Open Access Article
Open Access ArticleEnhancing mRNA stability and translational potential through tailored modifications at the 3′ end
Olga Perzanowskaab,
Joanna Kowalska*a and
Jacek Jemielity *b
*b
aDivision of Biophysics, Faculty of Physics, University of Warsaw, Pasteura 5, Warsaw, 02-093, Poland. E-mail: Joanna.Kowalska@fuw.edu.pl
bCentre of New Technologies, University of Warsaw, Banacha 2c, Warsaw, 02-097, Poland. E-mail: j.jemielity@cent.uw.edu.pl
First published on 17th April 2026
Abstract
Poly(A) tails regulate mRNA turnover and translation through multiple RNA–protein interactions, and deadenylation is the initiating step of the major cytoplasmic decay pathways. Here, we report a simple post-transcriptional strategy to protect the 3′ end of synthetic mRNA by enzymatically ligating short, chemically modified 5′-phosphorylated dinucleotides. Using T4 RNA ligase 1, we attached 2′-O-methyl and/or phosphorothioate containing A- or G-dinucleotides to the 3′ end of model oligoadenylates and to an IVT Gaussia luciferase (GLuc) mRNA bearing an ∼150-nt poly(A) tail. All ligated products showed strong resistance to human CNOT7-mediated deadenylation in vitro, whereas the unmodified control mRNA underwent poly(A) removal. In rabbit reticulocyte lysate, 3′-modified GLuc mRNAs translated comparably to the control, indicating minimal interference with the translational machinery. In mammalian cells (A549, JAWSII and HEK293), the protein output depended on the dinucleotide structure and cell type; two adenosine donors—pApAm and the D1 phosphorothioate stereoisomer pApsAm—consistently performed best, increasing the cumulative GLuc production up to 163% in HEK293 and up to 79% in A549. These results establish dinucleotide ligation as a minimal and modular 3′-end engineering approach that enhances resistance to deadenylation and can improve the translational performance of therapeutic mRNA candidates.
Introduction
The use of in vitro transcribed (IVT) mRNAs as potential drugs has been considered since the 1990s.1 Due to great technological advances, over the last decade, mRNA has become an increasingly promising tool in vaccine development2–4 and protein replacement therapies.5,6 The recent SARS-CoV-2 pandemic showcased the potential of mRNA-based vaccines in a spectacular fashion.7,8 The main advantages of mRNA virus vaccines over more conventional methods (such as killed/attenuated virus or DNA-based vaccines) are several: relatively low-cost and fast manufacturing, no risk of infection or insertional mutagenesis and, most importantly for this study, IVT mRNAs can be easily modified in various ways to improve their stability and translational potential in vivo.9–12 Recent work has also shown that engineering the poly(A) tail architecture itself (e.g., segmented poly(A) designs) can improve the robustness of IVT mRNA production and increase translatability.13,14 In addition to virus vaccines, mRNA cancer vaccines are being extensively studied as a non-toxic, highly specific alternative to conventional cancer treatments.15–17 Still, the core problem of mRNA-based drugs remains—mRNA is a highly unstable molecule due to omnipresent ribonucleases (RNases) that degrade RNA.18 Although significant achievements have been made in the realm of IVT mRNA proliferation, both at the point of delivery19 and further cellular metabolism, we still face a number of obstacles, the main of which is the efficiency of in vivo protein production after therapeutic mRNA administration.Regardless of the decay pathway, mRNA degradation in mammalian cells is initiated by deadenylation, which involves the removal of the poly(A) tail located at the 3′ end of most mature eukaryotic mRNAs.20 Deadenylation is predominantly executed by the CCR4-NOT complex and is increasingly recognized as a post-transcriptional hub that integrates mRNA turnover with translational control through multiple, regulated interactions with RNA-binding proteins and ribosomes.21–23 The poly(A) tail acts as one of the main regulators of mRNA metabolism24 by protecting the coding regions of mRNA from degradation25 and playing an important role in the translation initiation process through protein–RNA interactions.26,27 Beyond tail length, mounting evidence indicates that poly(A) tails can encode regulatory information through their composition and dynamics, including the presence and positioning of non-adenosine residues, providing a conceptual basis for minimal 3′-end interventions.28 Both of these features, combined with the non-coding nature of poly(A) tail, make it a perfect target for introducing advantageous modifications to increase the stability and translational potential of IVT mRNAs without hindering the binding of the ribosomal machinery. In recent years, several in-depth studies tackling poly(A) modifications for mRNA labeling and increasing mRNA stability have been reported.13,14,29–32 Transcriptome-wide poly(A) tail profiling has further revealed that non-A residues within and at the termini of poly(A) tails are widespread and dynamic across biological contexts, supporting the notion that modest compositional changes at the 3′ end can substantially affect RNA fate.33 In line with this, poly(A) tail engineering—ranging from chemical modifications to architectural redesign—has emerged as a practical route to improve IVT mRNA performance. This greatly inspired our studies, in addition to our previous work concerning the structure–activity relationship between modified poly(A) tail analogs and poly(A) binding protein (PABP).34 Among other functions, PABP promotes translation initiation by playing an intermediary role between the poly(A) tail and the rest of the translational machinery. Thus, we previously considered investigating the ligand specificity of PABP as an important step in further development of therapeutic mRNAs. In our research, we concluded that several of the investigated nucleotide modifications (2′-O-methyladenosine, 2′-O-fluoroadenosine and interchain phosphorothioates) had the ability to fully inhibit the poly(A) degradation process carried out by CNOT7 deadenylase. Additionally, by strategically placing these modifications within the poly(A) chain, such as at the 3′ end, any significant decrease in binding affinity for PABP could be avoided. Importantly, the specificity and rates of CCR4-NOT-mediated deadenylation can be tuned by using adaptor proteins and signaling inputs, implying that the impact of 3′-end engineering may vary across cellular contexts.35
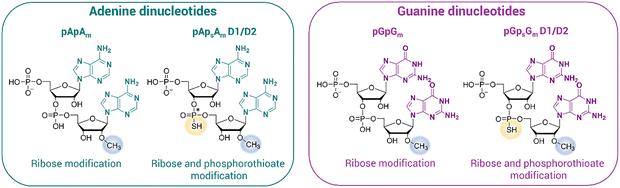
In this study, we tested a novel chemo-enzymatic approach for the modification of the mRNA 3′ end. We synthesized two sets of 5′-monophosphorylated dinucleotides, containing adenine (A) or guanine (G) nucleobases. In each set, there were three differently modified compounds, all containing 2′-O-methyl combined either with a regular phosphate (pApAm/pGpGm) or a phosphorothioate linkage (two pApsAm stereoisomers or two pGpsGm stereoisomers). First, we incorporated each of the dinucleotide compounds into the 3′ end of a short 12-mer oligoadenylate analog (A12 OA) via enzymatic ligation. The resulting 14-mer oligonucleotide products were then tested for susceptibility to specific degradation. The same dinucleotide compounds were then ligated to the 3′ end of mRNA encoding the Gaussia Luciferase reporter protein. Following ligation, all modified mRNAs were again tested for susceptibility to deadenylase activity. Protein expression experiments with the modified mRNAs were conducted both in the rabbit reticulocyte lysate (RRL) system and three different cell lines. This line of study led to the identification of mRNAs that yield significantly increased protein output compared to unmodified mRNAs.
Results and discussion
Short oligoadenylate analogs modified with dinucleotides at the 3′ end are fully resistant to deadenylation by recombinant CNOT7
Most of the 3′ end chemical modifications proposed to date require either co-transcriptional incorporation29 or the preparation of relatively long and complex compounds31 for the ligation approach. Poly(A) tail engineering is increasingly explored as a practical platform lever to improve IVT mRNA performance, including architectural redesign of tails while leaving the coding region unchanged.13 We aimed to achieve both increased enzymatic stability and translational potential of mRNA with small, easy-to-obtain compounds that could be incorporated to the 3′ end of any RNA chain, from short oligonucleotides to fully functional mRNAs. Drawing from our previous results,34 we selected two nucleotide modifications: 2′-O-methylation and 5′,3′-phosphorothioate to introduce into 5′-monophosphorylated 5′,3′-dinucleotides that could act as donors in an enzymatic ligation reaction. Our goal was to achieve a high level of enzymatic protection of the 3′ end, with an emphasis on protection against deadenylation, without hindering other important biological processes. The 3′ end of the RNA chain can be modified and, therefore, stabilized via enzymatic ligation carried out with T4 RNA Ligase 1. Due to the substrate specificity of T4 RNA Ligase 1, it is possible to attach chemically modified oligonucleotides of up to several dozen subunits in length to the 3′ end of any RNA precursor.31 The only limitation is the availability of the phosphate group at the 5′ end of the oligonucleotide donor and a free 3′-OH group at the 3′ end of the RNA substrate/acceptor.As all known eukaryotic deadenylases exhibit high substrate preference31,36 or indirect dependence37,38 on polyadenylate chains, we decided to design compounds containing either adenine or guanine nucleobases, in addition to the aforementioned chemical modifications. Incorporating G subunits at the 3′ end of the RNA chain could provide additional protection against deadenylation by further interfering with recognition and binding by deadenylases.39 Notably, non-adenosine residues at poly(A) tails are now recognized as widespread and dynamic, supporting the notion that modest 3′-end compositional changes can have disproportionate consequences for RNA fate.28,33 At the same time, altering the nucleobases of the terminal subunits of the poly(A) tail could impede cellular processes necessary for high mRNA translational output. In addition to achieving improved stability, we intended to identify compounds capable of increasing protein production, and so we considered it important to test different chemical modifications and nucleobase variations.
All 5′-monophosphorylated dinucleotides (pApAm, pApsAm) stereoisomers D1 and D2, pGpGm, and pGpsGm stereoisomers D1 and D2; (Fig. 1) were obtained by solid-phase synthesis on a solid support loaded with either 2′O-Me-A or 2′O-Me-G phosphoramidites. All compounds were purified using semi-preparative reversed-phase high-performance liquid chromatography (RP-HPLC) and characterized by high-resolution mass spectrometry (HRMS; for full characterization, see the SI). Based on previous reports linking phosphorus stereochemistry in dinucleotides with chromatographic mobility, we established that the stereoisomer D1 has RP configuration, while stereoisomer D2 has SP configuration.40,41
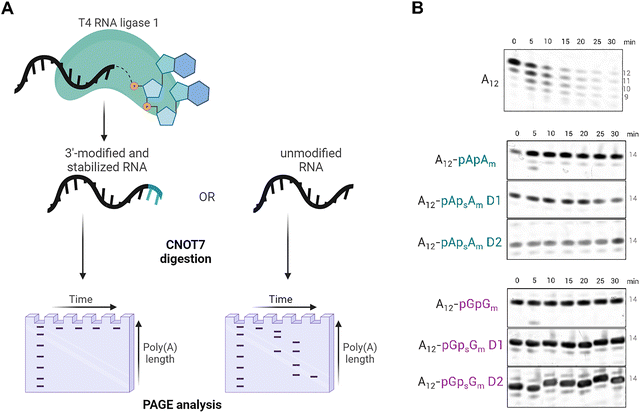
The ligation conditions and the enzymatic deadenylation protocol were optimized on a short oligoadenylate analog (A12) before attempting to introduce the modifications to the poly(A) tails of functional mRNA (Fig. 2A). For the ligation reaction with T4 RNA Ligase 1, a 5-fold molar excess of the dinucleotide donor over the oligonucleotide acceptor was enough to accomplish full conversion of the A12 substrate to the 14-mer product overnight at room temperature (Fig. S1A).43 We noticed that in the case of the pGpsGm D2 dinucleotide donor, a small amount of doubly ligated product was formed (Fig. S1B), which arises likely from the fact that the 2′-O-methylation of the ribose hinders, but does not completely block, the formation of the 3′,5′-phosphodiester bond and elongation of the chain.42 The main ligation products were in each case purified by RP-HPLC purification and freeze-dried.
To test the 14-mer ligation products for their susceptibility to enzymatic degradation, they were subjected to recombinant human CNOT7 deadenylase. CNOT7 is one of the catalytic subunits of the protein complex CCR4-NOT, which is mainly responsible for the removal of poly(A) tails from cytoplasmic eukaryotic mRNA.43,44 Beyond its catalytic role, CCR4-NOT functions as a regulatory hub coupling deadenylation with broader mRNP remodeling and translational control, making 3′-end protection strategies particularly attractive in therapeutic mRNA design. Modified oligonucleotides were incubated with CNOT7 for 30 minutes at 37 °C.21–23 Starting at 0 minutes, aliquots were taken from the reaction mixture every 5 minutes, and the reaction progress was quenched by dilution with a loading dye/EDTA mixture. Samples were analyzed using high-resolution urea PAGE (Fig. 2B). Unlike the unmodified A12 chain, which underwent gradual shortening as a result of enzymatic digestion, all modified 14-mers remained stable in the presence of CNOT7. These results confirmed our assumptions that incorporating selected, chemically modified dinucleotide structures at the 3′ end of the polyadenylate chain can protect it from the CNOT7 enzymatic activity and potentially other 3′–5′ exonucleases.
mRNAs modified with dinucleotides at the 3′ end are not susceptible to CNOT7 in vitro
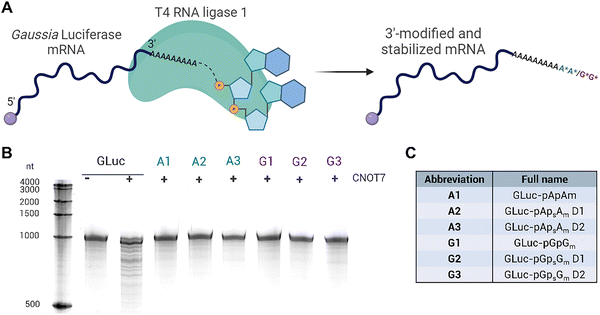
Given inevitable differences in the reaction conditions when working with full mRNA moieties rather than short oligonucleotides, we made necessary changes to our ligation protocol. The main concern was the lower concentration of mRNA in the solution, or rather the concentration of 3′-OH groups available as acceptors in the reaction. This required a substantial increase in the amount of dinucleotides in the ligation mixture, from a 5-fold molar excess to a 100-fold excess. Also, 15% PEG800 was added to the reaction mixture to increase molecular crowding and accelerate the ligation process.45 The incubation time was shortened from overnight to just 1 hour to avoid exposing vulnerable mRNAs to unfavorable conditions.To facilitate later biological studies, we chose mRNA encoding Gaussia Luciferase (GLuc) as a reporter protein for the 3′ end modification experiments.46 Following in vitro transcription, GLuc mRNA, capped at the 5′ end with cap-1 analog (m7GpppAmG) and containing an ∼150-long poly(A) tail at the 3′ end, was purified using RP-HPLC to increase the quality of the samples and to remove double-stranded RNA contaminants.47 Each ligation reaction was prepared according to the improved protocol (for the full ligation protocol see the Experimental section), including an unmodified control mRNA that was incubated without any 5′-monophosphorylated dinucleotides to ensure an identical preparation process for all mRNA samples (Fig. 3A and C). After 1 h of ligation, samples were purified using Monarch RNA cleanup spin columns. Similar to previously described 14-mer oligonucleotides, in order to test the effectiveness of the ligation procedure and examine the mRNAs for resistance to deadenylation, all mRNA samples were subjected to enzymatic degradation by CNOT7 deadenylase (Fig. 3B) for 30 min. As per previous results, ligated mRNAs remained stable, whereas the unmodified control sample was shortened due to the removal of the poly(A) tail.
Translation efficiency in cells is increased by 3′-end dinucleotide modifications of mRNA
Knowing that all ligated mRNAs were highly resistant to deadenylation, we hypothesized that at least some would lead to increased Gaussia Luciferase protein production in cells, at least in part by impeding poly(A) shortening and thereby potentially prolonging the functional mRNA lifetime.21–23 At the same time, we wanted to account for differences in translation mechanisms, mRNA decay rates, and transfection efficiencies in different cell lines. To this end, we chose three cell lines of significantly different morphologies and origins. A549 epithelial lung cancer cells are a popular model for testing anti-cancer therapies.48 JAWSII and other dendritic cells have long been used as models for cellular mediators in anti-cancer vaccinations with capability to induce an immune response against tumors.49,50 Lastly, HEK 293 kidney cells are highly versatile, easy to transfect, and characterized by an innate high level of translation and protein production.51Prior to cell line experiments, all modified mRNAs were also used in translation efficiency assays in the rabbit reticulocyte lysate (RRL) system. All exhibited translation levels similar to or slightly higher than those of unmodified GLuc mRNA (Fig. S2). Given the nature of the RRL system, in which the cellular machinery responsible for mRNA degradation is reduced to a minimum, these results were satisfactory. Since the mRNA modifications we introduced were intended to increase mRNA stability, this effect would not be detectable in the RRL system. Importantly, based on the result of the experiment, we could assume that none of the 3′ end modifications interfered significantly with the translational machinery, at least in the RRL system.
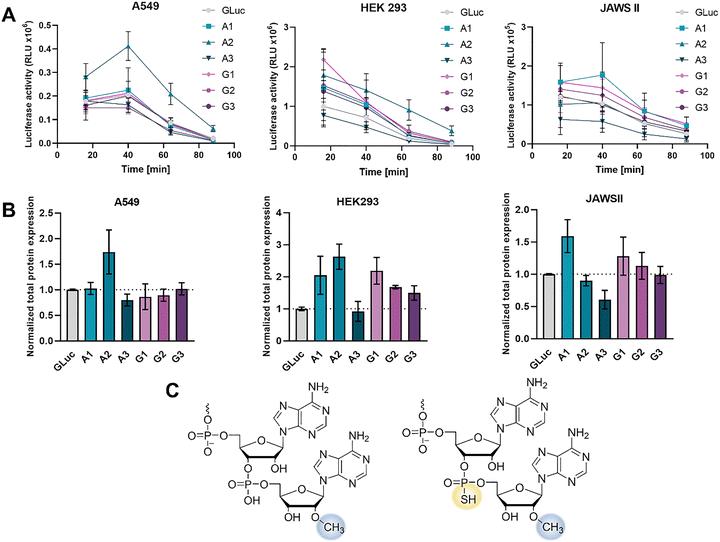
All ligated mRNAs were next transfected into A549, JAWSII and HEK293 cell lines, along with unmodified control mRNA. Gaussia Luciferase expression was monitored at several time points (16, 40, 64, and 88 h after transfection) by collecting cell culture medium and measuring the activity level of the secreted Gaussia Luciferase protein. Total protein expression levels of all modified mRNAs were normalized to the unmodified GLuc mRNA expression.
As expected, protein expression varied greatly depending on both mRNA type and cell line (Fig. 4). In adenocarcinomic cell line A549, only mRNA A2 exhibited translational potential higher than the control (79% increase in protein production). The expression of the rest was similar to that of control mRNA, indicating that either these modifications did not stabilize mRNA in this cell line or any translational enhancement brought about by increased stability was undercut by modifications of the 3′ end, negatively affecting the translation process (Fig. 4A and B). In the JAWSII cell line, we also identified one 3′ end modification that resulted in enhanced translation (mRNA A1, 70%). Interestingly, both D1 (mRNA A2) and D2 (mRNA A3) stereoisomers of pApsAm decreased the translation levels, in the case of D2 by 40%. In the case of mRNAs ligated with guanosine dinucleotides (mRNA G1, G2, G3), their translational output remained similar to that of the control. In HEK 293 cells, differences in protein expression were most pronounced. Both mRNA A1 and A2 led to significantly increased protein expression (by 105% and 163%, respectively), while modification in mRNA A3 once again proved to be the least beneficial in terms of improving the translational potential. mRNA G1 was also translated very efficiently (118% increase in protein production) and both G2 and G3 led to improved expression (by 65% and 49%, respectively). The pronounced cell-type dependence is consistent with mechanistic evidence that CCR4-NOT-mediated deadenylation rates can be tuned by adaptor recruitment and signaling inputs, which may modulate how strongly a given 3′-end modification shifts the balance between translation and decay.35 It is also possible that other general or cell-specific degradation mechanisms are responsible for the removal of the 3′-terminal dinucleotides from mRNA. Overall, two dinucleotides provided the best results in translational output enhancement: pApAm (mRNA A1) and pApsAm D1 (mRNA A2) (Fig. 4C). pApAm increased translation in both JAWSII and HEK293 cell lines, whereas pApsAm D1 provided translation enhancement in A549 and HEK293 cell lines. pGpGm modification also proved beneficial and could be of further interest considering its potentially even greater stability against deadenylases than in the case of adenosine dinucleotides.
3′-Terminal dinucleotide modification increases cytoplasmic stability of mRNA
We next aimed to investigate whether the 3′-terminal modification of mRNA affects its cytoplasmic stability. However, standard methods, such as transfection followed by qPCR, cannot accurately address this question as they are confounded by transfection kinetics and endosome-trapped mRNA. As such, we have microinjected mRNAs encoding a fluorescent reporter (eGFP) directly into cells’ cytoplasm and monitored their translational activity over time by confocal microscopy. Single A549 cells were injected with two mRNA variants: unmodified eGFP and modified with A2 (eGFP-pApsAm D1). The data on eGFP protein concentration over time were then analyzed using mathematical modeling to estimate protein and mRNA half-lives (tdp and tdm, respectively). To verify the assumption of a constant rate of mRNA degradation, and thus the possibility of using the eGFP reporter in the study, we used the simplest, first-order kinetic formulas:
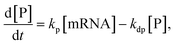 | (1) |
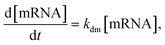 | (2) |
Based on the data collected from three independent replicates (Fig. S3), the estimated half-life was roughly twofold longer for modified eGFP-pApsAm D1 (tdm ∼ 66–71 h), when compared to that of its unmodified counterpart (tdm ∼ 27–28 h). This corresponds well with the overall increased protein output of GLuc-pApsAm D1 (A2) mRNA in A549 cells compared to the unmodified control mRNA and indicates that this type 3′-end modification may significantly increase the cytoplasmic half-life of mRNAs.
Conclusions
In summary, we developed a simple and efficient post-transcriptional strategy for engineering the 3′ end of synthetic mRNA using enzymatic ligation of short, chemically modified 5′-phosphorylated dinucleotides. We designed and synthesized six dinucleotide donors (A- and G-based), each bearing a 2′-O-methyl modification at the second nucleotide and, in selected cases, a stereodefined phosphorothioate linkage. Using T4 RNA ligase 1, these donors were appended to the 3′ end of a short oligoadenylate model (A12) and to an IVT Gaussia luciferase (GLuc) mRNA carrying an ∼150-nt poly(A) tail. In both formats, 3′-end dinucleotide installation conferred strong resistance to CNOT7-mediated deadenylation in vitro, whereas the unmodified mRNA control underwent poly(A) removal.In mammalian cells, the impact on protein output depended on the dinucleotide structure and the cellular context. Among the tested constructs, two adenosine-based donors (A1 and A2) yielded the most consistent increases in reporter production across cell lines. A2 modification also increased mRNA half-life after microinjection into A549 cells. Taken together, these results demonstrate that minimal 3′-end modifications can effectively protect the poly(A) tail from enzymatic shortening in vitro and, in favorable contexts, increase protein output in living cells without the need for altering the coding sequence. Future work should extend this approach to additional mRNA cargos and delivery settings, including in vivo models, and directly quantify mRNA abundance/functional lifetime in cells to further clarify the relationship between the susceptibility to deadenylation and protein output.
Experimental section
Synthesis of modified dinucleotides
Solid-phase synthesis was performed using an AKTA Oligopilot synthesizer. Reagents and solvents were sourced commercially. Five phosphoramidite equivalents were used for each coupling step. Oligonucleotides were synthesized on a 5 µmol scale.Cleavage from the solid support with simultaneous deprotection was performed using AMA (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 concentrated aqueous ammonia and concentrated aqueous methylamine) at 50 °C for 1 h, followed by filtration (to remove solid support particles) and freeze-drying. Final deprotection of the 2′-OH group was performed using triethylamine trihydrofluoride in DMSO at 60 °C for 4 h. After precipitation in n-butanol with sodium acetate (final concentration 75 µM), analogs were purified using an Agilent Tech. Series 1200 HPLC with a Clarity Oligo-RP C-18 column (150 × 10 mm, flow rate 5 mL min−1) in triethylammonium acetate (TEAAc) buffer with a linear acetonitrile gradient, coupled with UV-vis detection at 260 nm. The structure and homogeneity of the synthesized dinucleotides were confirmed using high-resolution mass spectrometry with negative electrospray ionization.
1 concentrated aqueous ammonia and concentrated aqueous methylamine) at 50 °C for 1 h, followed by filtration (to remove solid support particles) and freeze-drying. Final deprotection of the 2′-OH group was performed using triethylamine trihydrofluoride in DMSO at 60 °C for 4 h. After precipitation in n-butanol with sodium acetate (final concentration 75 µM), analogs were purified using an Agilent Tech. Series 1200 HPLC with a Clarity Oligo-RP C-18 column (150 × 10 mm, flow rate 5 mL min−1) in triethylammonium acetate (TEAAc) buffer with a linear acetonitrile gradient, coupled with UV-vis detection at 260 nm. The structure and homogeneity of the synthesized dinucleotides were confirmed using high-resolution mass spectrometry with negative electrospray ionization.
mRNA synthesis and purification
mRNA encoding Gaussia luciferase was obtained using pJET_T7_GLuc_31A plasmid, which was previously digested with AarI restriction enzyme (ThermoFisher Scientific). The plasmid was obtained by cloning the T7 promoter sequence and coding sequence of Gaussia luciferase into pJET_luc_128A.52 In vitro transcription reaction (120 µL) was incubated at 37 °C for 3 h in transcription buffer (40 mM Tris–HCl pH 7.9, 20 mM MgCl2, 1 mM DTT, and 2 mM spermidine) with 0.125 mg mL−1 T7 RNA polymerase, 1 U µL−1 RiboLock RNase Inhibitor, 4 mM ATP/UTP/CTP, 2 mM GTP, 4 mM m7GpppAmpG (cap 1 analog), 0.002 U µL−1 inorganic pyrophosphatase and 40 ng µL−1 of the template. After 3 h of incubation, 1 U µL−1 DNase was added to the reaction mixture and the incubation was continued for another 30 min at 37 °C. The reaction mixture was then diluted 2× and crude mRNA was purified using Monarch RNA Cleanup Columns T2050 (New England BioLabs). Quality of the transcript was evaluated on native 1% 1× Tris–borate–EDTA (TBE) agarose gel and the concentration of the sample was determined spectrophotometrically.The same in vitro transcription protocol was used to obtain the eGFP mRNA. Template DNA for eGFP was prepared by AarI digestion (Thermo) of the pJET1.2_T7_Egfp_3′utr-beta-globin_A128.
The poly(A) tail of mRNA was elongated to approximately 150 adenosines using the poly(A) Polymerase Tailing Kit (LuciGen) and following the standard protocol suggested by the manufacturer. The reaction (∼400 µL) was incubated for 30 min at 37 °C in a buffer (50 mM Tris–HCl, pH 8.0, 0.25 M NaCl, 10 mM MgCl2) with 1 mM ATP, 1 U µL−1 RiboLock RNase Inhibitor, 0.8 U µL−1 poly(A) polymerase and 0.6 µg µL−1 of mRNA. The reaction was stopped by adding EDTA to a final concentration of 11 mM and purified again using the same cleanup columns as previously mentioned. The quality of the mRNA was evaluated on an agarose gel as described previously. Final purification was performed on an Agilent Technologies Series 1200 HPLC using RNASep™ Prep – RNA Purification Column (ADS Biotec) at 55 °C. A linear gradient of buffer B (0.1 M triethylammonium acetate, pH 7.0, and 50% acetonitrile) from 20% to 25% in buffer A (0.1 M triethylammonium acetate, pH 7.0) over 20 minutes at a 0.9 mL min−1 flow rate was used. mRNA was later precipitated from the eluted fractions using 1.2 volumes of isopropanol with 0.3 M NaOAc (pH 5.2) and 1 µL mL−1 of glycogen, followed by a washing step with 80% ethanol. mRNA pellets were dried under reduced pressure and dissolved in RNase-free water. The concentration of the final sample was determined spectrophotometrically.
Ligation protocols
CNOT7 degradation assay
For 14-mer oligonucleotide products, CNOT7 digestion experiments were conducted as described previously.34For modified mRNA products, poly(A) tail degradation (deadenylation) reaction (5 µL) was carried out in a reaction buffer (20 mM Tris, 20 mM DTT, 10 mM MgCl2, 100 mM KCl, pH 8.0) with 60 ng of the chosen mRNA and 0.2 µg µL−1 of CNOT753 at 37 °C for 30 min. The reaction was stopped with EDTA (12.5 mM) and a 10× dilution with water. The reaction products were separated electrophoretically on 7% PAA polyacrylamide gel with urea in TBE buffer. The gel was later stained with SYBR Gold prior to scanning.
Translation efficiency experiments in the rabbit reticulocyte lysate system
Translation efficiency experiments were carried out in the RRL system (Promega). 4 µL of RRL was diluted with 5 µL of buffer (40 µM amino acid mixture D-cysteine, 40 µM amino acid mixture D-methionine, 2 mM MgCl2, and 380 mM potassium acetate) and incubated for 1 h at 30 °C. Next, 1 µL of the chosen modified Gaussia Luciferase mRNA dilution (7.8–500 pMol) was added to 9 µL of the RRL/buffer mixture and incubated for another 1 h at 30 °C. The reaction was quenched by freezing in liquid nitrogen. To detect luminescence, 50 µL of 10 ng mL−1 of h-coelenterazine (NanoLight) in PBS buffer was added to 10 µL of thawed translation mixture. Luminescence was measured using a Synergy H1 (BioTek) microplate reader. Linear regression was fitted to the points corresponding to the dependence of luminescence on mRNA concentration. The slope coefficients (a) of the linear equations obtained from the linear fit corresponded to the translation efficiency characteristic of a given mRNA molecule. All data were normalized to the average translation efficiency of unmodified GLuc mRNA.Protein expression studies
HEK-293 cells (ATCC CRL-1573) and A549 cells (ATCC CCL-185) were grown in DMEM (Gibco) supplemented with 10% FBS, GlutaMax and 1% penicillin/streptomycin (Gibco) at 5% CO2 and 37 °C. The murine immature dendritic cell line JAWS II (ATCC CRL-11904) was grown in RPMI 1640 (Gibco) supplemented with 10% FBS, sodium pyruvate (Gibco), 1% penicillin/streptomycin, and 5 ng mL−1 GM-CSF (PeproTech) at 5% CO2 and 37 °C. In all experiments, the passage number for the cells ranged from 5 to 25. On the day of transfection, 104 cells were seeded in 100 µL of medium per well, without antibiotics, in a 96-well plate. The transfection mixture for each well contained 0.3 µL of the Lipofectamine MessengerMAX Transfection Reagent (Invitrogen) and 5 ng of mRNA in 10 µL of OptiMEM (Gibco). Transfection lasted 16 h. Gaussia Luciferase expression was determined at multiple time points by fully removing the medium from each well and replacing it with fresh medium once every 24 h for 4 days. Luminescence was detected by adding 50 µL of 10 ng mL−1 h-coelenterazine in PBS to 10 µL of cell culture medium. Luminescence was measured using a Synergy H1 (BioTek) microplate reader. Total protein expression for each mRNA over 4 days was reported as a mean value ± SD normalized to unmodified Gaussia Luciferase mRNA.Kinetics of reporter protein and mRNA decay in single cells
A549 cells were microinjected with either eGFP or eGFP-pApsAm D1 mRNA. Next, the concentration of eGFP was measured at subsequent time points. The measurements were performed using a NIKON A1 confocal microscope with the LSM Upgrade KIT from PicoQuant, consisting of three pulse laser lines and two single photon avalanche diode (SPAD) detectors. The images of single cells were acquired and stored as the .ptu files; one file per cell per time point. The registered files were analysed using the smICA software.54 First, each PTU file was filtered to subtract the signal from each fluorescence channel. Next, using the calibration data, we applied the smICA software to estimate the concentrations of fluorescence-labelled dextrans and eGFP at each pixel in the imaged cells. The values for all pixels were averaged, and the mean value was stored. The procedure was repeated at every time point. Finally, the time evolution of the normalised mean values of the eGFP concentration was fitted with the kinetic model given by the formula (eqn (3)), derived directly from eqn (1) and (2):
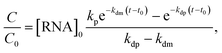 | (3) |
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting the findings of this study are available within the article and its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cb00033a.Additional data supporting the conclusions of this work are available from the corresponding authors upon reasonable request.
Acknowledgements
We thank Mariusz Czarnocki-Cieciura (International Institute of Molecular and Cell Biology, Warsaw, Poland) for providing purified recombinant human CNOT7 deadenylase. We also thank Karina Kwapiszewska, Grzegorz Bubak, Jarosław Michalski and Robert Hołyst (Institute of Physical Chemistry, Polish Academy of Sciences) for their help with mRNA degradation kinetics studies. This work was supported by the Virtual Research Institute Łukasiewicz Research Network—PORT Polish Center for Technology Development project “Horizon for Excellence in mRNA applications in immunOncology” [HERO] financed by the Polish Science Fund and the National Science Centre, Poland (2019/33/B/ST4/01843 to J. J. and 2018/31/B/ST5/03821 to J. K.).References
- J. Wolff, et al., Direct Gene Transfer into Mouse Muscle in Vivo, Science, 1990, 247, 1465–1468 CrossRef CAS PubMed.
- N. Pardi, M. J. Hogan, F. W. Porter and D. Weissman, mRNA vaccines—a new era in vaccinology, Nat. Rev. Drug Discovery, 2018, 17, 261–279 CrossRef CAS PubMed.
- B. Petsch, et al., Protective efficacy of in vitro synthesized, specific mRNA vaccines against influenza A virus infection, Nat. Biotechnol., 2012, 30, 1210–1216 CrossRef CAS PubMed.
- N. Pardi, et al., Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination, Nature, 2017, 543, 248–251 CrossRef CAS PubMed.
- L. Zangi, et al., Modified mRNA directs the fate of heart progenitor cells and induces vascular regeneration after myocardial infarction, Nat. Biotechnol., 2013, 31, 898–907 CrossRef CAS PubMed.
- C. G. Perez-Garcia, et al., Development of an mRNA replacement therapy for phenylketonuria, Mol. Ther. Nucleic Acids, 2022, 28, 87–98 CrossRef CAS PubMed.
- F. P. Polack, et al., Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine, N. Engl. J. Med., 2020, 383, 2603–2615 CrossRef CAS PubMed.
- L. R. Baden, et al., Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine, N. Engl. J. Med., 2021, 384, 403–416 CrossRef CAS PubMed.
- S. Holtkamp, et al., Modification of antigen-encoding RNA increases stability, translational efficacy, and T-cell stimulatory capacity of dendritic cells, Blood, 2006, 108, 4009–4017 CrossRef CAS PubMed.
- A. Thess, et al., Sequence-engineered mRNA Without Chemical Nucleoside Modifications Enables an Effective Protein Therapy in Large Animals, Mol. Ther., 2015, 23, 1456–1464 CrossRef CAS PubMed.
- J. Stepinski, C. Waddell, R. Stolarski, E. Darzynkiewicz and R. E. Rhoads, Synthesis and properties of mRNAs containing the novel ‘anti-reverse’ cap analogs 7-methyl(3′-O-methyl)GpppG and 7-methyl (3′-deoxy)GpppG, RNA, 2001, 7, 1486–1495 CAS.
- G. Kudla, L. Lipinski, F. Caffin, A. Helwak and M. Zylicz, High Guanine and Cytosine Content Increases mRNA Levels in Mammalian Cells, PLoS Biol., 2006, 4, e180 CrossRef PubMed.
- T. Spiewla, et al., PolyA tail segmentation improves the stability of the template DNA and increases the translatability of in vitro transcribed mRNA, Nucleic Acids Res., 2026, 54, gkaf1412 CrossRef CAS PubMed.
- Z. Trepotec, J. Geiger, C. Plank, M. K. Aneja and C. Rudolph, Segmented poly(A) tails significantly reduce recombination of plasmid DNA without affecting mRNA translation efficiency or half-life, RNA, 2019, 25, 507–518 CrossRef CAS PubMed.
- M. Diken, L. M. Kranz, S. Kreiter and U. Sahin, mRNA: A Versatile Molecule for Cancer Vaccines, Curr. Issues Mol. Biol., 2017, 22, 113–128 CrossRef PubMed.
- P. G. Coulie, B. J. Van den Eynde, P. van der Bruggen and T. Boon, Tumour antigens recognized by T lymphocytes: at the core of cancer immunotherapy, Nat. Rev. Cancer, 2014, 14, 135–146 CrossRef CAS PubMed.
- Ö. Türeci, et al., Targeting the Heterogeneity of Cancer with Individualized Neoepitope Vaccines, Clin. Cancer Res., 2016, 22, 1885–1896 CrossRef PubMed.
- N. B. Y. Tsui, E. K. O. Ng and Y. M. D. Lo, Stability of Endogenous and Added RNA in Blood Specimens, Serum, and Plasma, Clin. Chem., 2002, 48, 1647–1653 CrossRef CAS.
- A. Geall, et al., Nonviral delivery of self-amplifying RNA vaccines, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 14604–14609 CrossRef CAS PubMed.
- C.-Y. A. Chen, N. Ezzeddine and A.-B. Shyu, Messenger RNA half-life measurements in mammalian cells, Methods Enzymol., 2008, 448, 335–357 CAS.
- G. Caulier, J. Siblini, L. Sène, F. Mauxion and B. Séraphin, The CCR4–NOT complex: a multifaceted sensor of molecular signals instructing eukaryotic mRNA translation and stability, Nucleic Acids Res., 2025, 53, gkaf1401 CrossRef CAS PubMed.
- M. A. Collart, L. Audebert and M. Bushell, Roles of the CCR4-Not complex in translation and dynamics of co-translation events, Wiley Interdiscip. Rev.: RNA, 2024, 15, e1827 CrossRef CAS PubMed.
- T. Raisch and E. Valkov, Regulation of the multisubunit CCR4-NOT deadenylase in the initiation of mRNA degradation, Curr. Opin. Struct. Biol., 2022, 77, 102460 CrossRef CAS PubMed.
- Y. Liu, et al., Poly(A) tail length is a major regulator of maternal gene expression during the mammalian oocyte-to-embryo transition, bioRxiv, 2021, preprint, bioRxiv.2021.08.29.458052 DOI:10.1101/2021.08.29.458052.
- M. Dreyfus and P. Régnier, The Poly(A) Tail of mRNAs: Bodyguard in Eukaryotes, Scavenger in Bacteria, Cell, 2002, 111, 611–613 CrossRef CAS PubMed.
- S. Liu, et al., Classification and function of RNA–protein interactions, Wiley Interdiscip. Rev.: RNA, 2020, 11, e1601 CrossRef CAS PubMed.
- A. L. Nicholson and A. E. Pasquinelli, Tales of Detailed Poly(A) Tails, Trends Cell Biol., 2019, 29, 191–200 CrossRef CAS PubMed.
- J. Liu and F. Lu, Beyond simple tails: poly(A) tail-mediated RNA epigenetic regulation, Trends Biochem. Sci., 2024, 49, 846–858 CrossRef CAS PubMed.
- L. Anhäuser, S. Hüwel, T. Zobel and A. Rentmeister, Multiple covalent fluorescence labeling of eukaryotic mRNA at the poly(A) tail enhances translation and can be performed in living cells, Nucleic Acids Res., 2019, 47, e42 CrossRef PubMed.
- K. J. Westerich, et al., Bioorthogonal mRNA labeling at the poly(A) tail for imaging localization and dynamics in live zebrafish embryos, Chem. Sci., 2020, 11, 3089–3095 RSC.
- A. Aditham, et al., Chemically Modified mocRNAs for Highly Efficient Protein Expression in Mammalian Cells, ACS Chem. Biol., 2021, 17(12), 3352–3366 CrossRef PubMed.
- S. Son, M. Park, J. Kim and K. Lee, ACE mRNA (Additional Chimeric Element incorporated IVT mRNA) for Enhancing Protein Expression by Modulating Immunogenicity, Adv. Sci., 2024, 11, 2307541 CrossRef CAS PubMed.
- Y. Liu, et al., Remodeling of maternal mRNA through poly(A) tail orchestrates human oocyte-to-embryo transition, Nat. Struct. Mol. Biol., 2023, 30, 200–215 CrossRef CAS PubMed.
- O. Perzanowska, M. Smietanski, J. Jemielity and J. Kowalska, Chemically modified poly(A) analogs targeting PABP: structure activity relationship and translation inhibitory properties, Chem. – Eur. J., 2022, 28(42), e202201115 CrossRef CAS PubMed.
- J. A. W. Stowell, et al., Phosphorylation-dependent tuning of mRNA deadenylation rates, Nat. Struct. Mol. Biol., 2025, 1–8 Search PubMed.
- H. Wang, et al., Crystal structure of the human CNOT6L nuclease domain reveals strict poly(A) substrate specificity, EMBO J., 2010, 29, 2566–2576 CrossRef CAS PubMed.
- X. Zhang, et al., PARN deadenylase is involved in miRNA-dependent degradation of TP53 mRNA in mammalian cells, Nucleic Acids Res., 2015, 43, 10925–10938 CrossRef CAS PubMed.
- J. Wolf and L. A. Passmore, mRNA deadenylation by Pan2–Pan3, Biochem. Soc. Trans., 2014, 42, 184–187 CrossRef CAS PubMed.
- J. Lim, et al., Mixed tailing by TENT4A and TENT4B shields mRNA from rapid deadenylation, Science, 2018, 361, 701–704 CrossRef CAS PubMed.
- P. M. Burgers, F. Eckstein and D. H. Hunneman, Stereochemistry of hydrolysis by snake venom phosphodiesterase, J. Biol. Chem., 1979, 254, 7476–7478 CrossRef CAS PubMed.
- G. Slim and M. J. Gait, Configurationally defined phosphorothioate-containing oligoribonucleotides in the study of the mechanism of cleavage of hammerhead ribozymes, Nucleic Acids Res., 1991, 19, 1183–1188 CrossRef CAS PubMed.
- S. Maguire, G. J. S. Lohman and S. Guan, A low-bias and sensitive small RNA library preparation method using randomized splint ligation, Nucleic Acids Res., 2020, 48, e80 CrossRef CAS PubMed.
- E. Wahle and G. S. Winkler, RNA decay machines: deadenylation by the Ccr4-not and Pan2-Pan3 complexes, Biochim. Biophys. Acta, 2013, 1829, 561–570 CrossRef CAS PubMed.
- D. Mostafa, et al., Essential functions of the CNOT7/8 catalytic subunits of the CCR4-NOT complex in mRNA regulation and cell viability, RNA Biol., 2020, 17, 403–416 CrossRef CAS PubMed.
- B. Harrison and S. B. Zimmerman, Polymer-stimulated ligation: enhanced ligation of oligo- and polynucleotides by T4 RNA ligase in polymer solutions, Nucleic Acids Res., 1984, 12, 8235–8251 CrossRef CAS PubMed.
- B. A. Tannous, Gaussia luciferase reporter assay for monitoring biological processes in culture and in vivo, Nat. Protoc., 2009, 4, 582–591 CrossRef CAS PubMed.
- K. Karikó, H. Muramatsu, J. Ludwig and D. Weissman, Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA, Nucleic Acids Res., 2011, 39, e142 CrossRef PubMed.
- G. Rajivgandhi, et al., Enhanced anti-cancer activity of chitosan loaded Morinda citrifolia essential oil against A549 human lung cancer cells, Int. J. Biol. Macromol., 2020, 164, 4010–4021 CrossRef CAS PubMed.
- A. Heiser, et al., Autologous dendritic cells transfected with prostate-specific antigen RNA stimulate CTL responses against metastatic prostate tumors, J. Clin. Invest., 2002, 109, 409–417 CrossRef CAS PubMed.
- J. M. Jacobson, et al., Dendritic Cell Immunotherapy for HIV-1 Infection Using Autologous HIV-1 RNA: A Randomized, Double-Blind, Placebo-Controlled Clinical Trial, J. Acquired Immune Defic. Syndr., 2016, 72, 31–38 CrossRef CAS PubMed.
- P. Thomas and T. G. Smart, HEK293 cell line: a vehicle for the expression of recombinant proteins, J. Pharmacol. Toxicol. Methods, 2005, 51, 187–200 CrossRef CAS PubMed.
- M. Warminski, et al., Amino-Functionalized 5′ Cap Analogs as Tools for Site-Specific Sequence-Independent Labeling of mRNA, Bioconjugate Chem., 2017, 28, 1978–1992 CrossRef CAS PubMed.
- D. Strzelecka, et al., Phosphodiester modifications in mRNA poly(A) tail prevent deadenylation without compromising protein expression, RNA, 2020, 26, 1815–1837 CrossRef CAS PubMed.
- T. Kalwarczyk, et al., smICA: an open source repository for mapping the concentration of fluorescently labeled molecules in living cells on the basis of confocal imaging combined with time-correlated single-photon counting, arXiv, 2024, preprint, arXiv:2410.00532 DOI:10.48550/arXiv.2410.00532.
| This journal is © The Royal Society of Chemistry 2026 |