 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Beyond identification: inferring physical activity from fingerprint lipids using machine learning
Daphne R. Patten ,
Raven L. Buckman Johnson
,
Raven L. Buckman Johnson ,
Trevor T. Forsman†
and
Young Jin Lee
,
Trevor T. Forsman†
and
Young Jin Lee *
*
Department of Chemistry, Iowa State University, Ames, Iowa 50011, USA. E-mail: yjlee@iastate.edu
First published on 30th March 2026
Abstract
Fingerprints are a widely recognized form of forensic evidence, valued for their ability to link individuals with specific locations. Traditional fingerprint analysis relies on optical imaging to identify a match in a fingerprint database; however, where no match is found, the evidential value of a latent print is limited. Here, we present the first study to integrate matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS) with supervised machine learning to infer physical activity from fingerprint chemistry, expanding the utility of fingerprints beyond identification alone. Physical activity labels were derived from a validated questionnaire and converted into binary classes. Supervised machine learning algorithms were trained on the lipid features and evaluated against the survey-derived labels. The top-performing models were an ensemble algorithm based on multiple decision trees and a neural network, which classified physical activity with accuracies of 75 ± 8% and 73 ± 7%, respectively. These results demonstrate that fingerprint lipid chemistry encodes biologically meaningful information related to physical activity and establish a new approach for extracting lifestyle and behavioral indicators from trace evidence, with potential applications in forensic investigations and noninvasive fingerprint-based assessments in medicine.
Introduction
Fingerprints have long served as a cornerstone of forensic identification; however, their evidential value is constrained when ridge patterns cannot be matched to a database.1,2 As a result, unmatched fingerprints often serve only as confirmatory evidence once suspects are identified by other means. This limitation has motivated efforts to extract additional information from the chemical composition of latent prints, rather than treating them solely as physical impressions.Mass spectrometry (MS), one of the most discriminating analytical techniques in forensic science, has played a central role in this shift.3 In particular, matrix-assisted laser desorption/ionization MS (MALDI-MS) enables high-throughput, extraction-free analysis by rastering a laser across the fingerprint surface to generate desorption/ionization events that ionize the chemical compounds present.4–8 As a soft-ionization technique, MALDI-MS can detect a broad range of endogenous and exogenous chemicals in fingerprints, resulting in a complex mixture that undergoes physical and chemical changes after deposition, complicating data interpretations; thus, many have turned to advanced computational strategies to aid in data analysis.2,9
As fingerprint chemistry has come into focus, studies have increasingly examined endogenous compounds to infer subtle biological variations. Machine learning (ML) is well-suited to this task because it identifies multivariate patterns in complex datasets and builds predictive models from empirical data.10,11 A key subset of ML is supervised ML, in which models are trained on labeled data to improve the prediction accuracy.12 Several studies have demonstrated the potential of supervised ML to extract individual characteristics from fingerprint chemistry. For example, Ferguson et al. analyzed peptides and proteins in fingerprints from 80 people, using MALDI-MS and partial least squares discriminant analysis to determine sex with an accuracy of 85%.13 This work was later expanded to include 199 participants while still achieving 86% accuracy using a supervised ensemble model.14 Using the same fingerprint dataset, a parallel study by Bury et al. investigated whether a person's age could be distinguished; however, an ensemble model achieved only 66% accuracy on the same sample set.15 While insightful, peptide/protein analytes represent a small fraction of fingerprint residue.9,16,17 In contrast, lipids are more abundant and generally more stable in fingerprints, offering an analytically and biologically robust substrate for ML-based chemical profiling.9,16,17
One forensic characteristic that remains underexplored is physical activity (PA), which may offer insights into individuals' occupations or lifestyles. Numerous studies on lipid metabolism have demonstrated that even individuals who meet only the minimal recommended activity thresholds exhibit measurable alterations in serum lipid profiles. Therefore, we hypothesize that comparable lipid alterations may also be detectable in fingerprint lipids.18 Although criminal profilers may attempt to infer occupation type from crime scene evidence, such assessments are inherently subjective and vary in reliability.19,20 Empirically derived indicators of PA could strengthen behavioral profiling and assist investigations, particularly when other leads are limited.
Preliminary evidence by O'Neill et al. suggests that fingerprint lipids, particularly triacylglycerols (TGs), may reflect PA levels.21 However, their conclusions were based on a small cohort (n = 8) and a simple Boolean survey, both of which are susceptible to response bias. The present study expands upon that work by incorporating a larger participant group (n = 81) and using a validated PA assessment developed by Besson et al.22 This survey captures detailed information on the duration and frequency of participants' commuting, occupational activities, and other behaviors across 38 activity categories. Quantitative responses reduce the subjectivity common in qualitative surveys, enabling more reliable PA estimates without the need for invasive physiological measurements.22
In addition, this study examines wax esters (WEs), diacylglycerols (DGs), and TGs – the three major lipid classes present in sebaceous secretions and latent fingerprints – to evaluate their potential as biomarkers of physical activity. TGs and DGs play central roles in energy storage and metabolic regulation, with TGs being among the most extensively studied lipid classes in serum. In contrast, WEs are predominantly found in the epidermis; however, their biosynthesis depends on fatty acid availability, thereby indirectly linking all three lipid classes to lipid metabolism flux.23,24 Building on this foundation, the present study investigates PA as a forensic variable using a larger dataset, validated activity metrics, and chemically relevant lipid features. Supervised ML models are applied to MALDI-MS fingerprint data to classify lipid profiles into a binary PA category.
Experimental
Physical activity questionnaire
Participant PA status (i.e., active or inactive) was determined using a PA survey published by Besson et al.22 In their validation study, the survey-derived equations and corresponding PA metrics showed significant associations with empirical measurements obtained via the double-labeled water method and activity sensors, supporting the reliability of the approach for estimating PA. To tailor the survey to this study, additional questions were incorporated at the beginning to collect information on participants' sex, ethnicity, medication use, and dietary habits. These additions are detailed in the SI.Physical activity energy expenditure for a given activity, i, (PAEEi) was calculated using eqn (1) developed by Besson et al.:
 | (1) |
In this equation, DRA refers to the duration (in hours per day) of the activity, and MET is the metabolic equivalent of the task or activity. MET values were obtained from a previously published compendium of MET intensities, with 1 MET defined as energy consumption with no physical activity (e.g., sleeping) equivalent to 3.5 mL of oxygen consumption per kg (body weight) per min.25,26 Weight refers to the individual's self-reported body weight in kilograms. The conversion factor 4.263 represents the energy yield of aerobic respiration in kilojoules and is based on the assumption that one liter of oxygen consumption produces 20.3 kJ of energy.22 RMROxford denotes the individual's resting metabolic rate, calculated by using the Oxford equations published by Henry,27 also adopted by Besson et al., and is adjusted by the daily duration ratio (DRAi/24). MET values for each activity and an example of PAEE calculation can be found in the supplementary spreadsheet labeled Tables S1 and S2, respectively.
To estimate an individual's overall PA, eqn (2) was used to calculate the total physical activity energy expenditure per body weight (PAEEw):
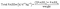 | (2) |
The sum of all PAEE values for each reported activity is combined with the PAEE value for unaccounted time (PAEEu), which has a MET value of 1.2. An MET value of 1.2 corresponds to the energy expenditure of being awake but mostly sedentary, such as during screen time or other low-intensity activities.22 Including the PAEEu component helps mitigate the underestimation of PAEE due to reporting bias related to time spent on electronic devices. The resulting total PAEE is then normalized by the individual's self-reported body weight (in kilograms).22
This work includes some deviations from the methodology used by Besson et al. Most notably, while Besson et al. assumed a fixed sleep duration of eight hours per night for all participants, the questionnaire used in the present study was modified to record each individual's self-reported sleep duration. Removing this assumption allows for a more accurate estimation of each subject's PAEEu. Additionally, Besson et al. applied PAEEu retroactively only to individuals who reported bicycling as a mode of transportation, assuming that such individuals engage in higher overall PA. In contrast, given the high number of college students in this study's participant pool, PAEEu was applied more broadly to account for low-intensity activities such as walking between classes and extended smartphone use.
Fig. 1 depicts the spread of PAEEw scores from the 81 participants used in this study. The box represents the interquartile range (IQR) with the median at 42 kJ (d−1 kg−1), and the 1st and 3rd quartiles at 32 and 62 kJ (d−1 kg−1), respectively. Whiskers extend to 1.5 times the IQR, ranging from 20 to 108 kJ (d−1 kg−1), which suggests a wide range of PAEEw values among the participants. Some outliers beyond the whiskers correspond to participants with unusually high levels of physical activity. The median value of 42 kJ (d−1 kg−1) matched the approximate PAEEw of an average adult with a sedentary occupation and thirty minutes of light walking.28–30 Therefore, 42 kJ (d−1 kg−1) was defined as the threshold for binary PA classification; those with a PAEEw < 42 kJ (d−1 kg−1) were labeled as inactive (n = 39), and those with PAEEw > 42 kJ (d−1 kg−1) were labeled as active (n = 42).
Sample collection and preparation
Recruitment and collection of fingerprints were completed in accordance with the Iowa State University Internal Review Board regulations that include obtaining informed consent. Volunteers, recruited through email solicitation and on-site collections at State Gym (Iowa State University, Ames, IA), were asked to complete a questionnaire as described above. Groomed fingerprints were acquired by having the donor swipe their finger across their forehead prior to deposition, ensuring sufficient sebaceous lipid collection for research-level MALDI-MS analysis, and were deposited onto precleaned glass microscope slides. Donors were asked to wash their hands before deposition, but this was not always possible during on-site collections. Fingerprints were stored in a desiccator for 1 to 2 days until analysis. Then, fingerprint samples were sprayed with 10 mM sodium acetate using a TM Sprayer (HTX Technologies LLC, Chapel Hill, NC, USA) to promote sodium adduct formation.31,32 A thin layer of gold was sputtered onto the surface (Cressington 108; Ted Pella, Redding, CA, USA) to ensure conductivity and serve as an inorganic matrix for MALDI-MS analysis (or Au-assisted-LDI-MS).21,31,33MALDI-MS acquisition
Mass spectrometry analysis was conducted using a Q-Exactive HF Orbitrap MS (Thermo Scientific, San Jose, CA, USA) equipped with a medium pressure (∼7 torr) matrix-assisted laser desorption/ionization (MALDI) source (Spectroglyph, Kennewick, WA, USA). Fingerprint samples were analyzed in positive ion mode with a mass resolution of 120![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 (at m/z 200) for a mass range of m/z 380–1100. An imaging raster step of 75–100 µm was used, and approximately 1000 spectra were collected and averaged. In accordance with the Iowa State University Internal Review Board guidelines, only a small portion of the fingerprints were analyzed to limit the total amount of identifiable information obtained by the instrument that can be imaged using MALDI-MS imaging.31
000 (at m/z 200) for a mass range of m/z 380–1100. An imaging raster step of 75–100 µm was used, and approximately 1000 spectra were collected and averaged. In accordance with the Iowa State University Internal Review Board guidelines, only a small portion of the fingerprints were analyzed to limit the total amount of identifiable information obtained by the instrument that can be imaged using MALDI-MS imaging.31
Data processing
A custom Python script was created to search for the theoretical masses of the lipids within a 3 ppm window and normalize the signal intensities to the summed saturated TGs.34 This included TGs with 46–52 carbons on the acyl chains and 0–3 levels of unsaturation, DGs with 22–43 carbons on the acyl chains and 0–3 levels of unsaturation, and WEs with acyl chains with 25–46 carbons and 0–3 levels of unsaturation were extracted. All lipid signal intensities were normalized to the sum of saturated TG signals to reduce variability due to deposition differences.21,31 Grooming increases the total amount of sebaceous material but is not expected to change the relative lipid composition, only total lipid loading.Mass spectra were internally calibrated to ensure a mass error of less than 3 ppm, allowing for the confident use of the Python script. Mass calibration was performed using two endogenous, abundant fingerprint lipids, TG 48:2 and WE 36:2, in a two-point calibration, following prior lipid assignments in a high-resolution mass spectrometry study.34 In total, 35 TGs, 88 DGs, and 88 WEs were used, alongside the biological sex and age of the participants, for a total of 213 possible features for ML. To ensure maximum data quality, a signal-to-noise ratio cutoff of three (S/N > 3) was implemented. A base-10 logarithmic transformation was then applied to lipid ratios. In preliminary ML, the log transformation yielded superior cross-validated performance for PA classification compared with alternative transformations in the CL app (data not shown). This aligns with prior work showing that log transformation is an effective pretreatment for MS-based metabolomics because it stabilizes variance, mitigates heteroscedasticity, and reduces the dominance of very high-intensity features—benefits that can translate into improved downstream statistical and ML performance.35,36 Averages were calculated from replicate fingerprints of the same donors. Due to the limited number of individuals with restricted diets, samples from individuals who reported restricted diets were removed due to the potential to interfere with lipids in fingerprints.21 Lastly, fingerprints from ten individuals were removed from the study due to the low quality of mass spectra attributed to high exogenous contamination and/or low lipid signals across all lipid analytes, bringing the final number of fingerprints from 106 to 81 participants.
Machine learning
All machine learning analysis was performed using MATLAB (ver. 2024b) with the Statistics and Machine Learning Toolbox™ (ver. 24.2). A workflow illustrating the analysis process is provided in Fig. 2. After data acquisition and preprocessing, the Classification Learner App (CL app) was used to explore learning algorithms best suited for fingerprint data; these models were trained on 100% of the processed data. Ten classification algorithms were available for testing in the CL app: decision tree (DT), discriminate analysis (DA), logistic regression (LR), naïve Bayes (NB), support vector machines (SVM), efficiently trained linear classifiers (EL), k-nearest neighbor (kNN), kernel approximation, ensemble (EN), and neural network (NN). The four model types with the best performance (accuracy ≥ 0.6) were selected for further testing/optimization, excluding the DT. The DT will be discussed more in the Results and Discussion section. Subsequent ML analysis was conducted using an in-house written MATLAB script; an example of the code is available on GitHub (https://github.com/drpatten/Fingerprint-Physical-Activity-ML.git). Hyperparameters from the CL app were transferred and further optimized to achieve the best accuracy. Processed data was randomized and split (70/30) for training and testing. Training data were assigned classification labels as active or inactive with the PAEEw score thresholds of 42 kJ (d−1 kg−1), as established in the Physical Activity Questionnaire section. This random selection, splitting, training, and testing portion of the ML analysis was conducted ten times to minimize bias and form informative ML metrics. All metrics presented here are the averages of the ten-replicate analysis with the associated standard deviation.Results and discussion
Fingerprints are complex chemical mixtures containing both natural endogenous compounds and exogenous contaminations. Endogenous chemicals are naturally secreted by the body and include amino acids, proteins, lipids, and other organic and inorganic compounds.1 Exogenous contaminations arise from routine contact with materials such as oils, insect repellent, cosmetics, and the use of drugs and alcohol.37,38 Among these, compounds that readily form positive ions can interfere with the detection of endogenous lipid analytes; this issue is especially pronounced when cosmetics contain plant-derived lipids. Such chemical complexity poses challenges in distinguishing endogenous from exogenous analytes. An example MALDI mass spectrum of a fingerprint is shown in Fig. 3.31,32,39 | ||
| Fig. 3 Example of a Fingerprint MALDI-Mass Spectrum. SQ: squalene, WE: wax ester, DG: diacylglycerol, TG: triacylglycerol. | ||
Exercise is widely recognized as essential to human health, and many studies have examined its physiological effects, including exercise-induced fluctuations in lipid metabolism. Given the known influence of PA on metabolism and lipid regulation, several lipid classes were selected as candidate biomarkers of PA in fingerprints: TGs, DGs, and WEs. TG levels in serum have been shown to vary with individual's exercise habits.21,40,41 DGs, produced during the hydrolysis of stored TGs, may similarly serve as indicators of PA.23 WEs – synthesized exclusively on human skin and functioning as protective hydrophobic barriers – were also evaluated based on the hypothesis that their secretion levels may shift with exercise.24,42
Regular exercise has additionally been shown to modulate androgen levels, altering the production of various metabolites.43 These androgen fluctuations can influence sebum lipid composition, although the underlying mechanisms remain incompletely understood.44,45 Other endogenous lipids of interest include fatty acids detected in the low-mass region (below m/z 400). This region, however, is frequently contaminated by exogenous compounds such as fingerprint enhancement powders, limiting its reliability.7 The squalene (SQ) base peak at m/z 433.3805 was not used in this study due to its high susceptibility to oxidation, both on the skin and after deposition, which limits its analytical utility.39,46,47
Binary classification of physical activity
The second step of this study is a preliminary ML analysis using all the participants (Fig. 2, bottom left). ML algorithms differ in their characteristics, strengths, and limitations, yet many published studies rely on a single algorithm without justification. To objectively identify suitable algorithms for classifying PA from fingerprint data, the CL app was used to compare the performance of all available supervised classification algorithms. Standardized performance metrics were applied to ensure fair comparison. Specifically, accuracy and the receiver operating characteristic area under the curve (ROC AUC) were evaluated to assess model performance. Accuracy represents the proportion of correct predictions, calculated as (true positive + true negative)/(true positive + true negative + false positive + false negative).48 ROC AUC quantifies a model's ability to distinguish different classes based on the relationship between true positive rate and false positive rate.49 Together, these metrics provide complementary insights: accuracy reflects overall classification success, and ROC AUC assesses class-separation robustness. Values closer to 1 indicate stronger model performance.Previous work by O'Neill et al. demonstrated statistically significant differences in TG levels between active and inactive men.21 Based on this, three feature lists were generated: TG only (37 features, including sex and age); TGs and WEs (125 features); and TGs, DGs, and WEs (213 features). Table S4 summarizes the performance of ten ML models in the CL app using these feature lists with a 10-fold cross-validation. All accuracies fell below 0.7 – well under acceptable scientific standards and unsuitable for forensic casework. One likely explanation for the poor performance is overfitting or modeling degradation due to irrelevant or noisy features. To improve model performance and exclude noninformative variables, Shapley feature selection was applied to retain only the top 15 features regardless of the lipid list used for training.50,51 Four algorithms achieving accuracy higher than 0.6 in the CL app were used for further optimization: SVM, kNN, EN, and NN. A limit of 15 features ensured a favorable feature-to-sample ratio, consistent with practice in high-dimensional fields such as genomics and metabolomics, which commonly face small sample sizes and the curse of dimensionality.52 In cases where fewer than 15 features had nonzero mean absolute Shapley values, only those features were retained.
Once optimized, the hyperparameters and selected features for each algorithm were incorporated into MATLAB for independent testing (Fig. 2, bottom right). Each model was retrained using a 70/30 train-test split to evaluate performance on unseen data, stimulating real-world forensic deployment. Fig. 4 presents boxplots comparing the four ML algorithms across the three feature lists with Shapley-reduced features. The best performance was achieved by the EN model trained on all three lipid classes (TGs, WEs, and DGs) plus participants' age, yielding an average accuracy of 75 ± 8% and ROC AUC of 0.84 ± 0.08, indicating strong discrimination between active and inactive groups. To our knowledge, this is the first study to classify PA from fingerprint lipid chemistry using supervised ML, limiting direct comparisons. Nevertheless, the magnitude of our metrics is consistent with prior MALDI-MS studies on different endpoints, such as Bury et al. at 66% for age predictions15 and Heaton et al. at 85% for predicting sex,14 both using peptide/protein analytes. Hence, this ML performance suggests the presence of biologically significant trends between fingerprint lipids and PA. This study has several limitations, as further discussed in the Limitations section, but there is a fundamental limitation in the binary reduction of a continuous phenotype such as PA. Heaton et al. noted similar limitations when reducing age from continuous to discrete age categories in their study.14 More discussion of the limitations can be found in the Limitations section. Table S6 lists the statistical values corresponding to the same models shown in Fig. 4.
The EN model's strong performance is partially attributed to the inclusion of participant age as a predictor, as previous studies have shown that biological age influences lipid production.2,16 Age may therefore correlate with the lipid profiles studied, although elucidating this biological relationship is beyond the scope of the present work. Despite its analytical usefulness, age is not ideal in forensic contexts because biological age cannot yet reliably inferred from fingerprints, as demonstrated by Bury et al.53 Accordingly, a second EN model was created that excluded age. As shown in Fig. S3, removal of age reduced the EN accuracy to 67 ± 8%, falling below the 0.7 threshold.
Among the remaining algorithms, the NN trained on TGs + WEs performed best without age, achieving an average accuracy of 73 ± 7% and ROC AUC of 0.79 ± 0.06 – comparable to EN with age. Thus, the NN algorithm currently represents the more appropriate option for forensic applications aimed at determining PA. Future improvements in biological age estimation from fingerprints may enhance EN performance and broaden its suitability. Understanding the behavior of each ML algorithm provides additional context for their performance. The EN algorithm achieved the highest metrics. The optimized EN model used an adaptive boosting approach, in which multiple decision trees (DTs) are sequentially built, each emphasizing corrections to the misclassification of the previous tree.54 This iterative learning strategy allows EN models to handle complex multivariate data such as fingerprint lipids. Although DTs performed similarly in the CL app, they were excluded due to their susceptibility to overfitting and reduced generalizability. ENs mitigate these limitations by aggregating multiple weak learners.12,54,55
The NN algorithm produced the second-highest metrics and was the best-performing model when age was excluded. NNs are well-suited to high-dimensional datasets because they can capture nonlinear relationships and interdependencies among features. However, they generally require larger datasets to avoid overfitting and demand substantial computational resources.12,55 Expanding the participant cohort would likely improve NN performance.
SVMs also handle redundant or interdependent features effectively, relying on finding the optimal separating hyperplanes to classify the data.12,55,56 However, datasets with only subtle biological variation, such as those used presented here, often result in poorly defined class boundaries, reducing SVM accuracy. Because all participants belong to the same species and exhibit overlapping PA levels the SVM likely could not establish adequate create margins.
Finally, kNN is an intuitive and transparent algorithm that classifies new data based on the most common class among the nearby training data.11,12,55 Its reliance on the full feature set makes it highly sensitive to irrelevant or noise variables, and interdependent features can further degrade performance, consistent with the results observed in this study.
Fig. 5 represents the confusion matrices for the best-performing EN and NN models. Correct predictions are shown in blue and misclassifications in orange. Both models show similar classification patterns. In forensic contexts, the false positive rate – false positives divided by (false positives + true negatives) – is of particular concern.48 Misclassifying inactive individuals as active could misdirect investigations and waste valuable time and resources. The EN model exhibited false positive rate is 22 ± 14% for the active class and 27 ± 17% for the inactive class. The NN yielded 32 ± 6% and 22 ± 14% for the same classes. While these values reflect encouraging trends, they remain insufficient for operational use. To confirm that observed patterns reflect genuine biological variation rather than ML artifacts, a permutation test was performed (Fig. S4).57 Activity labels were randomly reassigned five times and processed using the same ML pipeline. As expected, the ML accuracies dropped to 0.52 ± 0.10 (EN) and 0.51 ± 0.10 (NN), confirming that our model accuracy of 0.75 and 0.73 with EN and NN, respectively, is a meaningful outcome, and the classification patterns in Fig. 5 are not due to an ML artifact or random chance.
Limitations
The primary limitations of this study are (1) the limited sample size (n = 81), (2) broad PA distributions and (3) the arbitrary definition of active and inactive groups. In general, ML algorithms perform best when trained on large datasets, resulting in a common limitation (1) in fingerprint studies.58,59 Both EN and NN models would likely benefit from additional training data, as they rely on iterative learning. A larger dataset would also enable more refined modeling of PA levels. The broad distribution of PA levels (2) should also be considered; Fig. 1 shows a distribution in which the upper quartile (75–100%) shows a wider spread than the lower quartile (<50%), including some outliers. A more diverse population across a wide range of PA levels would enable more advanced classification labels or possibly regression modeling for continuous PA predictions. Another limitation (3) is imposed by the arbitrary definition of ‘active’ and ‘inactive’ groups. While this was appropriate for the goal of this work, we acknowledge that PA is not a binary trait, but rather a continuous one. Individuals whose true activity levels fall near the chosen cutoff are the most susceptible to misclassification by the ML models, due to limited biological separation. Future studies could incorporate multiple PA classes; however, this will require recruiting a larger cohort spanning a broader range of activity levels to ensure adequate representation of participants within each class. Regression modeling was also tested with the current dataset (Fig. S4). An EN regression model trained on TGs, DGs, and WEs achieved the strongest performance; however, the average R2 was only 0.17 (with 1 indicating a perfect prediction). The predicted versus true PAEEw plot shows systematic underprediction for participants with high PAEEw values. This poor performance may reflect the limited number of high-activity individuals in the dataset or constrained biological variability in lipid responses to physical activity. With the current data, it is not possible to determine which factor is more influential. Moreover, because the lipids examined here are common constituents of human physiology, individuals with very high activity levels may not exhibit markedly different lipid profiles compared to those with moderately high activity. While regression approaches did not perform well in this study, they are expected to improve with substantially larger and more demographically diverse datasets.While the present study is not yet suitable for operational deployment, it reveals a biological trend in fingerprint lipids in response to physical activity. To advance this empirically informed approach toward practical forensic application, external validation using independent cohorts is essential to evaluate robustness, reproducibility, and model generalizability. Future studies should therefore include larger and demographically diverse populations, incorporate expanded physical activity targets (e.g., multiclass classification or regression frameworks), and assess performance across varying environmental, physiological, and lifestyle conditions. In this study, we used groomed fingerprints to enhance data quality but this must be also tested with natural fingerprints in future studies. Such efforts will be critical to determine whether the observed lipid patterns can be reliably translated into a broadly applicable forensic tool.
Conclusions
ML has become an increasingly valuable tool for addressing complex analysis challenges, particularly in biological and forensic research, where subtle interpersonal variation can obscure meaningful patterns. When paired with chemical analysis, ML algorithms can uncover trends in trace-level compounds that are difficult to discern through manual interpretation. This study compared four supervised ML algorithms for classifying PA status based on fingerprint lipids. The EN and NN models achieved the highest classification performance, with average accuracies of 75% and 73%, respectively.These findings underscore the growing potential of ML to advance trace evidence analysis by extracting biologically relevant information from complex chemical data. Future work will prioritize expanding the sample size, refining classification schemes, and examining additional factors such as diet. Ultimately, the goal is to develop new tools for forensic investigations, including methods for extracting relevant information from the chemical composition of unknown latent fingerprints. With continued investment in both data collection and algorithm development, this study highlights a promising future for the integration of ML in forensic science.
Author contributions
Y. L. contributed to conceptualization, supervision, and project administration. D. P. performed funding acquisition, investigation, and validation. R. B. J. contributed to methodology development, MATLAB code creation, and validation. T. F. contributed to the investigation and resource management. D. P. wrote the original draft of the manuscript; Y. L. and R. B. J. reviewed and edited the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). The associated.xlsx file contains Tables S1–S3, data used in physical activity calculations and machine learning. An example of MATLAB code is available on GitHub: https://github.com/drpatten/Fingerprint-Physical-Activity-ML.git. Supplementary information: the survey addendum and additional results referenced in the manuscript. See DOI: https://doi.org/10.1039/d5ay02066b.Acknowledgements
This project was supported by Award no. 15PNIJ-23-GG-01950-RESS, awarded by the National Institute of Justice, Office of Justice Programs, U.S. Department of Justice. The opinions, findings, and conclusions or recommendations expressed in this publication are those of the authors and do not necessarily reflect those of the Department of Justice. The graphical abstract was constructed through BioRender.com.60References
- S. M. Bleay, R. S. Croxton and M. D. Puit, Fingerprint Development Techniques: Theory and Application, 2018 Search PubMed.
- J. De Alcaraz-Fossoul, ed. Technologies for Fingermark Age Estimations: A Step Forward, Springer International Publishing, Cham, 2021 Search PubMed.
- Scientific Working Group for the Analysis of Seized Drugs (SWGDRUG), SWGDRUG Recommendations, Version 8.0, 2019 Search PubMed.
- S. Francese, in Advances in MALDI and Laser-Induced Soft Ionization Mass Spectrometry, ed. R. Cramer, Springer International Publishing, Cham, 2016, pp. 93–128 Search PubMed.
- H. M. Brown, T. J. McDaniel, P. W. Fedick and C. C. Mulligan, Anal. Methods, 2020, 12, 3974–3997 RSC.
- K. C. O'Neill and Y. J. Lee, J. Forensic Sci., 2018, 63, 708–713 Search PubMed.
- P. Hinners and Y. J. Lee, J. Forensic Sci., 2019, 64, 1048–1056 CrossRef CAS PubMed.
- A. A. Frick, N. Kummer, A. Moraleda and C. Weyermann, Analyst, 2020, 145, 4212–4223 RSC.
- S. M. Bleay, R. S. Croxton and M. D. Puit, Fingerprint Development Techniques: Theory and Application, 2018 Search PubMed.
- V. Kotu and B. Deshpande, Predictive Analytics and Data Mining, Elsevier, 2015 Search PubMed.
- T. M. Mitchell, Machine Learning, McGraw-Hill, New York, 1997 Search PubMed.
- S. Badillo, B. Banfai, F. Birzele, I. I. Davydov, L. Hutchinson, T. Kam-Thong, J. Siebourg-Polster, B. Steiert and J. D. Zhang, Clin. Pharmacol. Therapeut., 2020, 107, 871–885 Search PubMed.
- L. S. Ferguson, F. Wulfert, R. Wolstenholme, J. M. Fonville, M. R. Clench, V. A. Carolan and S. Francese, Analyst, 2012, 137, 4686–4692 Search PubMed.
- C. Heaton, C. S. Bury, E. Patel, R. Bradshaw, F. Wulfert, R. M. Heeren, L. Cole, L. Marchant, N. Denison, R. McColm and S. Francese, J. Forensic Chem. Toxicol., 2020, 20, 100271 Search PubMed.
- C. S. Bury, C. Heaton, L. Cole, R. McColm and S. Francese, Anal. Methods, 2022, 14, 789–797 Search PubMed.
- S. Cadd, M. Islam, P. Manson and S. Bleay, Sci. Justice, 2015, 55, 219–238 CrossRef PubMed.
- A. Girod, R. Ramotowski and C. Weyermann, Forensic Sci. Int., 2012, 223, 10–24 CrossRef CAS PubMed.
- A. Muscella, E. Stefàno and S. Marsigliante, Am. J. Physiol. Heart Circ. Physiol., 2020, 319, H76–H88 CrossRef CAS PubMed.
- B. E. Turvey, Criminal Profiling: an Introduction to Behavioral Evidence Analysis, Academic Press, 2011 Search PubMed.
- R. A. B. Ribeiro and C. B. B. de M. Soeiro, Int. J. Law Psychiatr., 2021, 74, 101670 CrossRef PubMed.
- K. C. O'Neill, P. Hinners and Y. J. Lee, Anal. Methods, 2020, 12, 792–798 RSC.
- H. Besson, S. Brage, R. W. Jakes, U. Ekelund and N. J. Wareham, Am. J. Clin. Nutr., 2010, 91, 106–114 CrossRef CAS PubMed.
- M. I. Gurr, J. L. Harwood, K. N. Frayn, D. J. Murphy and R. H. Michell, Lipids: Biochemistry, Biotechnology and Health, John Wiley & Sons, Chichester, West Sussex ; Hoboken, NJ, 6th edn, 2016 Search PubMed.
- A. Pappas, Derm. Endocrinol., 2009, 1, 72–76 CrossRef CAS PubMed.
- B. E. Ainsworth, W. L. Haskell, A. S. Leon, D. R. Jacobs, H. J. Montoye, J. F. Sallis and R. S. Paffenbarger, Med. Sci. Sports Exerc., 1993, 25, 71 CrossRef CAS PubMed.
- B. E. Ainsworth, W. L. Haskell, M. C. Whitt, M. L. Irwin, A. M. Swartz, S. J. Strath, W. L. O'Brien, D. R. Bassett, K. H. Schmitz, P. O. Emplaincourt, D. R. Jacobs and A. S. Leon, Med. Sci. Sports Exerc., 2000, 32, S498–S504 CrossRef CAS PubMed.
- C. Henry, Public Health Nutr., 2005, 8, 1133–1152 CrossRef CAS PubMed.
- 10,000 steps a day: Too low? Too high?, https://www.mayoclinic.org/healthy-lifestyle/fitness/in-depth/10000-steps/art-20317391, accessed 25 November 2024.
- M. Friedrich, Census Bureau Estimates Show Average One-Way Travel Time to Work Rises to All-Time High, https://www.census.gov/newsroom/press-releases/2021/one-way-travel-time-to-work-rises.html, accessed 25 November 2024.
- N. Rafique, G. K. I. Alkaltham, L. A. A. Almulhim, L. I. Al-Asoom, A. A. AlSunni, R. Latif, M. H. AlSheikh, T. Yar, K. S. Al Ghamdi, A. S. Alabdulhadi, F. N. Saudagar and S. Wasi, J. Multidiscip. Healthc., 2022, 15, 2169–2176 CrossRef PubMed.
- P. Hinners, M. Thomas and Y. J. Lee, Anal. Chem., 2020, 92, 3125–3132 CrossRef CAS PubMed.
- D. R. Patten, A. E. Paulson, T. T. Forsman and Y. J. Lee, Anal. Chem., 2023, 95, 12047–12053 CrossRef CAS PubMed.
- F. Fournelle, N. Lauzon, E. Yang and P. Chaurand, Microchem. J., 2023, 185, 108294 CrossRef CAS.
- A. E. Paulson and Y. J. Lee, ACS Cent. Sci., 2022, 8, 1328–1335 CrossRef CAS PubMed.
- R. A. van den Berg, H. C. Hoefsloot, J. A. Westerhuis, A. K. Smilde and M. J. van der Werf, BMC Genom., 2006, 7, 142 Search PubMed.
- B. Li, J. Tang, Q. Yang, X. Cui, S. Li, S. Chen, Q. Cao, W. Xue, N. Chen and F. Zhu, Sci. Rep., 2016, 6, 38881 CrossRef CAS PubMed.
- P. Hinners, K. C. O'Neill and Y. J. Lee, Sci. Rep., 2018, 8, 5149 Search PubMed.
- G. Groeneveld, M. de Puit, S. Bleay, R. Bradshaw and S. Francese, Sci. Rep., 2015, 5, 11716 CrossRef CAS PubMed.
- A. E. Paulson and Y. J. Lee, ACS Cent. Sci., 2022, 8, 1328–1335 CrossRef CAS PubMed.
- N. Kokalas, A. Petridou, M. G. Nikolaidis and V. Mougios, Br. J. Nutr., 2005, 94, 698–704 CrossRef CAS PubMed.
- V. Mougios, C. Kotzamanidis, C. Koutsari and S. Atsopardis, Metabolism, 1995, 44, 681–688 CrossRef CAS PubMed.
- D. Mijaljica, J. P. Townley, F. Spada and I. P. Harrison, Prog. Lipid Res., 2024, 93, 101264 CrossRef CAS PubMed.
- C. G. Elliott, S. Vidal-Almela, P. Harvey, E. O'Donnell, J. L. Scheid, S. Visintini and J. L. Reed, CJC Open, 2023, 5, 54–71 CrossRef PubMed.
- D. Thiboutot, K. Gilliland, J. Light and D. Lookingbill, J. Invest. Dermatol., 1999, 135(9), 1041–1045 CAS.
- D. Thiboutot, J. Invest. Dermatol., 2004, 123, 1–12 CrossRef CAS PubMed.
- N. E. Archer, Y. Charles, J. A. Elliott and S. Jickells, Forensic Sci. Int., 2005, 154, 224–239 Search PubMed.
- B. N. Dorakumbura, F. Busetti and S. W. Lewis, J. Forensic Chem. Toxicol., 2020, 17, 100193 CAS.
- R. L. Buckman and A. Gundlach-Graham, J. Anal. At. Spectrom., 2023, 38, 1244–1252 RSC.
- C. Weis, A. Cuénod, B. Rieck, O. Dubuis, S. Graf, C. Lang, M. Oberle, M. Brackmann, K. K. Søgaard, M. Osthoff, K. Borgwardt and A. Egli, Nat. Med., 2022, 28, 164–174 CrossRef CAS PubMed.
- W. Gao, H. Li, J. Yang, J. Zhang, R. Fu, J. Peng, Y. Hu, Y. Liu, Y. Wang, S. Li and S. Zhang, Anal. Chem., 2024, 4c00741 Search PubMed.
- L. S. Shapley, in Contributions to the Theory of Games, ed. H. W. Kuhn and A. W. Tucker, Princeton University Press, 2016, pp. 307–318 Search PubMed.
- C. Ambroise and G. J. McLachlan, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 6562–6566 CrossRef CAS PubMed.
- C. S. Bury, C. Heaton, L. Cole, R. McColm and S. Francese, Anal. Methods, 2022, 14, 789–797 RSC.
- C. Zhang and Y. Ma, Ensemble Machine Learning: Methods and Applications, Springer US, Boston, MA, 2012 Search PubMed.
- S. B. Kotsiantis, Informatica, 2007, 31, 249–268 Search PubMed.
- M. Alloghani, D. Al-Jumeily, J. Mustafina, A. Hussain and A. J. Aljaaf, in Supervised and Unsupervised Learning for Data Science, ed. M. W. Berry, A. Mohamed and B. W. Yap, Springer International Publishing, Cham, 2020, pp. 3–21 Search PubMed.
- H. Desaire, J. Proteome Res., 2022, 21, 2071–2074 CrossRef CAS PubMed.
- A. Althnian, D. AlSaeed, H. Al-Baity, A. Samha, A. B. Dris, N. Alzakari, A. Abou Elwafa and H. Kurdi, Appl. Sci., 2021, 11, 796 Search PubMed.
- A. Rácz, D. Bajusz and K. Héberger, Molecules, 2021, 26, 1111 CrossRef PubMed.
- BioRender, https://app.biorender.com/illustrations/6937728b829698921e6bd2fc, accessed 8 December 2025.
Footnote |
| † Present address: Evotec, Princeton, New Jersey 08540, USA. |
| This journal is © The Royal Society of Chemistry 2026 |




