Open Access Article
Open Access ArticleInfluence of linker design on the stability, folding, and assembly of tethered collagen-mimetic peptides
Debasis
Ghosh
,
Anthony R.
Perez
 ,
S M Mobin
Sikder
,
S M Mobin
Sikder
 ,
Jesus R.
Vasquez
,
Jesus R.
Vasquez
 ,
Claire
Zhang
,
Claire
Zhang
 ,
Anjan
Maity
,
Tao
Ye
and
Andrea D.
Merg
,
Anjan
Maity
,
Tao
Ye
and
Andrea D.
Merg
 *
*
Department of Chemistry and Biochemistry, University of California, Merced, 5200 N. Lake Rd., Merced, CA 95343, USA. E-mail: amerg@ucmerced.edu
First published on 8th April 2026
Abstract
Covalently tethered collagen-mimetic peptides (CMPs) serve as synthetically programmable molecules for studying the collagen triple helix fold. Tethered CMPs, which overcome limitations that are inherent to their untethered counterparts (e.g., decreased stability, concentration-dependent folding, and slow folding kinetics), are constructed using a variety of templating strategies. Despite the plethora of reports of tethered CMPs in literature, there has been little exploration in determining the effects that the linker region, which connects the CMP sequence to the trivalent scaffold, has on the stability and assembly of tethered CMP triple helices. Here, we systematically study the influence of linker length and composition on the stability, folding, and assembly of covalently tethered CMPs. We synthesized a family of tethered CMPs comprising PEGylated linkers of different lengths (CTH-PEG2, CTH-PEG4, CTH-PEG6) to assess how linker length influences the properties of CMP triple helices. Moreover, we synthesized tethered CMPs comprising hydrophobic (CTH-HEX) and peptide-based linkers (CTH-GSG). All tethered CMPs possess a triblock sequence architecture that directs the assembly of resulting triple helices into nanostructures. Transmission electron microscopy (TEM) and atomic force microscopy (AFM) confirm that tethered CMPs assemble into nanosheets and nanoribbons. Circular dichroism (CD) spectroscopy reveals that increasing the length of the flexible linker systematically decreases the thermal stability of tethered CMP triple helices and alters their folding kinetics. Furthermore, CD data of CTH-HEX and CTH-GSG indicate that linker composition can play a role, though limited, in influencing the stability and folding properties of CMP triple helices. The presented work highlights how tuning the linker design – both length and composition – serves as a facile route towards fine-tuning the properties of CMP triple helices and their assemblies without perturbing the CMP sequence architecture, and will provide guidance to future researchers in choosing appropriate linkers for their own applications.
Introduction
Collagen-mimetic peptides (CMPs) represent synthetically accessible constructs for understanding the structural and folding properties of the collagen triple helix.1,2 Over the past several decades, studies involving CMP triple helices have elucidated their host–guest properties,3,4 afforded insights into their folding and unfolding kinetics,5 and have served as testing grounds for incorporating non-canonical residues and chemical moieties to modify, diversify, and stabilize the triple helix fold.2,6–9 Furthermore, inspired by the diversity and function of collagen-based architectures in biology,10 in which the collagen triple helix serves as the foundational structural motif, there has been significant progress in developing CMPs as synthetically programmable collagen-mimetic building blocks for fabricating artificial biomolecular assemblies for a broad array of potential applications in biotechnology.1,11–18The collagen triple helix fold, derived from CMPs, is defined by three peptide strands that intertwine to form a right-handed helix via staggered interstrand hydrogen bonding interactions. The peptide sequences comprise a three-residue repeat, Xaa-Yaa-Gly, in which Gly residues occupy every third position, while Pro and Hyp frequently occupy the Xaa and Yaa positions, respectively. Other amino acids can be incorporated in these positions, but this must be done judiciously as deviations from the canonical Pro-Hyp-Gly triad destabilizes the collagen triple helix fold.3,4
Because of the short sequence length of CMPs and the lack of covalent stabilization (a feature found in natural collagen), the stability of CMP triple helices is dependent on the CMP sequence length, amino acid composition, and CMP concentration in solution. Alterations to the peptide sequence outside the canonical Pro-Hyp-Gly repeat and working with CMPs under more dilute conditions, lowers the stability of the triple helix fold. Moreover, it is well-documented that CMPs exhibit slow rates of folding that can often take days and/or weeks.5,19 These limitations serve as barriers for employing CMPs and their supramolecular assemblies for biomedical applications. To overcome these challenges, template-assisted strategies have been developed to broaden the stability of CMP triple helices.20 Several approaches utilize trivalent organic scaffolds such as Kemp triacid (KTA),21,22 tris(2-aminoethyl) amine (TREN),23 cyclic hydropyran oligolides,24 cyclotriveratrylene (CTV),25 1,2,3-propanetricarboxylic acid,26 among others.20 In addition, there has been success in utilizing a branching strategy that involves employing orthogonally protected Lys residues, biorthogonal azide–alkyne cycloaddition, or disulfide bridges (e.g., “cysteine knots”).27–33 In all cases, covalent tethering of CMPs stabilizes triple helicity and enhances folding kinetics compared to their untethered counterparts.
Despite the plethora of reports on covalently tethered CMPs, little attention has been directed towards exploring the effects of the linker that connects the individual CMP strands to the trivalent template, and how the linker design influences the stability and folding of the resulting CMP triple helices. To our knowledge, only a single report investigated the role of linker length on the properties of tethered CMPs. In this work, conducted by Hojo and co-workers, they demonstrated that altering the linker length, via employing a combination of aminohexanoic acid (Ahx) and/or beta-alanine (βAla), was shown to influence the thermal stability of the triple helix.34 They determined that longer linkers, which possess greater conformational flexibility (thus increasing the entropic cost of folding), led to a decrease in the thermal stability of the tethered CMPs that were studied. Intrigued by this report and motivated by the lack of further exploration on the influence of tethered CMP linker designs in the literature, we set forth to provide a systematic investigation on the role that linker length and linker chemistry has in influencing the properties of CMP triple helices. Additional studies on linker design will provide researchers with guidelines for choosing appropriate linkers, including linker length and composition, for future applications involving tethered CMPs.
Here, we synthesized a family of tethered CMPs comprising different lengths and different linkage chemistry. Unlike the previous approach, we explore the effects of utilizing hydrophilic, PEGylated linkers of varying lengths, and compare them with tethered CMPs that utilize commonly employed hydrophobic linkers (Ahx) and peptide-based linkers (Gly-Ser-Gly). These studies are all within the context of employing a previously studied triblock CMP sequence design that directs the assembly of CMP triple helices into nanoscale architectures (Fig. 1a). The present work demonstrates that the stability and assembly of tethered CMPs can be fine-tuned and broadened without altering the CMP sequence architecture. Moreover, our results challenge the common assumption that extended linkers are advantageous for forming stable molecular triple helices. We demonstrate here that directly tethering CMPs off the ε-amine of Lys yields collagen triple helices with the greatest stability.
Results and discussion
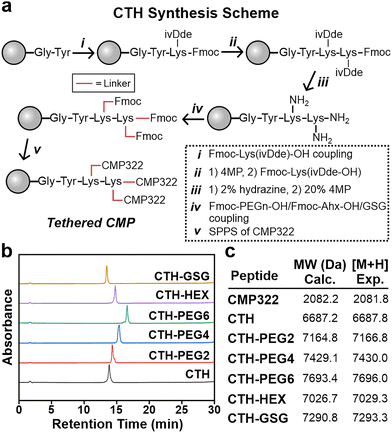
Tethered CMP Design and Synthesis. The parent CMP sequence, CMP322, consists of three Pro-Arg-Gly triads, two canonical Pro-Hyp-Gly triads, and two Glu-Hyp-Gly triads (Fig. 1c). This triblock sequence architecture, in which positive and negatively charged triads are segregated at opposite ends, has been previously shown to assemble into nanosheets (Fig. 1a).35–37 The CMP nanosheets comprise collagen triple helices that are packed in an antiparallel fashion, which affords favourable Coulombic interactions between adjacent helices (Fig. 1a). An additional positively charged triad (Pro-Arg-Gly) to the CMP sequence was included to bias the formation of single-layer nanosheets with a thickness that is commensurate with the length of a single triple helix, as demonstrated previously.38Tethered CMPs (CTH, CTH-PEG2, CTH-PEG4, CTH-PEG6, CTH-HEX, and CTH-GSG; Fig. 1c) were prepared using solid-phase peptide synthesis (SPPS) via a branched lysine strategy (Fig. 2a).29 Two Lys(ivDde) residues, which contain orthogonally protected ivDde-protecting group chemistry,39 are coupled sequentially near the C-terminus. Upon Fmoc and ivDde deprotection using 20% 4-methylpiperidine (4MP) and 2% hydrazine, respectively, the unprotected amines were coupled with various PEGylated acids (PEG2-OH, PEG4-OH, PEG6-OH for CTH-PEG2, CTH-PEG4, CTH-PEG6, respectively; PEG = polyethylene glycol), Ahx (CTH-HEX), or a Gly-Ser-Gly sequence (CTH-GSG) using standard SPPS chemistry. The CMP sequence (CMP322; Fig. 1c and S1) was then synthesized directly off the linkers via SPPS to yield tethered CMPs (Fig. 2a). For the synthesis of CTH, two CMP322 sequences were attached directly from the ε-amino groups of both Lys residues. Tyrosine was incorporated at the C-terminus to quantify the tethered CMPs via UV-vis spectroscopy. All tethered CMPs were successfully synthesized and isolated with product yields of 5.5%, 6.9%, 7.0%, and 7.2% for CTH, CTH-PEG2, CTH-PEG4, and CTH-PEG6, respectively (Fig. 2b, c and Fig. S2–S7). The lower yield of CTH-GSG (2.0%) may be a consequence of the sterically bulky Gly-Ser-Gly linker and the two additional coupling steps that are required.
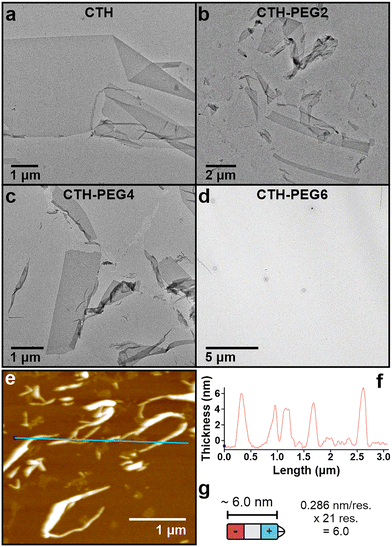
Effect of Linker Length on Tethered CMP Triple Helices. CTH, CTH-PEG2, CTH-PEG4, and CTH-PEG6 (200 µM) were dissolved in 20 mM 3-(Morpholin-4-yl)propane-1-sulfonic acid (MOPS) buffer (pH 7.0). The solutions were heated to 90 °C to dissociate any kinetically trapped aggregates and ensure a monomeric population of tethered CMPs. After heating, the solutions were slow-cooled to 4 °C (0.2 °C min−1) and incubated at 4 °C for at least one week. Transmission electron microscopy (TEM) images of CTH, CTH-PEG2, and CTH-PEG4 assembly solutions reveal the presence of sheets and nanoribbons of varying sizes and morphology (Fig. 3a–c and Fig. S8a–c), except for CTH-PEG6, in which no assemblies were observed (Fig. 3d and Fig. S8d). The physical dimensions of the assemblies extend from microns to tens of microns in size with varying aspect ratios. Atomic force microscopy (AFM) of CTH nanoribbons determine assembly thicknesses of 5.8 ± 2.0 nm (Fig. 3e, f and Fig. S9), which confirms that the assemblies do not comprise multiple stacked layers of CMPs. The average thickness is in line with the expected length of approximately 6.0 nm for the CMP triple helix domain (0.286 nm rise/res. for collagen triple helix; Fig. 3g).40,41 Unlike previously studied nanosheets, which were assembled from untethered CMPs, the nanostructures appear to be relatively flexible with a tendency to wrinkle and fold together. We attribute the observed increase in flexibility of the nanostructures to the shorter CMP length within tethered CMPs (21 residues) compared to previous untethered CMP designs that utilized CMP lengths ranging from 36 to 45 residues.35–38 We surmise that the decreased thicknesses of our tethered CMP assemblies give rise to less rigid nanostructures that are more susceptible to deformation. In addition, a uniform contrast of individual nanosheets, from the TEM data, further suggest that the tethered CMP assemblies comprise a single layer of CMPs. As described above, this is expected due to the additional positively charged triad within the sequence design.38 No definable assemblies were observed for CMP322 (600 µM) at comparable CMP concentrations, due to the significant decrease in triple helix stability of untethered CMP322 (vide infra; Fig. S10).
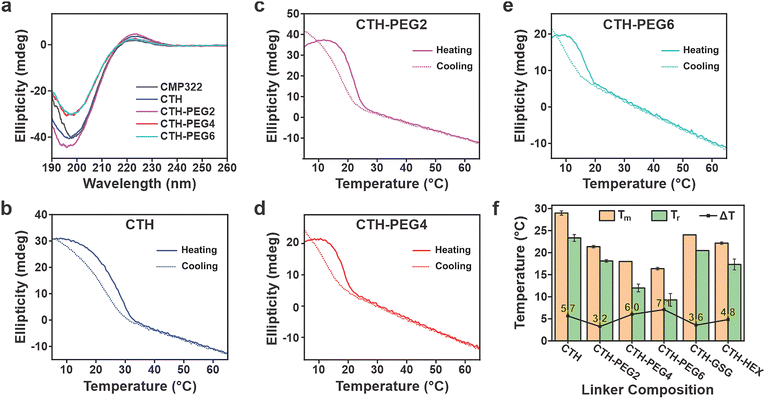
Circular dichroism (CD) spectroscopy confirms the presence of collagen triple helices for all tethered CMPs (Fig. 4a). CD spectra of the PEGylated CMP derivatives display the characteristic profile indicative of collagen triple helix formation, as evidenced by maxima and minima peaks at ∼224 nm and ∼198 nm, respectively (Fig. 4a). Rpn values, which report the ratio of these peak intensities, are useful in determining the population of collagen triple helices in solution. Rpn values of 0.09, 0.1, 0.09, and 0.08 for CTH, CTH-PEG2, CTH-PEG4, and CTH-PEG6, respectively, indicate the presence of stable collagen triple helices.42 These values contrast the calculated Rpn value of 0.05 for CMP322, which suggests that the increased population of folded triple helices is a result of covalent tethering. However, we note that light scattering and orientation effects, which stem from the large size and anisotropic nature of tethered CMP assemblies, can significantly distort CD measurements and thus prevent the use of Rpn as a conclusive method to determine collagen triple helix stability.
Prior reports have demonstrated that the stability of CMP nanosheets are determined by the melting temperature (Tm) of the collagen triple helices that comprise their assembled structure.36 CD thermal denaturation experiments, carried out in triplicate, of the tethered CMP assemblies confirm the decrease in stability of collagen triple helices as a function of increasing linker lengths (Fig. 4b–f and Fig. S11). We note that because triple helix formation proceeds intramolecularly, thus significantly reducing the association kinetics of trimerization, Tm values can provide insight regarding differences in thermal stabilities between tethered CMPs. Interestingly, CTH, in which CMP sequences are directly attached to lysine side chains, exhibits the greatest thermal stability (Tm = 29.0 ± 0.5 °C; Fig. 4b, f). These results imply that the inclusion of flexible linkers, which is routinely employed (e.g., Ahx) as a way to better accommodate the folding geometry of the triple helix structure, is unnecessary for enhancing the stability of collagen triple helices. The Lys side chain itself provides the necessary flexibility required for forming stable collagen triple helices. For CTH-PEG2, CTH-PEG4, and CTH-PEG6, we observe a decrease in Tm (21.4 ± 0.3 °C, 18.0 ± 0.04 °C, and 16.4 ± 0.3 °C, respectively) signifying that utilizing longer linkers destabilizes the triple helix fold (Fig. 4c–f). The low melting temperature of CTH-PEG6, which indicates its reduced triple helix stability, likely explains the lack of observable assemblies for CTH-PEG6. The decrease in thermal stability as a function of increasing linker lengths confirms the previous study (vide supra),34 which demonstrated that tethered CMPs with longer, hydrophobic linkers give rise to CMP triple helices with lower thermal stabilities. This is attributed to the increasing entropic penalty associated with longer, flexible linkers, without any enthalpic gains as the CMP length remains unchanged.34 In comparison to the tethered CMPs, no triple helix Tm was observed for CMP322 (Fig. S12). This is unsurprising since tethering CMP domains together lower the entropic and kinetic barriers for folding via increasing the effective local concentration and pre-organizing the alignment of CMP strands for triple helix formation. Both factors contribute to stabilizing the presence of the triple helical fold under the studied conditions. Altogether, these results demonstrate the ability to fine-tune the thermal stability of tethered CMPs simply through modulating the linker length. This contrasts conventional approaches that rely on manipulating the CMP sequence, either through extending the CMP length or altering the CMP amino acid composition.
The folding kinetics of CMPs were compared within the family of tethered PEGylated CMPs via assessing the hysteresis loop between their heating and cooling profiles.43 A significant hysteresis between the unfolding and refolding temperature is often observed for untethered CMPs due to the slow, rate-determining cis–trans amide isomerization step that is required for the formation of triple helices and the relative high kinetic barrier associated with bringing three strands together in the correct registry.44 In contrast, through lowering the kinetic barrier, intramolecular folding associated with tethered CMPs significantly minimize hysteresis, highlighting their faster fold recovery.45–47 From refolding experiments, we determine hysteresis (ΔT) values of 3.2 °C, 6.0 °C, and 7.1 °C for CTH-PEG2, CTH-PEG4, and CTH-PEG6, respectively, which suggests that longer PEGylated linker lengths lead to slower relative refolding rates (Fig. 4b–f). However, a ΔT of 5.7 °C is calculated for CTH, which implies that other factors (e.g., steric effects) may contribute to the refolding rate of our tethered CMPs. We also observe larger final CD signals for the PEGylated triple helices after conclusion of the refolding experiments. These results indicate that there is an increase in the fraction of folded collagen triple helices, which suggests that the folded population was not at equilibrium or comprised triple helices that were partially misfolded, prior to heating. Interestingly, we do not observe this behaviour for CTH. We speculate that this may be due to the improved stability and registry alignment of CTH triple helices, both of which can be attributed to the absence of the extended PEGylated linker. To our knowledge, this work provides the first characterization of unfolding/refolding hysteresis for end-capped CMPs. We note that the ΔT values are larger than reported values for covalently captured CMPs, in which CMPs are tethered via multiple, interstrand isopeptide bonds.45,46 The smaller hysteresis of covalently captured CMPs is likely due to slower heating/cooling rates that were employed, which more closely capture the true equilibrium transitions.43
Effect of Linker Composition. To assess the importance of the chemical make-up of the linker region on the thermal and folding properties of CMPs, we extended our study to include tethered CMPs having aliphatic and canonical peptide-based linkers (CTH-HEX and CTH-GSG, respectively; Fig. 1c). We hypothesized that these CMP designs may shed light on the effect of employing hydrophobic (CTH-HEX) and sterically bulky linkers (CTH-GSG). The contour lengths of the linkers within CTH-HEX and CTH-GSG (7.4 and 9.7 Å, respectively) are similar to the contour length of PEG2 within CTH-PEG2 (10.7 Å; Fig. S13).
Akin to the PEGylated derivatives, CTH-GSG and CTH-HEX assemble into nanostructures with heterogeneous size distributions (Fig. S14 and S15). CD spectra confirm the formation of collagen triple helices, as evidenced by the canonical collagen triple helical profile with maxima and minima at ∼224 and ∼198 nm, respectively (Fig. 5a). In comparison to CTH-PEG2 (Tm = 21.4 °C), CTH-GSG (Tm = 24.0 °C) and CTH-HEX (Tm = 22.2 °C) both possess slightly higher thermal stabilities (Fig. 4f and 5b,c). The higher Tm for CTH-GSG may be explained by the lower conformational freedom associated with the peptide-based linker, which decreases the entropic penalty for folding. Refolding experiments of CTH-GSG and CTH-HEX reveal ΔT values of 3.6 °C and 4.8 °C for CTH-GSG and CTH-HEX, respectively, which are similar to the ΔT value of CTH-PEG2 (3.2 °C; Fig. 4f and 5b, c). Altogether, these results suggest that the chemistry of the linker may have a subtle role in influencing the unfolding and folding properties of tethered CMP triple helices, however, additional studies involving a broader set of linkers will be needed to validate any potential trends.
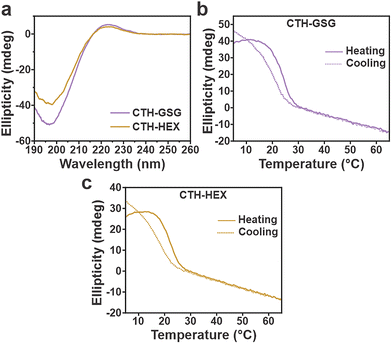 | ||
| Fig. 5 (a) CD spectra of CTH-GSG and CTH-HEX (200 µM each) in 20 mM MOPS buffer (pH 7.0) after one week. CD thermal denaturation and refolding curves for (b) CTH-GSG and (c) CTH-HEX (200 µM each). Tm, Tr and ΔT values for CTH-GSG and CTH-HEX are shown in Fig. 4f. | ||
Assembly under Dilute Conditions. A limitation of assembling biomaterials using CMP building blocks is that a relatively high concentration of CMPs is often required for assembly. Because of the concentration-independent folding of tethered CMPs, and that the formation of charge-segregated CMP triple helices is a necessary step for forming CMP nanosheets,36 we reasoned that our tethered CMPs should assemble under dilute conditions. CD spectrum of 50 µM CTH solution confirm the presence of stable collagen triple helices (Fig. 6a). Moreover, CD thermal denaturation and refolding experiments, carried out on CTH at 50 µM concentration, reveal approximate Tm of 27 °C, refolding temperature (Tr) of 21 °C, and ΔT value of 6 °C (Fig. 6b, c and Fig. S16). These values are similar to the data collected from 200 µM CTH solutions. TEM images of CTH, assembled at 50 µM reveal the presence of nanostructures, including nanoribbons and nanosheets (Fig. 6d), which confirm that tethered CMPs assemble independently of concentration. These results point to the ability to assemble tethered CMPs under dilute conditions, which overcomes limitations where relatively high concentrations are required for CMP self-assembly.
Conclusions
We systematically explored the effects of linker length and composition on the stability and folding of several tethered CMPs. We demonstrate that longer linkers give rise to tethered CMP triple helices with lower thermal stabilities when employing the same linker chemistry. Furthermore, longer linkers generally lead to slower relative fold recovery rates (i.e., larger hysteresis), however, there are some notable exceptions (e.g., CTH has larger hysteresis compared to CTH-PEG2). We also demonstrate that linker composition may play a role in altering the stability and folding of tethered CMPs, however, the changes we observe indicate that these effects are minor (at least for the limited number of linker designs that we study here).When employing a branched lysine template strategy, our results indicate that attaching a flexible linker off the lysine side chain is not a necessary requirement for forming stable triple helices. The use of Ahx, for example, as an additional linker is prevalent throughout the reported literature.20,29–32 In fact, the presented work indicates that tethered CMPs, in which CMP domains are directly attached to the ε-amino group of lysine, yield the most stable triple helices. These studies highlight how linker length can serve to “fine-tune” the stability of triple helices and their supramolecular assemblies. This strategy contrasts conventional approaches for tuning the properties of tethered CMP triple helices, which rely on altering the CMP sequence via incorporating amino acid substitutions or modifying the length of the CMP domain. Such changes to the CMP design may lead to a loss of target function, lower yields (due to additional SPPS coupling/deprotection steps), and decreases in the stability of triple helices and their assembled architectures. The ability to tailor the folding properties of tethered CMPs without perturbing the functional CMP sequence domain represents a useful tool that can be exploited for designing families of tethered CMP constructs with identical CMP sequences but having adjustable thermal stabilities.
Furthermore, in all cases, we demonstrate that the tethered CMPs, comprising charge-segregated sequences, self-assemble into nanostructures. In contrast, the untethered version, which cannot form stable triple helices at the conditions employed, are unable to self-assemble. Lastly, we confirm that the assembly of tethered CMPs is concentration-independent. We note that the presented conclusions are specific to the tethered system and CMP design presented (i.e., lysine-branched strategy and charge-segregated CMPs, respectively). Further studies are needed to determine the extent that this work translates with other CMP tethering strategies and CMP sequence architectures. In summary, we anticipate that the presented work will provide a starting point for designing future tethered CMPs in which researchers can weigh the utility of various linker designs for their intended applications.
Author contributions
D. G., A. R. P., S. M. M. S., A. M., and C. Z. synthesized, characterized, and purified CMPs, and collected HPLC, MALDI-TOF, TEM, and CD data. J. R. V. performed AFM characterization. J. R. V. and T. Y. analyzed AFM data. A. D. M. conceived the project and wrote the manuscript. All authors contributed to proofing and editing the final manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Additional requests for data are available from the corresponding author upon request.The data that support the findings of this study are available in the supplementary information (SI) of this article. See DOI: https://doi.org/10.1039/d5cb00298b.
Acknowledgements
This work was supported by the NSF (DMR-2534247, A.D.M.) and the University of California, Merced. A.R.P. acknowledges fellowship support from NSF-CREST: Center for Cellular and Biomolecular Machines (NSF-HRD-1547848 and NSF-HRD-2112675). J.R.V and T.Y. acknowledge support from U.S. Department of Energy, Office of Basic Energy Sciences (DE-SC0020961). We acknowledge access to TEM via the Imaging and Microscopy Facility at UC Merced.Notes and references
- J. A. Fallas, L. E. R. O'Leary and J. D. Hartgerink, Chem. Soc. Rev., 2010, 39, 3510–3527 RSC.
- S. A. H. Hulgan and J. D. Hartgerink, Biomacromolecules, 2022, 23, 1475–1489 CrossRef CAS PubMed.
- N. K. Shah, J. A. M. Ramshaw, A. Kirkpatrick, C. Shah and B. Brodsky, Biochemistry, 1996, 35, 10262–10268 CrossRef CAS PubMed.
- A. V. Persikov, J. A. M. Ramshaw, A. Kirkpatrick and B. Brodsky, Biochemistry, 2000, 39, 14960–14967 CrossRef CAS PubMed.
- C. C. Cole, D. R. Walker, S. A. H. Hulgan, B. H. Pogostin, J. W. R. Swain, M. D. Miller, W. Xu, R. Duella, M. Misiura, X. Wang, A. B. Kolomeisky, G. N. Philips and J. D. Hartgerink, Nat. Chem., 2024, 16, 1698–1704 CrossRef CAS PubMed.
- J. L. Kessler, G. Kang, Z. Qin, H. Kang, F. G. Whitby, T. E. Cheatham III, C. P. Hill, Y. Li and S. M. Yu, J. Am. Chem. Soc., 2021, 143, 10910–10919 CrossRef CAS PubMed.
- M. Goodman, G. Melacini and Y. Feng, J. Am. Chem. Soc., 1996, 118, 10928–10929 CrossRef CAS.
- Y. Feng, G. Melacini and M. Goodman, Biochemistry, 1997, 36, 8716–8724 CrossRef CAS PubMed.
- Y. Zhang, R. M. Malamakal and D. M. Chenoweth, J. Am. Chem. Soc., 2015, 137, 12422–12425 CrossRef CAS PubMed.
- D. E. Birk and P. Bruckner, in Collagen: Primer in Structure, Processing and Assembly, ed. J. Brinckmann, H. Notbohm and P. K. Müller, Springer Berlin Heidelberg, Berlin, Heidelberg, 2005, pp. 185–205 DOI:10.1007/b103823.
- D. Kalita and B. K. Sarma, Langmuir, 2025, 41, 9162–9185 CrossRef CAS PubMed.
- L. T. Yu, M. A. B. Kreutzberger, T. H. Bui, M. C. Hancu, A. C. Farsheed, E. H. Egelman and J. D. Hartgerink, Nat. Commun., 2024, 15, 10385 CrossRef CAS PubMed.
- L. E. R. O'Leary, J. A. Fallas, E. L. Bakota, M. K. Kang and J. D. Hartgerink, Nat. Chem., 2011, 3, 821–828 CrossRef PubMed.
- B. Sarkar, L. E. R. O’Leary and J. D. Hartgerink, J. Am. Chem. Soc., 2014, 136, 14417–14424 CrossRef CAS PubMed.
- F. W. Kotch and R. T. Raines, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 3028–3033 CrossRef CAS PubMed.
- E. M. Ford, A. M. Hilderbrand and A. M. Kloxin, J. Mater. Chem. B, 2024, 12, 9600–9621 RSC.
- S. Mukherjee, V. Varshashankari, A. Feba, N. Ayyadurai, K. Balamurugan and G. Shanmugam, Biomacromolecules, 2025, 26, 2171–2185 CrossRef CAS PubMed.
- I. C. Tanrikulu, L. Dang, L. Nelavelli, A. J. Ellison, B. D. Olsen, S. Jin and R. T. Raines, Adv. Sci., 2024, 11, 2303228 CrossRef CAS PubMed.
- N. J. Greenfield, Nat. Protoc., 2006, 1, 2527–2535 CrossRef CAS PubMed.
- M. Bhowmick and G. B. Fields, in Peptide Modifications to Increase Metabolic Stability and Activity, ed. P. Cudic, Humana Press, Totowa, NJ, 2013, pp. 167–194 DOI:10.1007/978-1-62703-652-8_11.
- M. Goodman, Y. Feng, G. Melacini and J. P. Taulane, J. Am. Chem. Soc., 1996, 118, 5156–5157 CrossRef CAS.
- Y. Li, X. Mo, D. Kim and S. M. Yu, Biopolymers, 2011, 95, 94–104 CrossRef CAS PubMed.
- J. Kwak, A. D. Capua, E. Locardi and M. Goodman, J. Am. Chem. Soc., 2002, 124, 14085–14091 CrossRef CAS PubMed.
- J.-C. Horng, A. J. Hawk, Q. Zhao, E. S. Benedict, S. D. Burke and R. T. Raines, Org. Lett., 2006, 8, 4735–4738 CrossRef CAS PubMed.
- E. T. Rump, D. T. S. Rijkers, H. W. Hilbers, P. G. de Groot and R. M. J. Liskamp, Chem. – Eur. J., 2002, 8, 4613–4621 CrossRef CAS PubMed.
- Y. Greiche and E. Heidemann, Biopolymers, 1979, 18, 2359–2361 CrossRef CAS.
- J. Ottl, R. Battistuta, M. Pieper, H. Tschesche, W. Bode, K. Kühn and L. Moroder, FEBS Lett., 1996, 398, 31–36 CrossRef CAS PubMed.
- J. Ottl and L. Moroder, J. Am. Chem. Soc., 1999, 121, 653–661 CrossRef CAS.
- C. G. Fields, C. M. Lovdahl, A. J. Miles, V. L. M. Hageini and G. B. Fields, Biopolymers, 1993, 33, 1695–1707 CrossRef CAS PubMed.
- W. Roth and E. Heidemann, Biopolymers, 1980, 19, 1909–1917 CrossRef CAS.
- S. Thakur, D. Vadolas, H.-P. Germann and E. Heidemann, Biopolymers, 1986, 25, 1081–1086 CrossRef CAS PubMed.
- H.-P. Germann and E. Heidemann, Biopolymers, 1988, 27, 157–163 CrossRef CAS PubMed.
- C. Byrne, P. A. McEwan, J. Emsley, P. M. Fischer and W. C. Chan, Chem. Commun., 2011, 47, 2589–2591 RSC.
- H. Hojo, Y. Akamatsu, K. Yamauchi, M. Kinoshita, S. Miki and Y. Nakamura, Tetrahedron, 1997, 53, 14263–14274 CrossRef CAS.
- T. Jiang, C. Xu, Y. Liu, Z. Liu, J. S. Wall, X. Zuo, T. Lian, K. Salaita, C. Ni, D. Pochan and V. P. Conticello, J. Am. Chem. Soc., 2014, 136, 4300–4308 CrossRef CAS PubMed.
- A. D. Merg, E. van Genderen, A. Bazrafshan, H. Su, X. Zuo, G. Touponse, T. B. Blum, K. Salaita, J. P. Abrahams and V. P. Conticello, J. Am. Chem. Soc., 2019, 141, 20107–20117 CrossRef CAS PubMed.
- A. D. Merg, G. Touponse, E. van Genderen, X. Zuo, A. Bazrafshan, T. Blum, S. Hughes, K. Salaita, J. P. Abrahams and V. P. Conticello, Angew. Chem., Int. Ed., 2019, 58, 13507–13512 CrossRef CAS PubMed.
- T. Jiang, O. A. Vail, Z. Jiang, X. Zuo and V. P. Conticello, J. Am. Chem. Soc., 2015, 137, 7793–7802 CrossRef CAS PubMed.
- S. R. Chhabra, A. N. Khan and B. W. Bycroft, Tetrahedron Lett., 1998, 39, 3585–3588 CrossRef CAS.
- K. Okuyama, Connect. Tissue Res., 2008, 49, 299–310 CrossRef CAS PubMed.
- K. Okuyama, X. Xu, M. Iguchi and K. Noguchi, Pept. Sci., 2006, 84, 181–191 CrossRef CAS PubMed.
- Y. Feng, G. Melacini, J. P. Taulane and M. Goodman, J. Am. Chem. Soc., 1996, 118, 10351–10358 CrossRef CAS.
- K. Mizuno, S. P. Boudko, J. Engel and H. P. Bächinger, Biophys. J., 2010, 98, 3004–3014 CrossRef CAS PubMed.
- H. P. Bächinger, J. Engel, P. Bruckner and R. Timpl, Eur. J. Biochem., 1978, 90, 605–613 CrossRef PubMed.
- S. A. H. Hulgan, A. A. Jalan, I. C. Li, D. R. Walker, M. D. Miller, A. J. Kosgei, W. Xu, G. N. Phillips Jr. and J. D. Hartgerink, Biomacromolecules, 2020, 21, 3772–3781 CrossRef CAS PubMed.
- C. C. Cole, L. T. Yu, M. Misiura, J. Williams III, T. H. Bui and J. D. Hartgerink, Biomacromolecules, 2023, 24, 5083–5090 CrossRef CAS PubMed.
- A. V. Persikov, Y. Xu and B. Brodsky, Protein Sci., 2004, 13, 893–902 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |