Open Access Article
Open Access ArticleGlutathione-activatable bola dendrimers mediate tumor-specific cytosolic siRNA delivery via dynamic thiol–disulfide exchange
Dandan
Zhu†
 a,
Huiling
Zhu†
a,
Yaoyun
Yu†
a,
Junyue
Zheng†
a,
Xianhui
Lin
a,
Huimin
Cheng
a,
Aoxue
Mei
a,
Ming
Chen
a,
Yun
Li
a,
Haijuan
Dong
b,
Jiehua
Zhou
c,
Juan
Liu
a,
Huiling
Zhu†
a,
Yaoyun
Yu†
a,
Junyue
Zheng†
a,
Xianhui
Lin
a,
Huimin
Cheng
a,
Aoxue
Mei
a,
Ming
Chen
a,
Yun
Li
a,
Haijuan
Dong
b,
Jiehua
Zhou
c,
Juan
Liu
 *d and
Xiaoxuan
Liu
*d and
Xiaoxuan
Liu
 *a
*a
aState Key Laboratory of Natural Medicines, Joint International Research Laboratory of Target Discovery and New Drug Innovation (Ministry of Education), Jiangsu Key Laboratory of Drug Discovery for Metabolic Diseases, Center of Advanced Pharmaceuticals and Biomaterials, China Pharmaceutical University, Nanjing 211198, China. E-mail: xiaoxuanliu@cpu.edu.cn
bThe Public Laboratory Platform, China Pharmaceutical University, Nanjing 211198, China
cLeaderna Therapeutics Ltd, Chengdu 610219, China
dHepato-Pancreato-Biliary Center, Beijing Tsinghua Changgung Hospital, Key Laboratory of Digital Intelligence Hepatology (Ministry of Education), School of Clinical Medicine, Tsinghua Medicine, Tsinghua University, Beijing 102218, China. E-mail: lja02720@btch.edu.cn
First published on 5th January 2026
Abstract
Effective delivery of small interfering RNA (siRNA) to the cytosol continues to pose a significant challenge in RNA interference (RNAi)-driven precision cancer therapy. In this study, we engineered glutathione (GSH)-responsive bola-amphiphilic peptide dendrimers (bola DS-Cn-K4) for tumor-specific cytosolic siRNA delivery. These dendrimers incorporate a hydrophilic polylysine dendron for efficient siRNA binding and a hydrophobic disulfide-bridged bola-lipid core with varying alkyl chain lengths, facilitating thiol-mediated cellular uptake and enabling siRNA release in response to intracellular higher GSH levels. Our structure–activity relationship studies revealed that bola DS-C6-K4, characterized by the shortest alkyl chain, exhibited superior siRNA delivery, which was attributed to optimized thiol-mediated cellular uptake and accelerated GSH-triggered siRNA release stemming from improved disulfide accessibility. Mechanistic investigations validated thiol-mediated uptake as the predominant cellular internalization pathway, effectively bypassing endosomal entrapment. The siRNA/bola DS-C6-K4 complexes efficiently downregulate oncoprotein expression, thereby impeding cancer cell proliferation, migration, and invasion, and simultaneously inducing apoptosis. In A549 xenograft models, intravenous administration of siPLK1/bola DS-C6-K4 achieved substantial reductions in tumor growth and PLK1 expression while exhibiting minimal systemic toxicity. This study highlights a synergistic approach utilizing bola-amphiphilic peptide dendrimers for tumor-specific and cytosolic siRNA delivery, leveraging membrane–thiol interactions and intracellular GSH-triggered siRNA release.
Introduction
Small interfering RNA (siRNA) has been proved as an effective and promising therapeutic modality for the prevention and treatment of human diseases.1,2 Through RNA interference (RNAi) mechanisms, siRNA effectively silences disease-related proteins at the post-transcriptional level.2,3 The high sequence specificity, design flexibility, abbreviated development timeline, and sustained therapeutic efficacy of siRNA have facilitated rapid clinical translation, with seven therapeutics receiving US Food and Drug Administration (FDA) approval since 2018.4,5 However, the therapeutic potential of unmodified siRNA is limited by its inherent instability against nuclease degradation, rapid reticuloendothelial clearance, and poor membrane permeability. Advanced delivery vectors have consequently been developed to enhance siRNA druggability, as exemplified by the lipid-based formulation Patisiran®, the first FDA-approved siRNA therapeutic for polyneuropathy.6 These delivery systems mitigate key obstacles through a stable complex formulation that shields siRNA from degradation, thereby enhancing RNAi efficiency.7 Despite these advances, current vectors frequently demonstrate suboptimal cytoplasmic siRNA release due to membrane barriers and endolysosomal entrapment during endocytic uptake.8,9 Consequently, the rational design of specialized vectors is essential to achieve efficient cytosolic delivery to target cells, ultimately expanding therapeutic efficacy and clinical translation.The pronounced hydrophobicity of cellular and organellar membranes constitutes a significant barrier to efficient cytosolic delivery. To surmount this challenge, strategic incorporation of phospholipids or branched alkyl chains into delivery vectors facilitates membrane integration and fusion, thereby enhancing cellular uptake and endosomal escape of siRNA. Furthermore, functionalization of vectors with membrane-permeable peptides enhances membrane interactions and modifies cellular internalization pathways.10 To fully exploit siRNA's therapeutic potential, vectors must additionally enable target cell-specific on-demand cargo release. Emerging thiol-responsive transporters represent an emerging alternative strategy, bypassing conventional endocytic routes via thiol-mediated mechanisms while enabling reduction-triggered cargo release to enhance cytosolic delivery efficiency.11–14 As representative redox-sensitive motifs, disulfide bonds undergo11 dynamic covalent exchange with overexpressed exofacial thiols on tumor cell membrane proteins. Critically, disulfide bonds maintain stability during systemic circulation yet undergo specific and rapid cleavage in response to tumor-overexpressed glutathione (GSH) in cancer cells, facilitating intracellular siRNA release and tumor-specific delivery.12,15,16 These attributes well established disulfide bond incorporation as a compelling approach to enhance thiol-mediated cellular uptake and cancer cell-specific delivery. Consequently, the rational engineering of disulfide-based vectors requires comprehensive mechanistic understanding to optimize thiol–disulfide exchange kinetics while preserving favorable pharmacokinetic profiles.
Amphiphilic dendrimers exhibit significant potential for precision siRNA delivery, leveraging their well-defined dendritic topology and multivalent cooperativity.17,18 Clinically, this is exemplified by MIR 19® (Russian Ministry of Health approval no. LP-006423), an amphiphilic dendrimer-based formulation delivering SARS-CoV-2-targeting siRNA for COVID-19 therapy.19 Such systems synergize the delivery merits of lipidic20 and polymeric21 platforms, currently the most advanced siRNA delivery technologies.17 Prior work from our group enhanced cytosolic delivery via phospholipid incorporation22 or arginine-functionalized termini23 to emulate cell-penetrating peptides. Bola-amphiphilic architectures, characterized by symmetric lipid chains flanking a central scaffold, provide enhanced structural stability and enable precise modulation of membrane interactions through chain length control.24–26 Previously, we utilized bola-amphiphilic dendrimers to develop cargo-selective nucleic acid delivery systems, which demonstrated promising results in tumor xenograft and metastasis models.27,28 Extending these developments, we engineered bola-amphiphilic dendrimers featuring an innovative disulfide-bridged lipid core designed to concurrently enable (i) thiol-mediated cellular uptake, (ii) glutathione-responsive siRNA release, and (iii) preserved biocompatibility through peptide-derived structural elements.
Herein, we report glutathione-activatable bola-amphiphilic peptide dendrimers (bola DS-Cn-K4; n = 6, 8, 11) engineered for tumor-targeted cytosolic siRNA delivery (Scheme 1). These dendrimers integrate three functional components: (i) hydrophilic polylysine dendrons mimicking protein transduction domains, (ii) hydrophobic disulfide-bridged bola-lipid cores with tunable alkyl spacers (C6, C8, C11), and (iii) multivalent cooperativity enabling self-assembly into siRNA-encapsulating nanoparticles. The disulfide moiety facilitates thiol-mediated cellular internalization while conferring glutathione-responsive cargo release. Rational alteration of the alkyl spacer length (C6, C8, C11) enabled precise modulation of the trade-off between nanoparticle stability and disulfide exchange kinetics. Structure–activity relationship analysis revealed bola-DS-C6-K4 as the optimal vector, exhibiting superior siRNA delivery efficiency and gene silencing potency. This enhancement stemmed from shortened alkyl spacers augmenting disulfide accessibility, thereby accelerating thiol-mediated internalization and improving membrane dynamics. Consequently, bola DS-C6-K4 achieved specific cytosolic delivery of siRNA in various cancer cells, suppressed the proliferation of non-small cell lung cancer A549 cells, induced apoptosis and simultaneously inhibited their migration and invasion in vitro. Notably, in A549 lung cancer xenografts, bola DS-C6-K4 mediated functional siRNA delivery, achieving robust oncogene silencing and significant antitumor efficacy without observable toxicity. This system thus establishes a synergistic strategy for conditional RNAi therapeutics via membrane–thiol interactions and intracellular glutathione-triggered siRNA release.
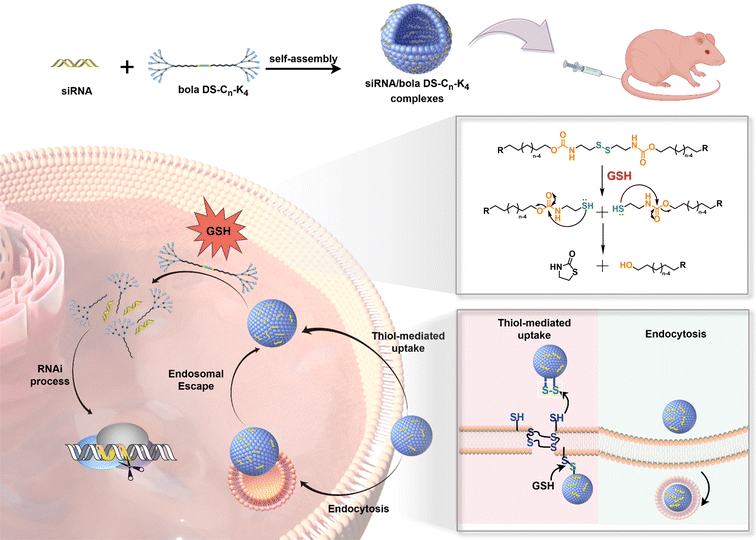 | ||
| Scheme 1 Cartoon illustration of bola amphiphilic peptide dendrimers bola DS-Cn-K4 for tumor cell-specific and cytosolic siRNA delivery. | ||
Results and discussion
Robust synthesis, physicochemical properties and biosafety profiling of bola DS-Cn-K4
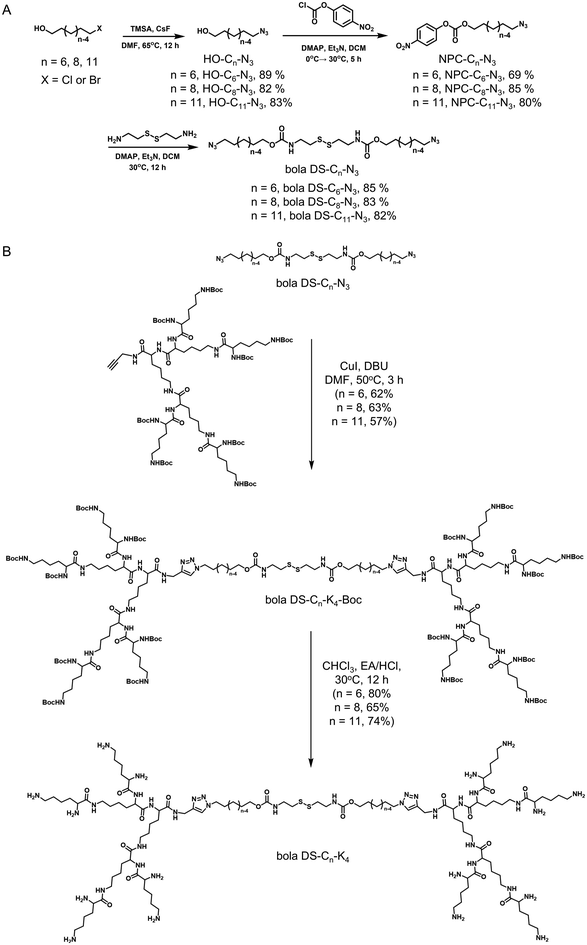
Bola-amphiphilic peptide dendrimers (bola DS-Cn-K4), comprising hydrophobic bola-lipid cores with varying chain lengths (n = 6, 8, 11), were synthesized via the synthetic route outlined in Scheme S1. Hydrophilic alkynyl-containing polylysine dendrons were synthesized following our previously established protocols,22,29 while disulfide- and azide-modified hydrophobic bola-lipid cores were prepared via the steps described in Scheme 2.Initially, halogenated alcohols served as the starting materials and were subjected to azidation with azide trimethylsilane to generate HO-Cn-N3 (n = 6, 8, 11) intermediates. These intermediates were subsequently reacted with 4-nitrophenyl chloroformate to produce PNC-Cn-N3, which was conjugated with freshly prepared cysteamine in the presence of the catalyst 4-dimethylaminopyridine (DMAP), resulting in the disulfide- and azide-functionalized bola-lipid cores (DS-Cn-N3) of varying chain lengths. The DS-Cn-N3 cores were then coupled with the alkynyl-containing polylysine dendrons through the Cu(I)-catalyzed azide–alkyne cycloaddition (CuAAC) click reaction to produce Boc-terminated peptide dendrimers (bola DS-Cn-K4-Boc) (Scheme 2 and Scheme S1). The Boc-protected dendrimers were subsequently deprotected, affording the amphiphilic peptide dendrimers bola DS-Cn-K4 with an impressive 95% yield and excellent quality (Fig. 1A). Comprehensive synthetic details are provided in the SI, and the chemical structures and purity of the newly synthesized bola DS-Cn-K4 dendrimers were thoroughly characterized and confirmed through 1H-NMR and 13C-NMR spectroscopy, high-resolution mass spectrometry (HRMS), and HPLC, as shown in Fig. S1–S3.
The bola DS-Cn-K4 dendrimers incorporate hydrophilic polylysine dendrons with protein-mimicking properties, prompting us to evaluate their secondary configurations through CD analysis. The CD spectra (Fig. 1B and Table S1) revealed that these dendrimers exhibit secondary structures analogous to those of proteins. Furthermore, the amphiphilic nature of bola DS-Cn-K4 dendrimers, conferred by their hydrophilic polylysine dendrons and hydrophobic bola-lipid cores, was further characterized by their self-assembly properties. CACs were determined via a fluorescent spectroscopic assay using pyrene, yielding values of 40, 44, and 49 µM for bola DS-C6-K4, bola DS-C8-K4, and bola DS-C11-K4, respectively (Fig. 1C). The amphiphilic nature of DS-Cn-K4 facilitated spontaneous assembly into nanoparticles in aqueous solution. These nanoparticles displayed hydrodynamic sizes ranging from 56 to 107 nm and zeta potentials of 23–30 mV, as measured by dynamic light scattering (Fig. 1D and Table S2). These findings suggest that the DS-Cn-K4 dendrimers exhibit comparable self-assembly capabilities.
The bola DS-Cn-K4 dendrimers, with their polylysine dendritic structure, are expected to exhibit a favorable safety profile, which is a critical requirement for developing novel siRNA delivery systems. To evaluate this, we assessed the metabolic toxicity of bola DS-Cn-K4 using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay to assess metabolic toxicity and a hemolysis assay to evaluate cell membrane damage. The MTT assays on normal cells, including mouse fibroblast L929 cells and Madin–Darby canine kidney MDCK cells, showed that all bola DS-Cn-K4 dendrimers have almost no toxicity within the tested concentration range (Fig. S4A and S4B). Notably, none of the bola DS-C6-K4, bola DS-C8-K4, or bola DS-C11-K4 dendrimers exhibited detectable hemolysis within the tested concentration range (Fig. S4C). These findings indicate that all three bola-amphiphilic peptide dendrimers exhibit favorable safety profiles.
Enhanced cellular uptake and GSH-responsive siRNA release via alkyl spacer-dependent thiol activation
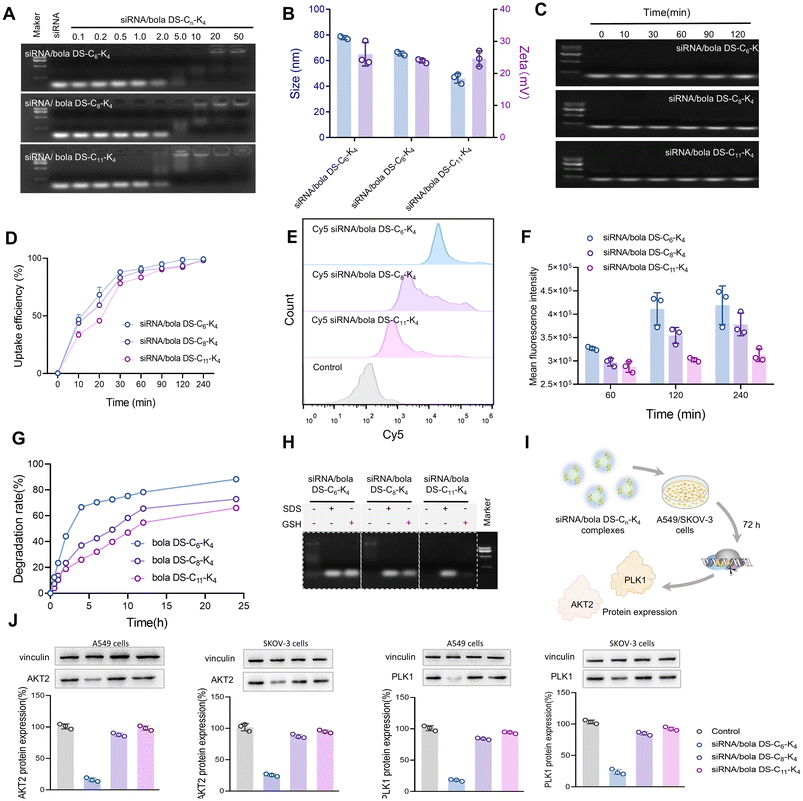
We next evaluated the siRNA delivery performance of bola DS-Cn-K4 by assessing siRNA binding capacity, cellular uptake, GSH-responsive disassembly, and siRNA release. A key property of effective vectors is their ability to condense siRNA. We initially evaluated this by assessing the binding efficiency of bola DS-Cn-K4 with siRNA using agarose gel electrophoresis. The results, as shown in Fig. 2A, demonstrate that all the dendrimers completely bound siRNA at an N/P ratio ≥ 5.0, indicating that bola DS-Cn-K4 can form stable complexes with siRNA. Transmission electron microscopy (TEM) analysis of the siRNA/bola DS-Cn-K4 complexes (Fig. S5) revealed the presence of uniform spherical nanoparticles of an optimal size of about 40–80 nm. Dynamic Light Scattering (DLS) analysis (Fig. 2B and Table S2) further corroborated that the size of the siRNA/bola DS-Cn-K4 complexes, ranging approximately from 77 nm to 48 nm, progressively decreased with the elongation of the dendrimer alkyl chain length from C6 to C11. Upon interaction with negatively charged siRNA, these dendrimer-based nanoassemblies undergo reorganization into nanomicellar structures that maximize their positive surface charges, thereby forming stable complexes with siRNA, according to previous research studies.30,31 This reduction in size is likely attributable to the enhanced self-assembly capability of bola DS-Cn-K4 at longer chain lengths, which strengthens the electrostatic interactions between the positively charged dendrimer-based nanoassemblies and the negatively charged siRNA, thereby leading to the formation of more compact complexes with reduced dimensions. Despite these size variations, the siRNA/bola DS-Cn-K4 complexes showed comparable surface potentials of +27 mV, +24 mV, and +25 mV, respectively (Fig. 2B and Table S2). The formed complexes of bola DS-Cn-K4 successfully protected siRNA from enzymatic degradation (Fig. 2C), unlike naked siRNA, which underwent rapid enzymatic degradation within minutes (Fig. S6). These findings collectively indicate that bola DS-Cn-K4 effectively encapsulates siRNA through electrostatic interactions, resulting in the formation of nanoscale and stable complexes.Notably, we hypothesized that bola DS-Cn-K4, bearing a disulfide bond-containing bola lipid, could facilitate enhanced cellular uptake through disulfide exchange with exofacial thiols on cell membranes and accelerate GSH-triggered degradation for efficient siRNA release. To validate this hypothesis, we evaluated the internalization of Cy5-labeled siRNA mediated by bola DS-Cn-K4 in A549 cells using flow cytometry. Although all three siRNA/bola DS-Cn-K4 complexes showed comparable cellular uptake rates due to their similar surface potentials (Fig. 2D), notably, A549 cells treated with siRNA/bola DS-C6-K4 exhibited significantly stronger Cy5 fluorescence signals compared to those treated with siRNA/bola DS-C8-K4 or siRNA/bola DS-C11-K4 (Fig. 2E). This observation indicates more effective intracellular accumulation of siRNA/bola DS-C6-K4 (Fig. 2F). The enhanced uptake of siRNA/bola DS-C6-K4 can be attributed to its shorter alkyl chain, which facilitates the compaction of siRNA into more flexible and dynamic complexes. This structural flexibility promotes thiol-mediated cellular internalization of siRNA/bola DS-C6-K4, outperforming complexes formed with bola DS-C8-K4 or bola DS-C11-K4.
To further investigate the superior siRNA delivery capabilities, we conducted investigations into the GSH-responsive disassembly of bola DS-Cn-K4 assemblies using the Ellman reagent. Notably, the degradation rate of bola DS-C6-K4 assemblies was significantly higher compared to those of the other dendrimers after exposure to dithiothreitol (DTT), as evidenced in Fig. 2G. This observation correlates with the enhanced siRNA delivery efficacy of bola DS-C6-K4 over bola DS-C8-K4 and bola DS-C11-K4, as siRNA complexes formed with bola DS-C6-K4 are more readily disassembled and release siRNA more efficiently upon GSH stimulation. To substantiate this, we examined GSH-triggered siRNA release from the siRNA/bola DS-Cn-K4 complexes through gel electrophoresis assays and DLS. As shown in Fig. 2H, the encapsulated siRNA within the siRNA/bola DS-C6-K4 complexes was released more rapidly upon exposure to GSH, in contrast to the slower release observed with the siRNA/bola DS-C8-K4 or siRNA/bola DS-C11-K4 complexes. DLS analysis (Fig. S7) further confirmed the better GSH-responsive disassembly of the siRNA/bola DS-C6-K4 complexes. Upon GSH treatment, the size of the siRNA/bola DS-C6-K4 complexes visibly increased significantly from approximately 80–100 nm to approximately 800 nm, whereas the siRNA/bola DS-C8-K4 or siRNA/bola DS-C11-K4 complexes showed comparatively smaller size changes. This discrepancy in release rates can be attributed to the greater stability of the siRNA complexes formed with bola DS-C8-K4 or bola DS-C11-K4, which impedes GSH-triggered cleavage of the disulfide linker. In contrast, the siRNA complexes formed with bola DS-C6-K4, bearing shorter alkyl chains, interact more readily with GSH, resulting in more efficient disassembly, improved siRNA efficacy, and enhanced gene silencing.
Subsequently, we explored the siRNA delivery efficiency of bola DS-Cn-K4 in non-small cell lung cancer A549 cells and ovarian cancer SKOV-3 cells using siRNA targeted against polo-like kinase 1 (PLK1)32,33 and protein kinase B (AKT2),34,35 both of which are crucial for tumor growth, proliferation, invasion, metastasis, and angiogenesis (Fig. 2I). As shown in Fig. 2J, the AKT2 protein expression levels noticeably decreased in SKOV-3 cells and A549 cells following treatment with siRNA/bola DS-C6-K4, while no significant changes were observed with siRNA/bola DS-C8-K4 or siRNA/bola DS-C11-K4. Further evaluation of PLK1 siRNA delivery by bola DS-Cn-K4 also confirmed that a potent gene silencing effect was only observed with siRNA/bola DS-C6-K4, whereas no prominent down-regulation of the PLK1 protein was noted in other complexes (Fig. 2J). Importantly, the siRNA/bola DS-C6-K4 complexes exhibited good serum stability and achieved a level of gene silencing comparable to that of the FDA-approved DLin-MC3-DMA (Fig. S8). These results underscore the exceptional siRNA delivery efficiency of bola DS-C6-K4.
Cancer cell-specific cytosolic siRNA delivery mediated by bola DS-C6-K4
The quest for tumor-specific siRNA delivery represents a cornerstone in the development of precision medicine. With this concept in mind, bola DS-C6-K4, featuring a GSH-sensitive disulfide group, was hypothesized to enable on-demand siRNA delivery to cancer cells in response to their elevated GSH levels. Indeed, the GSH-dependent siRNA delivery mediated by bola DS-C6-K4 was examined by modulating intracellular GSH levels in A549 cells using the GSH synthetase inhibitor L-buthionine-(s,r)-sulfoximine (BSO). After treatment with BSO, a significant reduction in intracellular GSH levels in A549 cells was documented (Fig. 3A) as detected by flow cytometry. Correspondingly, the gene silencing effect of the siRNA/bola DS-C6-K4 complexes dramatically decreased (Fig. 3B and C), highlighting the potential of bola DS-C6-K4 to efficiently and specifically deliver siRNA in GSH-enriched pathological environments. Further comparisons with the degradation product (OH-C6-K4) and GSH-nonresponsive bola C6-K4 revealed that only bola DS-C6-K4 effectively delivered siRNA therapeutics and produced specific gene silencing effects (Fig. S9). These results indicate that bola DS-C6-K4 enables efficient and cell-specific siRNA delivery in response to pathological GSH-enriched environments.Effective cellular uptake and endosomal escape of siRNA therapeutics into targeted cells are essential for optimizing delivery efficiency. Notably, bola DS-C6-K4 was hypothesized to enhance cellular uptake through disulfide exchange with exofacial thiols on cell membranes, contributing to the observed improvements in siRNA delivery efficiency. To assess these processes, the mechanism of bola DS-C6-K4-mediated cellular siRNA delivery was elucidated using various endocytic inhibitors, including cytochalasin D (micropinocytosis inhibitor), genistein (caveolae-mediated endocytosis inhibitor), chlorpromazine (clathrin-mediated endocytosis inhibitor), and 5,5′-dithiobis-2-nitrobenzoic acid (DTNB, thiol-mediated uptake inhibitor). The fluorescence intensity of the Cy5 siRNA/bola DS-C6-K4 complexes in A549 cells was markedly reduced upon incubation with these inhibitors (Fig. 3D). Notably, DTNB exerted the most significant inhibition of cellular uptake (Fig. 3D), with cytochalasin D, genistein, and chlorpromazine showing moderate effects. Confocal laser scanning microscopy (CLSM) further confirmed that the intracellular Cy5 fluorescence intensity was significantly diminished following DTNB treatment (Fig. 3E and Fig. S10). These findings demonstrate that the siRNA/bola DS-C6-K4 complexes predominantly internalize into tumor cells via thiol-mediated uptake mechanisms. Fig. 3F and Fig. S11 reveal that the distinct red fluorescence signals of the siRNA/bola DS-C6-K4 complexes remained separate from the green fluorescence signals of lysosomes across various incubation times, indicating negligible endosomal entrapment. We also visualized the co-localization of lysosomes with the siRNA/bola DS-C6-K4 complexes at various time points following treatment (Fig. S11B). Notably, the red fluorescence signals representing the siRNA/bola DS-C6-K4 complexes were distinctly separated from the green fluorescence signals of lysosomes, indicating that the complexes effectively bypass endosomal entrapment. CLSM imaging (Fig. S12) further confirmed that Cy5-labeled siRNA delivered by bola DS-C6-K4 was homogeneously distributed in A549 cells and gradually accumulated in the cytoplasm with increased incubation time. These observations support the ability of bola DS-C6-K4 to bypass intracellular barriers and facilitate siRNA translocation into the cytosol through thiol-mediated uptake, involving disulfide exchange between bola DS-C6-K4 and exofacial thiols on the cell surface, thereby enhancing interactions between cancer cell membranes and the complexes.
To investigate its specific gene silencing capabilities, bola DS-C6-K4-mediated siRNA delivery was evaluated in various cancer cell lines using siRNAs targeting PLK1 and AKT2. As depicted in Fig. S13, a significant decrease in PLK1 mRNA and protein levels was observed in A549 cells treated with the siPLK1/bola DS-C6-K4 complexes, whereas no notable gene silencing was observed with bola DS-C6-K4 alone, siPLK1 alone, or scramble siRNA/bola DS-C6-K4. Similar results were obtained in SKOV-3, HeLa, and HepG2 cells, where potent gene silencing was only achieved with the siPLK1/bola DS-C6-K4 complexes (Fig. 3G). AKT2 protein expression was substantially inhibited in A549 and SKOV-3 cells treated with the siAKT2/bola DS-C6-K4 complexes compared to other treatment groups (Fig. S14). Notably, neither the scramble siRNA/bola DS-C6-K4 complexes nor bola DS-C6-K4 alone induced appreciable metabolic cytotoxicity, membrane damage, or hemolytic toxicity (Fig. S15). No immunotoxic effects were observed in healthy mice after treatment with the scramble siRNA/bola DS-C6-K4 complexes or bola DS-C6-K4 alone (Fig. S16A) compared with the control, while the levels of inflammatory factors such as IL-1β, IL-6, TNF-α, and IFN-γ significantly increased in the mice treated with the positive control, lipopolysaccharide (LPS). Furthermore, no changes were detected in renal, hepatic, or blood lipid parameters after intravenous administration of the scramble siRNA/bola DS-C6-K4 complexes or bola DS-C6-K4 alone (Fig. S16B). Also, the normal tissue architecture and cellular morphology of major organs showed no detectable pathological alterations in the treated groups compared to the control group (Fig. S16C). These findings collectively demonstrate that bola DS-C6-K4 serves as an effective and safe mediator for siRNA delivery.
Bola DS-C6-K4 has garnered considerable attention as a promising siRNA delivery vector due to its unique integration of lipid and peptide dendrimer characteristics. To substantiate this premise, we evaluated the contributions of the hydrophilic polypeptide dendron and the hydrophobic bola-alkyl chain to the siRNA delivery activity of bola DS-C6-K4. Our findings revealed that neither the polypeptide dendron nor the bola-alkyl chain alone exhibited notable gene silencing activity compared to bola DS-C6-K4 (Fig. S17A). Furthermore, incorporation of 1,2-dioleoyl-sn-glycerol-3-phosphoethanolamine (DOPE), a helper lipid commonly used in lipid nanoparticles (LNPs) to enhance delivery efficacy by promoting membrane fusion,36,37 significantly improved the siRNA delivery efficiency and gene silencing of the siRNA/bola DS-C6-K4 complexes (Fig. S17B). These findings highlight the synergistic integration of lipid and peptide dendrimer characteristics in the protein-mimetic design of bola DS-C6-K4, enabling efficient and safe siRNA delivery.
Therapeutic efficacy against non-small cell lung cancer in vitro and in vivo
Building upon the exceptional siRNA delivery efficiency and tumor-specific targeting properties of bola DS-C6-K4, we investigated its therapeutic potential in the treatment of lung cancer. Lung cancer, particularly non-small cell lung cancer (NSCLC), which accounts for approximately 85% of all cases, remains the most prevalent and lethal malignancy worldwide, leading to approximately 350 fatalities each day.38,39 To evaluate the anti-tumor efficacy of the bola DS-C6-K4-mediated siRNA delivery system, we targeted the PLK1 gene in NSCLC A549 cells. PLK1, a critical multifunctional protein kinase, plays a significant role in tumor cell proliferation, invasion, metastasis, and angiogenesis, positioning it as a promising therapeutic target for lung cancer treatment. Our results revealed significant suppression of tumor cell proliferation in A549 cells treated with the siPLK1/bola DS-C6-K4 complexes compared to treatment with bola DS-C6-K4 alone or the scramble/bola DS-C6-K4 complexes (Fig. 4A). Furthermore, flow cytometry analysis validated the ability of the siPLK1/bola DS-C6-K4 complexes to markedly induce apoptosis in A549 cells (Fig. 4B and C). These results collectively underscore the promise of bola DS-C6-K4 to efficiently and safely deliver siRNA into cancer cells, thereby suppressing tumor proliferation and inducing apoptosis, highlighting its potential as an innovative therapeutic approach for lung cancer.Following the initial assessments, we conducted a comprehensive evaluation of the migration and invasion behaviors of A549 cells subsequent to PLK1 gene silencing. Initially, the migration capability was assessed using a wound healing assay. A549 cells exhibited rapid wound repair under untreated conditions and following treatment with the scramble/bola DS-C6-K4 complexes or bola DS-C6-K4 alone. In contrast, the migration capacity of A549 cells was significantly impaired following treatment with the siPLK1/bola DS-C6-K4 complexes (Fig. 4D and E). Subsequently, the invasion capability was investigated using a Matrigel-coated Transwell model. As shown in Fig. 4F and G, there was a pronounced reduction in the number of cancer cells penetrating the Matrigel barrier in the siPLK1/bola DS-C6-K4 pretreatment group compared to the control groups, including those treated with bola DS-C6-K4 alone and scramble/bola DS-C6-K4. These findings together suggest that effectively suppressing PLK1 expression using the bola DS-C6-K4-mediated siRNA delivery system significantly inhibits cancer cell proliferation and curtails the migration and invasion processes that are essential for tumor metastasis.
To comprehensively assess the in vivo delivery efficiency of the bola DS-C6-K4-mediated siRNA delivery system, we investigated the therapeutic efficacy of the siPLK1/bola DS-C6-K4 complexes in xenografted A549 tumor-bearing nude mice. As shown in Fig. 5A and B, the mice treated with the siPLK1/bola DS-C6-K4 complexes demonstrated markedly reduced tumor growth compared to those administered with PBS, bola DS-C6-K4 alone, or the scramble/bola DS-C6-K4 complexes. Moreover, the tumor weight of A549-bearing mice decreased by nearly 80% after treatment with the siPLK1/bola DS-C6-K4 complexes compared to other treatment groups (Fig. 5C). This substantial therapeutic effect is attributed to the enhanced in vivo siRNA delivery capability of bola DS-C6-K4, resulting in pronounced downregulation of PLK1 expression at both mRNA and protein levels (Fig. 5D–F and Fig. S18). In contrast, treatments with PBS, bola DS-C6-K4 alone, and the scramble/bola DS-C6-K4 complexes did not alter PLK1 mRNA or protein levels (Fig. 5D–F and Fig. S18). Immunohistochemical analysis of Ki67 further confirmed an obvious decrease in tumor cell proliferation in mice treated with the siPLK1/bola DS-C6-K4 complexes, whereas other treatment groups exhibited elevated proliferative activity (Fig. 5G and H). TUNEL staining demonstrated an increase in apoptosis among tumor cells within the siPLK1/bola DS-C6-K4 treatment group (Fig. 5I and J and Fig. S19), whereas no apoptotic effects were observed in the PBS, bola DS-C6-K4 alone, or scramble/bola DS-C6-K4 complex groups.
Notably, the body weights of the mice remained consistent throughout the duration of the experiment (Fig. S20A), and no significant pathological alterations were observed in the major serum biomarkers or vital organs (heart, liver, spleen, lungs, and kidneys) following treatment with any of the tested formulations (Fig. S20B and S20C). These results highlight the promise of the bola-amphiphilic peptide dendrimer (bola DS-C6-K4) as a safe, efficient, and highly specific platform for siRNA delivery in vivo, laying the foundation for precision oncology applications.
Conclusion
In this study, we engineered glutathione-responsive bola-amphiphilic peptide dendrimers for cancer cell-specific cytosolic siRNA delivery. These dendrimers consist of a polylysine dendron integrated with a disulfide-containing bola-hydrophobic core, effectively combining the advantages of lipid vectors and peptide dendrimers. Their amphiphilic nature imparts exceptional self-assembly capabilities. The hydrophilic and positively charged polylysine dendron units facilitate efficient siRNA binding, while the hydrophobic bola-lipid segment, which incorporates disulfide bonds, facilitates thiol-mediated cellular uptake and enables siRNA release in response to elevated intracellular GSH levels. Structure–activity relationship studies have indicated that the bola DS-C6-K4, distinguished by the shortest alkyl spacer, exhibited superior siRNA delivery efficiency. This enhanced performance is likely attributable to optimized thiol-mediated cellular uptake and accelerated GSH-triggered siRNA release, resulting from improved disulfide accessibility. Encouraged by these findings, further exploration of the versatility of bola DS-C6-K4 for delivering other nucleic acids is warranted. It is important to note that different nucleic acids, such as messenger RNA (mRNA) and antisense oligonucleotides (ASOs), vary significantly in size, structure, cellular targets, and mechanisms of action, presenting distinct challenges and opportunities for delivery.The siRNA delivered by bola DS-C6-K4 successfully downregulates oncoproteins in vitro, thereby inhibiting proliferation, migration, and invasion of lung cancer cells while inducing apoptosis. Importantly, bola DS-C6-K4 achieved cancer cell-specific cytosolic delivery of siRNA therapeutics and exhibited potent therapeutic efficacy in the A549 lung cancer cell model. This study presents a synergistic approach for the engineering of customized vector by harnessing membrane–thiol interactions and intracellular glutathione-responsive release mechanisms, effectively enhancing siRNA delivery to the cytoplasm and significantly boosting the translation efficiency. This strategy offers substantial potential as a clinically translatable platform for oncological applications.
Author contributions
DZ: writing – original draft, investigation, data curation, and funding acquisition. HZ: data curation, formal analysis, and methodology. YY: writing – original draft, resources, data curation, and validation. JyZ and AM: data curation and validation. XhL, HC, MC, and YL: resources and data curation. HD: formal analysis. JZ: writing – review & editing. JL: writing – review & editing, investigation, and visualization. XL: writing – review & editing, supervision, project administration, conceptualization, and funding acquisition. All authors read and proofed the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Ethical statement
All animal procedures were performed in accordance with the Guidelines for Care and Use of Laboratory Animals of China Pharmaceutical University and approved by the Animal Ethics Committee of China Pharmaceutical University.Data availability
All data supporting this study are present in this paper and the supplementary information (SI). Supplementary information is available. Materials, methods, and Supplementary Figures are included in the SI file. See DOI: https://doi.org/10.1039/d5bm01643f.Acknowledgements
This work was supported by the National Technology Innovation Center for Biopharmaceuticals (No. NCTIB2023XB02003), the National Natural Science Foundation of China (No. 81701815 and 82404545), the Natural Science Foundation of Jiangsu Province (No. BK20170735), the Youth Thousand-Talents Program of China, the National Key Research & Development Program of China for International S&T Cooperation Projects (No. 2018YFE0117800), and the Specialized Research Funds from the State Key Laboratory of Natural Medicines (China Pharmaceutical University, No. SKLNMZZ2024JS18). The authors also thank Jie Zhao from the Animal Experimental Center of China Pharmaceutical University for her help in animal experiments.References
- S. Wang, D. Weissman and Y. Dong, Nat. Rev. Drug Discovery, 2025, 24, 828–851 CrossRef CAS PubMed.
- R. L. Setten, J. J. Rossi and S. Han, The current state and future directions of RNAi-based therapeutics, Nat. Rev. Drug Discovery, 2019, 18(6), 421–446 CrossRef CAS PubMed.
- V. Jadhav, A. Vaishnaw, K. Fitzgerald and M. A. Maier, Nat. Biotechnol., 2024, 42, 394–405 CrossRef CAS PubMed.
- Q. Tang and A. Khvorova, Nat. Rev. Drug Discovery, 2024, 23(5), 341–364 CrossRef CAS PubMed.
- J. A. Kulkarni, D. Witzigmann, S. B. Thomson, S. Chen, B. R. Leavitt, P. R. Cullis and R. van der Meel, Nat. Nanotechnol., 2021, 16(6), 630–643 CrossRef CAS PubMed.
- S. M. Hoy, Patisiran: first global approval, Drugs, 2018, 78(15), 1625–1631 CrossRef CAS PubMed.
- B. Hu, L. Zhong, Y. Weng, L. Peng, Y. Huang, Y. Zhao and X.-J. Liang, Signal Transduction Targeted Ther., 2020, 5(1), 101 CrossRef CAS PubMed.
- G. Sahay, W. Querbes, C. Alabi, A. Eltoukhy, S. Sarkar, C. Zurenko, E. Karagiannis, K. Love, D. L. Chen, R. Zoncu, Y. Buganim, A. Schroeder, R. Langer and D. G. Anderson, Nat. Biotechnol., 2013, 31(7), 653–U119 CrossRef CAS PubMed.
- J. Gilleron, W. Querbes, A. Zeigerer, A. Borodovsky, G. Marsico, U. Schubert, K. Manygoats, S. Seifert, C. Andree, M. Stöter, H. Epstein-Barash, L. G. Zhang, V. Koteliansky, K. Fitzgerald, E. Fava, M. Bickle, Y. Kalaidzidis, A. Akinc, M. Maier and M. Zerial, Nat. Biotechnol., 2013, 31(7), 638–646 CrossRef CAS PubMed.
- M. M. Khan, N. Filipczak and V. P. Torchilin, J. Controlled Release, 2021, 330, 1220–1228 CrossRef CAS PubMed.
- Q. Laurent, R. Martinent, B. Lim, A. T. Pham, T. Kato, J. López-Andarias, N. Sakai and S. Matile, JACS Au, 2021, 1(6), 710–728 CrossRef CAS PubMed.
- Y. Wan, W. Wang, Q. Lai, M. Wu and S. Feng, Drug Discovery Today, 2023, 28(8), 103668 CrossRef CAS PubMed.
- J. Zhang, W. Sun, X. Wang, S. Chen, J. Li and H. Yang, Sci. China:Chem., 2024, 67(1), 383–389 CrossRef CAS.
- J. Zhou, Z. Shao, J. Liu, Q. Duan, X. Wang, J. Li and H. Yang, ACS Appl. Bio Mater., 2020, 3(5), 2686–2701 CrossRef CAS PubMed.
- Y. Zhu, M. Lin, W. Hu, J. Wang, Z. Zhang, K. Zhang, B. Yu and F. Xu, Angew. Chem., Int. Ed., 2022, 61(23), e202200535 CrossRef CAS PubMed.
- S. Bauhuber, C. Hozsa, M. Breunig and A. Göpferich, Adv. Mater., 2009, 21(32–33), 3286–3306 CrossRef CAS PubMed.
- J. Chen, D. Zhu, X. Liu and L. Peng, Acc. Mater. Res., 2022, 3(5), 484–497 CrossRef CAS PubMed.
- Z. Lyu, L. Ding, A. Tintaru and L. Peng, Acc. Chem. Res., 2020, 53, 2936–2949 Search PubMed.
- M. Khaitov, A. Nikonova, I. Kofiadi, I. Shilovskiy, V. Smirnov, O. Elisytina, A. Maerle, A. Shatilov, A. Shatilova, S. Andreev, I. Sergeev, D. Trofimov, T. Latysheva, N. Ilyna, A. Martynov, S. Rabdano, E. Ruzanova, N. Savelev, I. Pletiukhina, A. Safi, V. Ratnikov, V. Gorelov, V. Kaschenko, N. Kucherenko, I. Umarova, S. Moskaleva, S. Fabrichnikov, O. Zuev, N. Pavlov, D. Kruchko, I. Berzin, D. Goryachev, V. Merkulov, G. Shipulin, S. Udin, V. Trukhin, R. Valenta and V. Skvortsova, Allergy, 2023, 78(6), 1639–1653 Search PubMed.
- Y. Zhang, C. Sun, C. Wang, K. E. Jankovic and Y. Dong, Chem. Rev., 2021, 121(20), 12181–12277 CrossRef CAS PubMed.
- R. Kumar, C. F. Santa Chalarca, M. R. Bockman, C. Van Bruggen, C. J. Grimme, R. J. Dalal, M. G. Hanson, J. K. Hexum and T. M. Reineke, Chem. Rev., 2021, 121(18), 11527–11652 CrossRef CAS PubMed.
- Y. Dong, Y. Chen, D. Zhu, K. Shi, C. Ma, W. Zhang, P. Rocchi, L. Jiang and X. Liu, J. Controlled Release, 2020, 322, 416–425 CrossRef CAS PubMed.
- X. Liu, C. Liu, J. Zhou, C. Chen, F. Qu, J. Rossi, P. Rocchi and L. Peng, Nanoscale, 2015, 7(9), 3867–3875 RSC.
- A. H. Fuhrhop and T. Y. Wang, Chem. Rev., 2004, 104(6), 2901–2937 Search PubMed.
- M. Fariya, A. Jain, V. Dhawan, S. Shah and M. S. Nagarsenker, Adv. Pharm. Bull., 2014, 4(2), 483–491 Search PubMed.
- H. X. Zeng, M. E. Johnson, N. J. Oldenhuis, T. N. Tiambeng and Z. B. Guan, ACS Cent. Sci., 2015, 1(6), 303–312 CrossRef CAS PubMed.
- J. Chen, D. Zhu, B. Lian, K. Shi, P. Chen, Y. Li, W. Lin, L. Ding, Q. Long, Y. Wang, E. Laurini, W. Lan, Y. Li, A. Tintaru, C. Ju, C. Zhang, S. Pricl, J. Iovanna, X. Liu and L. Peng, Proc. Natl. Acad. Sci. U. S. A., 2023, 120(21), e2220787120 CrossRef CAS PubMed.
- X. Liu, Y. Wang, C. Chen, A. Tintaru, Y. Cao, J. Liu, F. Ziarelli, J. J. Tang, H. B. Guo, R. Rosas, S. Giorgio, L. Charles, P. Rocchi and L. Peng, Adv. Funct. Mater., 2016, 26(47), 8594–8603 CrossRef CAS.
- C. Ma, D. Zhu, Y. Chen, Y. Dong, W. Lin, N. Li, W. Zhang and X. Liu, Biophys. Rep., 2020, 6(6), 278–289 CrossRef CAS.
- V. Percec, D. A. Wilson, P. Leowanawat, C. J. Wilson, A. D. Hughes, M. S. Kaucher, D. A. Hammer, D. H. Levine, A. J. Kim, F. S. Bates, K. P. Davis, T. P. Lodge, M. L. Klein, R. H. DeVane, E. Aqad, B. M. Rosen, A. O. Argintaru, M. J. Sienkowska, K. Rissanen, S. Nummelin and J. Ropponen, Science, 2010, 328(5981), 1009–1014 CrossRef CAS PubMed.
- C. Chen, P. Posocco, X. Liu, Q. Cheng, E. Laurini, J. Zhou, C. Liu, Y. Wang, J. Tang, V. D. Col, T. Yu, S. Giorgio, M. Fermeglia, F. Qu, Z. Liang, J. Rossi, M. Liu, P. Rocchi, S. Pricl and L. Peng, Small, 2016, 12(27), 3667–3676 CrossRef CAS PubMed.
- S. Zitouni, C. Nabais, S. C. Jana, A. Guerrero and M. Bettencourt-Dias, Nat. Rev. Mol. Cell Biol., 2014, 15(7), 433–452 CrossRef CAS PubMed.
- K. Strebhardt and A. Ullrich, Nat. Rev. Cancer, 2006, 6(4), 321–330 CrossRef CAS PubMed.
- B. D. Manning and A. Toker, Cell, 2017, 169(3), 381–405 CrossRef CAS PubMed.
- Q. Wang, X. Chen and N. Hay, Br. J. Cancer, 2017, 117(2), 159–163 CrossRef CAS PubMed.
- X. W. Cheng and R. J. Lee, Adv. Drug Delivery Rev., 2016, 99(Pt A), 129–137 CrossRef CAS PubMed.
- Y. Hattori, S. Suzuki, S. Kawakami, F. Yamashita and M. Hashida, J. Controlled Release, 2005, 108(2–3), 484–495 CrossRef CAS PubMed.
- A. A. Thai, B. J. Solomon, L. Sequist, J. F. Gainor and R. S. Heist, Lancet, 2021, 398(10299), 535–554 CrossRef PubMed.
- L. E. L. Hendriks, J. Remon, C. Faivre-Finn, M. C. Garassino, J. V. Heymach, K. M. Kerr, D. S. W. Tan, G. Veronesi and M. Reck, Nat. Rev. Dis. Primers, 2024, 10(1), 71 CrossRef PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |