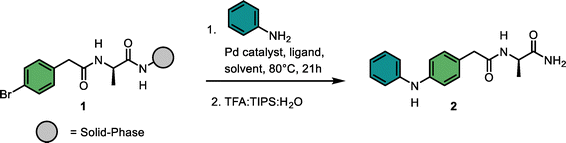Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Integration of palladium-catalyzed C–N coupling into self-encoded libraries for accelerated hit discovery
Edith
van der Nol†
 ab,
Zhenshuo
Luo†
ab,
Zhenshuo
Luo†
 a,
Qing Qing
Gao†
a,
Nils Alexander
Haupt
c,
Sebastian
Böcker
c and
Sebastian
Pomplun
a,
Qing Qing
Gao†
a,
Nils Alexander
Haupt
c,
Sebastian
Böcker
c and
Sebastian
Pomplun
 *ab
*ab
aLACDR, Leiden University, Leiden, 2333 CC, The Netherlands. E-mail: s.j.pomplun@lacdr.leidenuniv.nl
bOncode Institute, Utrecht, 3521 AL, The Netherlands
cChair for Bioinformatics, Institute for Computer Science, Friedrich Schiller University Jena, Jena, 07737, Germany
First published on 3rd March 2026
Abstract
Affinity screenings with encoded libraries are transformative tools for rapid hit discovery from vast compound collections. Yet the adaptation of established chemical reactions to DNA-encoded libraries (DELs) remains challenging due to DNA-compatibility constraints and mismatches between barcode and chemical structure in case of incomplete reactions or side product formation. Recently, we introduced self-encoded libraries (SELs) as a barcode-free alternative to DELs. The SEL platform offers unmatched flexibility in reaction conditions and decodes screening hits directly from their chemical structure, avoiding the problem of mismatched barcode-compound pairs. Here, we expand the SEL platform to Buchwald–Hartwig aminations, enabling the construction of new high diversity SELs. We performed a thorough reaction condition optimization and tested a scope of >170 different building blocks. We adapted our automated MS/MS-based decoding methodology SIRIUS-COMET to the resulting scaffolds, enabling accurate compound decoding from complex mixtures. A 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 725-member library was synthesized and screened all at once against carbonic anhydrase IX (CAIX), resulting in robust enrichment of hits with specific building block patterns and yielding several nanomolar-affinity binders. This work showcases the seamless integration of palladium-catalyzed cross-couplings into SELs, expanding the chemical space of this technology and accelerating hit discovery with high synthetic versatility.
725-member library was synthesized and screened all at once against carbonic anhydrase IX (CAIX), resulting in robust enrichment of hits with specific building block patterns and yielding several nanomolar-affinity binders. This work showcases the seamless integration of palladium-catalyzed cross-couplings into SELs, expanding the chemical space of this technology and accelerating hit discovery with high synthetic versatility.
Introduction
The identification of bioactive molecules is a crucial step in drug discovery, relying heavily on access to large and diverse collections of small molecules. Traditionally, high-throughput screening (HTS) involves the parallel screening of individual compounds but can be resource-intensive. Encoded-library techniques have emerged as a powerful alternative technology, enabling the rapid and cost-effective screening of millions or even billions of compounds simultaneously.1–3 Among these, DNA-encoded libraries (DELs) in particular has become a widespread technology in both industry and academia, leading to the discovery of numerous bioactive compounds across a broad range of targets.1,2,4–6Obtaining high-quality combinatorial libraries requires robust, high-yielding chemical reactions to maintain synthetic fidelity. Incorporating a broad range of chemical transformations grants access to a large chemical space and, in turn, increases the probability of discovering novel bioactive molecules. DELs have inherent challenges due to the synthesis of small molecules on DNA strands and although the number of DEL-compatible reactions continues to grow, it remains challenging to find robust conditions for several chemical transformations.7–9 Among the palladium-catalyzed cross-coupling reactions described for DELs, Suzuki–Miyaura, Sonogashira and Heck couplings have been successfully implemented under mild, DNA-compatible conditions.10–13
The Buchwald–Hartwig amination has become a cornerstone transformation in medicinal chemistry due to the prevalence of arylated amines in pharmaceuticals and natural products.10,14 However, its on-DNA implementation is hindered by several DNA degradation conditions, such as high temperatures, palladium-catalyst interactions, and exposure to organic solvents.12,15–18 Additionally, the presence of air or water can lead to catalyst inactivation, further complicating its use in DELs. The reported examples of Buchwald–Hartwig on DNA often suffer from low yields and limited substrate scope.15,19–23
In principle, the presence of incomplete reaction products is not inherently problematic for affinity selections, as also shown by Cernak et al., who used nanoscale synthesized libraries in affinity ranking experiments.24
However, in encoded libraries, such incompleteness leads to a mismatch between the molecular barcode and the corresponding truncated or deleted compound, resulting in misleading outcomes during hit decoding. Although strategies to mitigate these issues are under development, they inevitably introduce additional complexity to library synthesis.25
Recently, we introduced self-encoded libraries (SELs) as a barcode-free alternative to DELs.26 SELs are prepared by solid-phase synthesis and therefore offer exceptional flexibility in reaction conditions and catalyst compatibility with broad tolerance for organic reagents and elevated temperatures. In the SEL discovery workflow, combinatorial libraries are screened in affinity selections against a protein of interest. Enriched hit compounds are detected using tandem mass spectrometry (MS/MS) and decoded by an automated software (SIRIUS-COMET). SIRIUS-COMET enables the screening of very large libraries (up to 750![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 members) by accurately annotating compound structures from their fragmentation spectra, even in the presence of hundreds of isobaric compounds. Given the absence of DNA-compatibility constraints for library synthesis, the SEL platform enables the implementation of established medicinal chemistry transformations, including those challenging for DELs. Since the hit compounds are decoded directly from their own structure, there are no problems related to barcode-compound mismatches.
000 members) by accurately annotating compound structures from their fragmentation spectra, even in the presence of hundreds of isobaric compounds. Given the absence of DNA-compatibility constraints for library synthesis, the SEL platform enables the implementation of established medicinal chemistry transformations, including those challenging for DELs. Since the hit compounds are decoded directly from their own structure, there are no problems related to barcode-compound mismatches.
In this study, we implement the Buchwald–Hartwig reaction in our SEL platform. Building on previous reports on solid-phase based Buchwald–Hartwig aminations,27–29 we performed a thorough reaction condition optimization and a broad substrate scope. We expanded the SIRIUS-COMET decoding workflow to account for the resulting scaffolds and validated the annotation fidelity on ∼600 compounds. With the building blocks validated in the substrate scope, we built a 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 725 membered SEL and benchmarked the whole discovery workflow in affinity selections against carbonic anhydrase IX (CAIX). Robust enrichment of hits with experimentally validated nanomolar affinities were achieved, showcasing the compatibility of the SEL platform with this new library design using Buchwald–Hartwig aminations.
725 membered SEL and benchmarked the whole discovery workflow in affinity selections against carbonic anhydrase IX (CAIX). Robust enrichment of hits with experimentally validated nanomolar affinities were achieved, showcasing the compatibility of the SEL platform with this new library design using Buchwald–Hartwig aminations.
Results
Optimization of synthesis conditions for solid-phase Buchwald–Hartwig reactions
We started the condition optimization with the model reaction between solid-phase bound aryl bromide 1 and aniline (Table 1). For determining the conversion efficiency, we cleaved each compound from the solid support and analyzed the resulting mixture by LC-MS.25 In a first iteration we kept the Pd source (PdCl2), base (K2CO3) and solvent composition (DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O, 9
H2O, 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) constant and tested a set of different phosphine-based ligands. XPhos and tBuXPhos resulted in moderate conversions of 43% and 40%, respectively, to 2, while the other ligands resulted in poor yields (Table 1, entries 1–6). Changing the base to Na2CO3, DBU or KOtBu, decreased the reaction conversion (Table 1, entries 7–9). Notably, by switching the palladium source to Pd(OAc)2 a substantially better conversion was observed (67%), and could even be improved (80%), using higher catalyst loading (Table 1, entries 10–11). A small solvent screen showed that DMF, dioxane, toluene and anisole gave all comparably good conversions (Table 1, entries 12–14). Based on the conversions and green chemistry guidelines, we prioritized the most environmentally benign option, anisole.30 Performing solid-phase reaction under fully inert conditions, especially when aiming at a combinatorial set up, is very challenging. As a feasible set up we performed the reactions in sealed Eppendorf tubes, purged with N2. Compared to performing the reactions under air, the reproducibility of the reaction outcomes improved substantially with this set up (data not shown).
1) constant and tested a set of different phosphine-based ligands. XPhos and tBuXPhos resulted in moderate conversions of 43% and 40%, respectively, to 2, while the other ligands resulted in poor yields (Table 1, entries 1–6). Changing the base to Na2CO3, DBU or KOtBu, decreased the reaction conversion (Table 1, entries 7–9). Notably, by switching the palladium source to Pd(OAc)2 a substantially better conversion was observed (67%), and could even be improved (80%), using higher catalyst loading (Table 1, entries 10–11). A small solvent screen showed that DMF, dioxane, toluene and anisole gave all comparably good conversions (Table 1, entries 12–14). Based on the conversions and green chemistry guidelines, we prioritized the most environmentally benign option, anisole.30 Performing solid-phase reaction under fully inert conditions, especially when aiming at a combinatorial set up, is very challenging. As a feasible set up we performed the reactions in sealed Eppendorf tubes, purged with N2. Compared to performing the reactions under air, the reproducibility of the reaction outcomes improved substantially with this set up (data not shown).
| Entry | Pd catalyst (mol%) | Ligand (mol%) | Base (2 eq.) | Solvent (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) 1) |
Conversionb (%) |
|---|---|---|---|---|---|
a Reaction conditions: 1 (5 µmol, 1 eq.), aniline (10 µmol, 2 eq.), base (10 µmol, 2 eq.), solvent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O (9 H2O (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 100 µL), 80 °C, 21 h.
b Conversion was determined by LC-MS analysis thought peak integration of UV-absorption at 254 nm. 1, 100 µL), 80 °C, 21 h.
b Conversion was determined by LC-MS analysis thought peak integration of UV-absorption at 254 nm.
|
|||||
| 1 | PdCl2 (10) | Xphos (20) | K2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
43 |
| 2 | PdCl2 (10) | Johnphos (20) | K2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
11 |
| 3 | PdCl2 (10) | CyJohnphos (20) | K2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
2 |
| 4 | PdCl2 (10) | Davephos (20) | K2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
26 |
| 5 | PdCl2 (10) | tBuXphos (20) | K2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
40 |
| 6 | PdCl2 (10) | Sphos (20) | K2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
22 |
| 7 | PdCl2 (10) | Xphos (20) | Na2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :H2O :H2O |
26 |
| 8 | PdCl2 (10) | Xphos (20) | DBU | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
0 |
| 9 | PdCl2 (10) | Xphos (20) | KOtBu | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
17 |
| 10 | Pd(OAc)2 (10) | Xphos (20) | K2CO3 | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
67 |
| 11 | Pd(OAc) 2 (20) | Xphos (40) | K 2 CO 3 |
DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) H
2
O H
2
O
|
80 |
| 12 | Pd(OAc)2 (20) | Xphos (40) | K2CO3 | Toluene![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
78 |
| 13 | Pd(OAc)2 (20) | Xphos (40) | K2CO3 | Dioxane![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
76 |
| 14 | Pd(OAc) 2 (20) | Xphos (40) | K 2 CO 3 |
Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) H
2
O H
2
O
|
83 |
To evaluate the performance of the established conditions with aliphatic amines, we focused on the reaction with 1-methylpiperazine as a model substrate, given that piperidine and piperazine are among the most prevalent nitrogen heterocycles in FDA-approved drugs.31 When testing the reaction between model substrate 3 with 1-methylpiperazine under the optimized conditions for aniline, we obtained a conversion to product of only 3% (Table 2, entry 1) which required further improvement of reaction conditions. We first screened 10 different phosphine ligands (Table 2, entries 1–10). Nine of the ten ligands resulted in conversions between 0 and 7% and only tBuXPhos afforded a moderate conversion of 30% (Table 2, entry 10). Increasing the catalyst loading further improved the conversion to 41% (Table 2, entry 11). Another minor improvement to 46% was obtained by using Pd2(dba)3 (Table 2, entries 12–14). A subsequent solvent screening using different protic and aprotic solvents revealed toluene to be the optimal choice, leading to conversions of 61% (Table 2, entries 15–19). A similar performance was retained at slightly reduced catalyst loadings (Table 2, entry 20). While these results confirm the more challenging nature of Buchwald–Hartwig reactions for combinatorial library synthesis, the optimized conditions gave conversions over 50%, making it suitable for integration in our SEL platform.
| Entry | Pd catalyst (mol%) | Ligand (mol%) | Solvent (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) 1) |
Conversionb (%) |
|---|---|---|---|---|
a Reaction conditions: 1 (10 µmol, 1 eq.), 1-methylpiperazine (20 µmol, 2 eq.), solvent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O (9 H2O (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 300 µL), K2CO3 (20 µmol, 2 eq.), 80 °C, 18 h.
b Conversion was determined by LC-MS analysis through peak integration of UV-absorption at 254 nm. 1, 300 µL), K2CO3 (20 µmol, 2 eq.), 80 °C, 18 h.
b Conversion was determined by LC-MS analysis through peak integration of UV-absorption at 254 nm.
|
||||
| 1 | Pd(OAc)2 (20) | XPhos (40) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
3 |
| 2 | Pd(OAc)2 (20) | BINAP (20) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
0 |
| 3 | Pd(OAc)2 (20) | XantPhos (20) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
1 |
| 4 | Pd(OAc)2 (20) | SPhos (40) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
2 |
| 5 | Pd(OAc)2 (20) | RuPhos (40) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
4 |
| 6 | Pd(OAc)2 (20) | JohnPhos (40) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
7 |
| 7 | Pd(OAc)2 (20) | CyJohnPhos (40) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
1 |
| 8 | Pd(OAc)2 (20) | DavePhos (40) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
2 |
| 9 | Pd(OAc)2 (20) | BrettPhos (40) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
4 |
| 10 | Pd(OAc) 2 (20) | tBuXPhos (40) |
Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) H
2
O H
2
O
|
30 |
| 11 | Pd(OAc) 2 (50) | tBuXPhos (100) |
Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) H
2
O H
2
O
|
41 |
| 12 | PdCl2 (50) | tBuXPhos (100) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
18 |
| 13 | APC (25) | tBuXPhos (100) | Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
29 |
| 14 | Pd 2 (dba) 3 (25) | tBuXPhos (100) |
Anisole![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) H
2
O H
2
O
|
46 |
| 15 | Pd2(dba)3 (25) | tBuXPhos (100) | DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
59 |
| 16 | Pd2(dba)3 (25) | tBuXPhos (100) | Dioxane![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
7 |
| 17 | Pd2(dba)3 (25) | tBuXPhos (100) | THF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
14 |
| 18 | Pd2(dba)3 (25) | tBuXPhos (100) | Dibutylether![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O H2O |
51 |
| 19 | Pd 2 (dba) 3 (25) | tBuXPhos (100) |
Toluene![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) H
2
O H
2
O
|
61 |
| 20 | Pd 2 (dba) 3 (20) | tBuXPhos (80) |
Toluene![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) H
2
O H
2
O
|
60 |
Building block scope
With the optimized conditions for aromatic and aliphatic amines, we evaluated the reaction scope on 22 different solid-phase bound aryl bromides, reacting each with aniline and 1-methylpiperazine. Looking ahead to the planned combinatorial library synthesis, we defined a threshold of 50% conversion to consider a building block suitable for library incorporation. From the 22 aryl bromides investigated (Fig. 1), nine resulted in conversions >50% when reacted with aniline and five when reacted with 1-methylpiperazine. The majority of substituted benzoic acids (5a–f, 6a–f) gave unsatisfactory results. Bromophenylacetic acid variants (5g, 6g) and bromophenylpropanoic acids (5i–j, 6i–j) were well-tolerated. The bicyclic structures (5k–n, 6k–n) showed a general preference for the coupling with aromatic amines compared to the reaction with aliphatic amines. Furthermore, bromophenylalanine derivatives bearing different carboxylic acid decorators (5r–v, 6r–v) generally resulted in reasonable conversions.Next, we tested the reaction between substrate 1 and 156 aromatic and aliphatic amine building blocks (Fig. 2). A broad range of aromatic amines bearing alkyl, alkoxy, fluoro, trifluoromethyl, nitrile, ester, and acetyl substituents were converted to the desired products with conversions >50%. In contrast, amines containing alkyne, carboxylic acid, sulfonic acid, free sulfonamide, and nitro substituents mostly resulted in low conversions. From the heterocyclic amines, 2-aminopyridine and 2-aminopyrazine underwent efficient coupling, whereas the 3-amino- and 4-aminopyridine isomers or substituted derivatives gave poor conversion, likely due to their reduced nucleophilicity. Several piperazine analogs and a few other cyclic- and primary aliphatic amines met the conversion threshold. Among the variants that displayed lower conversions, steric hindrance seemed to be the most prominent common feature. Overall, 75 building blocks showed conversion over 50%, providing a set of diverse building blocks for library synthesis.
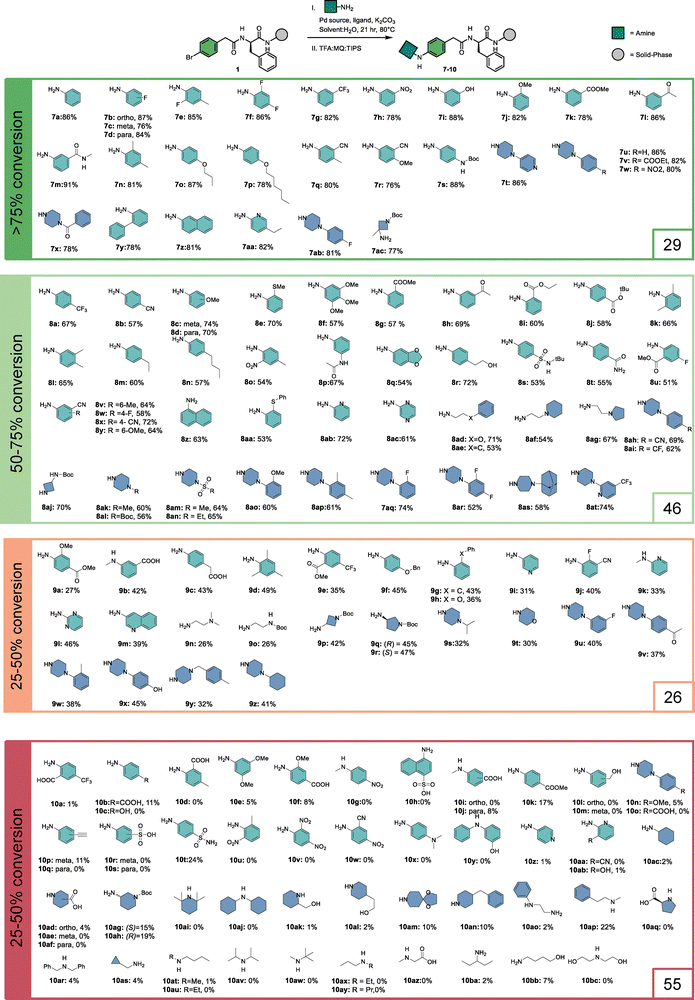 | ||
Fig. 2 Scope of 156 amines. a![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) The investigated amines are categorized based on conversion. b The investigated amines are categorized based on conversion. b![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) The turquoise colored amine used the reaction conditions from Table 1, entry 14. The dark blue amine used the reaction conditions from Table 2, entry 20. c The turquoise colored amine used the reaction conditions from Table 1, entry 14. The dark blue amine used the reaction conditions from Table 2, entry 20. c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Conversion was determined by LC-MS analysis through peak integration of UV-absorption at 254 nm. Conversion was determined by LC-MS analysis through peak integration of UV-absorption at 254 nm. | ||
Library decoding
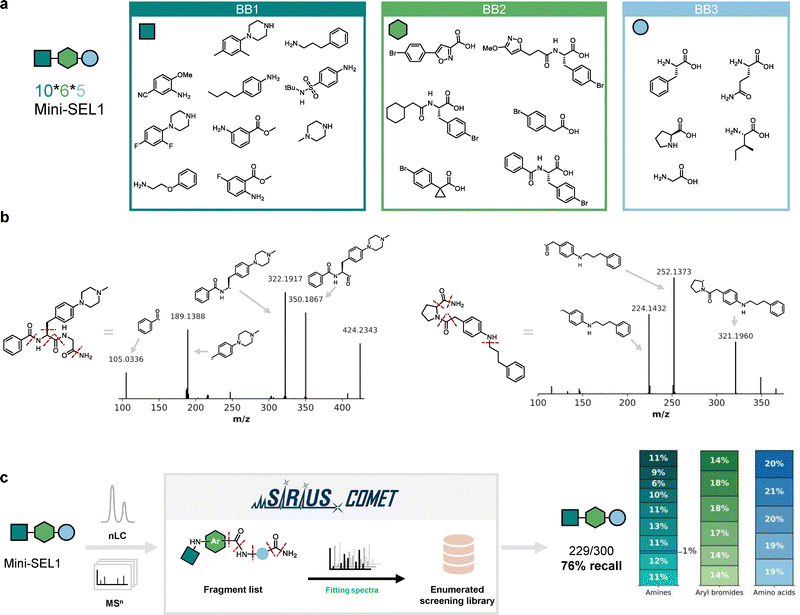
The SEL workflow entails screening a large library all at once against a target of interest via affinity selection. As opposed to classical AS-MS,32–35 no particular attention to the presence of isobaric compounds in a library is needed. This is possible thanks to our previously developed computational tool, SIRIUS-COMET, capable of annotating compounds from MS/MS data and annotate their structures.26In order to evaluate the decoding software and optimize parameters for this new Buchwald–Hartwig based library, we first synthesized two defined test libraries of 300 compounds each (miniSEL1 and miniSEL2, Fig. 3a and Fig. S1). Each library consists of a scaffold with three variable positions: an amino acid building blocks (BB3) coupled to an aryl bromides (BB2) and subsequently reacted with amines (BB1) using the established conditions. Each library was prepared via split-and-pool synthesis, bulk purified and then measured as a mixture by nanoLC–MS/MS. First, we analyzed a few selected spectra manually to observe overall trends in fragmentation of the library compounds. The most prominent and consistent fragmentation observed across scaffolds is the amide bond between the amino acid building block and the aryl bromide (see Fig. 3b for selected examples). Also, the terminal amide resulting from solid-phase synthesis was fragmented in all selected compounds. The C–N bond formed by the Buchwald–Hartwig coupling fragmented only sporadically with a low signal intensity. Increasing the fragmentation energy improved fragment intensity, but not enough to utilize it as a consistent fragment for library decoding (Fig. S2).
For the full decoding of our libraries, we used SIRIUS-COMET,26 a software platform that enables accurate annotation and decoding of combinatorial libraries and at the same time effectively reducing the number of false positive annotations. A fragment list based on the selected most prominent scaffold fragmentations is used to filter MS/MS scans based on a minimum number of matching peaks. The remaining spectra were then annotated using EPIMETHEUS, the ranking and substructure annotation module within the SIRIUS software, using the enumerated library database (represented as SMILES).
Analysis of the MS/MS spectra of miniSEL1 and 2 with SIRIUS-COMET resulted in the correct annotation of 229/300 library members for miniSEL1 (76%), and 218/300 (73%) for miniSEL2. An even distribution among the building blocks was observed, except for 4-butylaniline, indicating that combinations of the optimized building blocks were generally well tolerated (Fig. 3c and Fig. S3). These results confirm the successful synthesis under combinatorial settings of most library members, forming a robust basis for the preparation of larger libraries.
As SELs are decoded by their chemical structure, we investigated whether common side products could be detected and characterized. In this way, side products could even become part of the enumerated library and contribute to greater diversity. While analyzing the building block scope, we often observed conversions to the following recurring side products; unreacted starting material 1 and its hydrolyzed and proto-dehalogenated derivative. While we had previously confirmed the correct synthesis of the majority of all library compounds for miniSEL1 and 2, we wanted to assess whether we could also detect and annotate these combinatorial side products. For miniSEL1 74 out of 90 possible side products were identified in the MS/MS spectra (Fig. S5) and 16/45 for miniSEL2 (Fig. S6). These results highlight that SELs can, in principle, identify any structure formed during synthesis, provided that the underlying reaction pathways are known. Whether these side products will actually be productive and detectable in real screening campaigns will dependent on each individual quantity, but, notably, many of these structures would be incorrectly assigned in conventional DEL screenings and would require extensive off-DNA validation to determine the correct hit structure.
Library synthesis and selection against CAIX
With an established method for combinatorial synthesis and decoding of Buchwald–Hartwig based libraries, the preparation of a large SEL library was initiated. Using 49 amino acid building blocks, 7 aryl bromides and 75 amines, we assembled a SEL with 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 725 members (Fig. 4a). Depending on the type of amine, we used the respective optimized conditions for the Buchwald–Hartwig amination step. For specific exact masses the library contains up to 28 isobaric compounds but each of those features a distinct combination of building blocks, likely enabling structure annotation through MS/MS fragmentation (Fig. 4b). We computationally enumerated the entire library and visualized compliance with the Lipinski parameters: molecular weight (MW < 500 Da), X
725 members (Fig. 4a). Depending on the type of amine, we used the respective optimized conditions for the Buchwald–Hartwig amination step. For specific exact masses the library contains up to 28 isobaric compounds but each of those features a distinct combination of building blocks, likely enabling structure annotation through MS/MS fragmentation (Fig. 4b). We computationally enumerated the entire library and visualized compliance with the Lipinski parameters: molecular weight (MW < 500 Da), X![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (> 5), hydrogen bond donors (HBD < 5), hydrogen bond acceptors (HBA < 10), and topological polar surface area (TPSA < 140) (Fig. 4c).
P (> 5), hydrogen bond donors (HBD < 5), hydrogen bond acceptors (HBA < 10), and topological polar surface area (TPSA < 140) (Fig. 4c).
Finally, we wanted to validate the entire discovery workflow including affinity selection, tandem mass spectrometry and hit decoding. To this end, we performed an affinity selection experiment against CAIX. CAIX is a therapeutically relevant target for cancer treatment, particularly in hypoxic tumors, due to its role in tumor cell survival and pH regulation in the tumor microenvironment. High affinity CAIX ligands are enabling tools for radionuclide cancer therapy.36 Moreover, CAIX is a preferred scaffold for benchmarking library selections due to its predictable binding profile to aromatic sulfonamides.25,36 In addition, other members of the carbonic anhydrase family have frequently served as benchmark targets for the validation of ligand discovery strategies such as in situ click chemistry37 and direct to biology screening.38,39
The affinity selection was performed on biotin-CAIX, immobilized on streptavidin-coated magnetic beads and incubated with the library (using 100 fmol per member). After washing away non-binders, protein-bound hits were eluted under denaturing conditions and analyzed with our nanoLC-MS/MS SIRIUS-COMET workflow. The selection in parallel was performed on unfunctionalized magnetic streptavidin beads. Non-specific binders, appearing in both selections, were computationally removed from our analysis by performing a background subtraction. Only compounds displaying at least three-fold higher intensity in the CAIX sample were considered as hits.
After background subtraction and SIRIUS-COMET structure annotation, we obtained 21 candidate structures. When analyzing building block frequencies, we found a significant enrichment of 4-aminobenzenesulfonamide, appearing in 13 annotated structures as first ranked (Fig. 4d). Given the known affinity of 4-aminobenzenesulfonamide for CAIX, these 13 hits can be considered as real binders with high confidence. For four of the remaining eight candidates, the sulfonamide building block was still present, but proposed with a lower rank. This outcome indicates that optimizing the ranking algorithm could improve hit detection in the future. The remaining four candidates in the manual analysis showed unclear enrichment and/or overall poor structure annotation and were not considered as hits. Interestingly, the sulfonamide always occurred in combination with 5-(4-bromophenyl)isoxazole-3-carboxylic acid, indicating that this combination is important for binding (Fig. 4d). The building block enrichment (proportion of sulfonamide in hits vs. expected random distribution) has a strong statistical significance with a p value of p < 0.0001 (Fig. S5).
From the hits we selected two compounds for experimental binding validation (Fig. 4e). We prepared variants of both compounds (11 and 12) with a biotin tag and tested their binding via biolayer interferometry (BLI), which demonstrated low nanomolar binding affinities with KD values of 48 nM and 31 nM, respectively. We resynthesized the untagged version of compound 12 to further confirm and validate the results of our affinity selection. While the nanoLC retention time and MS/MS spectra matched the original hit perfectly (Fig. S18 and S19), NMR characterization was hindered by a proto-dehalogenated side product formed during synthesis. Despite multiple attempts, this byproduct could not be separated from the target compound (Fig. S20 and S21). This difficulty illustrates a common bottleneck in combinatorial chemistry, where the transition from a mixture-based “hit” to a purified, isolated compound is not always straightforward. Altogether, the CAIX selection demonstrates that Buchwald–Hartwig aminations can be incorporated in SELs and enable discovery campaigns from large libraries.
Discussion and conclusion
Here, we developed procedures for solid-phase Buchwald–Hartwig aminations, enabling the construction of large combinatorial libraries. We optimized reaction conditions for aromatic and aliphatic amines and tested a building block scope of 22 aryl bromides and 156 amines. Building blocks with incorporations >50% were included in a final SEL of 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 725 members. We adapted our previously developed SIRIUS-COMET software to enable the automated structure annotation of MS/MS spectra of complex samples of compounds resulting from the combinatorial Buchwald–Hartwig amination. Finally, we demonstrated the application in AS-MS/MS against CAIX, resulting in robust hit enrichment and two validated binders with nanomolar affinities (48 nM and 31 nM), highlighting the potential of SELs for discovering novel hits.
725 members. We adapted our previously developed SIRIUS-COMET software to enable the automated structure annotation of MS/MS spectra of complex samples of compounds resulting from the combinatorial Buchwald–Hartwig amination. Finally, we demonstrated the application in AS-MS/MS against CAIX, resulting in robust hit enrichment and two validated binders with nanomolar affinities (48 nM and 31 nM), highlighting the potential of SELs for discovering novel hits.
Compared to Suzuki cross couplings that we previously developed,26 establishing the Buchwald–Hartwig amination for combinatorial synthesis was more challenging, exhibited a narrower substrate scope and lower yields. The Buchwald–Hartwig amination reported here required tailored reaction conditions based on the type of amine, resulting in distinct reaction protocols for primary aromatic and cyclic aliphatic amines. Primary aromatic amines were generally well tolerated, while aliphatic amines showed poor conversion and probably due to their susceptibility to β-hydride elimination. Cyclic amines outperformed their acyclic counterparts. Both findings are consistent with previous reaction performed in DEL optimization.15,40
A remaining challenge for DELs is the identification of true hits after the selection arising from incorrectly encoded library members, due to truncations or side product formation. In contrast, SELs enable direct decoding of compounds from their own structure, thereby eliminating issues related to barcode-compound mismatches. Moreover, SELs also offer the potential to detect and characterize side products that may arise during the synthesis.
Future research will be needed to further expand the chemical space by incorporating a more diverse set of aryl bromides and amines. A significant variability in ligand performance in our optimizations would suggest that further optimization into newly developed ligands for the incorporation of primary amines,41 secondary amines42,43 and heterocycles44–46 would increase our substrate scope. Since we have no DNA constraints, we can likely directly apply novel reported reaction conditions to solid-support, offering a distinct advantage over DEL-based reaction optimization.
To decode isobaric compounds in larger libraries (beyond 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 members), additional MS/MS fragment information on the introduced C–N is required. Under the current conditions, this bond fragments only sporadically, limiting confident structural assignment for larger libraries, as information about the amine building block is missing. By further optimizing MS/MS-based fragmentation in terms of collision energy and structure annotations, we could potentially enhance the decoding accuracy.
000 members), additional MS/MS fragment information on the introduced C–N is required. Under the current conditions, this bond fragments only sporadically, limiting confident structural assignment for larger libraries, as information about the amine building block is missing. By further optimizing MS/MS-based fragmentation in terms of collision energy and structure annotations, we could potentially enhance the decoding accuracy.
The modest library size shown in this study still exceeds size the size of pools (typically 500–1000 members) that can be screened by classical affinity selection mass spectrometry screens, relying on identification by MS1. DELs usually achieve substantially larger library sizes. However, we are confident that our SEL platform continues expanding and multiple libraries with different scaffolds will overall contribute to a broad chemical diversity.
The selection against CAIX yielded 13 hits, where the enrichment was dependent on the combination of 4-aminobenzenesulfonamide and 5-(4-bromophenyl)isoxazole-3-carboxylic acid. CAIX is known to bind to aromatic sulfonamides, but adjacent BBs can play an important role.36 The dependence on these two building blocks showcase the importance of developing diverse chemistry of different library scaffolds.
From the 343 potential sulfonamide binders, only a fraction was identified. This outcome reflects the constraints arising from building-block orientation in combination with the applied washing protocol. While our washing steps efficiently remove non-binders also ligands with fast dissociation rates might be lost. Consequently, at 100 fmol per library member, the assay is biased toward the identification of low-nanomolar binders. Fine-tuning the library concentration and washing parameters could allow systematic modulation of this affinity threshold, an aspect to be explored in subsequent studies.
Metal residues in the library might pose a challenge in specific screening campaigns. The solid support based synthesis enables extensive washing after the cross-coupling step, likely removing the catalyst from the system. After acidic cleavage from the resin the library is also bulk purified on a C18 column, leading to further removal of undesired components. In this study we did not take additional steps for quantitative metal removal. We acknowledge that in specific cases the presence of palladium might represent a limitation and we suggest in these cases to perform additional resin washes using potent metal chelators.
Overall, this work expands the chemical space accessible to self-encoded libraries and illustrates how established reactions can be integrated into the SEL platform for early drug discovery workflows.
Author contributions
Conceptualization: EvdN, ZL, QQG, SP, synthesis: EvdN, ZL, QQG, methodology: EvdN, NAH, SB, SP, assays: EvdN, software: NAH, SB, visualization: EvdN, SP, supervision: EvdN, SP, funding acquisition: SP, SB, writing – original draft: EvdN, SP, writing – review & editing: EvdN, NAH, SB, SP.Conflicts of interest
SP, SB, EvdN and NAH have filed a patent application for the methodology described here. The remaining authors declare no competing interests.Data availability
The raw LC-MS/MS files and the files needed to run SIRIUS:COMET have been deposited in the Zenodo database (https://doi.org/10.5281/zenodo.17632652). Source data are provided with this paper. All other data is available in the main text, the supplementary information (SI) and from the corresponding author(s) upon request. Supplementary information is available. See DOI: https://doi.org/10.1039/d5cb00303b.Acknowledgements
SJP acknowledges funding via the ERC Starting Grant (101039354). The Pomplun Lab gratefully acknowledges financial support from Mr H. J. M. Roels through a donation to the Oncode Institute and KWF's financial support of the Oncode Institute. NAH and SB are supported by the Thüringer Ministerium für Wirtschaft, Wissenschaft und Digitale Gesellschaft (TMWWDG) with funds from the European Union as part of the European Regional Development Fund (ERDF, 2023 VFE 0003).References
- A. Gironda-Martínez, E. J. Donckele, F. Samain and D. Neri, DNA-Encoded Chemical Libraries: A Comprehensive Review with Succesful Stories and Future Challenges, ACS Pharmacol. Transl. Sci., 2021, 4, 1265–1279 CrossRef PubMed.
- A. A. Peterson and D. R. Liu, Small-molecule discovery through DNA-encoded libraries, Nat. Rev. Drug Discovery, 2023, 22, 699–722 CrossRef CAS PubMed.
- N. Favalli, G. Bassi, J. Scheuermann and D. Neri, DNA-encoded chemical libraries – achievements and remaining challenges, FEBS Lett., 2018, 592, 2168–2180 CrossRef CAS PubMed.
- X. Yang, J. L. Childs-Disney, M. Paegel and M. D. Disney, DNA-Encoded Libraries and Their Application to RNA, Isr. J. Chem., 2023, 63, e202300073 CrossRef CAS.
- Q. Chen and D. I. Israel, Affinity-based DEL Selections with Different Target Types: Overview and Achievements. in DNA-encoded Library Technology for Drug Discovery, Royal Society of Chemistry, 2025, pp. 1–16 10.1039/9781788016032-00001.
- R. E. Sibai, Z. E. M. Farahat, H. H. Qasem and H. Hassan, The power of DNA-encoded chemical libraries in the battle against drug-resistant bacteria, RSC Adv., 2025, 15, 14001–14029 RSC.
- P. Ma, et al., Evolution of chemistry and selection technology for DNA-encoded library, Acta Pharm. Sin. B, 2024, 14, 492–516 CrossRef CAS PubMed.
- Y. Gao, et al., Recent advances in DNA-encoded libraries, Chem. Commun., 2025, 61, 10952–10968 RSC.
- K. Götte, S. Chines and A. Brunschweiger, Reaction development for DNA-encoded library technology: From evolution to revolution?, Tetrahedron Lett., 2020, 61, 151889 CrossRef.
- P. Ruiz-Castillo and S. L. Buchwald, Applications of Palladium-Catalyzed C-N Cross-Coupling Reactions, Chem. Rev., 2016, 116, 12564–12649 CrossRef CAS PubMed.
- Y. Shi, Y. R. Wu, J. Q. Yu, W. N. Zhang and C. L. Zhuang, DNA-encoded libraries (DELs): a review of on-DNA chemistries and their output, RSC Adv., 2021, 11, 2359–2376 RSC.
- P. R. Fitzgerald and B. M. Paegel, DNA-Encoded Chemistry: Drug Discovery from a Few Good Reactions, Chem. Rev., 2021, 121, 7155–7177 CrossRef CAS PubMed.
- R. J. Fair, R. T. Walsh and C. D. Hupp, The expanding reaction toolkit for DNA-encoded libraries, Bioorg. Med. Chem. Lett., 2021, 51, 128339 CrossRef CAS PubMed.
- R. Dorel, C. P. Grugel and A. M. Haydl, The Buchwald–Hartwig Amination After 25 Years, Angew. Chem., Int. Ed., 2019, 58, 17118–17129 CrossRef CAS PubMed.
- X. Lu, S. E. Roberts, G. J. Franklin and C. P. Davie, On-DNA Pd and Cu promoted C–N cross-coupling reactions, MedChemComm, 2017, 8, 1614–1617 RSC.
- M. L. Malone and B. M. Paegel, What is a ‘DNA-Compatible’ Reaction?, ACS Comb. Sci., 2016, 18, 52 Search PubMed.
- T. Z. Liu, T. F. Lin, D. T. Y. Chiu, K. J. Tsai and A. Stern, Palladium or Platinum Exacerbates Hydroxyl Radical Mediated DNA Damage, Free Radical Biol. Med., 1997, 23, 155–161 CrossRef CAS PubMed.
- S. L. Rössler, N. M. Grob, S. L. Buchwald and B. L. Pentelute, Abiotic peptides as carriers of information for the encoding of small-molecule library synthesis, Science, 1979, 379(939–945), 2023 Search PubMed.
- H. Xu, et al., DNA-Encoded Libraries: Aryl Fluorosulfonates as Versatile Electrophiles Enabling Facile On-DNA Suzuki, Sonogashira, and Buchwald Reactions, Adv. Sci., 2019, 6, 1901551 CrossRef CAS PubMed.
- Y. C. Chen, et al., C-N Coupling of DNA-Conjugated (Hetero)aryl Bromides and Chlorides for DNA-Encoded Chemical Library Synthesis, Bioconjugate Chem., 2020, 31, 770–780 CrossRef CAS PubMed.
- E. De Pedro Beato, et al., Mild and Efficient Palladium-Mediated C-N Cross-Coupling Reaction between DNA-Conjugated Aryl Bromides and Aromatic Amines, ACS Comb. Sci., 2019, 21, 69–74 CrossRef CAS PubMed.
- P. R. Chheda, N. Simmons, D. P. Schuman and Z. Shi, Palladium-Mediated C−N Coupling of DNA-Conjugated (Hetero)aryl Halides with Aliphatic and (Hetero)aromatic Amines, Org. Lett., 2022, 24, 3401–3406 CrossRef CAS PubMed.
- J. S. Graham, J. H. Hunter and M. J. Waring, Micellar Buchwald–Hartwig Coupling of Aryl and Heteroarylamines for the Synthesis of DNA-Encoded Libraries, J. Org. Chem., 2021, 86, 17257–17264 CrossRef CAS PubMed.
- N. J. Gesmundo, et al., Nanoscale synthesis and affinity ranking, Nature, 2018, 557, 228–232 CrossRef CAS PubMed.
- M. Keller, et al., Highly pure DNA-encoded chemical libraries by dual-linker solid-phase synthesis, Science, 2024, 384, 1259–1265 CrossRef CAS PubMed.
- E. van der Nol, et al., Barcode-free hit discovery from massive libraries enabled by automated small molecule structure annotation, Nat. Commun., 2025, 16(1), 1–12 Search PubMed.
- C. A. Willoughby and K. T. Chapman, Solid phase synthesis of aryl amines, Tetrahedron Lett., 1996, 37, 7181–7184 CrossRef CAS.
- K. Weigand and S. Pelka, Palladium-Catalyzed Animation of Aryl Halides on Solid Support, Org. Lett., 2002, 4, 4689–4692 CrossRef CAS PubMed.
- Y. D. Ward and V. Farina, Solid phase synthesis of aryl amines via palladium catalyzed amination of resin-bound aromatic bromides, Tetrahedron Lett., 1996, 37, 6993–6996 CrossRef CAS.
- D. Prat, et al., CHEM21 selection guide of classical- and less classical-solvents, Green Chem., 2015, 18, 288–296 RSC.
- M. E. Vitaku, D. T. Smith and J. T. Njardarson, Analysis of the Structural Diversity, Substitution Patterns, and Frequency of Nitrogen Heterocycles among U.S. FDA Approved Pharmaceuticals, J. Med. Chem., 2014, 57, 10257–10274 CrossRef PubMed.
- R. N. Muchiri and R. B. van Breemen, Drug discovery from natural products using affinity selection-mass spectrometry, Drug Discovery Today Technol., 2021, 40, 59–63 CrossRef CAS PubMed.
- N. F. Rizvi, et al., Discovery of Selective RNA-Binding Small Molecules by Affinity-Selection Mass Spectrometry, ACS Chem. Biol., 2018, 13, 820–831 CrossRef CAS PubMed.
- R. Prudent, D. A. Annis, P. J. Dandliker, J. Y. Ortholand and D. Roche, Exploring new targets and chemical space with affinity selection-mass spectrometry, Nat. Rev. Chem., 2021, 5, 62–71 CrossRef CAS PubMed.
- D. A. Annis, E. Nickbarg, X. Yang, M. R. Ziebell and C. E. Whitehurst, Affinity selection-mass spectrometry screening techniques for small molecule drug discovery, Curr. Opin. Chem. Biol., 2007, 11, 518–526 CrossRef CAS PubMed.
- S. Oehler, et al., A DNA-encoded chemical library based on chiral 4-amino-proline enables stereospecific isozyme-selective protein recognition, Nat. Chem., 2023, 15, 1431–1443 CrossRef CAS PubMed.
- M. Kugler, et al., Identification of specific carbonic anhydrase inhibitors via in situ click chemistry, phage-display and synthetic peptide libraries: comparison of the methods and structural study, RSC Med. Chem., 2023, 14, 144–153 RSC.
- R. P. Thomas, et al., A direct-to-biology high-throughput chemistry approach to reactive fragment screening, Chem. Sci., 2021, 12, 12098–12106 RSC.
- R. P. Thomas, et al., Reactive fragments targeting carboxylate residues employing direct to biology, high-throughput chemistry, RSC Med. Chem., 2023, 14, 671–679 RSC.
- R. Li, Y. Ruff and F. Berst, Efficient copper-catalyzed amination of DNA-conjugated aryl iodides under mild aqueous conditions, MedChemComm, 2018, 9, 1188–1193 RSC.
- L. Zhang, et al., The terphenyl phosphine TRuPhos: a novel and efficient ligand for the palladium-catalyzed arylation of hindered primary alkyl amines with mechanism clarification, J. Catal., 2025, 451, 116391 CrossRef CAS.
- K. Feng, et al., Development of a Deactivation-Resistant Dialkylbiarylphosphine Ligand for Pd-Catalyzed Arylation of Secondary Amines, J. Am. Chem. Soc., 2024, 146, 26609–26615 CrossRef CAS PubMed.
- N. H. Park, E. V. Vinogradova, D. S. Surry and S. L. Buchwald, Design of New Ligands for the Palladium-Catalyzed Arylation of α-Branched Secondary Amines, Angew. Chem., Int. Ed., 2015, 54, 8259–8262 CrossRef CAS PubMed.
- M. Su, N. Hoshiya and S. L. Buchwald, Palladium-Catalyzed Amination of Unprotected Five-Membered Heterocyclic Bromides, Org. Lett., 2014, 16, 832–835 CrossRef CAS PubMed.
- M. D. Charles, P. Schultz and S. L. Buchwald, Efficient Pd-Catalyzed Amination of Heteroaryl Halides, Org. Lett., 2005, 7, 3965–3968 CrossRef CAS PubMed.
- F. Zhou, L. Zhang, W. Hu, B. Yuan and J. C. Shi, A general catalyst for Buchwald–Hartwig amination to prepare secondary five-membered heteroaryl amines with breaking the base barrier, J. Catal., 2023, 422, 36–42 CrossRef CAS.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |