 single-point and 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 double-point mutants for T. chitonophagus. Mutational hotspots are shown in blue/red (Fig. S9a and b), with arrows denoting predicted stabilizing mutants. From these, 10 double-point mutants of T. gammatolerans (Fig. 3a) and 5 single (Fig. S10) and 10 double-point mutants of T. chitonophagus (Fig. 3b) displayed enhanced predicted stability. Notably, the corresponding full-length enzyme mutants—highlighted in green in Fig. 3a and b, also showed high stabilization potential, indicating that domain-level improvements can, in part, be propagated to the full-length enzyme. Based on dStability scores and subsequent analyses, the double mutants P56L/S70L (T. gammatolerans) and K9I/K11L (T. chitonophagus) were prioritized for further analyses.
200 double-point mutants for T. chitonophagus. Mutational hotspots are shown in blue/red (Fig. S9a and b), with arrows denoting predicted stabilizing mutants. From these, 10 double-point mutants of T. gammatolerans (Fig. 3a) and 5 single (Fig. S10) and 10 double-point mutants of T. chitonophagus (Fig. 3b) displayed enhanced predicted stability. Notably, the corresponding full-length enzyme mutants—highlighted in green in Fig. 3a and b, also showed high stabilization potential, indicating that domain-level improvements can, in part, be propagated to the full-length enzyme. Based on dStability scores and subsequent analyses, the double mutants P56L/S70L (T. gammatolerans) and K9I/K11L (T. chitonophagus) were prioritized for further analyses.
In addition, stabilizing mutations identified in T. chitonophagus and T. gammatolerans using DDMut were transplanted into evolutionarily near and far homologs to evaluate their transferability.16 Computed ΔΔG scores (kcal mol−1) showed consistently positive values, thereby confirming their transferable stabilizing effects across evolutionary relatives (Table S2). Next, to evaluate the impact of identified stabilizing mutations on the full-length AMP phosphorylase—including its structural-functional integrity, contribution to interdomain interactions, and overall stabilization—we first performed molecular docking of the enzyme with its substrate AMP, followed by AAMD simulations of T. kodakarensis and the redesigned T. gammatolerans and T. chitonophagus mutants (3 µs total). A detailed methodology and respective analyses are provided in the SI. First, molecular docking results confirmed conserved AMP binding across all three species (Fig. S11), with 2D interaction maps showing key non-covalent interactions (Fig. S12a–c). AAMD simulations indicated relatively comparable global stability (RMSD, Fig. 3c), with redesigned T. gammatolerans AMP phosphorylase displaying the lowest RMSF and Rg values (Fig. S12d and e). PCA revealed that the redesigned T. gammatolerans and T. chitonophagus AMP phosphorylase mutants exhibited lower trace values compared to T. kodakarensis, indicating more restricted conformational sampling (Fig. 3d, e and Fig. S12f). Interdomain interaction analysis showed T. gammatolerans AMP phosphorylase had two hydrogen bonds in the static state, increasing to three during dynamics in its redesigned state (Fig. S12g and h), while T. chitonophagus AMP phosphorylase displayed two static and five hydrogen bonds in its dynamic state (Fig. S12i and j). Cation-π interactions were enriched, with T. gammatolerans AMP phosphorylase showing two static and three in its dynamic redesigned state (Fig. S12k and l), while T. chitonophagus AMP phosphorylase exhibited three and two, in its static and redesigned dynamic state, respectively (Fig. S12m and n). Ligand-protein interaction analysis revealed that both redesigned enzymes demonstrated higher or comparable frequencies of van der Waals and hydrogen-bond contacts compared to T. kodakarensis (Fig. S12o and p). Binding free energy calculations corroborated these observations, showing comparable energetics for redesigned enzymes (Fig. 3f, Table S3). These results signify that redesigned mutations enhance overall enzyme stability, maintain AMP binding and optimal molecular interactions.
Early studies on extremophiles from hydrothermal vents and acidic springs, such as Solfatara di Pozzuoli, established the compositional foundations of protein thermostability, revealing trends like enhanced salt-bridge density, proline enrichment, and compact hydrophobic cores that stabilize proteins under extreme conditions.17,18 These pioneering investigations laid the groundwork for understanding thermal adaptation; however, predicting stability directly from sequence alone remains difficult, and rational design approaches often fail to identify truly stabilizing mutations with high accuracy.19,20
Here, we developed an integrated, mechanism-based framework combining evolutionary mining, multiscale simulations, FEL analyses, and systematic mutagenesis. By examining DPBB domains both in isolation and within full-length AMP phosphorylase, we demonstrate how local domain stability propagates to overall enzyme robustness—a critical yet underexplored aspect of protein engineering. This framework yields experimentally testable hypotheses supported by quantified energetics and explicit structural coordinates, enhancing transparency and reproducibility. It advances thermostability research from descriptive observation to predictive design grounded in mechanistic sequence–structure–stability relationships. Our overall framework (Fig. 4) shows that T. kodakarensis possesses the most thermodynamically stable DPBB domain, exhibiting optimal structural metrics and conserved non-covalent networks across 300–373 K. Principal-component and FEL analyses reveal deeper minima and reduced conformational fluctuations compared with T. gammatolerans and T. chitonophagus. Guided by mutational mapping, rational redesign produced mutants with improved hydrogen-bonding, ionic, van der Waals, and π–π interactions. Despite the DPBB domains lying distal from the central and C-terminal regions (Fig. S13), these mutations propagate stabilizing effects throughout the enzyme, providing a structural–evolutionary–energetic blueprint for designing hyperstable, biotechnologically relevant enzymes.
While the present framework integrates sequence evolution, multiscale simulations, and rational redesign to elucidate DPBB and AMP phosphorylase thermostability, it remains primarily computational. Experimental validation—such as enzymatic activity assays, kinetic profiling, or stability assessment under physiological and industrial conditions—will be essential to confirm the predicted enhancements. Additionally, interdomain coupling and environmental effects (pH, ionic strength, crowding) could modulate the observed thermostability trends. These aspects represent key directions for extending the current findings.
This work was supported by the SERB (now ANRF), India (SRG/2023/000167), India. K. B. thanks the MHRD, GOI, for the research fellowship.
Conflicts of interest
The authors declare no conflict of interest.Data availability
All major analysis scripts, datasets, and structural models have been deposited in Figshare at https://figshare.com/s/ab70bf48d8aecc571ab4 and are also available within the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5cc05135e.References
- V. Alva, J. Söding and A. N. Lupas, eLife, 2015, 4, e09410 CrossRef PubMed.
- R. V. Eck and M. O. Dayhoff, Science, 1966, 152, 363–366 CrossRef CAS PubMed.
- D. Lang, R. Thoma, M. Henn-Sax, R. Sterner and M. Wilmanns, Science, 2000, 289, 1546–1550 CrossRef CAS PubMed.
- J. M. Thornton, C. A. Orengo, A. E. Todd and F. M. G. Pearl, J. Mol. Biol., 1999, 293, 333–342 CrossRef CAS PubMed.
- R. M. Castillo, K. Mizuguchi, V. Dhanaraj, A. Albert, T. L. Blundell and A. G. Murzin, Structure, 1999, 7, 227–236 CrossRef CAS PubMed.
- S. Yagi, A. K. Padhi, J. Vucinic, S. Barbe, T. Schiex, R. Nakagawa, D. Simoncini, K. Y. J. Zhang and S. Tagami, J. Am. Chem. Soc., 2021, 143, 15998–16006 CrossRef CAS PubMed.
- S. Westarp, F. Kaspar, P. Neubauer and A. Kurreck, Curr. Opin. Biotechnol., 2022, 78, 102829 CrossRef CAS.
- Y. Nishitani, R. Aono, A. Nakamura, T. Sato, H. Atomi, T. Imanaka and K. Miki, J. Mol. Biol., 2013, 425, 2709–2721 CrossRef CAS PubMed.
- S. Altschul, Nucleic Acids Res., 1997, 25, 3389–3402 CrossRef CAS.
- L. Fu, B. Niu, Z. Zhu, S. Wu and W. Li, Bioinformatics, 2012, 28, 3150–3152 CrossRef CAS PubMed.
- J. Galgonek, J. Vymětal, D. Jakubec and J. Vondrášek, Nucleic Acids Res., 2017, 45, W388–W392 CrossRef CAS PubMed.
- F. Sievers and D. G. Higgins, Protein Sci., 2018, 27, 135–145 CrossRef CAS PubMed.
- X. Robert and P. Gouet, Nucleic Acids Res., 2014, 42, W320–W324 CrossRef CAS PubMed.
- J. Laimer, J. Hiebl-Flach, D. Lengauer and P. Lackner, Bioinformatics, 2016, 32, 1414–1416 CrossRef CAS PubMed.
- A. K. Padhi and T. Tripathi, Briefings Funct. Genomics, 2023, 22, 195–203 CrossRef CAS PubMed.
- Y. Zhou, Q. Pan, D. E. V. Pires, C. H. M. Rodrigues and D. B. Ascher, Nucleic Acids Res., 2023, 51, W122–W128 CrossRef CAS PubMed.
- C. Vieille and G. J. Zeikus, Microbiol. Mol. Biol. Rev., 2001, 65, 1–43 CrossRef CAS PubMed.
- R. Sterner and W. Liebl, Crit. Rev. Biochem. Mol. Biol., 2001, 36, 39–106 CrossRef CAS PubMed.
- N. Tokuriki and D. S. Tawfik, Curr. Opin. Struct. Biol., 2009, 19, 596–604 CrossRef CAS.
- N. Berezovsky, K. B. Zeldovich and E. I. Shakhnovich, PLoS Comput. Biol., 2007, 3, e52 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |