Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Biofabrication of 3D-printed fibrous scaffolds for large muscle tissue engineering: enhancing scalability, myotube formation and surgical handling
Eva
Schätzlein
 a,
Oliver
Weeger
b,
Sebastian
Scholpp
a,
Oliver
Weeger
b,
Sebastian
Scholpp
 a,
Leon
Faulhaber
a,
Anna
Fritschen
a,
Leon
Faulhaber
a,
Anna
Fritschen
 a,
Jamina Sofie
Gerhardus
a,
Jamina Sofie
Gerhardus
 a,
Robin
Maatz
a,
Annabelle
Neuhäusler
a,
Souraj
Mandal
c,
Regine
von Klitzing
a,
Robin
Maatz
a,
Annabelle
Neuhäusler
a,
Souraj
Mandal
c,
Regine
von Klitzing
 c and
Andreas
Blaeser
*ad
c and
Andreas
Blaeser
*ad
aInstitute for BioMedical Printing Technologies, Technical University of Darmstadt, 64289 Darmstadt, Germany. E-mail: blaeser@idd.tu-darmstadt.de
bCyber-Physical Simulation Group, Department of Mechanical Engineering & Center for Computational Engineering, Technical University of Darmstadt, 64293 Darmstadt, Germany
cSoft Matter at Interfaces, Department of Physics, Technical University of Darmstadt, 64289 Darmstadt, Germany
dCentre for Synthetic Biology, Technical University of Darmstadt, 64289 Darmstadt, Germany
First published on 27th April 2026
Abstract
Volumetric muscle loss has a severe impact on patients’ quality of life, and current treatments often result in poor functional and aesthetic outcomes. This work aims to improve upcoming alternative treatments with tissue-engineered products, which currently are limited in size due to their production process. The methods used for achieving a cell-instructive growth milieu are designed for small volumes, lacking in nutrient supply structures and overall highly manual. To address these issues, we utilize a streamlined printing approach involving fused filament fabrication (FFF)-based spinning to produce fibrous muscle tissue engineering scaffolds. Similar to melt electro writing thin strands are drawn (around 100 µm), while retaining the advantages of FFF printing like fast and reliable production as well as a higher geometric freedom regarding patterns, shape, porosity and height tunable in the relevant range for volumetric muscle loss (width 0.5–15 cm, height 0.1–15 mm). These scaffolds are then combined with two bioinks which are infiltrated deeply into the scaffold with drop-on-demand printing. The first bioink consists of C2C12 myoblasts embedded in a collagen–matrigel matrix, while a second, sacrificial bioink is used to create vascular structures. Numerical simulations allowed for the scaffold design to be tailored, resulting in an anisotropic scaffold capable of repeated elastic deformation and spatio-temporal control of cell orientation, with up to 79.6% of aligned cells. This facilitates local isostatic conditioning, which is expressed in enhanced myotube formation. The tissue precursors simultaneously exhibit high biomechanical congruence (0.9–4.7 MPa), a high suture retention force (3.2 N per stitch) and shape retention (up to 80%), further augmented by the streamlined manufacturing process. These properties are pivotal for its prospective clinical translation.
1. Introduction
Volumetric muscle loss (VML) due to cancer as well as trauma and a following additional heterotopic ossification leads to a significant strength loss of patients. A loss of below 20% of the muscle can already lead to a strength deficit of 30–90%.1–6The self-healing process for VML is limited due to a missing re-vascularization, re-innervation and a missing basement membrane. This is evident from muscle stem cells (satellite cells) that lack contact guidance, secreted growth factors as well as mechanical support, resulting in limited cell division, migration into the wound and alignment.2,7,8
The current clinical gold standard for the treatment of VML are autologous transplants of muscle flaps for example the gracilis muscle flap or minced muscle grafts, however, these approaches have the large drawback of donor site morbidity and unsatisfactory functional and aesthetic recovery.1,9–12 To overcome the limitations of current clinical treatments tissue engineering approaches are explored, where the cell based transplant is pre-cultivated in vitro before implantation.13 Important cues for such tissue generation are the in vitro recapitulation of the in vivo niche,14 including cell types,15 scaffold topology, extracellular matrix (ECM) composition, soluble factors, mechanical properties, stimulation as well as sufficient nutrient supply.16–18
To recreate the natural environment collagen-based hydrogel scaffolds are often used,19–22 as muscle ECM mainly consists of collagen, fibronectin, and laminin.23
It was further shown that mechanical cues aid the formation of aligned and more mature myotubes during tissue formation, increasing the functionality of muscle tissue post implantation.24 For instance an elastic modulus of around 12 kPa, which corresponds with the natural muscle tissue stiffness, aids the differentiation into myotubes.21,25 During this differentiation, the myoblasts compact the tissue and apply a strain on the scaffold.26–28 Cells in scaffolds that withstand this strain exhibited a higher alignment and differentiation.29,30
This shows that many factors are required for functional muscle tissue generation in vitro. While the hydrogel environment is important, collagen-based materials are not stiff enough to offer the mechanical resistance needed to counteract the occurring compaction forces during tissue maturation to improve alignment and formation of more mature myotubes. To counteract these force other working groups used setups applicable mostly for small scale tissues of around 2 mm in diameter which furthermore are highly manual.18,31–33 While other show the need for pre-vascularization31–33 as well as lager supply structures,31 they lack the cell alignment and maturation31 which can be improved with an integrated scaffold in the hydrogel construct.
Various scaffold fabrication technologies exist such as textile engineering,34,35 melt electro writing,36 freeze drying,37 salt-leaching,38 or fused filament fabrication (FFF).39 Of these, FFF is an additive manufacturing technology that offers the highest degree of freedom regarding the patient specific and functional design of such scaffolds.
With FFF-based spinning, which we introduced in our previous work,40,41 the fiber orientation, thickness and pore sizes of the scaffold can be adjusted to actively direct cell alignment and tissue maturation by contact guidance and isometric strain on muscle cells.42,43 In comparison to FFF-printing, the spinning process utilizes increased printing speeds and lowered extrusion multiplicators to achieve fine strands.
The fiber and pore size influence cell orientation. Usually cells align along the fiber with a diameter greater than the cell size, however strands with much lower diameters in combination with a pore size smaller then cell size tend to lead to an orientation perpendicular to the fine strands.42 Fibers produced by FFF-spinning in the 100 µm range could foster alignment in the fiber direction while smaller fiber produced with melt electro writing can also lead to an unwanted alignment perpendicular to the fiber depending on the pore size.42,44
FFF-spinning offers a high surface-to-volume ratio and tunable porosity which is beneficial for cell tissue development and vascularization,45 and therefore improves FFF-printing while retaining the advantages. It is an easy and fast printing process, which allows for improved resulting heights and geometric freedom compared to melt electro writing,46,47 which is key when replacing large individual muscle defects. The materials used in FFF-spinning for tissue engineering are biodegradable thermoplastics such as polylactic acid (PLA) and polycaprolactone (PCL), which show excellent biocompatibility and a slow biodegradation.48–50 For FFF scaffolds to succeed in the translation into clinical practice, the approach must achieve a defined patient specific shape, maintain that shape, scale up for practical use, enable functional tissue formation in vitro pre-implantation and then, being suitable for transporting and fixation to the patient.51,52 Furthermore during the subsequent maturation step in vivo it should remain functional with sufficient nutrient supply and long term elastic deformation.23,48,52
This work addresses the challenges mentioned above by investigating a fiber-reinforced biofabrication approach, combining FFF-spinning and drop-on-demand (DoD)-bioprinting (Fig. 1). In short, a PCL scaffold with muscle mechanics mimicking properties is designed, fabricated and infiltrated with collagen–matrigel growth environment laden with myoblasts. The mechanical behaviour of the scaffold is predicted by a numerical model, which facilitates the computational design of tailorable scaffold properties. The hydrogel environment, coupled with the biomechanical and contact guidance-based cues of the scaffold, enables the targeted maturation of myoblasts into differentiated myotubes. The influence of the scaffolds’ geometric features on the 3D-alignment of cells, as well as their differentiation (Fig. 1). This work demonstrates high viability and good morphology of the cells after printing, as well as complex structuring of sacrificial bio-inks, which will enable the integration of vessels in the future. In addition, the samples showed improved shape and suture retention, underscoring their suitability for use in large muscle patches as well as their high clinical translation potential.
 | ||
| Fig. 1 Schematic overview of the biofabrication of 3D-printed fibrous scaffolds for muscle tissue engineering using a multi-modal printing setup. | ||
2. Results and discussion
To improve tissue engineered muscle for treating volumetric muscle loss, an elastic scaffold supporting muscle differentiation was fabricated and subsequently filled with a collagen–matrigel hydrogel containing myoblasts to match the native muscle mechanics.After scaffold customization, a combined bioprinting process allowing for easy production of large-scale tissues with integrated nutrient supply is introduced, in order to allow for translation to clinical practice. Finally, 3D culture and differentiation of the cells embedded in the collagen-based hydrogel within the scaffolds are analyzed, as well as the suitability for long-term use in vivo.
The scaffold is a FFF-printed grid-like structure from PCL, which is widely used as a printable scaffold material in tissue engineering due to its low melting point, low cytotoxicity and low stiffness compared to other thermoplastics such as PLA.48–50 C2C12 myoblasts are used in this study as a model for skeletal muscle tissue engineering.21,48,53 During differentiation, they fuse and form contracting myotubes.54 As an indicator of early differentiation desmin and for later stage differentiation myosin heavy chain was analyzed.17,55
Fabrication of PCL scaffolds
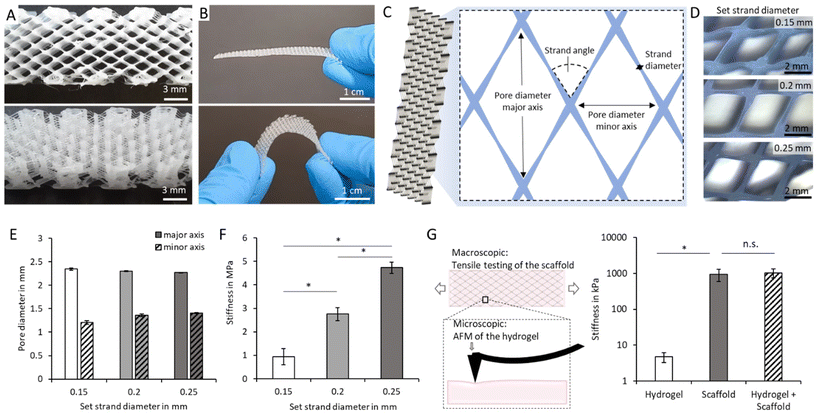
To enable subsequent integration into muscle tissue and to create optimized biomechanical conditions for muscle fiber differentiation, a highly porous PCL scaffold was created. A special modification of the FFF-printing process was used for this purpose (Fig. 2A and B). A combined extrusion and spin technique, called FFF-spinning, allows the creation of highly porous yet elastic structures with textile-like properties (Fig. 2A, B and Video S1).At the same time, the process enables precise adjustment of the relevant scaffold parameters, such as pore diameter, strand diameter, scaffold angle, and scaffold height (Fig. 2C). These parameters directly impact stiffness, elongation and anisotropic behavior of the resulting muscle differentiation supporting scaffold. Using the FFF-spinning technique, the strand diameters can be finely tuned as demonstrated in our previous work.40,41 In short, by using overhangs, increased printing speeds and lowered extrusion multiplicators, fine strands of distinct size can be drawn (Fig. 2D). The method enables controlled fabrication of structures with tailored strand diameters ranging from 160 µm down to 110 µm with little variation (Fig. S1). However, the nodal diameter at the strand intersections remains constant at around 320 µm (Fig. S1).
The scaffold was designed to match the biomechanics of native muscle and, as such, enable up to 10–15% elastic elongation along the major axis, which is known to improve myofiber size,56 alignment and force generation when used during maturation of the tissue.56–58 In addition, it is the expected deformation that the tissue will experience in vivo when implanted in the body allowing for use after implantation.30,59,60 The strand angle was set to 64° in order to achieve the described elongation, while keeping the angle as close to 0° as possible to facilitate cell alignment by contact guidance42 in the direction of the major axis. With this angle, theoretical pore sizes of 2.39, 2.34, 2.29 mm length along the major axis were created. Depending on the strand diameter, the pore sizes varied from 2.35 to 2.27 mm with not significant differences (Fig. 2E). Pore dimensions in the range of the perimysium in the z-axis and of the endomysium in the x- and y-axis were chosen in order to foster tissue maturation.61 At the same time, the pores enable post-fabrication filling of the scaffold with bioinks using drop-on-demand bioprinting.62 To demonstrate the platform character of the FFF-spinning technique, scaffolds with controllable stiffness ranging from 0.9–4.7 MPa were fabricated (Fig. 2F). In this way, scaffolds for different tissue engineering applications, such as cartilage,63 vasculature64–66 or skin tissue67 can be readily produced.68 The combination of the scaffold with the hydrogel filling provides effective means to support muscle differentiation. A strand diameter of 0.15 mm was chosen to achieve a scaffold with a stiffness of around 1 MPa, known to enable retention of compacting myoblasts during differentiation21,69,70 while the hydrogel offers a stiffness of 2.4 kPa (Fig. 2G).
To enable a comprehensive understanding of the reinforcing effect, the (gel-filled) PCL-scaffold was measured in a tensile testing setup, while the stiffness of the integrated hydrogel was assessed using atomic force microscopy (AFM) (Fig. 2G). The results underscore that the overall mechanics are dominated by the PCL-scaffold, which reached a stiffness of 0.9 MPa and 1.1 MPa (when filled with gel). The higher stiffness supports the overall surgical handling of the scaffolds without risking their damage. However, for cell development a bulk material with comparable stiffness would be rather hindering, since the high stiffness impedes differentiation of myoblasts into adult myotubes.25 At this point, the hybrid nature of the fiber reinforced structure can be exploited. While the macromechanics can be tuned with the scaffold, the local micromechanical properties are determined by the integrated hydrogel and offer a supportive microenvironment for myoblast differentiation25 This effect can be utilized to accustom the cellular environment within the range of the cells’ mechanosensing depth (approx. 100 µm).25 Using AFM, known as a suitable test for this purpose,68 the micromechanical stiffness of the gel that was integrated in the PCL scaffold was identified.
While the scaffolds enable the necessary retention at the same time, they offer a large surface area, which facilitates cell interaction and provides contact guidance cues to support cell alignment. Overall, the FFF-spinning process presented here is characterized by its ability to produce highly porous scaffolds from fine fibers while allowing the biomechanical properties to be adjusted (Fig. 2). Comparable results can be achieved, for example, with spacer fabrics71 or by means of melt electro writing.42 While melt electro writing is superior in terms of further reducing the fiber diameter (0.2–50 µm72,73), the process is limited in terms of the individualized shape and, above all, the height of the structures that can be produced (around 5 mm46) which are necessary for large scale defects such as volumetric muscle loss. The spacer fabrics produced using textile technology can be made more elastic and drapable due to the lack of bonds between the fiber intersections (nodal points). However, FFF-spinning enables the necessary geometric freedom to create complex 3D geometries and is economical for small production volumes, which allows its use in personalized medicine with adaptable biomechanical and geometric properties (Fig. 2).46,74,75
Multi-modal biofabrication process
A multi-modal biofabrication process was established which enables an infiltration of cells deep into the scaffolds. For this purpose, the previously described FFF-spinning technology was combined with drop-on-demand bioprinting in one device (Fig. 3A). The fabrication process is conducted successively and starts with the scaffold fabrication with the desired geometry (Fig. 3B, C, Videos S1 and S2).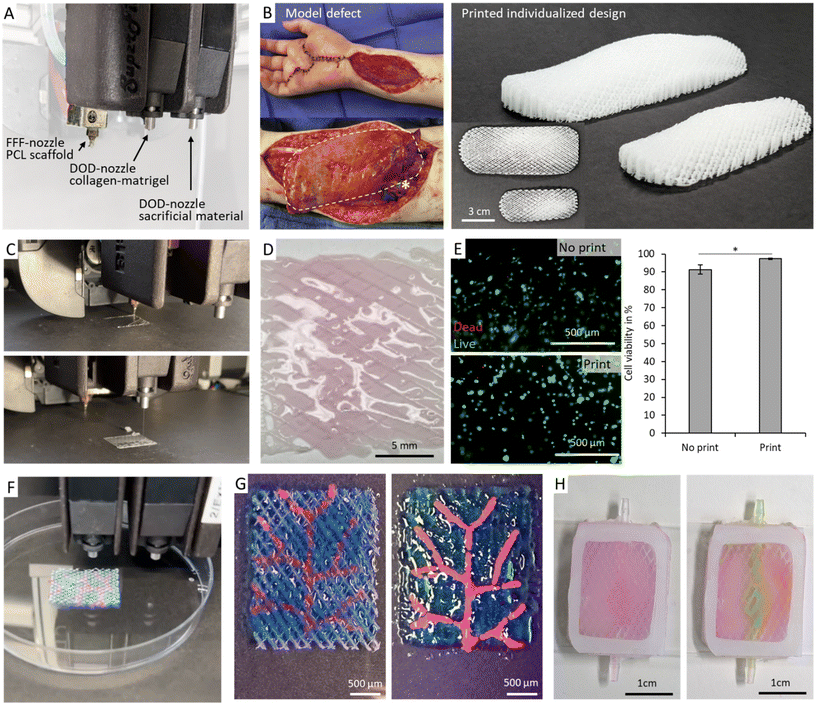 | ||
| Fig. 3 Integrated bioprinting system for manufacturing of the muscle tissues. (A) Overview of the multi-modal printing setup with the customized printer heads for fused filament fabrication (FFF) spinning of the PCL-scaffold as well as drop-on-demand (DoD) printing of cell laden collagen–matrigel hydrogels and DoD printing of sacrificial hydrogels. (B) Model defect of a resection of a synovial sarcoma with a resulting volumetric muscle loss and autologous transplant of the left gracilis muscle flap (highlighted in white)12 (reprinted from Journal of Hand Surgery Global Online, Volume 1, Issue 3, Power et al., ‘Restoration of Sensation and Thumb Opposition Using Nerve Transfers Following Resection of a Synovial Sarcoma of the Median Nerve’, Copyright 2019, with permission from Elsevier). Printed scaffold in two sizes based on the geometry of the model defect. (C) Images of the subsequential printing of the scaffold and hydrogels and (D) resulting printed sample. (E) Fluorescence images (live cells: green, dead cells: magenta) and quantification of the viability of printed and non printed C2C12 cells in collagen–matrigel 6 h after sample preparation (n = 4). (F, G) Subsequential printing of matrix and sacrificial hydrogel for integration of vascular structures in a scaffold and with out a scaffold. (H) Exemplary removal of a sacrificial linar channel of gelatin in a collagen–matrigel matrix in a scaffold and subsequent perfusion with water at 37 °C and 50 µl min−1. Statistical significance * for p < 0.05. | ||
For this we designed a scaffold after images of a case of a volumetric muscle loss after a resection of a synovial sarcoma (Fig. 3B).12 In the study the defect was treated with an autologous transplant of the left gracilis muscle flap with the size of around 4.5 × 10.5 cm. The design was printed in two sizes (1 × 3 × 7 cm, 1.5 × 4.5 × 10.5 cm) to show the robustness of the printing process for VML-relevant geometries.
The FFF-spinner was equipped with an air brush nozzle in order to generate fine grid structures and dissipate heat by radial radiation to prevent re-melting of previously printed layers.76
Precise temperature management was important in order to allow the fabrication of the elastic PCL scaffold and cell-laden hydrogels in one process. PCL was printed with a nozzle temperature of 100 °C, while the baseplate was kept at ambient conditions allowing subsequent infiltration of the scaffold with hydrogels and cells.77,78 The bioink consisting of a collagen–matrigel formulation and C2C12 myoblasts was prepared on ice and cooled to 10 °C in the printer head to prevent premature gelling. Due to its open porous structure (with pores ranging from 1.2 mm to 2.3 mm), the PCL scaffold could easily be infiltrated via drop-on-demand bioprinting, which achieves drop sizes ranging from 0.5 mm to 0.8 mm.62 The scaffold could be filled either homogeneously with a single bioink (Fig. 3C and D) or in a structured way using multiple materials (Fig. 3F, G and Video S5).
The process is well suited for the fabrication of low-viscous bioinks, such as the applied collagen–matrigel matrix.79,80 DoD represents a rather gentle dispensing process involving low shear forces.81 The latter could be demonstrated by the high post-printing viability of C2C12 myoblasts 6 h after printing (Fig. 3D and E). A common bottleneck of DoD, namely its low shape fidelity when processing low viscous bioinks,82 could be overcome by combining it with the FFF-spinning procedure. The scaffold structure stabilized the bioink and retained its shape after printing (Fig. 3G).
Furthermore, using multiple DoD printer heads, additional sacrificial materials or vasculature promoting bioinks can be integrated into the process allowing for sufficient nutrient supply in larger tissue constructs (Fig. 3G). For example, gelatin can be integrated in the collagen–matrigel matrix and flushed out at 37 °C to achieve a perfusable open channel (Fig. 3G, H and Fig. S2).
A combination of a bigger hollow structure with a smaller self-assembled vascular network previously showed a successful nutrient supply of larger structures for treatment of VML applications.31
In our approach the central structure is responsible for immediate nutrient supply after the removal of the sacrificial ink. For this the continuity of the printed construct is a necessity. The printing parameters can be adjusted to ensure a coalescence of the drops, for example by an increased nozzle opening time or higher number of drops per area.83 To achieve the smaller vasculature (roughly 35 µm in diameter), which can not be produced via DoD-printing due to its resolution, we presented a self assembly approach in our previous work.62 Here a combination of cells (dermal fibroblasts and umbilical vein endothelial cells) were placed in a fibrin matrix that supports vascular structure assembly.
Scaffold-hydrogel interactions under load
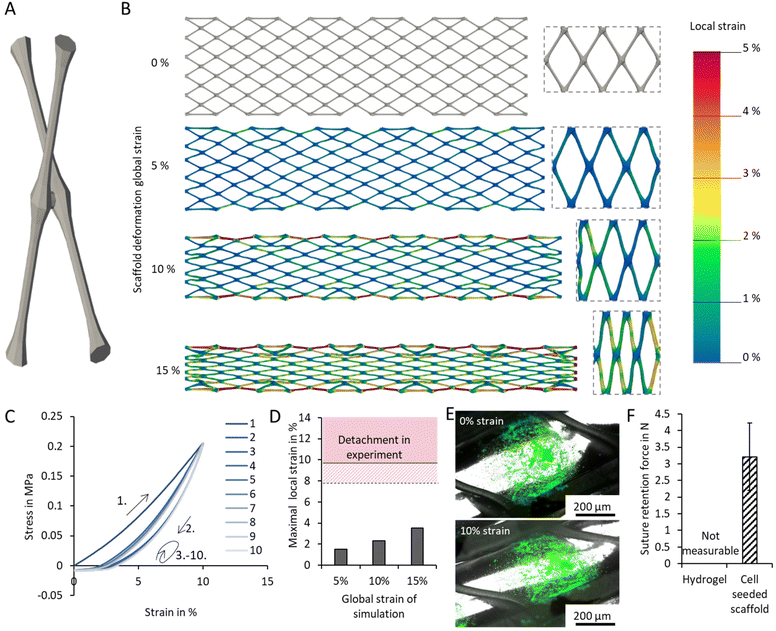
For potential application of the fabricated muscle precursor in future in vivo studies, the fiber-reinforced bioink structure needs to withstand the mechanical load during and after implantation.The scaffolds faces long term deformation of approx. 10–15% strain.56–58 A numerical simulation was carried out to verify that the deformation was in an elastic range of the scaffold and the plastic deformation was minimal. The printing process results in fine strands connected by thicker nodal points. This was translated in the simulation by a design with repetition of a biconical element with a cylindrical connection at the nodal point (Fig. 4A).
Elongating the overall structure by 0–15% resulted in simulated local strains ranging from 0 to 3.5% at the point where the structure bends the most (Fig. 4B). These lowered numbers result from the scaffold design that translates the stretch in the major axis also into stretch in the minor axis. With this, the expected strain lays in the range of the elastic deformation of the bulk material, which was measured experimentally to be in the range of 5% strain (Fig. S3A). In conclusion, the deformation of the printed scaffolds can be assumed to be mostly elastic (Fig. S3B). The simulation results were confirmed experimentally by cyclic tensile testing (Fig. 4C). Except from the first cycle, only little variations in the stress–strain curve could be detected in following load cycles, indicating elastic deformation behaviour of the non-degraded scaffold. We assume that minor plastic deformation occurring at the bridges as a printing artefact on the edge of the scaffold contribute to the initially elevated stress–strain profile. However, with degradation the mechanical properties of the scaffolds change.84,86
Even though PCL samples show no signs of degradation after 7 days in cell culture media in the fourier-transform infrared (FTIR) spectrum in our study (Fig. S3C), other studies were able to confirm beginning of degradation with a slight reduction of the band at 1165 cm−1.85,86 This band related to the amorphous area corresponds to a reduction of the symmetric C–O–C ether group85,86 increasing the overall crystallinity. The degradation speed of PCL is very slow with samples with similar geometry losing around 5% weight after 32 weeks in physiological conditions. Their mechanical properties change significantly with a decline of the peak load break at around 47% of the initial value.84,85 In sum, we expect the fiber-reinforced hydrogel construct suitable for future in vivo tests as the demonstrated elasticity provides the mechanical support during the in vitro maturation and initial in vivo phase followed by a degradation lowering an obstruction of the muscle movement.
To ensure full functionality and prevent delamination, sufficient adhesion of the hydrogel to the scaffold has to be ensured. In a first step, the maximal strain at the interface between scaffold and hydrogel was also determined via simulation (Fig. 4B). In parallel, the maximal strain at which the collagen–matrigel detached form the PCL substrate was experimentally determined to be 9.6% (Fig. 4D). As such, the maximal strain at the delamination point is higher than the expected local strain (3.5%) at maximal elongation of the scaffold.
A validation of the simulation in an experiment showed a simultaneous stretching of the gel with the scaffold (Fig. 4E), however in some parts a detachment of the gel from the scaffold can be noticed, which might be explained with irregularities in printing. In conclusion, the fiber-matrix adhesion in combination with enclosing from the scaffold shows promising results for repetitive deformation cycles.
Finally, suture retention was analyzed to test the sufficient fixation to the wound. A single stitch 3 mm from the edge was made and the maximal force was measured in a tensile testing machine (Fig. 4F and Fig. S4). The study reveals the significance of fiber-reinforcement at this point. The native hydrogel sample was shown to be too weak for suturing. The material was immediately cut by the suture at the slightest force (not even measurable). By reinforcing the gel with the PCL-scaffold, the material exhibited a suture retention force of about 3.2 N per needle stitch, which is approx. 1.5 times higher than shown for native muscle.87 Albeit, more sophisticated suturing techniques were shown to achieve retention strengths of up to 10 N per needle stitch.88 Overall, the occurring forces during suturing are reported to be in the range of 2–5 N,89,90 and during usage of muscles reach 22.5 N cm−2.91 In conclusion, for a hypothetical implantation of the described fiber-reinforced tissue precursor one suture every 0.14 cm2 would be required using the suturing method of this study. Other studies also report the use of biocompatible tissue glue to fix implants to the wound, which would also be possible with our approach. However this method comes with the drawback of a reduced direct contact area and limited cell migration from the surrounding tissue.92
Myotube formation in the hydrogel filled scaffold
C2C12 myoblasts embedded in a collagen–matrigel matrix were placed in the pores of the previously fabricated PCL scaffolds. The mixture of collagen and matrigel is known to provide both, good drop-on-demand printability93–95 and a muscle-like ECM that promotes myoblast differentiation.14,96 In a material screening assay, the beneficial nature of matrigel addition to the collagen with respect to the morphology of embedded C2C12 myoblasts could be demonstrated (Fig. 5A and Fig. S5A). The superior elongation of cells (Fig. 5A and Fig. S5C) as well as the increased metabolic activity (Fig. S5B) underscore the suitability of the chosen matrix as muscle tissue growth environment. The findings were not surprising as collagen61 and laminin97 resemble the most abundant proteins in native muscle tissue. In this regard, the presence of matrigel was an important co-factor for success. It comprises collagen IV, laminin as well as additional growth factors and was shown to support myogenesis,14,96 which we confirmed in our study (Fig. S5A).The FTIR spectra of collagen–matrigel show only slight variations to collagen (Fig. S5D) which is not surprising as the main component is collagen (50 vol%) and matrigel also consist of collagen.14,96,98 After two days of incubation in growth media the presence of matrigel and collagen was visible, indicating that no component was being flushed out. The spectra showed characteristic peaks for methylene-groups in matrigel (∼2930; ∼2870 cm−1)99 and amide groups visible for collagen and matrigel (∼1600–1700 cm−1; ∼1500–1580 cm−1; ∼1230–1300 cm−1)98,99 in the fresh collagen–matrigel gels as well as in gels incubated for 2 days. Minor differences such as relative intensity, shape and shifts of the peak due to local chemical environment indicate an adsorption of medium derived components to the gels.32
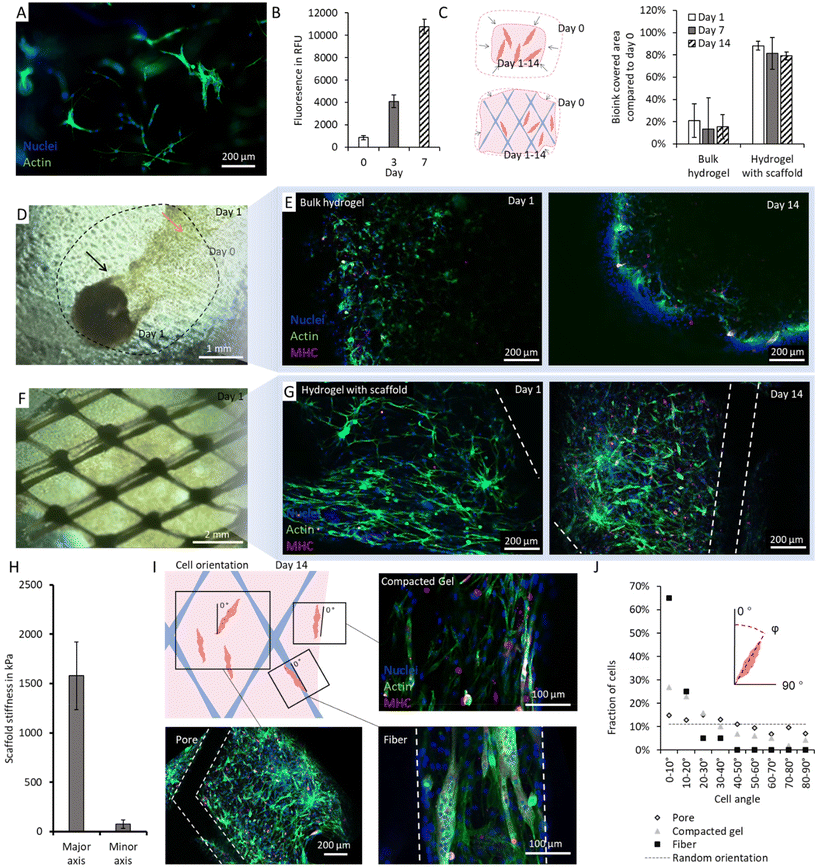
During in vitro tissue culture as well as after implantation, a high level of matrix diffusivity is essential to provide growth factors, nutrients and oxygen to the cells. This is important to bridge the nutrient supply gap before full scale vascularization by the surrounding host tissue is achieved. Using fluorescently labelled dextran the diffusion depth of particles into the hydrogel was measured. Depending on the molecular weight of the particles, diffusion depths ranging from 400–800 µm could be achieved within 120 min (Fig. S2E). The results set the limit for the overall thickness of the scaffold to be designed and show the necessity for integrated supply structures in larger tissue constructs. For this reason, the samples for the following cell studies were prepared with a thickness of approx. 500 µm. With the overall mesh design and biofabrication process in place, the final part of our work focused on the maturation and differentiation of the embedded myoblasts. The selected collagen–matrigel matrix supported C2C12 spreading and maintained a high proliferation potential of the cells in 3D (Fig. 5B).
Both, in the pure hydrogel as well as in the PCL-fiber reinforced structures successful myoblast differentiation could be induced. A dense tissue formation and myosin heavy chain-4 (MHC-4) expression, a myoblast differentiation marker,17,55 could be observed (Fig. 5).
As expected after 14 days in culture, the maturation of the cells leads to a strong compaction of the non-reinforced hydrogel patch (Fig. 5C and D).
Quantitative analysis (Fig. 5D) confirms the visual impression; the native hydrogel samples size decreased to less than 20% of their original size (Fig. 5E), while the scaffold-reinforced samples (Fig. 5G) retained around 80% of their original size after 14 days of culture. In both samples a (not significant) trend towards increased compaction over time, could be observed.
Interestingly, in bulk gels fewer MHC-4 expressing cells as well as a less stretched cell morphology compared to the reinforced samples are visible (Fig. 5F and H). Self-induced isometric conditioning could be a potential explanation for this. The scaffold's resistance to compaction, can result in an isometric tension tensor in the gel, which is known to support muscle cell elongation and myotube formation.21,27 The observed gel compaction is in accordance with literature, where it was found that (depending on the source) cells can exert contraction forces of up to 1 N per 106 cells.29,56,98 In theory, this force could lead to a theoretical deformation of the scaffold, by 5.5% along its major axis. However, visual analysis of the scaffold's compaction after 14 days did not show significant deformation compared to day 0 (Fig. S6).
During the maturation period of 2 and 4 weeks with its occurring contraction of the tissue, there was no delamination of the bioink from the scaffold visible, suggesting that the scaffold with its high porosity supports stable adhesion during long-term culture in vitro (Fig. S5F).
The formation of myofibers in PCL-reinforced samples was scrutinized (Fig. 5E and G). Confocal microscopy revealed a high degree of differentiated and fused myofibers, indicated by multi-nucleated cell bodies. Over the course of 14 days we could not detect a change in the expression of desmin, an early differentiation marker (Fig. S7A). However, an increase in the expression of myosin heavy chain, a later stage differentiation marker could be detected. In terms of size (16–42 µm) and morphology the myofibers were in a similar range, as its native counterpart, albeit at the lower end of its size range (10 and 100 µm) and not yet in perfect homogenously aligned form.56,61 To further increase the tissue functionality additional stimuli during muscle tissue maturation for instance electrical stimulation by external stimulators99 or innervation14 could be employed.59,100
Besides providing mechanical integrity to the overall muscle pre-cursor structure and preventing its shrinkage, the PCL scaffold also allows for cell alignment. The design of the scaffold enabled anisotropic material behavior (Fig. 5H). As such, the scaffold exhibited a preferential deformation axis under tension (stable major axis, compacted minor axis), which was thought to support cell alignment by contact guidance. Interestingly, the anisotropic nature of the scaffold mechanics resulted in spatio-temporally varying muscle cell development in terms of length and orientation (Fig. 5I and J). To analyze this effect, the angle of spread cells to the major axis of the scaffold was assessed. Cells embedded in the center of the pores exhibit only a slight tendency to align with the major axis at 0°. They have an average orientation angle of 38.9°. For reference, a random distribution would result in an average orientation of 45°. Cell cultivated in the gel facing the edge of the scaffold, indicate a higher degree of orientation with an average angle of 26.9°. This can be attributed to the compaction of the gel, which contracts in the minor axis while keeping its shape in the major axis. This effect is routinely used to promote isostatic myoblast differentiation and myotube formation.21,27 The highest degree of alignment could be observed for cells that were directly grown on top of the PCL filaments as it was expected for their strand size.42 The cells were almost parallel with the fibers (angle between fiber and the longitudinal cell axis: 9.2°) and a higher level of MHC is visible (Fig. S7B). The results highlight the importance of additional geometrical cues to provide contact guidance to the cells embedded in the hydrogel and indicate room for future optimization.
As intended, the anisotropic mechanical properties support the alignment of cells along the major axis of the scaffold (Fig. 5J). Even though the isostatic conditioning induces intracellular mechano-transduction processes, we could not detect significant differences in the percentage of nuclei in myosin positive cell depending on the location in the scaffolds (Fig. S7B). These mechano-transduction processes are important cofactors for myotube differentiation, characterized by spindle-like cell spreading, cellular fusion in nascent myotubes, and the formation of striations along the contractile myotubes.25,28,101 Still, the cells in the compacted areas exhibit a higher degree of alignment than the cells in the pores. This indicates a potential route for optimization of the scaffold. To further improve the alignment of the myotubes, the contraction of the tissue in the minor axis needs to be supported by lowering the stiffness in this axis, while maintaining the stiffness of around 1 MPa in the major axis.
3. Summary and outlook
The presented hybrid bioprinting approach offers a potential solution for the future biofabrication of large muscle tissue pre-cursors tailored to patient specific defect geometry (Fig. 6). The study focusses on an important translational challenge, provision of a large cell-instructive and at the same time mechanically supportive scaffold for muscle tissue engineering. The employed FFF-spinning approach allows fabrication of biodegradable scaffolds that match various tissues biomechanics and at the same time provide high suture retention strength. Compared with other fabrication techniques FFF-spinning offers many properties beneficial for treatment of VML. The spinning introduces a higher porosity of the scaffolds compared to FFF-printing resulting in a higher elasticity and perfusability. Furthermore, as presented in this study patient relevant sizes can be printed relatively fast (1.5 h) and are not limited in height by the physical properties of the setup like melt electro writing. Infiltrated with a collagen–matrigel based bioink, the scaffold enables differentiation of C2C12 myoblasts into MHC-expressing, multi-nucleated myotubes. | ||
| Fig. 6 Overview of the multi-modal-bioprinting process to fabricate muscle mechanics mimicking scaffolds with high biofunctionality for treatment of large muscle defects. | ||
In this respect, the anisotropic nature of the scaffold does not only promote myoblast differentiation, but also preserves the structural integrity of the tissue engineering constructs allowing for retention of the patient specific defect shape. In contrast, myoblasts cultivated in the bioink alone lead to deformation of the constructs due to cell compaction forces. Interestingly, the scaffold design was shown to spatio-temporally influence the quality and degree of myotube formation. The combination of lateral tissue compaction and simultaneous longitudinal anchoring in the scaffold contributes to this. The resulting isometric conditioning is particularly evident at the edges of the scaffold. In addition, the results of the study underscore the important role of contact guidance in influencing the alignment and orientation of muscle cells. The highest homogeneity of cellular alignment is found along the scaffold struts.
The demonstrated full integration of scaffold production and filling with hydrogel-cell mixture into a higher-level 3D-bioprinting process offers considerable promise for personalizing biofabrication.
Using the approach in this study individualizing to a patient specific defect is possible. It requires an additional first step where a 3D reconstruction via CT-Scan of the defect can be made as previously described.68 To include a shrinkage in the minor axis of the grid (that can be introduced to further increase alignment) the width of the 3D reconstruction needs to scaled up, which is easily done with commonly used slicers for FFF-printing (e.g. Ultimaker Cura or PrusaSlicer). Then the file needs to be sliced three times, first for the FFF-printing of the grid, second for the bioprinting of the patient specific myoblast laden hydrogels and depending on the size the sacrificial vasculature structures. The major axis of the grid needs to be aligned with the orientation of the muscle and the 3D reconstruction needs to be orientated in a way to have as few as possible overhangs, in order to insure a good print quality. After a subsequent maturation in vitro for 2–3 weeks,52,102 the individualized sample can be implanted in the defect zone of the patient.
By including sacrificial hydrogels nutrient supply structures can be integrated which are of utmost importance for large defects to avoid necrotic tissue cores.18,31
To ensure a long-term stability during perfusion of the printed nutrient supply structures the channels have to be endothelialized. For this, two techniques can be employed, an injection of primary endothelial cells such as human umbilical vein endothelial cells (HUVECs) after dissolving the sacrificial material103,104 or the direct printing of these cells in the sacrificial material.83
Looking ahead, further advancements can be made to both the material as well as the conditioning process. To improve the clinical relevancy primary cells or iPSc derived myocytes should to be included. While the integrated scaffold applies resistance to the contraction forces during the maturation phase of the myoblasts, the setup could be integrated in a more elaborate bioreactor to apply and measure the physiological forces produced by the muscle tissue to allow for an in-depth analysis of the contractility of the tissue.
Incorporation of electro-conductive fibers or neuro muscular junctions, opens up new avenues to promote tissue functionality.57,105 For instance, it was shown that a combined stimulation of the muscle tissue with both mechanical as well as electrical cues improves tissue maturation.16,17
To further strengthen the degree of cellular alignment and fostering myotube formation, the learning cases of this study could be utilized to improve the macro-architecture of the scaffold. For example, further reducing the stiffness of the minor axis of the PCL structure while maintaining the resistance along the main axis would support the tissue compaction effect described above. The lateral contractility made possible by this could extend the effect of improved cell alignment and differentiation, which was previously only observed at the edge of the scaffold, to the entire volume.
Besides providing electro-mechanical support, the scaffold material could also be enhanced by integration of bioactive molecules. Albeit focusing another tissue type, in previous work we already demonstrated the angiogenic effect of bioglass incorporated in PLA filaments.106,107 In case of muscle tissue engineering, vascular sprout supporting molecules could be added to promote vascularization of the construct after implantation.
In summary, it can be concluded that the approach presented in this study stands out due to its biochemical diversity and tailor-made mechanical properties. For this reason, the described procedure can also be understood as a platform technology for the production of other tissues for regenerative medicine. By adjusting the size, strand diameter, and material composition, it is possible, for example, to adapt the scaffold to the mechanical requirements of other tissues (e.g., tendons and ligaments, cartilage, or skin tissue). At the same time, its ease of manufacture, resistance to cyclic loading, inclusion of supply structures and dimensional stability, also offer potential for future clinical translation.
4. Material and methods
Fabrication of muscle-mechanics mimicking PCL-scaffolds using FFF-spinning
For manufacturing the scaffolds a STL model was sliced with the parameters of layer height: 0.2 mm, infill line distance 1.5 mm, infill line width 0.15–0.25 mm (set strand diameter) with the Ultimaker Cura Slicer (Version 4.7.0, Ultimaker; Utrecht, Netherlands), printed at 100 °C with PCL Filament (Facilan™ PCL 100-Filament, 3D4Makers, Haarlem, Netherlands) using a FFF print head equipped with an airbrush nozzle (airbrush-kit, youPrintin3D, Karlsruhe, Germany) a customized 3D-bioprinting system (Superfill, Blackdrop, Aachen, Germany).Scaffolds used for experiments involving cells were sterilized by submerging in 70% ethanol for 30 min, washing three times in PBS (RotiCell 10× PBS, Carl Roth GmbH + Co. KG, Karlsruhe Germany) and drying under the sterile workbench for two hours.
Optical analysis of scaffolds
Microscopic images of the scaffolds were taken (DM4000M, Leica Microsystems, Wetzlar, Germany) and pore sizes (n = 8, N = 3) and strand diameters (n = 7, N = 3) were analyzed using the measuring tool in the Leica Application Suite (LAS, 4.13.0, Leica Microsystems CMS, Switzerland).Mechanical characterization
For mechanical characterization the samples were stretched at room temperature until breaking point with a deformation rate of 10 mm min−1 in a tensile testing machine (Z050, ZwickRoell, Ulm, Germany) (n = 6) using the load cell (KAP-TC/50N, A.S.T. Angewandte System Technik GmbH, Mess und Regelungstechnik, Dresden, Germany) with a preload of 0.1 N. For the suture retention tests, glycolon monofilaments (Size 6–0, HR12, Resorba, Nürnberg, Germany) were chosen. A single stitch 3 mm from the edge of the sample was made. For samples that included a printed scaffold, two strands of the mesh were included in the stitch. The sample was clamped in the lower clamp and the suture thread was fixed in the upper clamp of the tensile testing machine (Fig. S3). The speed was set to 10 mm min−1 and maximal forces were measured (n = 3).To assess the mechanical properties of the collagen–matrigel hydrogel, atomic force microscopy indentation measurements were performed using a JPK NanoWizard 4xp system operating in quantitative imaging (QI) mode. Measurements were conducted at room temperature in Milli-Q water using BL-AC40TS cantilevers (NanoWorld, nominal spring constant k ≈ 0.09 N m−1, resonance frequency ≈ 110 kHz). The experimental spring constant (k ≈ 0.084 N m−1) was determined using the thermal noise method. Force–distance curves were acquired over 2 × 2 µm2 areas at a resolution of 32 × 32 pixels, with each pixel corresponding to an individual force curve. From each condition, 30 curves from three independent scan areas were analyzed. The Young's modulus was extracted by fitting the approach segment of each curve up to 1.2 nN using the Hertz/Sneddon contact model. Variations in trigger force within this range do not significantly influence the extracted modulus values, provided that smooth approach curves are obtained.
Adhesion of collagen–matrigel on PCL sheets was measured by a setup using two coated parallel plates with the hydrogel between them. First, the single use plates were coated with a thin PCL layer and installed in the rheometer (KINEXUS lab+, Malvern, Kassel, Germany). The loading temperature of the hydrogel was set to 10 °C, the plates were closed with a remaining gap of 1 mm. After a gelation time of 30 min at 37 °C the plates were moved apart with a speed of 0.3 mm min−1. The force and elongation perpendicular to the plates was and the maximal force and distance was analyzed.
Numerical modelling of scaffolds
The numerical simulations of the scaffold were performed by modeling it as a 3-dimensional beam structure. For this purpose, a geometric model of the scaffold was build using its as-fabricated parameters. The beam model consists of a representation of the centerline curves of each strut of the scaffold, as well as a description of the corresponding elliptical cross-sections, which vary conically from width 320 µm and height 120 µm at the nodes to width 190 µm and height 88 µm at the centers of the struts. The material behavior of the beams is modelled as elasto-plastic, using the experimentally identified Young's modulus of 350 MPa and yield stress of 18 MPa of the PCL fibers. The nonlinear 3D beam model that allows for large deformations is numerically discretized by an isogeometric collocation method (similar to a finite element analysis), which is described in detail108 and was also validated for other 3D printed polymer materials and scaffolds in.109Hydrogel composition
To prepare the stock solutions, agarose (low gelling temperature, Sigma-Aldrich, St. Louis, USA) was mixed with 1× PBS (RotiCell 10× PBS, Carl Roth GmbH + Co. KG, Karlsruhe Germany) and sterilized in the autoclave. Before use, it was reheated to 70 °C and maintained at 40 °C. For experiments 0.25 wt/vol% agarose was prepared by mixing 2 wt/vol% agarose with growth medium/Cell suspension.To prepare the stock solution of the collagen-based hydrogels 0.3 wt% collagen solution was prepared on ice by mixing collagen (Bovine skin collagen 6 mg ml−1, Sigma-Aldrich, St. Louis, USA), with 0.6 vol% NaOH solution (S2770, Sigma-Aldrich, St. Louis, USA), and 50 vol% growth media/cell suspension in growth medium. For collagen–matrigel containing gels the growth medium content was reduced and 17 vol% matrigel (Matrigel®Matrix Basement Membrane, Corning, NY, USA) was added on ice.
For sacrificial channels 6 wt% gelatin (G1890, Sigma-Aldrich, St. Louis, USA) in PBS was prepared. Before use the gel was reheated to 37 °C. For crosslinking the gel was cooled on ice for 15 min.
Fourier transform infrared spectroscopy (FTIR)
PCL degradation after 7 days and presence of bioink components after 2 days incubation in growth media at 37 °C was assessed using attenuated total reflection fourier-transformed infrared spectroscopy. FTIR spectra were recorded with a Spectrum 3 FTIR spectrometer (PerkinElmer, Waltham, MA, USA). Flat printed samples, were carefully washed in deionized water and dried for 1 min. Hydrogels were dried at room temperature for 1 day. Samples were placed directly onto the ATR crystal. Between individual measurements, the ATR crystal was thoroughly cleaned with acetone. Baseline correction was performed automatically using the Spectrum software (PerkinElmer).Perfusion
A printed scaffold (15 × 20 × 2 mm) was placed in a custom printed perfusion chamber, which was fixed to a glass slide with double-sided tape. First a 6% gelatin channel was pipetted in the chamber before casting the surrounding collagen–matrigel hydrogel. The chamber was sealed with a second glass slide and double-sided tape and stored in the incubator at 37 °C for 2 h for crosslinking of the collagen–matrigel and liquification of the gelatin. The channel was prepared with a needle to ensure a connection of the inlet and the gelatin core. Finally, a microfluidic pump (micro-pump HP, Medorex e.K, Nörten-Hardenberg, Germany) flushed out the gelatin core with fluorescence stained water (Createx, Kaltenkirchen, Germany) at 37 °C and 50 µl min−1.Cell culture
A mouse myoblast cell line (C2C12, CLS, Heidelberg Germany) at passage 31–40 was cultured in cell culture flasks in growth media. The growth media consists of DMEM high glucose (Gibco, Life Technologies Limited, Parsley, United Kingdom) supplemented with 20 vol% FBS (Thermo Fisher Scientific Inc. Waltham, MA, USA) and 1 vol% Penicillin/Streptavidin (Gibco, Life Technologies Limited, Parsley, United Kingdom) until about 80% confluency and then passaged every 2–3 days at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 ratio. To initiate differentiation, a differentiation media was used, containing DMEM high glucose, 2 vol% horse serum (H1270, Sigma-Aldrich, Taufkirchen, Germany), and 1 vol% Penicillin/Streptavidin.
15 ratio. To initiate differentiation, a differentiation media was used, containing DMEM high glucose, 2 vol% horse serum (H1270, Sigma-Aldrich, Taufkirchen, Germany), and 1 vol% Penicillin/Streptavidin.
Cells were removed from the flask by washing with PBS (RotiCell 10× PBS, Carl Roth GmbH + Co. KG, Karlsruhe, Germany) and trypsination for 5 min (TrypLE™ Express Enzym, Thermo Fisher Scientific Inc. Waltham, USA) and resuspended in growth media. For experiments involving differentiation of cells, the cells were cultured in a cell culture incubator (37 °C, 5% CO2, CO2 incubator, Binder, Tuttlingen, Germany) at 100% confluency for two days in growth media prior to harvesting to induce differentiation.
Cell assays
For 3D culture experiments, C2C12 cells were embedded in the different hydrogels. For this the preheated (37 °C) cell suspension and the agarose gel were mixed and immediately used for experiments. For collagen and collagen–matrigel bioinks the growth medium in the hydrogel formulation was replaced by pre-cooled cell suspension. The cell laden hydrogels were either pipetted or bioprinted in a well plate (VWR, Tissue Culture Plate, non-treated, VWR International LLC, PA, USA) and then cooled to 7 °C for 10 min for agarose gels or placed in the incubator at 37 °C for 30 min for collagen and collagen–matrigel hydrogels to allow for complete gelation. Afterwards the samples were covered with cell culture media and cultured in a cell culture incubator at 37 °C and 5% CO2.C2C12 morphology and metabolic activity in the hydrogels was assessed by culturing samples in growth media with a cell concentration of 0.5 × 106 cells per ml in the gels for a week with media exchange every three days (n = 3). Microscopic images for analysis of morphology (number of analyzed cells n > 130 per group) were taken by bright field, phase contrast and fluorescence microscopy with a light microscope (Echo Revolve, Discover Echo Inc. San Diego, USA). Metabolic activity was assessed with a CellTiter Blue assay (Promega Corporation, Fitchburg, USA). 100 µl of medium with 20 µl CellTiter Blue were added to each well and incubated at 37 °C for 2 h. Each day a new cell culture sample was used (n = 3). Fluorescence intensity of the supernatant was read with a plate reader (Infinite M Plex, Tecan Group AG, Männedorf, Switzerland).
Analysis of shape retention and differentiation was conducted using pre-differentiated cells with a concentration of 107 cells per ml in collagen–matrigel for two weeks. The cell laden hydrogel was pipetted into the scaffold or directly on to the well plate. After two days in growth media, the media was exchanged every three days with differentiation media. The area of cell laden hydrogel samples (n = 3) was monitored for two weeks microscopically (Echo Revolve, Discover Echo Inc. San Diego, USA), and successively analyzed using ImageJ (ImageJ 1.54f, National Institutes of Health, USA).
Cell differentiation was assessed using immunofluorescence staining. For this, cells were fixed in 4% paraformaldehyde (Carl Roth GmbH + Co. KG, Karlsruhe, Germany) for 1 h and permeabilized with 0.5% Triton X-100 in PBS (Carl Roth GmbH + Co. KG, Karlsruhe, Germany) for 30 min. For staining of myosin and desmin, samples were first incubated with 5 vol% goat serum (S26-M, Sigma-Aldrich, Taufkirchen, Germany) for 30 min, washed with PBS and then myosin was labeled with either the primary antibody anti-myosin 4 (MF20, eBioscience Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 in PBS or Anit-desmin (Monoclonal Antibody MA5–13259, eBioscience Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) diluted 1
500 in PBS or Anit-desmin (Monoclonal Antibody MA5–13259, eBioscience Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 in PBS. After incubation over night the samples were stained with the secondary antibody AlexaFluor594 goat anti-mouse (A11005, Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) or AlexaFluor488 goat anti-mouse (A11001, Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) for 30 min (1
100 in PBS. After incubation over night the samples were stained with the secondary antibody AlexaFluor594 goat anti-mouse (A11005, Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) or AlexaFluor488 goat anti-mouse (A11001, Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) for 30 min (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 dilution in PBS). Actin filaments were stained with Alexa Fluor 488 Phalloidin diluted 1
500 dilution in PBS). Actin filaments were stained with Alexa Fluor 488 Phalloidin diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 in PBS (A12379, Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) and nuclei for 30 min with DAPI diluted 1
400 in PBS (A12379, Invitrogen, Thermo Fisher Scientific Inc. Waltham, MA, USA) and nuclei for 30 min with DAPI diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 in PBS (MBD0015, Sigma Aldrich, St. Louis, USA). Microscopic images were taken by bright field, phase contrast and fluorescence mode using a light microscope (Echo Revolve, Discover Echo Inc. San Diego, USA). Cell orientation (N = 1, number of cells analysed: n > 180, except for cells on fibers n = 20) and percentage of nuclei in MHC-positive cells of cells differentiated (N = 1, n = 3, number of cells n > 146) for two weeks was assessed microscopically by MHC and actin filament staining and successively analyzed using ImageJ. For this purpose, both a fluorescence microscopy (Echo Revolve, Discover Echo Inc. San Diego, USA) as well as a confocal microscope (TCS SP8 Leica Microsystems, Mannheim, Germany) were employed.
800 in PBS (MBD0015, Sigma Aldrich, St. Louis, USA). Microscopic images were taken by bright field, phase contrast and fluorescence mode using a light microscope (Echo Revolve, Discover Echo Inc. San Diego, USA). Cell orientation (N = 1, number of cells analysed: n > 180, except for cells on fibers n = 20) and percentage of nuclei in MHC-positive cells of cells differentiated (N = 1, n = 3, number of cells n > 146) for two weeks was assessed microscopically by MHC and actin filament staining and successively analyzed using ImageJ. For this purpose, both a fluorescence microscopy (Echo Revolve, Discover Echo Inc. San Diego, USA) as well as a confocal microscope (TCS SP8 Leica Microsystems, Mannheim, Germany) were employed.
For the described bioprinting tests, C2C12 myoblasts at a concentration of 0.5 × 106 cells per ml suspended in collagen–matrigel were used. Cell viability was determined by staining for live cells with fluorescein diacetate (Sigma-Aldrich, St. Louis, USA) and dead cells with propidium iodide (Carl Roth GmbH + Co. KG, Karlsruhe, Germany) diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 60 in Ringer's solution 6 hours post-printing, while the control group was pipetted (n = 4, Cells analyzed n > 380 per group). Quantification of viable and dead cells was done in ImageJ using a threshold and the function “analyze particles”.
60 in Ringer's solution 6 hours post-printing, while the control group was pipetted (n = 4, Cells analyzed n > 380 per group). Quantification of viable and dead cells was done in ImageJ using a threshold and the function “analyze particles”.
Demonstration of DoD-bioprinting into prefabricated scaffolds
Bioprinting experiments were conducted with the same printer that was used for the scaffold structures equipped with drop-on-demand print heads (Black Drop Bioprinter GmbH, Aachen, Germany).All 3D bioprinting experiments were conducted with a 300 µm microvalve (Fritz Gyger, Gwatt, Switzerland), a valve opening time of 450 µs and a printing pressure of 0.4 bar.
To print the embedded vessel-like structures, two print heads were loaded with fluorescence-stained 6% gelatin. The printing temperature was set to 30 °C for the bulk structure (blue) and 37 °C for the vessel-like structures (pink). In the printing process, the bulk structure was printed first into the scaffold, followed by the vessel-like structure. Subsequently, the second bulk structure was printed on top, ensuring complete embedding of the vessel-like structure. A gelation time of 30 to 60 seconds at 7 °C was set between each printing step to ensure full sol–gel transition.
Diffusion in hydrogel
Diffusion in the hydrogel was analyzed as previously described.43 In brief, 350 µl of bioink were casted into a 3D-printed diffusion chamber. A 29 G injection needle was used to create a small microchannel inside the hydrogel. After gelation, the needle was removed and 500 µg ml−1 FITC-Dextran (4 kDa, 70 kDa or 150 kDa, Thermo Fisher Scientific Inc. Waltham, USA) was injected until the microchannel was filled. Microscopic pictures (Echo Revolve, Discover Echo Inc. San Diego, USA) were taken after 0, 10, 30, 60, and 120 min. In between measurements, the chambers were stored in a humid surrounding to ensure minimal evaporation at 23 °C. The distance at which only 50% of the dye was present was analyzed using a custom made MatLab algorithm (The MathWorks, Inc. Natick, MA, USA).Statistical analysis
Statistical significance was determined with a two-sample t-test and one-way ANOVA and Tukey HSD. Graphical data shows the mean with error bars as the standard deviation with significance depicted as n.s. for no significance, * for p < 0.05, with α = 0.05.Author contributions
Eva Schätzlein: conceptualization, data curation, formal analysis, funding acquisition, investigation project administration, validation, visualization, writing – original draft preparation, writing – review & editing, Oliver Weeger: formal analysis, funding acquisition, investigation, resources, software, visualization; Sebastian Scholpp: investigation; Leon Faulhaber: investigation; Anna Fritschen: writing – review & editing; Jamina Sofie Gerhardus: investigation, Robin Maatz: investigation, Annabelle Neuhäusler: investigation, software; Souraj Mandal: investigation; Regine von Klitzing: resources, supervision; Andreas Blaeser: conceptualization, funding acquisition project administration resources, supervision, validation, visualization, writing – review & editing.Conflicts of interest
Andreas Blaeser is co-founder of Black Drop Biodrucker GmbH.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5bm01465d.Acknowledgements
ES, OW and AB acknowledge the funding this project by LOEWE – Landes-Offensive zur Entwicklung Wissenschaftlich-ökonomischer Exzellenz, Förderlinie 5: Exploration (State Offensive for the Development of Scientific and Economic Excellence).We thank the working group Macromolecular Chemistry and Paper Chemistry at the TU Darmstadt for providing access and support to the confocal microscope.
We thank Robin Maatz for development of the collagen–matrigel hydrogel formulation.
Parts of this work were translated and rephrased with DeepL.
References
- S. Testa, E. Fornetti, C. Fuoco, C. Sanchez-Riera, F. Rizzo, M. Ciccotti, S. Cannata, T. Sciarra and C. Gargioli, Biomedicines, 2021, 9, 564 CrossRef PubMed.
- M. E. Carnes and G. D. Pins, Bioengineering, 2020, 7, 1–39 CrossRef PubMed.
- B. T. Corona, K. Garg, C. L. Ward, J. S. McDaniel, T. J. Walters and C. R. Rathbone, Am. J. Physiol., 2013, 305, C761–C775 CrossRef CAS PubMed.
- M. T. A. Li, N. J. Willett, B. A. Uhrig, R. E. Guldberg and G. L. Warren, J. Biomech., 2014, 47, 2013–2021 CrossRef PubMed.
- Y.-J. Choi, Y.-J. Jun, D. Y. Kim, H.-G. Yi, S.-H. Chae, J. Kang, J. Lee, G. Gao, J.-S. Kong, J. Jang, W. K. Chung, J.-W. Rhie and D.-W. Cho, Biomaterials, 2019, 206, 160–169 CrossRef CAS PubMed.
- S. E. Anderson, W. M. Han, V. Srinivasa, M. Mohiuddin, M. A. Ruehle, J. Y. Moon, E. Shin, C. L. San Emeterio, M. E. Ogle, E. A. Botchwey, N. J. Willett and Y. C. Jang, Tissue Eng. Part C Methods, 2019, 25, 59–70 CrossRef CAS PubMed.
- T. A. H. Järvinen, T. L. N. Järvinen, M. Kääriäinen, H. Kalimo and M. Järvinen, Am. J. Sports Med., 2005, 33, 745–764 CrossRef PubMed.
- B. Weining and N. Bursac, IEEE Eng. Med. Biol. Mag., 2008, 27, 109–113 Search PubMed.
- B. T. Corona, J. C. Rivera, J. C. Wenke and S. M. Greising, J. Exp. Orthop., 2017, 4, 36 CrossRef PubMed.
- S. J, D. Ms and D. Ss, Malays. Orthop. J., 2020, 14, 61–73 CrossRef PubMed.
- J.-Yi Guo, X. Chen and Y.-P. Zheng, in 2009 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, IEEE, 2009, pp. 193–196.
- H. A. Power, I. K. Fox, L. C. Kahn and S. E. Mackinnon, J. Hand Surg. Glob. Online, 2019, 1, 185–189 CrossRef.
- K. Garg, J. Brockhouse, C. M. McAndrew, A. J. Reiter, J. G. Owens, R. J. Mueller, G. Pena, A. Ridolfo and D. L. Johnson, J. Physiol., 2025, 1–19 Search PubMed.
- A. Ito, Y. Yamamoto, M. Sato, K. Ikeda, M. Yamamoto, H. Fujita, E. Nagamori, Y. Kawabe and M. Kamihira, Sci. Rep., 2014, 4, 4781 CrossRef PubMed.
- J. H. Kim, I. Kim, Y.-J. Seol, I. K. Ko, J. J. Yoo, A. Atala and S. J. Lee, Nat. Commun., 2020, 11, 1025 CrossRef CAS PubMed.
- C. Borselli, H. Storrie, F. Benesch-Lee, D. Shvartsman, C. Cezar, J. W. Lichtman, H. H. Vandenburgh and D. J. Mooney, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 3287–3292 CrossRef CAS PubMed.
- H. Park, R. Bhalla, R. Saigal, M. Radisic, N. Watson, R. Langer and G. Vunjak-Novakovic, J. Tissue Eng. Regen. Med., 2008, 2, 279–287 CrossRef CAS PubMed.
- S. S. Novakova, B. L. Rodriguez, E. E. Vega-Soto, G. P. Nutter, R. E. Armstrong, P. C. D. Macpherson and L. M. Larkin, Tissue Eng. Part A, 2020, 26, 837–851 CrossRef CAS PubMed.
- O. Aydin, X. Zhang, S. Nuethong, G. J. Pagan-Diaz, R. Bashir, M. Gazzola and M. T. A. Saif, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 19841–19847 CrossRef CAS PubMed.
- Y. Morimoto, H. Onoe and S. Takeuchi, APL Bioeng., 2020, 4, 026101 CrossRef CAS PubMed.
- R. Mestre, T. Patiño, X. Barceló, S. Anand, A. Pérez-Jiménez and S. Sánchez, Adv. Mater. Technol., 2018, 4, 1800631 CrossRef.
- R. Raman, C. Cvetkovic, S. G. M. Uzel, R. J. Platt, P. Sengupta, R. D. Kamm and R. Bashir, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 3497–3502 CrossRef CAS PubMed.
- D. Klumpp, R. E. Horch, U. Kneser and J. P. Beier, J. Cell. Mol. Med., 2010, 14, 2622–2629 CrossRef PubMed.
- E. Schätzlein and A. Blaeser, Commun. Biol., 2022, 5, 737 CrossRef PubMed.
- A. J. Engler, M. A. Griffin, S. Sen, C. G. Bönnemann, H. L. Sweeney and D. E. Discher, J. Cell Biol., 2004, 166, 877–887 CrossRef CAS PubMed.
- A. J. Engler, L. Richert, J. Y. Wong, C. Picart and D. E. Discher, Surf. Sci., 2004, 570, 142–154 CrossRef CAS.
- M. Furuhashi, Y. Morimoto, A. Shima, F. Nakamura, H. Ishikawa and S. Takeuchi, npj Sci. Food, 2021, 5, 1–8 Search PubMed.
- P. Heher, B. Maleiner, J. Prüller, A. H. Teuschl, J. Kollmitzer, X. Monforte, S. Wolbank, H. Redl, D. Rünzler and C. Fuchs, Acta Biomater., 2015, 24, 251–265 CrossRef CAS PubMed.
- A. S. T. Smith, S. Passey, L. Greensmith, V. Mudera and M. P. Lewis, J. Cell. Biochem., 2012, 113, 1044–1053 CrossRef CAS PubMed.
- S. Passey, N. Martin, D. Player and M. P. Lewis, Biotechnol. Lett., 2011, 33, 1513–1521 CrossRef CAS PubMed.
- E. O. Fischer, A. Tsukerman, M. Machour, M. Shuhmaher, A. Silverstein, M. Yaakov, O. Bar-Am, L. Debbi and S. Levenberg, Adv. Healthcare Mater., 2025, 14, 1–16 Search PubMed.
- J. Gilbert-Honick, S. R. Iyer, S. M. Somers, R. M. Lovering, K. Wagner, H. Q. Mao and W. L. Grayson, Biomaterials, 2018, 164, 70–79 CrossRef CAS PubMed.
- J. Gilbert-Honick and W. Grayson, Adv. Healthcare Mater., 2020, 9, 1–27 Search PubMed.
- J. Boublik, H. Park, M. Radisic, E. Tognana, F. Chen, M. Pei, G. Vunjak-Novakovic and L. E. Freed, Tissue Eng., 2005, 11, 1122–1132 CrossRef CAS PubMed.
- S. Scholpp, L. A. Hoffmann, E. Schätzlein, T. Gries, C. Emonts and A. Blaeser, Mater. Today Bio., 2025, 31, 19 Search PubMed.
- G. Cedillo-Servin, O. Dahri, J. Meneses, J. van Duijn, H. Moon, F. Sage, J. Silva, A. Pereira, F. D. Magalhães, J. Malda, N. Geijsen, A. M. Pinto and M. Castilho, Small, 2024, 20, 1–16 CrossRef PubMed.
- I. M. Basurto, G. C. Bandara, R. D. Boudreau, S. B. Shriver, S. A. Muhammad, G. J. Christ and S. R. Caliari, ACS Biomater. Sci. Eng., 2025, 11, 1598–1611 CrossRef CAS PubMed.
- L. Thorrez, J. Shansky, L. Wang, L. Fast, T. VandenDriessche, M. Chuah, D. Mooney and H. Vandenburgh, Biomaterials, 2008, 29, 75–84 CrossRef CAS PubMed.
- T. K. Merceron, M. Burt, Y.-J. Seol, H.-W. Kang, S. J. Lee, J. J. Yoo and A. Atala, Biofabrication, 2015, 7, 035003 CrossRef PubMed.
- E. Schätzlein, C. Kicker, N. Söhling, U. Ritz, J. Neijhoft, D. Henrich, J. Frank, I. Marzi and A. Blaeser, Polymers, 2022, 14, 2389 CrossRef PubMed.
- H. Siebert, D. Spiehl, V. Nienhaus, M. Krauße and E. Dörsam, in Rapid.Tech + FabCon 3.D International Hub for Additive Manufacturing: Exhibition + Conference + Networking, Carl Hanser Verlag GmbH & Co. KG, München, 2019, pp. 148–161 Search PubMed.
- C. Xie, Q. Gao, P. Wang, L. Shao, H. Yuan, J. Fu, W. Chen and Y. He, Mater. Des., 2019, 181, 108092 CrossRef CAS.
- A. Neuhäusler, K. Rogg, S. Schröder, D. Spiehl, H. Zora, E. Arefaine, J. Schettler, H. Hartmann and A. Blaeser, Biofabrication, 2025, 17, 015038 CrossRef PubMed.
- M. de Ruijter, A. Ribeiro, I. Dokter, M. Castilho and J. Malda, Adv. Healthcare Mater., 2019, 8, 1800418 CrossRef PubMed.
- C. Dazzi, K. F. Eichholz, F. E. Freeman, D. J. Kelly and S. Checa, Comput. Biol. Med., 2025, 195, 110574 CrossRef CAS PubMed.
- J. He, G. Hao, Z. Meng, Y. Cao and D. Li, Adv. Mater. Technol., 2022, 7, 9 Search PubMed.
- G. Hochleitner, M. Kessler, M. Schmitz, A. R. Boccaccini, J. Teßmar and J. Groll, Mater. Lett., 2017, 205, 257–260 CrossRef CAS.
- Y. Choi, T. G. Kim, J. Jeong, H. Yi, J. W. Park, W. Hwang and D. Cho, Adv. Healthcare Mater., 2016, 5, 2636–2645 CrossRef CAS PubMed.
- T. K. Merceron, M. Burt, Y.-J. Seol, H.-W. Kang, S. J. Lee, J. J. Yoo and A. Atala, Biofabrication, 2015, 7, 035003 CrossRef PubMed.
- S. Okhovatian, A. Shakeri, L. Davenport Huyer and M. Radisic, Biomacromolecules, 2023, 24, 4511–4531 CrossRef CAS PubMed.
- P. A. Massey, E. Dudoussat, C. Montgomery, W. Scalisi, H. McBride, R. Rutz and G. F. Solitro, Cartilage, 2024, 8 Search PubMed.
- R. K. Bour, G. T. Garner, S. M. Peirce and G. J. Christ, Tissue Eng. Part A, 2025, 31, 373–386 CrossRef CAS PubMed.
- J. Enrione, J. Blaker, D. Brown, C. Weinstein-Oppenheimer, M. Pepczynska, Y. Olguín, E. Sánchez and C. Acevedo, Materials, 2017, 10, 1404 CrossRef PubMed.
- M. L. P. Langelaan, K. J. M. Boonen, R. B. Polak, F. P. T. Baaijens, M. J. Post and D. W. J. van der Schaft, Trends Food Sci. Technol., 2010, 21, 59–66 CrossRef CAS.
- N. Orellana, E. Sánchez, D. Benavente, P. Prieto, J. Enrione and C. A. Acevedo, Foods, 2020, 9, 185 CrossRef CAS PubMed.
- C. A. Powell, B. L. Smiley, J. Mills and H. H. Vandenburgh, Am. J. Physiol., 2002, 283, C1557–C1565 CrossRef CAS PubMed.
- S. Rangarajan, L. Madden and N. Bursac, Ann. Biomed. Eng., 2014, 42, 1391–1405 CrossRef PubMed.
- C. S. Cheng, B. N. J. Davis, L. Madden, N. Bursac and G. A. Truskey, Exp. Biol. Med., 2014, 239, 1203–1214 CrossRef PubMed.
- K. J. M. Boonen, M. L. P. Langelaan, R. B. Polak, D. W. J. van der Schaft, F. P. T. Baaijens and M. J. Post, J. Biomech., 2010, 43, 1514–1521 CrossRef PubMed.
- G. P. Pappas, D. S. Asakawa, S. L. Delp, F. E. Zajac and J. E. Drace, J. Appl. Physiol., 2002, 92, 2381–2389 CrossRef PubMed.
- J. W. Wassenaar, G. R. Boss and K. L. Christman, Biomaterials, 2015, 64, 108–114 CrossRef CAS PubMed.
- A. Fritschen, N. Lindner, S. Scholpp, P. Richthof, J. Dietz, P. Linke, Z. Guttenberg and A. Blaeser, Adv. Healthcare Mater., 2024, 2304028, 1–11 Search PubMed.
- D. L. Robinson, M. E. Kersh, N. C. Walsh, D. C. Ackland, R. N. de Steiger and M. G. Pandy, J. Mech. Behav. Biomed. Mater., 2016, 61, 96–109 CrossRef CAS PubMed.
- A. Karimi, M. Navidbakhsh, A. Shojaei and S. Faghihi, Mater. Sci. Eng. C, 2013, 33, 2550–2554 CrossRef CAS PubMed.
- D. B. Camasão and D. Mantovani, Mater. Today Bio., 2021, 10, 100106 CrossRef PubMed.
- J. A. Chirinos, P. Segers, T. Hughes and R. Townsend, J. Am. Coll. Cardiol., 2019, 74, 1237–1263 CrossRef PubMed.
- M. L. Crichton, B. C. Donose, X. Chen, A. P. Raphael, H. Huang and M. A. F. Kendall, Biomaterials, 2011, 32, 4670–4681 CrossRef CAS PubMed.
- C. F. Guimarães, L. Gasperini, A. P. Marques and R. L. Reis, Nat. Rev. Mater., 2020, 5, 351–370 CrossRef.
- S. F. Eby, B. A. Cloud, J. E. Brandenburg, H. Giambini, P. Song, S. Chen, N. K. LeBrasseur and K.-N. An, Clin. Biomech., 2015, 30, 22–27 CrossRef PubMed.
- R. Souron, F. Bordat, A. Farabet, A. Belli, L. Feasson, A. Nordez and T. Lapole, J. Biomech., 2016, 49, 3534–3537 CrossRef PubMed.
- B. Schäfer, C. Emonts, N. Glimpel, T. Ruhl, A. S. Obrecht, S. Jockenhoevel, T. Gries, J. P. Beier and A. Blaeser, Materials, 2020, 13, 3518 CrossRef PubMed.
- K. Cao, F. Zhang, A. Zaeri, Y. Zhang, R. Zgeib, M. Calzolaio and R. C. Chang, Mater. Des., 2023, 226, 111618 CrossRef CAS.
- J. C. Kade and P. D. Dalton, Adv. Healthcare Mater., 2021, 10, 18 Search PubMed.
- K. M. A. Mueller, A. Hangleiter, S. Burkhardt, D. M. Rojas-González, C. Kwade, S. T. Pammer, S. Leonhardt and P. Mela, Small Sci., 2023, 3, 1–16 Search PubMed.
- S. Loewner, S. Heene, T. Baroth, H. Heymann, F. Cholewa, H. Blume and C. Blume, Front. Bioeng. Biotechnol., 2022, 10, 1–16 Search PubMed.
- R. F. Quero, G. Domingos da Silveira, J. A. Fracassi da Silva and D. P. De Jesus, Lab Chip, 2021, 21, 3715–3729 RSC.
- S. Bardiya, J. Jerald and V. Satheeshkumar, Mater. Today Proc., 2021, 39, 1362–1366 CrossRef CAS.
- C.-Y. Lee and C.-Y. Liu, Addit. Manuf., 2019, 25, 196–203 CAS.
- X. Lan, A. Adesida and Y. Boluk, Biomed. Mater., 2022, 17, 062005 CrossRef CAS PubMed.
- D. Janzen, E. Bakirci, A. Wieland, C. Martin, P. D. Dalton and C. Villmann, Adv. Healthcare Mater., 2020, 9, 6 Search PubMed.
- A. Blaeser, D. F. Duarte Campos, U. Puster, W. Richtering, M. M. Stevens and H. Fischer, Adv. Healthcare Mater., 2016, 5, 326–333 CrossRef CAS PubMed.
- A. Blaeser, D. F. Duarte Campos and H. Fischer, Curr. Opin. Biomed. Eng., 2017, 2, 58–66 CrossRef.
- J. Schöneberg, F. De Lorenzi, B. Theek, A. Blaeser, D. Rommel, A. J. C. Kuehne, F. Kießling and H. Fischer, Sci. Rep., 2018, 8, 1–13 Search PubMed.
- J. M. Anaya-Mancipe, A. C. de Figueiredo, L. G. Rabello, M. L. Dias and R. M. da Silva Moreira Thiré, J. Appl. Polym. Sci., 2024, 141, 1–16 CrossRef.
- C. X. F. Lam, M. M. Savalani, S. H. Teoh and D. W. Hutmacher, Biomed. Mater., 2008, 3, 034108 CrossRef PubMed.
- M. V. Deshpande, A. Girase and M. W. King, Polymers, 2023, 15, 3819 CrossRef CAS PubMed.
- M. He, S. J. Sebastin, A. W. T. Gan, A. Y. T. Lim and A. K. S. Chong, Ann. Plast. Surg., 2014, 73, 333–335 CrossRef CAS PubMed.
- J. F. Kragh, S. J. Svoboda, J. C. Wenke, J. A. Ward and T. J. Walters, J. Trauma, 2005, 59, 209–212 CrossRef PubMed.
- J. Oguma, S. Ozawa, Y. Morikawa, T. Furukawa, Y. Kitagawa, M. Ueda and M. Kitajima, J. Surg. Res., 2007, 140, 129–134 CrossRef PubMed.
- S. P. Rodrigues, T. Horeman, J. Dankelman, J. J. van den Dobbelsteen and F. W. Jansen, Surg. Endosc., 2015, 29, 1982–1989 CrossRef CAS PubMed.
- R. L. Lieber and S. R. Ward, Philos. Trans. R. Soc. B Biol. Sci., 2011, 366, 1466–1476 CrossRef PubMed.
- J. Gilbert-Honick, B. Ginn, Y. Zhang, S. Salehi, K. R. Wagner, H.-Q. Mao and W. L. Grayson, Cell Transplant., 2018, 27, 1644–1656 Search PubMed.
- A. Fritschen, M. Acedo Mestre, S. Scholpp and A. Blaeser, Front. Bioeng. Biotechnol., 2023, 11, 1–12 Search PubMed.
- M. Köpf, D. F. D. Campos, A. Blaeser, K. S. Sen and H. Fischer, Biofabrication, 2016, 8, 1–15 CrossRef PubMed.
- F. Kreimendahl, M. Köpf, A. L. Thiebes, D. F. Duarte Campos, A. Blaeser, T. Schmitz-Rode, C. Apel, S. Jockenhoevel and H. Fischer, Tissue Eng. Part C Methods, 2017, 23, 604–615 CrossRef CAS PubMed.
- S. Grefte, S. Vullinghs, A. M. Kuijpers-Jagtman, R. Torensma and J. W. Von den Hoff, Biomed. Mater., 2012, 7, 055004 CrossRef CAS PubMed.
- J. R. Sanes, J. Biol. Chem., 2003, 278, 12601–12604 CrossRef CAS PubMed.
- M. Kolodney and R. Wysolmerski, J. Cell Biol., 1992, 117, 73–82 CrossRef CAS PubMed.
- H. Fujita, T. Nedachi and M. Kanzaki, Exp. Cell Res., 2007, 313, 1853–1865 CrossRef CAS PubMed.
- J. J. Ross, M. J. Duxson and A. J. Harris, Development, 1987, 100, 395–409 CrossRef CAS PubMed.
- H. Stedman, H. Sweeney and A. Kelly, Nature, 1973, 246, 170 Search PubMed.
- M. A. Machingal, B. T. Corona, T. J. Walters, V. Kesireddy, C. N. Koval, A. Dannahower, W. Zhao, J. J. Yoo and G. J. Christ, Tissue Eng. Part A, 2011, 17, 2291–2303 CrossRef CAS PubMed.
- V. K. Lee, D. Y. Kim, H. Ngo, Y. Lee, L. Seo, S. Yoo, P. A. Vincent and G. Dai, Biomaterials, 2015, 35, 8092–8102 CrossRef PubMed.
- J. S. Miller, K. R. Stevens, M. T. Yang, B. M. Baker, D.-H. T. Nguyen, D. M. Cohen, E. Toro, A. A. Chen, P. A. Galie, X. Yu, R. Chaturvedi, S. N. Bhatia and C. S. Chen, Nat. Mater., 2012, 11, 768–774 CrossRef CAS PubMed.
- M. L. P. Langelaan, K. J. M. Boonen, K. Y. Rosaria-Chak, D. W. J. van der Schaft, M. J. Post and F. P. T. Baaijens, J. Tissue Eng. Regen. Med., 2011, 5, 529–539 CrossRef PubMed.
- E. Schätzlein, S. Al Zoghool, N. Söhling, D. Henrich, J. Frank, E. Dörsam, I. Marzi and A. Blaeser, in 3. Tag der Rhein-Main-Universitäten 13.09.2021, 2021.
- S. Cichos, E. Schätzlein, N. Wiesmann-Imilowski, A. Blaeser, D. Henrich, J. Frank, P. Drees, E. Gercek and U. Ritz, Int. J. Bioprint., 2023, 9, 751 CrossRef PubMed.
- O. Weeger, D. Schillinger and R. Müller, Comput. Methods Appl. Mech. Eng., 2022, 399, 115456 CrossRef.
- O. Weeger, I. Valizadeh, Y. Mistry and D. Bhate, Addit. Manuf. Lett., 2023, 4, 100111 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |