 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A tip-on-tip micro-solid-phase extraction HILIC-LC-MS/MS platform for the determination of urinary methylated nucleosides
C.
Mena-Iglesias
 ,
A.
Ballester-Caudet
,
A.
Ballester-Caudet
 ,
D.
García-Gómez
,
D.
García-Gómez
 ,
M.
Bustamante-Rangel
,
M.
Bustamante-Rangel
 *,
E.
Rodríguez-Gonzalo
*,
E.
Rodríguez-Gonzalo
 and
J. L. Pérez
Pavón
and
J. L. Pérez
Pavón

Department of Analytical Chemistry, Nutrition and Food Science, Faculty of Chemical Sciences, Universidad de Salamanca, Plaza de Los Caídos s/n, Salamanca, 37008, Spain. E-mail: mbr@usal.es; Fax: +34-923294483; Tel: +34-923294483
First published on 6th March 2026
Abstract
Methylated nucleosides (mNSs) excreted in urine are considered biomarkers associated with RNA turnover and disease states such as cancer and can be detected through non-invasive sampling. In this work, we report the development and validation of a sensitive hydrophilic interaction liquid chromatography coupled to tandem mass spectrometry (HILIC-LC-MS/MS) method for the simultaneous quantification of fifteen mNSs and related nucleobases in human urine. Due to the complexity of the biological matrix and the need to quantify low-abundance biomarkers, a sensitive and selective micro-scale solid-phase extraction (µ-SPE) using a tip-on-tip format with ENV+ sorbent is proposed as the most convenient strategy for sample preparation. This approach offers the advantages of rapid extraction, minimal solvent consumption, and simplified sample handling. The method is simple, effective, affordable, reproducible, and easily scalable. Additionally, HILIC-based chromatographic separation enabled effective retention and high resolution. According to international guidelines on endogenous analytes, the proposed method demonstrated excellent linearity (R2 > 0.99), low limits of detection (1–12 µg L−1), and acceptable intra- and inter-day precision (CV < 15%). Recovery values ranged from 85% to 120% across all analytes. This novel platform enables reliable and high-throughput profiling of urinary mNSs and offers strong potential for clinical diagnostics applications and population-based biomonitoring of RNA epigenetic signatures.
1. Introduction
In recent years, RNA chemical modifications have attracted increasing attention due to their central role in the post-transcriptional regulation of gene expression.1,2 Among these modifications, methylated nucleosides (mNSs), such as N6-methyladenosine (m6A), 1-methyladenosine (m1A), and 2′-O-methylcytidine (Cm), stand out for their functional relevance.3,4 These molecules are nucleosides/bases bearing a methyl group at specific positions, and they arise either from physiological nucleic acid turnover or as a result of pathological processes such as DNA damage induced by nitrosative stress.5,6Because most methylated nucleosides are not recycled through standard salvage pathways, they tend to accumulate intracellularly and are eventually excreted into biological fluids such as urine, blood, or saliva.7,8 In contrast to blood, which necessitates invasive sampling, urine and saliva can be collected non-invasively and are therefore particularly advantageous for biomarker investigations. Their presence in these matrices allows detection and quantification, positioning them as potential endogenous biomarkers for various chronic and hard-to-diagnose diseases, including cancer, neurodegenerative disorders, and renal dysfunction.9–11
Elevated levels of urinary methylated nucleosides have been associated with metabolic imbalances, inflammation, oxidative stress, and tumor presence,12 including breast,13,14 ovarian,15,16 and colorectal17 cancers. In some cases, these methylated compounds may offer higher sensitivity and specificity than traditional tumor biomarkers.18,19 Moreover, certain modifications—such as m6A—have been directly implicated in cancer-related pathways, including tumor progression,4,20 metastasis,21,22 drug resistance,23 and immune microenvironment remodeling,24 particularly in hematological malignancies like acute myeloid leukemia.25–27 Notably, 5-methyluridine and its derivatives contribute to RNA stability, translational regulation, and cellular homeostasis,28,29 and abnormal levels have been linked to pathological processes including cancer.28 Similarly, 3-methyladenine has been extensively studied due to its dual role as both an autophagy inhibitor30 and a modulator of cancer cell response to stress and treatment. It can prevent energy stress-induced necrotic death,31 influence cell survival mechanisms, and enhance susceptibility to anticancer drugs such as sorafenib in hepatocellular carcinoma.32 These findings highlight that such nucleoside modifications play a key role in cancer pathology and represent promising biomarker candidates. Importantly, their low concentrations in biological fluids are often near or below the detection limits of conventional analytical methods. Therefore, advanced sample preparation strategies, particularly microscale solid-phase extraction, are essential for achieving reliable enrichment and quantification in clinical and translational research applications.
Given their clinical significance, the accurate quantification of methylated nucleosides in biological matrices requires highly robust and sensitive analytical techniques.33–35 In this context, nanopore sequencing and immunoprecipitation have provided new insights into the structural and functional roles of RNA modification.10,36 However, issues such as isomer misidentification and matrix effects remain a challenge in routine analysis.37–39 In response to these analytical challenges, stable isotope dilution strategies combined with high-performance liquid chromatography-mass spectrometry have been implemented.40,41 Liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) has become the technique of choice for the determination of nucleosides in biological matrices, offering excellent separation efficiency, sensitivity and selectivity.42–44 Within this framework, hydrophilic interaction liquid chromatography-tandem mass spectrometry (HILIC-MS/MS) has proven particularly effective for the analysis of polar and hydrophilic compounds, including methylated nucleosides and related metabolites.33,35,38,45–47 Several studies have demonstrated the superior retention and chromatographic resolution achieved with HILIC in complex biological fluids such as urine or plasma.13,14,48
Despite the availability of well-established HILIC-MS/MS methods, comparative evaluations of sample pretreatment strategies remain scarce.19 Among the sample preparation strategies for hydrophilic analyte detection,45 solid-phase extraction (SPE)—using HLB, MCX, or PBA sorbents49—has become the most widely used technique due to its simplicity, versatility, and high efficiency in reducing matrix effects and concentrating analytes, in comparison with chemical derivatization46 or protein precipitation and/or liquid–liquid extraction (LLE), which often requires additional steps and may introduce variability.47,48
In addition, on-line SPE has been described as an attractive automated alternative, directly integrating sample cleanup into the chromatographic workflow to reduce manual handling and improve reproducibility.50 Effective on-line SPE-LC-MS/MS methods have been reported for the quantification of nucleosides and nucleobases in complex biological fluids such as urine and saliva.35 Nevertheless, these systems usually require dedicated instrumentation and may not be easily adaptable to diverse biofluids.
More recently, miniaturized SPE formats have emerged as high-throughput alternatives for bioanalytical applications.51 In particular, tip-on-tip micro solid-phase-extraction (TOT-µSPE) has attracted considerable interest. In this approach, the sorbent material is packed into pipette tips, enabling efficient low-volume extractions with minimal solvent consumption and high throughput using standard laboratory pipettes. Compared with conventional SPE, TOT-µSPE provides faster extraction, reduced solvent use, simplified sample handling and improved clean-up performance. These advantages have been demonstrated in various applications, including pipette-tip micro-SPE for small molecules in serum52 or tip-based pretreatment workflows in proteomics.53–55 In this study, we report the development and validation of an HILIC-MS/MS-based analytical approach for the simultaneous quantification of fifteen methylated nucleosides in human urine. The structures of these compounds are shown in the SI (Fig. S1). Tip-on-tip micro solid-phase-extraction is proposed as a more reliable sample pretreatment strategy for urine analysis compared to conventional methods. This approach provides fast extraction, requires minimal solvent consumption, and involves straightforward sample preparation. The procedure is easy to implement, efficient, reliable, and readily scalable. Method performance is evaluated in terms of linearity, sensitivity, precision, recovery, and matrix effects, aiming to contribute to the development of an efficient analytical platform for non-invasive epigenetic biomarker profiling. The proposed method is compared with a reference method consisting of diluting urine with acetonitrile (ACN) in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 (v/v) ratio and applying it for the analysis of methylated nucleosides in urine samples. Dilution is one of the simplest and most widely used sample preparation strategies for trace analysis, as it involves minimal sample handling and does not introduce additional extraction or enrichment steps.
50 (v/v) ratio and applying it for the analysis of methylated nucleosides in urine samples. Dilution is one of the simplest and most widely used sample preparation strategies for trace analysis, as it involves minimal sample handling and does not introduce additional extraction or enrichment steps.
2. Materials and methods
2.1. Chemicals and reagents
All analytical standards were of high purity (>96%). 1-Methyladenosine (m1A), 1-methylguanosine (m1G), 1-methylinosine (m1I), 3-methylcytidine (m3C), 3-methyladenine (m3A), 5-methylcytosine (m5Cs), 5-methyluridine (m5U), N6-methyladenosine (m6A) and 7-methylguanosine (m7Gs) were purchased from TargetMOL (Boston, MA). 2′-O-Methylcytidine (Cm) and 2′-O-methylguanosine (Gm) were supplied by Thermo Scientific (Waltham, Massachusetts). 6-O-Methylguanine (m6G) and 7-methylguanine (m7G) were obtained from Tokyo Chemical Industry (Tokyo, Japan), 5-methyluracil (m5Uc) was supplied by Fluorochem (Derbyshire, England) and 5-methylcytidine (m5C) was obtained from Glentham Life Sciences (Corsham, England). Creatinine (≥98%), methanol (CH3OH, HPLC grade), ammonium acetate (NH4Ac, ≥97%) and formic acid (HCOOH) were purchased from Sigma-Aldrich (Steinheim, Germany). Acetonitrile (ACN, 99.9%) was obtained from Fisher Scientific (Pittsburgh, USA). Ammonia (NH3) was supplied by Scharlau (Barcelona, Spain), and the ultra-high quality (UHQ) water used throughout the study was obtained with a Wasserlab Ultramatic water purification system (Nonain, Spain). For the preparation of the tip-on-tip devices, empty 50 µL DPX single-phase filtration tips, model DPX170247 (DPX Technologies; Columbia, USA), were used. The sorbent materials used to fill the tips were obtained from the following SPE cartridges: Oasis® MCX (3 mL, 60 mg) and Oasis® HLB (3 mL, 60 mg) from Waters (Barcelona, Spain); Bond Elut PBA (3 mL, 60 mg) from Agilent Technologies (Waldbrom, Germany); and Isolute® ENV+ (3 mL, 200 mg) from Biotage (Uppsala, Sweden). The sorbents were removed from the cartridges, and the appropriate amount of each material was weighed and transferred to the DPX tips.2.2. Standard solutions
Individual aqueous solutions of m1A, m1G, m1I, m3C, m3A, m5Cs, m5U, m6A, m7Gs, Cm, Gm, m6G, m7G, m5Uc and m5C were prepared at a concentration of 100 mg L−1 and stored at 4 °C. Working mixture solutions were freshly prepared each day by diluting stock solutions in water (Mili-Q) for TOT-µSPE treatment or in ACN in the case of reference method. Calibration solutions were prepared by diluting the working solutions to achieve the desired concentrations of analytes. Each calibration level was prepared in triplicate.2.3. Sample collection and pretreatment
Urine samples were collected from apparently healthy adults (25–48 years old) of both sexes and were directly stored at −20 °C until analysis. All participants were informed about the objective of the study, and written informed consent was obtained. Samples were used exclusively for the development and validation of this analytical method. The study was conducted in accordance with the Declaration of Helsinki and approved by the Clinical Research Ethics Committee of the Salamanca Health Area (protocol code: CTQ2013-47993-P/BQU; date of approval: 21 November 2013).After thawing at room temperature, samples were centrifuged at 1800 rpm for five minutes to remove precipitated proteins. Subsequently, the supernatants were subjected to the TOT-µSPE procedure (Fig. 1A), which was selected as the main pretreatment strategy due to its efficiency and reproducibility for the enrichment of mNSs. For this purpose, tip-on-tip microextraction devices were manually prepared by introducing approximately 15 mg of Isolute ENV+ sorbent, based on a hypercrosslinked polystyrene-divinylbenzene matrix that combines non-polar π–π interactions with polar retention sites, into DPX single phase filtration tips. The polymeric ENV+ phase provides a broad affinity range suitable for both purine and pyrimidine derivates. This balanced selectivity allows efficient extraction of chemically diverse nucleosides. The frit and barrier integrated in these tips prevent the sorbent from leaking out of the tip. These custom-made tips were used for a single extraction and discarded after use to avoid cross-contamination.
Following method optimization, the final extraction conditions were established as follows: conditioning with 150 µL of ACN and 150 µL of H2O, loading with 150 µL of urine sample, and elution with 500 µL of ACN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 (90
NH3 (90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, v/v). The eluate was evaporated under a nitrogen stream and reconstituted in 50 µL of ACN
10, v/v). The eluate was evaporated under a nitrogen stream and reconstituted in 50 µL of ACN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH4Ac (95
NH4Ac (95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, v/v), which was transferred to HPLC vials. A volume of 10 µL was injected into the LC-MS/MS system for analysis. During each aspiration step, a volume of solvent was drawn into the tip along with a similar volume of air, ensuring an effective agitation between the sample and sorbent that improved mass transfer between the sorbent and the liquid phase.
5, v/v), which was transferred to HPLC vials. A volume of 10 µL was injected into the LC-MS/MS system for analysis. During each aspiration step, a volume of solvent was drawn into the tip along with a similar volume of air, ensuring an effective agitation between the sample and sorbent that improved mass transfer between the sorbent and the liquid phase.
For comparison purposes, a reference pretreatment approach consisting of a simple 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 (v/v) dilution of urine samples with ACN was also evaluated. The total ion chromatograms obtained with the reference 1
50 (v/v) dilution of urine samples with ACN was also evaluated. The total ion chromatograms obtained with the reference 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution method and the proposed TOT-µSPE microextraction approach are shown in Fig. 1B. The figure provides a qualitative comparison of the signals obtained with both approaches, highlighting the improved matrix clean-up achieved with the TOT-µSPE procedure.
50 dilution method and the proposed TOT-µSPE microextraction approach are shown in Fig. 1B. The figure provides a qualitative comparison of the signals obtained with both approaches, highlighting the improved matrix clean-up achieved with the TOT-µSPE procedure.
Creatinine levels were measured using the Jaffé method56 and were used to normalize the concentrations of mNSs.
2.4. Instrumentation and HILIC-MS/MS analysis
The LC-MS/MS system consisted of an Agilent 1200 series HPLC system coupled to a 6410 LC/MS triple quadrupole (QqQ) mass spectrometer with an electrospray (ESI) ionization source from Agilent Technologies (Waldbrom, Germany). The HPLC chromatograph was equipped with a binary pump, a membrane degasser, an autosampler and a six-port valve. The MS was operated in positive mode, and the source nebulizer pressure and voltage were set at 35 psi and +3500 V, respectively. Nitrogen was used as the drying (12 L min−1 and 350 °C) and collision gas. Analyte quantification was carried out in Multiple Reaction Monitoring (MRM) mode. All the parameters related to instrumental configuration are listed in Table S1 in the SI. An InfinityLab Poroshell 120 HILIC-Z column (2.1 × 150 mm; 2.7 µm) from Agilent was used. Mobile phase A consisted of 100 mM NH4Ac aqueous solution and mobile phase B was ACN. The flow rate was 0.200 mL min−1. The chromatographic gradient program was as follows: 0–2 min, 10–15% A; 2–8 min, 15–20% A; 8–10 min, 20–30% A; 10–20 min, 30% A; 20–25 min, 30–10% A; 25–30 min, 10% A. These conditions were maintained for an additional 3 minutes to re-equilibrate the column. Each sample was measured in triplicate with an injection volume of 10 µL.2.5. Method validation
Method validation was performed in accordance with the guidelines outlined in Strategies for the Quantification of Small Molecules Endogenously Present in Biological Samples,57 covering parameters such as linearity range, limits of detection (LOD) and quantification (LOQ), intra- and inter-day coefficients of variation, recovery and matrix effects. Validation was performed concurrently for both the proposed and reference methods.Calibration curves were constructed from both pure solvent and natural urine samples, each using six concentration levels (specific for each analyte) and a blank sample (unspiked), with each level analyzed in triplicate. The calibration equations are expressed as S = βx + b, where β is the calibration slope, x is the analyte concentration and S represents the peak area of the analyte. LOD is defined as the lowest concentration spiked into the sample that produces a signal significantly different from that of the corresponding endogenous concentration. LOQ was determined as 3.33 times the estimated LOD.58
Method precision was assessed using urine samples spiked at three concentration levels, representing the lower, middle, and upper range of the calibration curve. 10 replicates of these levels were analyzed within a single day to assess intra-day precision (repeatability), and 10 replicates per day were analyzed across three different days to determine inter-day precision (reproducibility). Precision is expressed as the coefficient of variation (CV%) and considered acceptable if CV values are ≤ 20%.
Recovery (%R) was calculated using eqn (1) for the reference method.
 | (1) |
For the TOT-µSPE method, recovery was evaluated using a spiking-based approach with standard mixtures under three experimental conditions: (A) natural urine spiked before µ-SPE, (B) natural urine sample spiked after µ-SPE (spiking into the extract), and (C) non-spiked natural urine. The analytical signals obtained under each condition are denoted as SA, SB, and SC, respectively. Recoveries (%R) were then calculated using eqn (2), which corrects for the black signal (SC) and normalizes the measured responses according to the spiking and dilution volumes involved in the preparation of each sample:
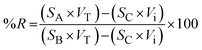 | (2) |
Matrix effects were subsequently evaluated by comparing the slopes of calibration curves prepared in spiked urine and in pure solvent, expressed as ion suppression or enhancement (0% for no effect and 100% for complete suppression).
3. Results and discussion
The analytical approach developed in this study is based on an LC-MS/MS system using a HILIC column, designed for the accurate quantification of polar analytes in biological matrices. A comprehensive evaluation of the method's suitability for routine quantitative analysis was performed, in which critical parameters influencing chromatographic separation, matrix effects, ionization efficiency, and overall sensitivity were systematically optimized. The main results obtained are outlined in the following sections.3.1. Tuning of mass spectrometry conditions
Multiple reaction monitoring (MRM) parameters were optimized by direct infusion of individual standard solutions corresponding to target analytes. All compounds were ionized in positive electrospray ionization mode (ESI+), with the predominant precursor ion corresponding to the [M + H]+ adduct. Key MS/MS parameters, such as fragmentor voltage and collision energy (CE), were individually optimized for each analyte to maximize the sensitivity and fragmentation efficiency. In the SI, Table S1 summarizes the selected precursor-to-product ion transitions and the optimized fragmentation conditions. The most abundant and stable product ions were selected for quantification, while additional transitions were used for confirmation purposes to ensure analyte identity.Additionally, ion source parameters were fine-tuned to further enhance the signal intensity. These included gas temperature (250–350 °C), nebulizer pressure (20–60 psi), drying gas flow (6–12 L min−1), and capillary voltage (2500–4000 V). Optimal values were established at 350 °C, 35 psi, 12 L min−1, and 3500 V, respectively.
3.2. Optimization of HILIC chromatographic separation
To optimize the stationary phase and considering the high polarity of the target analytes, different chromatographic columns were evaluated. These included non-polar reversed-phase columns (Kinetex C18), moderately polar columns (Luna PFP), and highly polar stationary phases such as XBridge Amide and HILIC-type columns. Among them, the zwitterionic hydrophilic interaction column (Z-HILIC), which incorporates a sulfobetaine-modified stationary phase, provided the best overall performance, yielding improved retention and enhanced chromatographic resolution. This type of zwitterionic phase carried both positively and negatively charged moieties within the same ligand, promoting a mixed-mode retention mechanism based on combined electrostatic and hydrophilic interactions. As a result, polar nucleosides exhibited stronger and more reproducible retention compared with amide-based HILIC columns, facilitating the separation of structurally related analytes. The advantages of sulfobetaine-modified zwitterionic phases for metabolite profiling and polar biomarker analysis have been highlighted in recent studies, which reported their high column efficiency, reproducibility and enhanced retention of polar analytes achieved with BEH-sulfobetaine columns, particularly in complex metabolomic applications.59–61Mobile phase composition was also investigated in both isocratic and gradient elution modes. Various mixtures of acetonitrile with aqueous solutions of ammonium acetate and formic acid were tested, with the aqueous phase ranging from 5% to 40% (v/v). Gradient elution using acetonitrile and 100 mM ammonium acetate led to significantly better separation and peak shapes. Fig. S2 shows the MRM chromatograms of 15 mNSs using a Z-HILIC column under optimized gradient conditions. Flow rates of 0.200, 0.300, and 0.400 mL min−1 were assessed. While no major differences were observed between 0.200 and 0.300 mL min−1 in terms of retention times or peak shapes, a noticeable increase in peak broadening and overlap was detected at 0.400 mL min−1. Since slightly higher signal intensities were obtained at 0.200 mL min−1 for most analytes, this value was selected as optimal. The signals for these three flow rates can be seen in Fig. S3.
The influence of the injection medium was also evaluated. Mixtures of analytes dissolved in acetonitrile and methanol were compared, with acetonitrile yielding narrower and better-resolved peaks. Due to the aqueous content in the mobile phase, different proportions of water in the injection medium (0–20%, v/v) were tested. The mixture of acetonitrile with 5% aqueous ammonium acetate (100 mM) (95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, v/v) provided the most favorable results in terms of peak shape, resolution and signal intensity. As shown in SI Fig. S4 and S5, this composition minimized peak broadening and baseline noise while enhancing analyte response. Therefore, this solvent mixture was selected as the optimal injection medium for all subsequent analyses.
5, v/v) provided the most favorable results in terms of peak shape, resolution and signal intensity. As shown in SI Fig. S4 and S5, this composition minimized peak broadening and baseline noise while enhancing analyte response. Therefore, this solvent mixture was selected as the optimal injection medium for all subsequent analyses.
Finally, injection volumes ranging from 2 to 50 µL were examined. While larger injection volumes resulted in higher signal intensities, they also led to increased peak broadening. A volume of 10 µL was selected as the compromise condition, providing sufficient sensitivity while maintaining acceptable peak shape and resolution. The influence of injection volume on signal intensity and peak shape is shown in Fig. S6 and S7. Fig. 2 shows the chromatograms obtained under optimized chromatographic conditions, illustrating the retention and separation achieved for all target analytes. It should be noted that multiple peaks are observed in some MRM chromatograms, which correspond to isobaric nucleosides/nucleobases sharing the same mass transitions (±1 u). This emphasizes the critical role of HILIC chromatographic separation in achieving accurate identification and quantification of structurally related methylated nucleosides.
3.3. Optimization of the tip-on-tip micro-solid phase extraction protocol
The tip-on-tip micro solid-phase-extraction approach (TOT-µSPE) was optimized to achieve efficient and reproducible extraction of methylated nucleosides and nucleobases from human urine while minimizing matrix interferences and solvent consumption. Unlike the conventional SPE, which typically requires multiple manual steps and larger sample and conditioning phases volumes, TOT-µSPE offers a microscale, low-volume alternative that is faster and more adaptable to high-throughput analysis according to sustainability requirements. This design represents the most innovative aspect of the study, providing a simplified and viable pretreatment strategy for urinary biomarker analysis.In the TOT-µSPE configuration, the sorbent material was confined within pipette tips, and extraction was performed by aspirating and dispensing small solvent volumes through the dispersed sorbent. Introducing a brief air gap during aspiration enhanced mixing and mass transfer, ensuring efficient analyte–sorbent contact and uniform wetting. The air-assisted mechanism accelerated extraction and improved reproducibility compared with conventional vacuum- or cartridge-based formats, explaining the superior enrichment and clean-up efficiency achieved.
Optimization experiments were carried out using a standard solution containing all fifteen target analytes, and recoveries were calculated relative to direct injections without extraction. Several parameters were systematically evaluated to identify the conditions that provided the highest recovery and reproducibility. Among the sorbents tested in the TOT configuration (Oasis MCX, Oasis HLB, Agilent PBA, and Isolute ENV+), ENV+ consistently exhibited the best performance and was therefore selected for further optimization. This superior performance was mainly reflected in higher recoveries and cleaner, more reproducible chromatographic profiles compared to other materials tested (Fig. S8). The effect of the amount of ENV+ packed in the tip was then investigated by varying the sorbent mass between 5 and 25 mg. Recoveries increased notably when the sorbent mass was increased from 5 to 15 mg; however, no further improvement was observed when using amounts above 15 mg, indicating that the available active surface was sufficient to quantitatively retain all analytes under the tested conditions. Additionally, the physical capacity of the pipette tip limited the maximum amount of sorbent that could be packed and the sample volume that could be efficiently processed. Taking into account these limitations, 15 mg of ENV+ was fixed as the optimal sorbent amount for subsequent experiments. Comparative chromatograms and recovery profiles obtained for different sorbents and sorbent amounts are provided in Fig. S8 and S9.
Next, the influence of the sample loading volume on the extraction performance was evaluated using 150, 300, and 500 µL aliquots of standard solution (Fig. S10). The highest recoveries, along with the most reproducible responses, were achieved with a loading volume of 150 µL. The use of larger sample volumes resulted in a slight decrease in extraction efficiency. This behavior can be attributed to partial analyte breakdown, as well as dilution effects within the packed tip, which could reduce effective interactions between the analytes and the sorbent. Based on these observations, 150 µL was established as the optimal sample volume. Afterwards, the effect of multiple loading cycles was examined by performing one, two, and three successive aspirations of 150 µL each through the same packed tip. The inclusion of additional loading cycles did not result in improved recoveries, confirming that a single aspiration step is sufficient to achieve exhaustive extraction. Furthermore, this approach offered the most efficient and straightforward workflow, minimizing the analysis time and operational complexity.
Different elution mixtures consisting of acetonitrile combined with formic acid or ammonia were evaluated to optimize analyte desorption. The solvent composition was investigated by varying the ratio of organic and aqueous solvent from 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 to 80
5 to 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 (v/v). The use of acetonitrile containing 10% ammonia (v/v) provided the highest recoveries for all analytes and yielded the most reproducible signal responses. Subsequently, the effect of elution volume was studied by testing total volumes between 100 and 500 µL (Fig. S11). Lower elution volumes resulted in incomplete desorption from the sorbent, an effect that was especially pronounced for more strongly retained compounds such as m1G and m7G. Increasing the elution volume above this range did not lead to further improvements in recovery but instead caused unnecessary dilution of the extracts, which is undesirable for sensitive detection. Based on these observations, an elution volume of 500 µL of acetonitrile
20 (v/v). The use of acetonitrile containing 10% ammonia (v/v) provided the highest recoveries for all analytes and yielded the most reproducible signal responses. Subsequently, the effect of elution volume was studied by testing total volumes between 100 and 500 µL (Fig. S11). Lower elution volumes resulted in incomplete desorption from the sorbent, an effect that was especially pronounced for more strongly retained compounds such as m1G and m7G. Increasing the elution volume above this range did not lead to further improvements in recovery but instead caused unnecessary dilution of the extracts, which is undesirable for sensitive detection. Based on these observations, an elution volume of 500 µL of acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ammonia (90
ammonia (90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, v/v) was selected as the optimal elution condition for subsequent analyses, as it ensured quantitative desorption while maintaining adequate analyte concentration and signal reproducibility.
10, v/v) was selected as the optimal elution condition for subsequent analyses, as it ensured quantitative desorption while maintaining adequate analyte concentration and signal reproducibility.
Finally, since eluates were evaporated to dryness prior to LC-MS/MS analysis, the reconstitution step was optimized by testing volumes between 25 and 150 µL using the same solvent composition as the initial chromatographic mobile phase (95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, v/v; acetonitrile
5, v/v; acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 mM ammonium acetate), which was also the solvent employed in the dilution-based method. This approach ensured full compatibility with the chromatographic conditions and minimized potential solvent-related effects on peak shape and ionization efficiency. As shown in Fig. S12, lower reconstitution volumes produced slightly higher signal intensities but poorer reproducibility. In contrast, larger reconstitution volumes led to a decrease in sensitivity as a consequence of analyte dilution. Overall, a reconstitution volume of 50 µL provided the best compromise between signal intensity, repeatability, and signal stability, and was therefore selected as the final reconstitution condition.
100 mM ammonium acetate), which was also the solvent employed in the dilution-based method. This approach ensured full compatibility with the chromatographic conditions and minimized potential solvent-related effects on peak shape and ionization efficiency. As shown in Fig. S12, lower reconstitution volumes produced slightly higher signal intensities but poorer reproducibility. In contrast, larger reconstitution volumes led to a decrease in sensitivity as a consequence of analyte dilution. Overall, a reconstitution volume of 50 µL provided the best compromise between signal intensity, repeatability, and signal stability, and was therefore selected as the final reconstitution condition.
3.4. Validation of the TOT-µSPE HILIC-MS/MS method
Linearity, limits of detection (LOD) and quantification (LOQ), intra- and inter-day precision, recovery, and matrix effects were evaluated for the proposed micro-scale TOT-µSPE protocol, and the results were compared with those obtained using a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution pretreatment strategy.
50 dilution pretreatment strategy.
Linearity was evaluated using standard solutions of analytes and spiked urine samples subjected to each of the following two strategies: 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution and TOT-µSPE. The results obtained for the urine samples are summarized in Table 1. Both methods exhibited excellent linearity, with correlation coefficients (R2) exceeding 0.990 for nearly all analytes. However, the 1
50 dilution and TOT-µSPE. The results obtained for the urine samples are summarized in Table 1. Both methods exhibited excellent linearity, with correlation coefficients (R2) exceeding 0.990 for nearly all analytes. However, the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution method showed a more pronounced matrix effect, characterized by significant signal suppression. This was evidenced by the notably lower slopes of the calibration curves prepared in urine compared to those prepared in pure solvent. This reduction in the slopes directly impacts the analytical sensitivity, which was found to be significantly higher when using the TOT-µSPE procedure due to the effective removal of interfering matrix components.
50 dilution method showed a more pronounced matrix effect, characterized by significant signal suppression. This was evidenced by the notably lower slopes of the calibration curves prepared in urine compared to those prepared in pure solvent. This reduction in the slopes directly impacts the analytical sensitivity, which was found to be significantly higher when using the TOT-µSPE procedure due to the effective removal of interfering matrix components.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution and TOT-µSPE sample pretreatment strategies in urine
50 dilution and TOT-µSPE sample pretreatment strategies in urine
| Analyte | Method | Calibration curve (µg L−1) | R 2 value | Linear range (mg L−1) | LOD (µg L−1) | LOQ (µg L−1) |
|---|---|---|---|---|---|---|
| m7Gs | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 21.711x + 1.1581 | 0.996 | 5–30 | 150 | 430 |
| TOT-µSPE | S = 0.6553x + 16.692 | 0.996 | 0.4–1.6 | 56 | 146 | |
| m1G | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 295.37x + 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 710 710 |
0.995 | 2.5–25 | 40 | 125 |
| TOT-µSPE |
S = 18.134x + 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 862 862 |
0.975 | 1.2–8 | 10 | 34 | |
| m1I | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 72.298x + 7064 | 0.992 | 5–30 | 600 | 1850 |
| TOT-µSPE |
S = 12.148x + 41![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 794 794 |
0.990 | 1–6 | 6 | 20 | |
| m6A | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 63.293x + 122.64 | 0.994 | 0.75–3 | 200 | 700 |
| TOT-µSPE | S = 20.138x + 304.13 | 0.993 | 0.1–0.4 | 6 | 20 | |
| m1A | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 2.5392x + 460.2 | 0.993 | 5–30 | 1250 | 3750 |
| TOT-µSPE | S = 0.117x + 703.46 | 0.992 | 1–6 | 44 | 146 | |
| m5U | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 33.782x − 35.805 | 0.995 | 0.5–3 | 150 | 500 |
| TOT-µSPE | S = 3.8594x − 55.467 | 0.993 | 0.1–0.4 | 8 | 26 | |
| m3C | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 66.255x + 3703 | 0.995 | 2.5–25 | 500 | 1650 |
| TOT-µSPE |
S = 2.4288x + 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 528 528 |
0.981 | 1–6 | 6 | 20 | |
| m5C | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 529.79x + 890.4 | 0.995 | 0.5–3 | 100 | 300 |
| TOT-µSPE | S = 30.707x + 242.33 | 0.996 | 0.1–0.4 | 3 | 10 | |
| m7G | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 287.81x + 27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 568 568 |
0.995 | 5–30 | 200 | 600 |
| TOT-µSPE |
S = 80.363x + 110![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 813 813 |
0.996 | 1–6 | 3 | 10 | |
| m6G | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
— | — | — | — | — |
| TOT-µSPE |
S = 18.326x + 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 372 372 |
0.993 | 0.3–1.2 | 6 | 20 | |
| m3A | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 822.85x − 1376 | 0.993 | 0.5–3 | 150 | 500 |
| TOT-µSPE | S = 256.39x + 1714 | 0.996 | 0.1–0.3 | 4 | 16 | |
| m5Uc | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 12.056x − 312.01 | 0.995 | 7.5–45 | 600 | 2000 |
| TOT-µSPE | S = 1.0403x + 438.59 | 0.993 | 1–6 | 20 | 66 | |
| m5Cs | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 28.619x − 11.095 | 0.996 | 5–30 | 40 | 125 |
| TOT-µSPE | S = 2.322x + 167.13 | 0.997 | 0.3–1.2 | 8 | 26 | |
| Cm | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 367.37x + 4125 | 0.997 | 1.5–9 | 500 | 1500 |
| TOT-µSPE |
S = 53.91x + 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 352 352 |
0.987 | 0.1–0.4 | 6 | 20 | |
| Gm | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution 50 dilution |
S = 305.78x + 1342 | 0.996 | 0.5–3 | 150 | 500 |
| TOT-µSPE | S = 36.502x + 3600 | 0.997 | 0.8–2.4 | 2 | 6 |
Sensitivity was also assessed for both methods. As can be seen in Table 1, the detection and quantification limits revealed that the TOT-µSPE method provided a substantial improvement in sensitivity for most analytes. Notably, the TOT-µSPE approach enabled significantly lower LODs for key biomarkers such as m6A, m5U, and m3A. In the case of m6G, this compound could not be detected at all using the dilution strategy, further highlighting the enhanced sensitivity of the TOT-µSPE method. This enhancement in analytical performance underscores the superior capability of the TOT-µSPE protocol to detect low-abundance compounds, an essential feature for the reliable application of these biomarkers in early-stage disease diagnostics and large-scale biomonitoring studies.
For precision and recovery studies, three different levels of each analyte were prepared as quality control (QC) samples. The concentration values of these samples for both pretreatment strategies are summarized in the SI.
Precision results are recapped in Table 2. Intra- and inter-day precision coefficients of variation (CVs) for the three concentration levels tested (low, medium, and high) were within acceptable limits (CV < 20%) for both methods. Specifically, the TOT-µSPE method exhibited greater variability for low-abundance analytes, particularly at low concentration levels. Despite this, the overall performance in terms of repeatability remained within acceptable thresholds for exploratory studies and validation procedures involving complex biological matrices.
| Reference method | TOT-µSPE method | |||||
|---|---|---|---|---|---|---|
| Low | Medium | High | Low | Medium | High | |
| (A) CV% intra-day | ||||||
| m7Gs | 7 | 7 | 6 | 19 | 9 | 3 |
| m1G | 7 | 7 | 4 | 5 | 12 | 11 |
| m1I | 7 | 7 | 5 | 14 | 10 | 17 |
| m6A | 5 | 5 | 6 | 12 | 13 | 14 |
| m1A | 7 | 7 | 5 | 15 | 12 | 10 |
| m5U | 9 | 9 | 10 | 14 | 18 | 20 |
| m3C | 8 | 8 | 5 | 18 | 8 | 14 |
| m5C | 12 | 12 | 7 | 13 | 9 | 20 |
| m7G | 9 | 9 | 5 | 18 | 13 | 15 |
| m6G | — | — | — | 5 | 17 | 11 |
| m3A | 9 | 9 | 5 | 11 | 13 | 20 |
| m5Uc | 6 | 6 | 5 | 14 | 17 | 19 |
| m5Cs | 8 | 8 | 5 | 18 | 11 | 16 |
| Cm | 4 | 4 | 3 | 15 | 12 | 9 |
| Gm | 4 | 4 | 7 | 16 | 14 | 18 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| (B) CV% inter-day | ||||||
| m7Gs | 10 | 13 | 11 | 20 | 18 | 3 |
| m1G | 8 | 9 | 9 | 9 | 12 | 17 |
| m1I | 10 | 10 | 9 | 19 | 13 | 18 |
| m6A | 9 | 6 | 9 | 13 | 16 | 14 |
| m1A | 9 | 10 | 11 | 15 | 18 | 10 |
| m5U | 14 | 9 | 12 | 17 | 19 | 20 |
| m3C | 8 | 10 | 9 | 16 | 13 | 14 |
| m5C | 8 | 12 | 12 | 18 | 11 | 12 |
| m7G | 9 | 10 | 9 | 17 | 15 | 15 |
| m6G | — | — | — | 20 | 16 | 10 |
| m3A | 11 | 10 | 9 | 13 | 13 | 19 |
| m5Uc | 12 | 10 | 7 | 19 | 19 | 18 |
| m5Cs | 13 | 11 | 9 | 18 | 17 | 16 |
| Cm | 6 | 8 | 9 | 16 | 12 | 11 |
| Gm | 7 | 8 | 12 | 15 | 15 | 18 |
The recovery rates for the QC samples at low, medium and high concentrations were evaluated, yielding values ranging from 85% to 120% as shown in Fig. 3. As previously stated, the m6G biomarker could only be detected and quantified using the TOT-µSPE method, likely due to the presence of multiple interfering substances in urine samples. In this regard, the matrix effect was evaluated by comparing the slopes of the calibration curves obtained by both pretreatment procedures in different matrices (pure solvent and urine). Fig. 4 illustrates the matrix effect (%ME), expressed as ion suppression, which ranges from 0% (no suppression) to 100% (complete suppression).62 The analysis revealed pronounced differences in matrix-related signal alteration across analytes and treatments. Overall, the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution method exhibited substantially higher matrix effects, with most analytes (13 out of 15) showing values exceeding 40%. This suggests an elevated level of co-eluting matrix components, which may enhance or suppress the ionization efficiency during mass spectrometric analysis. Notably, analytes such as m7Gs and m1I demonstrated significant signal enhancement, emphasizing the potential for compromised quantification accuracy when matrix effects are not adequately controlled. In contrast, TOT-µSPE-treated samples yielded consistently lower values across nearly all analytes. These results indicate a more effective removal of endogenous matrix constituents, leading to reduced ion suppression/enhancement. For example, compounds like m6A, m5C, and Cm showed markedly improved signal stability following TOT-µSPE, underscoring the robustness and cleaner background afforded by this approach.
50 dilution method exhibited substantially higher matrix effects, with most analytes (13 out of 15) showing values exceeding 40%. This suggests an elevated level of co-eluting matrix components, which may enhance or suppress the ionization efficiency during mass spectrometric analysis. Notably, analytes such as m7Gs and m1I demonstrated significant signal enhancement, emphasizing the potential for compromised quantification accuracy when matrix effects are not adequately controlled. In contrast, TOT-µSPE-treated samples yielded consistently lower values across nearly all analytes. These results indicate a more effective removal of endogenous matrix constituents, leading to reduced ion suppression/enhancement. For example, compounds like m6A, m5C, and Cm showed markedly improved signal stability following TOT-µSPE, underscoring the robustness and cleaner background afforded by this approach.
These findings highlight the superior matrix-cleanup efficiency of TOT-µSPE compared to direct dilution, particularly in complex biological matrices such as urine. While dilution protocols offer procedural simplicity and minimal sample handling, the elevated matrix effects observed in this study could compromise analytical reliability without the implementation of adequate calibration or internal standard correction. TOT-µSPE, therefore, represents a more analytically sound option for minimizing matrix-induced variability and improving data quality in quantitative LC-MS-based biomarker analysis.
Overall, the optimized TOT-µSPE protocol significantly improved analytical performance compared to the reference method, achieving high recoveries, minimizing matrix effects and substantially improving sensitivity for low-abundance analytes such as m6A, m5U, or m3A. These results demonstrate that careful optimization of the sorbent type, elution conditions and air aspiration step is key to obtaining reliable enrichment and accurate quantification of mNSs in complex biological matrices.
In summary, the optimized TOT-µSPE method provides a high-efficiency cleanup that is essential for the analysis of complex urine matrices. When compared to mainstream sample preparation techniques, this approach offers several practical advantages. Unlike the traditional cartridge-based SPE, which typically requires larger sample volumes and higher solvent consumption, TOT-µSPE operates at a micro-scale, reducing both environmental impact and cost per analysis. Furthermore, while online SPE systems provide a high degree of automation, they require costly specialized instrumentation and complex maintenance. The proposed tip-on-tip platform provides a well-balanced compromise between analytical performance and operational simplicity, making it a robust alternative for high-throughput routine clinical analysis. Additionally, the analytical figures of merit obtained with the TOT-µSPE platform, particularly sensitivity and recovery (85–120%), are highly competitive with those reported in previous studies using more laborious techniques like traditional SPE or liquid–liquid extraction (LLE), while significantly improving the results obtained by simple dilution.
3.5. Quantitative analysis of methylated nucleosides in urine
To evaluate the applicability of the developed method, the fifteen selected modified nucleosides and nucleobases were quantified in six human urine samples (U1–U6) using the TOT-µSPE sample preparation approach proposed in this study as well as by the reference method consisting of a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 sample dilution. The results are presented in Fig. 5.
50 sample dilution. The results are presented in Fig. 5.
 | ||
Fig. 5 Scatter plots illustrating the analysis of urine samples using the proposed TOT-µSPE pretreatment method in comparison to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution. 50 dilution. | ||
Compounds such as m1G, m1I, m1A, m3C, m7G, and Cm showed comparable concentrations between both methods, with differences under 10–15% in most cases. For example, for m1I in a representative urine sample, concentrations of 1610 ± 70 nM (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution) and 1690 ± 90 nM (TOT-µSPE) were obtained, indicating good reproducibility of the analyte with both procedures.
50 dilution) and 1690 ± 90 nM (TOT-µSPE) were obtained, indicating good reproducibility of the analyte with both procedures.
Slightly higher responses were observed for compounds such as m7Gs and Gm using the TOT-µSPE approach, which may reflect modest differences in extraction efficiency or reduced matrix suppression. For several low-abundance nucleosides, including m6A, m5U, m5C, m3A and m6G, detection was only possible after TOT-µSPE, confirming that this method primarily enhances detectability at concentration levels close to the instrumental detection limit. Representative chromatograms comparing both pretreatment strategies are displayed in Fig. 6.
 | ||
Fig. 6 Comparison of extracted ion chromatograms for the 15 targeted methylated nucleosides in urine samples prepared using 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 dilution (dashed line) and TOT-µSPE (solid line). 50 dilution (dashed line) and TOT-µSPE (solid line). | ||
Overall, TOT-µSPE proved to be a more sensitive and robust strategy for profiling a wide range of modified nucleosides in human urine. Meanwhile, direct dilution may still be suitable for the quantification of more abundant compounds or for rapid screening studies, due to its simplicity and speed of sample preparation.
4. Conclusions
In this study, we developed and validated a robust and sensitive HILIC-MS/MS method for the simultaneous quantification of fifteen methylated nucleosides and nucleobases in human urine. Sample pretreatment involved a tip-on-tip micro solid-phase-extraction (TOT-µSPE) for the effective extraction and preconcentration of methylated biomarkers. Its analytical performance in terms of sensitivity, precision, recovery, and matrix effects was compared to that of a direct 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 urine dilution. The validation process confirmed the reliability of both methods for routine analysis, with acceptable intra- and inter-day precision (CV < 20%), recovery rates between 85 and 120%, and excellent linearity across all analytes. Low-abundance compounds such as m6A, m5U, and m3A could only be detected by the TOT-µSPE method, enabling the detection of these analytes that remained undetectable or were significantly suppressed in diluted urine samples.
50 urine dilution. The validation process confirmed the reliability of both methods for routine analysis, with acceptable intra- and inter-day precision (CV < 20%), recovery rates between 85 and 120%, and excellent linearity across all analytes. Low-abundance compounds such as m6A, m5U, and m3A could only be detected by the TOT-µSPE method, enabling the detection of these analytes that remained undetectable or were significantly suppressed in diluted urine samples.
Moreover, TOT-µSPE exhibited significantly reduced matrix effects across nearly all compounds, offering greater reproducibility and analytical reliability in complex biological samples. These results strongly support the implementation of TOT-µSPE as the preferred sample preparation method when high sensitivity and selectivity are required, especially in biomarker studies, clinical validation studies, and population-scale biomonitoring programs.
Author contributions
M. Bustamante-Rangel: conceptualization, formal analysis, investigation, methodology, supervision, validation, writing – original draft, writing – review and editing. C. Mena-Iglesias: formal analysis, methodology, investigation, writing – original draft. A. Ballester-Caudet: formal analysis, investigation, methodology, supervision, validation, writing – review and editing. D. García-Gómez: conceptualization, formal analysis, investigation, methodology, supervision, writing – review and editing. E. Rodriguez-Gonzalo: conceptualization, supervision, validation, funding acquisition, project administration, resources, writing – review and editing. J. L. Pérez-Pavón: conceptualization, supervision, validation, funding acquisition, project administration, resources, writing – review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting the findings of this study are available within the article and its supplementary information. Additional data are available from the corresponding author upon reasonable request.Supplementary information (SI): Fig. S1–S12, Table S1 and additional details on analyte's characterization data, chromatographic optimization, tip-on-tip micro solid-phase extraction protocol, and quality control levels. See DOI: https://doi.org/10.1039/d6ay00083e.
Acknowledgements
This work was financially supported by the Spanish Ministry of Economy and Competitiveness (project No. PID2021-127679NB-100).References
- Z. D. Smith and A. Meissner, DNA methylation: roles in mammalian development, Nat. Rev. Genet., 2013, 14, 204–220 CrossRef CAS PubMed.
- W. Timp and A. P. Feinberg, Cancer as a dysregulated epigenome allowing cellular growth advantage at the expense of the host, Nat. Rev. Cancer, 2013, 13, 497–510 CrossRef CAS PubMed.
- X. Han, M. Wang, Y. L. Zhao, Y. Yang and Y. G. Yang, RNA methylations in human cancers, Semin. Cancer Biol., 2021, 75, 97–115 CrossRef CAS PubMed.
- L. He, H. Li, A. Wu, Y. Peng, G. Shu and G. Yin, Functions of N6-methyladenosine and its role in cancer, Mol. Cancer, 2019, 18, 176 Search PubMed.
- B. Thienpont, J. Steinbacher, H. Zhao, F. D'Anna, A. Kuchnio, A. Ploumakis, B. Ghesquière, L. Van Dyck, B. Boeckx, L. Schoonjans, E. Hermans, F. Amant, V. N. Kristensen, K. P. Koh, M. Mazzone, M. L. Coleman, T. Carell, P. Carmeliet and D. Lambrechts, Tumour hypoxia causes DNA hypermethylation by reducing TET activity, Nature, 2016, 537, 63–68 CrossRef CAS PubMed.
- R. Fang, L. Ye and H. Shi, Understanding the roles of N6-methyladenosine writers, readers and erasers in breast cancer, Neoplasia, 2021, 23, 551–560 CrossRef CAS PubMed.
- S. B. Murugaiyan, R. Ramasamy, M. Nakkeeran, V. Rangdhol, A. R. Srinivasan and G. Niranjan, Urinary 8-hydroxydeoxyguanosine as a marker of oxidative stress induced genetic toxicity in oral cancer patients, Indian J. Dent. Res., 2015, 26, 226–230 CrossRef PubMed.
- A. Seidel, P. Seidel, O. Manuwald and O. Herbarth, Modified nucleosides as biomarkers for early cancer diagnose in exposed populations, Environ. Toxicol., 2015, 30, 956–967 CrossRef CAS PubMed.
- M. Y. Chen, C. B. Qi, X. M. Tang, J. H. Ding, B. F. Yuan and Y. Q. Feng, Comprehensive profiling and evaluation of the alteration of RNA modifications in thyroid carcinoma by liquid chromatography-tandem mass spectrometry, Chin. Chem. Lett., 2022, 33, 3772–3776 CrossRef CAS.
- M. C. Lucas, L. P. Pryszcz, R. Medina, I. Milenkovic, N. Camacho, V. Marchand, Y. Motorin, L. Ribas de Pouplana and E. M. Novoa, Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing, Nat. Biotechnol., 2024, 42, 72–86 CrossRef CAS PubMed.
- T. Suzuki, The expanding world of tRNA modifications and their disease relevance, Cells, 2021, 22, 375–392 CAS.
- A. T. Godoy, M. N. Eberlin and A. V. C. Simionato, Targeted metabolomics: Liquid chromatography coupled to mass spectrometry method development and validation for the identification and quantitation of modified nucleosides as putative cancer biomarkers, Talanta, 2020, 210, 120640 CrossRef CAS PubMed.
- Z. Fang, Y. Hu, X. Hong, X. Zhang, T. Pan, C. Pan, S. Zheng and C. Guo, Simultaneous determination of methylated nucleosides by HILIC–MS/MS revealed their alterations in urine from breast cancer patients, Metabolites, 2022, 12, 973 CrossRef CAS PubMed.
- X. Zhang, Y. Hu, X. Hong, M. Wang, Z. Fang, X. Cao, K. Jiang and C. Guo, Determination of adenosine and its modifications in urine and plasma from breast cancer patients by hydrophilic interaction liquid chromatography-tandem mass spectrometry, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2022, 1209, 123428 CrossRef CAS PubMed.
- D. A. Mohl, S. Lagies, A. Lonzer, S. P. Pfäffle, P. Groß, M. Benka, M. Jäger, M. C. Huber, S. Günther, D. A. Plattner, I. Juhasz-Böss, C. Backhaus and B. Kammerer, On the quest for biomarkers: A comprehensive analysis of modified nucleosides in ovarian cancer cell lines, Cells, 2025, 14, 626 CrossRef CAS PubMed.
- J. Chen, L. Zhou, X. Zhang, X. Lu, R. Cao, C. Xu and G. Xu, Urinary hydrophilic and hydrophobic metabolic profiling based on liquid chromatography-mass spectrometry methods: Differential metabolite discovery specific to ovarian cancer, Electrophoresis, 2012, 33, 3361–3369 CrossRef CAS PubMed.
- C. Guo, Q. Chen, J. Chen, J. Yu, Y. Hu, S. Zhang and S. Zheng, 8-Hydroxyguanosine as a possible RNA oxidative modification marker in urine from colorectal cancer patients: Evaluation by ultra performance liquid chromatography-tandem mass spectrometry, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2020, 1136, 121931 CrossRef CAS PubMed.
- C. Yao, D. Lv, X. Zhou, P. Fu, W. Sun, J. Chen and H. Lin, Exploring urinary modified nucleosides as biomarkers for diabetic retinopathy: Development and validation of a ultra performance liquid chromatography-tandem mass spectrometry method, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2024, 1232, 123968 CrossRef CAS PubMed.
- W. Struck-Lewicka, R. Kaliszan and M. J. Markuszewski, Analysis of urinary nucleosides as potential cancer markers determined using LC-MS technique, J. Pharm. Biomed. Anal., 2014, 101, 50–57 CrossRef CAS PubMed.
- Q. Lan, P. Y. Liu, J. Haase, J. L. Bell, S. Huttelmaier and T. Liu, The critical role of RNA m6A methylation in cancer, Cancer Res., 2019, 79, 1285–1292 CrossRef CAS PubMed.
- M. Chen and C. M. Wong, The emerging roles of N6-methyladenosine (m6A) deregulation in liver carcinogenesis, Mol. Cancer, 2020, 19, 44 CrossRef CAS PubMed.
- T. Wang, S. Kong, M. Tao and S. Ju, The potential role of RNA N6-methyladenosine in cancer progression, Mol. Cancer, 2020, 19, 88 CrossRef CAS PubMed.
- Y. Zhao, Y. Shi, H. Shen and W. Xie, m6A-binding proteins: The emerging crucial performers in epigenetics, J. Hematol. Oncol., 2020, 13, 35 CrossRef CAS PubMed.
- Z. Liu, L. Gao, L. Cheng, G. Lv, B. Sun, G. Wang and Q. Tang, The roles of N6-methyladenosine and its target regulatory noncoding RNAs in tumors: Classification, mechanisms, and potential therapeutic implications, Exp. Mol. Med., 2023, 55, 487–501 CrossRef CAS PubMed.
- Z. Chen, Y. Guo, Z. Zhang, C. Li, L. Zhang, Y. Liu, G. Sun, F. Xiao, R. Feng and C. Zhang, mRNA m5C Alteration in azacitidine demethylation treatment of acute myeloid leukemia, Mol. Carcinog., 2025, 64, 502–512 CrossRef CAS PubMed.
- T. Hu, G. Wang, D. Wang, Y. Deng and W. Wang, m6A methylation modification: Potential pathways to suppress osteosarcoma metastasis, Int. Immunopharmacol., 2025, 145, 113759 CrossRef CAS PubMed.
- L. Wilkinson and T. Gathani, Understanding breast cancer as a global health concern, Sci. Rep., 2022, 95, 20211033 Search PubMed.
- M. S. H. Shaon, T. Karim, M. M. Ali, K. Ahmed, F. M. Bui, L. Chen and M. A. Moni, A robust deep learning approach for identification of RNA 5-methyluridine sites, Sci. Rep., 2024, 14, 25688 CrossRef CAS PubMed.
- K. Bartosik, E. Sochacka and G. Leszczynska, Post-synthetic conversion of 5-pivaloyloxymethyluridine present in a support-bound RNA oligomer into biologically relevant derivatives of 5-methyluridine, Org. Biomol. Chem., 2017, 15, 2097–2103 RSC.
- Y.-T. Wu, H.-L. Tan, G. Shui, C. Bauvy, Q. Huang, M. R. Wenk, C.-N. Ong, P. Codogno and H.-M. Shen, Dual role of 3-methyladenine in modulation of autophagy via different temporal patterns of inhibition on class I and III phosphoinositide 3-kinase, J. Biol. Chem., 2010, 285, 10850–10861 CrossRef CAS PubMed.
- M. Kosic, V. Paunovic, B. Ristic, A. Mircic, M. Bosnjak, D. Stevanovic, T. Kravic-Stevovic, V. Trajkovic and L. Harhaji-Trajkovic, 3-methyladenine prevents energy stress-induced necrotic death of melanoma cells through autophagy-independent mechanisms, J. Pharmacol. Sci., 2021, 147, 156–167 CrossRef CAS PubMed.
- F. Zhao, G. Feng, J. Zhu, Z. Su, R. Guo, J. Liu, H. Zhang and Y. Zhai, 3-methyladenine-enhanced susceptibility to sorafenib in hepatocellular carcinoma cells by inhibiting autophagy, Anti-Cancer Drugs, 2021, 32, 386–393 CrossRef CAS PubMed.
- E. Rodríguez-Gonzalo, L. Herrero-Herrero and D. García-Gómez, Development, validation and application of a fast analytical methodology for the simultaneous determination of DNA- and RNA-derived urinary nucleosides by liquid chromatography coupled to tandem mass spectrometry, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2016, 1019, 132–139 CrossRef PubMed.
- Z. Lu, Q. Wang, M. Wang, S. Fu, Q. Zhang, Z. Zhang, H. Zhao, Y. Liu, Z. Huang, Z. Xie, H. Yu and X. Gao, Using UHPLC Q-Trap/MS as a complementary technique to in-depth mine UPLC Q-TOF/MS data for identifying modified nucleosides in urine, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2017, 1051, 108–117 CrossRef CAS PubMed.
- Y. M. Liu, S. Wang, A. Dickenson, J. Mao, X. Bai and X. Liao, An on-line SPE-LC-MS/MS method for quantification of nucleobases and nucleosides present in biological fluids, Anal. Methods, 2024, 16, 2505–2512 RSC.
- Z. Wu, R. Zhou, B. Li, M. Cao, W. Wang and X. Li, Methylation modifications in tRNA and associated disorders: Current research and potential therapeutic targets, Cell Proliferation, 2024, 57, e13692 CrossRef CAS PubMed.
- X. Lin, Q. Zhang, Y. Qin, Q. Zhong, D. Lv, X. Wu, P. Fu and H. Lin, Potential misidentification of natural isomers and mass-analogs of modified nucleosides by liquid chromatography–triple quadrupole mass spectrometry, Genes, 2022, 13, 878 CrossRef CAS PubMed.
- C. Guo, C. Xie, Q. Chen, X. Cao, M. Guo, S. Zheng and Y. Wang, A novel malic acid-enhanced method for the analysis of 5-methyl-2′-deoxycytidine, 5-hydroxymethyl-2′-deoxycytidine, 5-methylcytidine and 5-hydroxymethylcytidine in human urine using hydrophilic interaction liquid chromatography-tandem mass spectrometry, Anal. Chim. Acta, 2018, 1034, 110–118 CrossRef CAS PubMed.
- H. P. Jiang, C. B. Qi, J. M. Chu, B. F. Yuan and Y. Q. Feng, Profiling of cis-diol-containing nucleosides and ribosylated metabolites by boronate-affinity organic-silica hybrid monolithic capillary liquid chromatography/mass spectrometry, Sci. Rep., 2015, 5, 7785 CrossRef CAS PubMed.
- L. Zhang, W. Zhang and H. Wang, Accurate quantification of ten mehtylated purine nucleosides by higly sensitive and stable isotope-diluted UHPLC-MS/MS, Anal. Chem., 2024, 96(28), 11366–11373 Search PubMed.
- R. Yin, J. Mo, M. Lu and H. Wang, Detection of human urinary 5-hydroxymethylcytosine by stable isotope dilution HPLC-MS/MS analysis, Anal. Chem., 2014, 87(3), 1846–1852 Search PubMed.
- J. F. Xiao, B. Zhou and H. W. Ressom, Metabolite identification and quantitation in LC-MS/MS-based metabolomics, TrAC, Trends Anal. Chem., 2012, 32, 1–14 CrossRef CAS PubMed.
- A. Kaufmann, High-resolution mass spectrometry for bioanalytical applications: Is this the new gold standard?, J. Mass Spectrom., 2020, 55, e4533 CrossRef CAS PubMed.
- A. Zhang, H. Sun, P. Wang, Y. Han and X. Wang, Modern analytical techniques in metabolomics analysis, Analyst, 2012, 137, 293–300 RSC.
- Y. Sakaguchi, K. Miyauchi, B. I. Kang and T. Suzuki, Nucleoside analysis by hydrophilic interaction liquid chromatography coupled with mass spectrometry, Methods Enzymol., 2015, 560, 19–28 CAS.
- Y. Tang, S. J. Zheng, C. B. Qi, Y. Q. Feng and B. F. Yuan, Sensitive and simultaneous determination of 5-methylcytosine and its oxidation products in genomic DNA by chemical derivatization coupled with liquid chromatography-tandem mass spectrometry analysis, Anal. Chem., 2015, 87, 3445–3452 CrossRef CAS PubMed.
- D. Moravcová, M. Haapala, J. Planeta, T. Hyötyläinen, R. Kostiainen and S. K. Wiedmer, Separation of nucleobases, nucleosides, and nucleotides using two zwitterionic silica-based monolithic capillary columns coupled with tandem mass spectrometry, J. Chromatogr. A, 2014, 1373, 90–96 CrossRef PubMed.
- R. Thakare, Y. S. Chhonker, N. Gautam, J. A. Alamoudi and Y. Alnouti, Quantitative analysis of endogenous compounds, J. Pharm. Biomed. Anal., 2016, 128, 426–437 CrossRef CAS PubMed.
- R. Jadda, S. Madhumanchi and R. Suedee, Novel adsorptive materials by adenosine 5′-triphosphate imprinted-polymer over the surface of polystyrene nanospheres for selective separation of adenosine 5′-triphosphate biomarker from urine, J. Sep. Sci., 2019, 42, 3662–3678 CrossRef CAS PubMed.
- R. Tuytten, F. Lemière, W. Van Dongen, E. Witters, E. L. Esmans, R. P. Newton and E. Dudley, Development of an on-line SPE-LC - ESI-MS method for urinary nucleosides: Hyphenation of aprotic boronic acid chromatography with hydrophilic interaction LC - ESI-MS, Anal. Chem., 2008, 80, 1263–1271 Search PubMed.
- S. Dugheri, G. Marrubini, N. Mucci, G. Cappelli, A. Bonari, I. Pompilio, L. Trevisani and G. Arcangeli, A review of micro-solid-phase extraction techniques and devices applied in sample pretreatment coupled with chromatographic analysis, Acta Chromatogr., 2021, 33, 99–111 Search PubMed.
- H. Vlčková, V. Pilařová, O. Novák, P. Solich and L. Nováková, Micro-SPE in pipette tips as a tool for analysis of small-molecule drugs in serum, Bioanalysis, 2017, 9, 887–901 Search PubMed.
- J. Rappsilber, M. Mann and Y. Ishihama, Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips, Nat. Protoc., 2007, 2, 1896–1906 CrossRef CAS PubMed.
- J. C. Kostas, M. Greguš, J. Schejbal, S. Ray and A. R. Ivanov, Simple and efficient microsolid-phase extraction tip-based sample preparation workflow to enable sensitive proteomic profiling of limited samples (200 to 10,000 Cells), J. Proteome Res., 2021, 20, 1676–1688 CrossRef CAS PubMed.
- A. M. Casas-Ferreira, M. del Nogal-Sánchez, Á. E. Arroyo, J. V. Vázquez and J. L. Pérez-Pavón, Fast methods based on mass spectrometry for peptide identification. Application to sex determination of human remains in tooth enamel, Microchem. J., 2022, 181, 107645 CrossRef CAS.
- M. Jaffe, Ueber den Niederschlag, welchen Pikrinsäure in normalem Harn erzeugt und Über eine neue Reaction des Kreatinins, Z. Physiol. Chem., 1886, 10, 391–400 Search PubMed.
- M. Nelis, P. Augustijns and D. Cabooter, Strategies for the quantification of endogenously present small molecules in biological samples, LCGC Europe, 2019, 32, 356–365 Search PubMed.
- A. Kruve, R. Rebane, K. Kipper, M. L. Oldekop, H. Evard, K. Herodes, P. Ravio and I. Leito, Tutorial review on validation of liquid chromatography-mass spectrometry methods: part I, Anal. Chim. Acta, 2015, 870, 29–44 CrossRef CAS PubMed.
- A. Lioupi, C. Virgiliou, T. H. Walter, K. M. Smith, P. Rainville, I. D. Wilson, G. Theodoridis and H. G. Gika, Application of a hybrid zwitterionic hydrophilic interaction liquid chromatography column in metabolic profiling studies, J. Chromatogr. A, 2022, 1672, 463013 CrossRef CAS PubMed.
- T. H. Walter, B. A. Alden, K. Berthelette, J. A. Field, N. L. Lawrence, J. McLaughlin and A. V. Patel, Characterization of a highly stable zwitterionic hydrophilic interaction chromatography stationary phase based on hybrid organic–inorganic particles, J. Sep. Sci., 2022, 45, 1389–1399 Search PubMed.
- F. Gritti, G. Izzo and R. Schaffer, Understanding retention and intra-particle diffusivity of alkylsulfobetaine-bonded ethylene bridged particles with different mesopore sizes for hydrophilic interaction liquid chromatography applications, J. Chromatogr. A, 2024, 1733, 465232 CrossRef CAS PubMed.
- D. L. Buhrman, P. I. Price and P. J. Rudewicz, Quantitation of SR 27417 in human plasma using electrospray liquid chromatography-tandem mass spectrometry: A study of ion suppression, J. Am. Soc. Mass Spectrom., 1996, 7, 1099–1105 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |




