 Open Access Article
Open Access ArticleClickable polyamidosaccharides: accessing bottlebrush inspired hyaluronic acid glycopolymers for CD44 targeting of breast cancer cells
Kaitlynn A.
Sockett†‡
 a,
Madeline K.
Loffredo†‡
a,
Madeline K.
Loffredo†‡
 b,
Christian D.
DeMoya
b,
Christian D.
DeMoya
 c,
Zoe G.
Garman
c,
Zoe G.
Garman
 c and
Mark W.
Grinstaff
c and
Mark W.
Grinstaff
 *abc
*abc
aDepartment of Chemistry, Boston University, Room 519, 590 Commonwealth Ave, Boston, MA 02215, USA. E-mail: mgrin@bu.edu; ksockett@bu.edu; Tel: +1 617-358-3429
bDivision of Material Science and Engineering, Boston University, Boston, MA 02215, USA. E-mail: msl@bu.edu
cDepartment of Biomedical Engineering, Boston University, Boston, MA 02215, USA. E-mail: cdemoya@bu.edu; zgarman@bu.edu
First published on 17th February 2026
Abstract
Hyaluronic acid (HA) binds the transmembrane glycoprotein cluster of differentiation-44 (CD44), a highly expressed surface receptor that plays a critical role in tumor growth, invasion, and metastasis. Approaches to target CD44 utilize biologically sourced HA which inherently suffers from molecular weight (MW) heterogeneity and biological contaminants. Fully synthetic approaches to HA are attractive and circumvent these biological contaminants; however, readily accessing oligomers of six monosaccharides or more, as is required for CD44 binding, is challenging. To this end, we report the synthesis of glycopolymers functionalized with HA disaccharide pendant chains. These well-defined and regioselective polymers consist of glucose monomers linked via α-1,2 amide bonds, termed polyamidosaccharides, functionalized with branched HA disaccharide moieties interspersed throughout via a strain-promoted azide–alkyne cycloaddition. Among these homopolymers and copolymers, two of the polymers bearing the highest HA disaccharide conjugation bind CD44 with nanomolar affinity. Assays using a rhodamine-labelled polymer reveal a positive relationship between cellular internalization and CD44 expression levels in breast cancer cells. Conjugation of paclitaxel to the polymer enhances paclitaxel potency in CD44-expressing cancer cells compared to free paclitaxel.
1. Introduction
Hyaluronic acid (HA) is a naturally occurring linear, non-sulfated glycosaminoglycan consisting of alternating disaccharide β-1,4-D-glucuronic acid and β-1,3-N-acetylglucosamine repeat units (Fig. 1A).1–4 It is a main component of the extracellular matrix and prominently found within synovial joints, providing structural support and water retention due to its anionic character at physiological pH.3–8 The structure of HA comprises a densely hydroxylated carbohydrate with carboxylate moieties which act as handles readily available for chemical modification. HA is biocompatible, biodegradable, and nontoxic, culminating in its wide-spread use in translational research and regulatory approved products.9–12 | ||
| Fig. 1 (A) Chemical structure of HA. (B) Example of a previously reported brush-like HA glycopolymer.13 (C) Examples of previously reported PASs with varied functionalities; glucose PAS,14 sulfate-PAS,15 and amino-PAS.16 (D) Proposed homopolymer and copolymer functionalized with a HA disaccharide pendant group. | ||
Additionally of importance, HA binds the transmembrane glycoprotein, cluster of differentiation-44 (CD44), which is upregulated on the surface of numerous cancer cell types including breast, prostate, pancreatic, gastrointestinal, lung, colorectal, and head and neck squamous cell carcinoma.17–20 CD44 is a cell surface adhesion receptor and, in homeopathic tissues, regulates cell growth, survival, cellular adhesion, signaling, and motility.21 In cancerous tissues, increased CD44 expression positively correlates with tumorigenesis and metastasis due to HA and CD44 binding interactions which activate signaling cascades that facilitate cancer progression and invasion.17,21,22 Numerous reports from the last decade employ HA as a targeting ligand on delivery systems to achieve tumor cell selectivity and localization of a chosen payload to enhance therapeutic efficacy and decrease off-target effects.23–31 These examples utilize HA isolated from biological sources; however, biologically sourced HA is highly heterogeneous in its molecular weight (MW) and often carries potentially harmful endotoxins and residual biological contaminants, all of which impact the biological function and experimental reproducibility of the biopolymer.10
Fully synthetic approaches to HA eliminate residual biological contaminants and endotoxin concerns; however, these syntheses are lengthy, given that the oligomer must contain at least six monosaccharide repeat units for effective CD44 binding,32,33 and limited by scalability due to the series of reactions necessary.34,35 Contrastingly, some synthetic polymers capitalize on multivalency through brush-like glycopolymers functionalized with low MW HA. For example, Carvalho et al. report the synthesis of a glycopolymer with a 24-monosaccharide multivalent HA targeting ligand on a hydrocarbon polymeric backbone, and demonstrate improved CD44 binding avidity with an association constant (KA) = (4.0 ± 6.0) × 1010 [M−1] compared to low MW HA (4.8 kDa) with a KA = (1.5 ± 0.9) × 107 [M−1], as determined by surface plasmon resonance (SPR) (Fig. 1B).13 Utilizing a similar polymeric architecture, Collis et al. describe hydrocarbon-based glycopolymers bearing N-acetyl glucosamine (GlcNAc) or glucuronic acid (GlcA) monosaccharides which bind CD44 with micromolar affinity (GlcNAc: 0.6 μM, GlcA: 0.9 μM, 5 kDa HA: 0.15 μM) via SPR to demonstrate CD44 binding with multivalent monosaccharide pendant moieties.36
Polyamidosaccharides (PASs) are a novel class of polysaccharide mimetics where α-1,2 amide linkages replace the native glycosidic bonds between the repeating sugar units. An anionic ring-opening polymerization of a reactive β-lactam monomer yields nontoxic PASs with controlled MW, narrow dispersity, and a helical secondary structure (Fig. 1C).14–16,37–42 PAS composition and structure afford opportunities to design functionally active polysaccharide mimetics which retain the biocompatibility, biodegradability, and dense functionality of their natural counterparts,15,16,37,41 and herein, we report the synthesis and biological activity of a HA-inspired PAS to target CD44-expressing cancer cells for therapeutic delivery. From a design perspective, we employ a “clickable” PAS backbone where a HA disaccharide links to the polymer in a multivalent fashion via a strain-promoted azide–alkyne cycloaddition (SPAAC). We describe the synthesis of a carbohydrate-based polymeric library consisting of HA disaccharide functionalized homopolymers and copolymers (Fig. 1D). The bottlebrush-like PAS armed with a synthetically accessible HA disaccharide binds to CD44 with nanomolar affinity, is internalized by CD44-expressing breast cancer cells, and exhibits potent cytotoxicity when functionalized with a chemotherapeutic against such breast cancer cells.
2. Results and discussion
2.1. Synthesis of the pendant HA disaccharide
Our strategy to the pendant functionalized polymer entails first synthesizing the HA derived pendant disaccharide targeting ligand and then employing a SPAAC reaction, due to its mild and catalyst free reaction conditions, orthogonality, and downstream biocompatibility, to link it to the PAS.43,44 Thus, we synthesized the HA pendant via preparation of the azido-armed glycosylation acceptor (Scheme 1A). Acetylation of D-glucose affords 1, followed by iodination of the anomeric center and subsequent treatment with thiourea and iodoethane to give thioglycoside 2 in excellent yields (85%).45 Saponification45 and subsequent benzylidene acetal protection46 of O4 and O6 affords diol 3. Functionalization of the O2 and O3 positions with benzoyl groups47 gives 4, which serves as the electrophilic partner in the trifluoromethanesulfonic acid-catalyzed nucleophilic addition of azidoethoxy ethanol at the anomeric position, furnishing 5.48 We confirmed the β-anomeric stereochemistry via1H nuclear magnetic resonance (NMR) J-value analysis as JH-1 = 7.8 Hz, indicating a diaxial proton orientation consistent with the β-configuration. Next, trifluoroacetic acid-catalyzed benzylidene acetal deprotection affords diol 6.49 Oxidation of the primary C6 hydroxyl to the corresponding carboxylic acid via a TEMPO oxidation followed by a chemoselective benzyl protection yields acceptor 7 (Scheme 1A).50For the glycosylation donor synthesis, a selective protection of glucosamine hydrochloride, affords imine 8 which is further acetylated to 9.45 Acidic hydrolysis of the imine gives the acetylated glucosamine salt 10 allowing subsequent installation of the N-trichloroacetyl (TCA) group to serve as the N-acetyl analog.45 Lewis acid mediated thiolation furnishes thioglycoside 11.51 Saponification followed by a benzylidene acetal protection of the O4 and O6 positions affords thioglycoside 12.51 Benzylation51 of O3 provides 13, which underwent a Lewis-acid catalyzed intramolecular cyclization47 to yield oxazoline donor 14 (Scheme 1B). With both the glycosylation acceptor and donor in hand, a triflate-mediated glycosylation provides the diastereopure azido-armed disaccharide 15 (Scheme 1C).47 Once again we used 1H NMR J-value analysis to confirm the stereochemistry of the isolated disaccharide. Due to spectrum overlap, we estimated the JH-1′ to be ∼8.7 Hz; however, the presence of the β-anomer is evident by the approximate J-value as well as the observed splitting pattern (Fig. S1).
2.2. Synthesis of the DBCO-functionalized linker
To functionalize the polymeric backbone, we next synthesized a linker bearing a terminal dibenzocyclooctyne (DBCO) to serve as the strained alkyne counterpart to the azido-armed disaccharide 15. Additionally, the DBCO-functionalized linker requires an electrophilic primary halide for nucleophilic substitution onto the PAS backbone, as wells as sufficient flexibility to reduce steric interactions which may impede the pendant disaccharide binding to CD44. To achieve this goal, we prepared linker 17 in good yields (Scheme 2A) by first substituting 2-(2-aminoethoxy)ethanol with 1-chloro-6-iodohexane to prepare 16, followed by a DBCO-N-hydroxysuccinimide coupling.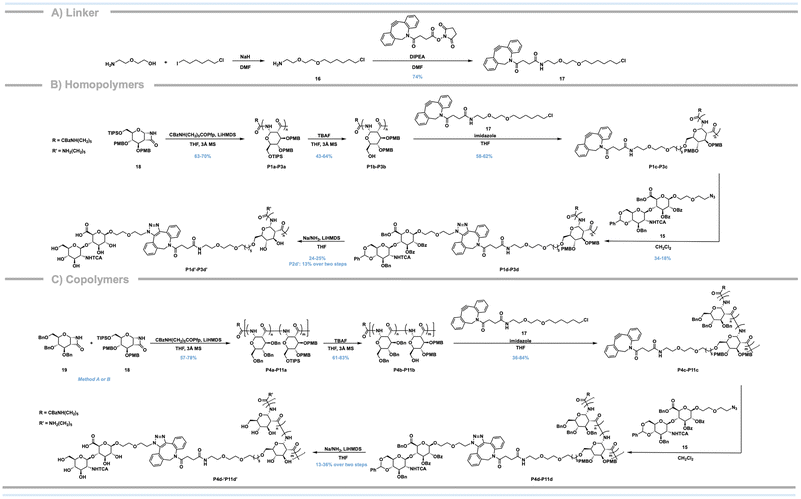 | ||
| Scheme 2 Synthesis of (A) DBCO-functionalized linker; (B) pendant disaccharide-functionalized homopolymers P1–P3; (C) pendant disaccharide-functionalized copolymers P4–P11. | ||
2.3. Homopolymer synthesis and characterization
To prepare the “clickable” polysaccharide mimetics, we first performed an anionic ring-opening polymerization of a protected β-lactam bicyclic monomer to afford the PAS backbone.14–16,37–42 For post-functionalization, we require a primary hydroxyl to serve as a handle for coupling with linker 17 and ultimately the pendant disaccharide 15, thus, we used selectively protected 6-O-triisopropylsilyl (TIPS) β-lactam 18 as the monomer.15 Lithium bis(trimethylsilyl) amide (LiHMDS) catalyzed polymerization of 18 in the presence of the initiator, Cbz-6-aminohexanoic acid pentafluorophenol ester, yields the PASs. Through adjusting the monomer to initiator ratio ([M]/[I] = 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 50
1, 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 25
1, 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) we obtain well defined homopolymers P1a–P3a with controlled MW (Scheme 2B). Gel permeation chromatography (GPC) equipped with a right-angle light scattering detector with THF as the mobile phase, reveals the measured degrees of polymerization (N) are in close agreement with the theoretical N (Ntheory) for P1a–P3a (Fig. 2A, Table 1, entries 1–3). Following polymerization, treatment with 1 M TBAF in THF selectively deprotects the TIPS group to yield the 6-OH polymers P1b–P3b (Scheme 2B, Fig. 2B, Table 1, entries 4–6). Coupling of the previously prepared linker 17 to the O6 position using imidazole in THF, followed by precipitation into cold n-pentane affords P1c–P3c (Scheme 2B, Fig. 2C, Table 1, entries 7–9). Based on the increase in absolute MW-weight-average molecular weight (Mw)-between polymers P1b–P3b to P1c–P3c, the conjugation efficiency is roughly 100% (Fig. 2C, Table 1, entries 4–9). 1H NMR analysis further corroborates the conjugation efficiency indicating a ∼1
1) we obtain well defined homopolymers P1a–P3a with controlled MW (Scheme 2B). Gel permeation chromatography (GPC) equipped with a right-angle light scattering detector with THF as the mobile phase, reveals the measured degrees of polymerization (N) are in close agreement with the theoretical N (Ntheory) for P1a–P3a (Fig. 2A, Table 1, entries 1–3). Following polymerization, treatment with 1 M TBAF in THF selectively deprotects the TIPS group to yield the 6-OH polymers P1b–P3b (Scheme 2B, Fig. 2B, Table 1, entries 4–6). Coupling of the previously prepared linker 17 to the O6 position using imidazole in THF, followed by precipitation into cold n-pentane affords P1c–P3c (Scheme 2B, Fig. 2C, Table 1, entries 7–9). Based on the increase in absolute MW-weight-average molecular weight (Mw)-between polymers P1b–P3b to P1c–P3c, the conjugation efficiency is roughly 100% (Fig. 2C, Table 1, entries 4–9). 1H NMR analysis further corroborates the conjugation efficiency indicating a ∼1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio of the polymeric H1 peak at ∼5.7 ppm to the amide proton peak of the linker at ∼6.0 ppm (Fig. S2). With the linker substituted polymers prepared, we next investigated the efficiency of the SPAAC between polymers P1c–P3c and the azido-armed disaccharide 15. The standard purification process of precipitation(s) in cold n-pentane does not isolate polymers from excess disaccharide 15; however, cold methanol precipitations remove the residual disaccharide with the exception of P2d which is soluble in cold methanol. Thus, we carried P2d forward to the next step without further purification (Scheme 2B, Fig. 2D, Table 1, entries 10–12).
1 ratio of the polymeric H1 peak at ∼5.7 ppm to the amide proton peak of the linker at ∼6.0 ppm (Fig. S2). With the linker substituted polymers prepared, we next investigated the efficiency of the SPAAC between polymers P1c–P3c and the azido-armed disaccharide 15. The standard purification process of precipitation(s) in cold n-pentane does not isolate polymers from excess disaccharide 15; however, cold methanol precipitations remove the residual disaccharide with the exception of P2d which is soluble in cold methanol. Thus, we carried P2d forward to the next step without further purification (Scheme 2B, Fig. 2D, Table 1, entries 10–12).
| Entry | Polymer | M n (kDa) | M w (kDa) | Đ | N | N theory | Yield (%) |
|---|---|---|---|---|---|---|---|
| N for P1c–P3c was calculated assuming 100% of Mw included linker-functionalized monomer units; N for P1d–P3d was calculated from disaccharide conjugation percentages determined via1H NMR analysis of deprotected polymers (eqn (S1)). P2d was unable to be purified and was thus carried forward to the next step without further purification. | |||||||
| 1 | P1a | 16.4 | 19.2 | 1.17 | 33 | 25 | 65 |
| 2 | P2a | 34.2 | 36.4 | 1.06 | 62 | 50 | 63 |
| 3 | P3a | 64.3 | 68.5 | 1.07 | 117 | 100 | 70 |
| 4 | P1b | 12.6 | 15.6 | 1.24 | 36 | 25 | 43 |
| 5 | P2b | 27.4 | 28.5 | 1.04 | 66 | 50 | 60 |
| 6 | P3b | 47.6 | 48.9 | 1.03 | 114 | 100 | 64 |
| 7 | P1c | 28.9 | 28.9 | 1.002 | 32 | 25 | 58 |
| 8 | P2c | 49.6 | 49.2 | 1.01 | 55 | 50 | 62 |
| 9 | P3c | 92.6 | 111.8 | 1.21 | 124 | 100 | 62 |
| 10 | P1d | 30.6 | 32.7 | 1.07 | 31 | 25 | 34 |
| 11 | P2d | 63.3 | 63.9 | 1.01 | 66 | 50 | — |
| 12 | P3d | 65.8 | 115.3 | 1.75 | 114 | 100 | 18 |
Sodium metal in liquid ammonia deprotection of polymers P1d–P3d followed by dialysis against deionized water, using an 8.0 kDa MW cutoff dialysis membrane, and lyophilization affords the final polymers P1d′–P3d′ as fluffy white solids in low to moderate yields (13–25%, Scheme 2B, Fig 4A, Table 4, entries 1–3). P1d′–P3d′ adopt a helical secondary structure as determined by circular dichroism (CD) (Fig. S3A).14,371H NMR spectroscopy confirms the removal of the aromatic protecting groups and analysis of the ratio of the polymeric H1 peak resolved at ∼5.75 ppm with the anomeric protons of the pendant HA disaccharide at ∼4.55 ppm and ∼4.50 ppm provides an approximation of the SPAAC reaction conjugation efficiency (Fig. 2E). The pendant conjugations range between 13–27% (Table 5, entries 1–3, Fig. S4A). These results are consistent with the increased absolute Mw observed for polymers P1d–P3d by GPC prior to deprotection. To calculate the degrees of polymerization (N), we used the conjugation efficiency determined by 1H NMR analysis of the deprotected polymers; the final N values are consistent with the calculated N from the previous steps, as seen with NP1a = 33 and NP1d = 31, for example (Table 1, entries 1 and 10). Equations for calculating N for each polymer class are listed in the SI. Our attempts to improve the conjugation efficiency by increasing the reaction temperature or increasing the equivalents of disaccharide 15 from 1.5 eq. to 2.0 eq. (relative to one monomer unit) were unsuccessful. Using aqueous GPC equipped with a right-angle light scattering detector and 1× PBS as the mobile phase, we determined the absolute Mw of P1d′–P3d′ which are in good agreement with the N calculated during the previous synthetic steps (Fig. 4A, Table 4, entries 1–3, Fig. S5A). Raw GPC traces are noisy due to the scattering effects of the salt; therefore, traces were fit with a moving average trendline (period of 50) for improved visualization (Fig. 4 and S5). The GPC analysis confirms the polymers remain intact throughout the deprotection and corroborates the approximate pendant conjugation seen with their protected polymer counterparts (Table 4, entries 1–11).
2.4. Copolymer synthesis and characterization
Given the ∼25% pendant conjugation efficiency of the homopolymers, we next investigated whether interspersing glucose monomer units among the HA pendant functionalized polymers impacts the SPAAC efficiency and downstream binding with CD44. Thus, we synthesized copolymers comprising both 6-O-TIPS lactam 18 and tri-O-Bn lactam 19 in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 monomeric ratio, respectively. We selected monomer 19 as the benzyl moieties are stable throughout the post-polymerization modifications, yet readily debenzylated in the final global deprotection.14,37 Due to the discrepancy between the polymerization kinetics of the chosen monomers (tri-O-Bn lactam 19 fully polymerizes in 5 min,14 6-O-TIPS-3,4-O-PMB lactam 18 fully polymerizes in 20 min15), we explored two different copolymerization methods as well as an additional higher MW polymer (Ntheory = 200; [M]/[I] = 200/1) to expand the MW scope of the copolymer library and to yield variable branching densities.
3 monomeric ratio, respectively. We selected monomer 19 as the benzyl moieties are stable throughout the post-polymerization modifications, yet readily debenzylated in the final global deprotection.14,37 Due to the discrepancy between the polymerization kinetics of the chosen monomers (tri-O-Bn lactam 19 fully polymerizes in 5 min,14 6-O-TIPS-3,4-O-PMB lactam 18 fully polymerizes in 20 min15), we explored two different copolymerization methods as well as an additional higher MW polymer (Ntheory = 200; [M]/[I] = 200/1) to expand the MW scope of the copolymer library and to yield variable branching densities.
Method A involves simultaneous monomer additions following the same polymerization procedure utilized for the homopolymerizations. LiHMDS polymerization of the monomers in the presence of Cbz-6-aminohexanoic acid pentafluorophenol ester yields copolymers, P4a–P7a, in good yields comparable to the homopolymers, and with N values that agree with Ntheory (Scheme 2C, Fig. 3A, Table 2, entries 1–4). To account for the 25% composition of the 6-O-TIPS monomer in the subsequent steps, we calculated the reagent equivalents based on monomer 18 content, while reaction concentrations were based on total mmol of monomers 18 and 19. This approach results in the successful selective deprotection of the TIPS group to afford polymers P4b–P7b in high yields. The observed Mw and calculated N6-OH for each copolymer are consistent with the calculated N6-TIPS copolymers (Scheme 2C, Fig. 3B, Table 2, entries 5–8, eqn (S2)). Addition of linker 17 affords polymers P4c–P7c in appreciable yields with a 100% conjugation efficiency relative to the 6-OH monomer units, as determined by the increase in absolute Mw from GPC analysis (Scheme 2C, Fig. 3C, Table 2, entries 9–12). For the SPAAC, we increased the equivalents of disaccharide 15 to 2.0 eq. from 1.5 eq. relative to the DBCO to drive the reaction to completion. Unfortunately, the disaccharide-functionalized copolymers do not precipitate into cold methanol, therefore, we carried the crude polymers forward to the next deprotection step. GPC analysis of P4d–P7d shows increased Mw for all polymers consistent with a successful SPAAC (Scheme 2C, Fig. 3D, Table 2, entries 13–16). Following the global deprotection using sodium metal in liquid ammonia, dialysis (8.0 kDa MW cutoff membrane) against deionized water removes any residual unconjugated disaccharide. Subsequent lyophilization gives polymers P4d′–P7d′ as fluffy white solids in moderate yields (17–36%, Scheme 2C, Table 4, entries 4–7). Polymers P4d′–P7d′ also adopt a helical secondary structure as determined via CD (Fig. S3B). 1H NMR analysis of the ratio of the anomeric H1 proton of copolymers P4d′–P7d′ to the corresponding anomeric H1 protons of the pendant disaccharide indicates a conjugation efficiency of approximately 11–15% relative to the full polymeric backbone, except for P4d′ (∼3% conjugation; Fig. 2E, Table 5, entries 4–7, Fig. S4B). Aqueous GPC analysis of polymers P4d′–P7d′ reveals degrees of polymerization N consistent with previous steps (Fig. 4B, Table 4, entries 4–7, Fig. S5B). Again, we calculated the N of the deprotected polymers using the relative conjugation efficiencies determined by 1H NMR analysis (eqn (S2)).
 | ||
| Fig. 3 GPC chromatograms of copolymers P4–P11. (A) P4a–P7a; (B) P4b–P7b; (C) P4c–P7c; (D) P4d–P7d; (E) P8a–P11a; (F) P8b–P11b; (G) P8c–P11c; (H) P8d–P11d. | ||
 | ||
| Fig. 4 GPC chromatograms fit with a moving average trendline (period of 50) of deprotected polymers (A) P1d′–P3d′; (B) P4d′–P7d′; (C) P8d′–P11d′. | ||
| Entry | Polymer | M n (kDa) | M w (kDa) | Đ | N | N theory | Yield (%) |
|---|---|---|---|---|---|---|---|
| N for P4c–P7c was calculated assuming 100% functionalization of linker; N for P4d–P7d was calculated from disaccharide conjugation percentages determined via1H NMR analysis of deprotected polymers (eqn (S2)). Polymers P4d–P7d were carried to the next step without further purification. | |||||||
| 1 | P4a | 17.6 | 17.7 | 1.01 | 37 | 25 | 78 |
| 2 | P5a | 24.5 | 24.5 | 1.001 | 51 | 50 | 63 |
| 3 | P6a | 49.2 | 49.6 | 1.01 | 101 | 100 | 57 |
| 4 | P7a | 95.7 | 102.0 | 1.05 | 210 | 200 | 57 |
| 5 | P4b | 15.9 | 16.0 | 1.002 | 35 | 25 | 76 |
| 6 | P5b | 23.1 | 23.3 | 1.01 | 52 | 50 | 67 |
| 7 | P6b | 40.6 | 42.7 | 1.05 | 95 | 100 | 62 |
| 8 | P7b | 89.4 | 94.9 | 1.06 | 210 | 200 | 61 |
| 9 | P4c | 17.6 | 18.7 | 1.06 | 36 | 25 | 64 |
| 10 | P5c | 24.4 | 25.3 | 1.04 | 48 | 50 | 36 |
| 11 | P6c | 58.5 | 58.8 | 1.01 | 112 | 100 | 61 |
| 12 | P7c | 85.5 | 109.4 | 1.28 | 209 | 200 | 74 |
| 13 | P4d | 20.1 | 21.0 | 1.04 | 40 | 25 | — |
| 14 | P5d | 30.2 | 30.9 | 1.02 | 57 | 50 | — |
| 15 | P6d | 58.8 | 59.2 | 1.01 | 107 | 100 | — |
| 16 | P7d | 97.0 | 118.6 | 1.22 | 218 | 200 | — |
We hypothesized the decreased pendant conjugation observed with copolymers P4d′–P7d′ relative to the homopolymers P1d′–P3d′ is a result of a more block-like polymeric sequence, which arises from the differences in the polymerization kinetics of lactams 18 and 19. This discrepancy likely affords polymers with increased steric restraints and dense branching similar to those of the homopolymers preventing full conjugation of the pendant disaccharide and, thus, we explored a second polymerization approach.
The second polymerization method, Method B, addresses the varying polymerization kinetics of our monomers to furnish a randomized sequence relative to polymers P4–P7. Herein we initiated the polymerization of the 6-O-TIPS lactam 18 alone, due to its slower polymerization kinetics (monomer fully consumed in 20 min15) followed by intermittent additions of the tri-O-benzyl lactam 19 in 0.5 mL aliquots over the course of 20 min (Scheme 2C). The aliquots ensure a polymerization concentration of ∼0.1 M relative to all monomer units in solution and a final 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 monomeric ratio of 6-O-TIPS lactam 18 to tri-O-benzyl lactam 19 as used in Method A. Again, LiHMDS polymerization of the monomers in the presence of Cbz-6-aminohexanoic acid pentafluorophenol ester gives the copolymers P8a–P11a in good yields with MW control. However, the dispersity increases relative to copolymers P4a–P7a, which we hypothesized is a consequence of the sequential additions of lactam 19 (Fig. 3E, Table 3, entries 1–4). We employed the same post-polymerization synthetic steps described for copolymers P4–P7 for copolymers P8–P11 (Scheme 2C). Silyl deprotections afford P8b–P11b in high yields with consistent N6-OH as compared to N6-TIPS (Fig. 3F, Table 3, entries 5–8, eqn (S2)). Addition of linker 17 provides copolymers P8c–P11c with full conjugation relative to each 6-OH monomer unit as determined by the increase in absolute Mwvia GPC analysis (Fig. 3G, Table 3, entries 9–12). The SPAAC with disaccharide 15 results in an increased Mw for all copolymers P8d–P11d (Fig. 3H, Table 3, entries 13–16) indicative of successful conjugations. As before, copolymers P8d–P11d do not precipitate in cold methanol and were carried through to the global deprotection step. Dialysis (8.0 kDa MW cutoff membrane) against deionized water removes any residual disaccharide. Following lyophilization, copolymers P8d′–P11d′ are white fluffy solids (moderate yields of 13–36%, Table 4, entries 8–11) with helical secondary structure as determined by CD (Fig. S3C). 1H NMR analysis of P8d′–P11d′ reveals improved pendant disaccharide conjugations relative to the corresponding copolymers synthesized via Method A, consistent with a decrease in overall steric hindrance and a decrease in localized branching density (Table 5, entries 4–11, Fig. S4C). For example, P10d′ conjugation efficiency of the pendant disaccharide relative to the clickable monomer units (Method B, Ntheory = 100) is 96% as compared to 72% for P6d′ (Method A, Ntheory = 100) and 20% for P3d′ (homopolymer, Ntheory = 100) (Table 5, entries 3, 6, and 10, Fig. 2E). Interestingly, the pendant disaccharide percent conjugation increases in correlation with an increase in N for all copolymers prepared via both methods up to N = 100, before decreasing slightly at N = 200 (Table 5, entries 4–11). Aqueous GPC once again confirms approximate disaccharide conjugations and absolute Mw for copolymers P8d′–P11d′ demonstrated by N which are consistent with the previous steps and based on disaccharide conjugation percentages as determined by 1H NMR (Fig. 4C, Table 4, entries 8–11, Fig. S5C, eqn (S2)).
3 monomeric ratio of 6-O-TIPS lactam 18 to tri-O-benzyl lactam 19 as used in Method A. Again, LiHMDS polymerization of the monomers in the presence of Cbz-6-aminohexanoic acid pentafluorophenol ester gives the copolymers P8a–P11a in good yields with MW control. However, the dispersity increases relative to copolymers P4a–P7a, which we hypothesized is a consequence of the sequential additions of lactam 19 (Fig. 3E, Table 3, entries 1–4). We employed the same post-polymerization synthetic steps described for copolymers P4–P7 for copolymers P8–P11 (Scheme 2C). Silyl deprotections afford P8b–P11b in high yields with consistent N6-OH as compared to N6-TIPS (Fig. 3F, Table 3, entries 5–8, eqn (S2)). Addition of linker 17 provides copolymers P8c–P11c with full conjugation relative to each 6-OH monomer unit as determined by the increase in absolute Mwvia GPC analysis (Fig. 3G, Table 3, entries 9–12). The SPAAC with disaccharide 15 results in an increased Mw for all copolymers P8d–P11d (Fig. 3H, Table 3, entries 13–16) indicative of successful conjugations. As before, copolymers P8d–P11d do not precipitate in cold methanol and were carried through to the global deprotection step. Dialysis (8.0 kDa MW cutoff membrane) against deionized water removes any residual disaccharide. Following lyophilization, copolymers P8d′–P11d′ are white fluffy solids (moderate yields of 13–36%, Table 4, entries 8–11) with helical secondary structure as determined by CD (Fig. S3C). 1H NMR analysis of P8d′–P11d′ reveals improved pendant disaccharide conjugations relative to the corresponding copolymers synthesized via Method A, consistent with a decrease in overall steric hindrance and a decrease in localized branching density (Table 5, entries 4–11, Fig. S4C). For example, P10d′ conjugation efficiency of the pendant disaccharide relative to the clickable monomer units (Method B, Ntheory = 100) is 96% as compared to 72% for P6d′ (Method A, Ntheory = 100) and 20% for P3d′ (homopolymer, Ntheory = 100) (Table 5, entries 3, 6, and 10, Fig. 2E). Interestingly, the pendant disaccharide percent conjugation increases in correlation with an increase in N for all copolymers prepared via both methods up to N = 100, before decreasing slightly at N = 200 (Table 5, entries 4–11). Aqueous GPC once again confirms approximate disaccharide conjugations and absolute Mw for copolymers P8d′–P11d′ demonstrated by N which are consistent with the previous steps and based on disaccharide conjugation percentages as determined by 1H NMR (Fig. 4C, Table 4, entries 8–11, Fig. S5C, eqn (S2)).
| Entry | Polymer | M n (kDa) | M w (kDa) | Đ | N | N theory | Yield (%) |
|---|---|---|---|---|---|---|---|
| N for P8c–P11c was calculated assuming 100% functionalization of linker; N for P8d–P11d was calculated from disaccharide conjugation percentages determined via1H NMR analysis of deprotected polymers (eqn (S2)). Polymers P8d–P11d were carried to the next step without further purification. | |||||||
| 1 | P8a | 7.0 | 12.7 | 1.82 | 26 | 25 | 66 |
| 2 | P9a | 25.3 | 27.6 | 1.01 | 57 | 50 | 70 |
| 3 | P10a | 72.9 | 73.8 | 1.01 | 152 | 100 | 73 |
| 4 | P11a | 65.1 | 116.0 | 1.78 | 239 | 200 | 73 |
| 5 | P8b | 10.2 | 13.1 | 1.28 | 29 | 25 | 83 |
| 6 | P9b | 15.7 | 26.6 | 1.78 | 59 | 50 | 71 |
| 7 | P10b | 62.8 | 70.7 | 1.13 | 156 | 100 | 69 |
| 8 | P11b | 61.9 | 107.0 | 1.73 | 237 | 200 | 78 |
| 9 | P8c | 11.0 | 14.7 | 1.33 | 28 | 25 | 36 |
| 10 | P9c | 27.7 | 30.5 | 1.10 | 58 | 50 | 84 |
| 11 | P10c | 63.0 | 77.4 | 1.23 | 148 | 100 | 69 |
| 12 | P11c | 69.8 | 113.8 | 1.63 | 217 | 200 | 78 |
| 13 | P8d | 12.9 | 15.8 | 1.22 | 30 | 25 | — |
| 14 | P9d | 25.6 | 32.7 | 1.27 | 60 | 50 | — |
| 15 | P10d | 52.4 | 84.4 | 1.61 | 152 | 100 | — |
| 16 | P11d | 87.6 | 126.1 | 1.44 | 232 | 200 | — |
| Entry | Polymer | M n (kDa) | M w (kDa) | Đ | N | N theory | Yield (%) |
|---|---|---|---|---|---|---|---|
| a Yields were calculated over two steps. Raw GPC traces (Fig. S5) were fit with a moving average trendline (period of 50). | |||||||
| 1 | P1d′ | 23.1 | 26.8 | 1.16 | 35 | 25 | 24 |
| 2 | P2d′ | 38.4 | 48.1 | 1.25 | 68 | 50 | 13a |
| 3 | P3d′ | 82.0 | 82.4 | 1.01 | 112 | 100 | 25 |
| 4 | P4d′ | 6.9 | 8.7 | 1.26 | 37 | 25 | 17a |
| 5 | P5d′ | 13.5 | 13.7 | 1.02 | 58 | 50 | 22a |
| 6 | P6d′ | 31.9 | 32.8 | 1.03 | 132 | 100 | 36a |
| 7 | P7d′ | 45.2 | 47.5 | 1.05 | 198 | 200 | 25a |
| 8 | P8d′ | 4.9 | 6.2 | 1.25 | 27 | 25 | 36a |
| 9 | P9d′ | 8.7 | 13.5 | 1.54 | 56 | 50 | 19a |
| 10 | P10d′ | 22.6 | 38.6 | 1.71 | 160 | 100 | 36a |
| 11 | P11d′ | 32.4 | 57.4 | 1.78 | 239 | 200 | 13a |
| Entry | Polymer | Relative conjugation to full polymeric backbone (%) | Relative conjugation to DBCO-linked monomer units (%) |
|---|---|---|---|
| 1 | P1d′ | 27 | 27 |
| 2 | P2d′ | 13 | 13 |
| 3 | P3d′ | 20 | 20 |
| 4 | P4d′ | 3 | 12 |
| 5 | P5d′ | 11 | 44 |
| 6 | P6d′ | 18 | 72 |
| 7 | P7d′ | 12 | 48 |
| 8 | P8d′ | 6 | 24 |
| 9 | P9d′ | 13 | 52 |
| 10 | P10d′ | 24 | 96 |
| 11 | P11d′ | 18 | 72 |
2.5. CD44 binding studies and cytotoxicity
With the polymer library prepared, we next measured the binding affinity of the HA disaccharide conjugated polymers to CD44 as compared to the binding affinity of native HA using biolayer interferometry (BLI). We immobilized CD44-Fc onto AHC Octet (Anti-Fc) sensors at 10 µg mL−1 in 1× PBS + 0.005% Tween-20. We then incubated the immobilized CD44 sensor in increasing concentrations of the polymer solutions to measure the association kinetics (kon) followed by an incubation in buffer to determine the dissociation kinetics (koff), from which we calculated the dissociation constant (KD). CD44 binding occurs for P1d′ and P10d′ with KD values of 3.79 ± 0.43 nM and 9.88 ± 0.96 nM, respectively (Fig. 5A, Table 6, entry 1; Table 6, entry 10, Fig. S6). The decrease in binding affinity as compared to native HA (8–15 kDa; KD = 0.123 ± 0.02 nM) is approximately 30 and 80-fold for P1d′ and P10d′, respectively. Interestingly, P1d′ and P10d′ also exhibit the highest pendant HA disaccharide conjugation of the polymer library, with 27% and 24%, respectively (Table 5). Additionally, we attempted to measure the binding affinity of high MW HA (1.5–1.75 MDa) to CD44; however, we were unable to complete the BLI experiment as HA forms a viscous gel-like material at concentrations of 10–100 µM. Using SPR as the measurement technique, other groups report a KD between 10–100 µM.52–54 As a negative control, glcOHPAS shows no binding to CD44 (Fig. S7). | ||
| Fig. 5 Association and dissociation BLI sensograms with immobilized CD44 of (A) P1d′ and (B) HA (8–15 kDa). | ||
| Entry | Polymer | K D (nM) |
|---|---|---|
| Polymers with no binding observed in the range tested are denoted as not detected (n.d.). | ||
| 1 | P1d′ | 3.79 ± 0.43 |
| 2 | P2d′ | n.d. |
| 3 | P3d′ | n.d. |
| 4 | P4d′ | n.d. |
| 5 | P5d′ | n.d. |
| 6 | P6d′ | n.d. |
| 7 | P7d′ | n.d. |
| 8 | P8d′ | n.d. |
| 9 | P9d′ | n.d. |
| 10 | P10d′ | 9.88 ± 0.96 |
| 11 | P11d′ | n.d. |
| 12 | HA | 0.123 ± 0.02 |
| 13 | glcOHPAS | n.d. |
After confirmation of CD44 binding activity, we assessed the cytotoxicity of P1d′ by treating NIH 3T3 mouse fibroblasts with polymer concentrations ranging from 0.0001–1.0 mg mL−1. We treated the cells with the polymer solutions for 24 hours before determining cell proliferation by measuring DNA content. HA (8–15 kDa) is not cytotoxic while P1d′ shows no cytotoxicity up to 0.5 mg mL−1, dropping slightly to 78% viability at 1.0 mg mL−1 (Fig. 6A).
2.6. Hyaluronidase degradation study
Hyaluronidase (HAase) readily degrades HA to give smaller molecular weight species which exhibit reduced in vivo residence times and different biological activity.12,55–57 For example, in the tumor microenvironment (TME) HAase concentrations are increased leading to an accumulation in LMW HA.57 Consequently, native HA CD44-targeting cancer delivery systems are suspectable to degradation and a loss of targeting efficiency. With this in mind, we conducted a HAase degradation study to discern potential enzymatic degradation, using one polymer from each synthetic method: P1d′, P4d′, and P8d′.We incubated the polymers and HA (8–15 kDa) in 1× PBS (pH 7.4) with or without HAase at 37 °C for 24 h.58,59 Next, we lyophilized and analyzed the polymers via GPC equipped with a refractive index detector and 1× PBS as the eluent. As expected, HAase degrades HA as evidenced by the increased GPC retention time (Fig. S8). In contrast, HAase does not degrade polymers P1d′, P4d′, and P8d′, as there is no shift in the GPC retention times (Fig. S9). The emergence of a slightly smaller MW peak is present with P4d′ and P8d′ incubated in PBS only, indicating some hydrolysis occurring in the copolymers, but not in the homopolymer. However, the lack of a shift between the PBS control samples and the HAase samples reveals the polymers are resistant to enzymatic degradation (Fig. S9B and S9C). This resistance to HAase degradation is likely due to the lack of glycosidic linkages and presence of an amide linkage contained within a rigid helical secondary structure shielding it from enzymatic degradation. In further support of this shielding mechanism, stapled glycans exhibit greater enzymatic stability compared to their linear counterparts.60
2.7. In vitro CD44-mediated internalization studies
Given the low nanomolar affinity of P1d′ for CD44, we further investigated its binding and subsequent cellular internalization in three breast cancer cell lines (MDA-MB-231, MCF-7, and MDA-MB-453) with varied surface CD44 expression (Fig. S10, data was downloaded from DepMap portal). Substantially higher CD44 expression is present in the highly invasive breast cancer cell line MDA-MB-231.61,62 Meanwhile, MCF-7 and MDA-MB-453 cell lines, possess moderate and low CD44 expression, respectively.46,63 First, we confirmed the CD44 expression of all three cell lines via immunofluorescence confocal microscopy. As expected, CD44 expression is highest on the MDA-MB-231 cell line followed by moderate expression apparent on the MCF-7 cell line and low expression on the MDA-MB-453 cells, albeit not significant (Fig. 6B and C).To determine potential cellular uptake, we fluorescently labeled P1d′ (rho-P1d′) for visualization by confocal microscopy with rhodamine labeled HA (rho-HA) and glcOHPAS (rho-glcOHPAS) as positive and negative controls, respectively. Following the global deprotection, the CBz protecting group from the polymerization initiator is removed to afford a functionalizable primary amine. Fluorescent labeling was achieved by reacting 5-carboxy-X-rhodamine N-succinimidyl ester with this terminal amine to afford rho-P1d′. Excess rhodamine was removed via dialysis against DI water with a 2.0 kDa MW cutoff dialysis membrane. We treated each cell line with 250 µg mL−1 of rho-P1d′, rho-HA, and rho-glcOHPAS for 6 hours. MDA-MB-231 cells treated with rho-P1d′ show polymer present within the cytoplasm and significantly increased fluorescence as compared to rho-glcOHPAS (Fig. S11). The relative fluorescence of P1d′ is highest within the MDA-MB-231 cells with significantly decreased fluorescence occurring in the MCF-7 and MDA-MB-453 cells (Fig. 6D and E). These data suggest a receptor mediated internalization of P1d′ dependent on CD44 cell surface expression.
The cellular internalization of P1d′ in CD44-positive cancer cells supported evaluation of its potential use as a therapeutic delivery system. Thus, we conjugated the chemotherapeutic, paclitaxel (PTX), to the polymeric construct and investigated its ability to deliver PTX to CD44-expressing breast cancer cells in an effort to enhance therapeutic efficacy and reduce off-target cell death. PTX is one of the most commonly used chemotherapeutic agents for treatment of breast cancer as it inhibits microtubule depolymerization causing cell cycle arrest and ultimately cell death.64,65 However, PTX suffers from significant limitations including poor solubility, requiring the use of polyethoxylated castor oil and dehydrated ethanol (Cremophore EL)66,67 or albumin (Abraxane)68 for delivery, and indiscriminate toxicity due to both healthy and cancerous cells being affected. Consequently, there is a need for alternative and innovative therapeutic modalities to improve its potency, selectivity, and/or specificity.69 Within the HA drug conjugation space, an HA–PTX conjugate (ONCOFID-F-B™) shows promise for clinical translation and is currently undergoing phase III clinical trials for bladder carcinoma (clinicaltrials.gov identifier: NCT05024773). However, HA-drug conjugates that employ native HA are hindered by widely dispersed MWs and HAase degradation, which limit therapeutic efficacy. Thus, we employed structurally defined P1d′ for PTX delivery given its CD44 targeting ability.
First, we coupled the terminal amine of P1d′ to commercially available NHS-functionalized PTX (MedChem Express LLC) to give the PTX–P1d′ conjugate linked via a newly formed amide. Dialysis against DI water using a 2.0 kDa MW cutoff dialysis membrane removes excess PTX and lyophilization of the polymer gives a white, fluffy solid. 1H NMR analysis of PTX–P1d′ shows the presence of the PTX aromatic proton (8.84 ppm) along with integral ratios between PTX and the polymeric H1 (5.69 ppm) that are consistent with the ratios of one PTX per P1d′ polymer chain (Fig. S13). We then incubated CD44 high expressing MDA-MB-231 cells with PTX–P1d′ or PTX alone (5 and 1 μg mL−1 of PTX) for 24 h before a washout and incubation in PTX-free media for two days. At 5 μg mL−1, PTX–P1d′ significantly decreases cell viability as compared to treatment with free PTX demonstrating the importance of the CD44 targeting for improved efficacy (Fig. 6F). This concentration of treatment is well below the threshold of cytotoxicity for P1d′ alone (5 µg mL−1 PTX on PTX–P1d′ is equivalent to 166 µg mL−1P1d′). We hypothesized that PTX–P1d′ is internalized via CD44-mediatd endocytosis and then hydrolyzed to release free PTX leading to cancer cell death, similar to other HA conjugate systems.70 Comparing these results to HA–PTX conjugates in literature, Mittapalli et al. report the treatment of MDA-MB-231 cells with a HA–PTX conjugate results in ∼20% viability following a 1 µM PTX treatment (∼0.9 µg mL−1) compared to a 50% viability observed for equivalent concentrations of free PTX.71 We report similar results with PTX–P1d′, albeit about a 5-fold decrease in cytotoxicity (∼20% viability in MDA-MB-231 cells following treatment with 5 µg mL−1 PTX equivalent). These results are promising when considering the resistance of P1d′ to HAase degradation and synthetic control over MW.
3. Conclusion
We describe the synthesis of a multivalent HA glycopolymer mimetic which targets the CD44 receptor present on cancer cells. The glycopolymer comprises a backbone of glucose repeating units containing amide linkages (termed a polyamidosaccharide; PAS) bearing pendant HA disaccharide moieties. A convergent approach provides the glycopolymer mimetic where a DBCO modified “clickable” PAS is functionalized with the azido-armed HA disaccharide. A series of glycosyl modifications affords the enantiopure glycosylation acceptor with the azido functionality at the anomeric position in good to high yields (74–86%). Similarly, a concise series of good to high yielding reactions (55–90%) give the thoughtfully protected glycosylation donor. A triflate-mediated glycosylation furnishes the diastereopure β-anomer disaccharide in good yields from these monosaccharide building blocks. Next, an anionic ring-opening polymerization of a bicyclic monomer(s) provides the enantiopure PAS homopolymers or copolymers in good to high yields with controlled MW and narrow dispersity. PASs with N repeat units from 25 to 200 are easily obtained. For the homopolymers, the monomer design contains an orthogonally protected silyl O6 position for subsequent installation of a DBCO linker, and generation of the first iteration of a “clickable” PAS. For the copolymerization, we use the aforementioned monomer along with a fully benzylated monomer at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 ratio, respectively, and describe two synthetic approaches given the differences in the polymerization kinetics of the two monomers. Both copolymerization methods followed by chemoselective deprotection and functionalization with the DBCO linker yield copolymers with 25% “clickable” PAS units.
3 ratio, respectively, and describe two synthetic approaches given the differences in the polymerization kinetics of the two monomers. Both copolymerization methods followed by chemoselective deprotection and functionalization with the DBCO linker yield copolymers with 25% “clickable” PAS units.
A SPAAC between the “clickable” polyamidosaccharide and the azido-armed HA disaccharide followed by global deprotection yields: homopolymers P1d′–P3d′ containing 13–27% pendant disaccharide conjugation; copolymers P4d′–P7d′, containing 3–18% disaccharide conjugation; and copolymers P8d′–P11d′ containing 6–24% pendant disaccharide. The range in disaccharide functionality among the copolymers suggests a modulation of the branching density depending on the polymerization method employed. Of the polymers in the library, P1d′ and P10d′ exhibit the highest percent conjugation of the HA disaccharide, and both bind CD44 with nanomolar affinity. Hyaluronidase does not degrade polymers, P1d′, P4d′, and P8d′, whereas HA is readily degraded. Breast cancer cells with high CD44 expression internalize rho-P1d′ while low CD44-expressing cells show decreased uptake, further corroborating the CD44 targeting of P1d′. Conjugating the chemotherapeutic, PTX, to the terminal amine of P1d′ results in decreased cell viability of CD44-expressing cancer cells compared to free PTX highlighting the importance of targeted delivery for improved chemotherapeutic efficacy. To our knowledge, this is one of the first examples of a multivalent HA disaccharide glycopolymer mimetic achieving CD44 targeting, and emphasizes the advantage of using a pendant polymer architecture to overcome the previous linear oligomer of at least six monosaccharides required for CD44 binding.32,33 This work highlights the potential of polysaccharide mimetics as tumor targeting modalities and provides insight into the chemical design space required for such polymers.
4. Experimental section
4.1. General methods and materials
Proton (1H), carbon (13C), and fluorine (19F) NMR spectra were acquired on a 600 MHz Bruker NMR Spectrometer at the Boston University Chemical Instrumentation Center (BU-CIC). All NMR spectra were acquired at ambient temperature with spectral references made to residual solvent peaks (CDCl3 = 7.26 ppm, DMSO = 2.50 ppm, and D2O = 4.79 ppm for 1H NMR, and CDCl3 = 77.16 ppm and DMSO = 39.52 ppm for 13C NMR). Lyophilization was carried out on a Labconco FreeZone 4.5 liter −50 °C benchtop freezer dryer. Gel permeation chromatography (GPC) spectra were acquired on an Agilent Infinity II 1260 instrument equipped with dual angle light scattering and refractive index detectors. THF was used as the eluent for all protected polymers at a flow rate of 1.0 mL min−1 through an Agilent PLgel MIXED-C column (300 × 7.5 mm) fitted with a PLgel 5 µm guard column. Deprotected polymers were characterized by GPC with 1× PBS as the eluent at a flow rate of 0.7 mL min−1 through an Agilent PL aquagel-OH MIXED-M column (300 × 7.5 mm) fitted with a PL aquagel-OH 8 µm guard column. GPC analysis was completed with Agilent GPC/SEC software version 2.2. The dn/dc was calculated via the software assuming 100% mass recovery and in conjunction with the concentration and refractive index detection of each sample. High resolution mass spectrometry (HRMS) results were acquired using a Waters QTOF API US. Circular dichroism (CD) spectra were recorded on an Applied Photophysics CS/2 Chirascan (NSF MRI grant: CHE-112654) with a standard xenon lamp. Column purification of small molecules was performed on a Teledyne combiflash nextgen 300+ automated chromatography system equipped with RediSep columns loaded with normal phase silica. Commercially available reagents and anhydrous solvents were purchased from Sigma Aldrich. All cells were purchased from ATCC and cultured in DMEM (Invitrogen) with 10% fetal bovine serum and 1× penicillin/streptomycin (100 IU mL−1 penicillin, 100 µg mL−1 streptomycin) in a 37 °C incubator under 5% CO2. GraphPad Prism 10 was used for analysis and generating GPC chromatograms and in vitro figures.4.2. Circular dichroism
Polymers P2d′, P5d′, and P9d′ were dissolved in DI water at a 0.2 mg mL−1 concentration. CD measurements were carried out at ambient temperature in a 1 mm path length cuvette.4.3. Biolayer interferometry
CD44-Fc was purchased from Sino Biological (12211-H02H). Biosensors were purchased from Sartorius (Anti Human Fc Capture (AHC); 18-5060). All BLI experiments were performed at ambient temperature on an Octet RED384 instrument and analyzed in Octet Analysis Studio 13.0 in the Center for Macromolecular Interactions (CMI) at Harvard Medical School. After equilibrating the AHC sensors in buffer (1× PBS + 0.005% Tween-20) and establishing a baseline, the sensors were transferred into wells containing 10 µg mL−1 CD44-Fc, or buffer (non-specific binding control), for immobilization onto the sensor. Loading was followed by transfer back into buffer to establish the second baseline. The loaded tips were then transferred into wells containing P1d′–P11d′ (0–100 µM) to allow for association for 240 s before transfer back into buffer wells for dissociation over 400 s. Native HA (8–15 kDa) and glcOHPAS were used as positive and negative controls, respectively. Association was controlled by subtraction of a trace where the tip was not loaded with protein before exposure to polymer ligand for each concentration tested, and dissociation was controlled via subtraction of a trace containing loaded CD44 but no exposure to ligand. Y-axis was aligned to the last five seconds of the second baseline and data was fit to a Mass Transport model.4.4. Cytotoxicity
NIH-3T3 fibroblast cells were seeded into a 96-well plate at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well and cultured for 24 h. Media was then removed and replaced with 0.100 mL of sterile filtered polymer solutions (P1d′ or HA) dissolved in media at concentrations from 0.0001–1.0 mg mL−1. Cells were incubated with polymer solutions for 24 h before removing media again and freezing. After freezing, 0.200 mL of CyQuant working reagent containing lysis buffer (prepared as described by manufacturer) was added to all wells and cells were incubated under light protection at room temperature with shaking for 5 min before mixing thoroughly and transferring 0.150 mL to a black flat-bottom 96-well plate and measuring fluorescence (480 excitation/520 emission). Wells containing no cells were considered 0% proliferative and wells containing cells treated with only media and no polymer were considered 100% proliferative, absorbance values were normalized to this range.
000 cells per well and cultured for 24 h. Media was then removed and replaced with 0.100 mL of sterile filtered polymer solutions (P1d′ or HA) dissolved in media at concentrations from 0.0001–1.0 mg mL−1. Cells were incubated with polymer solutions for 24 h before removing media again and freezing. After freezing, 0.200 mL of CyQuant working reagent containing lysis buffer (prepared as described by manufacturer) was added to all wells and cells were incubated under light protection at room temperature with shaking for 5 min before mixing thoroughly and transferring 0.150 mL to a black flat-bottom 96-well plate and measuring fluorescence (480 excitation/520 emission). Wells containing no cells were considered 0% proliferative and wells containing cells treated with only media and no polymer were considered 100% proliferative, absorbance values were normalized to this range.
4.5. Hyaluronidase degradation study
Bovine testes hyaluronidase (50 µg) was incubated with polymers, HA, P1d′, P4d′, or P8d′ (200 µg), in 0.2 µL of 1× PBS (pH 7.4) at 37 °C for 24 h. Polymer solutions were then lyophilized and redissolved in 0.2 µL of 1× PBS and filtered through a 0.22 µm filter before analyzing via GPC equipped with a refractive index detector. GraphPad Prism 10 was used to analyze GPC peak retention times.4.6. Immunofluorescence confocal microscopy
Confocal microscopy was performed on an Olympus FV3000 Inverted Confocal Microscope. Cells were seeded at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well and incubated in DMEM + 10% fetal bovine serum + 1× penicillin/streptomycin for 24 h before treatment or fixing with 4% paraformaldehyde for 15 min. Blocking was done using 5% goat serum. Human CD44 monoclonal antibody (IM7) (eBioscience, Fisher Scientific) was used as the primary antibody (1
000 cells per well and incubated in DMEM + 10% fetal bovine serum + 1× penicillin/streptomycin for 24 h before treatment or fixing with 4% paraformaldehyde for 15 min. Blocking was done using 5% goat serum. Human CD44 monoclonal antibody (IM7) (eBioscience, Fisher Scientific) was used as the primary antibody (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400), and goat anti-rat AlexaFluor488 (1
400), and goat anti-rat AlexaFluor488 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000) was used as the secondary antibody. Nuclear staining was performed with 1 µg mL−1 DAPI. Imaging settings were optimized to the highest expressing cells.
1000) was used as the secondary antibody. Nuclear staining was performed with 1 µg mL−1 DAPI. Imaging settings were optimized to the highest expressing cells.
4.7. Paclitaxel-P1d′ (PTX–P1d′) cytotoxicity assay
MDA-MB-231 cells were seeded at 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well and incubated for 24 h. Media was then removed and cells were treated with 0.100 mL of sterile solutions of 0, 1, or 5 µg mL−1 of PTX, or the equivalent amount of PTX viaPTX–P1d′; all concentrations of PTX–P1d′ were below the cytotoxicity threshold of P1d′ previously determined, and the final DMSO concentration was 0.1%. Cells were incubated at 37 °C, 5% CO2 for 24 h before removing treatment, replacing with 0.100 mL of fresh media, and incubating another 48 h for a total of 3 days post-treatment. After this time, media was removed and replaced with 0.100 mL of fresh media plus 20 µL of CellTiter 96® AQueous One Solution Cell Proliferation Assay (MTS) Reagent. Cells were incubated in this solution at 37 °C for 2–3 h before measuring absorbance at 490 nm. Wells containing no cells were considered 0% proliferative and wells containing cells treated with 0 µg mL−1 PTX or P1d′–PTX (0.1% DMSO) were considered 100% proliferative, absorbance values were normalized to this range.
000 cells per well and incubated for 24 h. Media was then removed and cells were treated with 0.100 mL of sterile solutions of 0, 1, or 5 µg mL−1 of PTX, or the equivalent amount of PTX viaPTX–P1d′; all concentrations of PTX–P1d′ were below the cytotoxicity threshold of P1d′ previously determined, and the final DMSO concentration was 0.1%. Cells were incubated at 37 °C, 5% CO2 for 24 h before removing treatment, replacing with 0.100 mL of fresh media, and incubating another 48 h for a total of 3 days post-treatment. After this time, media was removed and replaced with 0.100 mL of fresh media plus 20 µL of CellTiter 96® AQueous One Solution Cell Proliferation Assay (MTS) Reagent. Cells were incubated in this solution at 37 °C for 2–3 h before measuring absorbance at 490 nm. Wells containing no cells were considered 0% proliferative and wells containing cells treated with 0 µg mL−1 PTX or P1d′–PTX (0.1% DMSO) were considered 100% proliferative, absorbance values were normalized to this range.
Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information (SI): all additional figures, all synthetic methods, compound characterization, and NMR spectra. See DOI: https://doi.org/10.1039/d5bm01613d.Acknowledgements
Funding for this work is in part from the National Science Foundation Graduate Research Fellowship Program (DGE-1840990 C. D. D.), the National Institutes of Health (T32EB006359 C. D. D. and M. W. G.; F31CA298728 M. K. L.) and the William Fairfield Warren Professorship at Boston University. The content of this publication is the responsibility of the authors and does not necessarily represent the official views of the NIH. We acknowledge support from; the Boston University Chemical Instrumentation Center, namely Maria del Carmen Piqueras, for her support with obtaining our HRMS data; the Center for Macromolecular Interactions at Harvard Medical School for use of the BLI instrument; Alina Ringaci for help with collection of the CD data.Declaration of generative AI and AI-assisted technologies in the writing process: During the preparation of this work the author(s) used no tools or services.
References
- S. Garantziotis and R. C. Savani, Matrix Biol., 2019, 78–79, 1–10 CrossRef CAS PubMed.
- A. Fallacara, E. Baldini, S. Manfredini and S. Vertuani, Polymers, 2018, 10, 701 CrossRef PubMed.
- R. C. Gupta, R. Lall, A. Srivastava and A. Sinha, Front. Vet. Sci., 2019, 6, 192 CrossRef PubMed.
- V. C. Hascall, J. Biol. Chem., 2019, 294, 1690–1696 CrossRef CAS PubMed.
- G. Abatangelo, V. Vindigni, G. Avruscio, L. Pandis and P. Brun, Cells, 2020, 9, 1743 Search PubMed.
- T. Kobayashi, T. Chanmee and N. Itano, Biomolecules, 2020, 10, 1525 CrossRef CAS PubMed.
- A. Lierova, J. Kasparova, A. Filipova, J. Cizkova, L. Pekarova, L. Korecka, N. Mannova, Z. Bilkova and Z. Sinkorova, Pharmaceutics, 2022, 14, 838 CrossRef CAS PubMed.
- K. H. Lu, P. W. Lu, C. W. Lin, E. W. Lu and S. F. Yang, Matrix Biol., 2023, 117, 46–71 CrossRef CAS PubMed.
- H. Kim, M. Shin, S. Han, W. Kwon and S. K. Hahn, Biomacromolecules, 2019, 20, 2889–2903 Search PubMed.
- P. Snetkov, K. Zakharova, S. Morozkina, R. Olekhnovich and M. Uspenskaya, Polymers, 2020, 12, 1800 Search PubMed.
- H. Zhang, S. Huang, X. Yang and G. Zhai, Eur. J. Med. Chem., 2014, 86, 310–317 Search PubMed.
- K. A. Sockett, N.-C. Jiang, P. Ungolan, J. Niu and M. W. Grinstaff, Chem. Rev., 2026 DOI:10.1021/acs.chemrev.5c00790.
- A. M. Carvalho, J. Valcarcel, D. Soares da Costa, M. Gomes, J. A. Vazquez, R. L. Reis, R. Novoa-Carballal and I. Pashkuleva, ACS Appl. Mater. Interfaces, 2022, 14, 41779–41789 CrossRef CAS PubMed.
- E. L. Dane and M. W. Grinstaff, J. Am. Chem. Soc., 2012, 134, 16255–16264 CrossRef CAS PubMed.
- M. Varghese, F. Haque, W. Lu and M. W. Grinstaff, Biomacromolecules, 2022, 23, 2075–2088 CrossRef CAS PubMed.
- A. S. Balijepalli, R. C. Sabatelle, M. Chen, B. Suki and M. W. Grinstaff, Angew. Chem., Int. Ed., 2020, 59, 704–710 CrossRef CAS PubMed.
- C. Chen, S. Zhao, A. Karnad and J. W. Freeman, J. Hematol. Oncol., 2018, 11, 64 CrossRef PubMed.
- M. Hassn Mesrati, S. E. Syafruddin, M. A. Mohtar and A. Syahir, Biomolecules, 2021, 11, 1850 CrossRef CAS PubMed.
- H. Xu, Y. Tian, X. Yuan, H. Wu, Q. Liu, R. G. Pestell and K. Wu, OncoTargets Ther., 2015, 8, 3783–3792 CAS.
- R. Thapa and G. D. Wilson, Stem Cells Int., 2016, 2016, 2087204 CrossRef PubMed.
- H. Ponta, L. Sherman and P. A. Herrlich, Nat. Rev. Mol. Cell Biol., 2003, 4, 33–45 CrossRef CAS PubMed.
- S. Misra, V. C. Hascall, R. R. Markwald and S. Ghatak, Front. Immunol., 2015, 6, 201 Search PubMed.
- Y. Liang, Y. Wang, L. Wang, Z. Liang, D. Li, X. Xu, Y. Chen, X. Yang, H. Zhang and H. Niu, Bioact. Mater., 2021, 6, 433–446 CAS.
- C. Pornpitchanarong, T. Rojanarata, P. Opanasopit, T. Ngawhirunpat and P. Patrojanasophon, Colloids Surf., B, 2020, 196, 111279 CrossRef CAS PubMed.
- P. Thummarati, J. Suksiriworapong, K. Sakchaisri, T. Nawroth, P. Langguth, B. Roongsawang and V. B. Junyaprasert, J. Drug Delivery Sci. Technol., 2022, 77, 103883 Search PubMed.
- X. Zhang, M. Zhao, N. Cao, W. Qin, M. Zhao, J. Wu and D. Lin, Biomater. Sci., 2020, 8, 1885–1896 RSC.
- T. N. Le, C. J. Lin, Y. C. Shen, K. Y. Lin, C. K. Lee, C. C. Huang and N. V. Rao, ACS Appl. Bio Mater., 2021, 4, 8325–8332 Search PubMed.
- M. Bartkowski, V. Bincoletto, I. C. Salaroglio, G. Ceccone, R. Arenal, S. Nervo, B. Rolando, C. Riganti, S. Arpicco and S. Giordani, J. Colloid Interface Sci., 2024, 659, 339–354 CrossRef CAS PubMed.
- H. Y. Yang, M. S. Jang, X. S. Sun, C. L. Liu, J. H. Lee, Y. Li and Y. Fu, Colloids Surf., B, 2023, 228, 113395 CrossRef CAS PubMed.
- S. Sagbas Suner, B. Ari, F. C. Onder, B. Ozpolat, M. Ay and N. Sahiner, Int. J. Biol. Macromol., 2019, 126, 1150–1157 CrossRef CAS PubMed.
- Y. J. Jo, M. Gulfam, S. H. Jo, Y. S. Gal, C. W. Oh, S. H. Park and K. T. Lim, Carbohydr. Polym., 2022, 286, 119303 CrossRef CAS PubMed.
- P. Teriete, S. Banerji, M. Noble, C. D. Blundell, A. J. Wright, A. R. Pickford, E. Lowe, D. J. Mahoney, M. I. Tammi, J. D. Kahmann, I. D. Campbell, A. J. Day and D. G. Jackson, Mol. Cell, 2004, 13, 483–496 CrossRef CAS PubMed.
- J. Lesley, V. C. Hascall, M. Tammi and R. Hyman, J. Biol. Chem., 2000, 275, 26967–26975 CrossRef CAS PubMed.
- T. Slaghek, Y. Nakahara and T. Ogawa, Tetrahedron Lett., 1992, 33, 4971–4974 CrossRef CAS.
- X. Lu, M. N. Kamat, L. Huang and X. Huang, J. Org. Chem., 2009, 74, 7608–7617 CrossRef CAS PubMed.
- D. W. P. Collis, G. Yilmaz, Y. Yuan, A. Monaco, G. Ochbaum, Y. Shi, C. O'Malley, V. Uzunova, R. Napier, R. Bitton, C. R. Becer and H. S. Azevedo, RSC Chem. Biol., 2021, 2, 568–576 RSC.
- R. Xiao, J. Zeng and M. W. Grinstaff, ACS Macro Lett., 2018, 7, 772–777 CrossRef CAS PubMed.
- S. E. Stidham, S. L. Chin, E. L. Dane and M. W. Grinstaff, J. Am. Chem. Soc., 2014, 136, 9544–9547 CrossRef CAS PubMed.
- R. Xiao, E. L. Dane, J. Zeng, C. J. McKnight and M. W. Grinstaff, J. Am. Chem. Soc., 2017, 139, 14217–14223 CrossRef CAS PubMed.
- M. Varghese, K. A. Sockett, S. El-Arid, J. Korunes-Miller, J.-M. Guigner and M. W. Grinstaff, Macromolecules, 2022, 55, 5675–5684 CrossRef CAS.
- R. Xiao, J. Zeng, E. M. Bressler, W. Lu and M. W. Grinstaff, Nat. Commun., 2022, 13, 4661 CrossRef CAS PubMed.
- K. A. Sockett, M. Loffredo, J. Korunes-Miller, M. Varghese and M. W. Grinstaff, Carbohydr. Res., 2022, 522, 108697 CrossRef CAS PubMed.
- A. Battigelli, B. Almeida and A. Shukla, Bioconjugate Chem., 2022, 33, 263–271 CrossRef CAS PubMed.
- J. M. Baskin and C. R. Bertozzi, QSAR Comb. Sci., 2007, 26, 1211–1219 CrossRef CAS.
- M. Mende, M. Nieger and S. Brase, Chem. – Eur. J., 2017, 23, 12283–12296 CrossRef CAS PubMed.
- J. Kandasamy, F. Schuhmacher, H. S. Hahm, J. C. Klein and P. H. Seeberger, Chem. Commun., 2014, 50, 1875–1877 RSC.
- A. A. Grinkova, E. V. Sukhova, N. E. Ustyuzhanina and N. E. Nifantiev, Carbohydr. Res., 2022, 522, 108701 CrossRef CAS PubMed.
- R. Yadav, S. Leviatan Ben-Arye, B. Subramani, V. Padler-Karavani and R. Kikkeri, Org. Biomol. Chem., 2016, 14, 10812–10815 RSC.
- K. J. Loft, P. Bojarova, K. Slamova, V. Kren and S. J. Williams, ChemBioChem, 2009, 10, 565–576 CrossRef CAS PubMed.
- J. Dinkelaar, L. J. van den Bos, W. F. Hogendorf, G. Lodder, H. S. Overkleeft, J. D. Codee and G. A. van der Marel, Chem. – Eur. J., 2008, 14, 9400–9411 CrossRef CAS PubMed.
- A. A. Sherman, O. N. Yudina, Y. V. Mironov, E. V. Sukhova, A. S. Shashkov, V. M. Menshov and N. E. Nifantiev, Carbohydr. Res., 2001, 336, 13–46 CrossRef CAS PubMed.
- P. M. Wolny, S. Banerji, C. Gounou, A. R. Brisson, A. J. Day, D. G. Jackson and R. P. Richter, J. Biol. Chem., 2010, 285, 30170–30180 CrossRef CAS PubMed.
- S. Banerji, B. R. Hide, J. R. James, M. E. Noble and D. G. Jackson, J. Biol. Chem., 2010, 285, 10724–10735 CrossRef CAS PubMed.
- S. Banerji, A. J. Wright, M. Noble, D. J. Mahoney, I. D. Campbell, A. J. Day and D. G. Jackson, Nat. Struct. Mol. Biol., 2007, 14, 234–239 CrossRef CAS PubMed.
- J. Valcarcel, M. R. Garcia, U. R. Varela and J. A. Vazquez, Int. J. Biol. Macromol., 2020, 145, 788–794 CrossRef CAS PubMed.
- J. Lu, Z. Zhao, L. Pan, H. Wu, S. Wang, X. Tong and S. Wu, Mol. Biomed., 2025, 6, 50 CrossRef PubMed.
- W. Donelan, P. R. Dominguez-Gutierrez and S. Kusmartsev, Front. Immunol., 2022, 13, 971278 CrossRef CAS PubMed.
- S. K. Hahn, J. K. Park, T. Tomimatsu and T. Shimoboji, Int. J. Biol. Macromol., 2007, 40, 374–380 CrossRef CAS PubMed.
- G. W. Hong and K. H. Yi, Skin Res. Technol., 2024, 30, e13839 CrossRef PubMed.
- M. G. Ricardo, E. E. Reuber, L. Yao, J. Danglad-Flores, M. Delbianco and P. H. Seeberger, J. Am. Chem. Soc., 2022, 144, 18429–18434 CrossRef CAS PubMed.
- S. Hu, X. Shi, Y. Liu, Y. He, Y. Du, G. Zhang, C. Yang and F. Gao, Cancer Cell Int., 2020, 20, 563 CrossRef CAS PubMed.
- C. Sheridan, H. Kishimoto, R. K. Fuchs, S. Mehrotra, P. Bhat-Nakshatri, C. H. Turner, R. Goulet Jr., S. Badve and H. Nakshatri, Breast Cancer Res., 2006, 8, R59 CrossRef PubMed.
- A. K. Wege, T. F. Dreyer, A. Teoman, O. Ortmann, G. Brockhoff and H. Bronger, Cancers, 2021, 13, 2459 CrossRef CAS PubMed.
- M. Kaveh Zenjanab, S. Alimohammadvand, A. Doustmihan, S. Kianian, B. Sadeghzadeh Oskouei, M. Mazloomi, M. Akbari and R. Jahanban-Esfahlan, J. Drug Delivery Sci. Technol., 2024, 95, 105567 CrossRef CAS.
- T. M. Abu Samaan, M. Samec, A. Liskova, P. Kubatka and D. Busselberg, Biomolecules, 2019, 9, 789 CrossRef CAS PubMed.
- O. van Tellingen, M. T. Huizing, V. R. Panday, J. H. Schellens, W. J. Nooijen and J. H. Beijnen, Br. J. Cancer, 1999, 81, 330–335 CrossRef CAS PubMed.
- H. Gelderblom, J. Verweij, K. Nooter and A. Sparreboom, Eur. J. Cancer, 2001, 37, 1590–1598 CrossRef CAS PubMed.
- E. Miele, G. P. Spinelli, E. Miele, F. Tomao and S. Tomao, Int. J. Nanomed., 2009, 4, 99–105 CAS.
- H. A. Abouzeid, L. Kassem, X. Liu and A. Abuelhana, Cancer Treat. Res. Commun., 2025, 43, 100918 Search PubMed.
- G. Huang and H. Huang, J. Controlled Release, 2018, 278, 122–126 CrossRef CAS PubMed.
- R. K. Mittapalli, X. Liu, C. E. Adkins, M. I. Nounou, K. A. Bohn, T. B. Terrell, H. S. Qhattal, W. J. Geldenhuys, D. Palmieri, P. S. Steeg, Q. R. Smith and P. R. Lockman, Mol. Cancer Ther., 2013, 12, 2389–2399 CrossRef CAS PubMed.
Footnotes |
| † Co-first authors. |
| ‡ These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |



