Open Access Article
Open Access ArticleIntricacies in iron–sulfur cluster function and biogenesis: functional versatility, sulfur sources, and enzyme specificity
Sarah M.
Spigelmyer
and
Patricia C.
Dos Santos
 *
*
Department of Chemistry, Wake Forest University, Winston-Salem, NC 27106, USA. E-mail: dossanpc@wfu.edu
First published on 2nd April 2026
Abstract
Iron–sulfur (Fe–S) clusters are ancient inorganic cofactors ubiquitous across all domains of life. These cofactors associate with proteins through constitutive or transient coordination, expanding their chemistries and versatility in biological processes. Thus, Fe–S proteins participate in intricate and multifaceted chemistries critical to life on Earth. The biosynthesis of these cofactors has evolved to require complex machinery to catalyze cluster formation and subsequent transfer to target apo-proteins. Five Fe–S cluster biogenesis systems have been identified in prokaryotes, with varying degrees of complexity, including: iron–sulfur cluster (ISC), nitrogen fixation (NIF), sulfur mobilization (SUF), minimal iron–sulfur system (MIS), and SUF-like minimal system (SMS). Sulfur mobilization in the biosynthesis of Fe–S clusters is initiated, in most cases, by cysteine sulfurtransferases, also known as cysteine desulfurases. These enzymes use the amino acid cysteine as a sulfur source and require specific interactions with a sulfur acceptor to promote sulfur transfer. Physical interactions and coordination among biosynthetic components restrict their functions and guarantee the trafficking of reactive intermediates to proper destinations. As recently reported, the occurrence of alternate biosynthetic schemes using sulfide as the sulfur source bypasses the requirement for sulfurtransferases and provides alternate evolutionary strategies to construct Fe–S clusters.
1. Introduction to Fe–S clusters and their functions in biological systems
Iron–sulfur (Fe–S) clusters are ancient inorganic compounds that are prevalent in all forms of life. These clusters were originally recognized for their roles in electron transport, which are essential for processes such as cellular respiration.1 However, they are more versatile in biological systems, as they partake in critical roles in nitrogen fixation, radical biochemistry, genome maintenance, and photosynthesis.2 As a result, the presence of Fe–S clusters is fundamentally associated with life on Earth. Fe–S clusters can present themselves in multiple configurations and oxidation states. The most common of these configurations include planar [2Fe–2S], cuboidal [3Fe–4S], and cubic [4Fe–4S] (Fig. 1). The ability of these clusters to adopt different oxidation states enables efficient electron transport, as these cofactors can participate in single- or sequential multiple-electron transfer events.3–5 Fe–S clusters are mainly coordinated by cysteine thiol ligands; however, alternate coordination has also been observed, including histidine, aspartate, arginine, threonine, glutamate, serine, tyrosine, and methionine, as well as non-protein ligands including glutathione and water.6–8 Along with cluster ligands, the protein environment surrounding these cofactors modulates their electron potentials and reactivities, thereby expanding the functions of their associated proteins. Despite their diverse functionalities, Fe–S clusters can be classified into two distinct groups: constitutive clusters, considered integral components of Fe–S proteins, and transient clusters, which undergo transformation as an intrinsic feature of their function (Fig. 1).1.1 Constitutive Fe–S clusters
Constitutive Fe–S clusters have long been studied in their role in catalysis and electron transport. Fe–S clusters of this type have well-characterized roles in single-electron transfer reactions. In this capacity, efficient electron transfer results in redox cycling of the cluster, either localized to one Fe atom by cycling from ferric iron (Fe3+) to ferrous iron (Fe2+) or in a delocalized fashion where additional electrons are not residing solely on a single Fe of the cluster.8 Sequential steps of electron transfer in the electron transport chain (ETC) provide an excellent example of how the unique redox properties of metallocenters drive electron transfer across multiple sites.9 Mitochondrial complex I and bacterial NADH dehydrogenase, for instance, can have as many as nine distinct Fe–S clusters positioned across different subunits of this complex with differing nuclearity, ligands, and protein environment, promoting long-distance electron tunnelling.10–13 Complex II employs three types of cofactors, including [2Fe–2S], [3Fe–4S], and [4Fe–4S] clusters to mediate electron transfer from the flavin cofactor to ubiquinone (UQ) during aerobic metabolism, while having the capacity to operate in the reverse direction under anaerobic conditions.14,15 The single [2Fe–2S] cluster in complex III is a Rieske type with two His and two Cys ligands.16 This configuration enables bifurcation of electron transfer from UQ to the high-redox-potential Rieske cluster and the low-redox-potential cytochrome heme.9Constitutive Fe–S clusters display expanded chemical functionalities assisting chemical transformations, promoting Lewis acid catalysis, initiating radical-induced chemistry, and being directly involved in substrate binding and activation.17–19 Additionally, these Fe–S clusters have been reported to play a structural role, impacting the folding and stability of their associated partners, as in the cases of DNA primase,20 helicase,21 and helicase-nuclease enzymes.22
Diversification of the Fe–S clusters and expansion of the chemistries promoted by these cofactors is also observed through the incorporation of heterometals. Well-studied examples include nickel in NiFe hydrogenases and carbon monoxide dehydrogenases, molybdenum in Mo-dependent nitrogenase, and vanadium in V-dependent nitrogenase.23–25 These heterometallic Fe–S cofactors are involved in substrate binding and redox chemistry at the active site of their associated enzymes, and their synthesis and enzyme activation have been intensively explored in the literature.
Enzymes coordinating [4Fe–4S] clusters with three protein ligands provide a labile Fe site within the cluster, which serves as a site for substrate-activation and catalysis. Fe–S dehydratases, such as aconitase, nicotinic acid synthase, and dihydroxyacid dehydratases, utilize a [4Fe–4S] cluster coordinated to three cysteines, leaving an uncoordinated Fe site to serve as a Lewis acid.7 Enzymes displaying a three-Cys coordinated cluster have also been described as participants in non-redox thiolation reactions, including TtcA, TtuI, TtuA, NcsA, and C-type MnmA.26 These proteins are involved in post-transcriptional thio-modification of tRNA, catalyzing the substitution of an oxygen atom with a sulfur atom, to generate thionucleosides. They belong to the PP-loop ATP pyrophosphatase family with a characteristic SGGKDS motif, involved in ATP binding and tRNA adenylation, adding a suitable leaving group for insertion of sulfur to the correct position, while the Fe–S cluster is proposed to engage in sulfur insertion.27,28 In TtuA, for example, the cluster serves as a Lewis acid, forming a transient coordination to a fifth sulfur, resulting in a [4Fe–5S] cluster reaction intermediate.29,30 TtuA can undergo multiple turnovers while retaining its [4Fe–4S] cluster. However, other examples exhibit rapid cluster degradation,7,28 making it challenging to mechanistically assign their reaction paths and associated intermediates.
More recently, additional tRNA thiolating enzymes have been identified that coordinate Fe–S clusters and play roles similar to those of Lewis acid catalysts.26 The presence of Fe–S clusters associated with these enzymes was previously overlooked, and their associated reactions were initially thought to be performed by a persulfide attached to a catalytic cysteine. Unexpectedly, the E. coli D-type MnmA is capable of coordinating an Fe–S cluster, with an unusual (2Cys, 1Asp) coordination motif that departs from the previously studied Lewis-acid cluster catalysts with three Cys ligands.31 D-type MnmA is present in many bacterial species and is essential in some organisms, such as the model Gram-positive bacterium B. subtilis.32 Although the capacity to coordinate an Fe–S cluster has been demonstrated by several groups,33,34 their clusters are highly labile, making it difficult to investigate their biochemical functions. Nevertheless, the identification of new sequence motifs capable of coordinating Fe–S clusters hints at a potential for an even larger Fe–S proteome with broader roles in metabolism.
The diverse chemistries promoted by Fe–S proteins are best exemplified by the radical SAM enzyme superfamily.19,35 Enzymes of this group operate through activation of S-adenosylmethionine (SAM) by its characteristic [4Fe–4S] cluster in the reduced state (Fig. 1). This reaction generates a 5′-deoxyadenosyl radical, which, in most cases, promotes substrate activation via hydrogen-atom abstraction.36 The reactions catalyzed by these enzymes exhibit a mechanistic complexity that is difficult to predict from their protein sequences. The radical SAM enzyme superfamily comprises more than 700![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 unique sequences, and a few of its characterized members exemplify diverse chemical transformations.37 A few examples include S–C and C–C bond formation, formylation, dehydration, and complex rearrangements. The inventory of reactions promoted by these enzymes continues to expand rapidly, making the synthesis and functions of Fe–S clusters a relevant aspect of metabolism, from their emergence and evolution in the early history of life on Earth to their application in the design of novel catalysts.
000 unique sequences, and a few of its characterized members exemplify diverse chemical transformations.37 A few examples include S–C and C–C bond formation, formylation, dehydration, and complex rearrangements. The inventory of reactions promoted by these enzymes continues to expand rapidly, making the synthesis and functions of Fe–S clusters a relevant aspect of metabolism, from their emergence and evolution in the early history of life on Earth to their application in the design of novel catalysts.
1.2 Transient clusters
Transient cluster are defined as cofactors that are either consumed in reactions performed by their associated enzymes or undergo transformation or degradation as a signalling mechanism (Fig. 1). Exposure to oxygen can lead to cluster degradation, which in turn can alter protein stability and activity. Thus, the intrinsic labile nature of Fe–S clusters enables sensing of environmental changes.38,39 Several transcriptional factors utilize Fe–S as a signalling molecule to modulate transcription in response to changes in the status of Fe–S clusters (IscR and SufR), reactive nitrogen species (NsrR, SoxR), iron (RirA), and oxygen (FNR).40–46 These conditions elicit modifications or degradation of the cluster, resulting in conformational changes that impact DNA-binding affinities of these regulatory proteins. Another example of Fe–S proteins reporting cellular status is cytosolic aconitase.47 When iron levels are low, cytosolic aconitase loses its Fe–S cluster, triggering a signalling response to promote the incorporation of iron.48 Thus, the presence of Fe–S in these regulatory capacities is viewed as transient, as environmental factors can react with the cluster to elicit a biological response.Fe–S clusters that undergo self-destruction during a reaction cycle serve a sacrificial role in supplying Fe or S. In reported species of archaea, specialized proteins serve as storage entities for Fe and S. This proposed function was coined nearly 30 years ago by Beinert.49 Recent support for this role has been provided by the discovery of IssA, a protein that forms polymeric nanoparticles capable of coordinating linear FeS structures, denoted as thioferrates (Fig. 1). In Pyrococcus furiosus, these Fe–S units can be mobilized for the maturation of Fe–S proteins, such as ferredoxin.50,51 IssA orthologs are found in several archaeal species, indicating that utilization of transient Fe–S cluster-like structures as a storage and source of Fe, S, and perhaps Fe–S clusters can be spread across domains of life.52
Additional enzymes that promote thiolation reactions utilize their own Fe–S clusters as a source of sulfur in a sacrificial manner. Operating in this capacity, biotin synthase (BioB) and lipoyl synthase (LipA) coordinate two types of clusters: a constitutive cluster that promotes radical SAM chemistry and a transient cluster that serves as the sulfur source.53,54 In the case of lipoyl synthase, the reaction causes cluster destruction, and its reconstruction involves a partnership with the cluster carrier protein NfuA, leading to enzyme reactivation.55 Enzymes promoting methylthiolation, like MiaB, MtbA, and RimO, also contain two types of clusters: a constitutive (radical SAM) and a transient (auxiliary). Efforts to elucidate the purpose of the auxiliary cluster in this class of enzymes suggested that the auxiliary cluster's role was to aid in binding, coordination, and activation of exogenous sulfur, proposed to be hydrogen sulfide from endogenous polysulfides and persulfides, to the unclaimed coordination site.56–58 However, biochemical and structural analyses of these enzymes also provide evidence that a transient cluster is to be modified by methylation in the initial step of the reaction, and that it is later used for thiomethylation and the formation of a proposed damaged [3Fe–3S]1+ cluster.53,59 It is possible that different members of this enzyme family have evolved distinct roles for their auxiliary clusters, based on the protein environment surrounding each cluster and its location relative to the radical SAM cluster. To support these distinct functions, a new type of BioB has recently been reported that does not use a transient cluster but instead a [4Fe–5S] cluster intermediate to catalyze sulfur insertion in the final step of biotin synthesis.60
Lastly, transient clusters are also observed in cofactors that undergo transformation as a step in their catalytic cycle. One example is the molybdenum chaperone NifQ, which facilitates molybdenum delivery during the biosynthesis of the nitrogenase FeMo cofactor. Maturation of its cofactor is proposed to involve transfer of a [4Fe–4S] cluster from NifU directly to NifQ.61 Isolation of NifQ from its native organism, Azotobacter vinelandii, results in a mixture of species containing [Mo–3Fe–4S] and [3Fe–4S] clusters. Although the process is not fully understood, the active form of NifQ capable of participating in FeMo cofactor synthesis is a [Mo–3Fe–4S] cluster. Transfer of the Mo from this heterometallic cluster is only achieved in the presence of NifEN and NifH.62 Presumably, NifQ cycles through coordination of transient clusters [4Fe–4S] to [Mo–3Fe–4S] and [3Fe–4S] (Fig. 1).
Fe–S clusters, adopting permanent or transient roles as cofactors, are found in many aspects of metabolism. The few examples described in Fig. 1 illustrate their diverse functionalities and mechanisms for participating in biochemical processes. Thus, the assembly of these inorganic entities in biological systems is an area of investigation as interesting as the chemistries they promote. Biochemical pathways involving the assembly of Fe–S clusters are tightly regulated and essential for supplying reactive Fe and S sources, as well as for transporting reaction intermediates susceptible to damage by oxygen and other reactive species commonly found in cellular environments.
2. Formation of Fe–S clusters in biological systems
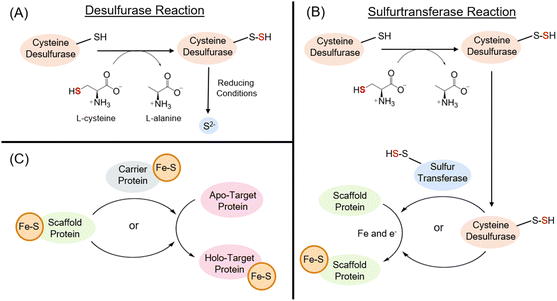
The biosynthesis of Fe–S clusters is not a spontaneous process and requires complex protein machineries. This biosynthetic process can be broken down into three general steps: mobilization of sulfur, cluster assembly on a scaffold protein, and cluster transfer to a recipient target. In most species, sulfur mobilization is characterized by the sulfur-activation step, catalyzed by cysteine sulfurtransferase, also known as cysteine desulfurase.63In vitro, the activity of these enzymes is typically assessed by measuring desulfurization, which yields sulfide, and this activity provides the rationale for their historical designation as desulfurases (Fig. 2A).64,65In vivo, these enzymes transfer sulfur, directly or indirectly, to scaffold proteins for the assembly of Fe–S clusters.66 From this biological functional perspective, these enzymes are classified as L-cysteine:acceptor sulfurtransferases (EC 2.8.1.7) (Fig. 2B). The scaffold protein requires an iron source and an electron input during cluster assembly to form S0 to S2−.66 The final main step is the transfer of pre-formed clusters to recipient targets, a process that involves Fe–S carriers (Fig. 2C). While individual pathway components are distinct, the general principles of the biological synthesis of Fe–S clusters are synonymous and include a sulfur donor, an iron donor, an electron donor, and the construction of the cluster onto a protein. The differences among these systems are the manner in which sulfur is mobilized through these pathways, the type of sulfur donor, and the presence or absence of a carrier protein.Several systems have been employed to generate Fe–S clusters in nature. To date, five systems have been described for the assembly of Fe–S cluster proteins in prokaryotes: iron–sulfur cluster (ISC), sulfur mobilization (SUF), nitrogen fixation (NIF), minimal iron–sulfur system (MIS), and SUF-like minimal system (SMS) (Fig. 3).67 No sulfurtransferase has yet been identified for the SMS system, raising questions about the source of sulfur in species where such cysteine desulfurases are absent and about how this simple system functions. Differences in biosynthetic components that perform equivalent functions, such as IscS, SufS, and NifS, and, in some cases, the absence of key components, such as a cysteine desulfurase or a carrier protein, raise questions about the mechanistic details of these pathways and the evolutionary factors underlying these differences. Analysis of over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 prokaryotic species shows that most genomes encode a single Fe–S cluster biogenesis system; however, some employ two or even three systems to meet the demand for Fe–S clusters under specific environmental and nutritional conditions.67 For instance, the SUF system shows a dominant presence across more than 6000 genomes analyzed, and of those, only 5% contain copies of the ISC or NIF systems. Eukaryotic pathways, although not covered in this review, have been recently described in the literature.68–71 While orthologues of ISC, SUF, and NIF enzymes are found in eukaryotes, these systems often require additional components, and individual steps may be confined to specific organelles. Nevertheless, despite differences, they share the general paradigm that Fe–S cluster synthesis in biological systems is a protein-assisted process in which reactive intermediates are protein-bound, thereby protecting them from damage and inadvertent degradation.
000 prokaryotic species shows that most genomes encode a single Fe–S cluster biogenesis system; however, some employ two or even three systems to meet the demand for Fe–S clusters under specific environmental and nutritional conditions.67 For instance, the SUF system shows a dominant presence across more than 6000 genomes analyzed, and of those, only 5% contain copies of the ISC or NIF systems. Eukaryotic pathways, although not covered in this review, have been recently described in the literature.68–71 While orthologues of ISC, SUF, and NIF enzymes are found in eukaryotes, these systems often require additional components, and individual steps may be confined to specific organelles. Nevertheless, despite differences, they share the general paradigm that Fe–S cluster synthesis in biological systems is a protein-assisted process in which reactive intermediates are protein-bound, thereby protecting them from damage and inadvertent degradation.
2.1 Synthesis of Fe–S clusters by the ISC system
The ISC system is the most well-studied Fe–S cluster biogenesis system and is initiated by the cysteine sulfurtransferase enzyme IscS. This enzyme uses pyridoxal-5′-phosphate to bind the substrate amino acid L-cysteine to enable cleavage of the C–S bond via the formation of an enzyme persulfide intermediate (IscS-SH).72 A strictly conserved Cys residue located in a flexible loop is involved in persulfide formation and transfer to the acceptor protein IscU. During catalysis, this Cys is proposed to swing into the active site for nucleophilic attack onto the Cys substrate thiol bound to the PLP cofactor. Following persulfide formation, the catalytic Cys residue moves away to interact with the S-acceptor protein. The characteristic loop structure is a defining element of class I cysteine sulfurtransferase, which was believed to be an element enabling facile transfer of the sulfur to the acceptor protein IscU for the synthesis of Fe–S clusters as well as other sulfur acceptors, including TusA in the synthesis of s2U tRNA and Mo-cofactor, ThiI in the synthesis of s4U and thiamine, FhdD in formate dehydrogenase maturation, and RhdA in cysteine redox regulation.73–76 The versatility of IscS in interacting with multiple acceptors was attributed to the flexibility of its active site Cys loop. However, structural and biochemical analyses of protein complexes with IscS revealed distinct interaction sites with S-acceptor partners, indicating the presence of specific sequence elements within the enzyme and its partners that serve as recognition points for sulfur transfer.73,77In Fe–S cluster synthesis, IscS interacts with IscU during sulfur transfer. The mechanistic steps leading to the synthesis of the [2Fe–2S] cluster by the E. coli ISC system have been explored extensively in vitro. IscU can bind zinc with high affinity (Kd 10−13 M), stabilizing a structured conformation of the protein.78 In fact, zinc overload affects Fe–S metabolism.79 Zinc loss leads to a disordered conformation and is promoted by IscS during sulfur transfer.80–82 Mechanistic studies have championed models for the early stages of the cluster assembly by aiming to define the order of binding events, as the “iron-first” or “sulfur-first” models. However, the interplay between zinc binding to IscU, its effect on IscU structure, IscS complex formation, and impact on Fe and S transfer adds layers of complexity to dissecting these pathways through the lens of physiologically relevant conditions.
In vitro experiments have shown that either Fe2+- or S0-bound IscU can initiate cluster assembly, depending on the reaction conditions. Experimental evidence supports either path, since persulfides can be observed on IscU in the absence of iron, and an Fe2+-bound IscU can also initiate cluster assembly.80 It is still not clear if Zn-bound IscU is a physiologically relevant form of the scaffold, since zinc is found at millimolar intracellular concentrations while cytoplasmic concentration of its free form is significantly lower (10−16 M).83 Zn-bound IscU is not capable of binding iron, as both metals are known to bind to the same site, but is capable of sulfur transfer, providing support for a sulfur-first mechanism.80,84 However, in vitro experiments probing the formation of Fe–S clusters starting from an apo-IscU are significantly more effective.80,84,85 Thus, mechanistic models supporting an Fe-first model have considered an apo-form of IscU as the starting point for cluster assembly.86
Zn-free IscU also binds iron with significant affinity (Kd of 10−6 M),80 enabling a concerted formation of a [2Fe–2S] cluster. The exact sequence of events in vivo is not known because the metalation state of IscU and the identity of the Fe donor remain elusive. In vitro, detection of reaction intermediates varies with reaction conditions. In cluster assembly reactions using an excess of iron, the formation of a [2Fe–2S] cluster is proposed to occur directly within a monomer.80 However, in experiments using a stoichiometric Fe-to-IscU ratio, the [1Fe–1S]–IscU intermediate dissociates from IscS, forming [2Fe–2S]–IscU2, which eventually dissociates, resulting in cluster coordination within a monomer of IscU.86 The physiological reductant for this reaction is provided by the [2Fe–2S] ferredoxin (Fdx) using NADPH-dependent ferredoxin reductase (FdxR) reaction.
IscU, in its role as the Fe–S cluster scaffold, interacts with multiple partners to enable cluster synthesis and transfer. Structures of the IscS2IscU2 complexes show that each IscU monomer engages with a functional active site monomer of IscS dimer (Fig. 4). The catalytic cysteine residue of IscS, positioned in a flexible loop, is not always resolved in structural models. However, its modeled location in the complex is at a possible distance from the IscU active-site cluster ligand Cys residues. The structure of the Archaeoglobus fulgidus cluster-bound [IscS-IscU D35A]2 complex shows that the coordination of the [2Fe–2S] cluster is provided by the three conserved cysteines of IscU and the Cys active site of IscS.87 While the role of the conserved Asp residue is not fully understood, substitution of this residue (D39 in E. coli IscU) with an alanine increases the stability of the Fe–S cluster.88–91 The structure of the IscU dimer from the thermophilic archaeon Methanothrix thermoacetophila shows that the Asp residue acts as a ligand to the [2Fe–2S] clusters coordinated at each monomer.92 Another residue involved in coordination is His (H105 in E. coli IscU). In the Aquifex aeolicus IscU, this His residue serves as the fourth ligand for the Fe–S cluster. Collectively, these structures indicate that while the three Cys residues appear to be invariant ligands, the fourth ligand varies between species or captures distinct stages of the assembly step. It is possible that these essential residues perform multiple functions during the assembly step, but also aid the cluster release mechanism by a “tug of war” mechanism previously described in the literature.93,94
In addition to residues involved in cluster coordination, the IscU surface contains critical residues for function. Essential non-cluster-coordinating residues provide contact points with IscS, orienting the protein in the correct position for sulfur transfer and subsequent interaction with biosynthetic partners, such as the chaperone HscA. Residues within the LPPVK motif (99–103 in E. coli IscU) enable interaction with the chaperone and co-chaperone HscA and HscB during cluster transfer of [2Fe–2S] clusters, where IscU shifts from an ordered to a disordered state, enabling release of the cluster to target proteins.95,96 The Tyr3 residue at the N-terminus of IscU in E. coli (Fig. 4) is essential for IscS sulfurtransferase activity by interacting with IscS at E311 via hydrogen bonding. In vitro experiments measuring the rate [2Fe–2S] cluster formation on E. coli IscU Y3A variant showed only slight defects,86 suggesting that the in vitro assay does not replicate the in vivo process or that the critical role of this residue lies at other steps of in vivo Fe–S assembly. Notably, in M. thermoacetophila holo IscU dimer structure, the equivalent residue Tyr7 forms a pi-stacking interaction with a conserved Phe residue (F61 in M. thermoacetophila) of the adjacent monomer.92 The engagement of conserved residues at the dimer interface (namely, Y7 with F61 and D40 with K104), positions the two clusters in the correct orientation for fusion. Thus, the dimerization of [2Fe–2S] IscU is proposed to be a step leading to the formation [4Fe–4S] clusters. This model is supported by the essentiality of equivalent residues in E. coli IscU (Y3, D39, K103)97 and earlier studies describing the reductive coupling of 2 × [2Fe–2S] on IscU dimer promoted by ferredoxin.98 Collectively, the working model describes the sequential formation of a [1Fe–1S] to [2Fe–2S] IscU, which is followed by the synthesis of [4Fe–4S] on an IscU dimer.
The bacterial ISC system also includes accessory proteins that support Fe–S cluster biogenesis, including IscA, IscX, CyaY. While IscS, IscU, Fdx, HscA, and HscB have defined and essential roles in Fe–S cluster formation as described above, genetic analysis of the studied E. coli and A. vinelandii models showed that inactivation of accessory proteins does not completely inactivate the ISC pathway.99 The recruitment of accessory proteins or even their presence varies across species.67,100 IscA, an A-type carrier (ATC), has been proposed to support cluster trafficking, and its apparent dispensability is explained by the presence of paralogous sequences, ErpA and SufA, that provide functional cross-talk.101,102 The functions of CyaY, a frataxin ortholog, and IscX remain not completely defined. Both proteins interact with IscS through a shared binding interface and modulate the interaction of IscS with IscU.77 Thus, CyaY and IscX emerge as regulators of sulfur transfer and Fe–S cluster formation under distinct iron concentrations.103 Interactions among other components of the ISC pathway provide excellent examples of the requirement for protein–protein interactions and how these interactions provide a point for regulating function.
2.2 Synthesis of Fe–S clusters by the SUF system
The SUF system is the most widespread system in bacteria.67 It is characterized by the presence of a two-component sulfur-mobilizing enzyme complex, SufSE or SufSU, and a three-component scaffold complex, SufBCD. The SUF system was first identified in E. coli through a genetic screen of pseudorevertants of isc-null strains.104 Mapping of suppressor mutations identified the sufABDCSE gene region, where mutations increased suf expression and, consequently, rescued the defects associated with inactivation of the isc genes. The SUF system operates in a supporting role for the ISC system, supplying Fe–S clusters under conditions that are detrimental to cluster synthesis in this organism. However, this duplication is one of a few exceptions, as analysis of sequenced genomes has shown that, in most cases, prokaryotes have evolved to contain either the ISC or SUF system.67 For instance, the SUF system is the main Fe–S cluster biosynthetic system and is essential for survival in Bacillus subtilis,105Staphylococcus aureus,106 and Mycobacterium tuberculosis.107 Therefore, the duplication of pathways first described in E. coli is considered an outlier among prokaryotic species.SufS initiates sulfur mobilization from Cys with a catalytic capacity equivalent to that of IscS. This enzyme also utilizes the PLP cofactor to promote the activation of the Cys substrate and the formation of a persulfide enzyme intermediate at a Cys residue.72 The active-site cysteine residue is located within a short, structured loop. Kinetic and structural characterization of the E. coli SufS–SufE reaction reveals that a structural element within SufS, designated the β-latch, impacts the reactivity of the enzyme.108,109 A hairpin within the β-latch from one subunit of the SufS dimer shields the active site Cys carrying the persulfide within the other subunit of SufS (Fig. 5). This structural motif prevents reaction with non-targets while facilitating complex formation with the physiological sulfur acceptor SufE and orienting the persulfide in the correct position for sulfur transfer. Together, the short active-site loop and the β-latch are characteristic of class II cysteine sulfurtransferases and are important elements that influence the selectivity of these enzymes and their dependencies on sulfur acceptor partners.
The reactivity of SufS and other class II sulfurtransferases is largely dependent on the engagement of a dedicated sulfur acceptor serving as an intermediate sulfurtransferase. Two types of sulfur acceptors have been described for SufS: SufE and SufU. Although these proteins lack amino acid sequence similarity, they share a similar architecture and perform equivalent functions in mediating sulfur-transfer reactions (Fig. 5). Sequence similarities between IscU and SufU and the ability of the SufSU reactions to accumulate Fe–S species in solution led to the initial proposal that SufU served as a scaffold protein.110,111 Subsequent biochemical and spectroscopic investigation of B. subtilis SufU, however, provided evidence that a restricted role of SufU is a zinc-dependent sulfurtransferase like SufE.112 This function was later supported by genetic complementation approaches in B. subtilis105 and analysis of sulfur transfer and Fe–S cluster assembly reactions involving SufS and SufU from M. tuberculosis and S. aureus.113–115
Genomic analysis shows that SufE and SufU are nearly mutually exclusive; that is, they rarely occur in the same species.116,117 The distribution of SUF-containing species that contain SufE is equivalent in numbers to those utilizing SufU. It is not clear what evolutionary factors impacted their distribution. From a sequence evolutionary perspective, SufU appears to have originated from a MisU ancestor sequence initially performing roles as a scaffold. However, the recruitment of a SufBCD scaffold and the tight binding of zinc to SufU can be considered factors that allowed this protein to specialize as a dedicated sulfurtransferase, in an equivalent capacity to that of SufE. In gammaproteobacteria, the presence of a sufE within the suf operon is often associated with the occurrence of another biosynthetic system in these genomes (87% of species),118 suggesting that the recruitment of SufE may have been justified in species where the SUF system operates in a specialized capacity, as in the case of E. coli. Moreover, these sulfur acceptors exhibit select reactivity towards their cognate SufS sulfurtransferases. For example, no functional cross-talk was observed between E. coli SufS and B. subtilis SufU and vice versa.105 However, functional sulfur-mobilizing complex E. coli SufSE could replace B. subtilis SufSU. Despite the high degree of sequence similarity between two SufS sequences, it is clear that these enzymes and respective sulfur acceptors display species-specific sequence elements restricting their functions.
In E. coli, SufE promotes protected sulfur transfer reactions from SufS to SufB via the conserved Cys51.112,119 The presence of SufE in reaction assays increases SufS turnover rate by up to 100-fold.112 The persulfide formed within SufS is protected from the action of reactive oxygen species, like H2O2, as well as reductants, such as DTT and TCEP. The binding of SufE induces conformational changes in SufS, enabling subsequent sulfur transfer. The presence of a persulfide within SufS or SufE enhances the binding affinity for the complex about 10-fold, indicating that SufS β-latch movement promotes the formation of a high-affinity closed complex (Fig. 5B).120 Sequential structural changes within SufS link the chemical steps in persulfide formation to protected sulfur transfer to SufE. The sulfuration state of SufE and subsequent persulfide transfer to SufB also affect the rate of sulfur transfer.112,121 The recent E. coli SufSE complex structure, while it does not show the β-latch in open conformation, provides insight into structural elements involved in binding and dissociation (Fig. 5).109 The conserved residue R119 of SufE is proposed to be involved in complex dissociation with SufS. A SufE R119A variant is still able to interact with SufS and serve as a sulfur acceptor, but does not elicit activity enhancement of SufS in vitro and is not a functional sulfur intermediate in cluster biogenesis in vivo.
Protein interactions between SufE and SufB are also critical for sulfur transfer.122 However, they appear not to be fully specific; E. coli SufE is capable of sulfur transfer to B. subtilis SufB, as evidenced by complementation studies of B. subtilis and E. coli SufB in both species.105 The SufB forms dynamic complexes with SufC and SufD proteins, and this interaction is critical for its function in cluster assembly in vivo. The different conformations of the three-component complex have been reported and include SufBC2D, SufB2C2, SufD2C2.123–125 While SufB and SufD share a high degree of sequence and structural similarity, in E. coli these proteins have diverged to perform distinct functions. Fe–S clusters are assembled within SufB and SufD interface,126 while SufD is proposed to serve as an intermediate site for iron delivery.127 FADH2 associates with the SufBC2D complex and is proposed to serve as the source of electrons in cluster assembly.128 SufC displays characteristic Walker A and Walker B ATPase motifs, but its structure includes a structural and regulatory element, the Q-loop, that positions the catalytic Glu171 residue away from the active site, resulting in low ATPase activity.127 Complex formation with SufBD proteins induces a conformational change in the Q loop, reorienting Glu171 toward the active site and thereby enhancing ATPase activity. The ATPase activity of the SufBC2D complex, however, is modulated through interactions with SufE.129 Computational models provided insight that residues Lys9 and Arg16 could interact with a patch of negative residues (D116, D119, D122) within SufC, providing the molecular basis of a non-competitive inhibitory effect of SufE on the ATPase activity of SufC. In fact, Ala substitution of K9 and R16 of SufE alleviates the inhibition of ATPase activity.129 Interestingly, these residues are also important for sulfur transfer, suggesting a coordinated sequence of events involving sulfur transfer and ATP hydrolysis during Fe–S cluster assembly by the E. coli SUF system.
A second type of SufS sulfur acceptor is provided by the zinc-dependent SufU (Fig. 5A). The role of SufU is equivalent to that of SufE in mediating sulfur transfer from SufS to SufB in several species, including the studied B. subtilis,130Staphylococcus aureus,114Mycobacterium tuberculosis,113 and Enterococcus faecalis.131 SufU shares sequence and structural similarity to IscU, where conserved Cys and Asp residues coordinate a tightly-bound zinc (Ka = 1017), therefore restricting the function of SufU as a dedicated sulfurtransferase of SufS.113,132 The mechanism for sulfur transfer is initiated by direct interaction between SufS and SufU, in which a conserved His residue of SufS (B. subtilis H342) transiently coordinates the zinc, displacing SufU Cys41 to allow for nucleophilic attack on the SufS Cys361 persulfide (Fig. 6).133 The ligand switch serves as a mechanistic feature to enhance the reactivity of Cys41 during sulfur transfer.133 Kinetic analysis supports this model, as the rate-limiting step of this reaction is determined by the availability of the deprotonated form of SufU, which controls the second half of the sulfurtransferase reaction.132
Recent structural analysis of B. subtilis SufU in its persulfurated form shows the engagement of Arg125 in stabilizing the persulfide sulfur, while the C41 bridging sulfur is bound to the zinc cofactor (Fig. 6).134 Arg125 is conserved across SufU sequences, and substitution to Ala or Lys impairs the ability of SufU to serve as a substrate of SufS. Together, the structures of SufU before reaction with SufS, the SufSU complex, and persulfurated SufU provide snapshots of the sulfur-transfer process and the involvement of several residues, guiding coordinated conformational changes and reactivity of both sulfur donors and acceptors (Fig. 6). Despite sequence differences, structural alignment of B. subtilis SufU and E. coli SufE reveals similar architectures and equivalent positions for Cys residues 41 and 51, respectively, involved in persulfide formation (Fig. 5).121,134 Interestingly, SufE R119 is also located in an equivalent position as SufU R125 (Fig. 5C). It is possible that these residues perform equivalent functions in stabilizing the persulfide bond and orienting the persulfide sulfur for subsequent transfer to SufB.
Distinct catalytic activities have been reported for SufSU reactions from different species. The catalytic turnover of SufS sulfurtransferase reaction in the presence of the sulfur acceptor is faster than the cysteine desulfurase reaction. That is, the half-reaction cycle leading to the formation of persulfide and reduction is very slow (0.2–0.4 min−1),130 likely due to the beta-latch structure protecting the active site from direct reduction of the persulfide by an artificial reductant like DTT. The rate of the complete sulfurtransferase reaction in the presence of SufU ranges from 1.5 to 45 min−1 across species.113,114,132,135 While reaction conditions such as pH and the SufS-to-SufU ratio affect the overall reaction rate, the low activity of the Mycoplasma penetrans SufS sulfurtransferase reaction has been attributed to the substitution of the conserved His residue (342 in B. subtilis), which is involved in transient zinc coordination.135 In B. subtilis, SufS H342 is essential for the sulfurtransferase activity;133 however, in M. penetrans, SufS Gln342 occupies this position (Fig. 7). Mutagenesis studies have shown that amino acid substitutions at this position yield varying degrees of activity. Variants containing residues that eliminate zinc binding (Ala, Tyr, Ser) showed no activity enhancement in the presence of SufU, whereas substitution with His, Glu, or Cys showed improved activity.135
It is unclear why most Mycoplasma species retain an apparently suboptimal residue, causing reduced efficiency of sulfurtransferase activity. It is worth noting that Mycoplasma species have reduced genomes and are missing components in multiple pathways. While the presence of SufS is consistent across species within this group, other SUF components are not always present.136 The lack of a complete SUF pathway, along with the absence of genes encoding several Fe–S enzymes, supports the hypothesis that some organisms have transitioned to a metabolism that is independent of Fe–S clusters.137 For example, the genomes of M. genitalium and M. pneumoniae contain SufSU copies, but lack genes coding for SufB, SufC, and SufD, as well as known Fe–S proteins.2 In M. pneumoniae, SufS (HapE) function is associated with haemolytic activity, likely due to its sulfide-release activity.137 Although these investigations have not probed the function of its counterpart SufU, species with condensed genomes have retained only a SufSU copy, with no other cysteine sulfurtransferase gene, suggesting their involvement in sulfur transfer reactions beyond Fe–S cluster biogenesis. The unique features of distinct SUF pathways, including mechanistic differences between the SufE and SufU components, not only explain their lack of functional cross-talk but also provide a point for exploration in the development of targeted inhibitors specific to subgroups of microbes.
2.3 Synthesis of Fe–S clusters by the NIF system
The NifU–NifS pair constitutes a two-component Fe–S cluster biosynthetic system first described in nitrogen-fixing organisms.138 NifS is the cysteine sulfurtransferase with sequence and mechanistic similarity to IscS and other class I cysteine desulfurases.66 NifU serves as a platform for the synthesis of [2Fe–2S] and [4Fe–4S] clusters.139 NifU is a modular protein containing three functional domains (Fig. 8).140 The N-terminal module is similar to IscU and is proposed to serve as the starting point for Fe–S cluster synthesis. The central module contains a [2Fe–2S] ferredoxin-like permanent cluster with proposed function in mediating electron transfer during cluster biogenesis. Lastly, the C-terminal region shares sequence similarity with Nfu cluster carrier proteins, thereby facilitating the delivery of newly formed clusters to target acceptors. Therefore, NifU combines three previously described functions in cluster synthesis: cluster scaffold, electron donor, and cluster carrier, all within a single polypeptide.Structural similarities between NifUS and IscSU pairs, and computational models, predict that the N-terminal domain of NifU directly interacts with NifS for cluster assembly, using a binding mode similar to that of IscU to IscS (Fig. 9).77 It is anticipated that Fe–S cluster assembly initiates on the N-terminal domain of NifU, using an equivalent mechanism to that employed by the IscSU pair. Residues involved in cluster coordination and shown to affect cluster synthesis on IscU are also conserved in NifU, including Cys35, Cys62, and Cys106, as well as Asp37 and His105. Structural models also predict that each of the three domains folds independently, supporting the proposal of three functions (Fig. 8). The sequence of the central domain is unique to NifU and can be used as a diagnostic for identifying the presence of the NifUS system across diverse species.138 The [2Fe–2S] central cluster has redox properties similar to ferredoxin-like [2Fe–2S] clusters,141 supporting the idea that the central domain performs functions equivalent to those of ferredoxin in the ISC system. Lastly, the C-terminal Nfu domains are connected to the protein through a variable linker. The sequence and length of this linker vary among species, and it is thought to grant flexibility of this domain for intra- and intermolecular interactions during cluster transfer.138
The NIF system was first characterized in the diazotroph A. vinelandii in functional studies of genes involved in nitrogen fixation.142 Subsequent characterization established that NifUS are involved in the synthesis of the three nitrogenase metallocofactors, the [4Fe–4S] cluster of Fe-protein (NifH), the P-cluster, and the FeMo cofactor of MoFe protein (NifDK).24 Recent studies have also established that NifU provides clusters for NifQ maturation and participates in Mo delivery for FeMo cofactor assembly.61 While a direct transfer of [4Fe–4S] cluster is sufficient for the maturation of NifH and NifQ, the synthesis of P-cluster and FeMo cofactor involves additional biosynthetic components for the conversion of [4Fe–4S] units into more elaborate metallic structures. NifUS are also recruited for the maturation of alternative Anf- and Vnf-nitrogenases in that organism.143 The maturation of these enzymes, however, has not been fully elucidated. It is presumed that NifUS are required for the synthesis of their active sites, FeV and FeFe cofactor, since they share a biosynthetic precursor NifB-co,144 which is known to receive [4Fe–4S] clusters from NifUS. Genomic studies showed that NifUS are predominantly present in aerobic diazotrophs, and their presence is rationalized by the increased demand for cluster synthesis and repair under conditions detrimental to Fe–S clusters.
The occurrence of NifUS sequences is not limited to diazotrophs or aerobic organisms. The presence of three-domain NifU-like sequences has been investigated in Helicobacter pylori and is also shown to coordinate the central cluster in its as-isolated form.145 Complementation experiments showed that H. pylori NifUS is able to replace E. coli ISC and SUF systems, demonstrating that this system can provide Fe–S clusters for housekeeping functions.146 The NifUS system has also been reported in eukaryotes, Entamoeba histolytica and Mastigamoeba malamutei.147,148 These organisms lack a standard ISC described in eukaryotes, and their mitochondria also lack the characteristic respiratory functions. Instead, these atypical organelles, defined as mitosomes, retain Fe–S cluster biosynthetic functions carried out by NifUS and specialized functions, such as those of the hydrogenosome, to support their anaerobic metabolism. The absence of genomic nitrogen-fixing neighborhood context led to the distinct classification of these NifU-like sequences as MisU, which also includes single-domain IscU-like sequences.67 However, biochemical data and the presence of three-domain NifU, with the characteristic central-domain sequence, are distinct from those of IscU sequences in both sequence and function. Notably, the diverse distribution of NifU and NifU-like sequences across taxonomic groups is also supported by analyses of sequenced genomes that cannot distinguish diazotrophs from non-diazotroph species in distinct branches.138 Overall, these sequences show strong genomic synteny for the occurrence of nifU and nifS genes, and the NifU sequences in nearly all cases contain conserved cluster ligand residues within its three domains. The NIF system offers an excellent model for studying the synthesis of Fe–S clusters because it is a two-component system yet harbors the functionality provided by multiple peptides in the ISC system. The three domains within NifU may be viewed as an optimized strategy for synthesizing initial building blocks that can undergo reductive couplings and subsequent transfer to acceptor targets.
2.4 Synthesis of Fe–S clusters by abbreviated systems: SMS and MIS
The SMS and MIS systems are recently classified truncated Fe–S biogenesis systems that have existed since the Last Universal Common Ancestor (LUCA).63,67 These systems are the least studied and research on the role of each component and the requirement for additional factors were unknown until recently. These systems lack several key components and associated functionalities that are present in the ISC, SUF, and NIF systems (Fig. 3). While the SMS and MIS systems remained in archaea, their diversification appeared in bacteria. The origin of the SUF system is likely the SMS system, whereas the origins of the ISC and NIF systems are likely the MIS system. Many evolutionary events have been hypothesized to have contributed to the development of these systems, including gene duplication, horizontal gene transfer, and diversification in response to environmental factors and the demand for multiple stable Fe–S clusters.63 For example, there is a clear divergence between SUF and SMS systems based on the presence of oxygen. The SUF system is primarily found in bacteria and certain groups of aerobic archaea, whereas the SMS system is primarily found in archaea and in select groups of anaerobic bacteria.The SMS system contains only the scaffold SmsCB. These proteins are homologous to the SUF scaffold proteins SufC and SufB. Recent characterization of this system across different organisms indicates that, in general, SmsC exhibits AAA+ ATPase activity that is enhanced by SmsB, in a manner similar to that described for the SUF system.67,149–151 The SmsC2B2 complex in Methanosarcina acetivorans, Methanocaldococcus jannaschii, and Methanococcus maripaludis was validated to function as an Fe–S cluster assembly scaffold and capable of transferring these clusters to apo-proteins.67,150 Sequence comparison between SufB and SmsB shows conservation of Cys residues implicated in cluster binding in SufB, but these residues do not affect cluster binding in the SmsB2C2 complex in Methanothermococcus thermolithotrophicus, suggesting that some species use an alternate site for cluster ligation.150
A subset of SmsC sequences contains a C-terminal region (CTR) to coordinate a Fe–S cluster. Intriguingly, the structure of M. jannaschii SufB2C2 shows the presence of a single [4Fe–4S] cluster, coordinated by three cysteine residues, on a single SmsC subunit within the CTR (Fig. 10). This region is disordered in the apo-SmsC structure and becomes structurally defined upon cluster formation.149 This type of cluster coordination was also validated through a mutagenesis approach in M. thermolithotrophicus.150 These two species, along with a number of species that contain the SMS system, exhibit sequence variation within the CTR of SmsC, but retain suitable cluster ligand residues.150 The occurrence of Cys residues within CTRs in SmsC provides novel sites for transient clusters coordination and leaves the role of SmsB in this species unresolved. The ability of SmsC to bind Fe–S clusters was unexpected and demonstrates a stark mechanistic difference from the SufBCD scaffold, which uses SufB as the cluster-binding site.149 In M. jannaschii SmsC, the binding of the Fe–S cluster and ATP was proposed to be mutually exclusive, based on experiments performed with non-hydrolyzable ATP.149 However, the ATPase activity of M. thermolithotrophicus SmsC was unaffected by the presence of the cluster.150 Therefore, it remains to be determined which steps in cluster synthesis require ATPase activity. It is worth noting that the occurrence of CTR is not universal to all SmsC sequences. For example, CTR is absent in Thermococcus kodakarensis SmsC, and the SmsC2B2 complex from this organism has been shown to be competent in Fe–S cluster assembly and in the maturation of the Fe–S target protein LipS.151 It is possible that the current definition of SMS systems encompasses all truncated Suf-like systems, with members at distinct evolutionary stages and performing varying functions that remain to be characterized.
Additional distinctive features of the SMS system include the absence of a cysteine desulfurase, a chaperone, or a cluster carrier protein within the genomic region (Fig. 3). Some SMS-containing species, such as T. kodakarensis, possess a sufS-like gene, which is dispensable in the presence of elemental sulfur.152 Several other species completely lack a cysteine sulfurtransferase gene.63 This observation was initially intriguing because this enzyme is required for the first step of sulfur mobilization in all other studied Fe–S biogenesis systems, raising questions about how sulfur incorporation occurs in organisms that lack this catalytic function. While the involvement of an alternative sulfurtransferase in these systems cannot be ruled out at this point, current evidence supports the idea that sulfide is directly added to the scaffold during Fe–S cluster synthesis. The SMS system is primarily found in archaeal species, including methanogens and hyperthermophiles. These organisms thrive in hot, anaerobic environments such as hydrothermal vents and volcanic marine sediments, where inorganic sulfur is plentiful.151 Earlier studies excluded cysteine as the sulfur source for Fe–S cluster biogenesis in M. maripaludis,153 supporting a model in which sulfide is directly used to assemble Fe–S clusters on the SMS scaffold proteins. Recently, additional support for this model was provided by interspecies complementation experiments showing that M. jannaschii smsBC can rescue the lethal phenotype caused by inactivation of the E. coli ISC and SUF systems when the growth medium is supplemented with sulfide, but not cysteine.149 Although it remains to be fully characterized, it is likely that the synthesis of the Fe–S cluster directly from sulfide involves a distinct mechanism, as it probably eliminates the need for reduction of a persulfide intermediate, like in other systems.
The MIS system is the least studied iron–sulfur cluster biogenesis system. This system contains a cysteine desulfurase (MisS) and scaffold pair (MisU) that are homologous to IscS and IscU.67 This system does not contain additional components such as a carrier protein or chaperones and has previously been mischaracterized as belonging to other biogenesis systems.154 Phylogenetic classification of MIS proposed by Garcia et al. defined NifUS-like proteins from non-diazotrophs as MisSU. This assignment was supported by clustering NifS coding sequences from nitrogen-fixation (nif) genomic regions into a distinct branch relative to MisS sequences from non-diazotrophs.67 Using this classification, assigned MisU sequences included IscU-like sequences and three-domain NifU-like sequences, which are known to be distinct in sequence and number of cluster coordination sites. From a catalytic mechanistic perspective, it is likely that these tentatively assigned three-domain MisU sequences operate as NifU-like, displaying chemical reactivity similar to that reported for NifU and were discussed in this review as NifU-like scaffolds. Likewise, the assembly of Fe–S clusters by IscU-like MisU sequences is proposed to follow steps analogous to those described for IscU. However, the self-contained functionality of MIS sequences displaying IscU-like sequences remains to be validated.
A number of archaeal species contain both SMS and MIS systems. M. acetivorans, for example, contains two copies of SMS and three copies of MIS. While both copies of SMS, SmsC1B1 and SmsC2B2, are competent for cluster assembly, a strain carrying inactivation of both copies remains viable.155 Likewise, inactivation of MisU2 also did not affect Fe–S cluster content in that organism.154 It is possible that the occurrence of multiple copies provides functional cross-talk among the various SMS and MIS systems in this organism and enables the organism to adapt its metabolism based on sulfur source. There is a clear separation between methanogens that contain both systems and those that have only the SMS system, with respect to their ability to utilize sulfide and cysteine as sources of Fe–S cluster formation. These minimal systems in archaea provide more insight into the origins and evolution of the present-day Fe–S biogenesis systems. This new and exciting area of exploration provides an opportunity to uncover mechanistic details of Fe–S cluster synthesis by these condensed systems and may reveal the involvement of novel factors supporting sulfur and iron acquisition, cluster assembly, and protein maturation as organisms containing these pathways are subjected to distinct environmental and metabolic challenges.
3. Concluding remarks
Fe–S clusters are ancient inorganic cofactors with a diverse set of functionalities. These ubiquitous cofactors require distinct pathways for the incorporation of sulfur, iron, electrons, and the formation of the clusters. The sulfur source has recently been a point of interest since the identification of minimal iron–sulfur systems that dispense the requirement of a cysteine sulfurtransferase. Sulfur has been an essential element since the beginning of time, when it was abundant alongside iron, resulting in mineral species such as pyrite (FeS2), mackinawite (FeS), and other metal sulfides. The iron–sulfur world hypothesis theorized that iron–sulfur species from hydrothermal systems served as catalysts in essential metabolic processes.156 It was previously postulated that iron and sulfur spontaneously formed Fe–S clusters in the anaerobic, high-temperature, and high-pressure environments accessible in early life. However, the presence of the Fe–S cluster biogenesis protein machinery, the MIS and SMS systems, in the Last Universal Common Ancestor (LUCA) challenges this hypothesis. Exogenous sulfur likely served as the sulfur source, and cluster formation was facilitated by protein machinery rather than occurring spontaneously.67 The recruitment of protein components for Fe–S cluster formation in biological systems has evolved from truncated systems that use elemental sulfur to intricate ones described here that utilize L-cysteine.While most Fe–S cluster biosynthetic systems follow similar general steps for sulfur mobilization, cluster assembly, and transfer to target proteins, their functional and biochemical characterization reveal key differences. The rise of different cysteine desulfurases, likely from the MisS, allowed for L-cysteine to serve as a sulfur donor for the biosynthesis of these clusters in present-day systems and in the biosynthesis of other sulfur-containing cofactors. Thus, cysteine sulfurtransferases, in the present, perform versatile roles in sulfur metabolism. In some organisms, such as B. subtilis, gene duplication has enabled these enzymes to perform dedicated roles as cysteine sulfurtransferases as described for SufS–SufU, NifZ–ThiI, YrvO–MnmA, and NifS–NadA sulfur donor–acceptor pairs.32,66,112,157 Specific protein–protein interactions between these enzymes and their dedicated sulfur acceptors restrict their reactivities and, consequently, biological functions.66 For instance, the BsNifS enzyme is directly involved in sulfur transfer to the final Fe–S target quinolinate synthase NadA, presumably via in situ assembly of the [4Fe–4S] on that enzyme.157 A distinct evolutionary strategy was observed in E. coli that utilizes a main cysteine sulfurtransferase to interact with multiple partners, as described for IscS and its sulfur acceptors IscU, TusA, and ThiI.73,77 EcIscS was previously coined as a promiscuous enzyme.72,158 Recent literature has challenged this paradigm by uncovering that specific sequence elements in these protein–protein interactions are essential for enzyme reactivity and function.
The versatile chemistries promoted by Fe–S proteins and their involvement in life-sustaining reactions justify the persistence of these fragile cofactors in the current oxygenated world. The occurrence and inventory of chemical functionalities associated with this class of proteins continues to expand. The recent discovery of novel cluster coordination motifs and improved computational models for structural predictions have directed research for reassignment of previously overlooked Fe–S sites. Furthermore, improved methodology for isolating these proteins under conditions that preserve Fe–S clusters in their intact and active forms has enabled validation of new in silico assignments. Fe–S clusters are embedded in the biochemistry of life. The involvement of Fe–S proteins in a multitude of biochemical processes opens the door to the application of their functionalities in biotechnology. Additionally, the specificity of their biosynthetic enzymes across microbial groups, along with their essentiality, suggests therapeutic potential for treating infectious agents.
Author contributions
SMS and PDS conceptualized the topic, wrote, and edited the review paper.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analyzed as part of this review.Acknowledgements
This study was supported by the National Science Foundation (MCB-2335999) and the Wake Forest Center for Molecular Signalling.Notes and references
- R. C. Valentine, Bacterial ferredoxin, Bacteriol. Rev., 1964, 28, 497–517, DOI:10.1128/br.28.4.497-517.1964.
- J. J. Braymer, S. A. Freibert, M. Rakwalska-Bange and R. Lill, Mechanistic concepts of iron-sulfur protein biogenesis in biology, Biochim. Biophys. Acta, Mol. Cell Res., 2021, 1868, 118863, DOI:10.1016/j.bbamcr.2020.118863.
- C. Ayala-Castro, A. Saini and F. W. Outten, Fe–S cluster assembly pathways in bacteria, Microbiol. Mol. Biol. Rev., 2008, 72, 110–125, DOI:10.1128/mmbr.00034-07.
- M. K. Raza, V. R. Jeyachandran and S. Bashir, Investigating iron-sulfur proteins in infectious diseases: a review of characterization techniques, Inorganics, 2024, 12, 25, DOI:10.3390/inorganics12010025.
- R. R. Mendel, T. W. Hercher, A. Zupok, M. A. Hasnat and S. Leimkühler, The requirement of inorganic Fe–S clusters for the biosynthesis of the organometallic molybdenum cofactor, Inorganics, 2020, 8, 43, DOI:10.3390/inorganics8070043.
- D. W. Bak and S. J. Elliott, Alternative FeS cluster ligands: tuning redox potentials and chemistry, Curr. Opin. Chem. Biol., 2014, 19, 50–58, DOI:10.1016/j.cbpa.2013.12.015.
- C. Vallières, O. Benoit, O. Guittet, M. E. Huang, M. Lepoivre, M. P. Golinelli-Cohen and L. Vernis, Iron-sulfur protein odyssey: exploring their cluster functional versatility and challenging identification, Metallomics, 2024, 16, mfae025, DOI:10.1093/mtomcs/mfae025.
- B. A. Skeel and D. L. M. Suess, Iron-sulfur clusters: the road to room temperature, J. Biol. Inorg. Chem., 2025, 30, 151–159, DOI:10.1007/s00775-025-02094-0.
- A. D. Read, R. E. T. Bentley, S. L. Archer and K. J. Dunham-Snary, Mitochondrial iron–sulfur clusters: structure, function, and an emerging role in vascular biology, Redox Biol., 2021, 47, 102164, DOI:10.1016/j.redox.2021.102164.
- D. Flemming, A. Schlitt, V. Spehr, T. Bischof and T. Friedrich, Iron-sulfur cluster N2 of the Escherichia coli NADH:ubiquinone oxidoreductase (complex I) is located on subunit NuoB, J. Biol. Chem., 2003, 278, 47602–47609, DOI:10.1074/jbc.M308967200.
- J. Hirst, Mitochondrial complex I, Annu. Rev. Biochem., 2013, 82, 551–575, DOI:10.1146/annurev-biochem-070511-103700.
- J. Zhu, K. R. Vinothkumar and J. Hirst, Structure of mammalian respiratory complex I, Nature, 2016, 536, 354–358, DOI:10.1038/nature19095.
- E. Gnandt, K. Dörner, M. F. J. Strampraad, S. de Vries and T. Friedrich, The multitude of iron–sulfur clusters in respiratory complex I, Biochim. Biophys. Acta, Bioenerg., 2016, 1857, 1068–1072, DOI:10.1016/j.bbabio.2016.02.018.
- T. M. Iverson, C. Luna-Chavez, G. Cecchini and D. C. Rees, Structure of the Escherichia coli fumarate reductase respiratory complex, Science, 1999, 284, 1961–1966, DOI:10.1126/science.284.5422.1961.
- T. M. Iverson, P. K. Singh and G. Cecchini, An evolving view of complex II—noncanonical complexes, megacomplexes, respiration, signaling, and beyond, J. Biol. Chem., 2023, 299, 104761, DOI:10.1016/j.jbc.2023.104761.
- M. Sarewicz, M. Dutka, S. Pintscher and A. Osyczka, Triplet State of the Semiquinone–Rieske Cluster as an Intermediate of Electronic Bifurcation Catalyzed by Cytochrome bc1, Biochemistry, 2013, 52, 6388–6395, DOI:10.1021/bi400624m.
- H. Beinert, M. C. Kennedy and C. D. Stout, Aconitase as Iron–Sulfur Protein, Enzyme, and Iron-Regulatory Protein, Chem. Rev., 1996, 96, 2335–2374, DOI:10.1021/cr950040z.
- A. McSkimming and D. L. M. Suess, Dinitrogen binding and activation at a molybdenum–iron–sulfur cluster, Nat. Chem., 2021, 13, 666–670, DOI:10.1038/s41557-021-00701-6.
- B. M. Hoffman, W. E. Broderick and J. B. Broderick, Mechanism of Radical Initiation in the Radical SAM Enzyme Superfamily, Annu. Rev. Biochem., 2023, 92, 333–349, DOI:10.1146/annurev-biochem-052621-090638.
- A. G. Baranovskiy, H. M. Siebler, Y. I. Pavlov and T. H. Tahirov, Iron–Sulfur Clusters in DNA Polymerases and Primases of Eukaryotes, Methods Enzymol., 2018, 599, 1–20, DOI:10.1016/bs.mie.2017.09.003.
- S. Pokharel and J. L. Campbell, Cross talk between the nuclease and helicase activities of Dna2: role of an essential iron–sulfur cluster domain, Nucleic Acids Res., 2012, 40, 7821–7830, DOI:10.1093/nar/gks534.
- Y. Wu and R. M. Brosh Jr., DNA helicase and helicase–nuclease enzymes with a conserved iron–sulfur cluster, Nucleic Acids Res., 2012, 40, 4247–4260, DOI:10.1093/nar/gks039.
- S. H. Newman-Stonebraker, T. J. Gerard and P. L. Holland, Opportunities for insight into the mechanism of efficient CO2/CO interconversion at a nickel-iron cluster in CO dehydrogenase, Chem, 2024, 10, 1655–1667, DOI:10.1016/j.chempr.2024.04.012.
- J. S. Martin del Campo, J. Rigsbee, M. Bueno Batista, F. Mus, L. M. Rubio, O. Einsle, J. W. Peters, R. Dixon, D. R. Dean and P. C. Dos Santos, Overview of physiological, biochemical, and regulatory aspects of nitrogen fixation in Azotobacter vinelandii, Crit. Rev. Biochem. Mol. Biol., 2022, 57, 492–538, DOI:10.1080/10409238.2023.2181309.
- R. G. Sawers, M. Hardelt, A. Haase and D. Lubek, Biosynthesis and assembly of hydrogenase [NiFe]-cofactor: recent advances and perspectives, Metallomics, 2025, 17, mfaf015, DOI:10.1093/mtomcs/mfaf015.
- O. Bimai, S. Arragain and B. Golinelli-Pimpaneau, Structure-based mechanistic insights into catalysis by tRNA thiolation enzymes, Curr. Opin. Struct. Biol., 2020, 65, 69–78, DOI:10.1016/j.sbi.2020.06.002.
- M. Fellner, R. P. Hausinger and J. Hu, A structural perspective on the PP-loop ATP pyrophosphatase family, Crit. Rev. Biochem. Mol. Biol., 2018, 53, 607–622, DOI:10.1080/10409238.2018.1516728.
- S. Gervason, S. Sen, M. Fontecave and B. Golinelli-Pimpaneau, [4Fe-4S]-dependent enzymes in non-redox tRNA thiolation, Biochim. Biophys. Acta, Mol. Cell Res., 2024, 1871, 119807, DOI:10.1016/j.bbamcr.2024.119807.
- S. Arragain, O. Bimai, P. Legrand, S. Caillat, J.-L. Ravanat, N. Touati, L. Binet, M. Atta, M. Fontecave and B. Golinelli-Pimpaneau, Nonredox thiolation in tRNA occurring via sulfur activation by a [4Fe-4S] cluster, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 7355–7360, DOI:10.1073/pnas.1700902114.
- S. Gervason, S. Sen, J.-L. Ravanat, S. Caillat, D. Hamdane and B. Golinelli-Pimpaneau, Deciphering the influence of the [4Fe–4S] cluster of tRNA thiolation enzymes on tRNA binding, RNA, 2025, 31, 735–742, DOI:10.1261/rna.080292.124.
- J. Zhou, M. Lénon, J. L. Ravanat, N. Touati, C. Velours, K. Podskoczyj, G. Leszcynska, M. Fontecave, F. Barras and B. Golinelli-Pimpaneau, Iron–sulfur biology invades tRNA modification: the case of U34 sulfuration, Nucleic Acids Res., 2021, 49, 3997–4007, DOI:10.1093/nar/gkab138.
- K. A. Black and P. C. Dos Santos, Abbreviated Pathway for Biosynthesis of 2-Thiouridine in Bacillus subtilis, J. Bacteriol., 2015, 197, 1952–1962, DOI:10.1128/jb.02625-14.
- S. Leimkühler, 2-Thiouridine formation in Escherichia coli: a critical review, J. Bacteriol., 2024, 207, e00420-24, DOI:10.1128/jb.00420-24.
- S. Gervason, S. Sen, J. Zhou, K. Podskoczyj, G. Leszczynska, S. Caillat, J. L. Ravanat, M. Fontecave and B. Golinelli-Pimpaneau, The thiolation of U34 at carbon 2 in tRNA by Escherichia coli MnmA precedes modification at carbon 5 and is dependent on a [4Fe-4S] cluster, J. Inorg. Biochem., 2026, 274, 113064, DOI:10.1016/j.jinorgbio.2025.113064.
- J. B. Broderick, W. E. Broderick and B. M. Hoffman, Radical SAM enzymes: Nature's choice for radical reactions, FEBS Lett., 2023, 597, 92–101, DOI:10.1002/1873-3468.14519.
- Y. Nicolet, Structure–function relationships of radical SAM enzymes, Nat. Catal., 2020, 3, 337–350, DOI:10.1038/s41929-020-0448-7.
- S. J. Booker and C. T. Lloyd, Twenty Years of Radical SAM! The Genesis of the Superfamily, ACS Bio Med Chem Au, 2022, 2, 538–547, DOI:10.1021/acsbiomedchemau.2c00078.
- A. E. Boncella, E. T. Sabo, R. M. Santore, J. Carter, J. Whalen, J. D. Hudspeth and C. N. Morrison, The expanding utility of iron-sulfur clusters: Their functional roles in biology, synthetic small molecules, maquettes and artificial proteins, biomimetic materials, and therapeutic strategies, Coord. Chem. Rev., 2022, 453, 214229, DOI:10.1016/j.ccr.2021.214229.
- F. Hammami, L. Tichit, B. Py, F. Barras, P. Mandin and E. Remy, Analysis of a logical regulatory network reveals how Fe–S cluster biogenesis is controlled in the face of stress, MicroLife, 2023, 4, uqad003, DOI:10.1093/femsml/uqad003.
- E. L. Mettert and P. J. Kiley, Fe–S cluster homeostasis and beyond: The multifaceted roles of IscR, Biochim. Biophys. Acta, Mol. Cell Res., 2024, 1871, 119749, DOI:10.1016/j.bbamcr.2024.119749.
- K. Kobayashi, Sensing Mechanisms in the Redox-Regulated, [2Fe–2S] Cluster-Containing, Bacterial Transcriptional Factor SoxR, Acc. Chem. Res., 2017, 50, 1672–1678, DOI:10.1021/acs.accounts.7b00137.
- R. Banerjee, I. Askenasy, E. L. Mettert and P. J. Kiley, Iron–sulfur Rrf2 transcription factors: an emerging versatile platform for sensing stress, Curr. Opin. Microbiol., 2024, 82, 102543, DOI:10.1016/j.mib.2024.102543.
- G. Shen, R. Balasubramanian, T. Wang, Y. Wu, L. M. Hoffart, C. Krebs, D. A. Bryant and J. H. Golbeck, SufR Coordinates Two [4Fe-4S]2+, 1+ Clusters and Functions as a Transcriptional Repressor of the sufBCDS Operon and an Autoregulator of sufR in Cyanobacteria, J. Biol. Chem., 2007, 282, 31909–31919, DOI:10.1074/jbc.M705554200.
- Y. Cheng, M. Lyu, R. Yang, Y. Wen, Y. Song, J. Li and Z. Chen, SufR, a [4Fe-4S] Cluster-Containing Transcription Factor, Represses the sufRBDCSU Operon in Streptomyces avermitilis Iron-Sulfur Cluster Assembly, Appl. Environ. Microbiol., 2020, 86, e01523-20, DOI:10.1128/AEM.01523-20.
- J. C. Crack and N. E. Le Brun, Mass Spectrometric Identification of [4Fe–4S](NO)x Intermediates of Nitric Oxide Sensing by Regulatory Iron–Sulfur Cluster Proteins, Chem. – Eur. J., 2019, 25, 3675–3684, DOI:10.1002/chem.201806113.
- M. T. Pellicer Martinez, J. C. Crack, M. Y. Stewart, J. M. Bradley, D. A. Svistunenko, A. W. Johnston, M. R. Cheesman, J. D. Todd and N. E. Le Brun, Mechanisms of iron- and O2-sensing by the [4Fe-4S] cluster of the global iron regulator RirA, eLife, 2019, 8, e47804, DOI:10.7554/eLife.47804.
- T. A. Rouault, The role of iron regulatory proteins in mammalian iron homeostasis and disease, Nat. Chem. Biol., 2006, 2, 406–414, DOI:10.1038/nchembio807.
- A. M. SantaMaria and T. A. Rouault, Regulatory and Sensing Iron–Sulfur Clusters: New Insights and Unanswered Questions, Inorganics, 2024, 12, 101, DOI:10.3390/inorganics12040101.
- H. Beinert, R. H. Holm and E. Münck, Iron-Sulfur Clusters: Nature's Modular, Multipurpose Structures, Science, 1997, 277, 653–659, DOI:10.1126/science.277.5326.653.
- B. J. Vaccaro, S. M. Clarkson, J. F. Holden, D. W. Lee, C. H. Wu, F. L. Poole II, J. J. H. Cotelesage, M. J. Hackett, S. Mohebbi, J. Sun, H. Li, M. K. Johnson, G. N. George and M. W. W. Adams, Biological iron-sulfur storage in a thioferrate-protein nanoparticle, Nat. Commun., 2017, 8, 16110, DOI:10.1038/ncomms16110.
- S. W. Vali, D. K. Haja, R. A. Brand, M. W. W. Adams and P. A. Lindahl, The Pyrococcus furiosus ironome is dominated by [Fe4S4]2+ clusters or thioferrate-like iron depending on the availability of elemental sulfur, J. Biol. Chem., 2021, 296, 100710, DOI:10.1016/j.jbc.2021.100710.
- C. Johnson, A. England, M. Munro-Ehrlich, D. R. Colman, J. L. DuBois and E. S. Boyd, Pathways of Iron and Sulfur Acquisition, Cofactor Assembly, Destination, and Storage in Diverse Archaeal Methanogens and Alkanotrophs, J. Bacteriol., 2021, 203, e00117-21, DOI:10.1128/jb.00117-21.
- N. D. Lanz and S. J. Booker, Auxiliary iron–sulfur cofactors in radical SAM enzymes, Biochim. Biophys. Acta, Mol. Cell Res., 2015, 1853, 1316–1334, DOI:10.1016/j.bbamcr.2015.01.002.
- E. L. McCarthy and S. J. Booker, Biochemical Approaches for Understanding Iron–Sulfur Cluster Regeneration in Escherichia coli Lipoyl Synthase During Catalysis, Methods Enzymol., 2018, 606, 217–239, DOI:10.1016/bs.mie.2018.06.006.
- E. L. McCarthy and S. J. Booker, Destruction and reformation of an iron-sulfur cluster during catalysis by lipoyl synthase, Science, 2017, 358, 373–377, DOI:10.1126/science.aan4574.
- E. Mulliez, V. Duarte, S. Arragain, M. Fontecave and M. Atta, On the Role of Additional [4Fe-4S] Clusters with a Free Coordination Site in Radical-SAM Enzymes, Front. Chem., 2017, 5, 17, DOI:10.3389/fchem.2017.00017.
- F. Forouhar, S. Arragain, M. Atta, S. Gambarelli, J. M. Mouesca, M. Hussain, R. Xiao, S. Kieffer-Jaquinod, J. Seetharaman, T. B. Acton, G. T. Montelione, E. Mulliez, J. F. Hunt and M. Fontecave, Two Fe–S clusters catalyze sulfur insertion by radical-SAM methylthiotransferases, Nat. Chem. Biol., 2013, 9, 333–338, DOI:10.1038/nchembio.1229.
- S. Kimura and T. Suzuki, Iron–sulfur proteins responsible for RNA modifications, Biochim. Biophys. Acta, Mol. Cell Res., 2015, 1853, 1272–1283, DOI:10.1016/j.bbamcr.2014.12.010.
- O. A. Esakova, T. L. Grove, N. H. Yennawar, A. J. Arcinas, B. Wang, C. Krebs, S. C. Almo and S. J. Booker, Structural basis for tRNA methylthiolation by the radical SAM enzyme MiaB, Nature, 2021, 597, 566–570, DOI:10.1038/s41586-021-03904-6.
- J. C. Lachowicz, D. Lennox-Hvenekilde, N. Myling-Petersen, B. Salomonsen, G. Verkleij, C. G. Acevedo-Rocha, B. Caddell, L. S. Gronenberg, S. C. Almo, M. O. A. Sommer, H. J. Genee and T. L. Grove, Discovery of a Biotin Synthase That Utilizes an Auxiliary 4Fe–5S Cluster for Sulfur Insertion, J. Am. Chem. Soc., 2024, 146, 1860–1873, DOI:10.1021/jacs.3c05481.
- E. Barahona, J. A. Collantes-García, E. Rosa-Núñez, J. Xiong, X. Jiang, E. Jiménez-Vicente, C. Echávarri-Erasun, Y. Guo, L. M. Rubio and M. González-Guerrero, Azotobacter vinelandii scaffold protein NifU transfers iron to NifQ as part of the iron-molybdenum cofactor biosynthesis pathway for nitrogenase, J. Biol. Chem., 2024, 300, 107900, DOI:10.1016/j.jbc.2024.107900.
- J. A. Hernandez, L. Curatti, C. P. Aznar, Z. Perova, R. D. Britt and L. M. Rubio, Metal trafficking for nitrogen fixation: NifQ donates molybdenum to NifEN/NifH for the biosynthesis of the nitrogenase FeMo-cofactor, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 11679–11684, DOI:10.1073/pnas.0803576105.
- P. S. Garcia, S. Gribaldo and F. Barras, When iron and sulfur met on an anoxic planet and eventually made clusters essential for life, Biochim. Biophys. Acta, Mol. Cell Res., 2024, 1871, 119730, DOI:10.1016/j.bbamcr.2024.119730.
- M. A. Addo, A. M. Edwards and P. C. Dos Santos, Methods to Investigate the Kinetic Profile of Cysteine Desulfurases, in Fe–S Proteins: Methods and Protocols, ed. P. C. Dos Santos, Springer US, New York, NY, 2021, ch. 10, pp. 173–189 DOI:10.1007/978-1-0716-1605-5_10.
- L. Zheng, R. H. White, V. L. Cash and D. R. Dean, Mechanism for the Desulfurization of L-Cysteine Catalyzed by the nifS Gene Product, Biochemistry, 1994, 33, 4714–4720, DOI:10.1021/bi00181a031.
- J. Swindell and P. C. Dos Santos, Interactions with sulfur acceptors modulate the reactivity of cysteine desulfurases and define their physiological functions, Biochim. Biophys. Acta, Mol. Cell Res., 2024, 1871, 119794, DOI:10.1016/j.bbamcr.2024.119794.
- P. S. Garcia, F. D’Angelo, S. Ollagnier de Choudens, M. Dussouchaud, E. Bouveret, S. Gribaldo and F. Barras, An early origin of iron–sulfur cluster biosynthesis machineries before Earth oxygenation, Nat. Ecol. Evol., 2022, 6, 1564–1572, DOI:10.1038/s41559-022-01857-1.
- N. Maio and T. A. Rouault, Mammalian iron sulfur cluster biogenesis: From assembly to delivery to recipient proteins with a focus on novel targets of the chaperone and co-chaperone proteins, IUBMB Life, 2022, 74, 684–704, DOI:10.1002/iub.2593.
- U. Mühlenhoff, O. Stehling and R. Lill, Biogenesis of Mitochondrial Iron–Sulfur Proteins, in Iron-Sulfur Clusters: Biogenesis and Biochemistry, ed. S. Leimkühler, G. Schwarz, O. Lenz and O. Einsle, Wiley-VCH GmbH, 1st edn, 2025, ch. 3, vol. 1, pp. 57–86 DOI:10.1002/9783527843596.ch3.
- J. J. Braymer and A. J. Pierik, Iron–Sulfur Protein Maturation in the Cytosol of Eukaryotes, in Iron-Sulfur Clusters: Biogenesis and Biochemistry, ed. S. Leimkühler, G. Schwarz, O. Lenz and O. Einsle, Wiley-VCH GmbH, 1st edn, 2025, ch. 4, vol. 1, pp. 87–134 DOI:10.1002/9783527843596.ch4.
- J. Przybyla-Toscano, J. Couturier, C. Remacle and N. Rouhier, Occurrence, Evolution and Specificities of Iron-Sulfur Proteins and Maturation Factors in Chloroplasts from Algae, Int. J. Mol. Sci., 2021, 22, 3175, DOI:10.3390/ijms22063175.
- K. A. Black and P. C. Dos Santos, Shared-intermediates in the biosynthesis of thio-cofactors: Mechanism and functions of cysteine desulfurases and sulfur acceptors, Biochim. Biophys. Acta, Mol. Cell Res., 2015, 1853, 1470–1480, DOI:10.1016/j.bbamcr.2014.10.018.
- P. Olivieri, M. Klabes, J. C. Crack, A. Lehmann, S. P. Bennett, N. E. Le Brun and S. Leimkühler, Binding of IscU and TusA to different but competing sites of IscS influences the activity of IscS and directs sulfur to the respective biomolecular synthesis pathway, Microbiol. Spectrum, 2024, 12, e00949-24, DOI:10.1128/spectrum.00949-24.
- C. T. Lauhon, Requirement for IscS in Biosynthesis of All Thionucleosides in Escherichia coli, J. Bacteriol., 2002, 184, 6820–6829, DOI:10.1128/jb.184.24.6820-6829.2002.
- R. Thomé, A. Gust, R. Toci, R. Mendel, F. Bittner, A. Magalon and A. Walburger, A Sulfurtransferase Is Essential for Activity of Formate Dehydrogenases in Escherichia coli, J. Biol. Chem., 2012, 287, 4671–4678, DOI:10.1074/jbc.M111.327122.
- F. Cartini, W. Remelli, P. C. Dos Santos, J. Papenbrock, S. Pagani and F. Forlani, Mobilization of sulfane sulfur from cysteine desulfurases to the Azotobacter vinelandii sulfurtransferase RhdA, Amino Acids, 2011, 41, 141–150, DOI:10.1007/s00726-010-0529-z.
- R. Shi, A. Proteau, M. Villarroya, I. Moukadiri, L. Zhang, J. F. Trempe, A. Matte, M. E. Armengod and M. Cygler, Structural Basis for Fe–S Cluster Assembly and tRNA Thiolation Mediated by IscS Protein–Protein Interactions, PLoS Biol., 2010, 8, e1000354, DOI:10.1371/journal.pbio.1000354.
- C. Iannuzzi, M. Adrover, R. Puglisi, R. Yan, P. A. Temussi and A. Pastore, The role of zinc in the stability of the marginally stable IscU scaffold protein, Protein Sci., 2014, 23, 1208–1219, DOI:10.1002/pro.2501.
- J. Li, X. Ren, B. Fan, Z. Huang, W. Wang, H. Zhou, Z. Lou, H. Ding, J. Lyu and G. Tan, Zinc Toxicity and Iron-Sulfur Cluster Biogenesis in Escherichia coli, Appl. Environ. Microbiol., 2019, 85, e01967-18, DOI:10.1128/AEM.01967-18.
- S. P. Bennett, J. C. Crack, R. Puglisi, A. Pastore and N. E. L. Brun, Native mass spectrometric studies of IscSU reveal a concerted, sulfur-initiated mechanism of iron–sulfur cluster assembly, Chem. Sci., 2023, 14, 78–95, 10.1039/D2SC04169C.
- J. H. Kim, M. Tonelli and J. L. Markley, Disordered form of the scaffold protein IscU is the substrate for iron–sulfur cluster assembly on cysteine desulfurase, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 454–459, DOI:10.1073/pnas.1114372109.
- C. W. Lin, S. D. Oney-Hawthorne, S. T. Kuo, D. P. Barondeau and D. H. Russell, Mechanistic Insights into IscU Conformation Regulation for Fe–S Cluster Biogenesis Revealed by Variable Temperature Electrospray Ionization Native Ion Mobility Mass Spectrometry, Biochemistry, 2022, 61, 2733–2741, DOI:10.1021/acs.biochem.2c00429.
- C. E. Outten and T. V. O’Halloran, Femtomolar Sensitivity of Metalloregulatory Proteins Controlling Zinc Homeostasis, Science, 2001, 292, 2488–2492, DOI:10.1126/science.1060331.
- C. W. Lin, J. W. McCabe, D. H. Russell and D. P. Barondeau, Molecular Mechanism of ISC Iron–Sulfur Cluster Biogenesis Revealed by High-Resolution Native Mass Spectrometry, J. Am. Chem. Soc., 2020, 142, 6018–6029, DOI:10.1021/jacs.9b11454.
- B. Srour, S. Gervason, M. H. Hoock, B. Monfort, K. Want, D. Larkem, N. Trabelsi, G. Landrot, A. Zitolo, E. Fonda, E. Etienne, G. Gerbaud, C. S. Müller, J. Oltmanns, J. B. Gordon, V. Yadav, M. Kleczewska, M. Jelen, M. B. Toledano, R. Dutkiewicz, D. P. Goldberg, V. Schünemann, B. Guigliarelli, B. Burlat, C. Sizun and B. D’Autréaux, Iron Insertion at the Assembly Site of the ISCU Scaffold Protein Is a Conserved Process Initiating Fe–S Cluster Biosynthesis, J. Am. Chem. Soc., 2022, 144, 17496–17515, DOI:10.1021/jacs.2c06338.
- S. Gervason, R. Dutkiewicz, K. Want, R. Benazza, R. Mor-Gautier, A. Grabinska-Rogala, C. Sizun, O. Hernandez-Alba, S. Cianferani, B. Guigliarelli, B. Burlat and B. D’Autréaux, The ISC machinery assembles [2Fe–2S] clusters by formation and fusion of [1Fe–1S] precursors, Nat. Chem. Biol., 2025, 21, 767–778, DOI:10.1038/s41589-024-01818-8.
- E. N. Marinoni, J. S. de Oliveira, Y. Nicolet, E. C. Raulfs, P. Amara, D. R. Dean and J. C. Fontecilla-Camps, (IscS-IscU)2 Complex Structures Provide Insights into Fe2S2 Biogenesis and Transfer, Angew. Chem., Int. Ed., 2012, 51, 5439–5442, DOI:10.1002/ange.201201708.
- M. C. Unciuleac, K. Chandramouli, S. Naik, S. Mayer, B. H. Huynh, M. K. Johnson and D. R. Dean, In Vitro Activation of Apo-Aconitase Using a [4Fe–4S] Cluster-Loaded Form of the IscU [Fe–S] Cluster Scaffolding Protein, Biochemistry, 2007, 46, 6812–6821, DOI:10.1021/bi6026665.
- M. W. Foster, S. S. Mansy, J. Hwang, J. E. Penner-Hahn, K. K. Surerus and J. A. Cowan, A Mutant Human IscU Protein Contains a Stable [2Fe–2S]2+ Center of Possible Functional Significance, J. Am. Chem. Soc., 2000, 122, 6805–6806, DOI:10.1021/ja000800+.
- Y. Shimomura, H. Kamikubo, Y. Nishi, T. Masako, M. Kataoka, Y. Kobayashi, K. Fukuyama and Y. Takahashi, Characterization and Crystallization of an IscU-type Scaffold Protein with Bound [2Fe–2S] Cluster from the Hyperthermophile, Aquifex aeolicus, J. Biochem., 2007, 142, 577–586, DOI:10.1093/jb/mvm163.
- G. Wu, S. S. Mansy, S. P. Wu, K. K. Surerus, M. W. Foster and J. A. Cowan, Characterization of an Iron–Sulfur Cluster Assembly Protein (ISU1) from Schizosaccharomyces pombe, Biochemistry, 2002, 41, 5024–5032, DOI:10.1021/bi016073s.
- K. Kunichika, R. Nakamura, T. Fujishiro and Y. Takahashi, The Structure of the Dimeric State of IscU Harboring Two Adjacent [2Fe–2S] Clusters Provides Mechanistic Insights into Cluster Conversion to [4Fe–4S], Biochemistry, 2021, 60, 1569–1572, DOI:10.1021/acs.biochem.1c00112.
- M. Adrover, B. D. Howes, C. Iannuzzi, G. Smulevich and A. Pastore, Anatomy of an iron-sulfur cluster scaffold protein: Understanding the determinants of [2Fe–2S] cluster stability on IscU, Biochim. Biophys. Acta, Mol. Cell Res., 2015, 1853, 1448–1456, DOI:10.1016/j.bbamcr.2014.10.023.
- J. L. Markley, J. H. Kim, Z. Dai, J. R. Bothe, K. Cai, R. O. Frederick and M. Tonelli, Metamorphic protein IscU alternates conformations in the course of its role as the scaffold protein for iron–sulfur cluster biosynthesis and delivery, FEBS Lett., 2013, 587, 1172–1179, DOI:10.1016/j.febslet.2013.01.003.
- S. Sato, Y. Matsushima, M. Kanazawa, N. Tanaka, T. Fujishiro, K. Kunichika, R. Nakamura, H. Tomioka, K. Wada and Y. Takahashi, Evidence for dynamic in vivo interconversion of the conformational states of IscU during iron–sulfur cluster biosynthesis, Mol. Microbiol., 2021, 115, 807–818, DOI:10.1111/mmi.14646.
- K. G. Hoff, J. R. Cupp-Vickery and L. E. Vickery, Contributions of the LPPVK Motif of the Iron-Sulfur Template Protein IscU to Interactions with the Hsc66-Hsc20 Chaperone System, J. Biol. Chem., 2003, 278, 37582–37589, DOI:10.1074/jbc.M305292200.
- N. Tanaka, E. Yuda, T. Fujishiro, K. Hirabayashi, K. Wada and Y. Takahashi, Identification of IscU residues critical for de novo iron–sulfur cluster assembly, Mol. Microbiol., 2019, 112, 1769–1783, DOI:10.1111/mmi.14392.
- K. Chandramouli, M. C. Unciuleac, S. Naik, D. R. Dean, B. H. Huynh and M. K. Johnson, Formation and Properties of [4Fe–4S] Clusters on the IscU Scaffold Protein, Biochemistry, 2007, 46, 6804–6811, DOI:10.1021/bi6026659.
- B. Py and F. Barras, ISC-Based Fe–S Protein Biogenesis in Bacteria, in Iron-Sulfur Clusters: Biogenesis and Biochemistry, ed. S. Leimkühler, G. Schwarz, O. Lenz and O. Einsle, Wiley-VCH GmbH, 1st edn, 2025, ch. 1, vol. 1, pp. 1–24 DOI:10.1002/9783527843596.ch1.
- P. S. Garcia, F. Barras and S. Gribaldo, Components of iron–Sulfur cluster assembly machineries are robust phylogenetic markers to trace the origin of mitochondria and plastids, PLoS Biol., 2023, 21, e3002374, DOI:10.1371/journal.pbio.3002374.
- F. Bonomi, S. Iametti and A. Barbiroli, Protein interactions in the biological assembly of iron–sulfur clusters in Escherichia coli: Molecular and mechanistic aspects of the earliest assembly steps, IUBMB Life, 2022, 74, 723–732, DOI:10.1002/iub.2622.
- J. Yang, J. P. Bitoun and H. Ding, Interplay of IscA and IscU in Biogenesis of Iron-Sulfur Clusters, J. Biol. Chem., 2006, 281, 27956–27963, DOI:10.1074/jbc.M601356200.
- P. Olivieri, J. C. Crack, A. Lehmann, N. E. Le Brun and S. Leimkühler, CyaY and TusA regulate ISC- and SUF-mediated l-cysteine desulfurase activity, RSC Chem. Biol., 2024, 5, 1165–1176, 10.1039/D4CB00225C.
- Y. Takahashi and U. Tokumoto, A Third Bacterial System for the Assembly of Iron-Sulfur Clusters with Homologs in Archaea and Plastids, J. Biol. Chem., 2002, 277, 28380–28383, DOI:10.1074/jbc.C200365200.
- N. Yokoyama, C. Nonaka, Y. Ohashi, M. Shioda, T. Terahata, W. Chen, K. Sakamoto, C. Maruyama, T. Saito, E. Yuda, N. Tanaka, T. Fujishiro, T. Kuzuyama, K. Asai and Y. Takahashi, Distinct roles for U-type proteins in iron–sulfur cluster biosynthesis revealed by genetic analysis of the Bacillus subtilis sufCDSUB operon, Mol. Microbiol., 2018, 107, 688–703, DOI:10.1111/mmi.13907.
- C. A. Roberts, H. M. Al-Tameemi, A. A. Mashruwala, Z. Rosario-Cruz, U. Chauhan, W. E. Sause, V. J. Torres, W. J. Belden and J. M. Boyd, The Suf Iron-Sulfur Cluster Biosynthetic System Is Essential in Staphylococcus aureus, and Decreased Suf Function Results in Global Metabolic Defects and Reduced Survival in Human Neutrophils, Infect. Immun., 2017, 85, e00100-17, DOI:10.1128/iai.00100-17.
- G. Huet, M. Daffé and I. Saves, Identification of the Mycobacterium tuberculosis SUF Machinery as the Exclusive Mycobacterial System of [Fe–S] Cluster Assembly: Evidence for Its Implication in the Pathogen's Survival, J. Bacteriol., 2005, 187, 6137–6146, DOI:10.1128/jb.187.17.6137-6146.2005.
- R. K. Gogar, F. Carroll, J. V. Conte, M. Nasef, J. A. Dunkle and P. A. Frantom, The β-latch structural element of the SufS cysteine desulfurase mediates active site accessibility and SufE transpersulfurase positioning, J. Biol. Chem., 2023, 299, 102966, DOI:10.1016/j.jbc.2023.102966.
- R. K. Gogar, N. Chhikara, M. Vo, N. C. Gilbert, J. A. Dunkle and P. A. Frantom, The structure of the SufS–SufE complex reveals interactions driving protected persulfide transfer in iron-sulfur cluster biogenesis, J. Biol. Chem., 2024, 300, 107641, DOI:10.1016/j.jbc.2024.107641.
- A. G. Albrecht, D. J. A. Netz, M. Miethke, A. J. Pierik, O. Burghaus, F. Peuckert, R. Lill and M. A. Marahiel, SufU Is an Essential Iron-Sulfur Cluster Scaffold Protein in Bacillus subtilis, J. Bacteriol., 2010, 192, 1643–1651, DOI:10.1128/jb.01536-09.
- G. P. Riboldi, J. S. de Oliveira and J. Frazzon, Enterococcus faecalis SufU scaffold protein enhances SufS desulfurase activity by acquiring sulfur from its cysteine-153, Biochim. Biophys. Acta, Proteins Proteomics, 2011, 1814, 1910–1918, DOI:10.1016/j.bbapap.2011.06.016.
- B. P. Selbach, P. K. Pradhan and P. C. Dos Santos, Protected Sulfur Transfer Reactions by the Escherichia coli Suf System, Biochemistry, 2013, 52, 4089–4096, DOI:10.1021/bi4001479.
- I. Elchennawi, P. Carpentier, C. Caux, M. Ponge and S. Ollagnier de Choudens, Structural and Biochemical Characterization of Mycobacterium tuberculosis Zinc SufU-SufS Complex, Biomolecules, 2023, 13, 732, DOI:10.3390/biom13050732.
- J. D. Hudspeth, A. E. Boncella, E. T. Sabo, T. Andrews, J. M. Boyd and C. N. Morrison, Structural and Biochemical Characterization of Staphylococcus aureus Cysteine Desulfurase Complex SufSU, ACS Omega, 2022, 7, 44124–44133, DOI:10.1021/acsomega.2c05576.
- E. Sabo, C. Nelson, D. Huntoon, J. Yang, M. Kirk, S. J. George, B. Bennett, J. M. Boyd and R. C. Holz, Structural and mechanistic insights into the sulfur transfer protein SufU from Staphylococcus aureus, J. Inorg. Biochem., 2026, 277, 113205, DOI:10.1016/j.jinorgbio.2025.113205.
- P. C. Dos Santos, Fe–S assembly in Gram-positive bacteria, in Iron-Sulfur Cluster in Chemistry and Biology: Biochemistry, Biosynthesis and Human Diseases, ed. T. Rouault, De Gruyter, 2nd edn, 2017, ch. 5, vol. 2, pp. 97–113 DOI:10.1515/9783110479850-005.
- J. Pérard and S. Ollagnier de Choudens, Iron–sulfur clusters biogenesis by the SUF machinery: close to the molecular mechanism understanding, J. Biol. Inorg. Chem., 2018, 23, 581–596, DOI:10.1007/s00775-017-1527-3.
- E. S. Boyd, K. M. Thomas, Y. Dai, J. M. Boyd and F. W. Outten, Interplay between Oxygen and Fe–S Cluster Biogenesis: Insights from the Suf Pathway, Biochemistry, 2014, 53, 5834–5847, DOI:10.1021/bi500488r.
- Y. Dai and F. W. Outten, The E. coli SufS–SufE sulfur transfer system is more resistant to oxidative stress than IscS–IscU, FEBS Lett., 2012, 586, 4016–4022, DOI:10.1016/j.febslet.2012.10.001.
- R. K. Gogar, J. V. Conte, N. Chhikara, J. A. Dunkle and P. A. Frantom, A Role for Two Conserved Arginine Residues in Protected Persulfide Transfer by SufE-Dependent SufS Cysteine Desulfurases, Biochemistry, 2025, 64, 2467–2475, DOI:10.1021/acs.biochem.4c00705.
- F. W. Outten, M. J. Wood, F. M. Muñoz and G. Storz, The SufE Protein and the SufBCD Complex Enhance SufS Cysteine Desulfurase Activity as Part of a Sulfur Transfer Pathway for Fe–S Cluster Assembly in Escherichia coli, J. Biol. Chem., 2003, 278, 45713–45719, DOI:10.1074/jbc.M308004200.
- G. Layer, S. A. Gaddam, C. N. Ayala-Castro, S. Ollagnier de Choudens, D. Lascoux, M. Fontecave and F. W. Outten, SufE Transfers Sulfur from SufS to SufB for Iron-Sulfur Cluster Assembly, J. Biol. Chem., 2007, 282, 13342–13350, DOI:10.1074/jbc.M608555200.
- K. Hirabayashi, E. Yuda, N. Tanaka, S. Katayama, K. Iwasaki, T. Matsumoto, G. Kurisu, F. W. Outten, K. Fukuyama, Y. Takahashi and K. Wada, Functional Dynamics Revealed by the Structure of the SufBCD Complex, a Novel ATP-binding Cassette (ABC) Protein That Serves as a Scaffold for Iron-Sulfur Cluster Biogenesis, J. Biol. Chem., 2015, 290, 29717–29731, DOI:10.1074/jbc.M115.680934.
- K. Wada, N. Sumi, R. Nagai, K. Iwasaki, T. Sato, K. Suzuki, Y. Hasegawa, S. Kitaoka, Y. Minami, F. W. Outten, Y. Takahashi and K. Fukuyama, Molecular Dynamism of Fe–S Cluster Biosynthesis Implicated by the Structure of the SufC2–SufD2 Complex, J. Mol. Biol., 2009, 387, 245–258, DOI:10.1016/j.jmb.2009.01.054.
- H. K. Chahal and F. W. Outten, Separate FeS scaffold and carrier functions for SufB2C2 and SufA during in vitro maturation of [2Fe2S] Fdx, J. Inorg. Biochem., 2012, 116, 126–134, DOI:10.1016/j.jinorgbio.2012.06.008.
- G. Veronesi, J. Pérard, M. Clémancey, C. Gerez, Y. Duverger, I. Kieffer, F. Barras, S. Gambarelli, G. Blondin and S. Ollagnier De Choudens, Multimodal Spectroscopic Analysis of the Fe–S Clusters of the as-Isolated Escherichia coli SufBC2D Complex, Inorg. Chem., 2024, 63, 8730–8738, DOI:10.1021/acs.inorgchem.4c00304.
- I. Elchennawi, C. E. Fisher, F. W. Outten and S. Ollagnier de Choudens, Unraveling the Complexity of the Suf-Based Fe–S Biogenesis, in Iron-Sulfur Clusters: Biogenesis and Biochemistry, ed. S. Leimkühler, G. Schwarz, O. Lenz and O. Einsle, Wiley-VCH GmbH, 1st edn, 2025, ch. 2, vol. 1, pp. 25–56 DOI:10.1002/9783527843596.ch2.
- S. Wollers, G. Layer, R. Garcia-Serres, L. Signor, M. Clemancey, J.-M. Latour, M. Fontecave and S. Ollagnier De Choudens, Iron-Sulfur (Fe–S) Cluster Assembly: The SufBCD Complex Is a New Type of Fe–S Scaffold with a Flavin Redox Cofactor, J. Biol. Chem., 2010, 285, 23331–23341, DOI:10.1074/jbc.M110.127449.
- N. Chhikara, G. Timilsina, Y. Wang, D. Reasons, F. W. Outten and P. A. Frantom, Characterization of a mechanistic connection between persulfide transfer and ATP hydrolysis in the SufBC2D scaffold of the Suf Fe–S cluster assembly pathway, Metallomics, 2025, 17, mfaf029, DOI:10.1093/mtomcs/mfaf029.
- B. Selbach, E. Earles and P. C. Dos Santos, Kinetic Analysis of the Bisubstrate Cysteine Desulfurase SufS from Bacillus subtilis, Biochemistry, 2010, 49, 8794–8802, DOI:10.1021/bi101358k.
- G. P. Riboldi, H. Verli and J. Frazzon, Structural studies of the Enterococcus faecalis SufU [Fe–S] cluster protein, BMC Biochem., 2009, 10, 3, DOI:10.1186/1471-2091-10-3.
- B. P. Selbach, A. H. Chung, A. D. Scott, S. J. George, S. P. Cramer and P. C. Dos Santos, Fe–S Cluster Biogenesis in Gram-positive Bacteria: SufU Is a Zinc-Dependent Sulfur Transfer Protein, Biochemistry, 2014, 53, 152–160, DOI:10.1021/bi4011978.
- T. Fujishiro, T. Terahata, K. Kunichika, N. Yokoyama, C. Maruyama, K. Asai and Y. Takahashi, Zinc-Ligand Swapping Mediated Complex Formation and Sulfur Transfer between SufS and SufU for Iron–Sulfur Cluster Biogenesis in Bacillus subtilis, J. Am. Chem. Soc., 2017, 139, 18464–18467, DOI:10.1021/jacs.7b11307.
- T. Terahata, Y. Shimada, C. Maki, S. Muroga, R. Sakurai, K. Kunichika and T. Fujishiro, Cysteine-Persulfide Sulfane Sulfur-Ligated Zn Complex of Sulfur-Carrying SufU in the SufCDSUB System for Fe–S Cluster Biosynthesis, Inorg. Chem., 2024, 63, 19607–19618, DOI:10.1021/acs.inorgchem.4c02654.
- D. Ma, H. Yao, Y. Liu, W. Gong, Y. Zhao, R. Wang, C. Wu, W. Wang and H. Wang, The reduced interaction between SufS and SufU in Mycoplasma penetrans results in diminished sulfotransferase activity, Int. J. Biol. Macromol., 2025, 284, 138181, DOI:10.1016/j.ijbiomac.2024.138181.
- A. Perálvarez-Marín, E. Baranowski, P. Bierge, O. Q. Pich and H. Lebrette, Metal utilization in genome-reduced bacteria: Do human mycoplasmas rely on iron?, Comput. Struct. Biotechnol. J., 2021, 19, 5752–5761, DOI:10.1016/j.csbj.2021.10.022.
- S. Großhennig, T. Ischebeck, J. Gibhardt, J. Busse, I. Feussner and J. Stülke, Hydrogen sulfide is a novel potential virulence factor of Mycoplasma pneumoniae: characterization of the unusual cysteine desulfurase/desulfhydrase HapE, Mol. Microbiol., 2016, 100, 42–54, DOI:10.1111/mmi.13300.
- J. S. Martin del Campo, S. Shaybani, D. R. Dean and P. C. Dos Santos, NifUS is a two-component toolbox involved in assembling Fe–S clusters associated with nitrogen fixation and beyond, in Iron-Sulfur Clusters: Biogenesis and Biochemistry, ed. S. Leimkühler, G. Schwarz, O. Lenz, and O. Einsle, Wiley-VCH GmbH, 1st edn, 2025, ch. 9, vol. 1, pp. 257–286 DOI:10.1002/9783527843596.ch9.
- A. D. Smith, G. N. L. Jameson, P. C. Dos Santos, J. N. Agar, S. Naik, C. Krebs, J. Frazzon, D. R. Dean, B. H. Huynh and M. K. Johnson, NifS-mediated assembly of [4Fe–4S] clusters in the N- and C-terminal domains of the NifU scaffold protein, Biochemistry, 2005, 44, 12955–12969, DOI:10.1021/bi051257i.
- J. N. Agar, P. Yuvaniyama, R. F. Jack, V. L. Cash, A. D. Smith, D. R. Dean and M. K. Johnson, Modular organization and identification of a mononuclear iron-binding site within the NifU protein, J. Biol. Inorg. Chem., 2000, 5, 167–177, DOI:10.1007/s007750050361.
- W. Fu, R. F. Jack, T. V. Morgan, D. R. Dean and M. K. Johnson, nifU gene product from Azotobacter vinelandii is a homodimer that contains two identical [2Fe–2S] clusters, Biochemistry, 1994, 33, 13455–13463, DOI:10.1021/bi00249a034.
- M. R. Jacobson, V. L. Cash, M. C. Weiss, N. F. Laird, W. E. Newton and D. R. Dean, Biochemical and genetic analysis of the nifUSVWZM cluster from Azotobacter vinelandii, Mol. Gen. Genet., 1989, 219, 49–57, DOI:10.1007/BF00261156.
- C. Kennedy and D. Dean, The nifU, nifS and nifV gene products are required for activity of all three nitrogenases of Azotobacter vinelandii, Mol. Gen. Genet., 1992, 231, 494–498, DOI:10.1007/BF00292722.
- A. Pérez-González, E. Jimenez-Vicente, J. Gies-Elterlein, A. Salinero-Lanzarote, Z. Y. Yang, O. Einsle, L. C. Seefeldt and D. R. Dean, Specificity of NifEN and VnfEN for the assembly of nitrogenase active-site cofactors in Azotobacter vinelandii, mBio, 2021, 12, e01568-21, DOI:10.1128/mbio.01568-21.
- J. W. Olson, J. N. Agar, M. K. Johnson and R. J. Maier, Characterization of the NifU and NifS Fe–S cluster formation proteins essential for viability in Helicobacter pylori, Biochemistry, 2000, 39, 16213–16219, DOI:10.1021/bi001744s.
- U. Tokumoto, S. Kitamura, K. Fukuyama and Y. Takahashi, Interchangeability and distinct properties of bacterial Fe–S cluster assembly systems: functional replacement of the isc and suf operons in Escherichia coli with the nifSU-like operon from Helicobacter pylori, J. Biochem., 2004, 136, 199–209, DOI:10.1093/jb/mvh104.
- V. Ali, Y. Shigeta, U. Tokumoto, Y. Takahashi and T. Nozaki, An intestinal parasitic protist, Entamoeba histolytica, possesses a non-redundant nitrogen fixation-like system for iron–sulfur cluster assembly under anaerobic conditions, J. Biol. Chem., 2004, 279, 16863–16874, DOI:10.1074/jbc.M313314200.
- E. Nývltová, R. Šuták, K. Harant, M. Šedinová, I. Hrdý, J. Pačes, Č. Vlček and J. Tachezy, NIF-type iron–sulfur cluster assembly system is duplicated and distributed in the mitochondria and cytosol of Mastigamoeba balamuthi, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 7371–7376, DOI:10.1073/pnas.1219590110.
- M. Dussouchaud, M. Martinez-Carranza, P. S. Garcia, M. Clémancey, G. Blondin, J. M. Betton, A. Haouz, S. Gribaldo, S. Ollagnier De Choudens, L. Sauguet, A. Mechaly and F. Barras, Ancestral [Fe–S] biogenesis system SMS has a unique mechanism of cluster assembly and sulfur utilization, PLoS Biol., 2025, 23, e3003223, DOI:10.1371/journal.pbio.3003223.
- C. Zhao, N. Shao, N. Bryer, H. Chen, W. B. Whitman, D. J. Vinyard and Y. Hu, A minimal SufB2C2 complex functions as a [4Fe–4S] cluster scaffold in methanogenic archaea, Microbiol. Spectrum, 2025, 13, e02134-25, DOI:10.1128/spectrum.02134-25.
- J. Jin, T. Sato and H. Atomi, SmsB-SmsC machinery in Thermococcus kodakarensis functions as an archaeal scaffold mediating Fe–S cluster assembly, Appl. Environ. Microbiol., 2025, 91, e01438-25, DOI:10.1128/aem.01438-25.
- R. Hidese, T. Inoue, T. Imanaka and S. Fujiwara, Cysteine desulphurase plays an important role in environmental adaptation of the hyperthermophilic archaeon Thermococcus kodakarensis, Mol. Microbiol., 2014, 93, 331–345, DOI:10.1111/mmi.12662.
- Y. Liu, M. Sieprawska-Lupa, W. B. Whitman and R. H. White, Cysteine is not the sulfur source for iron-sulfur cluster and methionine biosynthesis in the methanogenic archaeon Methanococcus maripaludis, J. Biol. Chem., 2010, 285, 31923–31929, DOI:10.1074/jbc.M110.152447.
- T. M. Deere, D. Prakash, F. H. Lessner, E. C. Duin and D. J. Lessner, Methanosarcina acetivorans contains a functional ISC system for iron-sulfur cluster biogenesis, BMC Microbiol., 2020, 20, 323, DOI:10.1186/s12866-020-02014-z.
- J. Saini, T. M. Deere and D. J. Lessner, The minimal SUF system is not required for Fe–S cluster biogenesis in the methanogenic archaeon Methanosarcina acetivorans, Sci. Rep., 2023, 13, 15120, DOI:10.1038/s41598-023-42400-x.
- G. Wächtershäuser, Groundworks for an evolutionary biochemistry: The iron-sulphur world, Prog. Biophys. Mol. Biol., 1992, 58, 85–201, DOI:10.1016/0079-6107(92)90022-X.
- T. Jones, Z. Fang, S. M. Spigelmyer and P. C. Dos Santos, Direct assembly of the iron-sulfur cluster on quinolinate synthase NadA by the Bacillus subtilis cysteine sulfurtransferase NifS, J. Inorg. Biochem., 2026, 274, 113095, DOI:10.1016/j.jinorgbio.2025.113095.
- M. Nuth and J. A. Cowan, Iron–sulfur cluster biosynthesis: characterization of IscU–IscS complex formation and a structural model for sulfide delivery to the [2Fe–2S] assembly site, J. Biol. Inorg. Chem., 2009, 14, 829–839, DOI:10.1007/s00775-009-0495-7.
| This journal is © The Royal Society of Chemistry 2026 |