Open Access Article
Open Access ArticleBackbone N-heteroatom substitution as a strategy to enhance peptide proteolytic stability
Avraz F. Anwar,
Natalia Cano-Sampaio and
Juan R. Del Valle *
*
Department of Chemistry & Biochemistry, University of Notre Dame, Notre Dame, IN 46556, USA. E-mail: jdelvalle@nd.edu
First published on 23rd April 2026
Abstract
Peptide-based ligands are well-suited to engage large biomolecular surfaces but are often limited by rapid proteolytic degradation in biological environments. Backbone modification offers a direct means to disrupt protease recognition while preserving side chain functionality; however, many established approaches impose conformational constraints that compromise biological activity. Here, we evaluate backbone N-amino and N-hydroxy substitution as a strategy for enhancing peptide proteolytic stability. Using a defined chymotrypsin substrate, we demonstrate that backbone N-amination confers pronounced, position-dependent protection when introduced at or adjacent to the scissile bond. Extending these findings to a β-sheet-forming antimicrobial peptide, we show that poly-N-amination dramatically enhances serum stability while preserving or enhancing conformation-dependent antibacterial activity. Together, these results expand the repertoire of peptide backbone modifications that mitigate proteolytic degradation while retaining the conformational and functional features required for the design of peptide- and protein-based biological probes.
Introduction
Peptides represent a chemically versatile platform for engaging expansive and shallow biomolecular surfaces that are difficult to target with small molecules.1,2 However, their use in cellular and in vivo studies is often undermined by rapid proteolytic degradation. Endogenous proteases efficiently process canonical α-peptide backbones, which generally adopt β-strand-like local conformation prior to enzymatic cleavage.3 The short lifetime of peptides in tissue and serum thus poses a major obstacle to their development as chemical probes and therapeutics.4,5Numerous chemical strategies have been developed to address the problem of peptide proteolysis, including main chain macrocyclization, the incorporation of noncanonical residues, and backbone amide substitution (Fig. 1).6–11 Amide N-methylation has emerged as an extensively studied modification, with previous work demonstrating strong positional effects on proteolytic susceptibility that arise from disruption of hydrogen-bonding interactions required for protease binding.8,12–14 However, N-methylation and related approaches can impose conformational constraints that compromise biological activity.15–17 This trade-off is particularly problematic for peptides whose function depends on extended backbone conformations that are predisposed to interact with proteases.3 There remains a need for additional backbone editing approaches that can inform the design of conformationally and proteolytically stable peptidomimetics.
Hydrazide and hydroxamate bonds differ fundamentally from amides in their electronic properties, hydrogen-bonding behavior, and conformational bias. Recent studies from our laboratory have shown that backbone N-amination and N-hydroxylation can stabilize β-sheet conformations through cooperative noncovalent interactions, suggesting that N-heteroatom substitution may offer a unique combination of conformational control and side chain preservation.18–21 Despite considerable insights into the folding preferences of N-heteroatom-substituted peptides, the impact of these backbone modifications on proteolytic stability has not been systematically explored.22
In this study, we employ a chymotrypsin substrate sequence to investigate how backbone N-amino and N-hydroxy substitutions influence proteolytic stability as a function of position relative to the scissile bond. We then apply these insights to a β-sheet-forming antimicrobial peptide, demonstrating that poly-N-amination confers exceptional serum stability while preserving or enhancing conformation-dependent antibacterial activity. Together, these results establish backbone N-heteroatom substitution as a chemically distinct, minimalist strategy for stabilizing bioactive peptides.
Results and discussion
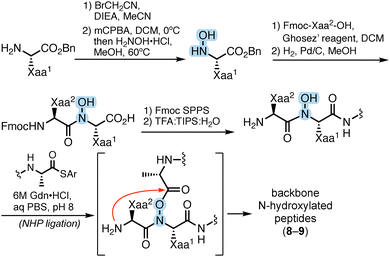
To assess the proteolytic susceptibility of peptides featuring backbone N-heteroatom substituents, we employed a sequence variant of a model originally reported by Horne and co-workers (1, Fig. 2A).8 Peptide 1 contains a single chymotrypsin cleavage site between Tyr (P1) and Lys (P1′) and lacks any appreciable secondary structure (Fig. S1). This design enables direct evaluation of how a single backbone modification at or near the scissile bond influences the rate of proteolysis, independent of conformational effects that could otherwise complicate interpretation.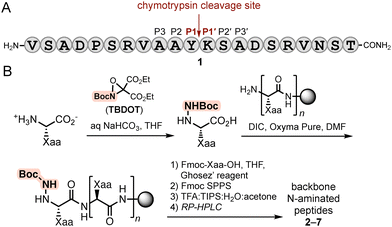 | ||
| Fig. 2 (A) Sequence and cleavage site of a model chymotrypsin substrate (1). (B) Synthesis of N-amino peptide variants of 1. | ||
We synthesized a series of N-aminated substrates (2–7) in which a single residue in positions P3 through P3′ was replaced with its corresponding α-hydrazino acid. The N-amino peptides (NAPs) were prepared using our previously reported monomer-based synthetic strategy (Fig. 2B).23 Briefly, zwitterionic amino acids were selectively aminated at the α-nitrogen using the oxaziridine reagent TBDOT24 to afford the corresponding α-hydrazino acids. These monomers were incorporated into the peptide sequence via standard Fmoc solid-phase peptide synthesis (SPPS), with subsequent Boc hydrazide acylation achieved through acid chloride-mediated coupling. Following elongation and cleavage from the resin, all peptides were purified by RP-HPLC and characterized by HRMS and analytical HPLC (Table 1).
| Peptide | P3 | P2 | P1 | P1′ | P2′ | P3′ | t1/2 (min) |
|---|---|---|---|---|---|---|---|
| 1 | Ala | Ala | Tyr | Lys | Ser | Ala | 2.0 |
| 2 | aAla | Ala | Tyr | Lys | Ser | Ala | 5.3 |
| 3 | Ala | aAla | Tyr | Lys | Ser | Ala | 3.5 |
| 4 | Ala | Ala | aTyr | Lys | Ser | Ala | 300 |
| 5 | Ala | Ala | Tyr | aLys | Ser | Ala | 202 |
| 6 | Ala | Ala | Tyr | Lys | aSer | Ala | 6.2 |
| 7 | Ala | Ala | Tyr | Lys | Ser | aAla | 2.5 |
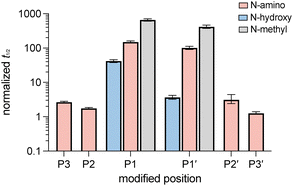
With peptides 1–7 in hand, their proteolytic stability was evaluated by incubation with chymotrypsin and quantification of intact peptide over time by analytical HPLC. For all peptides examined, data was fit to a single-phase exponential decay model to derive relative half-lives (Fig. 3 and Fig. S2). Notably, peptides 4 and 5, bearing N-amino substituents immediately preceding (P1) and at the scissile bond (P1′), respectively, exhibited half-lives that were 100–150-fold greater than that of the unmodified control peptide 1. In contrast, peptides 2, 3, 6, and 7, which feature N-amino modifications more distal to the cleavage site, exhibited half-lives within one order of magnitude of peptide 1. This positional dependence is consistent with a prior study showing that backbone N-methylation confers the greatest protection when introduced at amides that would otherwise engage chymotrypsin as H-bond donors (P3, P1, P2′).8 These results establish the protective effect of N-amination at or near the chymotrypsin cleavage site. This effect likely arises from a combination of altered substrate binding, reduced acyl–enzyme formation, and local conformational changes; however, the current data do not distinguish among these possibilities.
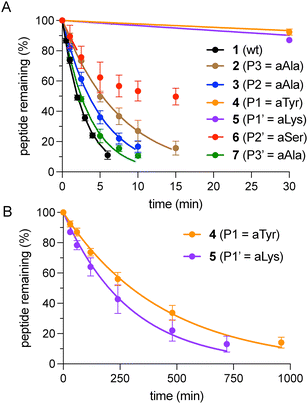 | ||
| Fig. 3 (A) Chymotrypsin-mediated degradation of 1–7 over 30 min, monitored by RP-HPLC (λ = 220 nm). Curves are derived from non-linear regression and error bars represent SD for each data point (n = 3). In the case of peptide 6, curve-fitting is based only on the initial 3 time points due to apparent chymotrypsin inactivation by the substrate.8 (B) Extended time scale showing degradation of 4 and 5. | ||
On the basis of these results, we sought to compare N-amination at P1 and P1′ directly to N-hydroxylation and N-methylation using the same parent sequence. Our group recently reported the synthesis of N-hydroxy peptides (NHPs) via chemoselective ligation of thioesters with peptide segments featuring a hydroxamate at the penultimate residue.25 We used this protocol to prepare 8 and 9, which incorporate N-hydroxy Tyr (hTyr) at P1 or N-hydroxy Lys (hLys) at P1′, respectively (Fig. 4). We also synthesized N-methyl peptides (NMPs) 10 and 11 by conventional SPPS for comparison (Table 2).
| Peptide | P3 | P2 | P1 | P1′ | P2′ | P3′ | t1/2 (min) |
|---|---|---|---|---|---|---|---|
| 8 | Ala | Ala | hTyr | Lys | Ser | Ala | 83 |
| 9 | Ala | Ala | Tyr | hLys | Ser | Ala | 7.3 |
| 10 | Ala | Ala | MeTyr | Lys | Ser | Ala | 1311 |
| 11 | Ala | Ala | Tyr | MeLys | Ser | Ala | 832 |
These peptides were evaluated using the same chymotrypsin stability assay described above (Fig. 5 and Fig. S2). Introduction of an N-hydroxy substituent at the P1 position (8) resulted in approximately 40-fold increase in proteolytic stability relative to 1. Although this enhancement was lower than that observed for the corresponding N-amino (4) and N-methyl (10) peptides, it nonetheless demonstrates that backbone N-hydroxylation at P1 can significantly slow proteolysis. In contrast, hydroxamate substitution at the P1′ position (9) resulted in only a modest increase in stability relative to 1, with proteolysis occurring significantly faster than N-amino variant 5 or N-methyl variant 11. Greater electrophilicity of the hydroxamate carbonyl at the scissile bond likely increases its susceptibility to hydrolysis relative to the hydrazide and tertiary amide variants, contributing to its weaker stabilization. Random coil CD signatures for each of the peptides tested (1–11) suggest that the observed stability trends are not the result of global conformational changes, although local perturbations, which are not distinguished in CD, may have an effect (Fig. S1).
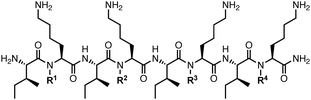
While backbone amide substitution represents a convenient and effective strategy to enhance resistance to proteolysis, the potential for significant conformational perturbation can limit such strategies in the context of bioactive peptides. We thus sought to investigate whether N-amination could enhance the stability of a peptide whose biological activity depends on conformation. Toward this end, we selected a known β-sheet-forming antimicrobial peptide (AMP) composed of four repeating Ile-Lys dipeptides (12, Table 3).26,27 Peptide 12 exhibits poor serum stability but possesses broad-spectrum antibacterial activity linked to its amphipathic β-sheet structure upon binding to the bacterial membrane. Our group previously showed that, in contrast to N-methylation, backbone N-amination of Cα-substituted amino acid residues can significantly increase the folded population of β-sheet model peptides.18,19 We therefore hypothesized that N-amination of 12 would impart resistance to proteolysis while maintaining or reinforcing conformation-dependent antibacterial activity.
We prepared a series of N-aminated and N-methylated octapeptides in which one to four Lys residues were replaced with aLys or MeLys (13–20, Table 3). The growth inhibitory activity of these peptides was evaluated against Escherichia coli (E. coli) and methicillin-resistant Staphylococcus epidermidis (MRSE), as representative Gram-negative and Gram-positive pathogens, respectively. Minimum inhibitory concentrations (MICs) were determined using a standard microbroth dilution assay. The unmodified parent peptide 12 exhibited MIC values of 50 µM against E. coli and 12.5 µM against MRSE. Peptides 13–15, containing one to three α-hydrazino acid substitutions, displayed antibacterial activities comparable to that of peptide 12, whereas tetra-N-aminated analogue 16 exhibited a two-fold enhancement in activity against both pathogens. Mono-NMP 17 exhibited similar antibacterial activity against both pathogens compared to the unmodified peptide 12, while analogues 18–20, containing two to four MeLys substitutions, exhibited significantly reduced activity. This result is consistent with a previous report showing that N-methylation of amphipathic β-sheet peptides significantly reduces their membrane lytic activity.28 Encouraged by the antibacterial activity of tetra-NAP 16, we synthesized a tetra-N-hydroxylated analogue (21) using a dipeptide building block approach. In contrast to 16, tetra-NHP 21 exhibited significantly weaker activity against both pathogens relative to 12.
Having identified 16 as the most potent inhibitor of bacterial growth, we evaluated its stability against trypsin and in human serum (37 °C) relative to the unmodified peptide 12. The fraction of intact peptide over time was monitored by analytical LC-MS. As expected, peptide 12 was rapidly degraded by trypsin within 20 minutes. Conversely, analogue 16 remained completely intact over 24 hours under identical conditions (Fig. 6A and Fig. S3A). Similarly, 12 underwent near complete degradation within 3 hours in human serum, whereas tetra-NAP 16 remained fully intact over the course of 400 hours as determined by LC-MS (Fig. 6B and Fig. S3B). This result demonstrates that poly-N-amination can dramatically enhance peptide serum stability, particularly when employed at every other backbone amide. Moreover, the incorporation of one or more non-canonical α-hydrazino acids into the parent octapeptide served to maintain or improve antibacterial activity in every case, unlike N-methylation, which substantially reduced activity in most cases.
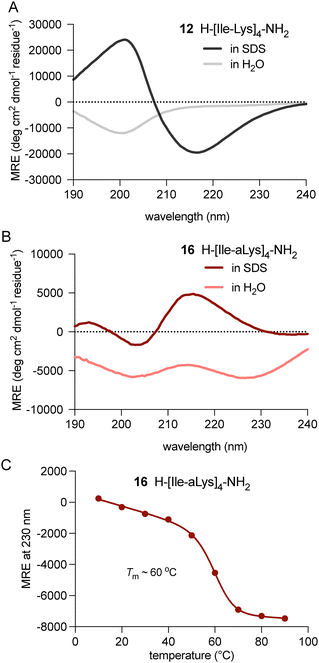
Finally, we assessed whether the antibacterial activity of peptide 16 is the result of altered secondary structure in a membrane environment. Circular dichroic (CD) spectra were thus obtained for peptides 12 and 16 in pure water and in the presence of 25 mM SDS as a membrane-mimicking surfactant. As previously reported, peptide 12 exhibited a random coil signature in water and a clear β-sheet structure in the presence of SDS, indicating environment-dependent folding (Fig. 7A).26 In contrast, 16 displayed anomalous CD signatures in pure water and aq SDS (Fig. 7B). These spectra likely arise from perturbed electronic transitions, owing to the presence of four hydrazide bonds within the strand.29 However, the clearly distinct signatures for 16 in water versus aq SDS are also indicative of environment-dependent adoption of secondary structure.
Interestingly, the CD spectrum of unmodified peptide 12 in aq SDS remained unchanged even at 90 °C, suggesting strong membrane association with a highly stable β-sheet-like structure (Fig. S4A). In contrast, thermal denaturation of peptide 16 in aq SDS revealed a well-defined isodichroic point indicative of cooperative unfolding (Fig. S4B). Fitting to a two-state denaturation model revealed a Tm of ∼60 °C (Fig. 7C).30 Although the precise conformation of 16 in aq SDS is unknown, our results indicate that tetra-N-amination provides complete protection from degradation in serum while maintaining the ability to adopt a biologically active fold.
Conclusions
Here, we systematically examined how backbone N-heteroatom substitution influences peptide proteolytic stability as a function of position relative to a defined cleavage site. N-Amination confers pronounced protection when introduced at the P1 and P1′ positions of the chymotrypsin substrate, consistent with disruption of interactions required for protease recognition. Direct comparison with N-hydroxylation and N-methylation reveals that these backbone modifications exert distinct effects on proteolytic susceptibility, possibly reflecting differences in the electronic properties and H-bonding capacities of the modified amide bonds. Notably, N-amination at the scissile bond (P1′) significantly enhances proteolytic stability despite the increased electrophilicity of the hydrazide carbonyl.The protective effects of N-amination were extended to an amphipathic antimicrobial octapeptide whose biological activity is linked to its β-sheet-like structure. Poly-N-amination dramatically enhances serum stability while improving antibacterial activity, whereas poly-N-methylation reduces membrane lytic activity. Circular dichroism studies indicate that tetra-N-amination preserves the ability of the peptide to adopt a membrane-associated folded state despite an unnatural residue content of 50%.
Together, these results establish backbone N-heteroatom substitution as a chemically distinct and minimally perturbing strategy for enhancing the proteolytic stability of bioactive peptides. In conjunction with their known conformational preferences, we anticipate that backbone N-amination and N-hydroxylation will find utility in the design of folded peptide and protein mimics for studies in cells and in vivo.
Author contributions
A. F. A. and N. C.-S. conducted the experimental work. A. F. A., N. C.-S., and J. R. D. conceptualized the project, analyzed the data, and prepared the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cb00061d.Acknowledgements
This work was supported by a grant from the National Science Foundation (CHE2109008). We gratefully acknowledge fellowship support to A. F. A. from NIH grant T32GM145773, the Berthiaume Institute for Precision Health, and the Welter Family Foundation. We thank Prof. Yichun Wang for access to CD instrumentation. We thank Prof. Christian Melander and Dr Ansley Nemeth for providing materials and bacteria used to determine antibacterial activity.Notes and references
- L. Nevola and E. Giralt, Chem. Commun., 2015, 51, 3302–3315 RSC.
- L.-G. Milroy, T. N. Grossmann, S. Hennig, L. Brunsveld and C. Ottmann, Chem. Rev., 2014, 114, 4695–4748 CrossRef CAS PubMed.
- J. D. A. Tyndall, T. Nall and D. P. Fairlie, Chem. Rev., 2005, 105, 973–999 CrossRef CAS PubMed.
- K. Fosgerau and T. Hoffmann, Drug Discovery Today, 2015, 20, 122–128 CrossRef CAS PubMed.
- X. Lai, J. Tang and M. E. H. ElSayed, Expert Opin. Drug Discovery, 2021, 16, 1467–1482 CrossRef CAS PubMed.
- L. Gentilucci, R. De Marco and L. Cerisoli, Curr. Pharm. Des., 2010, 16, 3185–3203 CrossRef CAS PubMed.
- A. Bhat, L. R. Roberts and J. J. Dwyer, Eur. J. Med. Chem., 2015, 94, 471–479 CrossRef CAS PubMed.
- H. M. Werner, C. C. Cabalteja and W. S. Horne, ChemBioChem, 2016, 17, 712–718 CrossRef CAS PubMed.
- M. Erak, K. Bellmann-Sickert, S. Els-Heindl and A. G. Beck-Sickinger, Bioorg. Med. Chem., 2018, 26, 2759–2765 CrossRef CAS PubMed.
- O. Al Musaimi, L. Lombardi, D. R. Williams and F. Albericio, Pharmaceuticals, 2022, 15, 1283 CrossRef CAS PubMed.
- C. Lamers, Future Drug Discovery, 2022, 4(2), FDD75, DOI:10.4155/fdd-2022-0005.
- J. Chatterjee, C. Gilon, A. Hoffman and H. Kessler, Acc. Chem. Res., 2008, 41, 1331–1342 CrossRef CAS PubMed.
- S. V. Fiacco and R. W. Roberts, ChemBioChem, 2008, 9, 2200–2203 CrossRef CAS PubMed.
- J. Chatterjee, F. Rechenmacher and H. Kessler, Angew. Chem., Int. Ed., 2013, 52, 254–269 CrossRef CAS PubMed.
- Y. C. Koay, N. L. Richardson, S. S. Zaiter, J. Kho, S. Y. Nguyen, D. H. Tran, K. W. Lee, L. K. Buckton and S. R. McAlpine, ChemMedChem, 2016, 11, 881–892 CrossRef CAS PubMed.
- H. Ogawa, G. T. Burke, J. D. Chanley and P. G. Katsoyannis, Int. J. Pept. Protein Res., 1987, 30, 460–473 CrossRef CAS PubMed.
- S. Sagan, P. Karoyan, O. Lequin, G. Chassaing and S. Lavielle, Curr. Med. Chem., 2004, 11, 2799–2822 CrossRef CAS PubMed.
- M. P. Sarnowski, C. W. Kang, Y. M. Elbatrawi, L. Wojtas and J. R. Del Valle, Angew. Chem., Int. Ed., 2017, 56, 2083–2086 CrossRef CAS PubMed.
- M. P. Sarnowski, K. P. Pedretty, N. Giddings, H. L. Woodcock and J. R. Del Valle, Bioorg. Med. Chem., 2018, 26, 1162–1166 CrossRef CAS PubMed.
- M. P. Sarnowski and J. R. Del Valle, Org. Biomol. Chem., 2020, 18, 3690–3696 RSC.
- S. K. Starnes, W. S. Horne and J. R. Del Valle, J. Org. Chem., 2025, 90, 17214–17220 CrossRef CAS PubMed.
- I. J. Angera, M. M. Wright and J. R. Del Valle, Acc. Chem. Res., 2024, 57, 1287–1297 CrossRef CAS PubMed.
- A. F. Anwar and J. R. Del Valle, J. Org. Chem., 2025, 90, 6084–6089 CrossRef CAS PubMed.
- A. Armstrong, L. H. Jones, J. D. Knight and R. D. Kelsey, Org. Lett., 2005, 7, 713–716 CrossRef CAS PubMed.
- N. Cano-Sampaio and J. R. Del Valle, Chem. – Eur. J., 2026, e70850 CrossRef PubMed.
- Z. Y. Ong, S. J. Gao and Y. Y. Yang, Adv. Funct. Mater., 2013, 23, 3682–3692 CrossRef CAS.
- Z. Y. Ong, J. Cheng, Y. Huang, K. Xu, Z. Ji, W. Fan and Y. Y. Yang, Biomaterials, 2014, 35, 1315–1325 CrossRef CAS PubMed.
- S. E. Miller, K. Tsuji, R. P. M. Abrams, T. R. Burke Jr and J. P. Schneider, J. Am. Chem. Soc., 2020, 142, 19950–19955 CrossRef CAS PubMed.
- M. M. Wright, B. H. Rajewski, T. A. Gerrein, Z. Xu, L. J. Smith, W. Seth Horne and J. R. Del Valle, Commun. Chem., 2025, 8, 76 CrossRef CAS PubMed.
- D. Shortle, A. K. Meeker and E. Freire, Biochemistry, 1988, 27, 4761–4768 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |