 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Post-translational modifications of silk proteins
Kota
Nomura
a and
Keiji
Numata
 *ab
*ab
aDepartment of Material Chemistry, Graduate School of Engineering, Kyoto University, Kyoto 615-8510, Japan. E-mail: keiji.numata@riken.jp
bBiomacromolecules Research Team, RIKEN Center for Sustainable Resource Science, Saitama 351-0198, Japan
First published on 5th March 2026
Abstract
Post-translational modifications (PTMs) endow silk proteins with chemical diversity that governs their higher-order assembly, hydration, and covalent connectivity. This review highlights the principal PTMs that define silk protein function, including hydroxylation, glycosylation, phosphorylation, and covalent crosslinking. We also describe their contributions to protein structural stability and mechanical properties. Recent advances in proteomics have begun to reveal low-abundance PTMs, whereas synthetic biology and bioorthogonal chemistry enable the programmed installation of modifications to tune physicochemical properties. Understanding and harnessing these chemistries provides a foundation for the predictive design of next-generation protein-based materials at the interface of chemical biology and materials science.
1. Introduction
Silk proteins are fibrous structural biomacromolecules produced by diverse organisms, including spiders, silkworms, and caddisflies, and are well known for forming fibers with exceptional mechanical performance, such as high tensile strength, toughness, and environmental resilience.1–3 These properties arise from the hierarchical organization of silk proteins, spanning amino acid sequence design, self-assembly, and higher-order structural transitions during fiber formation.Importantly, the performance of silk fibers is governed not only by their primary amino acid sequences but also by the chemical diversity imparted by post-translational modifications (PTMs),4 such as phosphorylation,5,6 proline hydroxylation,7 and dityrosine cross-links.8 By modulating intermolecular interactions, hydration states, higher-order folding, and fibril assembly, these PTMs play critical roles in directing silk protein assembly and tuning the final mechanical and functional properties of the resulting materials.
In this review, we summarize PTMs identified in silk proteins and discuss how these modifications contribute to silk-material properties. Integrating biochemical, structural, and materials-science perspectives, we highlight silk PTMs as a distinctive class of modifications that bridge molecular chemistry and macroscopic material function.
2. Classification of PTMs in silk proteins
2.1. Hydroxylation
Enzymatic hydroxylation is a hallmark PTM of silk proteins (Fig. 1A).9 Prolyl 4-hydroxylase (P4H) converts certain Pro residues to 4-hydroxyproline (Hyp).10 These enzymes reside within the lumen of the rough endoplasmic reticulum (ER) and act on nascent polypeptides. The Hyp residues introduced by ER-localized P4H greatly stabilize fibrillar architectures via strengthened hydrogen bonding interactions and increased hydration.11 In some structural or adhesive contexts, tyrosine (Tyr) residues are hydroxylated to 3,4-dihydroxyphenylalanine (DOPA).12,13 This reaction is catalyzed by phenoloxidase enzymes such as tyrosinase or insect tyrosine hydroxylase in the secretory pathway or extracellularly.13,14 The resulting DOPA provides a catechol functional group that enables metal coordination and redox-driven curing, as in marine mussel adhesives.15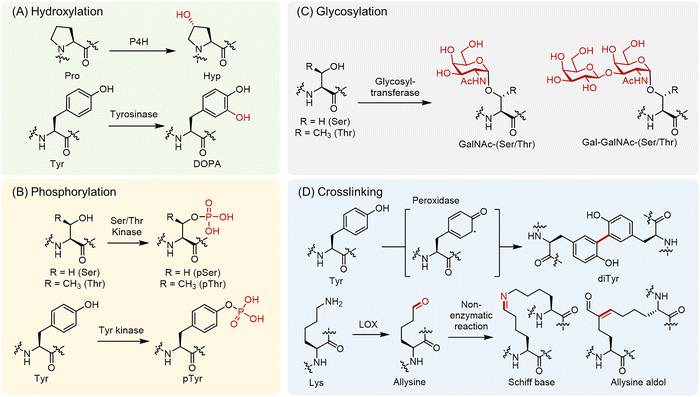 | ||
| Fig. 1 Structures of representative PTMs of silk proteins. Panel A shows hydroxylation reactions, including the conversion of Pro to Hyp by P4H and Tyr to DOPA by tyrosinase.9–15 Panel B depicts phosphorylation, in which the Ser/Thr hydroxyl group or the phenolic group of Tyr are modified by protein kinases.16,17 Panel C illustrates O-glycosylation, including the addition of simple sugars such as GalNAc, Gal-GalNAc to Ser/Thr.18,19 Panel D presents cross-linking reactions, including peroxidase-mediated diTyr formation and the LOX-mediated oxidation of Lys to allysine.20–23 | ||
2.2. Phosphorylation
Serine/threonine (Ser/Thr) protein kinases introduce phosphate groups to yield phosphoserine (pSer) or phosphothreonine (pThr).16 Similarly, Tyr kinases introduce phosphate groups to produce phosphotyrosine (pTyr) (Fig. 1B). In proteins, phosphorylation typically occurs after translation in the secretory pathway, for example, by a Golgi-resident kinase.17 The fibroin heavy chain of the silkworm Bombyx mori contains multiple pSer residues in its repeat regions,5 and in spider silk,6 phosphorylation of the terminal domain is observed.8 In silk fibroin, phosphorylation is thought to modulate the storage solubility of the protein dope (the highly concentrated aqueous silk-protein solution stored in the silk gland prior to spinning) and to bias β-sheet formation during spinning. Changes in phosphorylation status act as a reversible switch, maintaining solubility in the gland prior to spinning and promoting assembly during spinning.2.3. Glycosylation
Glycosylation generally occurs in two main forms, with O-linked glycosylation occurring on Ser/Thr residues and N-linked glycosylation occurring on Asn residues. N-Linked glycosylation is less frequently highlighted in the context of silk proteins, as O-linked modifications are more prominent18 (Fig. 1C). For example, O-glycosylation, such as in the saccharides α/β-D-N-acetyl galactosamine (GalNAc) and glucosamine (GlcNAc), occurs at Ser and Thr residues on certain adhesive proteins, such as aggregate silk spidroin.19 Glycosylation contributes to the hydration and adhesion properties of the substrate.2.4. Enzymatic crosslinking
Lysyl oxidase (LOX) is a Cu-dependent amine oxidase secreted into the extracellular space as a proenzyme that is activated outside the cell (Fig. 1D).20–22 It acts post-translationally in the extracellular matrix (ECM) by oxidatively deaminating Lys residues to yield reactive aldehydes (e.g., allysine), which can form covalent cross-links through spontaneous condensation reactions, including Schiff base formation with Lys residues and aldol condensation with other allysine residues.232.5. Oxidative modifications
Oxidative coupling of Tyr side chains yields dityrosine (diTyr) and trityrosine (triTyr) cross-links (Fig. 1D).24,25 Such cross-links can form nonenzymatically upon interaction with reactive oxygen species, but in biological systems, they are often formed enzymatically by oxidases (e.g., peroxidases or laccase-type phenoloxidases) secreted into the ECM.26 In insect cuticles, resilin, and fibroin, peroxidase oxidizes Tyr residues to form Tyr radicals that couple to form diTyr cross-links.27 These oxidative Tyr linkages are integral to the unique elastic recovery of resilin, effectively creating an amorphous rubber-like network.28,292.6. Other nonenzymatic modifications
A variety of spontaneous chemical degradation products accumulate over time and with environmental exposure. Examples include deamidated asparagine and glutamine (Asn/Gln) residues,30 oxidized methionine (Met) to sulfoxide,31 and disulfide bond reshuffling or cleavage of cysteine (Cys) residues.32 These products are formed in the presence of physicochemical stimuli (pH, concentration, reactive oxygen species, and UV irradiation). These changes are typically detrimental to material performance. For instance, deamidation alters the charge and conformation of the protein, leading to protein brittleness, and Met oxidation or unwanted disulfide changes in Cys can reduce protein elasticity or stability.333. PTMs in silk proteins and their function
3.1. PTMs in spider silks
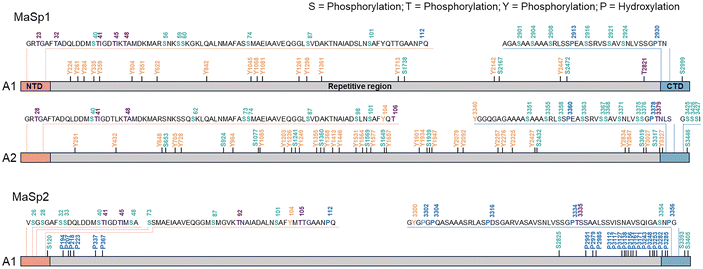 | ||
| Fig. 2 Comprehensive map of the PTMs of MaSp1 and MaSp2 of Trichonephila clavata. The numbered residues correspond to phosphorylated Ser (S, light green), Thr (T, purple), or Tyr (Y, orange) residues and hydroxylated Pro residues (P, blue). NTD, N-terminal domain; CTD, C-terminal domain.6 Adapted from ref. 4 under the Creative Commons CC BY 4.0 license. | ||
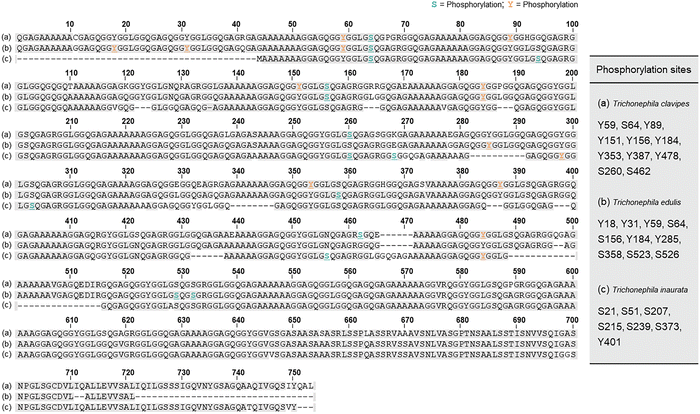 | ||
| Fig. 3 Multiple alignment of the primary sequences of MaSp1 from (a) Trichonephila clavipes, (b) Trichonephila edulis, and (c) Trichonephila inaurata. The numbered residues correspond to phosphorylated Ser residues (S, light green) and phosphorylated Tyr residues (Y, orange). PTMs of MaSp1 revealed by MASCOT.8 Adapted with permission from ref. 8 Copyright 2016, Elsevier. | ||
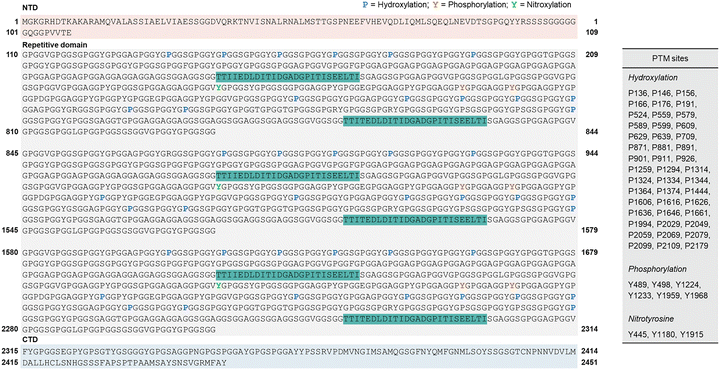 | ||
| Fig. 4 Representative sequences of flagelliform silk from Trichonephila clavipes. The numbered residues correspond to phosphorylated Tyr residues (Y, orange) and hydroxylated Pro residues (P, blue); the spacer domain is indicated in green. The NTD domain is indicated in light red. The repetitive domain is indicated in gray. The CTD domain is indicated in light blue. PTMs of flagelliform silk revealed by MASCOT.39–41 Adapted from ref. 41 under the Creative Commons CC BY 4.0 license. | ||
Tubuliform silk in the outer egg sac layer is characterized by a very high Ser content and low Gly content relative to those of other spider silks.60 This distinctive amino acid profile is thought to account for its propensity to form rigid fibers through extensive β-sheet crystallization. Given the high Ser content, it is plausible that the principal egg sac proteins harbor pSer. However, definitive evidence of PTMs is lacking. For example, proteomic studies on the egg sac silk of Latrodectus hesperus have identified TuSp1 as the major fibroin together with additional egg case proteins (ECPs), but evidence of phosphorylation or other modifications has not yet been reported.62,63 Even so, because the egg sac presents a robust barrier that protects eggs from UV radiation and microbes, it has been hypothesized that diTyr cross-links occur after egg sac formation to stabilize the fibers. Evidence for such oxidative crosslinking and pigmentation has been reported in silkworm egg sacs and cocoons.64,65
Piriform silk is a specialized protein that provides strong adhesion between silk threads and substrates. Rather than being a thread, it is secreted as a pad-like deposit and must harden and set extremely quickly.61 However, there is currently no direct evidence of PTMs in PiSp proteins.
The role of the aciniform silk in enveloping prey and lining the inner layer of the egg sacs suggests a balance between extensibility and moderate strength.59 Aciniform fibroins possess Gly/Ala-rich repetitive motifs akin to those in dragline silk. Some analyses have reported a slightly greater proportion of acidic residues in aciniform silk than in dragline. It was shown that an aciniform spidroin (AcSp1) is a key component of the small diameter fibers in egg sacs as well as in prey wrap silk, emphasizing that aciniform silk contributes to both prey capture and protection.59,66 Recent research on Argiope argentata has shown that the dope adopts an α-helical beads-on-a-string conformation in which Ser/Ala/Gly-rich linkers adopt β-sheet structures upon spinning67 and that AcSp2 yields recombinant fibers extending to approximately 160%, underscoring sequence-level control of AcSp mechanics.68
3.2. PTMs in silkworm (Bombyx mori) silk
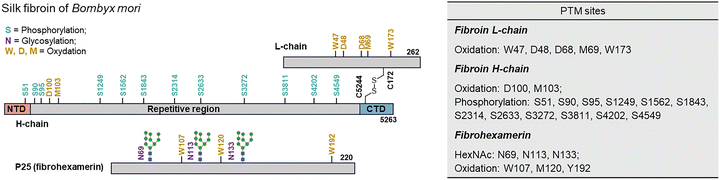 | ||
| Fig. 5 The modifications of silkworm silk fibroin from Bombyx mori. The numbered residues correspond to phosphorylated Ser residues (S, light green), N-glycosylated Asn residues (N, purple) and oxidized Trp, Asp, and Met residues (W, N, M, yellow). The NTD domain is indicated in light red. The repetitive domain is indicated in gray. The CTD domain is indicated in light blue. PTMs of silk protein revealed by MASCOT.67,68,73 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P25 ratio of 6
P25 ratio of 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 still exists, but because the P25 glycans are exposed, most of it is degraded by quality control machinery. In the normal system where the L-chain is present, the glycans of P25 are masked by the L-chains, which both protects them from proteases and maintains the higher-order structure of the complex. Thus, N-glycosylation of P25 is essential for the formation of the fibroin unit, and interactions between its glycans and the L-chains likely contribute to the intracellular assembly and secretion of the large complex.75 Another interesting point about P25 is that multiple isoforms exist. These differences have been reported to arise not only from subtle variations in glycan structure but also from reversible phosphorylation events.
1 still exists, but because the P25 glycans are exposed, most of it is degraded by quality control machinery. In the normal system where the L-chain is present, the glycans of P25 are masked by the L-chains, which both protects them from proteases and maintains the higher-order structure of the complex. Thus, N-glycosylation of P25 is essential for the formation of the fibroin unit, and interactions between its glycans and the L-chains likely contribute to the intracellular assembly and secretion of the large complex.75 Another interesting point about P25 is that multiple isoforms exist. These differences have been reported to arise not only from subtle variations in glycan structure but also from reversible phosphorylation events.
Sericins are another important group of proteins that make up the cocoon, functioning as both an adhesive and a protective coating for the silk filament.76 Sericins consist of multiple subunits of different sizes (Ser1, Ser2, Ser3, etc.), all of which are characterized by high contents of hydroxy amino acids and repetitive sequences. Sericins are secreted and deposited around fibroin in the middle section of the silk gland and harbor typical O-linked glycans (α-Gal and α-GalNAc).76 These glycans enhance the adhesive properties of the sericin, strengthening adhesion between the cocoon and surrounding twigs after spinning. A notable PTM of sericins is the formation of cross-links. In the domesticated silkworm, phenoloxidases are released immediately after cocoon formation; thus, domesticated sericin remains uncrosslinked within the cocoon and can be readily extracted with warm water. In contrast, in wild silkmoths, sericins sometimes undergo partial oxidative polymerization during cocooning or cocoon sheet formation, producing an insoluble, resin-like matrix. For example, in saturniid cocoons, laccase-like enzymes are secreted at the final larval instar, oxidizing the Tyr residues of sericin and exogenous tannins to form quinone crosslinks, thereby causing the cocoon to darken and become insoluble.
3.3. PTMs in caddisfly silk
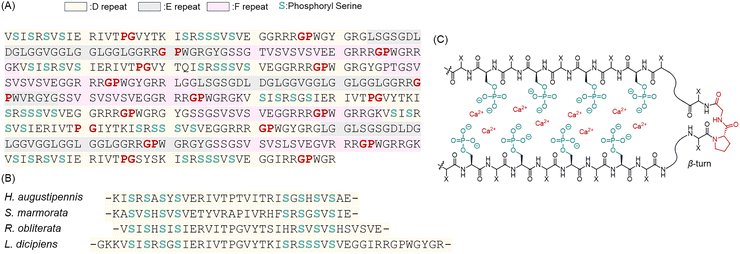 | ||
| Fig. 6 (A) Partial amino acid sequences of heavy-chain fibroin protein from the caddisfly silk. Phosphorylated (SX)4 motifs found in the D-repeat (shown in light yellow) are proposed to exist in calcium-phosphoserine sheets via pSer (light green). In support of the sheet model, proline residues often found in β-turn secondary structures appear shortly after every (SX)4 region. (B) Sequence correlation of D-repeat motifs across four distinct species of caddisfly (Hydropsyche augustipennis, Stenopsyche marmorata, Rhyacophila obliterata, and Limnephilus decipiens).78 Adapted with permission from ref. 78 Copyright 2013, American Chemical Society. (C) Schematic illustration of calcium–phosphate interactions within phosphorylated β-sheets.79,80 Adapted with permission from ref. 79 Copyright 2010, American Chemical Society. | ||
3.4. PTMs of other insect silks
Beyond spiders and silkworms, many other insects produce silk-like protein fibers.91 Bagworm larvae build tubular cases by stitching together small twigs and leaf fragments with their own silk. Bagworm silk presumably consists of large proteins, but the details have not been well studied. Recent transcriptomic analysis of Eumeta variegata revealed an extremely long, repetitive silk fibroin gene (19 kbp) encoding a Gly/Ala-rich protein, supporting the presence of heavy-chain fibroin analogs in bagworms.92 Given that it is used to build shelters and is exposed to wind and rain in terrestrial environments, it is possible that bagworm silk undergoes phenolic crosslinking. Indeed, the interior of bagworm cases sometimes shows brown discoloration and hardening of the threads, which may result from reactions between plant-derived tannins or microbially degraded lignin components.93Many hymenopteran larvae, including numerous species of bees, wasps, and ants, spin cocoons during their final instar stage before pupation.94,95 These Hymenopteran silks have distinct origins from Lepidopteran fibroin and can possess properties akin to those of cuticular proteins.96 Particularly noteworthy is the cocoon thread of the willow sawfly, Nematus oligospilus, which was recently reported to be an entirely new type of silk that forms a collagen-like triple helix.97 Sutherland and colleagues reported that the N. oligospilus cocoon comprises three small collagen proteins that carry Gly-Pro-Pro repeats and self-assemble into triple-helical fibers. Strikingly, despite being an insect, Hyp was detected at Pro residues in this type of collagen silk. The resulting willow-sawfly cocoon is a novel material that combines tensile strength with flexibility. However, few detailed analyses have been performed on other Hymenopteran silks, but these proteins tend to be relatively large and rich in Gly and Tyr.98 For instance, the cocoon silks of nonsocial wasps (Ichneumonoidea) are fibroin-like and composed largely of Gly, Ala, and Ser residues.98
Overall, PTMs of insect silks are extraordinarily diverse and have been optimized for each ecological niche. Unique PTM strategies not found in silkworms and spiders have also evolved; for instance, the adoption of a collagen in Hymenoptera and hybrid crosslinking with plant polyphenols in bagworms and caddisflies. A comprehensive survey of PTMs in less-studied silks should allow a holistic understanding of the gamut of material strategies devised by life and their translation into human use.
4. Applications to materials science
4.1. Introducing PTMs via synthetic biology and chemical synthesis approaches
By strategically manipulating the PTMs of structural proteins, we can finely tune their material properties, including mechanical strength, extensibility, thermal stability and heat resistance, water resistance, and self-healing.99 To append PTMs to material proteins, a combination of synthetic-biological and chemical approaches is indispensable.100 Advances in genetic engineering have allowed the introduction of artificial modification acceptor sites into protein primary sequences. A classic example is the incorporation of unnatural amino acids (UAAs) via genetic code expansion and amber codon reassignment.101,102 Using such techniques, we can insert UAAs bearing novel functional groups at defined positions in a protein. Examples include alkenes, azides, alkynes, ketones, and phenylalanine derivatives. This approach endows the material with chemical reactivities that are absent in biological systems. Notably, natural protein materials often contain nonstandard amino acids that are PTM products, which are crucial for function. However, conventional expression hosts such as E. coli often cannot install these PTMs. Introducing artificial PTMs by genetically encoding UAAs has thus become a powerful approach. Recent work has achieved genetic code expansion to install DOPA into protein polymers.103 Catechols are the adhesive residues in mussel adhesive proteins. By introducing catechols into engineered proteins, we can design high-performance materials with underwater adhesion and self-healing capabilities.104 In this way, genetic methods for incorporating UAAs and PTM-like modifications are dramatically expanding the repertoire of protein–material design. Bioorthogonal chemistry also provides powerful artificial modification routes. Cell-permeable reagents can be conjugated to UAAs via click reactions, enabling covalent attachment of polymers. For instance, p-azidophenylalanine introduced by genetic encoding can undergo a click reaction with alkyne-bearing compounds to append polyethylene glycol (PEG) chains, improving the durability and water retention of protein hydrogels (Fig. 7I).105 Other reports introduce UAAs bearing spiro moieties or thiol groups and induce gelation by the specific crosslinking with externally added crosslinkers.106 While the applications of these techniques to the modification of silk proteins remain limited, they are expected to become important tools.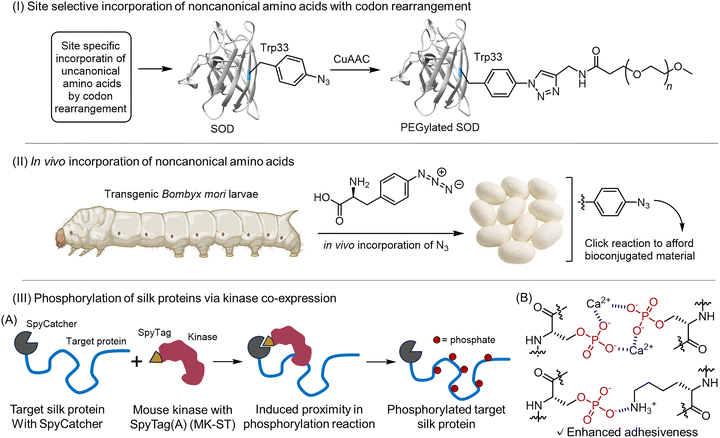 | ||
| Fig. 7 Strategies for engineering PTMs into structural proteins. (I) Site-selective incorporation using codon rearrangement: genetic codon editing introduces azido-phenylalanine at Trp residues, followed by click conjugation with PEG.105 (II) In vivo incorporation of noncanonical amino acids: transgenic silkworms fed azido-phenylalanine produce silk fibroin carrying azido groups.107 (III) (A) Induced proximity approach for silk protein phosphorylation. Schematic representation of aggregate spider silk. (B) Scheme of a possible adhesion mechanism of phosphorylated silk supplemented with Ca2+ ions.108 Adapted from ref. 108 under the Creative Commons CC BY 4.0 license. | ||
Beyond amber codon reassignment, residue-specific UAA incorporation has also been realized in multicellular systems. For example, Kojima and Teramoto demonstrated that transgenic silkworms fed phenylalanine analogs could produce silk fibroin carrying azide groups, yielding clickable silks that survive degumming and enable versatile bioorthogonal modifications (Fig. 7-II).107
Another recent approach uses proximity-assisted enzymatic PTM installation. Using a SpyCatcher/SpyTag system, silk proteins expressed in E. coli can be selectively recruited to a coexpressed kinase, greatly enhancing on-target phosphorylation while minimizing global toxicity (Fig. 7III(A)).108 This simple design enables high-density, multisite phosphorylation of dragline and aggregate silk, yielding recombinant phosphorylated silks with measurably improved adhesion and Ca2+-dependent cohesion properties (Fig. 7III(B)). Because the scaffold is modular, the same strategy could be adapted for other PTM enzymes to program PTM patterns in engineered fibrous proteins.
Furthermore, dehydroalanine (Dha) chemistry has been shown to be a versatile tag-and-modify platform for installing a wide spectrum of PTMs. These include phosphorylation, glycosylation, ubiquitination, and acetylation, all of which can be introduced into recombinant proteins under mild conditions (Fig. 8I).109 Such chemical methods are extremely powerful because they endow materials with functional groups that biology does not natively create, bridging synthetic polymer chemistry with protein material approaches.
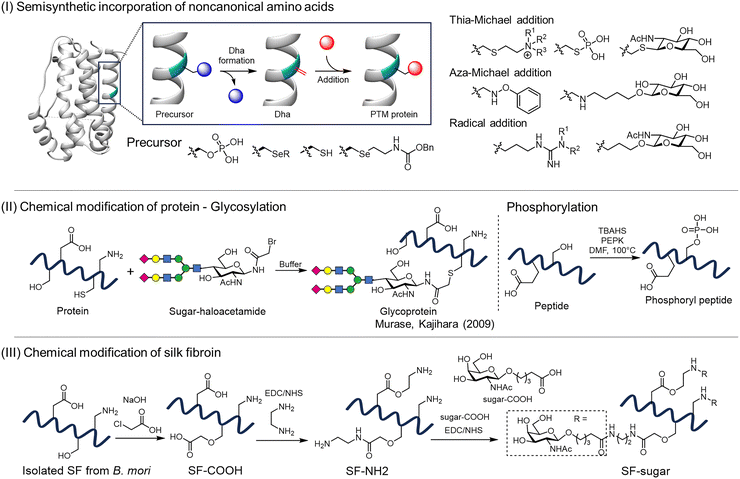 | ||
| Fig. 8 Chemical strategies for introducing PTMs into proteins. (I) Semisynthetic incorporation of noncanonical amino acids: precursor residues are genetically introduced and subsequently converted to dehydroalanine (Dha), which serves as a reactive handle for Michael addition or radical-based modification to install diverse PTMs.107 (II) Chemical modification of proteins and peptides: (a) site-selective glycosylation via the coupling of haloacetamide-activated glycans to Cys residues;112,113 and (b) chemical phosphorylation targeting Ser/Thr residues to overcome the challenge of competing nucleophiles such as Lys.114,115(III) Chemical modification of silk fibroin: global conjugation of sugars to carboxyl, amino, or hydroxyl groups on fibroin, yielding glycosylated silk materials.116 | ||
Rapidly advancing cell-free systems and secretion-based expression platforms are also useful for PTM installation. Cell-free protein synthesis (CFPS)110 using cell extracts or purified transcription-translation machinery eliminates the constraints of cell walls and viability, enabling synthesis with high concentrations of UAAs.111 CFPS also allows the coaddition of enzymes to install PTMs concurrently with translation. Indeed, studies using CFPS to macrocyclize RiPPs (ribosomally synthesized and post-translationally modified peptides) or to append artificial glycans to proteins have screened many enzyme variants within hours and identified mutants with superior glycosylation efficiency.112 By combining CFPS with high-throughput analysis, the process of PTM installation can now be rapidly optimized.113 Conversely, secretion using eukaryotic cell culture can favor spontaneous disulfide formation and near-native glycosylation by routing products through the secretory pathway. Efforts are underway to secrete spider silk proteins from yeast to obtain materials in which the stabilizing crosslinks required for higher-order structures are correctly formed.37
In addition, natural protein materials can be chemically modified after expression or synthesis. Elegant examples include the work of Kajihara and colleagues, who developed haloacetamide-based Cys-selective coupling to construct homogeneous glycoproteins with defined saccharide structures (Fig. 8II, left panel).114,115 Moreover, Kanai and colleagues recently reported catalytic phosphorylation of unprotected Ser/Thr and Tyr residues (Fig. 8II, right panel).116,117 Similarly, mucin-inspired engineering of silk fibroin revealed that covalent conjugation of sugars such as GalNAc to the side chain functionalities of Lys, Ser, Thr, Asp, and Glu inhibited Streptococcus mutans biofilm formation while preserving bacterial growth, effectively reproducing the antivirulence effect of mucin (Fig. 8III).118 This approach imparts new functions without re-expression, although the lack of site specificity and chemoselectivity yield heterogeneous modifications. Collectively, combining synthetic biological tools with chemical methods makes freely programmable PTM of silk protein design a practical reality. This approach opens paths to materials whose functions are absent in nature and to improved biomaterials.
Despite these advances, several limitations remain in current strategies for installing PTMs in silk proteins. Genetic code expansion and synthetic biology approaches can be constrained by incorporation efficiency and scalability, and may impose metabolic burden or toxicity in host organisms, particularly when introducing multiple or high-density modifications. Chemical modification strategies introduce heterogeneity in modification patterns and are difficult to control at precise stoichiometry. Addressing these challenges will be essential for translating programmable PTM strategies toward scalable and structurally well-defined silk-based materials.
4.2. Discovering unknown protein PTMs
The discovery and comprehensive understanding of novel PTMs directly propel the field of structural protein materials. Recent advances in proteomics have established highly sensitive MS methods for identifying low-abundance and previously unknown PTMs that were once difficult to detect.119 In particular, data-independent acquisition (DIA) enables comprehensive capture of peptide fragments in a sample with precise quantification and is being vigorously applied to PTM analysis.120 Because DIA acquires spectra without bias, modified peptides are recorded exhaustively, allowing site-specific global analyses of the phosphoproteome or glycoproteome.121,122 By combining DIA with deep learning-based in silico spectral libraries and scoring, the accuracy of PTM identification has improved dramatically.123 This now permits high-throughput, high-accuracy workflows that span detection, site localization, and characterization of combined modifications.Furthermore, accelerated and parallelized cross-linking mass spectrometry (XL-MS) is transforming the study of structural proteins.124 XL-MS covalently crosslinks proximal residues within proteins using reactive reagents for MS-based identification of the resulting cross-linked peptides. This approach reveals protein structures and interaction networks in solution. Recent work has succeeded in crosslinking entire cells to obtain proteome-wide distance constraints under native intracellular conditions. Bartolec reported the largest dataset to date, identifying 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 910 unique residue pairs derived from 91
910 unique residue pairs derived from 91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 709 cross-linked spectral matches across 4084 human proteins and 2110 protein–protein interactions.125 Importantly, these crosslinks capture proteins with their natural sequences, PTM states, subcellular locations, and cofactor-bound forms. They provide physiological snapshots that cannot be obtained with purified proteins alone. XL-MS data also serve to validate and refine machine learning structure predictions such as AlphaFold2,126 ushering in an era of systematic structural proteomics. Together, DIA-based proteomics and XL-MS provide a powerful platform for discovering unknown PTMs and clarifying their structural roles.
709 cross-linked spectral matches across 4084 human proteins and 2110 protein–protein interactions.125 Importantly, these crosslinks capture proteins with their natural sequences, PTM states, subcellular locations, and cofactor-bound forms. They provide physiological snapshots that cannot be obtained with purified proteins alone. XL-MS data also serve to validate and refine machine learning structure predictions such as AlphaFold2,126 ushering in an era of systematic structural proteomics. Together, DIA-based proteomics and XL-MS provide a powerful platform for discovering unknown PTMs and clarifying their structural roles.
To identify unknown PTMs, it is also crucial to learn from diverse organisms and ecologies. Modifications rare in a single species may emerge through cross-species comparisons. Researchers worldwide are accumulating PTM data from many organisms, and PTM databases are being continuously updated to organize this vast amount of information. A recent example is dbPTM 2025, a comprehensive resource that integrates >2.79 million experimentally validated PTM sites from 48 data sources and over 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 publications.127 dbPTM 2025 supports searches by species, PTM type, and modified residue. With an eye to disease research, the database also integrates large-scale cancer phosphoproteomics and kinase activity profiles, supporting PTM network analyses and comparisons across species and diseases. The development of such cross-species, comprehensive PTM databases is expected to greatly accelerate the identification of unusual PTMs and the understanding of their evolutionary conservation and diversity.
000 publications.127 dbPTM 2025 supports searches by species, PTM type, and modified residue. With an eye to disease research, the database also integrates large-scale cancer phosphoproteomics and kinase activity profiles, supporting PTM network analyses and comparisons across species and diseases. The development of such cross-species, comprehensive PTM databases is expected to greatly accelerate the identification of unusual PTMs and the understanding of their evolutionary conservation and diversity.
Applications of artificial intelligence (AI) to PTM research are likewise advancing rapidly. While deep learning structure prediction revolutionized the inference of protein structures, it initially ignored PTMs.128 Current efforts are underway to overlay PTM data onto predicted structures to evaluate changes in stability and to visualize structural fluctuations in PTM-bearing regions. In protein sequence design with generative AI, attention is turning to PTMs. The recently reported PTM-aware protein language model “PTM-Mamba,” trained on approximately 79![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 modified protein sequences (based on >3
000 modified protein sequences (based on >3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 PTM records), successfully embeds PTM-bearing sequences into high-dimensional spaces that capture modification-specific features.129 Unlike earlier language models that masked or ignored modified residues, PTM-Mamba encodes modified amino acids as dedicated tokens and integrates embeddings from large models such as ESM-2, enabling the model to represent functional and structural changes associated with PTMs. Visualization of the embedding space shows modified and unmodified versions of a protein located nearby yet separable, with clustering by PTM type. This is the first demonstration that deep learning can learn and represent the effects of PTMs on structure and function, opening the door to applications such as designing stabilizing mutations contingent upon a target PTM or predicting crosstalk among multiple PTMs. Complementary efforts include prompting Transformer-type models to predict PTM sites (PTMGPT) and, conversely, generating sequences expected to realize specified PTM patterns.130 AI excels at discovering hidden regularities in massive datasets. Data-driven PTM design support is now at the forefront, enabling discovery of unknown PTMs and prediction of surprising PTM combinations and their functions. However, applications of these AI-based approaches to silk PTMs remain limited at present, and most current efforts are still at a conceptual or exploratory stage. Future AI-driven approaches will become particularly valuable in addressing unresolved questions in silk PTMs. For example, quantitative prediction of phosphorylation site occupancy and stoichiometry from sequence and proteomic data could help distinguish structurally dominant modifications from low-abundance modifications. In addition, integrative computational frameworks combining proteomics, structural modeling, and materials simulations could clarify how specific PTMs influence solubility and assembly during spinning.
000 PTM records), successfully embeds PTM-bearing sequences into high-dimensional spaces that capture modification-specific features.129 Unlike earlier language models that masked or ignored modified residues, PTM-Mamba encodes modified amino acids as dedicated tokens and integrates embeddings from large models such as ESM-2, enabling the model to represent functional and structural changes associated with PTMs. Visualization of the embedding space shows modified and unmodified versions of a protein located nearby yet separable, with clustering by PTM type. This is the first demonstration that deep learning can learn and represent the effects of PTMs on structure and function, opening the door to applications such as designing stabilizing mutations contingent upon a target PTM or predicting crosstalk among multiple PTMs. Complementary efforts include prompting Transformer-type models to predict PTM sites (PTMGPT) and, conversely, generating sequences expected to realize specified PTM patterns.130 AI excels at discovering hidden regularities in massive datasets. Data-driven PTM design support is now at the forefront, enabling discovery of unknown PTMs and prediction of surprising PTM combinations and their functions. However, applications of these AI-based approaches to silk PTMs remain limited at present, and most current efforts are still at a conceptual or exploratory stage. Future AI-driven approaches will become particularly valuable in addressing unresolved questions in silk PTMs. For example, quantitative prediction of phosphorylation site occupancy and stoichiometry from sequence and proteomic data could help distinguish structurally dominant modifications from low-abundance modifications. In addition, integrative computational frameworks combining proteomics, structural modeling, and materials simulations could clarify how specific PTMs influence solubility and assembly during spinning.
Outlook
Protein–materials science centered on PTMs holds immense promise at the interface of life science and materials science. PTMs are molecular switches through which cells rapidly alter protein states in response to their environment. Leveraging them in material design enables new functional materials to respond to external cues. Although PTM research is rapidly expanding across molecular and cellular biology, our understanding of PTMs in silk proteins remains incomplete. Their diversity, context dependence, and often low stoichiometry make systematic characterization difficult, leaving many fundamental questions unresolved. Precisely controlling and observing PTM reactions within artificial materials could deepen our understanding of in vivo PTM networks. Through collaboration between data-driven computation and experimentation, the creation of intelligent protein materials with PTMs woven in at will could lead to transformative breakthroughs. Consequently, the fusion of materials and life sciences will serve as the driving force behind future advances in this field, establishing a new interdisciplinary frontier.Author contributions
K. N. and K. N. wrote and reviewed the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This research was financially supported by the MEXT Data Creation and Utilization-type MaTerial R&D project (JPMXP1122714694) (K. N.) and JST-COI-NEXT (JPMJPF2114) (K. N.).Notes and references
- K. Numata, Biomacromolecules, 2025, 26, 1393–1403 CrossRef CAS PubMed.
- K. Numata and D. L. Kaplan, Adv. Mater., 2025, 37, e2411256 CrossRef PubMed.
- T. Schiller and T. Scheibel, Commun. Mater., 2024, 5, 56 CrossRef CAS.
- A. Miserez, J. Yu and P. Mohammadi, Chem. Rev., 2023, 123, 2049–2111 CrossRef CAS PubMed.
- W. Q. Chen, H. Priewalder, J. P. John and G. Lubec, Proteomics, 2010, 10, 369–379 CrossRef CAS PubMed.
- H. C. Craig, A. D. Malay, F. Hayashi, M. Mori, K. Arakawa and K. Numata, MRS Bull., 2024, 49, 1192–1204 CrossRef CAS.
- J. E. Jenkins, M. S. Creager, E. B. Butler, R. V. Lewis, J. L. Yarger and G. P. Holland, Chem. Commun., 2010, 46, 6714–6716 RSC.
- J. R. dos Santos-Pinto, G. Lamprecht, W. Q. Chen, S. Heo, J. G. Hardy, H. Priewalder, T. R. Scheibel, M. S. Palma and G. Lubec, J. Proteomics, 2014, 105, 174–185 CrossRef CAS PubMed.
- A. M. Salo and J. Myllyharju, Exp. Dermatol., 2021, 30, 38–49 CrossRef CAS PubMed.
- J. Myllyharju, Matrix Biol., 2003, 22, 15–24 CrossRef CAS PubMed.
- J. Bella, B. Brodsky and H. M. Berman, Structure, 1995, 3, 893–906 CrossRef CAS PubMed.
- X. Ou, B. Xue, Y. Lao, Y. Wutthinitikornkit, R. Tian, A. Zou, L. Yang, W. Wang, Y. Cao and J. Li, Sci. Adv., 2020, 6, eabb7620 CrossRef CAS PubMed.
- H. G. Silverman and F. F. Roberto, Mar. Biotechnol., 2007, 9, 661–681 CrossRef CAS PubMed.
- S. Kim, G. Bae, M. Shin, E. Kang, T. Y. Park, Y. S. Choi and H. J. Cha, Biotechnol. J., 2021, 16, 2100216 CrossRef CAS PubMed.
- Y. Ding, Y. Cui, H. Xie, J. Liang, H. Liu, F. Yuan and T. Liu, Insect Sci., 2025, 1–12, DOI:10.1111/1744-7917.70065.
- S. S. Taylor, E. Radzio-Andzelm and T. Hunter, FASEB J., 1995, 9, 1255–1266 CrossRef CAS PubMed.
- V. S. Tagliabracci, S. E. Wiley, X. Guo, L. N. Kinch, E. Durrant, J. Wen, J. Xiao, J. Cui, K. B. Nguyen, J. L. Engel, J. J. Coon, N. Grishin, L. A. Pinna, D. J. Pagliarini and J. E. Dixon, Cell, 2015, 161, 1619–1632 CrossRef CAS PubMed.
- M. He, X. Zhou and X. Wang, Signal Transduct. Target. Ther., 2024, 9, 194 CrossRef PubMed.
- K. Vasanthavada, X. Hu, T. Tuton-Blasingame, Y. Hsia, S. Sampath, R. Pacheco, J. Freeark, A. M. Falick, S. Tang, J. Fong, K. Kohler, C. La Mattina-Hawkins and C. Vierra, J. Biol. Chem., 2012, 287, 35986–35999 CrossRef CAS PubMed.
- H. M. Kagan and W. Li, J. Cell. Biochem., 2003, 88, 660–672 CrossRef CAS PubMed.
- W. Li, K. Nellaiappan, T. Strassmaier, L. Graham, K. M. Thomas and H. M. Kagan, Proc. Natl. Acad. Sci. U. S. A., 1997, 94, 12817–12822 CrossRef CAS PubMed.
- L. I. Smith-Mungo and H. M. Kagan, Matrix Biol., 1998, 16, 387–398 CrossRef CAS PubMed.
- A. Sethi, R. J. Wordinger and A. F. Clark, Exp. Eye Res., 2012, 104, 97–98 CrossRef CAS PubMed.
- I. K. Amma, R. S. J. Ingrole, G. K. Venkatesa Prabhu, R. Dominquez, D. Kong, S. C. H. Mangalara, G. B. McKenna and H. S. Gill, Biomacromolecules, 2025, 26, 1580–1594 CrossRef PubMed.
- C. Giulivi and K. J. A. Davies, Methods in Enzymology, Academic Press, 1994, vol. 233, pp. 363–371 Search PubMed.
- S. O. Andersen, Insect Biochem. Mol. Biol., 2010, 40, 166–178 CrossRef CAS PubMed.
- R. J. Suderman, N. T. Dittmer, M. R. Kanost and K. J. Kramer, Insect Biochem. Mol. Biol., 2006, 36, 353–365 CrossRef CAS PubMed.
- C. M. Elvin, A. G. Carr, M. G. Huson, J. M. Maxwell, R. D. Pearson, T. Vuocolo, N. E. Liyou, D. C. Wong, D. J. Merritt and N. E. Dixon, Nature, 2005, 437, 999–1002 CrossRef CAS PubMed.
- D. R. Sell and V. M. Monnier, Connect. Tissue Res., 1989, 19, 77–92 CrossRef CAS PubMed.
- Y. Ying and H. Li, Methods, 2022, 200, 42–57 CrossRef CAS PubMed.
- J. Q. Bettinger, M. Simon, A. Korotkov, K. A. Welle, J. R. Hryhorenko, A. Seluanov, V. Gorbunova and S. Ghaemmaghami, J. Proteome Res., 2022, 21, 1495–1509 CrossRef CAS PubMed.
- S. Chaturvedi, S. Bawake and N. Sharma, Rapid Commun. Mass Spectrom., 2024, 38, e9713 CrossRef CAS PubMed.
- J. Rosenberg, V. Sundaresh, J. Gililland, A. Osman, W. Woolley and C. Acevedo, JOM, 2024, 76, 5684–5691 CrossRef CAS.
- C. A. Michal, A. H. Simmons, B. G. Chew, D. B. Zax and L. W. Jelinski, Biophys. J., 1996, 70, 489–493 CrossRef CAS PubMed.
- A. D. Malay, H. C. Craig, J. Chen, N. A. Oktaviani and K. Numata, Biomacromolecules, 2022, 23, 1827–1840 CrossRef CAS PubMed.
- V. Sahni, T. A. Blackledge and A. Dhinojwala, Sci. Rep., 2011, 1, 41 CrossRef PubMed.
- M. Ramezaniaghdam, N. D. Nahdi and R. Reski, Front. Bioeng. Biotechnol., 2022, 10, 2022 Search PubMed.
- J. R. Santos-Pinto, H. A. Arcuri, G. Lubec and M. S. Palma, Biochim. Biophys. Acta, 2016, 1864, 1444–1454 CrossRef PubMed.
- N. A. Oktaviani, A. D. Malay, A. Matsugami, F. Hayashi and K. Numata, Biomacromolecules, 2023, 24, 1604–1616 CrossRef CAS PubMed.
- H. C. Craig, S. J. Blamires, M. A. Sani, M. M. Kasumovic, A. Rawal and J. M. Hook, Chem. Commun., 2019, 55, 4687–4690 RSC.
- J. R. A. Dos Santos-Pinto, H. A. Arcuri, F. G. Esteves, M. S. Palma and G. Lubec, Sci. Rep., 2018, 8, 14674 CrossRef PubMed.
- C. Y. Hayashi and R. V. Lewis, J. Mol. Biol., 1998, 275, 773–784 CrossRef CAS PubMed.
- N. Becker, E. Oroudjev, S. Mutz, J. P. Cleveland, P. K. Hansma, C. Y. Hayashi, D. E. Makarov and H. G. Hansma, Nat. Mater., 2003, 2, 278–283 CrossRef CAS PubMed.
- F. Vollrath and D. P. Knight, Nature, 2001, 410, 541–548 CrossRef CAS PubMed.
- M. Heim, D. Keerl and T. Scheibel, Angew. Chem., Int. Ed., 2009, 48, 3584–3596 CrossRef CAS PubMed.
- A. Rising and J. Johansson, Nat. Chem. Biol., 2015, 11, 309–315 CrossRef CAS PubMed.
- J. R. dos Santos-Pinto, H. A. Arcuri, H. Priewalder, H. C. Salles, M. S. Palma and G. Lubec, J. Proteome Res., 2015, 14, 3859–3870 CrossRef CAS PubMed.
- O. Choresh, B. Bayarmagnai and R. V. Lewis, Biomacromolecules, 2009, 10, 2852–2856 CrossRef CAS PubMed.
- E. Tillinghast, M. Townley, T. Wight, G. Uhlenbruck and E. Janssen, MRS Proc., 1992, 292, 9–23 CrossRef.
- M. A. Collin, T. H. Clarke, 3rd, N. A. Ayoub and C. Y. Hayashi, Sci. Rep., 2016, 6, 21589 CrossRef CAS PubMed.
- G. Amarpuri, V. Chaurasia, D. Jain, T. A. Blackledge and A. Dhinojwala, Sci. Rep., 2015, 5, 9030 CrossRef CAS PubMed.
- G. Amarpuri, C. Zhang, C. Diaz, B. D. Opell, T. A. Blackledge and A. Dhinojwala, ACS Nano, 2015, 9, 11472–11478 CrossRef CAS PubMed.
- B. D. Opell, S. E. Karinshak and M. A. Sigler, J. Exp. Biol., 2011, 214, 2988–2993 CrossRef PubMed.
- V. Sahni, T. A. Blackledge and A. Dhinojwala, Nat. Commun., 2010, 1, 19 CrossRef PubMed.
- N. A. Ayoub, K. Friend, T. Clarke, R. Baker, S. M. Correa-Garhwal, A. Crean, E. Dendev, D. Foster, L. Hoff, S. D. Kelly, W. Patterson, C. Y. Hayashi and B. D. Opell, Integr. Comp. Biol., 2021, 61, 1459–1480 CAS.
- V. Sahni, T. Miyoshi, K. Chen, D. Jain, S. J. Blamires, T. A. Blackledge and A. Dhinojwala, Biomacromolecules, 2014, 15, 1225–1232 CrossRef CAS PubMed.
- S. Singla, G. Amarpuri, N. Dhopatkar, T. A. Blackledge and A. Dhinojwala, Nat. Commun., 2018, 9, 1890 CrossRef PubMed.
- C. Heinritz, X. J. Ng and T. Scheibel, Adv. Funct. Mater., 2024, 34, 2303609 CrossRef CAS.
- C. Y. Hayashi, T. A. Blackledge and R. V. Lewis, Mol. Biol. Evol., 2004, 21, 1950–1959 CrossRef CAS PubMed.
- M. Tian and R. V. Lewis, Biochemistry, 2005, 44, 8006–8012 CrossRef CAS PubMed.
- J. O. Wolff, I. Grawe, M. Wirth, A. Karstedt and S. N. Gorb, Soft Matter, 2015, 11, 2394–2403 RSC.
- J. E. Garb and C. Y. Hayashi, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 11379–11384 CrossRef CAS PubMed.
- W. Huang, Y. Zhang, Y. Chen, Y. Wang, W. Yuan, N. Zhang, T. J. Lam, Z. Gong, D. Yang and Z. Lin, Sci. Rep., 2017, 7, 13354 CrossRef PubMed.
- C. Solazzo, J. M. Dyer, S. Deb-Choudhury, S. Clerens and P. Wyeth, Photochem. Photobiol., 2012, 88, 1217–1226 CrossRef CAS PubMed.
- B. Tulachan, S. Srivastava, T. S. Kusurkar, N. K. Sethy, K. Bhargava, S. K. Singh, D. Philip, A. Bajpai and M. Das, Sci. Rep., 2016, 6, 21915 CrossRef CAS PubMed.
- K. Vasanthavada, X. Hu, A. M. Falick, C. La Mattina, A. M. Moore, P. R. Jones, R. Yee, R. Reza, T. Tuton and C. Vierra, J. Biol. Chem., 2007, 282, 35088–35097 CrossRef CAS PubMed.
- K. Chalek, D. Onofrei, J. E. Aldana, H. R. Johnson, D. Stengel, A. T. Chau, A. R. Domingo, E. Alcala, E. Tsui, F. Botello, A. Soni, B. Addison and G. P. Holland, Adv. Funct. Mater., 2024, 35, 2409321 CrossRef.
- R. Wen, K. Wang and Q. Meng, Acta Biomater., 2020, 115, 210–219 CrossRef CAS PubMed.
- C. Z. Zhou, F. Confalonieri, N. Medina, Y. Zivanovic, C. Esnault, T. Yang, M. Jacquet, J. Janin, M. Duguet, R. Perasso and Z. G. Li, Nucleic Acids Res., 2000, 28, 2413–2419 CrossRef CAS PubMed.
- S. Inoue, K. Tanaka, F. Arisaka, S. Kimura, K. Ohtomo and S. Mizuno, J. Biol. Chem., 2000, 275, 40517–40528 CrossRef CAS PubMed.
- C.-Z. Zhou, F. Confalonieri, M. Jacquet, R. Perasso, Z.-G. Li and J. Janin, Proteins: Struct., Funct., Bioinf., 2001, 44, 119–122 CrossRef CAS PubMed.
- J. Song, J. Che, Z. You, L. Ye, J. Li, Y. Zhang, Q. Qian and B. Zhong, J. Proteomics, 2016, 148, 194–201 CrossRef CAS PubMed.
- J. Song, J. Che, Z. You, L. Ye, J. Li, Y. Zhang, Q. Qian and B. Zhong, J. Proteomics, 2016, 148, 194–201 CrossRef CAS PubMed.
- N. Johari, L. Moroni and A. Samadikuchaksaraei, Eur. Polym. J., 2020, 134, 109842 CrossRef CAS.
- P. Zhang, K. Yamamoto, Y. Aso, Y. Banno, D. Sakano, Y. Wang and H. Fujii, Biosci., Biotechnol., Biochem., 2005, 69, 2086–2093 CrossRef CAS PubMed.
- R. I. Kunz, R. M. Brancalhão, L. F. Ribeiro and M. R. Natali, BioMed Res. Int., 2016, 2016, 8175701 Search PubMed.
- J. Heckenhauer, R. J. Stewart, B. Ríos-Touma, A. Powell, T. Dorji, P. B. Frandsen and S. U. Pauls, iScience, 2023, 26, 107253 CrossRef CAS PubMed.
- R. J. Stewart and C. S. Wang, Biomacromolecules, 2010, 11, 969–974 CrossRef CAS PubMed.
- J. B. Addison, N. N. Ashton, W. S. Weber, R. J. Stewart, G. P. Holland and J. L. Yarger, Biomacromolecules, 2013, 14, 1140–1148 CrossRef CAS PubMed.
- J. B. Addison, W. S. Weber, Q. Mou, N. N. Ashton, R. J. Stewart, G. P. Holland and J. L. Yarger, Biomacromolecules, 2014, 15, 1269–1275 CrossRef CAS PubMed.
- N. N. Ashton, D. R. Roe, R. B. Weiss, T. E. Cheatham, III and R. J. Stewart, Biomacromolecules, 2013, 14, 3668–3681 CrossRef CAS PubMed.
- C. S. Wang and R. J. Stewart, J. Exp. Biol., 2012, 215, 351–361 CrossRef PubMed.
- N. N. Ashton, D. R. Roe, R. B. Weiss, T. E. Cheatham, 3rd and R. J. Stewart, Biomacromolecules, 2013, 14, 3668–3681 CrossRef CAS PubMed.
- N. N. Ashton and R. J. Stewart, Soft Matter, 2015, 11, 1667–1676 RSC.
- H.-J. Kim, Y. Sun and M.-J. Moon, Appl. Microsc., 2020, 50, 16 CrossRef PubMed.
- C. S. Wang, N. N. Ashton, R. B. Weiss and R. J. Stewart, Insect Biochem. Mol. Biol., 2014, 54, 69–79 CrossRef CAS PubMed.
- J. B. Addison, N. N. Ashton, W. S. Weber, R. J. Stewart, G. P. Holland and J. L. Yarger, Biomacromolecules, 2013, 14, 1140–1148 CrossRef CAS PubMed.
- R. J. Stewart, Appl. Microbiol. Biotechnol., 2011, 89, 27–33 CrossRef CAS PubMed.
- A. Kirillova, C. Kelly, N. von Windheim and K. Gall, Adv. Healthcare Mater., 2018, 7, e1800467 CrossRef.
- S. Standring, J. Heckenhauer, R. J. Stewart and P. B. Frandsen, Trends Genet., 2025, 41, 537–546 CrossRef CAS.
- T. D. Sutherland, J. H. Young, S. Weisman, C. Y. Hayashi and D. J. Merritt, Annu. Rev. Entomol., 2010, 55, 171–188 CrossRef CAS PubMed.
- N. Kono, H. Nakamura, R. Ohtoshi, M. Tomita, K. Numata and K. Arakawa, Commun. Biol., 2019, 2, 148 CrossRef PubMed.
- J. J. Halvorson, J. M. Gonzalez and A. E. Hagerman, Soil Biol. Biochem., 2011, 43, 1139–1147 CrossRef CAS.
- O. Wasserman, J. J. Morley, M.-K. F. Williams, B. E. Bell, D. L. Cox-Foster and J. A. Jones, PLoS One, 2025, 20, e0318918 CrossRef CAS PubMed.
- S. Tragust, L. V. Ugelvig, M. Chapuisat, J. Heinze and S. Cremer, BMC Evol. Biol., 2013, 13, 225 CrossRef PubMed.
- P. M. Campbell, H. E. Trueman, Q. Zhang, K. Kojima, T. Kameda and T. D. Sutherland, Insect Biochem. Mol. Biol., 2014, 48, 40–50 CrossRef CAS PubMed.
- T. D. Sutherland, Y. Y. Peng, H. E. Trueman, S. Weisman, S. Okada, A. A. Walker, A. Sriskantha, J. F. White, M. G. Huson, J. A. Werkmeister, V. Glattauer, V. Stoichevska, S. T. Mudie, V. S. Haritos and J. A. M. Ramshaw, Sci. Rep., 2013, 3, 2864 CrossRef PubMed.
- D. L. J. Quicke and M. R. Shaw, Biol. J. Linnean Soc., 2004, 81, 161–170 CrossRef.
- M. Terajima, Y. Taga, W. A. Cabral, Y. Liu, M. Nagasawa, N. Sumida, Y. Kayashima, P. Chandrasekaran, L. Han, N. Maeda, I. Perdivara, S. Hattori, J. C. Marini and M. Yamauchi, PLoS Genet., 2019, 15, e1008196 CrossRef CAS PubMed.
- D. M. Pinkas, S. Ding, R. T. Raines and A. E. Barron, ACS Chem. Biol., 2011, 6, 320–324 CrossRef CAS PubMed.
- C. H. Kim, J. Y. Axup and P. G. Schultz, Curr. Opin. Chem. Biol., 2013, 17, 412–419 CrossRef CAS PubMed.
- J. W. Chin, T. A. Cropp, J. C. Anderson, M. Mukherji, Z. Zhang and P. G. Schultz, Science, 2003, 301, 964–967 CrossRef CAS PubMed.
- Y. S. Jeong, B. Yang, B. Yang, M. Shin, J. Seong, H. J. Cha and I. Kwon, Biotechnol. Bioeng., 2020, 117, 1961–1969 CrossRef CAS PubMed.
- J. Choi, S. Lee, Y. Lee and D. S. Hwang, Nat. Commun., 2025, 16, 233 CrossRef PubMed.
- A. Deiters, T. A. Cropp, D. Summerer, M. Mukherji and P. G. Schultz, Bioorg. Med. Chem. Lett., 2004, 14, 5743–5745 CrossRef CAS PubMed.
- S. A. Costa, J. R. Simon, M. Amiram, L. Tang, S. Zauscher, E. M. Brustad, F. J. Isaacs and A. Chilkoti, Adv. Mater., 2018, 30, 1704878 CrossRef PubMed.
- H. Teramoto and K. Kojima, Biomacromolecules, 2014, 15, 2682–2690 CrossRef CAS PubMed.
- N. B. Möttönen, R. Fan, S. Aspholm-Tsironi, S. Keskitalo, A. Tuhkala, M. Varjosalo and A. S. Aranko, Biomacromolecules, 2025, 26(12), 8594–8605 CrossRef PubMed.
- J. Dadová, S. R. Galan and B. G. Davis, Curr. Opin. Chem. Biol., 2018, 46, 71–81 CrossRef PubMed.
- M. G. Casteleijn, U. Abendroth, A. Zemella, R. Walter, R. Rashmi, R. Haag and S. Kubick, Chem. Rev., 2025, 125, 1303–1331 CrossRef CAS PubMed.
- W. Gao, E. Cho, Y. Liu and Y. Lu, Front. Pharmacol., 2019, 10, 611 CrossRef CAS PubMed.
- D. A. Wong, Z. M. Shaver, M. D. Cabezas, M. Daniel-Ivad, K. F. Warfel, D. V. Prasanna, S. E. Sobol, R. Fernandez, F. Tobias, S. K. Filip, S. W. Hulbert, P. Faull, R. Nicol, M. P. DeLisa, E. P. Balskus, A. S. Karim and M. C. Jewett, Nat. Commun., 2025, 16, 7215 CrossRef CAS PubMed.
- Z. Sun and J. L. Brodsky, J. Cell Biol., 2019, 218, 3171–3187 CrossRef CAS PubMed.
- T. Murase, T. Tsuji and Y. Kajihara, Carbohydr. Res., 2009, 344, 762–770 CrossRef CAS PubMed.
- T. Murase and Y. Kajihara, Carbohydr. Res., 2010, 345, 1324–1330 CrossRef CAS PubMed.
- K. Domon, M. Puripat, K. Fujiyoshi, M. Hatanaka, S. A. Kawashima, K. Yamatsugu and M. Kanai, ACS Cent. Sci., 2020, 6, 283–292 Search PubMed.
- B. D. A. Shennan, T. Fukuta, M. Yamane, T. Koyama, H. Mitsunuma and M. Kanai, J. Am. Chem. Soc., 2025, 147, 6349–6354 CrossRef CAS PubMed.
- C. A. Werlang, J. K. Sahoo, G. Cárcarmo-Oyarce, C. Stevens, D. Uzun, R. Putnik, O. Hasturk, J. Choi, D. L. Kaplan and K. Ribbeck, J. Am. Chem. Soc., 2024, 146, 34661–34668 CrossRef CAS PubMed.
- F. Yu, G. C. Teo, A. T. Kong, S. E. Haynes, D. M. Avtonomov, D. J. Geiszler and A. I. Nesvizhskii, Nat. Commun., 2020, 11, 4065 CrossRef CAS PubMed.
- T. Pradita, Y. J. Chen, T. H. Su, K. H. Chang, P. J. Chen and Y. J. Chen, J. Proteome Res., 2024, 23, 3571–3584 CrossRef CAS PubMed.
- D. B. Bekker-Jensen, O. M. Bernhardt, A. Hogrebe, A. Martinez-Val, L. Verbeke, T. Gandhi, C. D. Kelstrup, L. Reiter and J. V. Olsen, Nat. Commun., 2020, 11, 787 CrossRef CAS PubMed.
- D. Chang and J. Zaia, Mass Spectrom. Rev., 2022, 41, 922–937 CrossRef CAS PubMed.
- Y. Yang, X. Liu, C. Shen, Y. Lin, P. Yang and L. Qiao, Nat. Commun., 2020, 11, 146 CrossRef CAS PubMed.
- A. Leitner, M. Faini, F. Stengel and R. Aebersold, Trends Biochem. Sci., 2016, 41, 20–32 CrossRef CAS PubMed.
- T. K. Bartolec, X. Vázquez-Campos, A. Norman, C. Luong, M. Johnson, R. J. Payne, M. R. Wilkins, J. P. Mackay and J. K. K. Low, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2219418120 CrossRef CAS PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. A. A. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A. W. Senior, K. Kavukcuoglu, P. Kohli and D. Hassabis, Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- C. R. Chung, Y. Tang, Y. P. Chiu, S. Li, W. K. Hsieh, L. Yao, Y. C. Chiang, Y. Pang, G. T. Chen, K. C. Chou, Y. S. Paik, P. L. Tran, C. P. Lin, Y. M. Kao, Y. J. Chen, W. C. Chang, J. B. Hsu, J. T. Horng and T. Y. Lee, Nucleic Acids Res., 2025, 53, D377–D386 CrossRef CAS PubMed.
- I. Bludau, S. Willems, W.-F. Zeng, M. T. Strauss, F. M. Hansen, M. C. Tanzer, O. Karayel, B. A. Schulman and M. Mann, PLoS Biol., 2022, 20, e3001636 CrossRef CAS PubMed.
- F. Z. Peng, C. Wang, T. Chen, B. Schussheim, S. Vincoff and P. Chatterjee, Nat. Methods, 2025, 22, 945–949 CrossRef CAS PubMed.
- P. Shrestha, J. Kandel, H. Tayara and K. T. Chong, Nat. Commun., 2024, 15, 6699 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |
