Open Access Article
Open Access ArticleIgalan attenuates sepsis-induced inflammation through covalent targeting of the NLRP3 inflammasome pathway
Chengchen Hou a,
Yang Chena,
Wenjie Bib,
Lin Gaob,
Simiao Yua,
Xiaowen Zhangac,
Zekun Chenad,
Tiantian Weia,
Dilnoza E. Dusmatova
a,
Yang Chena,
Wenjie Bib,
Lin Gaob,
Simiao Yua,
Xiaowen Zhangac,
Zekun Chenad,
Tiantian Weia,
Dilnoza E. Dusmatova e,
Rimma F. Mukhamatkhanovae,
Ildar D. Sham’yanove,
Liwen Hanb,
Zhiyuan Lu
e,
Rimma F. Mukhamatkhanovae,
Ildar D. Sham’yanove,
Liwen Hanb,
Zhiyuan Lu b,
Hua Wang*fg,
Nilufar Z. Mamadalieva
b,
Hua Wang*fg,
Nilufar Z. Mamadalieva *e and
Kewu Zeng
*e and
Kewu Zeng *ad
*ad
aState Key Laboratory of Natural and Biomimetic Drugs, School of Pharmaceutical Sciences, Peking University, Beijing 100191, China. E-mail: ZKW@bjmu.edu.cn
bKey Laboratory of Chemical Biology (Ministry of Education), School of Pharmaceutical Sciences, Cheeloo College of Medicine Shandong University, Jinan 250012, China
cBeijing Research Institute of Chinese Medicine, Beijing University of Chinese Medicine, Beijing 102488, China
dDepartment of Integration of Chinese and Western Medicine, School of Basic Medical Sciences, Peking University, Beijing 100191, China. E-mail: ZKW@bjmu.edu.cn
eInstitute of the Chemistry of Plant Substances, Uzbekistan Academy of Sciences, Tashkent 100170, Uzbekistan. E-mail: nmamadalieva@yahoo.com
fDepartment of Oncology, the First Affiliated Hospital of Anhui Medical University, Hefei 230036, China. E-mail: wanghua@ahmu.edu.cn
gKey Laboratory of Anti-inflammatory and Immune Medicine (Anhui Medical University), Ministry of Education, Hefei, 230032, China. E-mail: wanghua@ahmu.edu.cn
First published on 23rd April 2026
Abstract
Macrophages play a critical role in sepsis, a life-threatening systemic inflammatory syndrome that necessitates urgent therapeutic intervention. Unfortunately, no approved drugs currently target this specific pathological mechanism. In this study, we identify Igalan, a natural compound that selectively disrupts macrophage-mediated inflammatory cascades in macrophages. Mechanistic studies demonstrate that Igalan alleviates oxidative stress and maintains mitochondrial integrity. Crucially, we reveal that Igalan covalently modifies the NLRP3 (NLR family pyrin domain containing 3) NACHT domain, thereby irreversibly suppressing inflammasome activation and subsequent pro-inflammatory signaling. In vivo studies demonstrate potent therapeutic effects of Igalan, mitigating systemic inflammation in LPS-challenged zebrafish and providing protection against pulmonary injury and intestinal barrier dysfunction in murine sepsis models. Collectively, our work establishes Igalan as a covalent NLRP3 inhibitor with translational potential for sepsis treatment.
Introduction
Sepsis is a life-threatening organ dysfunction caused by a dysregulated host response to infection.1 Current therapies for sepsis-associated lung injury, including corticosteroids and TNF-α/IL-6-targeting monoclonal antibodies, are limited by the risks of systemic immunosuppression and provide only marginal survival benefits.2–4 In recent years, macrophages have been increasingly reported to play a pivotal role in the pathogenesis of sepsis.2,3,5 The modulation of aberrant macrophage activation constitutes a vital therapeutic strategy. Thus, there is an urgent need to identify novel druggable targets that selectively regulate macrophage activation to advance innovative drug development.6Recent studies have revealed that NLRP3 inflammasome-mediated macrophage dysregulation disrupts immune homeostasis, promoting pathogenic polarization and systemic inflammation in sepsis.7,8 The activation of NLRP3 not only amplifies pro-inflammatory cytokine cascades but also disrupts immune homeostasis, thereby exacerbating systemic inflammation and multi-organ dysfunction, a hallmark of sepsis lethality.9–11 Mechanistically, NLRP3 oligomerization recruits the adaptor protein ASC, which subsequently facilitates the activation of caspase-1. This process leads to the proteolytic maturation of interleukin-1 beta (IL-1β) and interleukin-18 (IL-18), culminating in the induction of pyroptotic cell death.12,13 Of note, the feed-forward loop of cytokine release and gasdermin D-mediated pyroptosis not only amplifies inflammatory signaling but also disrupts tissue barriers, enhances damage-associated molecular pattern (DAMP) release, and further perpetuates systemic inflammatory cascades.14–16
Natural products are a significant source of new drugs due to the extensive pharmacological activities.17–20 Sesquiterpene lactones (SLs) are a class of bioactive plant-derived metabolites with diverse biological activities, including anti-inflammatory, anticancer, antimicrobial, antioxidant and immunomodulatory effects.21–23 Igalan, a sesquiterpene lactone present in Inula species, is derived from plants traditionally used to manage inflammatory diseases.24,25 Previous studies have identified Igalan as possessing diverse biological effects. For instance, a previous study demonstrated the efficacy of Igalan in improving oxidative stress response through the activation of Nrf2 signaling, thereby enhancing cellular antioxidant capacity.26 Moreover, another study revealed the role of Igalan in enhancing anti-tumor immunity by promoting the expression of genes related to immune cell activation and facilitating the response to PD-1 blockade in colorectal cancer.27 However, research on the anti-inflammatory properties of Igalan, particularly in mitigating sepsis-induced inflammatory pathology, remains limited.
In this study, we identified Igalan as an anti-inflammatory compound targeting sepsis-associated pulmonary injury and intestinal injury. Mechanistically, Igalan inhibits oxidative stress-mediated mitochondrial damage and suppresses the release of pro-inflammatory cytokines. Notably, Igalan directly targets the NLRP3 inflammasome by blocking ATPase activity in the NACHT domain, thereby preventing conformational changes required for inflammasome assembly. Moreover, Igalan reduces systemic reactive oxygen species (ROS) levels in vivo, as demonstrated in zebrafish models. Importantly, in murine sepsis models, Igalan not only alleviates pulmonary tissue injury and monocyte infiltration but also mitigates intestinal mucosal damage and barrier dysfunction. In summary, our study unveils an anti-inflammatory mechanism of Igalan, distinguishing it from conventional NLRP3 inhibitors and offering a new strategy for developing selective therapeutics.
Methods
Chemicals and reagents
Igalan (C15H20O2; molecular weight 232.318) was kindly provided by Prof. Nilufar Z. Mamadalieva, Uzbekistan Academy of Sciences. The lipopolysaccharide (LPS) employed in the study was obtained from Sigma-Aldrich (L2630, St. Louis, MO, USA), derived from Escherichia coli strain O111:B4. Bovine serum albumin (BSA) was obtained from VivaCell (C04001, Shanghai, China). High-glucose Dulbecco's modified Eagle's medium (DMEM), phosphate-buffered saline (PBS), trypsin, and penicillin–streptomycin were acquired from Macgene (Beijing, China). The Cell Counting Kit-8 (CCK-8) was purchased from New Cell & Molecular Biotech (Suzhou, China). Dexamethasone was purchased from J&K Scientific (308890, Beijing, China). Ethanol (100092683), xylene (10023418) and neutral gum (10004160) were obtained from Sinopharm Chemical Reagent (Beijing, China). The hematoxylin–eosin staining kit (G1003) was purchased from Wuhan Servicebio Technology (Wuhan, Hubei, China). 4% paraformaldehyde (P1110) and a DAB substrate kit were obtained from Solarbio (Beijing, China). The following primary antibodies were used for immunoblotting (diluted at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000): anti-NLRP3 (Cell Signaling Technology, Cat# 15101), anti-IL-1β (Proteintech, Cat# 16806-1-AP), anti-GSDMD (Abcam, Cat# ab209845), and the loading control anti-α-tubulin (Proteintech, Cat# 66031-1-Ig). For immunohistochemistry (IHC) analysis, the anti-TNF-α primary antibody was obtained from Abcam (Cat# ab1793, diluted at 1
1000): anti-NLRP3 (Cell Signaling Technology, Cat# 15101), anti-IL-1β (Proteintech, Cat# 16806-1-AP), anti-GSDMD (Abcam, Cat# ab209845), and the loading control anti-α-tubulin (Proteintech, Cat# 66031-1-Ig). For immunohistochemistry (IHC) analysis, the anti-TNF-α primary antibody was obtained from Abcam (Cat# ab1793, diluted at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200).
200).
The secondary antibodies used were horseradish peroxidase (HRP)-linked anti-rabbit IgG (Cell Signaling Technology, Cat# 7074, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2000) and HRP-linked anti-mouse IgG (Cell Signaling Technology, Cat# 7076, 1
2000) and HRP-linked anti-mouse IgG (Cell Signaling Technology, Cat# 7076, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2000). All general chemicals and reagents were of standard analytical grade.
2000). All general chemicals and reagents were of standard analytical grade.
Cell culture and stimulation
RAW264.7 murine macrophages were obtained from the Cell Bank of the Peking Union Medical College (Beijing, China). The immortalized mouse bone marrow-derived macrophages (iBMDMs) were generously gifted by Prof. Fuping You's lab at the School of Basic Medical Sciences, Peking University. Both cell lines were cultured in DMEM supplemented with 10% fetal bovine serum (FBS), 100 U mL−1 penicillin, and 100 µg mL−1 streptomycin, and maintained in a humidified incubator at 37 °C with 5% CO2 and 95% air. To assess the anti-inflammatory activity of Igalan, iBMDM and RAW264.7 cells were stimulated with LPS (1 µg mL−1) in the presence or absence of Igalan for 24 h. Dex was included as a positive control. Igalan and dexamethasone were dissolved in DMSO to prepare stock solutions. For all in vitro assays, the final concentration of DMSO in the cell culture medium was strictly maintained at 0.1% (v/v) across all experimental and vehicle control groups. To assess the inhibitory effect of NLRP3 inflammasome, iBMDM and RAW264.7 cells were primed with LPS (1 µg mL−1) for 3 h. Subsequently, Igalan or MCC950 was administered for 1 h, followed by treatment with Nigericin (10 µM) for 1 h. Western blot analysis was performed for subsequent assessment.Cell survival assay
Cell viability was assessed using the CCK-8 assay. Briefly, CCK-8 working solution (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 dilution) was added to each well, mixed gently, and incubated at 37 °C for 1 h in the dark. Absorbance was measured at 450 nm using a Sunrise-Basic microplate reader (Tecan, Männedorf, Zürich, Switzerland). Data were processed and visualized using GraphPad Prism (version 10.1).
10 dilution) was added to each well, mixed gently, and incubated at 37 °C for 1 h in the dark. Absorbance was measured at 450 nm using a Sunrise-Basic microplate reader (Tecan, Männedorf, Zürich, Switzerland). Data were processed and visualized using GraphPad Prism (version 10.1).
Nitric oxide (NO) assay
Nitric oxide (NO) production was measured using a commercial NO assay kit based on the Griess reaction. After 24 h of co-treatment with LPS (1 µg mL−1) and Igalan or Dex, culture supernatants were collected and mixed with Griess reagent at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio, followed by incubation at room temperature for 10 min. Absorbance was measured at 540 nm using an ELX800 UV universal microplate reader (BioTek, Winooski, VT, USA).
1 ratio, followed by incubation at room temperature for 10 min. Absorbance was measured at 540 nm using an ELX800 UV universal microplate reader (BioTek, Winooski, VT, USA).
Enzyme-linked immunosorbent assay (ELISA)
Levels of IL-1β, TNF-α, and IL-6 in the cell culture supernatant and serum were quantified using commercial ELISA kits (ExCell Bio). The cells were treated as described above and the culture media were collected and centrifuged at 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g for 15 min at 4 °C. The resulting cell culture supernatants were diluted 1
000 × g for 15 min at 4 °C. The resulting cell culture supernatants were diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 and subjected to subsequent analyses. Samples and standards were incubated in antibody-coated plates at 37 °C for 2 h, washed five times, and then incubated with HRP-conjugated detection antibodies at 37 °C for 1 h. After washing, chromogenic substrates A/B were added simultaneously, and color development was carried out for 10 min at room temperature under light-protected conditions. Reactions were stopped, and the optical density was quantified at 450 nm (reference wavelength: 570 nm) utilizing a Sunrise microplate reader (Tecan, Männedorf, Zürich, Switzerland). For serum preparation, whole blood was allowed to clot at room temperature for 60 min and centrifuged at 3000 × g for 10 min at 4 °C. The supernatant serum was diluted 1
2 and subjected to subsequent analyses. Samples and standards were incubated in antibody-coated plates at 37 °C for 2 h, washed five times, and then incubated with HRP-conjugated detection antibodies at 37 °C for 1 h. After washing, chromogenic substrates A/B were added simultaneously, and color development was carried out for 10 min at room temperature under light-protected conditions. Reactions were stopped, and the optical density was quantified at 450 nm (reference wavelength: 570 nm) utilizing a Sunrise microplate reader (Tecan, Männedorf, Zürich, Switzerland). For serum preparation, whole blood was allowed to clot at room temperature for 60 min and centrifuged at 3000 × g for 10 min at 4 °C. The supernatant serum was diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 before analysis, and all subsequent procedures were performed as described above.
5 before analysis, and all subsequent procedures were performed as described above.
Western blotting
Cells were lysed in ice-cold RIPA buffer supplemented with protease inhibitors. Protein concentrations were determined using an enhanced BCA protein assay kit (TransGen Biotech). Lysates were separated on 8–12% SDS-PAGE gels and transferred to polyvinylidene fluoride (PVDF) membranes (Merck) using a wet transfer system (Bio-Rad Laboratories, Hercules, CA, USA). After blocking with 5% (w/v) skimmed milk for 1 h at room temperature, the membranes were probed with primary antibodies (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution) at 4 °C overnight. Then, the membranes were incubated with HRP-conjugated secondary antibodies (1
1000 dilution) at 4 °C overnight. Then, the membranes were incubated with HRP-conjugated secondary antibodies (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000) for 1 h at room temperature. Protein bands were visualized using enhanced chemiluminescence (ECL) reagents, and images were captured with a Tanon 5200 Imaging System (Tanon, Shanghai, China). Densitometric quantification of band intensities was conducted using ImageJ software.
1000) for 1 h at room temperature. Protein bands were visualized using enhanced chemiluminescence (ECL) reagents, and images were captured with a Tanon 5200 Imaging System (Tanon, Shanghai, China). Densitometric quantification of band intensities was conducted using ImageJ software.
Detection of total cellular ROS and mitochondrial ROS
Total cellular ROS and mitochondrial ROS were measured using fluorescent probes DCFH-DA (Tauto Biotech) and MitoSOX™ Red (Thermo Fisher Scientific). After 24 h of co-treatment with LPS (1 µg mL−1) and Igalan or Dex, the cells were incubated with DCFH-DA (10 µM) or MitoSOX (5 µM) in a serum-free medium at 37 °C for 30 min in the dark. After three washes with serum-free medium, fluorescence was recorded using an IX73 fluorescence microscope (Olympus, Tokyo, Japan). DCFH-DA fluorescence was measured at excitation/emission wavelengths of 490 nm/530 nm, whereas MitoSOX was detected at excitation/emission wavelengths of 510 nm/580 nm. Image quantification was performed using Fiji/ImageJ software to calculate the numbers of DCFH-DA-positive cells, MitoSOX-positive cells, and total cell counts.Detection of mitochondrial membrane potential
Mitochondrial membrane potential was assessed using the fluorescent probe JC-1 (Beyotime). Briefly, after 24 h of co-treatment with LPS (1 µg mL−1) and Igalan or Dex, the culture medium was removed, and cells were incubated with JC-1 (10 µg mL−1) diluted in serum-free medium at 37 °C for 20 min in the dark. Then, the cells were washed three times with serum-free medium. Fluorescence was measured using an IX73 fluorescence microscope (Olympus, Tokyo, Japan). JC-1 monomers (green) were detected at 514 nm (excitation), whereas JC-1 aggregates (red) were observed at 590 nm (emission). The number of JC-1-positive cells with green and red fluorescence was quantified using Fiji/ImageJ software, and the green/red ratio was calculated to evaluate mitochondrial membrane potential.Sample preparation and LC-MS/MS analysis
iBMDM cells were stimulated with LPS (1 µg mL−1), lysed in RIPA buffer on ice for 1 h, and clarified by centrifugation (20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g, 4 °C). Equal protein aliquots were treated with either Igalan (2 µM) or DMSO at room temperature for 1 h with gentle agitation. The samples were processed by filter-aided sample preparation (FASP) using Vivacon-500 centrifugal filters (10 kDa MWCO, Sartorius) with centrifugation at 14
000 × g, 4 °C). Equal protein aliquots were treated with either Igalan (2 µM) or DMSO at room temperature for 1 h with gentle agitation. The samples were processed by filter-aided sample preparation (FASP) using Vivacon-500 centrifugal filters (10 kDa MWCO, Sartorius) with centrifugation at 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g for 15 min per wash step. The retentate was washed three times with 8 M urea/50 mM NH4HCO3, reduced with TCEP, alkylated with iodoacetamide (30 min, dark), and washed three times with 50 mM NH4HCO3. Trypsin digestion (1
000 × g for 15 min per wash step. The retentate was washed three times with 8 M urea/50 mM NH4HCO3, reduced with TCEP, alkylated with iodoacetamide (30 min, dark), and washed three times with 50 mM NH4HCO3. Trypsin digestion (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50, enzyme
50, enzyme![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) protein) was carried out overnight at 37 °C, and eluted peptides were vacuum-dried.
protein) was carried out overnight at 37 °C, and eluted peptides were vacuum-dried.
LC-MS/MS analysis was performed on an Orbitrap Exploris 480 mass spectrometer coupled to a Vanquish Neo UHPLC system (Thermo Fisher Scientific). Peptides were loaded onto a PepMap Neo trap column and separated on an in-house C18 analytical column (450 nL min−1) using a 90 min gradient from 4% to 99% mobile phase B (0.1% formic acid in 80% acetonitrile). The instrument was operated in positive ion, data-dependent acquisition (DDA) mode with a FAIMS Pro interface (CVs: −45 and −65 V). Full MS scans were acquired at 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 resolution (m/z 350–1500; AGC target 300%), followed by HCD-based MS/MS fragmentation (NCE 30; resolution 15
000 resolution (m/z 350–1500; AGC target 300%), followed by HCD-based MS/MS fragmentation (NCE 30; resolution 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000; isolation window 1.6 m/z; AGC target 100%). Raw data were searched against the UniProt mouse proteome database using the Andromeda engine in MaxQuant (v1.6.3.4). Covalent modification by Igalan was searched as a variable modification on cysteine residues, with a mass shift of +232.15 Da. An additional variable modification of +250.33 Da was included to assess potential lactone ring-opening hydrolysis.
000; isolation window 1.6 m/z; AGC target 100%). Raw data were searched against the UniProt mouse proteome database using the Andromeda engine in MaxQuant (v1.6.3.4). Covalent modification by Igalan was searched as a variable modification on cysteine residues, with a mass shift of +232.15 Da. An additional variable modification of +250.33 Da was included to assess potential lactone ring-opening hydrolysis.
KEGG and GO enrichment analysis
Functional annotation of Igalan-modified proteins was conducted using Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis and Gene Ontology (GO) enrichment analysis on the Bioinformatics.com.cn platform (Shanghai NewCore Biotechnology). Enriched terms with adjusted P values < 0.05 (Benjamini–Hochberg correction) were considered significant. Data visualization was performed using GraphPad Prism to generate bubble plots and bar charts representing enriched pathways and biological processes.Molecular docking analysis
Molecular docking simulations were performed using Schrödinger Release 2018 software: Maestro (Schrödinger, New York, NY, USA). The 3D structure of human NLRP3 (PDB ID: 6NPY) was downloaded from the RCSB Protein Data Bank. Since the Cys704 residue is located within an unresolved loop region in PDB: 6NPY, the missing loop was reconstructed using the Prime module within the Schrödinger software suite prior to docking. The completed structure was then prepared with the Protein Preparation Wizard, including bond order assignment, addition of hydrogen atoms, and removal of small molecules. Energy minimization was carried out under positional restraints using the OPLS3 force field. The grid box around the active site was generated with the Receptor Grid Generation module. Subsequently, NLRP3 was docked with Igalan using the Covalent Dock protocol implemented in the Schrödinger software suite. The reaction type of Michael addition was set. Scoring was performed with Glide XP, owing to its high accuracy in predicting binding modes.Animals
Wild-type AB zebrafish were randomly assigned to experimental groups at 4 days post-fertilization (4 dpf) and maintained in 6-well plates at 28 °C under a 14 h light/10 h dark photoperiod. All zebrafish experiments were conducted in full compliance with the relevant national regulations of China and the institutional guidelines for the care and use of laboratory animals of the Institute of Materia Medica, Shandong Academy of Medical Sciences. All procedures were reviewed and approved by the Experimental Animal Ethics and Welfare Committee of the Institute of Materia Medica, Shandong Academy of Medical Sciences (Approval no. W20250828850), and all experiments were performed in accordance with ARRIVE 2.0 guidelines.Male BALB/c mice (6–8 weeks old, 18–22 g) were obtained from the Department of Laboratory Animal Science, Peking University Health Science Center. The mice were housed under a controlled 12 h light/12 h dark cycle with ad libitum access to food and water. All mouse experiments were carried out in strict accordance with the relevant national laws and the institutional guidelines for the care and use of laboratory animals in China. All procedures were approved by the Biomedical Ethics Committee of Peking University (Approval no. DLASBD0156) and performed in compliance with ARRIVE 2.0 guidelines, with all efforts made to minimize animal suffering.
LPS-induced zebrafish inflammation model
Zebrafish were treated with LPS (70 µg mL−1) alone, co-treated with Igalan (1, 2, or 5 µg mL−1), or with the positive control dexamethasone (3 µg mL−1). After 24 h of exposure, cellular ROS levels were assessed. The zebrafish were incubated with 10 µM DCFH-DA at 37 °C for 10 min in the dark, followed by three washes with E3 embryonic medium (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2, and 0.33 mM MgSO4, supplemented with 0.1% methylene blue, pH 7.2). Fluorescence was observed using an IX73 fluorescence microscope (Olympus, Tokyo, Japan), with excitation/emission wavelengths of 490/530 nm. The fluorescence intensity of DCFH-DA in individual zebrafish was quantified using Fiji/ImageJ software.LPS-induced mouse sepsis model
Sepsis was induced in BALB/c mice by intraperitoneal injection of LPS (5 mg kg−1). Dexamethasone (DEX) was employed as the positive control to evaluate the in vivo therapeutic efficacy of Igalan. Mice in the treatment groups received intraperitoneal injections of Igalan (5, 10, or 20 mg kg−1) or dexamethasone (5 mg kg−1) 1 h before LPS challenge and again 6 h after the challenge. Following treatment, all animals were returned to their cages with ad libitum access to food and water. After 12 h of LPS challenge, blood samples were collected for ELISA. Intestinal and lung tissues were harvested, fixed in 4% paraformaldehyde (PFA), embedded in paraffin, and processed for hematoxylin and eosin (H&E) staining and immunohistochemical (IHC) analysis.Histology and immunohistochemistry
For hematoxylin and eosin (H&E) staining, fresh mouse lung tissues were fixed in 4% paraformaldehyde for 48 h, dehydrated through graded ethanol and xylene, embedded in paraffin, and sectioned at 5 µm using a rotary microtome (RM2106, Leica Biosystems, Wetzlar, Germany). After washing, sections were stained with 0.1% HCl–ethanol for 30 s, counterstained with eosin for 2 min, mounted with neutral gum, and examined for pathological changes under an Eclipse E100 light microscope (Nikon, Tokyo, Japan).For immunohistochemistry (IHC), paraffin sections (5 µm) were dewaxed in xylene, rehydrated through graded ethanol, and incubated in 3% H2O2 to quench endogenous peroxidase activity. After washing with TBST (five times), the sections were incubated overnight at 4 °C with a primary antibody against TNF-α (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500), followed by incubation with a goat anti-rabbit secondary antibody for 1 h at 25 °C. Visualization was achieved using a DAB substrate kit, and nuclei were counterstained with hematoxylin. The stained slides were imaged using a NanoZoomer-SQ digital slide scanner (Hamamatsu Photonics, Shizuoka, Japan).
500), followed by incubation with a goat anti-rabbit secondary antibody for 1 h at 25 °C. Visualization was achieved using a DAB substrate kit, and nuclei were counterstained with hematoxylin. The stained slides were imaged using a NanoZoomer-SQ digital slide scanner (Hamamatsu Photonics, Shizuoka, Japan).
Statistical analysis
All data are presented as mean ± standard deviation (SD) from at least three independent experiments. Statistical analyses were performed using GraphPad Prism 10.1.2 (GraphPad Software, San Diego, CA). Student's t-test was used to determine statistical significance, and P < 0.05 was considered statistically significant.Results
Igalan inhibits LPS-induced macrophage inflammatory responses
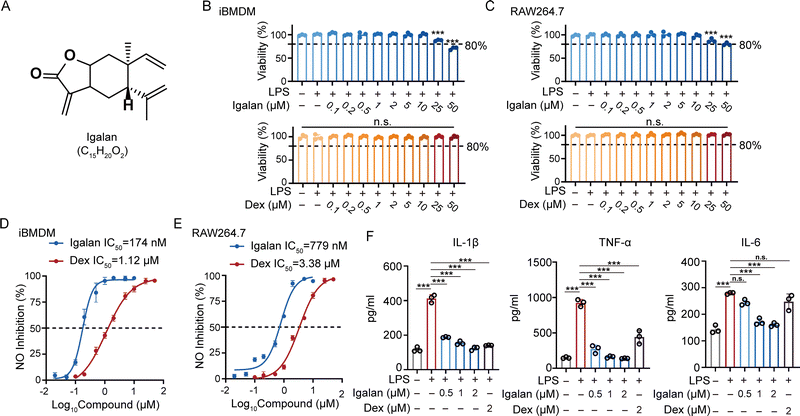
To assess the anti-inflammatory properties of Igalan (Fig. 1A), an inflammatory response was provoked in iBMDM cells using LPS (1 µg mL−1), co-treated with varying concentrations of Igalan (0.1–50 µM), or with dexamethasone as a positive control. The non-cytotoxic concentration range for Igalan was established using the CCK-8 assay (Fig. 1B and C), while nitric oxide (NO) levels in the culture supernatant were quantified to assess the anti-inflammatory effects. As a result, lgalan demonstrated potent inhibition of NO production, with IC50 values of 174 nM in iBMDM cells (Fig. 1D). We also detected the NO-inhibitory effects of Igalan with an IC50 of 779 nM in RAW264.7 macrophages (Fig. 1E). Furthermore, intracellular levels of the cytokines IL-1β, TNF-α, and IL-6 in LPS-stimulated iBMDMs co-treated with Igalan (0.5, 1, and 2 µM) were measured by ELISA. As shown in Fig. 1F, Igalan markedly reduced the synthesis of pro-inflammatory cytokines in a concentration-dependent manner. Collectively, these findings suggest that Igalan exerts potent anti-inflammatory effects by downregulating several inflammatory mediators.Igalan attenuates LPS-induced mitochondrial dysfunction by alleviating oxidative stress
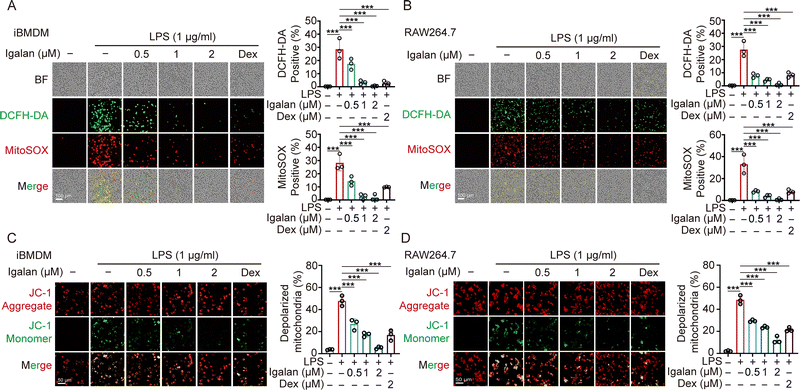
The intracellular accumulation of ROS serves as a hallmark of inflammatory responses, where excessive ROS exacerbates mitochondrial membrane disruption and subsequent cellular energy homeostasis dysregulation. To evaluate the effect of Igalan on LPS-induced intracellular ROS production, total ROS levels were measured using the fluorescent probe DCFH-DA. Meanwhile, mitochondrial superoxide generation was quantified using MitoSOX staining. As shown in Fig. 2A, Igalan decreased the proportion of DCFH-DA-positive cells in iBMDMs in a concentration-dependent manner and likewise reduced MitoSOX-positive cells, indicating the inhibition of LPS-induced total and mitochondrial ROS. Consistently, Igalan demonstrated a similar ROS-suppressive effect in RAW264.7 cells (Fig. 2B), further confirming the antioxidative capacity. To further investigate mitochondrial function, we assessed alterations in mitochondrial membrane potential using JC-1 staining. Quantitative analysis revealed that Igalan potently attenuated LPS-induced ΔΨm collapse in iBMDM cells (Fig. 2C), indicating preservation of mitochondrial integrity. Furthermore, this protective effect was consistently replicated in RAW264.7 macrophages (Fig. 2D). Collectively, these findings demonstrate that Igalan effectively suppresses ROS overproduction in activated macrophages, thereby preserving mitochondrial functionality under inflammatory conditions.Igalan suppresses macrophage activation by covalently modifying NLRP3 to block inflammasome signaling
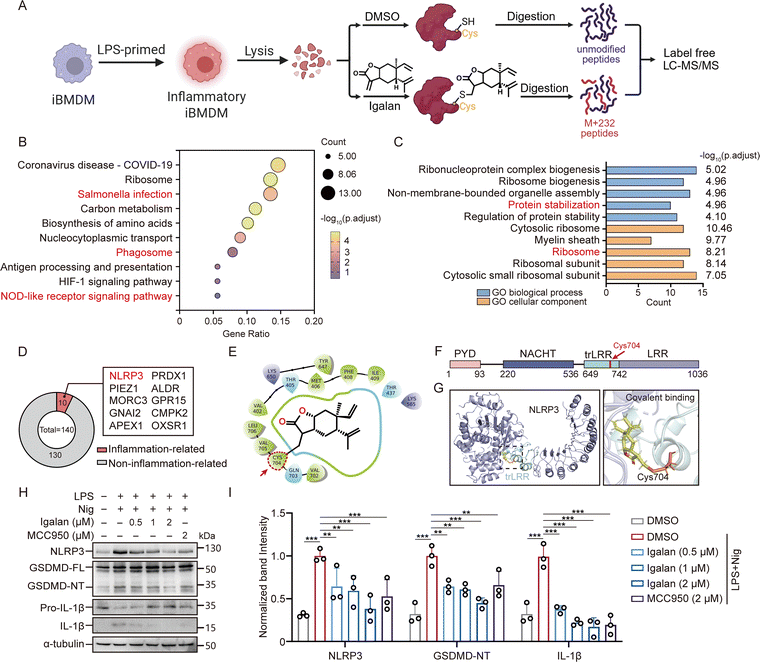
Structural analysis of Igalan (Fig. 1A) revealed an α,β-unsaturated ketone moiety, which enables Michael addition to cysteine residues of cellular target proteins. We hypothesized that the Igalan-mediated covalent modification may induce functional changes in the target protein, resulting in anti-inflammatory effects. To this end, we performed LC-MS/MS profiling of Igalan-modified proteins (Fig. 3A). A total of 140 proteins were identified, all verified to contain peptides with a +232.15 Da mass shift on cysteine residues (Table S1). KEGG pathway analysis showed significant enrichment in Salmonella infection, phagosome, and NOD-like receptor signaling pathways (Fig. 3B). Gene Ontology (GO) enrichment analysis further demonstrated that Igalan predominantly regulated biological processes such as protein stabilization and cellular components, including the ribosome (Fig. 3C). Correlation analysis of the modified proteome (Fig. 3D) revealed that 10 of the 140 proteins were directly implicated in inflammatory regulation.To elucidate the underlying mechanism, we focused on NLRP3, a critical component of the inflammasome, and conducted molecular docking studies with Igalan. The results indicated that Igalan covalently binds to Cys704 within the transition Leucine-Rich Repeat (trLRR) domain of NLRP3 (Fig. 3E–G). Furthermore, high-resolution LC-MS/MS analysis of the Igalan-modified NLRP3 peptide confirmed Cys704 as the specific covalent modification site (Fig. S1). The observed mass shift of +232.15 Da, with no corresponding +250.33 Da shift indicative of lactone ring-opening hydrolysis, is consistent with a Michael addition mechanism in which the lactone ring of Igalan remains intact upon adduction to Cys704. The trLRR domain bridges the NACHT and LRR domain (Fig. 3F), which is critical for NLRP3 inflammasome assembly and activation. Upon binding to Cys704, Igalan stabilizes the interface between these two domains, thereby preventing the conformational changes in the NACHT domain that are essential for inflammasome activation. These structural rearrangements are required for ATP binding and hydrolysis as a critical step that drives inflammasome assembly. Subsequently, western blotting was performed to assess the effect of Igalan on NLRP3 inflammasome activation, GSDMD cleavage, and IL-1β maturation. As shown in Fig. 3H, Igalan potently inhibited the activation of NLRP3 inflammasome, thereby reducing GSDMD cleavage and the secretion of IL-1β. Meanwhile, the inhibitory effects of Igalan were comparable to those observed with MCC950 (Fig. 3I). To further validate that Cys704 is the functional target of Igalan, a mutagenesis rescue assay was performed. The expression of NLRP3 was depleted in iBMDMs by siRNA and reconstituted with WT or C704S mutant NLRP3, followed by measurement of IL-1β secretion upon Igalan treatment. As shown in Fig. S2, Igalan potently suppressed IL-1β release in WT-reconstituted cells, whereas this effect was abrogated in C704S mutant-expressing cells, demonstrating that covalent modification of Cys704 is indispensable for Igalan-mediated inflammasome inhibition. Collectively, these findings demonstrate that Igalan strongly suppresses NLRP3 inflammasome activation and subsequent inflammatory responses.
Igalan inhibits LPS-induced inflammation in zebrafish and mouse models of sepsis
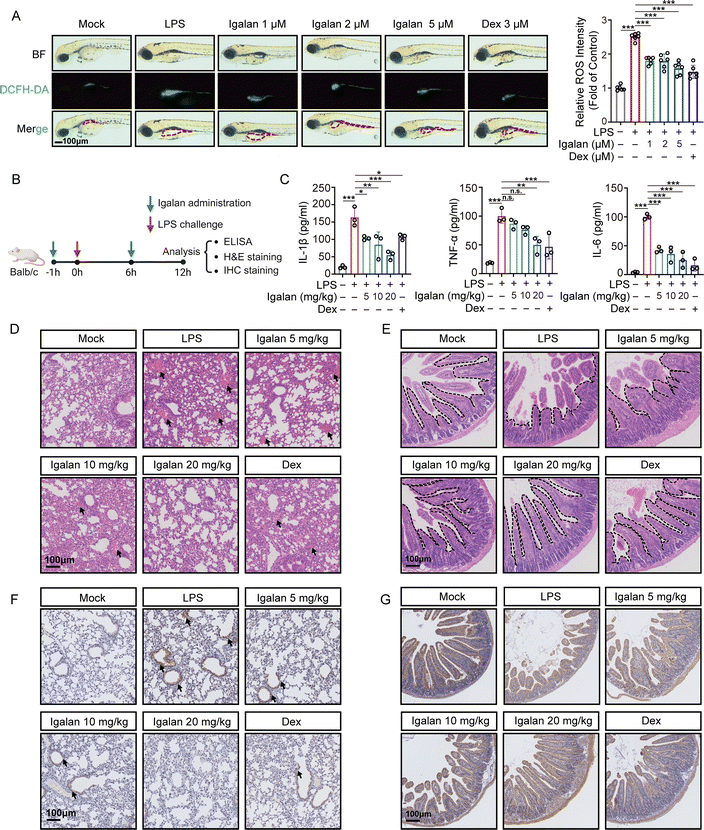
To evaluate the anti-inflammatory effect of Igalan in vivo, we established a zebrafish model of systemic inflammation by LPS treatment. DCFH-DA staining demonstrated that LPS induced a marked increase in pro-inflammatory ROS, which was potently attenuated by Igalan (Fig. 4A). Moreover, a septic model was established in Balb/c mice by intraperitoneal injection of LPS (Fig. 4B). ELISA analysis revealed that Igalan markedly inhibited the LPS-induced release of pro-inflammatory cytokines (IL-1β, TNF-α, and IL-6), with the high-dose group (20 mg kg−1) exhibiting robust suppression comparable to that of the positive control, dexamethasone (Fig. 4C). Histopathological examination of lung tissues (H&E staining) revealed that LPS provoked mononuclear macrophage infiltration, which was markedly alleviated by Igalan treatment (Fig. 4D). Furthermore, H&E staining demonstrated that Igalan significantly alleviated LPS-induced intestinal mucosal damage (Fig. 4E). Of note, immunohistochemical (IHC) staining confirmed that Igalan reduced TNF-α expression in lung tissues, thereby attenuating pulmonary inflammation (Fig. 4F). Furthermore, Igalan reversed LPS-induced downregulation of the tight junction protein ZO-1 in intestinal epithelia, indicating a protective effect on gut barrier integrity (Fig. 4G). Collectively, these findings demonstrate that Igalan effectively attenuates LPS-induced systemic inflammation, suggesting its potential as a therapeutic candidate for sepsis management.Discussion
Sepsis pathophysiology is driven by dysregulated innate immune activation, in which NLRP3 inflammasome signaling plays a major role.28,29 This study identifies Igalan as a novel anti-inflammatory agent that directly suppresses NLRP3 inflammasome activation through covalent modification, thereby suppressing oxidative stress as a downstream consequence. Mechanistically, we demonstrate that Igalan induces a covalent electrophile-driven post-translational modification, which restricts ATP hydrolysis-dependent conformational changes in the NLRP3 NACHT domain. This perturbation may disrupt the structural rearrangements required for inflammasome oligomerization, ASC speck formation, caspase-1 activation, and IL-1β maturation. Given that the inhibition of ATP-driven conformational activation of NLRP3 is recognized as a well-established preclinical mechanistic strategy,30,31 our findings identify Igalan as a direct inhibitor of this pathway. Thus, this study positions Igalan as a promising next-generation therapeutic candidate for the treatment of inflammasome-mediated pathologies.Igalan is a sesquiterpene lactone containing an α,β-unsaturated carbonyl moiety, that can undergo Michael addition with nucleophilic cysteines.32 Then, we performed unbiased chemoproteomic profiling and identified a subset of inflammation-associated proteins, ultimately determining NLRP3 for further investigation. Although this initial profiling was performed in cell lysates to assess intrinsic electrophilic reactivity and avoid metabolic sequestration,33,34 we subsequently validated its physiological relevance through downstream functional assays in intact macrophages and in vivo models.
Proteomic analysis revealed that structural proteins (e.g., ribosomes) were modified at 2 µM, while Igalan exhibited no cytotoxicity at 25 µM. This aligned with paradigms where massively abundant proteins acted as nucleophilic sinks,35,36 yielding high hit counts but serving as functionally silent off-targets. In contrast, signal transduction nodes like the NOD-like receptor (NLR) pathway yielded fewer hits due to their inherently low abundance. Crucially, ligandability did not inherently dictate druggability.37 Therefore, the low-abundance NLR network was prioritized as the functionally pivotal driver of macrophage innate immune activation, while robust compensatory mechanisms allowed cells to tolerate silent off-target modifications without overt cytotoxicity.38
Critically, molecular docking simulations demonstrate that Igalan forms a covalent adduct with Cys704 in the trLRR domain. The covalent modification of Cys704 establishes a steric “lock” mechanism, characterized by a calculated burial surface. Meanwhile, the modification not only rigidifies the trLRR architecture but also allosterically impedes the conformational transitions essential for the biological activity of NLRP3. Thus, the adjacent NACHT domain fails to undergo de-repression and is maintained in its functionally auto-inhibited state, which subsequently suppresses the intrinsic ATPase activity of the NACHT domain. Igalan treatment concentration-dependently reduced NLRP3 protein levels without altering α-tubulin expression. The observed reduction may be attributed to the Igalan-dependent covalent modification of Cys704, which could lead to NLRP3 destabilization by exposing degrons for subsequent proteasomal or lysosomal degradation.38–41 Further investigation is warranted to fully elucidate the underlying molecular mechanisms involved.
Collectively, these findings broaden the utility of natural products in the development of covalent NLRP3 inhibitors, wherein electrophilic species mediate inflammatory cascades via site-specific cysteine modifications.
Systems-level analysis revealed that Igalan attenuates the inflammatory response in LPS-stimulated macrophages through the covalent modification of NLRP3 inflammasome, while reducing mitochondrial and cellular oxidative damage.42,43 Moreover, our study demonstrates that Igalan mitigates LPS-induced pulmonary inflammatory histopathology and simultaneously preserves intestinal barrier integrity. Given that inflammatory injury and oxidative stress are hallmark pathological features of numerous immune-mediated diseases including rheumatoid arthritis, neuroinflammation, and inflammatory liver injury,18,44–46 we propose that Igalan may exert protective effects across these conditions. To validate this hypothesis and accelerate translational develop, future studies should systematically evaluate the effect of Igalan in diverse preclinical models of inflammation.
Igalan showed divergent pharmacological effects (lower IC50 in iBMDMs vs. RAW264.7 cells), reflecting critical phenotypic differences between macrophage models. Because RAW264.7 cells lack PYCARD (ASC) expression and cannot assemble the canonical NLRP3 inflammasome, our core mechanistic assays were strictly conducted in iBMDMs, a standard model for innate immune signaling.47 However, we acknowledge recent findings indicating that iBMDMs may display an altered, somewhat anti-inflammatory profile relative to primary isolated BMDMs.48 Although in vivo data strongly support the therapeutic potential of Igalan in sepsis, future studies employing primary BMDMs or alternative ASC-competent cell lines will be necessary to fully elucidate the complex, context-dependent dynamics of inflammasome inhibition.
While current data establish NLRP3 as a primary functional target, mapping the complete off-target landscape of Igalan remains an ongoing objective. Future studies utilizing biotinylated probes for affinity pull-down assays will be highly valuable to comprehensively evaluate its proteome-wide specificity. In summary, Igalan serves as a covalent agent for the selective inhibition of inflammasomes by reducing stress. In particular, the structural and functional properties position Igalan as a promising scaffold for sepsis drug development.
Conclusions
Our study identifies Igalan as a promising therapeutic candidate for sepsis. Specifically, we reveal a previously unrecognized mechanism whereby it suppresses NLRP3 inflammasome activation by inhibiting ATPase activity in the NACHT domain, thereby preventing the conformational changes required for inflammasome assembly. These findings collectively establish Igalan as a valuable agent that targets the inflammatory cascades central to sepsis pathogenesis.Author contributions
Kewu Zeng designed the project. Chengchen Hou performed the major experiments, analyzed the data, and drafted the manuscript. Yang Chen carried out the mouse sepsis animal experiments and assisted in data analysis. Wenjie Bi and Lin Gao performed the zebrafish experiments under the guidance of Liwen Han and Zhiyuan Lu. Simiao Yu assisted in biochemical and cellular assays. Xiaowen Zhang and Zekun Chen performed molecular biology and cell imaging experiments. Dilnoza E. Dusmatova, Rimma F. Mukhamatkhanova, and Ildar D. Sham’yanov provided natural products and performed structural characterization. Hua Wang, Nilufar Z. Mamadalieva, and Kewu Zeng performed overall supervision and critically reviewed the final manuscript. All authors approved the final version.Conflicts of interest
There are no conflicts to declare.Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.Supplementary information (SI) includes the high-resolution MS/MS spectrum of the Igalan-modified NLRP3 peptide, mutagenesis validation of Cys704, and a chemoproteomic profiling table of 140 target proteins. See DOI: https://doi.org/10.1039/d5cb00280j.
Acknowledgements
This work was financially supported by the Beijing Municipal Natural Science Foundation – the Key Research Project of Daxing (L246029), the National Natural Science Foundation of China (82174008), the Jinan New 20 Policies for Higher Education Funding (202228048), the Guizhou Provincial Workstation for Leading Science & Technology Innovation Talents in Traditional Chinese Medicine Chemical Biology (QianKeHe Platform-KXJZ[2025]033), and the Special Fund for Taishan Scholars Project in Shandong Province (tstp20230633). The authors acknowledge the S.Y. Yunusov Institute of the Chemistry of Plant Substances of the Academy of Sciences of the Republic of Uzbekistan for institutional support and provision of the compound. Contributions of the institute's staff are reflected in the authorship of this paper.References
- M. Singer, C. S. Deutschman, C. W. Seymour, M. Shankar-Hari, D. Annane, M. Bauer, R. Bellomo, G. R. Bernard, J.-D. Chiche and C. M. Coopersmith, et al., JAMA, 2016, 315, 801–810 CrossRef CAS PubMed.
- T. van der Poll, M. Shankar-Hari and W. J. Wiersinga, Immunity, 2021, 54, 2450–2464 CrossRef CAS PubMed.
- B. Evrard, P. Sinha, K. Delucchi, C. M. Hendrickson, K. N. Kangelaris, K. D. Liu, A. Willmore, N. Wu, L. Neyton and E. Schmiege, et al., Crit. Care, 2024, 28, 164 CrossRef PubMed.
- R. Xie, D. Tan, B. Liu, G. Xiao, F. Gong, Q. Zhang, L. Qi, S. Zheng, Y. Yuan and Z. Yang, et al., MedComm, 2025, 6, e70074 CrossRef CAS PubMed.
- E. M. O'Brien and K. L. Spiller, J. Leukocyte Biol., 2022, 111, 989–1000 CrossRef PubMed.
- C. N. Castro, T. Sahner, C. Obodozie, P. Metzger and H. Weber, Cancer Res., 2024, 84, 6538 CrossRef.
- G. Li and H.-X. Lou, Med. Res. Rev., 2018, 38, 1255–1294 CrossRef PubMed.
- D. Tahri and F. Elhouiti, Phytochem. Rev., 2025, 24, 5019–5044 CrossRef CAS.
- J. Beltran-Garcia, R. Osca-Verdegal, J. L. Garcia-Gimenez and F. V. Pallardo, Redox Exp. Med., 2022, 2022, R84–R95 Search PubMed.
- Y. Liao, Y. Kong, H. Chen, J. Xia, J. Zhao and Y. Zhou, Int. Immunopharmacol., 2025, 146, 113821 CrossRef CAS PubMed.
- Y. Zhao, Y. Wu, L. Qu, Y. Hu, S. Sun, R. Yao and R. Li, Inflammation, 2025, 48, 2395–2406 CrossRef CAS PubMed.
- N. J. Makoni and M. R. Nichols, Arch. Biochem. Biophys., 2021, 699, 108753 CrossRef CAS PubMed.
- L. Boermel, S. Kluge, S. Liao, T. Schubert, S. Lorkowski and M. Wallert, Appl. Res., 2024, 3, e202300036 CrossRef CAS.
- F. A. Fischer, K. W. Chen and J. S. Bezbradica, Front. Immunol., 2021, 12, 661162 CrossRef CAS PubMed.
- Y. Bai, Y. Pan and X. Liu, Nat. Rev. Mol. Cell Biol., 2025, 26, 501–521 CrossRef CAS PubMed.
- N. Zhang and D. Xu, Dev. Cell, 2025, 60, 994–1007 CrossRef CAS PubMed.
- W. Wang, L. Liu, Z. Yang, C. Lu, P. Tu and R. Zhao, Chin. J. Nat. Med., 2024, 22, 127–136 CAS.
- K. W. Zeng, Q. Yu, L. X. Liao, F. J. Song, H. N. Lv and Y. Jiang, J. Cell. Biochem., 2015, 116, 1286–1299 CrossRef CAS PubMed.
- K. W. Zeng, X. Fu, Y. Jiang and P. F. Tu, Eur. J. Pharmacol., 2015, 748, 18–29 CrossRef CAS PubMed.
- Y.-Q. Zhao, X. Li, H.-Y. Guo, Q.-K. Shen, Z.-S. Quan and T. Luan, Molecules, 2023, 28(18), 6478 CrossRef CAS PubMed.
- M. Chadwick, H. Trewin, F. Gawthrop and C. Wagstaff, Int. J. Mol. Sci., 2013, 14, 12780–12805 CrossRef PubMed.
- J. S. Camara, R. Perestrelo, R. Ferreira, C. V. Berenguer, J. A. M. Pereira and P. C. Castilho, Molecules, 2024, 29(16), 3861 CrossRef CAS PubMed.
- J. Li, X. Li and H. Liu, Front. Pharmacol., 2025, 16, 1551115 CrossRef CAS PubMed.
- T. T. P. Dao, K. Song, J. Y. Kim and Y. S. Kim, Inflamm. Res., 2020, 69, 309–319 CrossRef CAS PubMed.
- K.-M. Lee, J. M. Shin, J. Chun, K. Song, C. W. Nho and Y. S. Kim, J. Biochem. Mol. Toxicol., 2019, 33, e22297 CrossRef PubMed.
- K. M. Lee, J. M. Shin, J. Chun, K. Song, C. W. Nho and Y. S. Kim, J. Biochem. Mol. Toxicol., 2019, 33, e22297 CrossRef PubMed.
- T. T. P. Dao, K. Song, J. Y. Kim and Y. S. Kim, Inflamm. Res., 2020, 69, 309–319 CrossRef CAS PubMed.
- W. He, H. Dong, C. Wu, Y. Zhong and J. Li, Int. Immunopharmacol., 2023, 115, 109697 CrossRef CAS PubMed.
- F. Belicard, K. Tarte and J.-M. Tadie, J. Leukocyte Biol., 2024, 115, 999–1001 CrossRef CAS PubMed.
- J. A. Duncan, D. T. Bergstralh, Y. Wang, S. B. Willingham, Z. Ye, A. G. Zimmermann and J. P.-Y. Ting, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 8041–8046 CrossRef CAS PubMed.
- R. C. Coll, J. R. Hill, C. J. Day, A. Zamoshnikova, D. Boucher and N. L. Massey, et al., Nat. Chem. Biol., 2019, 15, 556–559 CrossRef CAS PubMed.
- Q. V. Vu, S. Sayama, M. Ando and T. Kataoka, Molecules, 2024, 29(8), 1866 CrossRef CAS PubMed.
- K. M. Backus, B. E. Correia, K. M. Lum, S. Forli, B. D. Horning, G. E. González-Páez, S. Chatterjee, B. R. Lanning, J. R. Teijaro, A. J. Olson, D. W. Wolan and B. F. Cravatt, Nature, 2016, 534, 570–574 CrossRef CAS PubMed.
- R. E. Moellering and B. F. Cravatt, Chem. Biol., 2012, 19, 11–22 CrossRef CAS PubMed.
- K. M. Backus, B. E. Correia, K. M. Lum, S. Forli, B. D. Horning, G. E. Gonzalez-Paez, S. Chatterjee, B. R. Lanning, J. R. Teijaro, A. J. Olson and B. F. Cravatt, Nature, 2016, 534, 570–574 CrossRef CAS PubMed.
- E. Weerapana, C. Wang, G. M. Simon, F. Richter, S. Khare, M. B. D. Dillon, D. A. Bachovchin, K. Mowen, D. Baker and B. F. Cravatt, Nature, 2010, 468, 790–795 CrossRef CAS PubMed.
- K. K. Hallenbeck, D. M. Turner, A. R. Renslo and M. R. Arkin, Curr. Top. Med. Chem., 2017, 17, 4–15 CrossRef CAS PubMed.
- L. Boike, N. J. Henning and D. K. Nomura, Nat. Rev. Drug Discovery, 2022, 21, 881–898 CrossRef CAS PubMed.
- M. Békés, D. R. Langley and C. M. Crews, Nat. Rev. Drug Discovery, 2022, 21, 181–200 CrossRef PubMed.
- L. Boike, A. G. Cioffi, F. C. Majewski, J. Su, S. Klaeger, S. E. Hernandez, N. J. Henning and D. K. Nomura, Cell Chem. Biol., 2021, 28, 4–13 CrossRef CAS PubMed.
- J. N. Spradlin, X. Hu, C. C. Ward, S. M. Brittain, M. D. Jones, L. Ou, M. To, A. Proudfoot, E. Ornelas, M. Woldegiorgis, J. A. Olzmann, D. E. Bussiere, J. R. Thomas, J. A. Tallarico, J. M. McKenna, M. Schirle, T. J. Maimone and D. K. Nomura, Nat. Chem. Biol., 2019, 15, 747–755 CrossRef CAS PubMed.
- P. Pradhan, V. Vijayan, K. Cirksena, F. F. R. Buettner, K. Igarashi, R. Motterlini, R. Foresti and S. Immenschuh, Redox Biol., 2022, 51, 102265 CrossRef CAS PubMed.
- S. Han, X. Li, N. Xia, Y. Zhang, W. Yu, J. Li, C. Jiao, Z. Wang and L. Pu, Int. J. Mol. Sci., 2023, 24(7), 6348 CrossRef CAS PubMed.
- K. Weisel, N. E. Scott, D. J. Tompson, B. J. Votta, S. Madhavan, K. Povey, A. Wolstenholme, M. Simeoni, T. Rudo, L. Richards-Peterson, T. Sahota, J. G. Wang, J. Lich, J. Finger, A. Verticelli, M. Reilly, P. J. Gough, P. A. Harris, J. Bertin and M.-L. Wang, Pharmacol. Res. Perspect, 2017, 5(6), e00365 CrossRef PubMed.
- J. Fu, X. Chen, J. Li and L. Peng, Front. Med., 2025, 12, 1569732 CrossRef PubMed.
- J. Li, K.-W. Zeng, S.-P. Shi, Y. Jiang and P.-F. Tu, Fitoterapia, 2012, 83, 896–900 CrossRef CAS PubMed.
- D. De Nardo, D. V. Kalvakolanu and E. Latz, Methods Mol. Biol., 2018, 35–49 CrossRef CAS PubMed.
- M. Herb, V. Schatz, K. Hadrian, D. Hos, B. Holoborodko, J. Jantsch and N. Brigo, Front. Cell. Infect. Microbiol., 2024, 14, 1457323 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |