 Open Access Article
Open Access ArticleDevelopment and application of a tetra-ARMS PCR assay for detecting indel polymorphisms in the bovine PRNP gene
Ye
Xu
a,
Siyu
Yang
a,
Siling
Ding
a,
Ting
Xu
a,
Jian
Ge
a,
Weiming
Xiao
b,
Like
Zhu
b and
Feng
Guan
 *a
*a
aCollege of Life Sciences, China Jiliang University, Hangzhou 310018, China. E-mail: guanfengzgjl@163.com
bKey Laboratory of Dairy Cow Genetic Improvement and Milk Quality Research of Zhejiang Province, Wenzhou 325000, China
First published on 11th February 2026
Abstract
Insertion/deletion (indel) polymorphisms in the promoter region (23 bp) and intron 1 (12 bp) of the bovine PRNP gene influence gene expression and susceptibility to bovine spongiform encephalopathy (BSE). Conventional detection strategies dependent on DNA sequencing are cumbersome and costly. A tetra-primer amplification refractory mutation system PCR (tetra-ARMS PCR) assay was developed which can enable efficient and cost-effective genotyping of each indel locus. Optimized tetra-ARMS PCR primers and multiplex conditions allowed electrophoretic genotyping of the 23 bp indel via the size and presence of 422 bp, 259 bp, and 193 bp amplicons, whereas the 12 bp indel was genotyped based on the size and presence of 598 bp, 472 bp, and 153 bp fragments. Furthermore, the compatibility of these primer sets was preliminarily investigated, demonstrating the potential for co-amplification in a single-tube multiplex format. Validation against Sanger sequencing using 62 randomly selected cattle-derived retail samples demonstrated complete concordance. This straightforward, specific and cost-effective method requires only conventional PCR instrumentation, thereby establishing a robust and accessible platform for PRNP-assisted breeding and cattle product genotyping.
1 Introduction
Bovine spongiform encephalopathy (BSE), commonly known as ‘mad cow disease’, is a fatal neurodegenerative disorder resulting from conformational aberrations in the scrapie prion protein (PrPSc).1 Initially identified in the United Kingdom in 1986, BSE later underwent rapid global dissemination, causing devastating livestock losses and precipitating an international public health crisis.2–4 Although the global incidence of BSE has substantially declined following the implementation of control measures (e.g., prohibitions on ruminant-derived feed), elucidating its pathogenesis and establishing effective management strategies remain significant challenges.5–7 The pathogenic isoform PrPSc exhibits pronounced resistance to conventional disinfection and thermal decontamination.8–10 Furthermore, multiple transmission routes of BSE are documented, including milk and meat products,11,12 while environmental prion contamination may constitute secondary vectors.13,14 Collectively, these effective factors impede eradication efforts for this disease.13,15–17 As the primary determinant of prion protein expression, polymorphisms in the bovine prion protein gene (PRNP) modulate its susceptibility to BSE and its heritability.18–22 Following the bovine PRNP gene sequencing in 2001,6,20,23,24 studies have demonstrated that 23 bp indel in the promoter region and 12 bp indel in the intronic region are significantly associated with BSE resistance, thereby informing DNA marker assisted selection of resistant individuals.7,18,22,25–27 Current strategies encompass PRNP gene editing to enhance resistance in cattle breeding.28 Consequently, the detection of PRNP polymorphisms in any locus for BSE risk assessment provides assisted molecular markers for breeding programs, enabling strategic selection of resistant populations and guiding containment policies—imperative for safeguarding livestock industries and public health.Current DNA genotyping of the 23 bp and 12 bp indel polymorphisms in the bovine PRNP gene primarily employs Sanger sequencing,22,29–34 PCR-RFLP analysis,35 quantitative PCR (qPCR),36 next-generation sequencing (NGS),37 and microarray genotyping.38 These methodologies are constrained by protracted processing times, elevated costs, and technical limitations, notably incomplete restriction enzyme digestion. Specifically, Sanger sequencing, despite providing precise sequence resolution, incurs high per-sample costs; PCR-RFLP is susceptible to interpretive errors owing to partial enzymatic cleavage, compromising its reliance on restriction site specificity; NGS, although permitting high-throughput multi-locus analysis, requires substantial instrumentation investment and sophisticated bioinformatics assay, thereby precluding routine diagnostics in resource-limited settings. Compared to the conventional detection methods mentioned above, amplification refractory mutation system PCR (ARMS-PCR) exploits deliberate primer–template mismatches and constitutes a robust DNA analysis platform featuring operational simplicity, elimination of enzymatic digestion/fluorescent probes, cost-effectiveness, broad applicability, visual electrophoretic resolution, and inherent heterozygote discrimination.39–43 Subsequent to its inception, the tetra-primer ARMS-PCR incorporated internal control primers to increase its accuracy and has been widely applied to species identification and SNP genotyping across diverse biological systems,44–49 thus showing exceptional procedural efficiency and accessibility for routine genetic screening.
A tetra-ARMS PCR-based DNA mismatch detection system was established in this study through optimized designed primers for the rapid and accurate genotyping of the 23 bp and 12 bp indel polymorphisms in the bovine PRNP gene. This study developed and validated two independent tetra-ARMS PCR assays, and explored their potential for integration into a multiplex system. This assay enhances genotyping efficiency, broadens methodological capabilities for indel screening, and thereby provides technical support for DNA marker-assisted breeding programs and disease risk assessment in cattle populations.
2 Materials and methods
2.1 Sample sources and DNA extraction
A total of 230 genomic DNA samples were stored in the laboratory and were isolated from blood samples of Holstein cattle (n = 200), Yellow cattle (n = 10), and Wenzhou water buffalo (n = 20). Additionally, commercial specimens (fresh milk, beef, milk powder, cheese; n = 62) were purchased from retail markets, representing diverse production batches and dates. The blood samples of Holstein cattle and Wenzhou water buffalo were provided by the Key Laboratory of Dairy Cow Genetic Improvement and Milk Quality Research of Zhejiang Province, while those of Yellow cattle were from the Dairy Cattle Breeding Farm of Shandong Academy of Agricultural Sciences. Genomic DNA was extracted using a commercial Animal Tissue DNA Extraction Kit (Hangzhou Xinjing Biotechnology Co., China). All animal-related procedures were approved by the Animal Experiment Ethics Committee of China Jiliang University (Approval No. 2021-005).Genomic DNA was extracted from commercial specimens using the TakaRa MiniBEST Universal Genomic DNA Extraction Kit (Takara Bio, China) according to the manufacturer's instructions. Approximately 25 mg aliquots underwent manufacturer-specified pretreatment, followed by genomic DNA isolation according to the instructions. Extracted DNA was quantified via NanoDrop 2000 spectrophotometry (Thermo Fisher Scientific) and assessed for integrity through 1.5% agarose gel electrophoresis with GelRed staining.
2.2 Sequencing analysis of two PRNP polymorphic loci
Sequencing and validation analysis of the 12 bp and 23 bp indel polymorphisms was performed using primer pairs reported by Imran et al.,50 with the bovine PRNP gene sequence (GenBank accession no AJ298878.1) as the reference template. The polymorphic sequences were defined as 5′-GGGGGCCGCGGC-3′ (12 bp) and 5′-TCTCAGATGTCTTCCCAACAGCA-3′ (23 bp).All primers were synthesized by Hangzhou Tsingke Biotechnology Co. (China), with sequences detailed in Table 1.
| Polymorphic locus | Primer | Primer sequence (5′ → 3′) |
|---|---|---|
| 12 bp indel | 12indel F | GTGCTCGTTGGTTTTTGC |
| 12indel R | TCCTACACACCACCCACATA | |
| 23 bp indel | 23indel F | AGCCAGGTAAGAAGCTCATC |
| 23indel R | CATGAATTGTGTAGGCCAAA |
PCR amplification targeting the 12 bp and 23 bp indel fragments in the bovine PRNP gene was performed separately using two independent primer sets under identical reaction conditions. The 20 µL reaction system followed the same PCR profile: 95 °C for 5 min; 35 cycles of 94 °C for 30 s, 60 °C for 30 s, and 72 °C for 45 s; 72 °C for 10 min; and storage at 4 °C. Amplification products were electrophoresed on 1.5% agarose gels, and fragments matching predicted sizes were purified and transferred to Hangzhou Tsingke Biotechnology (China Hangzhou) for bidirectional Sanger sequencing.
2.3 Tetra-ARMS PCR primer design
Tetra-ARMS PCR primers targeting the 12 bp and 23 bp indel polymorphic loci were designed based on the bovine PRNP gene reference sequence (GenBank accession No AJ298878.1) and sequencing data described in Section 2.2, following primer designed principles39,45,47 with the expected amplicon sizes constrained to 150–600 bp for clear electrophoretic resolution. The 12 bp inner primer (12-IF2) has 5-nucleotide complementarity at the 3′-terminus to the insertion sequence, and the 23 bp inner primer (23-IFA4) has 22-nucleotide complementarity at the 3′-terminus to the insertion sequence. To enhance the PCR specificity, an additional T–G weak mismatch was introduced at nucleotide position 5 (from the 5′-terminus) in the upstream inner primer 23-IFA4 for the 23 bp polymorphic locus, and a T–G mismatch at nucleotide position 3 (from the 3′-terminus) in the downstream inner primer 23-AR2. The primer design strategy is illustrated in Fig. 1, with sequences provided in Table 2. All primers were synthesized by Hangzhou Tsingke Biotechnology Co., Ltd (China), diluted to 10 µM, and stored at 4 °C for future use.| Polymorphic locus | Primer | Primer sequence (5′ → 3′) | Fragment length (bp) |
|---|---|---|---|
| a In the tetra-ARMS PCR primer system: in primer nomenclature, “I” denotes the upstream inner primer for the insertion sequence, and “D” denotes the downstream inner primer for the deletion sequence. Lowercase letters in “I” primers indicate bases complementary to the insertion sequence, while letters enclosed in boxes represent intentionally introduced mismatch bases. | |||
| 12 bp indel | 12-PF3 | ACCTTGACCGTGAGTAGGGCTGGG | |
| 12-IF2(I) | ACTCGGAATGTGGGCggggg | 472(I) | |
| 12-Rin(D) | GGGGGACCAGCCAGCCCA | 153(D) | |
| 12-LR1 | CGCCAGGTGGCCCATGATT | 598/610 | |
| 23 bp indel | 23-PF1 | GCAATGCAGCTTCCATGTTGTTTACTGA | |
| 23-IFA4(I) |
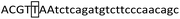
|
193(I) | |
| 23-AR2(D) |
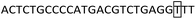
|
259(D) | |
| 23-PR3 | CGTGAGGGTTTGAGGGAACGAAATGAC | 422/445 | |
2.4 Tetra-ARMS PCR amplification
The basic PCR program used was as follows: initial denaturation at 95 °C for 8 min; 35 cycles of 94 °C for 35 s, annealing at 57 °C, 60 °C, 63 °C, 66 °C, 69 °C and 72 °C for 40 s, 72 °C for 1 min; followed by a final extension at 72 °C for 10 min, and finally the PCR products were maintained at 4 °C in the end. Optimal reaction conditions were determined through systematic optimization of annealing temperature (57–72 °C gradient) and cycle number (30, 32, 35, and 38 cycles). Subsequent optimizations included Mg2+ concentration (1.5, 2.0, 2.5, and 3.0 mM) and primer concentration ratios (outer![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) inner from 5
inner from 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 1
1 to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5). PCR products were resolved by 2.0% agarose gel electrophoresis, with optimal reaction conditions selected based on product specificity and amplification efficiency.
5). PCR products were resolved by 2.0% agarose gel electrophoresis, with optimal reaction conditions selected based on product specificity and amplification efficiency.
2.5 Limit of detection (LOD) assessment for tetra-ARMS PCR
To further explore the detection limit of ARMS-PCR, the optimized detection system was used to assess it for different genotypes using continuous gradient dilution of DNA. Genomic DNA samples from individuals with three genotypes (Ins, Del, and Hetero) for both the 12 bp and 23 bp loci were selected (two samples per genotype) as the template. DNA samples of identical genotypes were mixed and subjected to 10-fold serial dilution for detection limit assay.2.6 Genotyping of commercial cattle-derived products
The PRNP genotypes of 62 commercial cattle-derived products (raw milk, milk powder, beef and cheese) were determined using an optimized single-locus tetra-ARMS PCR system for both the 12 bp and 23 bp polymorphic sites. Concurrently, three samples per genotype were randomly selected for bidirectional sequencing verification according to the method in Section 2.2.Based on ARMS-PCR electrophoretic profiles, allele and genotype frequencies for both loci in the bovine PRNP gene were calculated.
3 Results and analysis
3.1 DNA extraction and quality
The analysis of 62 extracted DNA samples revealed that the DNA concentration for meat samples was 46.2 ± 9.55 ng µL−1 and the OD260/280 ratio was 1.93 ± 0.07; for raw milk samples, the concentration was 2.69 ± 1.09 ng µL−1 and OD260/280 was 1.66 ± 0.21; for milk powder samples, the concentration was 6.93 ± 2.65 ng µL−1 and OD260/280 was 1.74 ± 0.23. PCR products showed that all the DNA samples could amplify the specific PRNP gene containing the 12 bp and 23 bp polymorphic sites, demonstrating that the DNA concentrations and purity meet the PCR analysis requirements.3.2 Sequencing-based genotyping for the two PRNP polymorphic loci
Electrophoretic analysis of PRNP amplicons indicated the obtained fragment sizes were 119/131 bp for 12 bp locus and 245/268 bp for 23 bp locus and are shown in Fig. 2, consistent with expected sizes. However, the close proximity of fragment sizes impeded definitive genotyping by electrophoresis. Sequence alignment using EditSeq and MegAlign software confirmed the presence of three genotypes per locus, including homozygous deletion (Del genotypes), homozygous insertion (Ins genotypes), and insertion-deletion heterozygote (Hetero genotypes), as shown in Fig. 3.3.3 ARMS-PCR genotyping systems
| 12 bp Genotype | 23 bp Genotype | Fragment length (bp) | Total bands |
|---|---|---|---|
| a Symbols ‘+’ and ‘−’ represent insertion and deletion at the gene locus, respectively. | |||
| + + | + + | 610, 472, 445, 193 | 4 |
| + + | + − | 610, 472, 445, 259, 193 | 5 |
| + + | − − | 610, 472, 422, 259 | 4 |
| + − | + + | 610, 472, 445, 193, 153 | 5 |
| + − | + − | 610, 472, 445, 259, 193, 153 | 6 |
| + − | − − | 610, 472, 422, 259, 153 | 5 |
| − − | + + | 598, 445, 193, 153 | 4 |
| − − | + − | 598, 445, 259, 193, 153 | 5 |
| − − | − − | 598, 422, 259, 153 | 4 |
3.4 Limit of detection for ARMS-PCR
Using the verified DNA templates of three genotypes at the 12 bp and 23 bp polymorphic loci as the test samples, the detection limits of the individual single-locus tetra-ARMS PCR assays were assessed using 10-fold serial dilutions. Following PCR amplification with the two separate tetra-primer systems, agarose gel electrophoresis results in Fig. 6 revealed that locus-specific LOD values were as follows: for the 12 bp locus, the minimum detectable concentration of the genomic DNA template with Del genotypes was 4.53 ng µL−1 (0.6795 ng µL−1 in the PCR system), that for Hetero genotypes was 5.39 ng µL−1 (0.8085 ng µL−1 in the PCR system), and that for Ins genotypes was 5.01 ng µL−1 (0.7515 ng µL−1 in the PCR system). For the 23 bp locus: the minimum concentration of the DNA template with Del genotypes was 4.14 ng µL−1 (0.621 ng µL−1 in the PCR system), that for Hetero genotypes was 4.15 ng µL−1 (0.6225 ng µL−1 in the PCR system), and that for Ins genotypes was 0.516 ng µL−1 (0.0774 ng µL−1 in the PCR system). The overall detection limit ranged from 0.516 to 5.39 ng µL−1 (0.0774–0.8085 ng µL−1 in the PCR system). To contextualize the relative sensitivity level of the developed assay, its detection limits were compared with those reported for other applications of the tetra-ARMS PCR technique in the literature, as summarized in Table 4. The 23 bp Ins genotypes showed 10-fold higher sensitivity, potentially attributable to the lower GC content in insertion-specific primers enhancing amplification efficiency.| Method | Target locus | LOD | Reference |
|---|---|---|---|
| Tetra-ARMS PCR | Bovine PRNP gene 12/23 bp indel | 0.516–5.39 ng µL−1 | This work |
| Multi ARMS-PCR | Apolipoprotein E genotyping | 10 ng | Lian et al., 2016 (ref. 51) |
| Tetra-ARMS PCR | SARS-CoV-2 | 78.91 copies per µL | Wang et al., 2022 (ref. 52) |
| TaqMan ARMS-PCR | Mitochondrial DNA (mtDNA) 1555A>G | 0.1 ng µL−1 | Tan et al., 2024 (ref. 53) |
3.5 Commercial product genotyping
The two aforementioned primer sets were used to amplify the 62 commercially available bovine-derived products. The results for the 12 bp locus were 18 Del, 13 Ins, and 31 Hetero; for the 23 bp locus, they were 12 Del, 9 Ins, and 41 Hetero. Among these, the identification results of 24 samples are shown in Fig. 7. The electrophoresis detection results were completely consistent with DNA sequencing, demonstrating that this study provides an accurate and intuitive detection method for genotyping the bovine PRNP gene. Based on electrophoretic genotyping results, allele and genotype frequencies were calculated for the 12 bp and 23 bp loci in the bovine PRNP gene. For the 12 bp locus, 21.9% of samples belonged to the Ins genotype, 49.2% belonged to the Hetero genotype and 28.9% belonged to the Del genotype. For the 23 bp locus, 20.9% belonged to the Ins genotype, 51.2% belonged to the Hetero genotype, and 27.9% belonged to the Del genotype. Polymorphism differences between loci are detailed in Tables 4 and 5.4 Discussion
Since the completion of bovine PRNP gene sequencing in 2001, extensive research has characterized its genetic polymorphisms.23 Among 60 polymorphic sites identified in the gene sequence, the 23 bp indel polymorphism in the promoter region showed significant association with BSE susceptibility, with deletion homozygotes and heterozygotes exhibiting higher disease risk.2 Subsequent studies revealed the 12 bp indel polymorphism in the intron region similarly correlates with BSE resistance/susceptibility. Individuals carrying both 23 bp and 12 bp insertions demonstrate the strongest resistance to BSE, while double-deletion homozygotes show maximal susceptibility.54 These polymorphic sites regulate prion protein (PrP) expression levels and conformational stability, directly impacting pathological accumulation and transmission efficiency of PrPSc.6,21,55,56 Molecular analyses indicated that 12 bp deletion abolishes transcription factor SP1 binding sites, reducing transcriptional efficiency, while 23 bp deletion interferes with 58 kDa repressor protein (RP58) binding, altering chromatin conformation. Both mechanisms converge on PRNP gene regulatory networks, ultimately affecting prion protein metabolic homeostasis.57–59 Additionally, the PRNP gene contains other polymorphic sites such as E211K that influence the prion protein structure.60,61 Nevertheless, the 12 bp and 23 bp polymorphic sites remained central to disease-resistant breeding applications and BSE safety assessments in cattle.28,62–64| Locus | n | Allele frequency | Genotype frequency | |||
|---|---|---|---|---|---|---|
| Insertion allele | Deletion allele | +/+ | +/− | −/− | ||
| 12 bp+/− | 62 | 0.460 | 0.540 | 0.219 | 0.492 | 0.289 |
| 23 bp+/− | 0.465 | 0.535 | 0.209 | 0.512 | 0.279 | |
The critical impact and significance of PRNP gene polymorphisms in animal breeding have driven the extensive development of detection methodologies, including Sanger sequencing,22,29–34 PCR-RFLP analysis,35 quantitative PCR (qPCR),36 next-generation sequencing (NGS),37 and microarray genotyping.38 However, these platforms and developed methods are constrained by expensive instrumentation, high reagent costs, operational complexity, and prolonged processing times. In contrast, ARMS-PCR provides a simple, cost-effective solution for detecting single nucleotide polymorphisms (SNPs) and small indels. Currently, this technology is used in medical diagnostics for the precision genotyping of breast cancer susceptibility alleles,65 genetic risk assessment for mosquito-borne diseases,66 and polygenic analysis of colorectal cancer.67 The applications in the agricultural field include screening for bovine complex vertebral malformation (CVM) susceptibility68 and authentication of Thai bird's nest products.69 Within livestock genetics, the rapid genotyping of leptin receptor mutations in rats70 and ASB-3 gene polymorphisms affect bovine growth traits.71 Extensive applications demonstrated that optimized ARMS-PCR with refined primer design and reaction conditions delivered visual electrophoretic results within 2–3 hours. With minimal costs and scalability for large-scale screening, this method provides an efficient and economical solution for genetic analysis.
This study established two streamlined, independent genotyping assays for bovine PRNP 12 bp and 23 bp indel polymorphisms based on tetra-ARMS PCR technology. The developed single-locus assays can efficiently identify PRNP 12 bp and 23 bp indel genotypes of cattle and cattle-derived products with simplicity, low cost, and high efficiency. ARMS-PCR amplifies target sequences only with perfect 3′-end complementarity to template DNA, blocking amplification otherwise. Incorporating outer primers as positive controls creates a tetra-primer system.39,45 This design enhances accuracy by reducing false negatives and ambiguous results in practical applications.
This study designed an optimized tetra-ARMS PCR system targeting 12 bp and 23 bp indel polymorphisms in the bovine PRNP gene, which provided a better method for PRNP genotyping technology. It is worth mentioning that the tetra-ARMS PCR primers designed in this study are different from other SNP genotyping primers, and all the primers were designed only for inserting genotype sequences with multiple bases complementary to the insertion sequence at the 3′ end of the inner primer which had higher accuracy and sensitivity than other common tetra-ARMS PCR single base mismatches. Critically, inner primer sequences fully featured extended 3′-terminal complementarity to insertion sequences, specifically, 5 nucleotides in 12-IF2 and 22 nucleotides in 23-IFA4 were complete pairing the 12 bp and 23 bp insertion sequences, which ensures the specificity of PCR amplification. This design achieved detection limits as low as 0.516 ng µL−1 genomic DNA and is comparable to that of other established tetra-ARMS PCR applications, including the assay reported for apolipoprotein E genotyping.51 Moreover, the specificity was enhanced through an introduced T–G mismatch at position 5 (from 5′-terminus) in primer 23-IFA4 and position 3 (from 3′-terminus) in primer 23-AR2. These strategically positioned weak mismatches suppress non-specific amplification while improving allelic discrimination,39,72 aligning with established principles for enhancing ARMS-PCR accuracy.
The concentration ratio of primers in the tetra-ARMS PCR reaction system is also a key factor affecting specificity and sensitivity. Due to the presence of multiple mismatch sites in the inner specific primer, the amplification efficiency of the outer primers is higher than that of the inner primer, as they are completely matched. Therefore, adjusting primer concentration ratios was adopted to counterbalance amplification efficiency disparities between fully matched outer primers and mismatch-containing inner primers.73 Specifically, by increasing inner primer concentrations, this study resolved amplification imbalances.
Furthermore, the annealing temperature is another critical factor influencing PCR amplification, and its effect in the tetra-ARMS PCR system indicates that as the annealing temperature gradually increases, the specificity of the two sets of primers will also be correspondingly enhanced. This study has demonstrated this phenomenon that annealing temperature optimization addressed the problem of low PCR efficiency due to the high GC content (73%) in the 12 bp insertion sequence (GGGGGCCGCGGC). At 69 °C, exceeding conventional PCR Tm values, specific target amplification was achieved while eliminating non-specific products.
The principal value of the assay developed herein is its potential to significantly reduce the economic and temporal barriers associated with screening for these important PRNP indels. Compared to the gold-standard Sanger sequencing, which is accurate but costly and slower for batch processing, our method requires only a conventional PCR thermocycler and standard gel electrophoresis apparatus, instruments accessible in most molecular biology laboratories. Reagent costs are minimal, and the workflow from DNA to genotype can be completed within approximately 3 hours, enabling high-throughput analysis. This makes it a viable and attractive option for large-scale screening within breeding programs aimed at increasing BSE resistance in cattle populations. By facilitating the cost-effective identification of resistant (Ins/Ins) individuals, the method directly supports DNA marker-assisted selection. Additionally, its successful application to commercial products (milk, beef, and cheese) demonstrates utility in the genetic traceability and risk assessment of animal-derived foods, contributing to safety monitoring in the food chain.
Nevertheless, this study has certain limitations. First, the developed multiplex detection system requires further systematic optimization to improve its stability and reproducibility across different laboratory conditions. Second, although validated using commercial processed products (e.g., milk powder and cheese), the performance of the method on highly degraded or low-quality DNA,74 or with extreme sample matrices containing complex inhibitory substances,75 remains to be specifically evaluated in future work. Finally, as a technique based on end-specific amplification, its accuracy depends on precise primer design and stringent PCR condition control, which requires careful attention to the experimental technique.
Additionally, the ARMS-PCR framework demonstrates potential prospects for integration with advanced platforms. Combining it with lateral flow chromatography enabled the SARS-CoV-2 detection limit to 1.90 copies per µL,61 and coupling with digital PCR achieved a detection limit of 0.308 copies per µL for epidermal growth factor receptor mutations.76 Such integrations highlight its promise for future ultra-sensitive diagnostics.
In summary, the tetra-ARMS PCR method established in this study offers significant advantages of low cost and high efficiency, over existing techniques for analysing 12 bp and 23 bp indel polymorphisms in the bovine PRNP gene. It provides a viable option for large-scale screening of disease-resistant animal individuals. Furthermore, disease resistance breeding represents an important safeguard for animal husbandry safety and public health, holding significant scientific value and societal value.
5 Conclusions
This study successfully developed and validated two independent tetra-ARMS PCR assays for the efficient genotyping of 12 bp and 23 bp indel polymorphisms in the bovine PRNP gene. Furthermore, a preliminary multiplex configuration was demonstrated, outlining a clear pathway for future development toward a single-tube, high-throughput screening tool. The established single-locus method offers cost-effective, high-throughput screening, addressing limitations of existing methods. This approach significantly advances disease-resistant cattle DNA marker assisted breeding and safety assessment of animal-derived food products.Ethical statement
The blood samples used for genomic DNA isolation were originally collected as part of routine herd health monitoring programs by the collaborating farms and research institutes. All animal-related procedures in this study were conducted in strict compliance with the national regulations and guidelines of the People's Republic of China, including the “Regulations for the Administration of Affairs Concerning Experimental Animals” and the “Guideline on the Humane Treatment of Laboratory Animals”. The protocol for the retrospective genetic analysis of these pre-existing DNA samples was reviewed and formally approved by the Animal Experiment Ethics Committee of China Jiliang University (Approval No. 2021-005). This research did not involve human participants.Author contributions
Ye Xu: conceptualization, methodology, investigation, writing – original draft. Siyu Yang: conceptualization, methodology. Siling Ding and Ting Xu: investigation. Jian Ge: investigation, formal analysis. Weiming Xiao: resources. Like Zhu: funding acquisition, project administration. Feng Guan: supervision, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2025) in National Genomics Data Center (Nucleic Acids Res 2025), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA: CRA030580) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa/search?searchTerm=CRA030580.Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (31672394), Zhejiang Science and Technology Major Program on Agricultural New Variety Breeding (2021C02068-6), Zhejiang Province Agricultural Major Technology Collaborative Promotion Plan (2023ZDXT16-3), the Project for Zhejiang Province Basic Public Welfare Research Program (LTGN24C050001, 2026C02A3025), China Agriculture Research System (CARS-39-07), Young Science and Technology Top Talent Program of Tianshan Talent Training Program in Xinjiang Province (2022TSYCCX0124), Xinjiang Agriculture Research System (XJARS-09-26), and Project of Corps Science and Technology in Key Areas (2024AB017).Notes and references
- S. B. Prusiner, Science, 1982, 216, 136–144 CrossRef CAS PubMed.
- P. Sander, H. Hamann, I. Pfeiffer, W. Wemheuer, B. Brenig, M. H. Groschup, U. Ziegler, O. Distl and T. Leeb, Neurogenetics, 2004, 5, 19–25 CrossRef CAS PubMed.
- U. Rasheed, M. Khalid, A. Noor, U. Saeed, R. Uppal and S. Zafar, Prion, 2024, 18, 1–7 Search PubMed.
- M. C. Jurcau, A. Jurcau, R. G. Diaconu, V. O. Hogea and V. S. Nunkoo, Neurol. Int., 2024, 16, 1039–1065 CrossRef CAS PubMed.
- S. Liu, S. E. Heumüller, A. Hossinger, S. A. Müller, O. Buravlova, S. F. Lichtenthaler, P. Denner and I. M. Vorberg, Nat. Commun., 2023, 14, 5034 CrossRef CAS PubMed.
- L. Orge, C. Lima, C. Machado, P. Tavares, P. Mendonça, P. Carvalho, J. Silva, M. L. Pinto, E. Bastos, J. C. Pereira, N. Gonçalves-Anjo, A. Gama, A. Esteves, A. Alves, A. C. Matos, F. Seixas, F. Silva, I. Pires, L. Figueira, M. Vieira-Pinto, R. Sargo and M. D. A. Pires, Biomolecules, 2021, 11, 1–29 CrossRef PubMed.
- S. Gilch and H. M. Schatzl, Cell Tissue Res., 2023, 392, 1–5 CrossRef PubMed.
- S. A. Madsen-Bouterse, M. A. Highland, R. P. Dassanayake, D. Zhuang and D. A. Schneider, PLoS One, 2018, 13, e0204281 CrossRef PubMed.
- C. Ligios, M. G. Cancedda, A. Carta, C. Santucciu, C. Maestrale, F. Demontis, M. Saba, C. Patta, J. C. DeMartini, A. Aguzzi and C. J. Sigurdson, J. Virol., 2025, 99, e0073125 CrossRef PubMed.
- T. Onodera and A. Sakudo, in Prions: Current Progress in Advanced Research, 2nd edn, 2019, DOI:10.21775/9781910190951.01.
- M. A. Tranulis and M. Tryland, Foods, 2023, 12, 824 CrossRef CAS PubMed.
- A. M. Zaharioiu, C. Şandru, E. I. Ionete, F. Marin, R. E. Ionete, A. Soare, M. Constantinescu, F. Bucura and V. C. Niculescu, Materials, 2022, 15, 6593 CrossRef CAS PubMed.
- C. M. Carlson, S. Thomas, M. W. Keating, P. Soto, N. M. Gibbs, H. Chang, J. K. Wiepz, A. G. Austin, J. R. Schneider, R. Morales, C. J. Johnson and J. A. Pedersen, iScience, 2023, 26, 108428 CrossRef CAS PubMed.
- A. Sakudo, Curr. Issues Mol. Biol., 2020, 36, 13–22 CrossRef PubMed.
- R. A. Williamson, D. Peretz, N. Smorodinsky, R. Bastidas, H. Serban, I. Mehlhorn, S. J. DeArmond, S. B. Prusiner and D. R. Burton, Proc. Natl. Acad. Sci. U. S. A., 1996, 93, 7279–7282 CrossRef CAS PubMed.
- K. J. Park, H. C. Park, Y. R. Lee, G. Mitchell, Y. P. Choi and H. J. Sohn, mSphere, 2025, 10, e0086624 CrossRef PubMed.
- T. A. Nichols, B. Pulford, A. C. Wyckoff, C. Meyerett, B. Michel, K. Gertig, E. A. Hoover, J. E. Jewell, G. C. Telling and M. D. Zabel, Prion, 2014, 3, 171–183 CrossRef PubMed.
- E. Y. Teferedegn, H. Can, S. Erkunt Alak and C. Ün, Anim. Biotechnol., 2023, 34, 1931–1936 CrossRef CAS PubMed.
- X. He, S. Memon, D. Yue, J. Zhu, Y. Lu, X. Liu, H. Xiong, G. Li, W. Deng and D. Xi, Animals, 2023, 13, 3627 CrossRef PubMed.
- A. Gurgul, M. P. Polak, M. Larska and E. Slota, J. Appl. Genet., 2012, 53, 337–342 CrossRef CAS PubMed.
- E. D. Cassmann, A. J. Frese, K. A. Becker and J. J. Greenlee, Front. Vet. Sci., 2023, 10, 1301998 CrossRef PubMed.
- A. C. Adeola, S. F. Bello, A. M. Abdussamad, R. A. M. Adedokun, S. C. Olaogun, N. Abdullahi, A. I. Mark, A. B. Onoja, O. J. Sanke, G. F. Mangbon, J. Ibrahim, P. M. Dawuda, A. E. Salako, S. Kdidi and M. H. Yahyaoui, BMC Genom., 2024, 25, 177 CrossRef CAS PubMed.
- D. Hills, S. Comincini, J. Schlaepfer, G. Dolf, L. Ferretti and J. L. Williams, Anim. Genet., 2001, 32, 231–232 CrossRef CAS PubMed.
- S. Mead, S. Lloyd and J. Collinge, Annu. Rev. Genet., 2019, 53, 117–147 CrossRef CAS PubMed.
- I. S. Roh, Y. C. Kim, H. J. Kim, S. Y. Won, M. J. Jeong, J. Y. Hwang, H. E. Kang, H. J. Sohn and B. H. Jeong, Vet. Rec., 2022, 190, e940 CrossRef PubMed.
- K. Moazami-Goudarzi, O. Andréoletti, J. L. Vilotte and V. Béringue, Vet. Res., 2021, 52, 128 CrossRef PubMed.
- Y. R. Lee, Y. C. Kim, S. Y. Won, M. J. Jeong, K. J. Park, H. C. Park, I. S. Roh, H. E. Kang, H. J. Sohn and B. H. Jeong, Vet. Res., 2023, 54, 48 CrossRef CAS PubMed.
- J. Ren, T. Hai, Y. Chen, K. Sun, Z. Han, J. Wang, C. Li, Q. Wang, L. Wang, H. Zhu, D. Yu, W. Li and S. Zhao, Sci. China Life Sci., 2024, 67, 555–564 CrossRef CAS PubMed.
- M. L. Clawson, M. P. Heaton, J. W. Keele, T. P. Smith, G. P. Harhay, J. A. Richt and W. W. Laegreid, BMC Res. Notes, 2008, 1, 32 CrossRef PubMed.
- S. Y. Won, Y. C. Kim, K. Do and B. H. Jeong, Animals, 2021, 11, 2574 CrossRef PubMed.
- X. He, S. Memon, D. Yue, J. Zhu, Y. Lu, X. Liu, H. Xiong, G. Li, W. Deng and D. Xi, Anim. Biotechnol., 2023, 34, 2433–2440 CrossRef CAS PubMed.
- J. C. Pereira, N. Gonçalves-Anjo, L. Orge, M. A. Pires, S. Rocha, L. Figueira, A. C. Matos, J. Silva, P. Mendonça, P. Carvalho, P. Tavares, C. Lima, A. Alves, A. Esteves, M. L. Pinto, I. Pires, A. Gama, R. Sargo, F. Silva, F. Seixas, M. Vieira-Pinto and E. Bastos, Prion, 2023, 17, 75–81 CrossRef CAS PubMed.
- A. C. Adeola, S. F. Bello, A. M. Abdussamad, A. I. Mark, O. J. Sanke, A. B. Onoja, L. M. Nneji, N. Abdullahi, S. C. Olaogun, L. D. Rogo, G. F. Mangbon, S. L. Pedro, M. P. Hiinan, M. M. Mukhtar, J. Ibrahim, H. Saidu, P. M. Dawuda, R. K. Bala, H. L. Abdullahi, A. E. Salako, S. Kdidi, M. H. Yahyaoui and T. T. Yin, Gene, 2023, 855, 147121 CrossRef CAS PubMed.
- M. R. Gurau, E. Negru, T. Ionescu, A. A. Udriste, C. P. Cornea and S. Baraitareanu, Genes, 2022, 13, 1316 CrossRef CAS PubMed.
- A. Psifidi, Z. Basdagianni, C. I. Dovas, G. Arsenos, E. Sinapis, M. Papanastassopoulou and G. Banos, Anim. Genet., 2011, 42, 406–414 CrossRef CAS PubMed.
- M. Van Poucke, J. Vandesompele, M. Mattheeuws, A. Van Zeveren and L. J. Peelman, BMC Infect. Dis., 2005, 5, 13 CrossRef PubMed.
- V. V. Giau, E. Bagyinszky, Y. S. Yang, Y. C. Youn, S. S. A. An and S. Y. Kim, Sci. Rep., 2019, 9, 8368 CrossRef PubMed.
- C. Ferrari, C. Punturiero, R. Milanesi, A. Delledonne, A. Bagnato and M. G. Strillacci, Vet. Res., 2024, 55, 99 CrossRef CAS PubMed.
- S. Ye, S. Dhillon, X. Ke, A. R. Collins and I. N. Day, Nucleic Acids Res., 2001, 29, E88 CrossRef CAS PubMed.
- A. Mukherjee and T. Chattopadhyay, Methods Mol. Biol., 2023, 2638, 315–325 CrossRef CAS PubMed.
- S. A. Song, J. G. Shin and S. H. Oh, Ann. Clin. Lab. Sci., 2024, 54, 519–524 CAS.
- A. Khalil, H. N. Khan, M. Wasim, H. Ayesha and F. R. Awan, Nucleosides Nucleotides Nucleic Acids, 2024, 43, 517–529 CrossRef CAS PubMed.
- M. Hussain, H. N. Khan, S. Abbas, A. Ali, M. N. Aslam and F. R. Awan, Nucleosides Nucleotides Nucleic Acids, 2023, 1–16, DOI:10.1080/15257770.2023.2181972.
- S. Torki, M. Nezhadali, M. Hedayati, H. Karimi, S. A. Razavi and L. Najd Hassan Bonab, Nucleosides Nucleotides Nucleic Acids, 2024, 1–12, DOI:10.1080/15257770.2024.2354427.
- P. K. Dubey, S. Dubey, S. Singh, P. D. Bhat, S. Pogwizd and P. Krishnamurthy, PLoS One, 2024, 19, e0293105 CrossRef CAS PubMed.
- L. Najd-Hassan-Bonab, M. Hedayati, S. A. Shahzadeh Fazeli and M. S. Daneshpour, Heliyon, 2023, 9, e21102 CrossRef CAS PubMed.
- Y. Ke-Xin, C. Xiang, H. Qing-Qing, Y. Yi-An, W. Xiao-Ming, X. Ai-Chun, G. Jian and G. Feng, Anal. Sci., 2023, 39, 1947–1956 CrossRef PubMed.
- H. Yang, S. Yang, X. Xia, R. Deng, H. Gao and Y. Dong, J. Agric. Food Chem., 2022, 70, 8451–8457 CrossRef CAS PubMed.
- J. Chen, X. Xu, P. Dalhaimer and L. Zhao, Animals, 2022, 12, 2680 CrossRef PubMed.
- M. Imran, S. Mahmood, M. E. Babar, R. Hussain, M. Z. Yousaf, N. B. Abid and K. P. Lone, Gene, 2012, 505, 180–185 CrossRef CAS PubMed.
- T. Lian, W. Hui, X. Li, C. Zhang, J. Zhu, R. Li, Y. Wan and Y. Cui, Mol. Med. Rep., 2016, 14, 4153–4161 CrossRef CAS PubMed.
- Y. Wang, H. Chen, H. Wei, Z. Rong and S. Wang, Lab Chip, 2022, 22, 1531–1541 RSC.
- J. Tan, X. Zhang, X. Wei and M. Ding, Curr. Issues Mol. Biol., 2024, 46, 5454–5466 CrossRef CAS PubMed.
- K. Juling, H. Schwarzenbacher, J. L. Williams and R. Fries, BMC Biol., 2006, 4, 33 CrossRef PubMed.
- L. Orge, M. Lurdes Pinto, P. Cristovão, P. Mendonça, P. Carvalho, C. Lima, H. Santos, A. Alves, F. Seixas, I. Pires, A. Gama and M. Dos Anjos Pires, J. Visualized Exp., 2023, 195, e64560 Search PubMed.
- H. Zhao, S.-Q. Wang, L.-L. Qing, L.-L. Liu and Y.-P. Zhang, Sci. Bull., 2016, 61, 1–7 Search PubMed.
- P. Sander, H. Hamann, C. Drögemüller, K. Kashkevich, K. Schiebel and T. Leeb, J. Biol. Chem., 2005, 280, 37408–37414 CrossRef CAS PubMed.
- C. L. Haigh, J. A. Wright and D. R. Brown, J. Mol. Biol., 2007, 368, 915–927 CrossRef CAS PubMed.
- K. Kashkevich, A. Humeny, U. Ziegler, M. H. Groschup, P. Nicken, T. Leeb, C. Fischer, C.-M. Becker and K. Schiebel, Faseb. J., 2007, 21, 1547–1555 Search PubMed.
- C. R. Troth, A. Burian, Q. Mauvisseau, M. Bulling, J. Nightingale, C. Mauvisseau and M. J. Sweet, Sci. Total Environ., 2020, 748, 141394 Search PubMed.
- E. M. Nicholson, B. W. Brunelle, J. A. Richt, M. E. Kehrli Jr and J. J. Greenlee, PLoS One, 2008, 3, e2912 CrossRef PubMed.
- A. Masone, C. Zucchelli, E. Caruso, G. Musco and R. Chiesa, Neural Regener. Res., 2025, 20, 1009–1014 CrossRef CAS PubMed.
- C. S. Han, S. Y. Won, S. H. Park and Y. C. Kim, Animals, 2025, 15, 220 CrossRef PubMed.
- R. Benavente, F. Brydon, F. Bravo-Risi, P. Soto, J. H. Reed, M. Lockwood, G. Telling, M. A. Barria and R. Morales, Emerg. Infect. Dis., 2025, 31, 363–367 CrossRef PubMed.
- W. E. Nouh, E. F. El Azab, E. A. Oraby, S. M. Ahmed, M. A. El-Eshmawy, H. K. Badawy, E. I. A. Shaaban, N. S. El-Beltagy, H. A. Alrub, E. Wahsh, H. A. M. Elmashad, A. M. Elsaid, A. E. T. M. Elhassan, E. Toraih, R. M. Elshazli, A. I. Alalawy and Z. R. Attia, Gene, 2025, 942, 149259 Search PubMed.
- R. Basri, M. B. Aqeel, F. M. Awan, S. N. Khan, A. Obaid, R. Parveen, M. Mohsin, W. Akhtar, A. H. Shah, T. S. Afghan, A. Alam, S. Khan and A. Naz, Sci. Rep., 2025, 15, 3478 Search PubMed.
- M. Ijaz, C. C. Chen, R. K. Iqbal, H. A. Farooq, R. Mehmood, M. Asif, A. Akbar, A. Khan, W. Ijaz, M. B. Said, G. F. Wondmie, S. Ibenmoussa, M. K. Okla and F. Iqbal, Sci. Rep., 2024, 14, 20464 CrossRef CAS PubMed.
- R. R. Alyethodi, U. Singh, S. Kumar, R. Alex, G. S. Sengar, T. V. Raja, R. Deb and B. Prakash, BMC Biotechnol., 2021, 21, 36 Search PubMed.
- D. Lv, Y. Fan, W. Zhong, P. Lonan, K. Liu, M. Wu, Y. Wu, Y. Liang, X. Lai, G. Li and L. Yu, Front. Genet., 2021, 12, 632232 CrossRef CAS PubMed.
- X. Xu, X. Hu, G. Ma, T. Wang, J. Wu, X. Zhu, G. Chen, L. Zhao and J. Chen, Heliyon, 2023, 9, e20159 Search PubMed.
- A. Qi, J. Yan, Y. Yang, J. Tang, W. Ru, X. Jiang, C. Lei, X. Sun and H. Chen, Gene, 2022, 838, 146700 CrossRef CAS PubMed.
- X. Chaobo, G. Feng, W. Pingya, Z. Jin and L. Yuanyuan, Int. Food Res. J., 2017, 24, 159–163 CAS.
- A. Garcés-Claver, S. M. Fellman, R. Gil-Ortega, M. Jahn and M. S. Arnedo-Andrés, Theor. Appl. Genet., 2007, 115, 907–916 CrossRef PubMed.
- S. Qu, L. Liu, S. Gan, H. Feng, J. Zhao, J. Zhao, Q. Liu, S. Gao, W. Chen, M. Wang, Y. Jiang and J. Huang, Clin. Biochem., 2016, 49, 287–291 CrossRef CAS PubMed.
- A. Markou, E. Tzanikou, I. Ladas, G. M. Makrigiorgos and E. Lianidou, Anal. Chem., 2019, 91, 13105–13111 Search PubMed.
- X. Song, J. Gong, X. Zhang, X. Feng, H. Huang, M. Gao and L. Chu, Br. J. Cancer, 2020, 123, 1437–1444 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |







