Open Access Article
Open Access ArticleMutagenesis-based evolution enables rapid retargeting of L-aptamers for structured RNA recognition
Xuan
Han
 a,
Tyler
Guilbault
a and
Jonathan T.
Sczepanski
a,
Tyler
Guilbault
a and
Jonathan T.
Sczepanski
 *ab
*ab
aDepartment of Chemistry, Texas A&M University, College Station, Texas 77843, USA. E-mail: jon.sczepanski@chem.tamu.edu
bDepartment of Biochemistry and Biophysics, Texas A&M University, College Station, Texas 77843, USA
First published on 25th March 2026
Abstract
Mirror-image L-(deoxy)ribose nucleic acid aptamers (L-aptamers) enable high-affinity, selective recognition of native structured RNAs via shape-based, cross-chiral interactions rather than Watson–Crick base pairing. However, achieving this level of specificity has typically required de novo selection from fully randomized libraries for each new target. Here, we report the first application of mutagenesis-based evolution to an existing cross-chiral L-aptamer to rapidly reprogram target specificity. Starting from a parental aptamer recognizing the HIV-1 TAR RNA hairpin, we generated two evolved aptamers that, together with the parent, form a set of orthogonal binders capable of discriminating RNA hairpins differing by a single nucleotide. Deep sequencing revealed that retargeting requires extensive remodeling of the aptamer fold rather than incremental sequence changes, reflecting the dependence of cross-chiral interactions on global RNA architecture rather than local sequence complementarity. Finally, RNA pulldown experiments demonstrate that these orthogonal L-aptamers retain strict selectivity in a competitive and structurally complex context, including extended flanking sequences and multiple RNA targets. Taken together, this work establishes mutagenesis-based evolution as a versatile strategy for rapidly retargeting cross-chiral L-aptamers and reinforces the unique advantages of structure-based RNA recognition, providing a framework for developing adaptable and multiplexable RNA-targeting strategies.
Introduction
Due to its single-stranded nature, RNA can form complex three-dimensional structures driven by intramolecular base pairing interactions.1 These structures, which include helices, loops, and bulges, facilitate interactions with proteins and other RNAs, enabling participation in critical regulatory and gene expression processes. Recent research has increasingly demonstrated relationships between structured RNAs and diverse disease states, including cancer,2–4 viral infection,5–7 neurological disorders,8–10 and immune responses.11,12 These discoveries have made structured RNAs attractive targets for both diagnostic and therapeutic applications.13–15Traditionally, RNA is targeted using hybridization-based technologies, such as antisense oligonucleotides (ASOs) or small interfering RNAs (siRNAs); however, the inaccessibility of sequences within pre-existing structures presents significant challenges to these methods.16 Structural-based approaches and helper strands can be implemented to increase potency, but these strategies still do not fully overcome barriers posed by more stable RNA structures. Small molecules have emerged as a promising platform for targeting RNA structures. In recent years, small molecules targeting distinct structural elements with specific sequences have progressed from laboratory research to FDA-approved drugs.17 Nevertheless, the discovery of RNA-binding small molecules with high affinity and structural specificity remains challenging due to RNA's negatively charged backbone, inherent flexibility, and structural redundancy. Limited structural data for nucleic acid further limits the structure-based approach using small molecules.15
A promising alternative is the use of aptamers—nucleic acid sequences selected to bind target ligands with high specificity and affinity. The process used to generate aptamers, systematic evolution of ligands by exponential enrichment (SELEX),18 allows for the discovery of aptamers against targets ranging from single metal ions19 to whole cells.20,21 Notably, work from our group and others22–29 has shown that aptamers composed of L-DNA or L-RNA, which are the synthetic enantiomers of natural D-nucleotides, can be evolved to bind tightly to RNA structures, and may be particularly well suited for this purpose. Although D- and L-oligonucleotides share identical physical and chemical properties, they cannot form contiguous Watson–Crick base pairs with one another. As a result, so-called “cross-chiral” interactions between D- and L-oligonucleotides must occur independently of primary sequence. This unique feature is leveraged during in vitro selection to evolve L-aptamers that adaptively bind structured D-RNA targets through tertiary, shape-driven interactions rather than sequence complementarity.30 We have demonstrated that cross-chiral recognition occurs with low-nanomolar affinity and high selectivity.22–26,31 Notably, L-aptamers can distinguish their cognate D-RNA targets from those with single-nucleotide substitutions or small structural perturbations.22,24,30 Combined with the inherent nuclease resistance of L-oligonucleotides,32 these properties position L-aptamers as versatile and powerful reagents for targeting structured RNA.
Mutagenesis-based aptamer selection is a strategy used to evolve the target specificity of an existing aptamer by deliberately introducing mutations into its sequence and then rescreening the resulting library. This differs from the standard SELEX process, which starts with a fully random library. The mutagenesis-based approach allows for fine-tuning of an aptamer's binding properties to discriminate between closely related targets or to evolve new specificity entirely.33,34 For example, Huang et al. used this approach to alter an ATP aptamer to bind GTP, with aptamers binding GTP emerging after five rounds of selection from a mutagenized library against a GTP analogue.33 Given the high selectivity of cross-chiral interactions between L-aptamers and RNA, along with their unique structure-based mode of recognition, we were curious whether mutagenesis-based selection could be used to evolve L-aptamers with altered substrate specificity. Although mutagenesis-based selection has not previously been attempted with cross-chiral aptamers, such a strategy could greatly expand the toolkit of mirror-image technologies. Not only would this provide a convenient and rapid strategy for generating L-aptamers against new RNA targets, but the results may also reveal important insights into the relationship between specificity and structure, aiding in the design of future cross-chiral L-aptamers.
In this study, we introduce mutagenesis-based SELEX as a rapid strategy for retargeting cross-chiral L-aptamers to new structured RNA targets. We show that closely related RNA structures previously disfavored by the parental aptamer can be selectively recognized without restarting selection from fully randomized libraries. Deep sequencing reveals that altered target specificity arises from extensive remodeling of the aptamer fold rather than incremental sequence adjustments, highlighting the structure-driven nature of cross-chiral recognition. Enrichment analysis further demonstrates the rapid emergence of orthogonal, highly specific L-aptamer–RNA pairs, providing a foundation for scalable and multiplexable RNA-targeting strategies.
Results and discussion
Library and target design
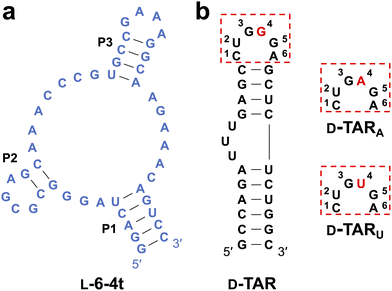
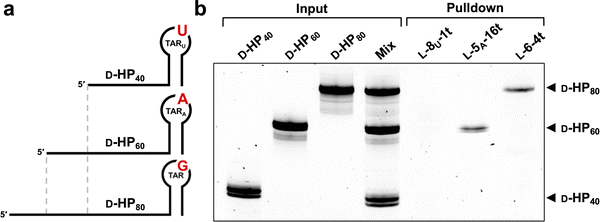
The aptamer chosen for this study was L-6-4t (Fig. 1a), which binds tightly (Kd = 100 ± 30 nM) to the trans-activation response element RNA (D-TAR) from Human Immunodeficiency Virus type 1 (HIV-1), shown in Fig. 1b.23 Previous in-line hydrolysis probing showed that L-6-4t binds D-TAR through its 6-nucleotide (nt) distal loop.23 Importantly, using a library of TAR hairpins containing all possible 6-nucleotide loops, we previously showed that L-6-4t binds with high selectively to the 5′-(C/U)UGGG(A/G)-3′ loop motif (Fig. 1b).25 This strict sequence requirement provided an ideal opportunity to test whether we could evolve altered target specificity. Therefore, we chose to incorporate either a transition mutation (D-TARA) or transversion mutation (D-TARU) at position G4 within the invariable 6-nt loop domain (Fig. 1b). Because the distal loop constitutes the primary aptamer recognition interface, mutations within the stem region were not expected to directly impact binding specificity and were therefore not examined in this study.23,31 We hypothesized that the overall structural change for a transition mutation would be less significant than for a transversion and, thus, would require fewer compensatory mutations within the aptamer to accommodate. We prepared L-RNA versions of these hairpins (L-TARA and L-TARU) in which the target loop sequences were grafted onto a minimum stem scaffold to reinforce loop-based interaction (Fig. S1a and S2). As expected, an electrophoretic mobility shift assay (EMSA) revealed that D-6-4t was unable to bind the mutated L-TAR targets (Fig. S1c).We next prepared a library of D-6-4t aptamers (D-Lib14) such that the original random sequence domain was mutagenized to a degeneracy of 14% at each position (Fig. S1b), yielding a population carrying an average of 6 mutation per member (see Methods and Fig. S3a and b). The binomial distribution strongly favors low mutation counts; consequently, unique sequences with fewer mutations are more abundant in the pool. We expected that favoring a lower number of mutations would drive repurposing of the existing structural motif rather than major structural reorganization. To facilitate the SELEX process, the primer binding site sequences used for isolating the parent 6-4t aptamer were appended to the ends of the mutagenized library (Fig. S1b). As with the parent aptamer, D-Lib14 did not bind to either of the mutated L-TAR hairpins (Fig. S1c).
Mutagenesis-based SELEX alters the substrate specificity of cross-chiral interaction
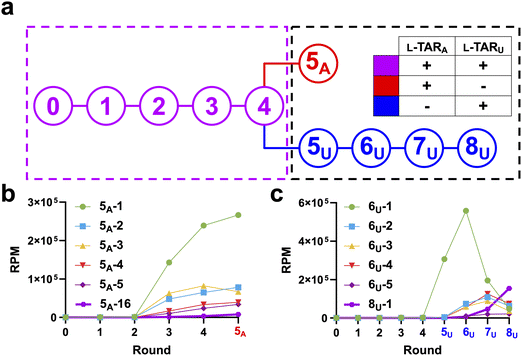
Similar to the isolation of the parent aptamer L-6-4t, we carried out cross-chiral mutagenesis-based SELEX using the “selection reflection” technique.35–37 In this process, a D-nucleic acid library (i.e., D-Lib14) is selected against the enantiomer of the target ligand (i.e., L-TARA or L-TARU), which facilitates enzymatic amplification of the library after the selection step. Once the SELEX process is complete and lead D-aptamers are identified, they can be chemically synthesized in the L-form and used to bind the intended D-RNA ligand.35,37 Accordingly, D-Lib14 was incubated with 3′-biotinylated L-TARA or L-TARU (L-TARA-BIO and L-TARU-BIO, respectively; Fig. S1a and Table S1) at room temperature in the presence of 50 mM NaCl, 10 mM MgCl2, 0.1% (v/v) Tween-20, and Tris (pH = 7.6). Sequences bound to a biotinylated hairpin target were immobilized on magnetic streptavidin beads. The beads were then washed thoroughly with the same buffer and bound sequences were eluted by denaturation with NaOH. Eluted sequences were reverse transcribed and amplified by PCR. The resulting dsDNA libraries were subjected to in vitro transcription to generate the corresponding ssRNA library to be used in the next round of SELEX.During the first four rounds, selection of D-Lib14 was performed against L-TARA and L-TARU simultaneously to enhance SELEX efficiency. After round 4, an EMSA revealed that the enriched library had substantial affinity for L-TARA, with no detectable binding to L-TARU (Fig. S4a). To avoid further skewing the library away from the L-TARU target, all subsequent rounds were carried out using individual targets (referred to as “deconvolution” rounds) (Fig. 2a). Because the library was already highly enriched for L-TARA-binding sequences after round 4, only one round (round 5A) of deconvolution was performed to further enrich sequences that bound to L-TARA. However, L-TARU required four additional deconvolution rounds (rounds 5U through 8U, 8 in total) before appreciable binding was observed by EMSA (Fig. S4b). Nevertheless, these results indicated that we had successfully obtained cross-chiral aptamers against both mutant targets.
High-throughput sequencing analysis
The amplified eluates isolated from each SELEX round were subjected to next-generation sequencing (NGS), resulting in over 105 processed reads per round. This high-level sequence analysis enabled quantitative tracking of sequence enrichment throughout the SELEX process. Initially, read per million (RPM) values were used to rank the relative abundance of each individual sequence emerging from a round. After verifying the sequence distribution of the round 0 library (Fig. S3a–c), we first considered the top five most abundant sequences observed after the single deconvolution round for target L-TARA (Round 5A). These sequences, referred to as 5A-1 through 5A-5, constituted nearly half (48.5%) of the total reads found in the final pool while being considerably diverse (Fig. 2b and Fig. S5a). In addition, four out of these five sequences were further enriched in deconvolution round 5A, which is in agreement with the observed prominent L-TARA affinity of the overall pool after the initial four rounds. By tracking the abundance of these sequences through each round, we also found that the top three sequences by RPM in round 5A (5A-1, 5A-2, and 5A-3) were already highly enriched by round 3, differing from round 5A only in their relative ranking (Fig. 2b and Fig. S6a). Not only did this result show that the population converged rapidly, but it also indicated that new cross-chiral aptamers with altered selectivity could be effectively identified within as few as three rounds of mutagenesis-based SELEX for this target.We performed the same analysis with the L-TARU target. The top-5 most enriched sequences in round 8U, referred to as 8U-1 through 8U-5, demonstrated the same dominance in relative abundance, accounting for 48.8% of the total reads (Fig. S5c). Interestingly, the most abundant sequences in the final round (round 8U) were also highly enriched in round 6U before plateauing or declining in later rounds (Fig. 2c and Fig. S6b, c). In fact, many of the TARU-binding sequences that became enriched early on during the deconvolution process (e.g., by round 6U) decreased in abundance by round 8U. A plausible explanation is that sequences appearing early in the deconvolution process were simply more competitive against L-TARA-binding sequences during the mixed-pool phase of selection, even if they were not the highest-affinity L-TARU binders. If correct, we would expect sequences becoming more enriched during later rounds to have higher affinity towards L-TARU.
The library's unique bias towards L-TARA in early rounds makes it an interesting case for studying specificity of cross-chiral interactions. During the deconvolution rounds for L-TARU, the absence of the L-TARA target resulted in a drastic and rapid decrease in the abundance of the most enriched L-TARA-binding sequences (Fig. S7). This observation suggested that these sequences are highly selective for L-TARA, being able to distinguish the single nucleotide difference between the two hairpin targets. Unfortunately, the comparable analysis was not possible for L-TARU binding sequences because the most abundant sequences in rounds 6U and 8U were not detectable by sequencing during the earlier rounds (Fig. S6b and c). However, comparison of the top enriched sequences from round 5A and 8U revealed low sequence similarities (Fig. S5a and c), adding more evidence for the specificity of emerged sequences. The selectivity of specific sequences is examined later in the study.
Comparative sequence analysis between mutant aptamers and parent L-6-4t
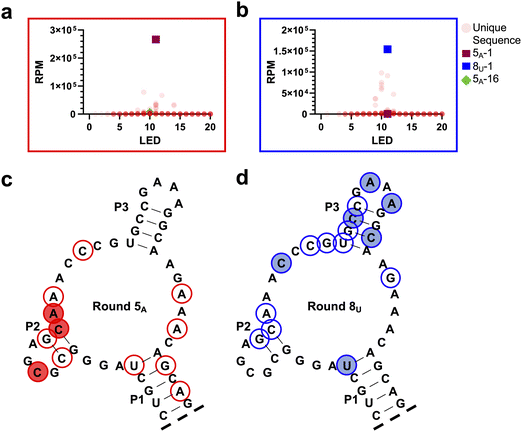
The mutagenized library D-Lib14 was designed to favor a low mutation rate (Fig. S3) due to our initial hypothesis that specific compensatory mutations would be able to accommodate the single-base mutation on the target hairpin. Therefore, we determined the Levenshtein edit distance (LED) values for each sequence compared to the parent aptamer L-6-4t. LED values measure the minimum number of single-nucleotide edits (insertions, deletions, or mutations) needed to transform one sequence into another. As deletions and insertions are only introduced through synthetic impurities and enzymatic errors, this value can be approximated to mutations in this work.Scatter plots of individual sequences with RPM versus LED allows clear visualization of changes in mutation numbers among emerging sequences (Fig. 3a, b and Fig. S8). As predicted via our sequence distribution model, the abundance of each unique sequence is in a negative relationship to its mutation number, as seen in the starting library and enriched round 1 pool (Fig. S3a and b). However, starting at round 2, sequences with a high number of mutations (>10) started to emerge from the population, which continued through rounds 3 and 4 (Fig. S8). In the deconvolution round for L-TARA, round 5A, sequences with 11 mutations were the most abundant, accounting for 49.9% of the total population (Fig. 3a and Fig. S9a). This included 5A-1, the most abundant sequence overall for binding to L-TARA (Fig. S5a). A similar trend was observed during the deconvolution rounds for L-TARU (Fig. S8). During this process, sequences with different LED values were rapidly enriched, while remaining distributed around 11. In the terminal round for L-TARU, round 8U, sequences with LED values of 10 and 11 were most abundant, attributing to 40% and 34% of the population, respectively (Fig. 3b and Fig. S9b). The most abundant sequence in round 8U, 8U-1, has an LED of 11 (Fig. S5c).
We next examined how mutations accumulated along the parental 6-4t aptamer sequence during selection. Although comparison of the most abundant sequences from key deconvolution rounds revealed clear differences between selections (Fig. S5), this approach captures only a limited snapshot of the evolving population. To obtain a more global view of mutational patterns, we generated consensus sequences for the round 5A and round 8U pools using the top 1000 most abundant sequences from each round, together accounting for over 90% of total sequencing reads. For each position, the consensus nucleotide was assigned as the base with the highest cumulative RPM, provided its frequency exceeded that of the second most abundant base by at least twofold; positions failing to meet this criterion were designated as ambiguous. This analysis revealed a concentration of mutation in or around the P2 stem-loop within the round 5A population (Fig. 3c). The P2 stem–loop was previously shown to be essential for binding of the parent aptamer L-6-4t to D-TAR, indicating that this region forms key contacts with the TAR hairpin. Accordingly, the single-base mutation in L-TARA appears to have necessitated substantial reorganization of the P2 domain to preserve high-affinity binding. In contrast, far fewer mutations accumulated near P2 in the round 8U population (Fig. 3d), despite the corresponding position also being mutated in the L-TARU target. Instead, mutations in round 8U were more concentrated around the P3 stem-loop, a region that in the parent aptamer serves primarily as a structural element and is not thought to engage TAR directly. This shift suggests that the selection process recruited or remodeled the P3 domain to create a new functional interface capable of accommodating the transversion mutation present in L-TARU. Notably, these divergent mutational solutions highlight how selections against L-TARA and L-TARU, which differ by only a single nucleotide, yielded aptamer families with markedly different sequence architectures.
Characterization of individual aptamers
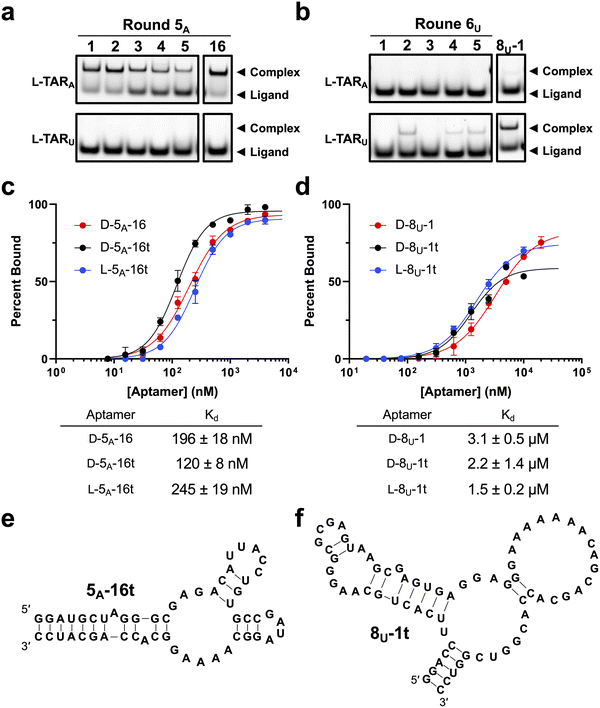
To identify lead aptamer candidates for each target, we screened the most abundant sequences emerging from the respective deconvolution rounds using EMSA (Fig. 4a and b). All sequences tested from round 5A (D-5A-1 through D-5A-5) bound tightly to L-TARA, with a clear correlation between sequence abundance and binding affinity. Importantly, no cross-reactivity with L-TARU was observed (Fig. 4a), confirming the target selectivity inferred from sequencing analyses (Fig. S7).We next evaluated sequences enriched during selection against L-TARU. Candidates from round 6U (D-6U-1–D-6U-5), together with a representative sequence from round 8U (D-8U-1), were examined by EMSA (Fig. 4b). Sequences were intentionally chosen from multiple rounds to test the hypothesis that continued selection leads to improved binding performance. Because the most abundant sequences in round 8U were highly similar (Fig. S5c), only D-8U-1 was selected for screening. Consistent with this rationale, although D-6U-1 through D-6U-5 displayed strong selectivity for L-TARU, their binding affinities were weaker than that of D-8U-1 (Fig. 4b).
In addition to final sequence abundance, we also examined enrichment dynamics across selection rounds to identify candidates that expanded most rapidly during selection. This analysis highlighted sequence 5A-16, which exhibited the largest fold enrichment between rounds 1 and 5A despite remaining at substantially lower abundance in the round 5A pool relative to other top-ranking sequences (Fig. S5a). Assuming a conservative baseline corresponding to a singleton in round 1, 5A-16 increased by at least ∼1200-fold by round 5A; because this sequence was not detected until round 3, its true enrichment is likely substantially greater. Notably, 5A-16 enriched 7.7-fold between rounds 3 and 5A, representing the largest enrichment over this interval among the top-ranking sequences. In contrast, the most abundant sequence, 5A-1, exhibited a smaller (∼380-fold) enrichment over rounds 1 to 5A. Despite its lower final abundance, D-5A-16 appeared to bind slightly more tightly to L-TARA than 5A-1 (Fig. 4a). While this observation is based on a single sequence and is limited by sequencing depth, it nonetheless supports prior conclusions that enrichment dynamics, rather than final abundance alone, provide a more reliable indicator of functional aptamer performance.38,39
Based on binding performance, subsequent analyses focused on sequences 5A-16 and 8U-1 (Fig. S5a and c). In vitro transcribed clone D-5A-16 bound L-TARA with a dissociation constant (Kd) of 196 ± 18 nM, whereas clone D-8U-1 bound L-TARU with substantially weaker affinity (Kd = 3.1 ± 0.5 µM; Fig. 4c and d). Although both aptamers exhibited reduced affinity relative to the parental L-6-4t aptamer binding to D-TAR (Kd = 100 ± 30 nM),23 the loss in binding strength was far more pronounced for D-8U-1, which displayed a >30-fold increase in Kd. This pronounced difference likely reflects the distinct structural consequences of the target mutations. The transversion mutation present in L-TARU is expected to introduce a larger perturbation to the RNA hairpin geometry than the corresponding transition mutation in L-TARA, posing a greater challenge for shape-based, cross-chiral recognition. Consistent with this interpretation, sequences binding L-TARA dominated the early rounds of selection, suggesting that smaller structural deviations from the parental TAR hairpin are more readily accommodated. Notably, analysis of predicted secondary structures revealed that neither evolved aptamer retains appreciable similarity to the parental L-6-4t structure (Fig. S10a and e). Together, these findings suggest that even single-nucleotide changes in the target RNA can necessitate extensive remodeling of the aptamer fold, highlighting the complexity of cross-chiral, structure-based recognition and the need for coordinated, multi-site mutational networks to restore high-affinity binding.
Preparation of truncated L-RNA aptamers
We sought to remove non-essential nucleotides from 5A-16 and 8U-1, being 79-nt and 78-nt in length, to improve downstream synthesis and potentially increase affinity due to reduced steric effect.22–24 Rational truncation of both aptamers was performed based on predicted secondary structures (Fig. S10a–h). Both sequences tolerated only mild truncations of potentially non-functional duplex-forming sequence at both ends. The resulting truncations yielded sequences 5A-16t and 8U-1t (Fig. 4e and f). D-5A-16t and D-8U-1t bound to their corresponding L-RNA targets with Kd values of 120 ± 5 nM and 2.2 ± 1.4 µM, respectively (Fig. 4c and d). The affinity of both aptamers improved after truncation, similar to what has been observed with prior cross-chiral aptamers.22–24,26 Interestingly, aptamer D-8U-1t displayed self-inhibitory activity at higher concentration, which was not observed for the full-length sequence (Fig. 4d). We speculated that the truncation removed structural constraints present in the full-length aptamer, promoting formation of nonproductive structures (e.g., dimers) at higher concentrations.Both truncated aptamers were then prepared in the L-RNA form through solid-phase chemical synthesis using L-ribonucleoside phosphoramidites (Fig. S11). L-5A-16t bound to D-TARA with a Kd of 245 ± 19 nM, while L-8U-1t bound to D-TARU with a Kd of 1.5 ± 0.2 µM. Due to the reduced quality of synthetic L-RNA compared to enzymatically prepared D-RNA, it is not uncommon for L-aptamers to have slight increase in Kd values relative to their D-RNA counterparts.22–24,26 The apparent increase in affinity observed for L-8U-1t is likely an artifact of the self-inhibitory behavior that complicates accurate binding quantification. Notably, cross-reactivity was evaluated by EMSA across all aptamer–ligand combinations. No binding was observed to non-cognate targets, and L-aptamers bound exclusively to the corresponding D-RNA targets, with no detectable interaction with the L-RNA counterparts (Fig. S12).
Orthogonal recognition of structured RNA targets by cross-chiral aptamers
A central goal of this study was to generate aptamers that exhibit high specificity and orthogonality toward closely related structured RNA targets. To evaluate aptamer performance beyond EMSA, we employed an RNA pulldown experiment to directly assess target discrimination in a competitive and more structurally complex context. A mixture of three RNA transcripts, D-HP80, D-HP60, and D-HP40, was prepared, each containing a terminal hairpin structure with distal loop sequences corresponding to TAR, TARA, and TARU, respectively (Fig. 5a). These transcripts differed in overall length, allowing them to be readily distinguished by denaturing PAGE. 5′-Biotinylated L-aptamers (L-5A-16t, L-8U-1t, and L-6-4t) were individually immobilized on streptavidin-coated magnetic beads and incubated with the RNA mixture. Following washing, the captured RNA was analyzed by denaturing PAGE, with transcript identity resolved by length (Fig. 5b). The pulldown results were fully consistent with EMSA-based binding measurements. Both L-5A-16t and L-6-4t selectively recovered RNA transcripts containing their cognate target sequences, with no detectable capture of non-cognate RNAs. Under the stringency of this competitive assay, L-8U-1t did not recover measurable RNA, consistent with its weaker affinity for D-TARU determined by EMSA, and did not exhibit any nonspecific interactions. Importantly, this assay was performed in the presence of extended flanking sequences and direct competition among multiple structured RNA targets. The retention of strict target selectivity under these conditions highlights the robustness of the evolved aptamers and underscores their potential as highly specific and orthogonal RNA-binding reagents.Conclusion
In this study, we demonstrate that mutagenesis-based SELEX can be successfully applied to cross-chiral aptamers to rapidly reprogram target specificity while maintaining high selectivity. Starting from a single parental L-aptamer, this approach yielded two evolved aptamers, L-5A-16t and L-8U-1t, which together with the original L-6-4t form a set of three orthogonal RNA-binding reagents capable of discriminating RNA hairpins that differ by only a single nucleotide. By combining deep sequencing with enrichment analysis, we show that this process is rapid and highly informative. Cross-chiral aptamers with altered target selectivity can be identified within three or fewer selection rounds, underscoring the speed and efficiency of this approach. At the same time, the sequencing data reveal that retargeting efficiency is sensitive to the degree of structural perturbation in the target RNA, as illustrated by the distinct evolutionary trajectories observed for the TARU variant. Importantly, analysis of sequence evolution and predicted secondary structures revealed that accommodation of even a single-nucleotide change in the target does not arise from local or incremental adjustments to the parental aptamer. Instead, both transition and transversion mutations in the target RNA necessitated extensive restructuring of the aptamer fold through complex, distributed mutation networks. These observations underscore a defining advantage of cross-chiral aptamers: their ability to recognize RNA targets through global structure and tertiary architecture rather than sequence complementarity. As a result, cross-chiral aptamers can achieve levels of specificity and single-nucleotide discrimination that are difficult to realize with hybridization-based approaches such as antisense oligonucleotides, which rely primarily on Watson–Crick base pairing and are often insensitive to subtle but functionally important structural differences.40,41 Structure-based recognition enables selective targeting of conformationally distinct RNA states, providing a powerful means to discriminate between closely related RNAs that differ minimally in sequence but meaningfully in shape.While the present conclusions are derived from a defined set of cross-chiral aptamer–hairpin pairs, the broader implications extend well beyond the TAR system. RNA hairpins and stem–loop motifs are ubiquitous in biology, serving as regulatory elements, protein-binding platforms, and conserved functional structures in viral and cellular RNAs.1,42 Unlike hybridization-based approaches such as antisense oligonucleotides or siRNAs, which rely on Watson–Crick base pairing and therefore require access to linear sequence elements, cross-chiral aptamers recognize RNA targets through tertiary structure and global shape complementarity.16 This feature may enable selective targeting of structured RNA motifs that are difficult to access using traditional hybridization-based technologies. The ability to rapidly retarget an existing aptamer to closely related structural variants offers a practical route to generate new reagents against evolving RNA targets, such as viral genomes that accumulate point mutations during strain evolution while preserving overall secondary structure. In this context, mutagenesis-based evolution enables parallel diversification of aptamer specificity from a common scaffold, allowing rapid adaptation to emerging variants without restarting the selection process. Future applications of this framework could extend beyond the single-nucleotide loop mutations examined here to explore larger sequence and structural perturbations, further defining the scope and adaptability of cross-chiral aptamer evolution. While mutagenesis-based selection is inherently biased toward sequence space proximal to the parental aptamer, the substantial mutational divergence and structural remodeling observed here indicate that this approach can access a diverse range of sequence and structural solutions. Nonetheless, de novo selection from fully randomized libraries may yield alternative aptamer architectures with distinct and potentially improved binding properties, and these strategies should therefore be viewed as complementary depending on the intended application. Importantly, even within the limited sequence space examined here, the robustness of the evolved aptamers is already evident. RNA pulldown experiments demonstrated that the evolved aptamers retain strict target selectivity in a competitive and structurally complex environment that includes extended flanking sequences and multiple RNA species. The maintenance of orthogonality under these conditions underscores their robustness and highlights their potential for diverse applications, including RNA-selective biosensors, diagnostic tools, and affinity reagents for isolating specific structured RNAs from complex biological samples.30 An important future direction will be structural characterization of cross-chiral aptamer–RNA complexes to better understand the molecular basis of these interactions. While current computational prediction tools are largely optimized for natural biopolymers, advances in RNA structural biology and modeling may eventually enable high-resolution characterization of cross-chiral recognition mechanisms.
Recent discussions surrounding mirror-life have emphasized the possibility that mirror biopolymers could be largely orthogonal to natural biological systems.43 However, cross-chiral aptamers provide clear experimental examples of selective molecular recognition between mirror-image nucleic acids. The results presented here further demonstrate that such cross-chiral interactions can evolve to discriminate structured RNA targets that differ by only a single nucleotide, highlighting the precision achievable through tertiary, structure-based recognition. These findings illustrate that while mirror biopolymers are incompatible with canonical Watson–Crick hybridization, highly specific interactions between chiral systems can nevertheless arise through alternative structural recognition mechanisms.
Taken together, this work establishes mutagenesis-based SELEX as a versatile strategy for rapidly reprogramming cross-chiral L-aptamers and reinforces the unique advantages of structure-based RNA recognition for discriminating closely related structured RNA targets. More broadly, it provides a framework for developing adaptable RNA-targeting reagents capable of keeping pace with dynamic and mutating RNA landscapes, positioning cross-chiral aptamers as powerful tools in chemical biology and RNA-focused biotechnology.
Author contributions
Xuan Han: methodology, investigation, formal analysis, visualization, writing – original draft. Tyler Guilbault: methodology, investigation, formal analysis, visualization. Jonathan T. Sczepanski: conceptualization, formal analysis, writing – original draft, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Fig. S1–S19 and Tables S1, S2, reagents and experimental methods, oligonucleotides sequences and structures, mass spectra of oligonucleotides, gel electrophoresis images, and DNA sequencing data. See DOI: https://doi.org/10.1039/d6cb00057f.Acknowledgements
This work was supported by the National Institute of General Medical Sciences (R35GM124974) of the National Institutes of Health. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.References
- X. Cao, Y. Zhang, Y. Ding and Y. Wan, Identification of RNA structures and their roles in RNA functions, Nat. Rev. Mol. Cell Biol., 2024, 25(10), 784–801 CrossRef CAS PubMed.
- Y. Zhang, J. Qian, C. Gu and Y. Yang, Alternative splicing and cancer: a systematic review, Signal Transduction Targeted Ther., 2021, 6(1), 78 CrossRef CAS PubMed.
- Q. Peng, Y. Zhou, L. Oyang, N. Wu, Y. Tang, M. Su, X. Luo, Y. Wang, X. Sheng, J. Ma and Q. Liao, Impacts and mechanisms of alternative mRNA splicing in cancer metabolism, immune response, and therapeutics, Mol. Ther., 2022, 30(3), 1018–1035 CrossRef CAS PubMed.
- Y. Peng and C. M. Croce, The role of MicroRNAs in human cancer, Signal Transduction Targeted Ther., 2016, 1(1), 15004 CrossRef PubMed.
- B. Malone, N. Urakova, E. J. Snijder and E. A. Campbell, Structures and functions of coronavirus replication–transcription complexes and their relevance for SARS-CoV-2 drug design, Nat. Rev. Mol. Cell Biol., 2022, 23(1), 21–39 CrossRef CAS PubMed.
- T. Osawa, M. Aoki, H. Ehara and S.-I. Sekine, Structures of dengue virus RNA replicase complexes, Mol. Cell, 2023, 83(15), 2781–2791.e4 CrossRef CAS PubMed.
- R. G. Huber, X. N. Lim, W. C. Ng, A. Y. L. Sim, H. X. Poh, Y. Shen, S. Y. Lim, K. B. Sundstrom, X. Sun, J. G. Aw, H. K. Too, P. H. Boey, A. Wilm, T. Chawla, M. M. Choy, L. Jiang, P. F. de Sessions, X. J. Loh, S. Alonso, M. Hibberd, N. Nagarajan, E. E. Ooi, P. J. Bond, O. M. Sessions and Y. Wan, Structure mapping of dengue and Zika viruses reveals functional long-range interactions, Nat. Commun., 2019, 10(1), 1408 CrossRef PubMed.
- P. K. Todd and H. L. Paulson, RNA-mediated neurodegeneration in repeat expansion disorders, Ann. Neurol., 2010, 67(3), 291–300 CrossRef CAS PubMed.
- A. Baud, M. Derbis, K. Tutak and K. Sobczak, Partners in crime: proteins implicated in RNA repeat expansion diseases, WIREs RNA, 2022, 13(4), e1709 CrossRef CAS PubMed.
- Y. Sun, H. Dai, X. Dai, J. Yin, Y. Cui, X. Liu, G. Gonzalez, J. Yuan, F. Tang, N. Wang, A. E. Perlegos, N. M. Bonini, X. W. Yang, W. Gu and Y. Wang, m1A in CAG repeat RNA binds to TDP-43 and induces neurodegeneration, Nature, 2023, 623(7987), 580–587 CrossRef CAS PubMed.
- A. Chini, P. Guha, V. S. Malladi, Z. Guo and S. S. Mandal, Novel long non-coding RNAs associated with inflammation and macrophage activation in human, Sci. Rep., 2023, 13(1), 4036 CrossRef CAS PubMed.
- S. R. Nallagatla, R. Toroney and P. C. Bevilacqua, Regulation of innate immunity through RNA structure and the protein kinase PKR, Curr. Opin. Struct. Biol., 2011, 21(1), 119–127 CrossRef CAS PubMed.
- S. Ando, S. Suzuki, S. Okubo, K. Ohuchi, K. Takahashi, S. Nakamura, M. Shimazawa, K. Fuji and H. Hara, Discovery of a CNS penetrant small molecule SMN2 splicing modulator with improved tolerability for spinal muscular atrophy, Sci. Rep., 2020, 10(1), 17472 CrossRef CAS PubMed.
- R. Aguilar, K. B. Spencer, B. Kesner, N. F. Rizvi, M. D. Badmalia, T. Mrozowich, J. D. Mortison, C. Rivera, G. F. Smith, J. Burchard, P. J. Dandliker, T. R. Patel, E. B. Nickbarg and J. T. Lee, Targeting Xist with compounds that disrupt RNA structure and X inactivation, Nature, 2022, 604(7904), 160–166 CrossRef CAS PubMed.
- J. L. Childs-Disney, X. Yang, Q. M. R. Gibaut, Y. Tong, R. T. Batey and M. D. Disney, Targeting RNA structures with small molecules, Nat. Rev. Drug Discovery, 2022, 21(10), 736–762 CrossRef CAS PubMed.
- B. S. E. Heale, H. S. Soifer, C. Bowers and J. J. Rossi, siRNA target site secondary structure predictions using local stable substructures, Nucleic Acids Res., 2005, 33(3), e30–e30 CrossRef PubMed.
- H. Ratni, R. S. Scalco and A. H. Stephan, Risdiplam, the First Approved Small Molecule Splicing Modifier Drug as a Blueprint for Future Transformative Medicines, ACS Med. Chem. Lett., 2021, 12(6), 874–877 CrossRef CAS PubMed.
- C. Tuerk and L. Gold, Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase, Science, 1990, 249(4968), 505–510 CrossRef CAS PubMed.
- H. Qu, A. T. Csordas, J. Wang, S. S. Oh, M. S. Eisenstein and H. T. Soh, Rapid and Label-Free Strategy to Isolate Aptamers for Metal Ions, ACS Nano, 2016, 10(8), 7558–7565 CrossRef CAS PubMed.
- M. Guérin, M. Vandevenne, A. Matagne, W. Aucher, J. Verdon, E. Paoli, J. Ducrotoy, S. Octave, B. Avalle, I. Maffucci and S. Padiolleau-Lefèvre, Selection and characterization of DNA aptamers targeting the surface Borrelia protein CspZ with high-throughput cross-over SELEX, Commun. Biol., 2025, 8(1), 632 CrossRef PubMed.
- K. Sefah, D. Shangguan, X. Xiong, M. B. O'Donoghue and W. Tan, Development of DNA aptamers using Cell-SELEX, Nat. Protoc., 2010, 5(6), 1169–1185 CrossRef CAS PubMed.
- J. T. Sczepanski and G. F. Joyce, Specific Inhibition of MicroRNA Processing Using L-RNA Aptamers, J. Am. Chem. Soc., 2015, 137(51), 16032–16037 CrossRef CAS PubMed.
- J. T. Sczepanski and G. F. Joyce, Binding of a Structured D-RNA Molecule by an L-RNA Aptamer, J. Am. Chem. Soc., 2013, 135(36), 13290–13293 CrossRef CAS PubMed.
- J. Li and J. T. Sczepanski, Targeting a conserved structural element from the SARS-CoV-2 genome using L-DNA aptamers, RSC Chem. Biol., 2022, 3(1), 79–84 RSC.
- A. M. Kabza and J. T. Sczepanski, An L-RNA Aptamer with Expanded Chemical Functionality that Inhibits MicroRNA Biogenesis, ChemBioChem, 2017, 18(18), 1824–1827 CrossRef CAS PubMed.
- S. Dey and J. T. Sczepanski, In vitro selection of L-DNA aptamers that bind a structured D-RNA molecule, Nucleic Acids Res., 2020, 48(4), 1669–1680 CrossRef CAS PubMed.
- M. I. Umar, C.-Y. Chan and C. K. Kwok, Development of RNA G-quadruplex (rG4)-targeting L-RNA aptamers by rG4-SELEX, Nat. Protoc., 2022, 17(6), 1385–1414 Search PubMed.
- S. Y. Lam, M. I. Umar, H. Zhao, J. Zhao and C. K. Kwok, Capture of RNA G-quadruplex structures using an L-RNA aptamer, RSC Chem. Biol., 2024, 5(10), 1045–1051 RSC.
- D. Ji, J.-H. Yuan, S.-B. Chen, J.-H. Tan and C. K. Kwok, Selective targeting of parallel G-quadruplex structure using L-RNA aptamer, Nucleic Acids Res., 2023, 51(21), 11439–11452 CrossRef PubMed.
- V. Shearer, C.-H. Yu, X. Han and J. T. Sczepanski, The clinical potential of L-oligonucleotides: challenges and opportunities, Chem. Sci., 2024, 15(44), 18239–18258 RSC.
- A. T. Piwko, X. Han, A. M. Kabza, S. Dey and J. T. Sczepanski, Inverse In Vitro Selection Enables Comprehensive Analysis of Cross-Chiral L-Aptamer Interactions, ChemBioChem, 2022, 23(24), e202200520 CrossRef CAS PubMed.
- C. Olea, J. Weidmann, P. E. Dawson and G. F. Joyce, An L-RNA Aptamer that Binds and Inhibits RNase, Chem. Biol., 2015, 22(11), 1437–1441 CrossRef CAS PubMed.
- Z. Huang and J. W. Szostak, Evolution of aptamers with a new specificity and new secondary structures from an ATP aptamer, RNA, 2003, 9(12), 1456–1463 CrossRef CAS PubMed.
- F. Wachowius, B. T. Porebski, C. M. Johnson and P. Holliger, Emergence of ATP- and GTP-Binding Aptamers from Single RNA Sequences by Error-Prone Replication and Selection, ChemSystemsChem, 2023, 5(5), e202300006 CrossRef CAS PubMed.
- S. Klußmann, A. Nolte, R. Bald, V. A. Erdmann and J. P. Fürste, Mirror-image RNA that binds D-adenosine, Nat. Biotechnol., 1996, 14(9), 1112–1115 CrossRef PubMed.
- K. P. Williams, X.-H. Liu, T. N. M. Schumacher, H. Y. Lin, D. A. Ausiello, P. S. Kim and D. P. Bartel, Bioactive and nuclease-resistant L-DNA ligand of
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) vasopressin, Proc. Natl. Acad. Sci. U. S. A., 1997, 94(21), 11285–11290 CrossRef CAS PubMed.
vasopressin, Proc. Natl. Acad. Sci. U. S. A., 1997, 94(21), 11285–11290 CrossRef CAS PubMed. - A. Nolte, S. Klußmann, R. Bald, V. A. Erdmann and J. P. Fürste, Mirror-design of L-oligonucleotide ligands binding to L-arginine, Nat. Biotechnol., 1996, 14(9), 1116–1119 CrossRef CAS PubMed.
- T. Singh, R. Fogel and J. Limson, NGS-Guided Aptamer Re-Selection for Improved Sensor Applications: Biotin as a Modification Tag in the Amplification of Enriched Pools, Eng. Proc., 2025, 109(1), 7 Search PubMed.
- R. Didarian, H. K. Ozbek, V. C. Ozalp, O. Erel and N. Yildirim-Tirgil, Enhanced SELEX Platforms for Aptamer Selection with Improved Characteristics: A Review, Mol. Biotechnol., 2025, 67(8), 2962–2977 CrossRef CAS PubMed.
- J. Scharner, W. K. Ma, Q. Zhang, K.-T. Lin, F. Rigo, C. F. Bennett and A. R. Krainer, Hybridization-mediated off-target effects of splice-switching antisense oligonucleotides, Nucleic Acids Res., 2020, 48(2), 802–816 CrossRef CAS PubMed.
- P. Andersson, S. A. Burel, H. Estrella, J. Foy, P. H. Hagedorn, T. A. Harper, S. P. Henry, J.-C. Hoflack, E. M. Holgersen, A. A. Levin, E. Morrison, A. Pavlicek, L. Penso-Dolfin and U. Saxena, Assessing Hybridization-Dependent Off-Target Risk for Therapeutic Oligonucleotides: Updated Industry Recommendations, Nucleic Acid Ther., 2025, 35(1), 16–33 CrossRef CAS PubMed.
- P. Svoboda and A. D. Cara, Hairpin RNA: a secondary structure of primary importance, Cell. Mol. Life Sci., 2006, 63(7), 901–908 CrossRef CAS PubMed.
- K. P. Adamala, D. Agashe, Y. Belkaid, D. M. D. C. Bittencourt, Y. Cai, M. W. Chang, I. A. Chen, G. M. Church, V. S. Cooper, M. M. Davis, N. K. Devaraj, D. Endy, K. M. Esvelt, J. I. Glass, T. W. Hand, T. V. Inglesby, F. J. Isaacs, W. G. James, J. D. G. Jones, M. S. Kay, R. E. Lenski, C. Liu, R. Medzhitov, M. L. Nicotra, S. B. Oehm, J. Pannu, D. A. Relman, P. Schwille, J. A. Smith, H. Suga, J. W. Szostak, N. J. Talbot, J. M. Tiedje, J. C. Venter, G. Winter, W. Zhang, X. Zhu and M. T. Zuber, Confronting risks of mirror life, Science, 2024, 386(6728), 1351–1353 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |