Open Access Article
Open Access ArticleDevelopment of PROTACs for targeted degradation of oncogenic TRK fusions
Saurav
Kumar†
 a,
Jiewei
Jiang†
a,
Jiewei
Jiang†
 b,
Mia S.
Donald-Paladino
b,
Mia S.
Donald-Paladino
 a,
Joy
Chen
a,
Andrea
Gutierrez
c,
Alexander J.
Federation
c,
Frank
Szulzewsky
de,
Eric C.
Holland
a,
Fleur M.
Ferguson
*bf and
Behnam
Nabet
a,
Joy
Chen
a,
Andrea
Gutierrez
c,
Alexander J.
Federation
c,
Frank
Szulzewsky
de,
Eric C.
Holland
a,
Fleur M.
Ferguson
*bf and
Behnam
Nabet
 *ag
*ag
aHuman Biology Division, Fred Hutchinson Cancer Center, Seattle, WA 98109, USA. E-mail: bnabet@fredhutch.org
bDepartment of Chemistry and Biochemistry, University of California San Diego, La Jolla, CA 92093, USA. E-mail: fmferguson@ucsd.edu
cTalus Bioscience, Inc, Seattle, WA 98122, USA
dHuntsman Cancer Institute, University of Utah, Salt Lake City, UT 84112, USA
eDepartment of Neurosurgery, Clinical Neurosciences Center, University of Utah, Salt Lake City, UT 84112, USA
fSkaggs School of Pharmacy and Pharmaceutical Sciences, University of California San Diego, La Jolla, CA 92093, USA
gDepartment of Pharmacology, University of Washington, Seattle, WA 98195, USA
First published on 18th March 2026
Abstract
Chromosomal translocations leading to the fusion of tropomyosin receptor kinases (TRKs) with diverse partner proteins have been identified as oncogenic drivers in many adult and pediatric cancers. While first-generation TRK kinase inhibitors, such as entrectinib and larotrectinib, have shown positive responses in TRK fusion-positive cancers, resistance mutations against these inhibitors in the kinase domain limit their efficacy. Second-generation inhibitors are in clinical evaluation, highlighting a need for novel therapeutic modalities to achieve durable suppression of the oncogenic activity of TRK fusions. Here, we developed heterobifunctional small molecule degraders (PROTACs) to achieve targeted degradation of TRK fusions. By conjugating entrectinib to thalidomide, we identified JWJ-01-378 as a potent and selective cereblon (CRBN)-recruiting degrader of the TPM3-TRKA fusion. JWJ-01-378 induced TPM3-TRKA degradation through the ubiquitin-proteasome system and proteomics analysis confirmed the acute selectivity of JWJ-01-378 for achieving TPM3-TRKA degradation with minimal off-target effects. Importantly, JWJ-01-378 did not degrade CRBN neosubstrates that are targeted by existing TRK PROTACs including CG-428. While JWJ-01-378 was also able to degrade wild-type TRK, it was unable to degrade TRK inhibitor resistant mutants and ALK fusions. TPM3-TRKA degradation by JWJ-01-378 suppressed downstream signaling and reduced cancer cell viability, with improved responses compared to a heterobifunctional control compound that cannot degrade TPM3-TRKA. Together, our study expands the toolbox of selective compounds for evaluating targeted degradation of TRK fusions in diseases including cancer.
Introduction
The neurotrophic receptor tyrosine kinase (NTRK) gene family, which is comprised of NTRK1, NTRK2, and NTRK3, encodes the tropomyosin receptor kinase proteins TRKA, TRKB, and TRKC, respectively.1 During normal development, these receptor tyrosine kinases are predominantly expressed in neuronal tissues and activated in response to ligands including NGF, BDNF, NT-3 and NT-4. Ligand binding to their extracellular domains induces receptor dimerization, autophosphorylation of the tyrosine kinase domain, and activation of downstream signaling including MAPK, PI3K-AKT, mTOR, and PLCγ pathways that are essential for neuronal development, survival, and differentiation.1,2 Alterations in NTRK genes, which predominately occur through chromosomal translocations and less commonly include point mutations and deletions, have been identified as driver events in several cancers.3–6NTRK fusions are recurrent events (>80%) in rare cancers, such as secretory breast cancer and infantile fibrosarcoma, while less frequent (<25%) in pediatric and adult cancers, such as high-grade glioma, melanoma, colorectal cancer, and lung cancer.6In vivo expression of TRK fusions in mice is sufficient to induce tumor formation, further highlighting their role as strong oncogenic drivers.7,8Most commonly, these chromosomal translocations involve fusion of the 3′ region of an NTRK gene, which encodes the functional tyrosine kinase domain, to the 5′ region of another gene partner.9,10 Many of these 5′ fusion partners contribute oligomerization domains that induce ligand-independent receptor dimerization, leading to constitutive activation of the TRK fusions.6,10 TRK fusions then coordinate aberrant downstream signaling and promote uncontrolled cell proliferation and survival, which are hallmark features of cancer.11 As such, targeting TRK fusions has emerged as a tractable drug discovery approach, and several TRK inhibitors have advanced into clinical trials.
TRK-targeting compounds include broad-spectrum tyrosine kinase inhibitors such as entrectinib, lestaurtinib, and crizotinib, which demonstrate activity against multiple kinases including TRK, ALK, and ROS1.12–15 Inhibitors with improved specificity towards TRK proteins have also been developed such as larotrectinib and BPI-28592.16,17 Notably, entrectinib and larotrectinib received FDA approval in 2018 and 2019, respectively, for the treatment of cancers harboring TRK mutations and TRK fusions.18,19 Both entrectinib and larotrectinib have shown high overall response rates in TRK fusion-positive cancers,20–22 but the emergence of drug resistance to these inhibitors poses significant clinical challenges.6,23,24 Common resistance mechanisms involve on-target mutations in the TRK kinase domain, including the gatekeeper residue, solvent front and activation loop. Second-generation TRK inhibitors designed to overcome these resistance mutations including selitrectinib, repotrectinib, and ONO-5390556 are currently under clinical evaluation.25–28 However, on-target mutations including secondary mutations in TRK confer resistance to these inhibitors.29 Therefore, continued advancement of novel therapeutic approaches is necessary to target TRK fusions in cancer.
Rather than inhibiting protein activity, small molecule degraders such as proteolysis-targeting chimeras (PROTACs) are a powerful chemical strategy to target proteins for degradation through the ubiquitin-proteasome system.30,31 PROTACs are heterobifunctional molecules that induce proximity between a target protein and an E3 ubiquitin ligase, leading to ubiquitination and subsequent proteasomal degradation. PROTACs offer several advantages compared to small molecule inhibitors, including their catalytic nature and ability to achieve rapid and sustained elimination of all protein functions including non-enzymatic roles.30 We32,33 and others34,35 have developed PROTACs targeting kinases, demonstrating their improved activity in dampening aberrant downstream signaling and biological responses compared to kinase inhibitors. Given the oncogenic role of TRK fusions in many pediatric and adult cancers and the challenges associated with resistance mutations that limit the effectiveness of TRK inhibitors, we aimed to develop PROTACs to target TRK fusions for degradation.
To study TRK fusion protein degradation, we focused on the TPM3-TRKA fusion and synthesized PROTACs by conjugating entrectinib to thalidomide to recruit the E3 ubiquitin ligase cereblon (CRBN). We identified several compounds as effective degraders, with JWJ-01-378 nominated as the most potent degrader of the TPM3-TRKA fusion. JWJ-01-378 achieved rapid and durable degradation of TPM3-TRKA, comparable to a previously described TRK PROTAC (Cullgen's CG-428).36 JWJ-01-378 required the ubiquitin-proteasome system for achieving degradation and reduced viability of colorectal cancer cells dependent on TPM3-TRKA, with enhanced effects on downstream signaling and cell viability compared to a heterobifunctional control compound that cannot degrade TRK fusions. Proteomics analyses at an acute time-point confirmed TPM3-TRKA degradation with minimal off-target and secondary responses. Crucially, we observed that JWJ-01-378 displayed an improved selectivity profile over CG-428 and did not degrade CRBN neosubstrates including IKZF1, IKZF3, and GSPT1 that were targeted by CG-428. In addition, we observed that JWJ-01-378 degraded wild-type (WT) TRK and was unable to degrade TRK inhibitor resistant mutants or ALK fusions. Together, we developed an expanded toolbox of selective compounds for the interrogation of TRK fusion proteins in cancer.
Results and discussion
Synthesis of entrectinib-derived PROTACs for targeted degradation of TRK fusions
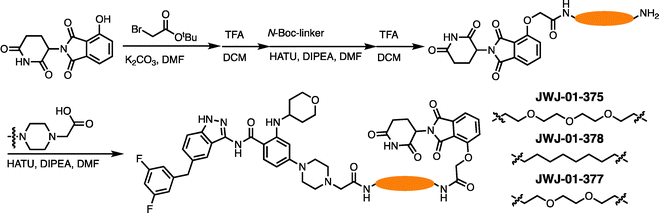
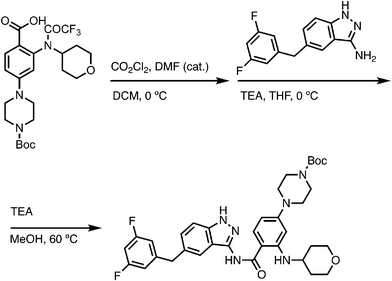
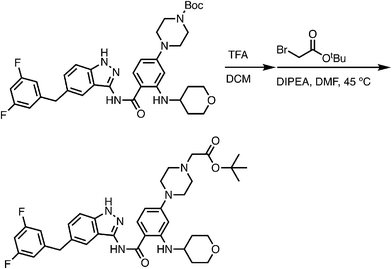
In this study, we aimed to develop TRK PROTACs to evaluate targeted degradation of TRK fusions. As the TRK-binding moiety, we selected entrectinib, which is a potent FDA-approved pan-TRK inhibitor for treating TRK-dependent tumors.37 In addition, entrectinib's synthetic simplicity, commercial availability of an intermediate, and lack of requirement for chiral separation, made it an attractive starting point for PROTAC development. Entrectinib binds to the active conformation of TRK by occupying ATP-binding sites in the kinase domain to prevent phosphorylation of its downstream substrates. To functionalize entrectinib and develop CRBN-recruiting PROTACs, we evaluated the reported co-crystal structure in complex with the TRKA kinase domain (PDB ID: 5KVT) and decided to use the solvent exposed N-methyl piperazine as a convenient anchor point (Fig. 1A and B). Importantly, we also aimed to recruit CRBN using a thalidomide-derivative linked via an ether bond at the 4-position to avoid off-target degradation of CRBN neosubstrates.38As depicted in Scheme 1, the key carboxylate intermediate was generated within ten steps. Briefly, methyl 2-amino-4-bromobenzoate underwent a reductive amination to introduce the tetrahydropyran ring. A Buchwald-Hartwig coupling installed 1-Boc protected piperazine at the para position of the aryl ring. After the hydrolysis, the secondary aniline was protected by trifluoroacetic anhydride, followed by the oxalyl chloride-mediated amidation with the commercially available indazole moiety. The trifluoroacetic and Boc protecting groups were removed by TEA and TFA, respectively. The displacement between the piperazine and tert-butyl bromoacetate and sequential TFA treatment yielded the target carboxylate.
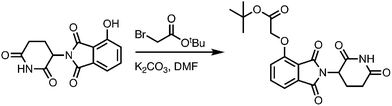
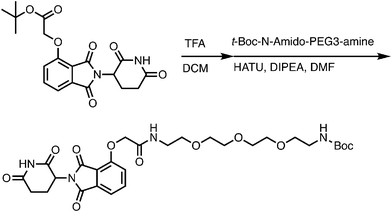
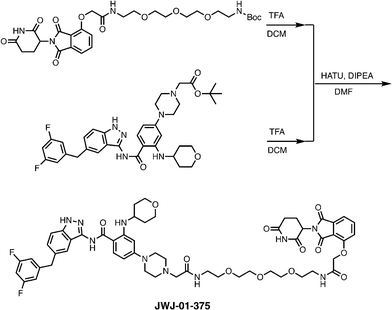
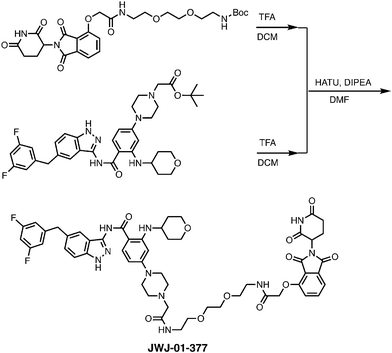
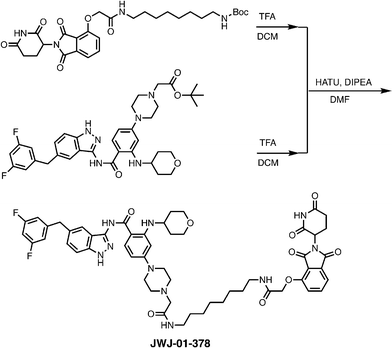
Meanwhile as shown in Scheme 2, 4-hydroxy thalidomide was treated with tert-butyl bromoacetate at the presence of potassium carbonate to selectively alkylate at phenolic site. After the TFA treatment, the carboxylate was coupled with various N-Boc protected linkers. Another Boc deprotection yielded the terminal amine to be coupled with the acid-functionalized entrectinib and give the desired PROTAC molecules. Together, we synthesized several entrectinib-derived PROTACs for evaluation (Fig. 1C and Fig. S1–S3).
Entrectinib-derived PROTACs induce rapid and potent degradation of TRK fusions
To evaluate the activity of our entrectinib-derived PROTACs, we utilized KM12 cells, a colorectal cancer cell line that harbors an endogenous TPM3-TRKA fusion (Fig. S5A). This fusion arises from an intrachromosomal rearrangement at 1q22-23 between the N-terminal domain of TPM3 (codons 1-221) and the C-terminal portion of TRKA (codons 419-end) that encodes the transmembrane and intracellular tyrosine kinase domain.39 Prior work has shown that the viability of KM12 cells is substantially reduced upon knockdown of TPM3-TRKA,40 making these cells a powerful model to study the consequences of TPM3-TRKA disruption. This is further supported by the Cancer Dependency Map (DepMap)41 where TPM3 scores as a genetic dependency (Fig. S5B). While NTRK1 does not score as a significant genetic dependency (Fig. S5C), this may be due to the sgRNA location relative to the fusion breakpoint, which may fail to capture a true dependency on a partner gene.42 Notably, KM12 cells do not express ALK or ROS1, the most potent off-targets of entrectinib12 (Fig. S5D and E).To characterize our PROTACs, we evaluated TPM3-TRKA degradation with extensive dose-responses and time-course assessments, performed on-mechanism and selectivity studies to characterize lead compounds, and determined the phenotypic consequences of TRK fusion degradation. We also compared our entrectinib-derived PROTACs to Cullgen's TRK-targeting PROTAC (CG-428),36 which differs in its TRK-targeting inhibitor (GNF-8625) and uses a 4-anilino attachment chemistry, mimicking pomalidomide for the CRBN-recruiting ligand (Fig. 1D). First, treatment of KM12 cells with JWJ-01-375 (3-PEG linker), JWJ-01-377 (2-PEG linker), and JWJ-01-378 (8-carbon linker) led to rapid and pronounced degradation of TPM3-TRKA and suppression of downstream phosphorylated ERK (pERK) levels, a pathway that is aberrantly activated by TRK fusions43 (Fig. 2A and B). Notably, of the PROTACs tested, JWJ-01-378 achieved the most potent degradation at doses as low as 10 nM and was prioritized for further evaluation. As expected, CG-428 and entrectinib decreased pERK levels, and degradation of TPM3-TRKA was observed upon treatment with CG-428 (Fig. 2A and B).
Given the similar degradation profiles between JWJ-01-378 and CG-428, we examined the binding modes of these PROTACs in more detail. Evaluating the co-crystal structure of entrectinib and GNF-8625, we observed a similar binding mode with the TRKA kinase domain (PDB ID: 5KVT) (Fig. S6A). We also sought to investigate the binding mode upon formation of a ternary complex using molecular docking. We started from the reported co-crystal structures of CRBN (PDB ID: 4CL1) and TRKA (PDB ID: 5KVT 4YNE). By using the three-body-search method from MOE,44 we were able to predict the binding modes of both JWJ-01-378 and CG-428, which form a stable ternary complex between CRBN and TRKA through a similar protein–protein interface (Fig. S6B). One noticeable difference from the superimposition is that the exit vector on the thalidomide moiety was flipped, which could be due to the different exit vector angles on the TRK binders. When overlaying two predicted complexes based on the TRKA protein, we noticed that the CRBN-recruiting ligand in JWJ-01-378 had minor shifts, compared to that with CG-428. The observed thalidomide orientation thus likely retains the desired protein–protein interactions between CRBN and TRKA.
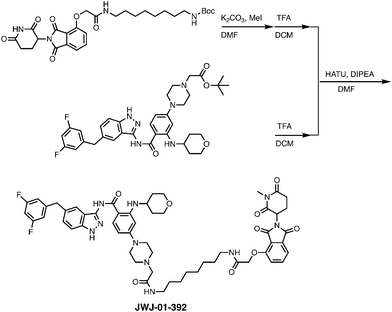
Next, to understand and evaluate JWJ-01-378 activity and biological effects, we synthesized a heterobifunctional control compound (JWJ-01-392) consisting of an N-methylated glutarimide analog that disables CRBN binding45 (Fig. 1C and Fig. S4). JWJ-01-392 failed to induce TPM3-TRKA degradation and did not suppress pERK to the levels of JWJ-01-378 (Fig. 2C and D). To study the kinetics of TPM3-TRKA degradation and impacts on downstream pERK signaling, we performed time-course experiments. JWJ-01-378 treatment diminished downstream TPM3-TRKA protein levels and abolished pERK signaling, with substantially improved suppression of pERK compared to JWJ-01-392 (Fig. 2E). Entrectinib treatment also diminished signaling similarly to JWJ-01-378, albeit with faster kinetics, which may be due to improved cell permeability and/or consequent enhanced off-target inhibitory effects from additional known kinases targeted by entrectinib.13,46–48 Our time-course analysis revealed partial recovery of pERK levels by 24 hours of treatment with the TRK PROTACs or inhibitors. To understand the responses beyond 24 hours, we evaluated the TPM3-TRKA and pERK levels over the course of five days, following a single treatment. JWJ-01-378 and CG-428 displayed similar partial recovery kinetics of TPM3-TRKA levels, which was delayed compared to pERK. Interestingly, pERK recovered to baseline levels after five days (Fig. 2F). The pERK reactivation may be due to the recovery of TPM3-TRKA levels, the emergence of compensatory signaling, or cellular rewiring to reactivate the pathway.
Following confirmation of the disruption of downstream signaling, we next evaluated the mechanism-of-action of JWJ-01-378. Pretreatment with proteasomal inhibitor carfilzomib or the NEDD8 inhibitor MLN4924 successfully rescued TPM3-TRKA degradation upon JWJ-01-378 treatment, confirming its dependence on the ubiquitin-proteasome system (Fig. 2G). In addition, pretreatment with lenalidomide to saturate CRBN binding or pretreatment with entrectinib or JWJ-01-392 to saturate TRK-kinase domain binding also rescued the degradation of TPM3-TRKA by JWJ-01-378 (Fig. 2G). By contrast, inhibition of autophagy with bafilomycin A1 failed to rescue TPM3-TRKA loss upon JWJ-01-378 treatment, further validating that JWJ-01-378 induces TRK degradation through the proteasome (Fig. 2G). Together these findings demonstrate that JWJ-01-378 induces rapid loss of the TPM3-TRKA fusion protein through a targeted protein degradation mechanism, leading to impairment of downstream signaling.
To evaluate the proteome-wide specificity of JWJ-01-378, we performed data-independent acquisition (DIA) proteomic profiling of KM12 cells treated with 100 nM of JWJ-01-378 for two hours (Fig. 2H and Dataset S1). JWJ-01-378 led to significant TRK fusion degradation with minimal decreases in other proteins. Proteins that displayed alterations included NDUFS5 (NADH: Ubiquinone Oxidoreductase Subunit S5), KRR1 (KRR1 Small Subunit Processome Component), SARNP (SAP Domain Containing Ribonucleoprotein), COX16 (Cytochrome C Oxidase Assembly Protein COX16), MAP1LC3B (Microtubule Associated Protein 1 Light Chain 3 Beta) and GTF2E2 (General Transcription Factor IIE Subunit 2), which may have resulted as a consequence of TPM3-TRKA fusion degradation and rapid changes in downstream signaling that are observed as early as one hour after treatment. None of these proteins have been reported as entrectinib binders or CRBN neosubstrates. Additionally, JWJ-01-378 did not alter the levels of known kinase off-targets of entrectinib that were detected including JAK1, PTK2, SYK, LCK, ABL1, TYK2, and FGFR3 (Fig. 2H and Dataset S1). Together, these data confirm that JWJ-01-378 is a fast-acting, selective, and potent degrader of TPM3-TRKA.
Entrectinib-derived PROTACs degrade wild-type TRK but not TRK inhibitor resistance mutants or EML4-ALK
As entrectinib has been shown to inhibit both WT TRK and TRK fusions,49 we sought to evaluate WT TRK degradation in human erythroleukemia (HEL) cells, which endogenously express WT TRKA (Fig. S5A) and NIH/3T3 cells ectopically expressing WT TRKA. Consistent with prior work characterizing CG-428,36 treatment of HEL cells with JWJ-01-378 resulted in a significant reduction in WT TRKA protein levels, without impacting cell viability (Fig. 3A and Fig. S7A and B). The reduction of WT TRKA by JWJ-01-378 and CG-428 was confirmed in NIH/3T3 cells ectopically expressing WT TRKA (Fig. 3B). As expected, JWJ-01-392 did not degrade WT TRKA (Fig. 3A and B).We next evaluated whether these PROTACs were able to degrade TRK-inhibitor resistance mutants. We expressed the G595R point mutation, which confers resistance to entrectinib,50 in NIH/3T3 cells and compared the degradation to TPM3-TRKA. While we observed degradation of TPM3-TRKA, JWJ-01-378 and CG-428 failed to induce degradation of TPM3-TRKA-G595R (Fig. 3C and D). In addition, a reported off-target of entrectinib is ALK,13 and we evaluated whether entrectinib-derived PROTACs were able to degrade ALK fusions in H3122 cells, which endogenously express EML4-ALK, and NIH/3T3 cells ectopically expressing EML4-ALK or a drug-refractory mutant EML4-ALK-G1202R.51 At the doses and time-points assessed, we did not observe degradation of EML4-ALK or EML4-ALK-G1202R upon treatment with JWJ-01-378 or CG-428 (Fig. 3E–G). Together, these findings indicate JWJ-01-378 and CG-428 degrade WT TRK and TPM3-TRKA and are unable to degrade TPM3-TRKA-G595R, EML4-ALK, or EML4-ALK-G1202R.
JWJ-01-378 does not degrade cereblon neosubstrates that are targeted by CG-428
Prior work has shown that kinase-targeting PROTACs may have off-target activity towards several CRBN neosubstrates, leading to loss of viability through mechanisms unrelated to degradation of the kinase.52,53 To further understand the selectivity profile of JWJ-01-378, we surveyed CRBN neosubstrates including the translation termination factor GSPT1 and the transcription factors IKZF1 and IKZF3. Off-target degradation of GSPT1 is particularly important as it is a pan-essential gene,41 and degraders such as CC-885 have shown toxicity in clinical trials.54 We also evaluated the activity of CG-428, which was not previously explored in depth for CRBN neosubstrate activity.36 Upon evaluation of KM12 cells, which express GSPT1 and IKZF3, and are genetically dependent on GSPT1 (Fig. S8A–C), we did not observe changes in GSPT1 or IKZF3 levels in response to JWJ-01-378 treatment (Fig. 4A and B). However, we observed substantial degradation of GSPT1 and IKZF3 by CG-428 at doses and time-points that were not previously investigated36 (Fig. 4A and B).This result prompted us to validate these observations in MM1.S cells, a multiple myeloma cell line that expresses and is genetically dependent on GSPT1, IKZF1 and IKZF3 (Fig. S8A–C). MM1.S cells do not express WT TRK or TRK fusions (Fig. S5A–C). Consistent with KM12 cells, we did not observe changes in GSPT1, IKZF1 or IKZF3 levels in response to JWJ-01-378 but observed substantial reductions in each of these proteins in response to CG-428 (Fig. 4C and D). This degradation led to reduced viability of MM1.S cells upon CG-428 treatment, with increased activity compared to pomalidomide (Fig. 4E). Importantly, JWJ-01-378 did not reduce the viability of MM1.S cells (Fig. 4E). We further confirmed the degradation of IKZF1 and IKZF3 by CG-428 in K-562 cells, a lymphoblast cell line that expresses IKZF1 and IKZF3 (Fig. S8A, B, and D). Together, these data support the improved specificity of JWJ-01-378 with respect to CRBN neosubstrate degradation, relative to CG-428.
JWJ-01-378 diminishes viability of TRK fusion-positive cells
To assess the biological consequences of PROTAC-mediated degradation of TPM3-TRKA, we evaluated the antiproliferative effects of our compounds in 2D-adherent monolayer and 3D-spheroid cultures. 3D-spheroids are valuable models for providing predictive insights into in vivo efficacy.8,32 KM12 cells were treated with a panel of TRK inhibitors (entrectinib and GNF-8625), PROTACs (JWJ-01-375, JWJ-01-377, JWJ-01-378, and CG-428), and a heterobifunctional control (JWJ-01-392) (Fig. 5A–D and Table S1). In line with the observed degradation effects, JWJ-01-378 demonstrated more potent antiproliferative activity in both 2D-adherent monolayer and 3D-spheroid cultures, compared to JWJ-01-375 and JWJ-01-377, with a substantial shift in IC50 compared to JWJ-01-392. While CG-428 led to a more pronounced antiproliferative effect compared to JWJ-01-378, this is likely due to the degradation of CRBN neosubstrates including GSPT1 by CG-428 (Fig. 4A and B and Fig. S8C).Compared to the parental inhibitors, JWJ-01-378 was less effective than entrectinib and GNF-8625, which may be due to several factors including higher cell permeability, contributions from polypharmacology, or compound efflux. In particular, drug efflux has emerged as a common mechanism of diminishing PROTAC activity.55,56 To test whether ABCB1 transporters were contributing to these differences, we co-treated entrectinib, JWJ-01-378 or CG-428 with the ABCB1-efflux pump inhibitors tariquidar (Fig. S9A) and zosuquidar (Fig. S9B). We also evaluated whether a second treatment of entrectinib, JWJ-01-378, or CG-428 at 72 hours improved responses (Fig. S9C). Co-treatment with tariquidar or zosuquidar or a second treatment of the compounds did not substantially improve the antiproliferative effects of the compounds. Together, these data suggest that JWJ-01-378 diminishes the proliferation of TPM3-TRKA fusion-positive cells, with improved effects compared to JWJ-01-392, its matched heterobifunctional control compound.
Conclusions
TRK fusions are oncogenic drivers of several adult and pediatric cancers. First-generation and second-generation TRK inhibitors have demonstrated efficacy in clinical trials in patients with TRK-driven malignancies including SCOUT (NCT02637687) and NAVIGATE (NCT02576431).6,57 However, these inhibitors have off-target effects and the development of acquired resistance limits their long-term clinical utility. Targeted protein degradation is a promising approach to circumvent these limitations by promoting the complete removal of wild-type and mutant proteins.58–61 In this study, we aimed to develop PROTACs to target TRK fusion proteins for degradation. Using structure-guided design, we conjugated entrectinib to thalidomide and nominated JWJ-01-378 as a potent TRK-targeting PROTAC. JWJ-01-378 induces the degradation of WT TRK and TRK fusions within 2 hours of treatment at a 10 nM dose, diminishes levels of downstream pERK signaling, and requires the ubiquitin-proteasome system for degradation. Proteomics analysis confirmed the specificity of TPM3-TRKA degradation at two hours. In-depth characterization of the levels of CRBN neosubstrates including GSPT1, IKZF1, and IKZF3 confirmed the specificity of JWJ-01-378 and uncovered liabilities of CG-428, which significantly degraded these targets.We also compared the biological responses upon JWJ-01-378 treatment to inhibitors (entrectinib and GNF-8625), CG-428, and a heterobifunctional control compound (JWJ-01-392). While JWJ-01-378 achieved substantial TPM3-TRKA degradation, comparable to CG-428, it did not diminish the viability of KM12 cells to levels observed with entrectinib, GNF-8625, or CG-428. Importantly, the improved antiproliferative effects upon CG-428 treatment may be due to off-target degradation of CRBN neosubstrates including GSPT1, which is a pan-essential genetic dependency. The improved antiproliferative effects upon entrectinib treatment may be due to differential impacts on downstream pathways or off-target effects62 that are spared by JWJ-01-378. We confirmed that the differences in potency with entrectinib were not due to ABCB1-mediated efflux and were not improved with a second administration of TRK PROTACs. Critically, JWJ-01-378 had improved antiproliferative activity in 2D-monolayer and 3D-spheroids compared to JWJ-01-392, supporting the enhanced degradation-dependent effects over a matched heterobifunctional control compound. Notably, JWJ-01-378 also degrades WT TRK, which presents an interesting opportunity for an application towards chronic pain management, as WT TRK is a known mediator of chronic pain and nociception.63
With the compounds developed in this study, there are several future directions to pursue that were not addressed in this work. First, it will be important to determine how TRK fusion degradation impacts downstream signaling networks to nominate potential drug combination strategies. In our time-course experiments with JWJ-01-378 and entrectinib, we observed reactivation of pERK signaling as early as 24 hours after treatment, returning to the baseline levels by five days. Additionally, we observed partial recovery of TPM3-TRKA levels by five days, which may be due to several factors including compound stability, transcriptional feedback, or alterations in the levels or activity of the CRBN complex. This increase in TPM3-TRKA levels is delayed compared to pERK reactivation but may contribute to the recovery of pERK signaling. In addition, compensatory rewiring of the MAPK pathway is a well-documented mechanism of resistance to TRK inhibitors and may involve reactivation of alternative receptor tyrosine kinases such as EGFR24 or secondary mutations including BRAFV600E or KRASG12D, or amplification of MET.50 Evaluation of combinatorial strategies with MAPK or PI3K pathway inhibitors to enhance the durability of TRK degradation effects is necessary, which has been previously shown to be effective in the context of TRK inhibitors.24,64
Next, it will be important to define the mechanisms of resistance to TRK PROTACs. Although JWJ-01-378 efficiently degraded WT and TPM3-TRKA fusion proteins, it failed to degrade the TPM3-TRKA-G595R resistance mutant. This finding aligns with previous studies showing that the G595R mutation in the kinase domain of TRK confers resistance to TRK inhibitors, such as entrectinib.23 Furthermore, identifying the spectrum of TRK fusions targeted by JWJ-01-378 due to its binding to the TRK kinase domain will be important to assess its ability to tackle the EML4-TRK, ETV6-TRK, and SQSTM1-TRK fusions that are observed in many cancers. Interestingly, while ALK is a known off-target of entrectinib, JWJ-01-378 did not degrade EML4-ALK or EML4-ALK-G1202R, suggesting selectivity for TRK over ALK proteins, and the necessity of these evaluations. Finally, continued evaluation of TRK-targeting PROTACs in mouse models of cancer is necessary to understand toxicities and potential as a therapeutic approach in cancer.
In conclusion, JWJ-01-378 represents a potent TRK-targeting PROTAC that efficiently reduces TPM3-TRKA fusion protein levels, displays an improved specificity profile compared to existing TRK PROTACs that degrade CRBN neosubstrates, and suppresses cancer cell proliferation. However, its inability to degrade TRK resistance mutants and the activation of compensatory MAPK signaling responses highlight the need for further optimization and combinatorial treatment strategies. These findings provide a strong foundation for the continued development of TRK-targeting PROTACs as therapeutic agents for TRK fusion-driven cancers.
Methods
Synthetic procedures
Methyl 4-bromo-2-((tetrahydro-2H-pyran-4-yl)amino)benzoate
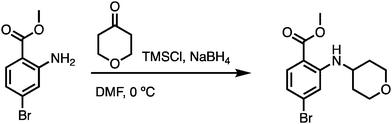
According to reported literature,67 a dry 100 mL two-neck round bottom flask with magnetic stirring bar was charged consecutively with methyl 2-amino-4-bromobenzoate (1.0 equiv.), tetrahydro-4h-pyran-4-one (1.1 equiv.), TMSCl (2.5 equiv.), and dry DMF in a nitrogen atmosphere. The reaction mixture was cooled to 0 °C and NaBH4 (1.0 equiv.) was added in one portion. The reaction mixture was kept stirring at 0 °C until full conversion of the arylamine was detected by UPLC. The vigorously stirred reaction mixture was treated with saturated NaHCO3 solution, followed by EtOAc extraction. The two-phase mixture was kept stirring until the gas evolution had ceased. The phases were separated, and the aqueous layer was further extracted with EtOAc once. The combined organic layers were dried over anhydrous MgSO4. The drying agent was removed by filtration, and the solvent was removed under reduced pressure. The crude product was loaded on the silica gel and purified by chromatography, which gave the title compound as pale-yellow solid. 1H NMR (500 MHz, DMSO-d6) δ 7.79 (d, J = 7.8 Hz, 1H), 7.70 (dd, J = 8.6, 1.1 Hz, 1H), 7.07 (d, J = 1.8 Hz, 1H), 6.74 (dt, J = 8.6, 1.5 Hz, 1H), 3.83 (dt, J = 11.6, 3.7 Hz, 2H), 3.79 (d, J = 1.1 Hz, 3H), 3.77–3.69 (m, 1H), 3.53–3.45 (m, 2H), 1.96–1.88 (m, 2H), 1.44–1.34 (m, 2H). MS (ESI) m/z 314.2 (M+H)+.
tert-Butyl 4-(4-(methoxycarbonyl)-3-((tetrahydro-2H-pyran-4-yl)amino)phenyl)piperazine-1-carboxylate
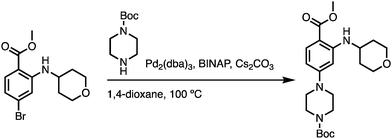
To a two-neck round bottom flask equipped with a nitrogen balloon and a septum, the resulting reductive amination intermediate (1.0 equiv.), BINAP (0.2 equiv.), Pd2(dba)3 (0.1 equiv.), Cs2CO3 (3.0 equiv.), and tert-butyl piperazine-1-carboxylate (1.1 equiv.) were charged. Anhydrous 1,4-dixoane was then added and the system was flushed with nitrogen gas three times. The mixture was heated to 100 °C and stirred overnight. Upon complete consumption of reductive amination intermediate monitored by UPLC, 50 mL EtOAc was added and filtered through a pad of Celite. The filtrate was saved and washed with brine twice. The organic layer was dried over anhydrous MgSO4, concentrated under reduced pressure, and purified via flash column chromatography, which provided the title compound as pale-yellow solid. 1H NMR (500 MHz, DMSO-d6) δ 7.76 (d, J = 7.7 Hz, 1H), 7.63 (dd, J = 9.2, 2.6 Hz, 1H), 6.21 (dd, J = 9.3, 2.5 Hz, 1H), 6.08 (d, J = 2.4 Hz, 1H), 3.83 (dt, J = 11.7, 3.5 Hz, 2H), 3.71 (d, J = 2.5 Hz, 4H), 3.53–3.47 (m, 2H), 3.43 (s, 4H), 3.27 (t, J = 5.5 Hz, 4H), 1.96 (t, J = 12.1 Hz, 2H), 1.42 (d, J = 2.6 Hz, 11H). MS (ESI) m/z 420.5 (M+H)+.
4-(4-(tert-Butoxycarbonyl)piperazin-1-yl)-2-((tetrahydro-2H-pyran-4-yl)amino)benzoic acid
The methyl ester intermediate (1.0 equiv.) was dissolved in methanol before the addition of 1 M NaOH solution (3.0 equiv.). The mixture was stirred for 8 hours at 60 °C, which resulted in a brown clear solution. The solution was then chilled on ice and acidified with 1M HCl solution to pH 2–3, which generated a lot of solid. The cloudy mixture was further stirred for 30 minutes before the filtration to collect the solid. The pale-white solid was dried under reduced pressure to yield the title acid intermediate without further purification. 1H NMR (500 MHz, DMSO-d6) δ 11.93 (s, 1H), 7.93 (d, J = 7.6 Hz, 1H), 7.61 (d, J = 9.0 Hz, 1H), 6.18 (dd, J = 9.1, 2.3 Hz, 1H), 6.07 (d, J = 2.4 Hz, 1H), 3.83 (dt, J = 11.7, 3.8 Hz, 2H), 3.70 (d, J = 11.0 Hz, 1H), 3.49 (td, J = 11.2, 2.4 Hz, 2H), 3.43 (d, J = 10.6 Hz, 4H), 3.24 (t, J = 5.3 Hz, 4H), 1.94 (d, J = 12.9 Hz, 2H), 1.41 (s, 9H), 1.35 (ddd, J = 13.8, 8.6, 3.9 Hz, 2H). MS (ESI) m/z 406.6 (M+H)+.
4-(4-(tert-Butoxycarbonyl)piperazin-1-yl)-2-(2,2,2-trifluoro-N-(tetrahydro-2H-pyran-4-yl)acetamido)benzoic acid
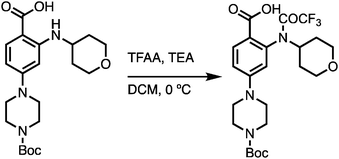
To the acid intermediate (1.0 equiv.) solution in DCM, TEA (2.0 equiv.) was added. The solution was cooled to 0 °C before the addition of TFAA (1.5 equiv.). The reaction was kept stirring at 0 °C until full conversion of the acid was detected by UPLC. The solvent was removed under reduced pressure, which yielded the desired protected intermediate as yellow oil with no further purification. MS (ESI) m/z 502.3 (M+H)+.
tert-Butyl 4-(4-((5-(3,5-difluorobenzyl)-1H-indazol-3-yl)carbamoyl)-3-((tetrahydro-2H-pyran-4-yl)amino)phenyl)piperazine-1-carboxylate
According to reported literature,19 to a suspension of acid trifluoroacetate (1.0 equiv.) in dry DCM, oxalyl chloride (1.5 equiv.) and DMF (1-2 drops, catalytic amount) were added. The mixture was stirred at room temperature for 2 hours then evaporated to dryness. The resulting crude acyl chloride was taken-up with toluene, evaporated, and then dissolved in dry THF at 0 °C. A solution of 5-(3,5-difluoro-benzyl)-1H-indazol-3-ylamine (1.0 equiv.) and DIPEA (4.0 equiv.) in dry THF was added to the cooled acyl chloride solution. The mixture was stirred at 0 °C for 4 hours then quenched by adding DI H2O/EtOAc. The organic phase was washed with a saturated NaHCO3 solution, dried over MgSO4, and evaporated to dryness. Crude reaction mixture is dissolved in methanol in the presence of triethylamine (3.0 equiv.) and stirred at 65 °C for 2 hours. The solvent was removed under reduced pressure and the residue treated with DI H2O/EtOAc. Organic layer was dried over anhydrous MgSO4 and evaporated to dryness. The crude product was further purified by chromatography over silica gel to generate the title compound as a pale yellow solid. 1H NMR (500 MHz, DMSO-d6) δ 12.66 (s, 1H), 10.12 (s, 1H), 8.29 (d, J = 7.6 Hz, 1H), 7.81 (d, J = 9.0 Hz, 1H), 7.48 (s, 1H), 7.41 (d, J = 8.6 Hz, 1H), 7.25 (dd, J = 8.6, 1.6 Hz, 1H), 7.00 (ddd, J = 15.5, 7.8, 4.4 Hz, 3H), 6.24 (dd, J = 9.0, 2.3 Hz, 1H), 6.15 (d, J = 2.4 Hz, 1H), 4.05–4.02 (m, 2H), 3.81 (dt, J = 11.8, 3.8 Hz, 2H), 3.68 (d, J = 6.6 Hz, 1H), 3.51–3.41 (m, 6H), 3.25 (t, J = 5.3 Hz, 4H), 1.94 (d, J = 12.8 Hz, 2H), 1.43 (s, 9H), 1.33 (q, J = 13.5, 11.6 Hz, 2H). MS (ESI) m/z 647.5 (M+H)+.
tert-Butyl 2-(4-(4-((5-(3,5-difluorobenzyl)-1H-indazol-3-yl)carbamoyl)-3-((tetrahydro-2H-pyran-4-yl)amino)phenyl)piperazin-1-yl)acetate
To the Boc-protected intermediate (1.0 equiv.) in DCM, TFA (2.0 equiv.) was added. The resulting solution was stirred for 1 hour to fully remove the Boc group, indicated by UPLC. The solvent was then removed under reduced pressure before re-dissolving in dry DMF. tert-Butyl bromoacetate (1.1 equiv.) and DIPEA (1.5 equiv.) were added to the solution and stirred for another 1 hour at 45 °C. Upon complete consumption of free piperazine intermediate monitored by UPLC, DI H2O/EtOAc were added. The organic layer was dried over anhydrous MgSO4, concentrated under reduced pressure, and purified via flash column chromatography, which provided the title compound as pale-yellow solid. 1H NMR (500 MHz, DMSO-d6) δ 12.65 (s, 1H), 10.10 (s, 1H), 8.28 (d, J = 7.6 Hz, 1H), 7.79 (d, J = 9.0 Hz, 1H), 7.48 (s, 1H), 7.40 (d, J = 8.6 Hz, 1H), 7.25 (dd, J = 8.6, 1.6 Hz, 1H), 7.03–6.96 (m, 3H), 6.23 (dd, J = 9.0, 2.3 Hz, 1H), 6.13 (d, J = 2.3 Hz, 1H), 4.03 (s, 2H), 3.81 (dt, J = 11.6, 3.9 Hz, 2H), 3.71–3.64 (m, 1H), 3.52–3.46 (m, 2H), 3.27 (t, J = 4.9 Hz, 4H), 3.17 (s, 2H), 2.62 (t, J = 5.0 Hz, 4H), 1.93 (d, J = 12.8 Hz, 2H), 1.43 (s, 9H), 1.33 (td, J = 13.6, 9.9 Hz, 2H).
tert-Butyl 2-((2-(2,6-dioxopiperidin-3-yl)-1,3-dioxoisoindolin-4-yl)oxy)acetate
According to reported literature,68 2-(2,6-dioxopiperidin-3-yl)-4-hydroxyisoindoline-1,3-dione (1.0 equiv.) was dissolved in DMF at room temperature before the addition of K2CO3 (1.5 equiv.) and tert-butyl bromoacetate (1.0 equiv.). After 2 hours, the mixture was charged with DI H2O/EtOAc. The organic layer was dried over anhydrous MgSO4, concentrated under reduced pressure, and purified via flash column chromatography, which provided the title compound.
tert-Butyl (1-((2-(2,6-dioxopiperidin-3-yl)-1,3-dioxoisoindolin-4-yl)oxy)-2-oxo-6,9,12-trioxa-3-azatetradecan-14-yl)carbamate
To the Boc-protected intermediate (1.0 equiv.) in DCM, TFA (2.0 equiv.) was added. The resulting solution was stirred for 1 hour to fully remove the Boc group, indicated by UPLC. The solvent was then removed under reduced pressure before re-dissolving in dry DMF. t-Boc-N-amido-PEG3-amine (1.1 equiv.), HATU (1.5 equiv.), and DIPEA (3.0 equiv.) were successively charged to the reaction solution. After 2 hours, the reaction was charged with DI H2O/EtOAc. The organic layer was dried over anhydrous MgSO4, concentrated under reduced pressure, and purified via flash column chromatography, which provided the title compound.
N-(5-(3,5-Difluorobenzyl)-1H-indazol-3-yl)-4-(4-(17-((2-(2,6-dioxopiperidin-3-yl)-1,3-dioxoisoindolin-4-yl)oxy)-2,16-dioxo-6,9,12-trioxa-3,15-diazaheptadecyl)piperazin-1-yl)-2-((tetrahydro-2H-pyran-4-yl)amino)benzamide
To the Boc-protected intermediate (1.0 equiv.) in DCM, TFA (2.0 equiv.) was added. The resulting solution was stirred for 1 hour to fully remove the Boc group, indicated by UPLC. The solvent was then removed under reduced pressure before re-dissolving in dry DMF. Meanwhile, the tert-butyl ester was dissolved in DCM before the addition of TFA (3.0 equiv.). The solution was stirred for 5 hours to fully remove the ester group, monitored by UPLC. The solvent as well as excess TFA were evaporated under reduced pressure. The freshly generated acid (1.0 equiv.) in DMF, HATU (1.5 equiv.), and DIPEA (3.0 equiv.) were successively added to Boc-deprotected amine solution. Upon the completion of the amide coupling reaction indicated by UPLC, the residue was purified by prep HPLC to yield the desired product (8.1 mg, 17%) with 94% purity. 1H NMR (500 MHz, DMSO-d6) δ 12.70 (s, 1H), 11.13 (s, 1H), 10.18 (s, 1H), 8.68 (t, J = 5.6 Hz, 1H), 8.01 (t, J = 5.8 Hz, 1H), 7.82 (dd, J = 17.1, 8.7 Hz, 2H), 7.51 (d, J = 7.3 Hz, 1H), 7.46 (s, 1H), 7.41 (dd, J = 8.6, 5.9 Hz, 2H), 7.26 (d, J = 8.6 Hz, 1H), 7.04–6.94 (m, 3H), 6.27 (d, J = 8.8 Hz, 1H), 6.18 (s, 1H), 5.12 (dd, J = 12.8, 5.4 Hz, 1H), 4.79 (s, 2H), 4.03 (d, J = 9.1 Hz, 8H), 3.81 (dd, J = 9.4, 5.6 Hz, 4H), 3.52 (s, 10H), 3.47 (d, J = 5.7 Hz, 4H), 3.32 (q, J = 5.7 Hz, 4H), 3.22 (s, 4H), 2.90 (ddd, J = 17.6, 13.6, 5.4 Hz, 1H), 2.65–2.51 (m, 2H), 2.08–2.01 (m, 1H), 1.93 (d, J = 12.7 Hz, 2H), 1.34 (q, J = 13.0, 11.2 Hz, 2H). MS (ESI) m/z 1093.6 (M+H)+.
N-(5-(3,5-Difluorobenzyl)-1H-indazol-3-yl)-4-(4-(14-((2-(2,6-dioxopiperidin-3-yl)-1,3-dioxoisoindolin-4-yl)oxy)-2,13-dioxo-6,9-dioxa-3,12-diazatetradecyl)piperazin-1-yl)-2-((tetrahydro-2H-pyran-4-yl)amino)benzamide
By employment of the similar procedure to JWJ-01-375, from the corresponding carboxylic acid (1.0 equiv.), the title compound was prepared (6.7 mg, 17%) with 99% purity. 1H NMR (500 MHz, DMSO-d6) δ 12.70 (s, 1H), 11.13 (s, 1H), 10.17 (s, 1H), 8.68 (s, 1H), 8.02 (s, 1H), 7.87–7.79 (m, 2H), 7.51 (d, J = 7.3 Hz, 1H), 7.46 (s, 1H), 7.41 (d, J = 8.6 Hz, 2H), 7.28 (s, 1H), 7.05–6.96 (m, 3H), 6.29 (s, 1H), 6.18 (s, 1H), 5.12 (dd, J = 12.8, 5.4 Hz, 2H), 4.80 (s, 2H), 4.03 (d, J = 8.0 Hz, 6H), 3.82 (d, J = 11.2 Hz, 2H), 3.69 (s, 1H), 3.57–3.52 (m, 5H), 3.47 (td, J = 5.7, 2.3 Hz, 7H), 3.33 (q, J = 5.5 Hz, 4H), 3.22 (s, 4H), 2.94–2.86 (m, 1H), 2.63–2.53 (m, 2H), 2.07–2.00 (m, 1H), 1.93 (d, J = 12.3 Hz, 2H), 1.34 (d, J = 11.0 Hz, 2H). MS (ESI) m/z 1049.7 (M+H)+.
N-(5-(3,5-Difluorobenzyl)-1H-indazol-3-yl)-4-(4-(2-((8-(2-((2-(2,6-dioxopiperidin-3-yl)-1,3-dioxoisoindolin-4-yl)oxy)acetamido)octyl)amino)-2-oxoethyl)piperazin-1-yl)-2-((tetrahydro-2H-pyran-4-yl)amino)benzamide
By employment of the similar procedure to JWJ-01-375, from the corresponding carboxylic acid (1.0 equiv.), the title compound was prepared (6.3 mg, 16%) with 92% purity. 1H NMR (500 MHz, DMSO-d6) δ 12.69 (s, 1H), 11.13 (s, 1H), 10.18 (s, 1H), 8.54 (t, J = 5.7 Hz, 1H), 7.95 (t, J = 5.8 Hz, 1H), 7.87–7.77 (m, 2H), 7.51 (d, J = 7.3 Hz, 1H), 7.46 (s, 1H), 7.40 (t, J = 7.8 Hz, 2H), 7.27 (s, 1H), 7.05–6.95 (m, 3H), 6.28 (d, J = 8.7 Hz, 1H), 6.19 (s, 1H), 5.12 (dd, J = 12.8, 5.4 Hz, 1H), 4.77 (s, 2H), 4.02 (d, J = 18.5 Hz, 6H), 3.90–3.80 (m, 4H), 3.48 (t, J = 10.8 Hz, 4H), 3.32–3.06 (m, 8H), 2.96–2.88 (m, 1H), 2.64–2.51 (m, 2H), 2.07–2.01 (m, 1H), 1.93 (d, J = 12.4 Hz, 2H), 1.44 (t, J = 6.9 Hz, 4H), 1.34 (d, J = 10.6 Hz, 2H), 1.26 (s, 8H). MS (ESI) m/z 1045.7 (M+H)+.
N-(5-(3,5-Difluorobenzyl)-1H-indazol-3-yl)-4-(4-(2-((8-(2-((2-(1-methyl-2,6-dioxopiperidin-3-yl)-1,3-dioxoisoindolin-4-yl)oxy)acetamido)octyl)amino)-2-oxoethyl)piperazin-1-yl)-2-((tetrahydro-2H-pyran-4-yl)amino)benzamide
By employment of the similar procedure to JWJ-01-375, from the corresponding carboxylic acid (1.0 equiv.), the title compound was prepared (6.9 mg, 18%) with 99% purity. 1H NMR (500 MHz, DMSO-d6) δ 12.71 (s, 1H), 10.19 (s, 1H), 8.55 (d, J = 5.8 Hz, 1H), 7.97 (t, J = 5.9 Hz, 1H), 7.88–7.78 (m, 2H), 7.51 (d, J = 7.2 Hz, 1H), 7.46 (s, 1H), 7.40 (dd, J = 8.6, 6.0 Hz, 2H), 7.26 (d, J = 8.5 Hz, 1H), 7.00 (dd, J = 20.8, 8.7 Hz, 3H), 6.27 (d, J = 9.0 Hz, 1H), 6.18 (s, 1H), 5.19 (dd, J = 13.0, 5.4 Hz, 1H), 4.77 (s, 2H), 4.02 (d, J = 15.0 Hz, 6H), 3.82 (d, J = 8.4 Hz, 4H), 3.69 (s, 2H), 3.48 (t, J = 10.8 Hz, 4H), 3.21 (s, 2H), 3.14 (q, J = 6.6 Hz, 4H), 3.02 (s, 3H), 2.97–2.91 (m, 1H), 2.80–2.74 (m, 1H), 2.55 (d, J = 8.5 Hz, 1H), 2.05 (d, J = 12.7 Hz, 1H), 1.93 (d, J = 12.5 Hz, 2H), 1.43 (t, J = 6.9 Hz, 4H), 1.34 (d, J = 10.4 Hz, 2H), 1.26 (s, 8H). MS (ESI) m/z 1059.9 (M+H)+.
Molecular docking
Ternary complex docking was conducted using the MOE2024.0601 suite. In brief, the reported co-crystal structures of CRBN and NTRK with corresponding small molecules were first examined by the QuickPrep module to fix the missing residues. The PROTACs were imported and saved as individual files. Next, the prepared binary complex and the PROTACs were imported to the three-body conformational search method to run the ternary complex docking. After the conformational search, the top clustered complexes undergo the energy minimization for the predicted binding mode of each PROTAC molecule.Additional compounds
Entrectinib (Selleck Chemicals, #S7998), GNF-8625 (MedChemExpress, #HY-131706A), CG-428 (Tocris, #7425), carfilzomib (MedChemExpress, #HY-10455), MLN4924 (Calbiochem, #5054770001), lenalidomide (Millipore Sigma, #901558), bafilomycin A1 (Cambridge Bioscience, #b0025) and pomalidomide (MedChemExpress, #HY-10984) were purchased from the indicated sources.Cell line culturing
KM12 (kindly provided by the MD Anderson Cancer Center, Houston, TX), HEL 92.1.7 (ATCC, #TIB-180), and MM.1S (ATCC, #CRL-2974) cells were cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum, 100 U mL−1 penicillin, and 100 µg mL−1 streptomycin at 37 °C with 5% CO2. K-562 (ATCC, #CCL-243) cells were cultured in Iscove's modified Dulbecco's medium (IMDM) supplemented with 10% fetal bovine serum, 100 U mL−1 penicillin, and 100 µg mL−1 streptomycin at 37 °C with 5% CO2. DF-1 cells (ATCC, #CRL-12203) were cultured in Dulbecco's modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum, 100 U mL−1 penicillin, and 100 µg mL−1 streptomycin at 39 °C with 5% CO2. NIH/3T3-tv-a cells69 were cultured in DMEM supplemented with 10% calf serum, 100 U mL−1 penicillin, and 100 µg mL−1 streptomycin at 37 °C with 5% CO2. Cell lines were confirmed for mycoplasma negative monthly using the MycoProbe Mycoplasma detection kit (R&D systems, #CUL001B).Viral production and stable cell line generation
For the generation of NIH/3T3 cells expressing TRK fusions, DF-1 cells were transfected with the indicated RCAS plasmid using X-tremeGENE 9 DNA transfection reagent (Roche, #XTG9-RO) according to the manufacturer's protocol. Viral supernatant from DF-1 cultures was sterile-filtered and added to NIH/3T3-tv-a cells every eight hours for three cycles.Immunoblotting
Immunoblotting was performed as previously described.70 Cell lines were lysed in RIPA buffer containing cOmplete protease inhibitor (Millipore Sigma, #11873580001), PhosSTOP phosphatase inhibitor (Millipore Sigma, #4906837001), and 0.1% Benzonase (Millipore Sigma, #712063) on the ice for 1 hour. Lysates were centrifuged at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 10 minutes at 4 °C and clarified supernatants were transferred to the pre-chilled tubes. Protein lysates were quantified and loaded in equal amounts on 4–12% SDS-PAGE gradient gels, followed by transfer onto nitrocellulose membranes using an iBlot3 (Thermo Fisher). The membranes were incubated overnight with the indicated primary antibodies: pan-TRK (Cell Signaling Technology, #92991), β-Actin (Cell Signaling Technology, #3700), p44/42 MAPK (ERK1/2) (Cell Signaling Technology, #4696), Phospho-p44/42 MAPK (ERK1/2) (Thr202/Tyr204) (Cell Signaling Technology, #4370), Ikaros (Cell Signaling Technology, #14859), Aiolos (Cell Signaling Technology, #15103), eRF3a (Cell Signaling Technology, #70152), and ALK (Cell Signaling Technology, #3633). Membranes were washed the next day with TBS-T (TBS with 0.1% Tween), incubated with fluorescently labeled infrared secondary antibodies (Li-COR) at room temperature for 1 hour, and imaged using the LI-COR Odyssey® Imaging System.
000 rpm for 10 minutes at 4 °C and clarified supernatants were transferred to the pre-chilled tubes. Protein lysates were quantified and loaded in equal amounts on 4–12% SDS-PAGE gradient gels, followed by transfer onto nitrocellulose membranes using an iBlot3 (Thermo Fisher). The membranes were incubated overnight with the indicated primary antibodies: pan-TRK (Cell Signaling Technology, #92991), β-Actin (Cell Signaling Technology, #3700), p44/42 MAPK (ERK1/2) (Cell Signaling Technology, #4696), Phospho-p44/42 MAPK (ERK1/2) (Thr202/Tyr204) (Cell Signaling Technology, #4370), Ikaros (Cell Signaling Technology, #14859), Aiolos (Cell Signaling Technology, #15103), eRF3a (Cell Signaling Technology, #70152), and ALK (Cell Signaling Technology, #3633). Membranes were washed the next day with TBS-T (TBS with 0.1% Tween), incubated with fluorescently labeled infrared secondary antibodies (Li-COR) at room temperature for 1 hour, and imaged using the LI-COR Odyssey® Imaging System.
Analysis of dependency map data
The expression levels and dependency scores for genes-of-interest were analyzed using the Dependency Map (DepMap Public 25Q3, https://depmap.org/)41 and plotted using GraphPad PRISM v10.Analysis of cell viability in 2D-adherent monolayer and 3D-spheroid culture
2D-adherent monolayer and 3D-spheroid culture experiments were performed as previously described32,70 with minor modifications. For 2D-adherent viability assays, KM12 cells were plated in white 384-well culture plates (Corning, #3570) at a density of 500 cells per well in 50 µL media. For ultra-low adherent 3D-spheroid viability assays, KM12 cells were plated in PrimeSurface 384-well 3D culture spheroid plates (S-bio, #MS-9384WZ) at a density of 500 cells per well in 50 µL media. HEL 92.1.7 cells, which are suspension cultures, were plated in white 384-well culture plates (Corning, #3570) at a density of 1000 cells per well in 50 µL media. MM1.S cells, which are semi-adherent cultures, were plated in white 384-well culture plates (Corning, #3570) at a density of 1000 cells per well in 50 µL media. The next day, cells were treated with compounds using a D300e digital dispenser (HP), and all wells were normalized to 100 nL total DMSO. For the drug efflux inhibitor experiments, cells were treated simultaneously with test compounds and zosuquidar 3HCl (Selleck Chemicals, #S1481) or tariquidar (MedChemExpress, #HY-10550) and incubated for 120 hours. For experiments where a second dose of compound was administered, cells were treated with indicated compounds again at 72 hours. In all experiments, after 120 hours from the start of treatment, cells were equilibrated at room temperature for 30 minutes, followed by the addition of 10 µL CellTiter-Glo (Promega, #G7570) per well and shaking for 5 minutes at room temperature. After incubation at room temperature for 20 minutes, luminescence was measured on the CLARIOstar Plus Plate Reader (BMG LabTech). The data was normalized to DMSO-treated control wells and analyzed using GraphPad PRISM v10.Data-independent acquisition (DIA)-based quantitative proteomics
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well in 200 µL of media in 96 well plates (Greiner, #655180) using a Viaflo liquid handler (Integra). After 24 hours, cells were treated with compounds using D300e digital dispenser (Tecan) in triplicate. Cells were treated using Integra Viaflo by transferring media off of cells, dispensing and mixing on a compound plate to resuspend them, and then transferring the media plus drug back onto cells prior to incubation. Cells were treated with 100 nM of JWJ-01-378 for two hours followed by protein fractionation, digestion, and mass spectrometry DIA analysis.
000 cells per well in 200 µL of media in 96 well plates (Greiner, #655180) using a Viaflo liquid handler (Integra). After 24 hours, cells were treated with compounds using D300e digital dispenser (Tecan) in triplicate. Cells were treated using Integra Viaflo by transferring media off of cells, dispensing and mixing on a compound plate to resuspend them, and then transferring the media plus drug back onto cells prior to incubation. Cells were treated with 100 nM of JWJ-01-378 for two hours followed by protein fractionation, digestion, and mass spectrometry DIA analysis.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 enzyme to protein ratio. Proteins were digested into peptides for approximately 16–20 hours with trypsin at 37 °C.
10 enzyme to protein ratio. Proteins were digested into peptides for approximately 16–20 hours with trypsin at 37 °C.
Author contributions
SK, MSDP, and JC performed all the biological experiments. JJ performed compound design and synthesis and docking modeling. AG and AJF performed proteomics studies and AG, AJF, and SK analyzed the data. FS and ECH provided resources and analyzed and interpreted the data. FMF and BN conceptualized the study, designed the experiments, analyzed and interpreted the data, acquired funding, and supervised the study. SK and BN wrote the manuscript with input from all authors.Conflicts of interest
AG and AJF are employees of Talus Bioscience, Inc. FMF is a scientific cofounder and equity holder in Proximity Therapeutics, has served as a science advisory board member to Triana Biomedicines, and has served as a consultant and/or has received speaker honoraria from RA Capital, Triana Biomedicines, Eli Lily and Co., Sorrento Pharma, Plexium Inc., Tocris Biotechne, Neomorph Inc., Bristol-Myers Squibb, Novartis, and Amgen. The Ferguson lab receives or has received research funding or resources in-kind from Ono Pharmaceuticals Ltd., Merck & Co., Eli Lilly and Co., Promega, Mirati, and J&J. These interests have been reviewed and approved by the University of California San Diego in accordance with its conflict-of-interest policies. The Nabet laboratory receives or has received research funding from Mitsubishi Tanabe Pharma America, Inc. No disclosures were reported by the other authors.Data availability
The raw proteomics datasets generated during this study are available at the ProteomeXchange Consortium via the PRIDE partner repository65,66 with the dataset identifier PXD065035. Normalized and scaled proteomics data are provided in Dataset S1. Publicly available data was used to determine the expression levels and dependency scores for genes-of-interest using the Dependency Map (DepMap Public 25Q3, https://depmap.org/).41 Supplementary information (SI) includes tables, figures, and 1H NMR spectra. Uncropped immunoblots are provided in Fig. S10–S18. See DOI: https://doi.org/10.1039/d6cb00008h.Acknowledgements
The authors thank members of the Nabet and Ferguson laboratories for feedback on this study and manuscript. This work was supported by: Hartwell Innovation Fund – Swim Across America (SK), NSF CAREER CHE-2339705 (FMF), NIH/NCI K22 CA258805 (BN), NIH/NCI Cancer Center Support Grant P30 CA015704 (BN), NIH/NCI U01 CA282109 (BN), and NIH/NCATS R43 TR004221 (AJF and BN). This research was supported by the Cellular Imaging Shared Resource, RRID:SCR_022609 of the Fred Hutch/University of Washington/Seattle Children's Cancer Consortium (P30 CA015704).References
- A. Amatu, A. Sartore-Bianchi, K. Bencardino, E. G. Pizzutilo, F. Tosi and S. Siena, Ann. Oncol., 2019, 30, viii5–viii15 CrossRef CAS PubMed.
- M. E. Cunningham and L. A. Greene, EMBO J., 1998, 17, 7282–7293 CrossRef CAS PubMed.
- F. Coulier, R. Kumar, M. Ernst, R. Klein, D. Martin-Zanca and M. Barbacid, Mol. Cell. Biol., 1990, 10, 4202–4210 CAS.
- S. K. Joshi, K. Qian, W. H. Bisson, K. Watanabe-Smith, A. Huang, D. Bottomly, E. Traer, J. W. Tyner, S. K. McWeeney, M. A. Davare, B. J. Druker and C. E. Tognon, Blood, 2020, 135, 2159–2170 CrossRef PubMed.
- T.-C. Cheong, A. Jang, Q. Wang, G. C. Leonardi, B. Ricciuti, J. V. Alessi, A. Di Federico, M. M. Awad, M. K. Lehtinen, M. H. Harris and R. Chiarle, Nat. Commun., 2024, 15, 5110 CrossRef CAS PubMed.
- E. Cocco, M. Scaltriti and A. Drilon, Nat. Rev. Clin. Oncol., 2018, 15, 731–747 CrossRef CAS PubMed.
- B. Oldrini, Á. Curiel-García, C. Marques, V. Matia, Ö. Uluçkan, O. Graña-Castro, R. Torres-Ruiz, S. Rodriguez-Perales, J. T. Huse and M. Squatrito, Nat. Commun., 2018, 9, 1466 CrossRef PubMed.
- S. Schmid, Z. R. Russell, A. S. Yamashita, M. E. West, A. G. Parrish, J. Walker, D. Rudoy, J. Z. Yan, D. C. Quist, B. N. Gessesse, N. Alvinez, K. D. Hill, L. W. Anderson, P. J. Cimino, D. K. Kumasaka, R. E. Parchment, E. C. Holland and F. Szulzewsky, Cell Rep., 2024, 43, 114829 CrossRef CAS PubMed.
- A. Vaishnavi, A. T. Le and R. C. Doebele, Cancer Discovery, 2015, 5, 25–34 CrossRef CAS PubMed.
- A. M. Schram, M. T. Chang, P. Jonsson and A. Drilon, Nat. Rev. Clin. Oncol., 2017, 14, 735–748 CrossRef CAS PubMed.
- D. Hanahan, Cancer Discovery, 2022, 12, 31–46 CrossRef CAS PubMed.
- Q. Jiang, M. Li, H. Li and L. Chen, Biomed. Pharmacother., 2022, 150, 112974 CrossRef CAS PubMed.
- E. Ardini, M. Menichincheri, P. Banfi, R. Bosotti, C. De Ponti, R. Pulci, D. Ballinari, M. Ciomei, G. Texido, A. Degrassi, N. Avanzi, N. Amboldi, M. B. Saccardo, D. Casero, P. Orsini, T. Bandiera, L. Mologni, D. Anderson, G. Wei, J. Harris, J.-M. Vernier, G. Li, E. Felder, D. Donati, A. Isacchi, E. Pesenti, P. Magnaghi and A. Galvani, Mol. Cancer Ther., 2016, 15, 628–639 CrossRef CAS PubMed.
- R. Iyer, A. E. Evans, X. Qi, R. Ho, J. E. Minturn, H. Zhao, N. Balamuth, J. M. Maris and G. M. Brodeur, Clin. Cancer Res., 2010, 16, 1478–1485 CrossRef CAS PubMed.
- S. Peters, D. R. Camidge, A. T. Shaw, S. Gadgeel, J. S. Ahn, D.-W. Kim, S.-H. I. Ou, M. Pérol, R. Dziadziuszko, R. Rosell, A. Zeaiter, E. Mitry, S. Golding, B. Balas, J. Noe, P. N. Morcos and T. Mok, N. Engl. J. Med., 2017, 377, 829–838 CrossRef CAS PubMed.
- A. F. Farago, L. P. Le, Z. Zheng, A. Muzikansky, A. Drilon, M. Patel, T. M. Bauer, S. V. Liu, S.-H. I. Ou, D. Jackman, D. B. Costa, P. S. Multani, G. G. Li, Z. Hornby, E. Chow-Maneval, D. Luo, J. E. Lim, A. J. Iafrate and A. T. Shaw, J. Thorac. Oncol., 2015, 10, 1670–1674 CrossRef CAS PubMed.
- J. Sheng, H. Chen, B. Fu, H. Pan, J. Wang and W. Han, npj Precis. Oncol., 2024, 8, 198 CrossRef CAS PubMed.
- R. C. Doebele, L. E. Davis, A. Vaishnavi, A. T. Le, A. Estrada-Bernal, S. Keysar, A. Jimeno, M. Varella-Garcia, D. L. Aisner, Y. Li, P. J. Stephens, D. Morosini, B. B. Tuch, M. Fernandes, N. Nanda and J. A. Low, Cancer Discovery, 2015, 5, 1049–1057 CrossRef CAS PubMed.
- M. Menichincheri, E. Ardini, P. Magnaghi, N. Avanzi, P. Banfi, R. Bossi, L. Buffa, G. Canevari, L. Ceriani, M. Colombo, L. Corti, D. Donati, M. Fasolini, E. Felder, C. Fiorelli, F. Fiorentini, A. Galvani, A. Isacchi, A. L. Borgia, C. Marchionni, M. Nesi, C. Orrenius, A. Panzeri, E. Pesenti, L. Rusconi, M. B. Saccardo, E. Vanotti, E. Perrone and P. Orsini, J. Med. Chem., 2016, 59, 3392–3408 CrossRef CAS PubMed.
- J. J. Carlson, A. Italiano, M. S. Brose, N. Federman, U. Lassen, S. Kummar and S. D. Sullivan, Am. J. Manage. Care, 2022, 28, S26–s32 Search PubMed.
- G. Demetri, L. Paz-Ares, A. Farago, S. Liu, S. Chawla, D. Tosi, E. Kim, C. Blakely, J. Krauss, D. Sigal, L. Bazhenova, T. John, B. Besse, J. Wolf, T. Seto, E. Chow-Maneval, P. S. Multani, A. D. Johnson, B. Simmons and R. C. Doebele, Ann. Oncol., 2018, 29, viii713 CrossRef.
- A. Drilon, T. W. Laetsch, S. Kummar, S. G. DuBois, U. N. Lassen, G. D. Demetri, M. Nathenson, R. C. Doebele, A. F. Farago, A. S. Pappo, B. Turpin, A. Dowlati, M. S. Brose, L. Mascarenhas, N. Federman, J. Berlin, W. S. El-Deiry, C. Baik, J. Deeken, V. Boni, R. Nagasubramanian, M. Taylor, E. R. Rudzinski, F. Meric-Bernstam, D. P. S. Sohal, P. C. Ma, L. E. Raez, J. F. Hechtman, R. Benayed, M. Ladanyi, B. B. Tuch, K. Ebata, S. Cruickshank, N. C. Ku, M. C. Cox, D. S. Hawkins, D. S. Hong and D. M. Hyman, N. Engl. J. Med., 2018, 378, 731–739 CrossRef CAS PubMed.
- G. Harada and A. Drilon, Cancer Genet., 2022, 264–265, 33–39 CrossRef CAS PubMed.
- A. Vaishnavi, M. T. Scherzer, C. G. Kinsey, G. L. Parkman, A. Truong, P. Ghazi, S. Schuman, B. Battistone, I. Garrido-Laguna and M. McMahon, Cell Rep., 2020, 32(5), 107994 CrossRef CAS PubMed.
- Z. Liu, P. Yu, L. Dong, W. Wang, S. Duan, B. Wang, X. Gong, L. Ye, H. Wang and J. Tian, J. Med. Chem., 2021, 64, 10286–10296 CrossRef CAS PubMed.
- A. Drilon, R. Nagasubramanian, J. F. Blake, N. Ku, B. B. Tuch, K. Ebata, S. Smith, V. Lauriault, G. R. Kolakowski, B. J. Brandhuber, P. D. Larsen, K. S. Bouhana, S. L. Winski, R. Hamor, W.-I. Wu, A. Parker, T. H. Morales, F. X. Sullivan, W. E. DeWolf, L. A. Wollenberg, P. R. Gordon, D. N. Douglas-Lindsay, M. Scaltriti, R. Benayed, S. Raj, B. Hanusch, A. M. Schram, P. Jonsson, M. F. Berger, J. F. Hechtman, B. S. Taylor, S. Andrews, S. M. Rothenberg and D. M. Hyman, Cancer Discovery, 2017, 7, 963–972 CrossRef CAS PubMed.
- A. Drilon, S. I. Ou, B. C. Cho, D. W. Kim, J. Lee, J. J. Lin, V. W. Zhu, M. J. Ahn, D. R. Camidge, J. Nguyen, D. Zhai, W. Deng, Z. Huang, E. Rogers, J. Liu, J. Whitten, J. K. Lim, S. Stopatschinskaja, D. M. Hyman, R. C. Doebele, J. J. Cui and A. T. Shaw, Cancer Discovery, 2018, 8, 1227–1236 CrossRef CAS PubMed.
- R. Kozaki, T. Yoshizawa, K. Tsukamoto, H. Kato and K. Kawabata, Cancer Res., 2016, 76, 2954A–2954A CrossRef.
- S. Xiang and X. Lu, Acta Pharm. Sin. B, 2024, 14, 517–532 CrossRef CAS PubMed.
- M. Békés, D. R. Langley and C. M. Crews, Nat. Rev. Drug Discovery, 2022, 21, 181–200 CrossRef PubMed.
- K. M. Sakamoto, K. B. Kim, A. Kumagai, F. Mercurio, C. M. Crews and R. J. Deshaies, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 8554–8559 CrossRef CAS PubMed.
- E. Koide, M. L. Mohardt, Z. M. Doctor, A. Yang, M. Hao, K. A. Donovan, C. C. Kuismi, A. J. Nelson, K. Abell, M. Aguiar, J. Che, M. P. Stokes, T. Zhang, A. J. Aguirre, E. S. Fischer, N. S. Gray, B. Jiang and B. Nabet, ChemBioChem, 2023, 24, e202300141 CrossRef CAS PubMed.
- K. A. Donovan, F. M. Ferguson, J. W. Bushman, N. A. Eleuteri, D. Bhunia, S. Ryu, L. Tan, K. Shi, H. Yue, X. Liu, D. Dobrovolsky, B. Jiang, J. Wang, M. Hao, I. You, M. Teng, Y. Liang, J. Hatcher, Z. Li, T. D. Manz, B. Groendyke, W. Hu, Y. Nam, S. Sengupta, H. Cho, I. Shin, M. P. Agius, I. M. Ghobrial, M. W. Ma, J. Che, S. J. Buhrlage, T. Sim, N. S. Gray and E. S. Fischer, Cell, 2020, 183, 1714–1731.e1710 CrossRef CAS PubMed.
- G. M. Burslem, B. E. Smith, A. C. Lai, S. Jaime-Figueroa, D. C. McQuaid, D. P. Bondeson, M. Toure, H. Dong, Y. Qian, J. Wang, A. P. Crew, J. Hines and C. M. Crews, Cell Chem. Biol., 2018, 25, 67–77.e63 CrossRef CAS PubMed.
- S. O. Alhassan, Z. Y. Abd Elmageed, Y. Errami, G. Wang, J. A. Abi-Rached, E. Kandil and M. Zerfaoui, Clin. Transl. Med., 2025, 15, e70251 CrossRef CAS PubMed.
- L. Chen, Y. Chen, C. Zhang, B. Jiao, S. Liang, Q. Tan, H. Chai, W. Yu, Y. Qian, H. Yang, W. Yao, J. Yu, Y. Luo, M. Plewe, J. Wang, H. Xiao-Ran and J. Liu, J. Med. Chem., 2020, 63, 14562–14575 CrossRef CAS PubMed.
- R. C. Doebele, A. Drilon, L. Paz-Ares, S. Siena, A. T. Shaw, A. F. Farago, C. M. Blakely, T. Seto, B. C. Cho, D. Tosi, B. Besse, S. P. Chawla, L. Bazhenova, J. C. Krauss, Y. K. Chae, M. Barve, I. Garrido-Laguna, S. V. Liu, P. Conkling, T. John, M. Fakih, D. Sigal, H. H. Loong, G. L. Buchschacher Jr., P. Garrido, J. Nieva, C. Steuer, T. R. Overbeck, D. W. Bowles, E. Fox, T. Riehl, E. Chow-Maneval, B. Simmons, N. Cui, A. Johnson, S. Eng, T. R. Wilson and G. D. Demetri, Lancet Oncol., 2020, 21, 271–282 CrossRef CAS PubMed.
- T. M. Nguyen, V. Sreekanth, A. Deb, P. Kokkonda, P. K. Tiwari, K. A. Donovan, V. Shoba, S. K. Chaudhary, J. A. M. Mercer, S. Lai, A. Sadagopan, M. Jan, E. S. Fischer, D. R. Liu, B. L. Ebert and A. Choudhary, Nat. Chem., 2024, 16, 218–228 CrossRef CAS PubMed.
- D. Martin-Zanca, S. H. Hughes and M. Barbacid, Nature, 1986, 319, 743–748 CrossRef CAS PubMed.
- A. Vaishnavi, M. Capelletti, A. T. Le, S. Kako, M. Butaney, D. Ercan, S. Mahale, K. D. Davies, D. L. Aisner, A. B. Pilling, E. M. Berge, J. Kim, H. Sasaki, S. Park, G. Kryukov, L. A. Garraway, P. S. Hammerman, J. Haas, S. W. Andrews, D. Lipson, P. J. Stephens, V. A. Miller, M. Varella-Garcia, P. A. Jänne and R. C. Doebele, Nat. Med., 2013, 19, 1469–1472 CrossRef CAS PubMed.
- R. Arafeh, T. Shibue, J. M. Dempster, W. C. Hahn and F. Vazquez, Nat. Rev. Cancer, 2025, 25, 59–73 CrossRef CAS PubMed.
- R. Gillani, B. K. A. Seong, J. Crowdis, J. R. Conway, N. V. Dharia, S. Alimohamed, B. J. Haas, K. Han, J. Park, F. Dietlein, M. X. He, A. Imamovic, C. Ma, M. C. Bassik, J. S. Boehm, F. Vazquez, A. Gusev, D. Liu, K. A. Janeway, J. M. McFarland, K. Stegmaier and E. M. Van Allen, Cancer Res., 2021, 81, 3971–3984 CrossRef CAS PubMed.
- T. Zhu, J. Xie, H. He, H. Li, X. Tang, S. Wang, Z. Li, Y. Tian, L. Li, J. Zhu and G. Zhu, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2219589120 CrossRef CAS PubMed.
- M. L. Drummond, A. Henry, H. Li and C. I. Williams, J. Chem. Inf. Model., 2020, 60, 5234–5254 CrossRef CAS PubMed.
- B. Jiang, Y. Gao, J. Che, W. Lu, I. H. Kaltheuner, R. Dries, M. Kalocsay, M. J. Berberich, J. Jiang, I. You, N. Kwiatkowski, K. M. Riching, D. L. Daniels, P. K. Sorger, M. Geyer, T. Zhang and N. S. Gray, Nat. Chem. Biol., 2021, 17, 675–683 CrossRef CAS PubMed.
- V. G. Klein, C. E. Townsend, A. Testa, M. Zengerle, C. Maniaci, S. J. Hughes, K.-H. Chan, A. Ciulli and R. S. Lokey, ACS Med. Chem. Lett., 2020, 11, 1732–1738 CrossRef CAS PubMed.
- G. E. Winter, A. Mayer, D. L. Buckley, M. A. Erb, J. E. Roderick, S. Vittori, J. M. Reyes, J. di Iulio, A. Souza, C. J. Ott, J. M. Roberts, R. Zeid, T. G. Scott, J. Paulk, K. Lachance, C. M. Olson, S. Dastjerdi, S. Bauer, C. Y. Lin, N. S. Gray, M. A. Kelliher, L. S. Churchman and J. E. Bradner, Mol. Cell, 2017, 67, 5–18.e19 CrossRef CAS PubMed.
- Y. Takumi, S. Arai, C. Suzuki, K. Fukuda, A. Nishiyama, S. Takeuchi, H. Sato, K. Matsumoto, K. Sugio and S. Yano, Cancer Med, 2023, 12, 5809–5820 CrossRef CAS PubMed.
- A. Drilon, Ann. Oncol., 2019, 30, viii23–viii30 CrossRef CAS PubMed.
- E. Cocco, A. M. Schram, A. Kulick, S. Misale, H. H. Won, R. Yaeger, P. Razavi, R. Ptashkin, J. F. Hechtman, E. Toska, J. Cownie, R. Somwar, S. Shifman, M. Mattar, S. D. Selçuklu, A. Samoila, S. Guzman, B. B. Tuch, K. Ebata, E. de Stanchina, R. J. Nagy, R. B. Lanman, B. Houck-Loomis, J. A. Patel, M. F. Berger, M. Ladanyi, D. M. Hyman, A. Drilon and M. Scaltriti, Nat. Med., 2019, 25, 1422–1427 CrossRef CAS PubMed.
- J. J. Lin, V. W. Zhu, S. Yoda, B. Y. Yeap, A. B. Schrock, I. Dagogo-Jack, N. A. Jessop, G. Y. Jiang, L. P. Le, K. Gowen, P. J. Stephens, J. S. Ross, S. M. Ali, V. A. Miller, M. L. Johnson, C. M. Lovly, A. N. Hata, J. F. Gainor, A. J. Iafrate, A. T. Shaw and S. I. Ou, J. Clin. Oncol., 2018, 36, 1199–1206 CrossRef CAS PubMed.
- M. Ishoey, S. Chorn, N. Singh, M. G. Jaeger, M. Brand, J. Paulk, S. Bauer, M. A. Erb, K. Parapatics, A. C. Müller, K. L. Bennett, G. F. Ecker, J. E. Bradner and G. E. Winter, ACS Chem. Biol., 2018, 13, 553–560 CrossRef CAS PubMed.
- V. Vetma, L. C. Perez, J. Eliaš, A. Stingu, A. Kombara, T. Gmaschitz, N. Braun, T. Ciftci, G. Dahmann, E. Diers, T. Gerstberger, P. Greb, G. Kidd, C. Kofink, I. Puoti, V. Spiteri, N. Trainor, H. Weinstabl, Y. Westermaier, C. Whitworth, A. Ciulli, W. Farnaby, K. McAulay, A. B. Frost, N. Chessum and M. Koegl, ACS Chem. Biol., 2024, 19, 1484–1494 CrossRef CAS PubMed.
- Q. Lin, W. Liu, W. Lu, M. Zhao, L. Cao, Z. Li, J. Wang, X. Xu and H. Wu, Adv. Sci., 2025, 12, e11789 CrossRef CAS PubMed.
- C. E. Powell, Y. Gao, L. Tan, K. A. Donovan, R. P. Nowak, A. Loehr, M. Bahcall, E. S. Fischer, P. A. Jänne, R. E. George and N. S. Gray, J. Med. Chem., 2018, 61, 4249–4255 CrossRef CAS PubMed.
- J. Popow, W. Farnaby, A. Gollner, C. Kofink, G. Fischer, M. Wurm, D. Zollman, A. Wijaya, N. Mischerikow, C. Hasenoehrl, P. Prokofeva, H. Arnhof, S. Arce-Solano, S. Bell, G. Boeck, E. Diers, A. B. Frost, J. Goodwin-Tindall, J. Karolyi-Oezguer, S. Khan, T. Klawatsch, M. Koegl, R. Kousek, B. Kratochvil, K. Kropatsch, A. A. Lauber, R. McLennan, S. Olt, D. Peter, O. Petermann, V. Roessler, P. Stolt-Bergner, P. Strack, E. Strauss, N. Trainor, V. Vetma, C. Whitworth, S. Zhong, J. Quant, H. Weinstabl, B. Kuster, P. Ettmayer and A. Ciulli, Science, 2024, 385, 1338–1347 CrossRef CAS PubMed.
- L. Mascarenhas, S. G. DuBois, C. M. Albert, S. Bielack, D. Orbach, N. Federman, B. Geoerger, R. Nagasubramanian, Y. Zhang, J. Chisholm, S. G. Melcon, H. Goto, D. A. Morgenstern, C. Owens, A. S. Pappo, S. Perreault, J. H. Schulte, N. Shukla, C. M. Zwaan, N. Neu, V. Bernard-Gauthier, E. D. L. Cuesta, C. M. V. Tilburg and T. W. Laetsch, J. Clin. Oncol., 2025, 43, 1180–1187 CrossRef CAS PubMed.
- S. Alabi, S. Jaime-Figueroa, Z. Yao, Y. Gao, J. Hines, K. T. G. Samarasinghe, L. Vogt, N. Rosen and C. M. Crews, Nat. Commun., 2021, 12, 920 CrossRef CAS PubMed.
- A. D. Buhimschi, H. A. Armstrong, M. Toure, S. Jaime-Figueroa, T. L. Chen, A. M. Lehman, J. A. Woyach, A. J. Johnson, J. C. Byrd and C. M. Crews, Biochemistry, 2018, 57, 3564–3575 CrossRef CAS PubMed.
- G. Posternak, X. Tang, P. Maisonneuve, T. Jin, H. Lavoie, S. Daou, S. Orlicky, T. Goullet de Rugy, L. Caldwell, K. Chan, A. Aman, M. Prakesch, G. Poda, P. Mader, C. Wong, S. Maier, J. Kitaygorodsky, B. Larsen, K. Colwill, Z. Yin, D. F. Ceccarelli, R. A. Batey, M. Taipale, I. Kurinov, D. Uehling, J. Wrana, D. Durocher, A.-C. Gingras, R. Al-Awar, M. Therrien and F. Sicheri, Nat. Chem. Biol., 2020, 16, 1170–1178 CrossRef CAS PubMed.
- S. Montoya, J. Bourcier, M. Noviski, H. Lu, M. C. Thompson, A. Chirino, J. Jahn, A. K. Sondhi, S. Gajewski, Y. S. Tan, S. Yung, A. Urban, E. Wang, C. Han, X. Mi, W. J. Kim, Q. Sievers, P. Auger, H. Bousquet, N. Brathaban, B. Bravo, M. Gessner, C. Guiducci, J. N. Iuliano, T. Kane, R. Mukerji, P. J. Reddy, J. Powers, M. Sanchez Garcia de los Rios, J. Ye, C. Barrientos Risso, D. Tsai, G. Pardo, R. Q. Notti, A. Pardo, M. Affer, V. Nawaratne, T. M. Totiger, C. Pena-Velasquez, J. M. Rhodes, A. D. Zelenetz, A. Alencar, L. E. Roeker, S. Mehta, R. Garippa, A. Linley, R. K. Soni, S. S. Skånland, R. J. Brown, A. R. Mato, G. M. Hansen, O. Abdel-Wahab and J. Taylor, Science, 2024, 383, eadi5798 CrossRef CAS PubMed.
- A. M. Kurimchak, C. Herrera-Montávez, S. Montserrat-Sangrà, D. Araiza-Olivera, J. Hu, R. Neumann-Domer, M. Kuruvilla, A. Bellacosa, J. R. Testa, J. Jin and J. S. Duncan, Sci. Signaling, 2022, 15, eabn2707 CrossRef CAS PubMed.
- A. Liu, M. A. Mohr, J. M. Hope, J. Wang, X. Chen and B. Cui, ACS Chem. Biol., 2024, 19, 1626–1637 CrossRef CAS PubMed.
- R. Romero, V. I. Sayin, S. M. Davidson, M. R. Bauer, S. X. Singh, S. E. LeBoeuf, T. R. Karakousi, D. C. Ellis, A. Bhutkar, F. J. Sánchez-Rivera, L. Subbaraj, B. Martinez, R. T. Bronson, J. R. Prigge, E. E. Schmidt, C. J. Thomas, C. Goparaju, A. Davies, I. Dolgalev, A. Heguy, V. Allaj, J. T. Poirier, A. L. Moreira, C. M. Rudin, H. I. Pass, M. G. Vander Heiden, T. Jacks and T. Papagiannakopoulos, Nat. Med., 2017, 23, 1362–1368 CrossRef CAS PubMed.
- Y. Perez-Riverol, C. Bandla, D. J. Kundu, S. Kamatchinathan, J. Bai, S. Hewapathirana, N. S. John, A. Prakash, M. Walzer, S. Wang and J. A. Vizcaíno, Nucleic Acids Res., 2025, 53, D543–d553 CrossRef CAS PubMed.
- Y. Perez-Riverol, J. Bai, C. Bandla, D. García-Seisdedos, S. Hewapathirana, S. Kamatchinathan, D. J. Kundu, A. Prakash, A. Frericks-Zipper, M. Eisenacher, M. Walzer, S. Wang, A. Brazma and J. A. Vizcaíno, Nucleic Acids Res., 2022, 50, D543–d552 CrossRef CAS PubMed.
- J. Pletz, B. Berg and R. Breinbauer, Synthesis, 2016, 1301–1317 CAS.
- D. Remillard, D. L. Buckley, J. Paulk, G. L. Brien, M. Sonnett, H. S. Seo, S. Dastjerdi, M. Wühr, S. Dhe-Paganon, S. A. Armstrong and J. E. Bradner, Angew. Chem., Int. Ed., 2017, 56, 5738–5743 CrossRef CAS PubMed.
- T. Ozawa, M. Riester, Y. K. Cheng, J. T. Huse, M. Squatrito, K. Helmy, N. Charles, F. Michor and E. C. Holland, Cancer Cell, 2014, 26, 288–300 CrossRef CAS PubMed.
- D. Li, K. Geng, Y. Hao, J. Gu, S. Kumar, A. T. Olson, C. C. Kuismi, H. M. Kim, Y. Pan, F. Sherman, A. M. Williams, Y. Li, F. Li, T. Chen, C. Thakurdin, M. Ranieri, M. Meynardie, D. S. Levin, J. Stephens, A. Chafitz, J. Chen, M. S. Donald-Paladino, J. M. Powell, Z.-Y. Zhang, W. Chen, M. Ploszaj, H. Han, S. S. Gu, T. Zhang, B. Hu, B. A. Nacev, M. E. Kaiza, A. H. Berger, X. Wang, J. Li, X. Sun, Y. Liu, X. Zhang, T. C. Bruno, N. S. Gray, B. Nabet, K.-K. Wong and H. Zhang, J. Clin. Invest., 2024, 135, e174249 CrossRef PubMed.
- C. S. Hughes, S. Moggridge, T. Müller, P. H. Sorensen, G. B. Morin and J. Krijgsveld, Nat. Protoc., 2019, 14, 68–85 CrossRef CAS PubMed.
- V. Demichev, C. B. Messner, S. I. Vernardis, K. S. Lilley and M. Ralser, Nat. Methods, 2020, 17, 41–44 CrossRef CAS PubMed.
- P. Di Tommaso, M. Chatzou, E. W. Floden, P. P. Barja, E. Palumbo and C. Notredame, Nat. Biotechnol., 2017, 35, 316–319 CrossRef CAS PubMed.
- C. Dai, J. Pfeuffer, H. Wang, P. Zheng, L. Käll, T. Sachsenberg, V. Demichev, M. Bai, O. Kohlbacher and Y. Perez-Riverol, Nat. Methods, 2024, 21, 1603–1607 CrossRef CAS PubMed.
Footnote |
| † SK and JJ contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |